Abstract
Epstein–Barr virus nuclear antigen 1 (EBNA1) contains two arginine-glycine (RG) repeats that contain symmetric/asymmetric dimethylarginine (SDMA/ADMA) and monomethylarginine (MMA) residues. We generated mouse monoclonal antibodies directed against a monomethylated GRGRGG-containing repeat located between amino acids 328 and 377 of EBNA1. In addition to detecting MMA-modified EBNA1, we also had the goal of identifying cellular proteins that bind to MMA-modified EBNA1 in EBV-positive Raji cells. Furthermore, we hypothesized that antibodies against MMA-modified EBNA1 might also recognize cell factors that use an MMA-modified surface structure similar to that of EBNA1 to bind to their common targets. Using a combination of immunoprecipitation and mass spectrometry, we identified a number of such cellular proteins, including SNRPD1-3, ALY/REF, RPS15, DIDO1, LSM12, LSM14A, DAP3, and CPSF1. An NACA complex protein that was shown previously to bind to the glycine-alanine repeat of EBNA1 was also identified. The proteins identified in this study are involved in splicing, tumorigenesis, transcriptional activation, DNA stability, and RNA processing or export.
Supplementary Information
The online version contains supplementary material available at 10.1007/s00705-024-06172-7.
Introduction
The oncogenic Epstein–Barr virus (EBV) has been detected in nasopharyngeal carcinoma (NPC), nasal NK/T-cell lymphoma (NKTL), Burkitt’s lymphoma (BL), Hodgkin’s lymphoma (HL), gastric carcinoma (GC), and posttransplant lymphoproliferative disease (PTLD) samples from immunosuppressed patients [16] and appears to play a role in multiple sclerosis [20]. Epstein–Barr virus nuclear antigen 1 (EBNA1) is essential for cell transformation and maintenance of episomal EBV DNA in infected cells and is the only viral protein present in all types of infected cells [6].
EBNA1 contains a glycine-alanine (GA) repeat that varies in length among viral strains and is important for immune evasion during primary infection [5]. It also contains arginine-methylated arginine-glycine (RG) repeats that are involved in RNA and DNA binding [19] (Supplementary Fig. S1). The RG repeats are located between the N-terminal amino acids 34–52 (LR1) and the C-terminal amino acids 328–377 (LR2), which play a critical role in replication of the viral episome by targeting EBNA1 to the origin recognition complex (ORC) [14].
Posttranslational arginine methylation in proteins plays a role in RNA binding, protein‒protein interactions, transcriptional regulation, signal transduction, chromatin remodeling, and DNA repair. The arginine methyl transferases PRMT-1, -2, -3, -4 (CARM1), -6, and − 8 generate either monomethylarginine (MMA) or asymmetric dimethylarginine (ADMA) residues, PRMT5 and PRMT9 generate either MMA or symmetric dimethylarginine (SDMA) residues, and PRMT7 appears to preferentially generate MMA residues [26].
Previously, we showed that a monoclonal antibody (mAb) directed against the MMA-modified RG repeat of EBV-encoded nuclear antigen 2 (EBNA2) also reacted with the N-terminal RG repeat of EBNA1 (aa 34–52) [1]. While the EBNA2 RG repeat and the N-terminal RG repeat of EBNA1 mainly consist of RGRG sequences, the C-terminal RG repeat of EBNA1 (aa 328–377) also contains RGRGG sequences. We therefore generated monoclonal antibodies against two synthetic MMA-modified peptides derived from the C-terminal RG repeat of EBNA1 to identify cellular factors that bind to MMA-modified EBNA1. As methylated RG repeats serve as contact surfaces to bind to target proteins and thereby affect downstream pathways [26], an additional hypothesis of this study was that these antibodies would react with cellular proteins whose function is mimicked by EBNA1. The detection of such proteins might help to identify previously unknown pathways that are targeted during EBV-mediated transformation.
Materials and methods
Cell lines and transfection
HEK 293E1 cells, which express EBNA1 of EBV strain B95-8, were obtained from Aloys Schepers, Helmholtz Zentrum München, Munich, Germany, and were maintained in DMEM (Sigma‒Aldrich, #D0822, Merck, Darmstadt, Germany) supplemented with 0.1 mg of G418 (Sigma‒Aldrich #A1720, Merck, Darmstadt, Germany) per mL [14]. The EBV-positive Raji (ATCC: CCL-86), M-ABA, and P3HR-1 (ATCC: HTB-62) [1] cells were grown in RPMI-1640 medium (Sigma‒Aldrich #R8758, Merck, Darmstadt, Germany) supplemented with 10% fetal bovine serum (FBS; #S0615, Biochrom, Berlin, Germany) and antibiotics (40 IU of penicillin and 50 µg of streptomycin [Sigma‒Aldrich #P4333, Merck, Darmstadt, Germany], 1 IU of neomycin sulfate [#1405-10-3; Roth, Karlsruhe, Germany], and 90 IU of nystatin [#700114-0006; Fagrom, Barsbüttel, Germany]) per mL. The EBV-transformed marmoset cell line M-ABA was originally obtained from Beverly E. Griffin, Imperial College of Medicine, London, UK [7].
Animals and antibodies
C57BL6J mice were maintained at the animal facility at the faculty of Biology, LMU Munich, in accordance with German Animal Welfare Legislation and that of the Government of Upper Bavaria, Germany (Gz: 55.2-1-54-2532.0-12-2016). For immunization, the peptides Cys-grgrggsggrgrggsggrgrggsggr and Cys-GGsggrgrggsggrrgrgrerARGGSRE containing monomethylarginine (R, MMA) residues corresponding to aa 328–377 (grgrggsggrgrggsggrgrggsggrrgrgrerARGGSRE) were coupled to ovalbumin via their Cys residues. Monoclonal antibodies were produced as described previously [9] and screened by ELISA for their ability to recognize the unmethylated (NMA) peptide or the methylated peptide. Clones 5C7, 7C10, and 1H7 (mouse IgG2c), which reacted exclusively with the MMA-containing peptides, were used for further analysis. Because these antibodies reacted with both of the methylated peptides described above, peptides corresponding to the overlapping GRGRGG sequence were used for release experiments (see below). All of the peptides used in this study were synthesized by Peps4LS GmbH (Heidelberg, Germany). The rat mAb 1H4, which recognizes EBNA1, was described previously [8]. Mouse mAb (IgG2c) against HSV-1 ICP8 (unpublished) was used as a control.
Cell lysis and immunoprecipitation
Immunoprecipitation with the mAbs 5C7, 7C10, and 1H7 was performed as described previously [1]. In brief, protein A Sepharose beads (Protein A Sepharose 4 FastFlow #17-5280, GE Healthcare, Freiburg, Germany) were incubated overnight with 1.6 mL of mAb 5C7 or 1.6 ml of the isotype control antibody at 4°C and washed with lysis buffer (LB) (25 mM Tris-HCl [pH 7.4], 150 mM KCl, 2 mM EDTA, 0.5% IGEPAL-CA360 [Sigma‒Aldrich #18896, Merck, Darmstadt, Germany], and protease inhibitors [Roche Complete mini, #4693159001, Merck, Darmstadt, Germany]). A total of 1.5 × 107 Raji cells were suspended in 1 ml of LB for 30 min, centrifuged at 20,000 × g for 30 min at 4°C, washed with a buffer containing 50 mM Tris-HCl (pH 7.4), 300 mM KCl, 1 mM MgCl2, and 0.5% IGEPAL, and analyzed by immunoblotting and enhanced chemiluminescence (ECL) (#6883, Cell Signaling Technology, Danvers, Massachusetts, USA) detection [1]. For release experiments with the unmethylated GRGRGG and MMA-modified GRGRGG peptide, mAb-5C7-conjugated beads loaded with Raji cell extracts were incubated with the peptides at 0.1 mg/mL in 10 µL of washing buffer (see above), 10% of the released fraction was subjected to Western blotting [1], and the remainder was used for mass spectrometry (see below).
Mass spectrometry analysis
Samples from the protein release experiments were boiled with an equal amount of 2x Laemmli buffer and loaded onto a polyacrylamide gel (TGX Stain-Free Precast Gels 4–20%, Bio-Rad, Hercules, CA, USA, # 4568091), and proteins were separated by the application of 200 V for 3 min. The gels were stained with InstantBlue (#AB119211, Abcam, Cambridge, UK), and the protein-containing regions were excised. For reduction and alkylation, the gel pieces were incubated first in 45 mM dithiothreitol (30 min, 55°C) and then in 100 mM iodoacetamide (2 × 15 min, at room temperature). The gel slices were minced, and 70 ng of modified porcine trypsin (#V5111, Promega, Fitchburg, WI, USA) was added. After overnight digestion at 37°C, peptides were extracted with 70% acetonitrile and dried in a SpeedVac vacuum concentrator. The samples were dissolved in 0.1% formic acid (FA), and LC‒MS/MS was performed using an UltiMate 3000 Nano liquid chromatography system (Thermo Fisher Scientific, Dreieich, Germany) coupled online to a Q Exactive HF‒X mass spectrometer (Thermo Fisher Scientific, Dreieich, Germany). Separation was performed at 250 nl/min on an EasySpray column (PepMap RSLC C18, 50 cm length, 75 µm ID, # ES903, Thermo Fisher Scientific, Dreieich, Germany). Solvent A was 0.1% FA in water, and solvent B was 0.1% FA in acetonitrile. A 30-minute gradient from 3–25% solvent B followed by a 5-minute gradient from 25–40% solvent B, was used for peptide separation. Spectra were collected via a top-12 acquisition method. For protein identification and quantification, MaxQuant V1.6.1 [23] was used in combination with the human and EBV subsets of the UniProt database. Statistical validation and volcano plot generation were performed using Perseus V1.5.3.2. Enrichment analysis was performed using WebGestalt [10]. The mass spectrometry proteomics data have been deposited into the ProteomeXchange Consortium database via the PRIDE [15] partner repository under the dataset identifier PXD051181.
Results
Generation of monoclonal antibodies against monomethylarginine (MMA)-modified EBNA1
Using monoclonal antibodies, we showed previously that Epstein–Barr virus nuclear antigen 2 (EBNA2) contains SDMA, ADMA, and MMA-modified residues within its RGRG repeat, but these antibodies also reacted with the N-terminal RG repeat (LR1) of nuclear antigen 1 (EBNA1) [1]. Therefore, in this study, we generated mAbs against the large C-terminal RGRGG-containing repeat (LR2) of EBNA1 to determine whether these mAbs would recognize EBNA1 as well as similarly modified cellular proteins.
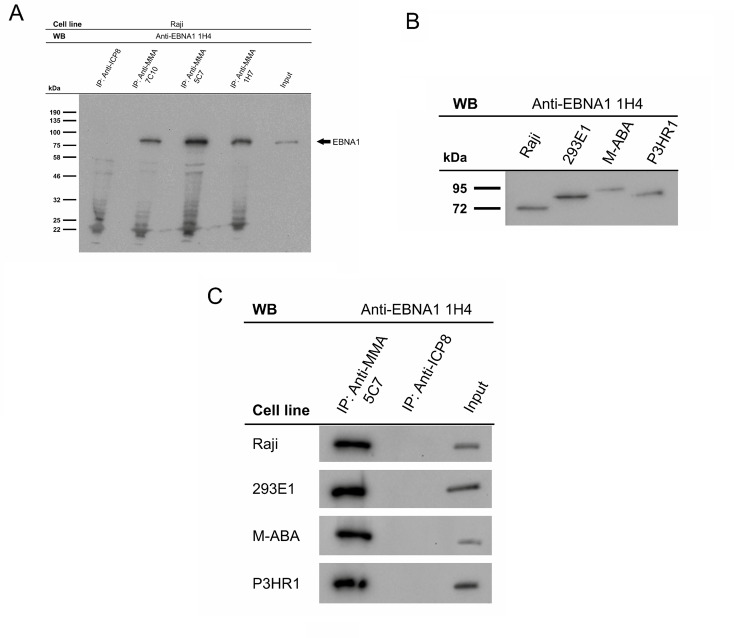
Clones 5C7, 7C10, and 1H7 were used for immunoprecipitation of EBNA1, as shown in Fig. 1A. Equal loading was confirmed (Supplementary Fig. S2A). mAb 5C7 was used in the subsequent analyses for the precipitation of EBNA1 from EBV-containing Raji, M-ABA, and P3HR1 cells and from B95.8-strain-containing HEK 293E1 cells. The corresponding EBNA1 proteins contain GA repeats of varying length, resulting in slightly different mobility on SDS-PAGE gels, as shown in Fig. 1B. P3HR1 cells harbor type 2 EBV, and the other three lines harbor type 1 EBV. The precipitation of the EBNA1 protein by mAb 5C7 is shown in Fig. 1B, and the corresponding control reactions are shown in Fig. 1C. mAb 5C7 reacted with both peptides used for immunization. We determined that the sequence GRGRGG was the most likely overlapping sequence. We used two peptides, one with, and one without MMA modification, for release experiments. As shown in Fig. 2A, only the MMA-modified peptide was able to release the precipitated EBNA1, indicating that mAb 5C7 recognizes MMA-modified EBNA1. The Raji cell extract was used for immunoprecipitation with mAb 5C7 in a peptide release assay as described above. The proteins from four independent experiments and the corresponding controls (Supplementary Fig. S2B) were analyzed via mass spectrometry. A list of proteins that were identified in the protein release experiments but were not present in the control precipitates is shown in Table 1. EBNA1 was not present in any of the corresponding control precipitates.
Fig. 1.
Characterization of mouse monoclonal antibodies (mAbs) directed against MMA-modified EBNA1 via Western blotting. (A) Extracts of the BL cell line Raji (EBV-positive) were precipitated with the mAbs 7C10, 5D7, and 1H7 as well as the control antibody “anti-ICP8” (isotype IgG control). Bound EBNA1 was analyzed by SDS-PAGE and visualized by immunoblotting with the rat monoclonal antibody 1H4 and the appropriate horseradish-peroxidase-conjugated mouse anti-rat IgG secondary antibody using enhanced chemiluminescence (ECL). (B) Extracts of the indicated cell lines were analyzed by Western blotting. EBNA1 was visualized by ECL after immunoblotting using the rat mAb 1H4 and the appropriate secondary antibodies. The cell lines expressed EBNA1 proteins of different sizes due to strain variability. (C) Extracts of the indicated cell lines were used for precipitation with mAb 5C7 or the isotype control antibody “anti-ICP8”. The precipitated EBNA1 was visualized by immunoblotting with mAb 1H4
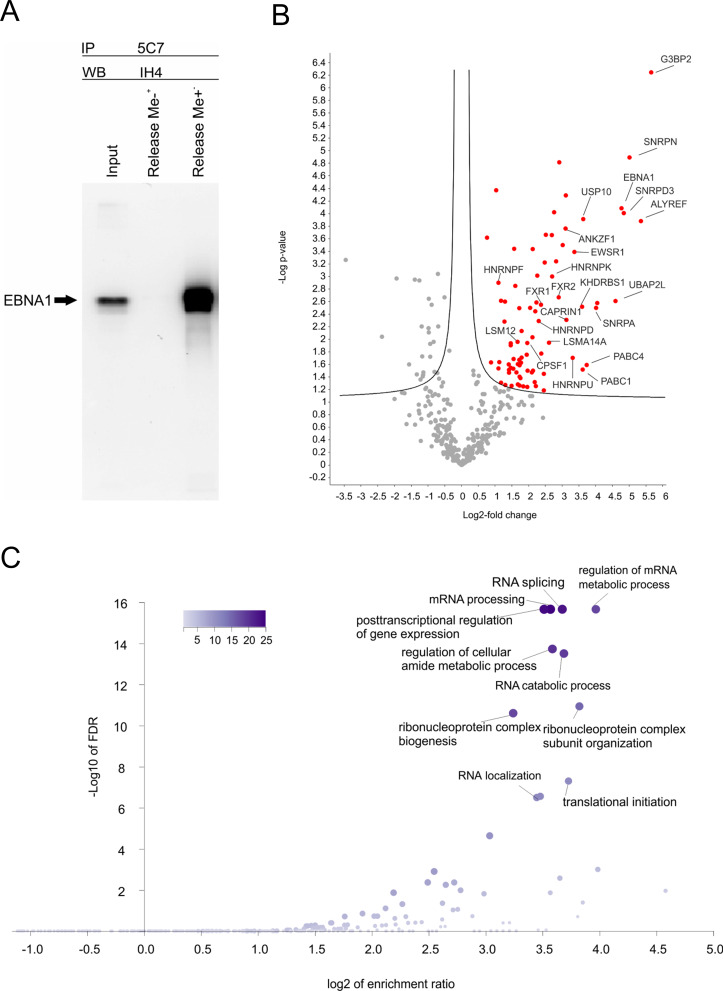
Fig. 2.
Peptide release of EBNA1 and its interactors. (A) Extracts of Raji cells were subjected to immunoprecipitation using the mouse mAb 5C7. The resin was incubated with the peptide RGRGRGG containing either MM-modified arginine residues (“Release Me+”) or unmodified arginine residues (“Release Me-”). The released EBNA1 was visualized by ECL after immunoblotting using the rat mAb 1H4. A 10-µL aliquot of the cell extract was run in an adjacent lane. (B) Volcano plot analysis of mass-spectrometry-based label-free quantification (LFQ) values of proteins released with the MM-modified and unmodified RGRGRGG peptides. Proteins that were more abundant in the release experiment with the MM-modified peptide are highlighted in red. (C) Overrepresentation analysis of this set of proteins was performed with WebGestalt (www.webgestalt.org). The volcano plot includes the enriched functional categories. The size and color of the dots in the scale indicate the sizes of the gene sets
Table 1.
Proteins that were identified in protein release experiments
| Gene | Protein name | Oncogenic function (PMID) |
|---|---|---|
| SNRPD3 | Small nuclear ribonucleoprotein SmD3 | |
| ALYREF | THO complex subunit 4 | 37537569 |
| EBNA1 | EBNA1 | |
| RPS15A | 40S ribosomal protein S15a | 30661291 |
| DIDO1 | Death-inducer obliterator 1 | 22469980 |
| LSM12 | Protein LSM12 homolog | 37394011 |
| LSM14A | Protein LSM14 homolog A | |
| DAP3 | 28S ribosomal protein S29, mitochondrial | 7499268 |
| CPSF1 |
Cleavage and polyadenylation specificity factor subunit 1 |
36381317 |
| NACA |
Nascent polypeptide- associated complex subunit alpha |
|
| KHDRBS1 |
KH domain-containing, RNA-binding, signal transduction-associated protein 1 |
37208334 |
| MIA3 | Melanoma inhibitory activity protein 3 | 37948019 |
| ERH | Enhancer of rudimentary homolog | 37997813 |
| PGAM5 | Serine/threonine-protein phosphatase PGAM5 | 33370650 |
| CHERP |
Calcium homeostasis endoplasmic reticulum protein |
12656674 |
| SLC25A3 | Phosphate carrier protein, mitochondrial | |
| HNRNPCL1 |
Heterogeneous nuclear ribonucleoprotein C-like 1 |
34386035 |
| AKAP8 | A-kinase anchor protein 8 | 31980632 |
| RPL29 | 60S ribosomal protein L29 | 22868929 |
| SNRPN |
Small nuclear ribonucleoprotein- associated protein N |
26261020 |
To identify less prominently enriched proteins, volcano plot analysis was performed (Fig. 2B). The entire set of significantly enriched proteins is listed in Supplementary Table S1. Overrepresentation analysis of this set of proteins revealed significant enrichment of RNA binding or processing proteins (Fig. 2C). RNA binding of EBNA1 via its RG repeats has been demonstrated experimentally [12]. Notably, many of the proteins identified in this study have RG repeats, e.g., SmD3. References (PMIDs) to possible or known roles of the significantly enriched proteins in tumor progression or inhibition or to potential roles in apoptosis are listed in Table 1. The references corresponding to the PMIDs and the known or possible roles of the proteins in tumorigenesis are listed in Supplementary Table S2. For example, DIDO1 promotes the progression of melanoma and inhibits the apoptosis of melanoma cells [3]. In dengue-virus-infected cells, DIDO1 supports virus production by interfering with the interferon response [4]. DIDO1 forms a complex with HNRNPK [17], another confirmed EBNA1 interactor [1] that was also detected in the present study.
Discussion
Viral proteins often affect cellular regulatory pathways by binding to cellular proteins to interfere with or alter their function. For example, by binding via its TrpTrpPro (WWP) motif, nuclear antigen 2 of EBV (EBNA2) alters the function of the repressor JBPjK to activate otherwise silent genes [11]. Using monoclonal antibodies against SDMA-modified EBNA2 [2], we showed that this modification confers binding ability to the survival motor neuron (SMN) protein, possibly by interfering with SMN’s cellular partner protein DDX20 (DP103/Gemin-3) [24].
As mentioned above, mAbs against MMA-modified EBNA2 reacted not only with EBNA2 and various other cellular proteins but also with EBNA1 [1]. Two of the goals of the experiments presented here were to determine whether EBNA1 contains MMA residues within its C-terminal RG repeat and to identify proteins that share similarly modified surface epitopes. In addition, we predicted that cellular proteins that interact with EBNA1 might be identified by coimmunoprecipitation with mAbs. It is therefore noteworthy that NACA (nascent polypeptide-associated complex subunit alpha), which is one of the proteins shown in Table 1, was shown recently to be an EBNA1-binding protein. The association of EBNA1 with NACA is mediated via the glycine-alanine (GAr) stretch of EBNA1 rather than its RG repeats [29]. Some proteins that were precipitated with antibodies against MMA-modified EBNA2 were also precipitated with the mAbs used in this study: ALY/REF, SNRPD3, and some proteins of ribosomal subunits were identified in both analyses.
Some proteins that reacted only with mAb 5C7 may play a role in EBV-mediated transformation. For example, activation of the proto-oncogene c-MYC via chromosomal translocation is a hallmark of Burkitt’s lymphoma [18]. Three of the proteins identified in this study interact with c-MYC RNA: CAPRIN1 binds c-MYC mRNA via its C-terminal RGG repeats [21]; CPSF1 stabilizes c-MYC mRNA, thus increasing the c-MYC level [13]; and the FXR1 protein, together with FXR2, forms a complex with CAPRIN1 [22]. ALY/REF, in turn, is activated by c-MYC and drives cancer cell proliferation [25].
In summary, we have identified various MMA-modified proteins that bind to the novel antibody 5C7, either directly or indirectly via MMA-modified EBNA1, and these proteins are involved in various cellular functions, including gene regulation, RNA processing, export, transcriptional activation, and splicing. The identification of these interacting proteins will serve as a starting point for future studies of EBV function and EBNA1-associated factors involved in cellular transformation or multiple sclerosis.
Electronic Supplementary Material
Below is the link to the electronic supplementary material
Acknowledgments
F.A.G. wishes to dedicate this paper to the memory of the late Gernot Walter, UCSD, San Diego, CA, USA.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Data availability
No datasets were generated or analysed during the current study.
Declarations
Conflict of interest
The authors declare no conflicts of interest.
Footnotes
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
- 1.Ayoubian H, Frohlich T, Pogodski D, Flatley A, Kremmer E, Schepers A, Feederle R, Arnold GJ, Grasser FA (2017) Antibodies against the mono-methylated arginine-glycine repeat (MMA-RG) of the Epstein-Barr virus nuclear antigen 2 (EBNA2) identify potential cellular proteins targeted in viral transformation. J Gen Virol 98:2128–2142 [DOI] [PubMed] [Google Scholar]
- 2.Barth S, Liss M, Voss MD, Dobner T, Fischer U, Meister G, Grasser FA (2003) Epstein-Barr virus nuclear antigen 2 binds via its methylated arginine-glycine repeat to the survival motor neuron protein. J Virol 77:5008–5013 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Braig S, Bosserhoff AK (2013) Death inducer-obliterator 1 (Dido1) is a BMP target gene and promotes BMP-induced melanoma progression. Oncogene 32:837–848 [DOI] [PubMed] [Google Scholar]
- 4.Caraballo GI, Rosales R, Viettri M, Castillo JM, Cruz R, Ding S, Greenberg HB, Ludert JE (2022) The Dengue Virus Nonstructural Protein 1 (NS1) Interacts with the Putative Epigenetic Regulator DIDO1 to Promote Flavivirus Replication in Mosquito Cells. J Virol 96:e0070422 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Dantuma NP, Sharipo A, Masucci MG (2002) Avoiding proteasomal processing: the case of EBNA1. Curr Top Microbiol Immunol 269:23–36 [DOI] [PubMed] [Google Scholar]
- 6.Frappier L (2015) Ebna1. Curr Top Microbiol Immunol 391:3–34 [DOI] [PubMed] [Google Scholar]
- 7.Gao Y, Xue S, Griffin BE (1999) Sensitivity of an epstein-barr virus-positive tumor line, Daudi, to alpha interferon correlates with expression of a GC-rich viral transcript. Mol Cell Biol 19:7305–7313 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Grässer FA, Murray PG, Kremmer E, Klein K, Remberger K, Feiden W, Reynolds G, Niedobitek G, Young LS, Mueller Lantzsch N (1994) Monoclonal antibodies directed against the Epstein-Barr virus-encoded nuclear antigen 1 (EBNA1): immunohistologic detection of EBNA1 in the malignant cells of Hodgkin's disease. Blood 84:3792–3798 [PubMed] [Google Scholar]
- 9.Jakob T, Kollisch GV, Howaldt M, Bewersdorff M, Rathkolb B, Muller ML, Sandholzer N, Nitschke L, Schiemann M, Mempel M, Ollert M, Neubauer A, Soewarto DA, Kremmer E, Ring J, Behrendt H, Flaswinkel H (2008) Novel mouse mutants with primary cellular immunodeficiencies generated by genome-wide mutagenesis. J Allergy Clin Immunol 121:179–184 e177 [DOI] [PubMed] [Google Scholar]
- 10.Liao Y, Wang J, Jaehnig EJ, Shi Z, Zhang B (2019) WebGestalt 2019: gene set analysis toolkit with revamped UIs and APIs. Nucleic Acids Res 47:W199–W205 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Ling PD, Hayward SD (1995) Contribution of conserved amino acids in mediating the interaction between EBNA2 and CBF1/RBPJk. J Virol 69:1944–1950 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Lu CC, Wu CW, Chang SC, Chen TY, Hu CR, Yeh MY, Chen JY, Chen MR (2004) Epstein-Barr virus nuclear antigen 1 is a DNA-binding protein with strong RNA-binding activity. J Gen Virol 85:2755–2765 [DOI] [PubMed] [Google Scholar]
- 13.Mayro B, Hoj JP, Cerda-Smith CG, Hutchinson HM, Caminear MW, Thrash HL, Winter PS, Wardell SE, McDonnell DP, Wu C, Wood KC, Pendergast AM (2023) ABL kinases regulate the stabilization of HIF-1alpha and MYC through CPSF1. Proc Natl Acad Sci U S A 120:e2210418120 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Norseen J, Thomae A, Sridharan V, Aiyar A, Schepers A, Lieberman PM (2008) RNA-dependent recruitment of the origin recognition complex. EMBO J 27:3024–3035 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Perez-Riverol Y, Bai J, Bandla C, Garcia-Seisdedos D, Hewapathirana S, Kamatchinathan S, Kundu DJ, Prakash A, Frericks-Zipper A, Eisenacher M, Walzer M, Wang S, Brazma A, Vizcaino JA (2022) The PRIDE database resources in 2022: a hub for mass spectrometry-based proteomics evidences. Nucleic Acids Res 50:D543–D552 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Rickinson AB, Kieff E (2007) Epstein-Barr Virus. In: Knipe D, Griffin DE, Lamb RA, Strauss SE, Howley PM, Martin MA, Roizman B (eds) Fields Virology. Lippincott Williams & Wilkins, Philadelphia, pp 2655–2700 [Google Scholar]
- 17.Rual JF, Venkatesan K, Hao T, Hirozane-Kishikawa T, Dricot A, Li N, Berriz GF, Gibbons FD, Dreze M, Ayivi-Guedehoussou N, Klitgord N, Simon C, Boxem M, Milstein S, Rosenberg J, Goldberg DS, Zhang LV, Wong SL, Franklin G, Li S, Albala JS, Lim J, Fraughton C, Llamosas E, Cevik S, Bex C, Lamesch P, Sikorski RS, Vandenhaute J, Zoghbi HY, Smolyar A, Bosak S, Sequerra R, Doucette-Stamm L, Cusick ME, Hill DE, Roth FP, Vidal M (2005) Towards a proteome-scale map of the human protein-protein interaction network. Nature 437:1173–1178 [DOI] [PubMed] [Google Scholar]
- 18.Sakaguchi AY, Lalley PA, Naylor SL (1983) Human and mouse cellular myc protooncogenes reside on chromosomes involved in numerical and structural aberrations in cancer. Somatic Cell Genet 9:391–405 [DOI] [PubMed] [Google Scholar]
- 19.Shire K, Kapoor P, Jiang K, Hing MN, Sivachandran N, Nguyen T, Frappier L (2006) Regulation of the EBNA1 Epstein-Barr virus protein by serine phosphorylation and arginine methylation. J Virol 80:5261–5272 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Soldan SS, Lieberman PM (2023) Epstein-Barr virus and multiple sclerosis. Nat Rev Microbiol 21:51–64 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Solomon S, Xu Y, Wang B, David MD, Schubert P, Kennedy D, Schrader JW (2007) Distinct structural features of caprin-1 mediate its interaction with G3BP-1 and its induction of phosphorylation of eukaryotic translation initiation factor 2alpha, entry to cytoplasmic stress granules, and selective interaction with a subset of mRNAs. Mol Cell Biol 27:2324–2342 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Sylvestersen KB, Horn H, Jungmichel S, Jensen LJ, Nielsen ML (2014) Proteomic analysis of arginine methylation sites in human cells reveals dynamic regulation during transcriptional arrest. Mol Cell Proteom 13:2072–2088 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Tyanova S, Temu T, Cox J (2016) The MaxQuant computational platform for mass spectrometry-based shotgun proteomics. Nat Protoc 11:2301–2319 [DOI] [PubMed] [Google Scholar]
- 24.Voss MD, Hille A, Barth S, Spurk A, Hennrich F, Holzer D, Mueller-Lantzsch N, Kremmer E, Grasser FA (2001) Functional cooperation of Epstein-Barr virus nuclear antigen 2 and the survival motor neuron protein in transactivation of the viral LMP1 promoter. J Virol 75:11781–11790 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Wang J, Li Y, Xu B, Dong J, Zhao H, Zhao D, Wu Y (2021) ALYREF Drives Cancer Cell Proliferation Through an ALYREF-MYC Positive Feedback Loop in Glioblastoma. Onco Targets Ther 14:145–155 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Xu J, Richard S (2021) Cellular pathways influenced by protein arginine methylation: Implications for cancer. Mol Cell 81:4357–4368 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Zheng AJL, Thermou A, Daskalogianni C, Malbert-Colas L, Karakostis K, Le Senechal R, Trang Dinh V, Tovar Fernandez MC, Apcher S, Chen S, Blondel M, Fahraeus R (2022) The nascent polypeptide-associated complex (NAC) controls translation initiation in cis by recruiting nucleolin to the encoding mRNA. Nucleic Acids Res 50:10110–10122 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
No datasets were generated or analysed during the current study.