Abstract
Purpose
Our research aimed to evaluate whether proto-oncogene serine/threonine-protein kinase Pim-1 (Pim1) inactivation could attenuate asthma by promoting runt-related transcription factor 3 (Runx3) expression and explore the underlying molecular mechanism.
Method
Phorbol 12-myristate 13-acetate (PMA, 50 nM) was used to induce inflammation in BEAS-2B human airway epithelial cells. ELISA and immunofluorescence double staining confirmed inflammation modelling and differential expression of Pim1 and Runx3. Pim1 inhibitor (SGI-1776) and Runx3 siRNA (siRunx3) were used in this study. Apoptosis, inflammation, MUC5AC protein expression, Pim1 kinase and Runx3 protein expression, and PI3K/AKT/nuclear factor-κB (NF-κB) pathway-associated protein expression were also assessed by flow cytometry, immunofluorescence and western blot. The effects of Pim1 inactivation on airway inflammation, pathological injury and mucus secretion in wild-type and Runx3 knockout mice were observed by in vivo experiments.
Results
The results of the in vitro experiments showed that PMA stimulation causes BEAS-2B cell apoptosis and promotes the MUC5AC expression. In addition, PMA stimulation activated the PI3K/AKT/NF-κB pathway. SGI-1776 treatment partially reversed these effects, whereas siRunx3 attenuated the effects of SGI-1776 on PMA-stimulated BEAS-2B cells. In vivo experiments showed that in Runx3-KO asthmatic mice, inhibition of Pim1 kinase had less effect on airway inflammation, pathological injury and mucus secretion. Meanwhile, Pim1 kinase expression was higher in Runx3-KO asthmatic mice than in wild-type asthmatic mice. Furthermore, inhibition of Pim1 kinase inhibited activation of the PI3K/AKT/NF-κB pathway, whereas these effects were attenuated in Runx3-KO mice.
Conclusion
Our results suggest that Pim1 inactivation can ameliorate airway inflammation and mucus hypersecretion through upregulation of Runx3 and the effect could be mediated through modulation of the PI3K/AKT/NF-κB pathway.
Keywords: Airway Epithelium, Asthma
WHAT IS ALREADY KNOWN ON THIS TOPIC
Pim1 inactivation exerted the protective effect on allergen-induced airway hyper-responsiveness and inflammation. Current reports confirmed that Pim1 inactivation upregulates RUNX3 expression in certain diseases, such as peanut allergy, salivary adenoid cystic carcinoma and breast cancer.
WHAT THIS STUDY ADDS
Our in vitro and in vivo studies revealed that inactivation of Pim1 kinase promotes Runx3 expression, which ameliorates airway inflammation and mucus hypersecretion.
HOW THIS STUDY MIGHT AFFECT RESEARCH, PRACTICE OR POLICY
Further clarification of the mechanism of action for Pim1 in asthma is related to Runx3, which provides strong theoretical support for Pim1 to become a clinical therapeutic target for asthma.
Introduction
Allergic asthma (AA) is well known as a common respiratory inflammatory disease characterised by chronic bronchial airway inflammation, obstruction and hyper-responsiveness.1 In recent years, allergic mechanism had been recognised as the main trigger of asthma in genetically predisposed individuals that result from allergen exposure, and inflammatory responses are characterised by eosinophils, Th2 cells, Th2-driven inflammation and elevated IgE.2 3 Th2 cells are involved in allergic inflammation through the production of the cytokines interleukin (IL)-4, IL-5, IL-9 and IL-13. Therefore, Th2 cells are believed to play an important role in the development and exacerbation of AA. Besides these relevant cytokines, several studies have demonstrated that the development of AA was also regulated by multiple signalling pathways including PI3K/AKT, nuclear factor-κB (NF-κB). Antiasthmatic effects through inhibition of inflammatory cytokines and NF-κB activation in an OVA-induced AA mouse model. In addition, the PI3K/AKT regulates the NF-κB signalling pathway to influence the inflammation-related cytokines including eosinophil production in mice model of AA.
Pim1 kinase is a family of three serine/threonine kinases that controls cell survival, proliferation, differentiation and apoptosis.4 5 A recent study showed that Pim1 kinase promotes T-cell survival.6 Pim1 inactivation attenuates virus/allergen-induced airway hyper-responsiveness (AHR) and inflammation.7 In studies of pulmonary fibrosis, Pim1 kinase was found to directly phosphorylate p65/RelA and promote inflammatory cytokine production by activating NF-κB transcription and enhancing IL-6 production; based on these findings, several new properties have been attributed to Pim1, which may be applicable in AA pathogenesis.
Transcription factor 3 (Runx3), a member of the Runt-related transcription factor family, has attracted considerable attention because of its important role in the development and function of the immune system, especially in T cell differentiation.8 9 Accumulating evidence has shown that Runx3 is involved in T cell development, lineage specification and differentiation, including congenital lymphoid cells, Treg cells and dendritic cells, suggesting that Runx3 may contribute to the development of AA.10,12 A recent study showed inhibition of Pim1 Kinase protected against peanut allergy by potentiating Runx3 expression and suppressing Th 2 and Th 17 differentiation.13 In addition, Pim1 Kinase inactivation reduces the stem-like characteristics of breast cancer cells by inducing the nuclear localisation of Runx3.14 However, whether Pim1 kinase inactivation promotes Runx3 expression to ameliorate airway inflammation and mucus hypersecretion has not been reported.
Therefore, we conducted in vitro and in vivo studies to examine whether Pim1 inactivation promotes Runx3 expression to attenuate asthma and to explore the possible mechanisms.
Methods
Chemicals and cell culture
Phorbol-12-myristate-13-acetate (PMA) and Pim-1 Kinase inhibitor (SGI-1776) were purchased from MedChemExpress (Shanghai, China). Human airway epithelial cells BEAS-2B (Procell Life Science & Technology Co., China) were cultured in a humidified 37°C, 5% CO2 incubator with DMEM high glucose medium (Gibco Life Technologies, USA) containing fetal bovine serum.
Cells grouping and treatment
Cells (2×105) were grown in 96 well plates for 12 hours. The cells were treated with 50 nM PMA. To explore the effect of Pim1 on airway inflammation and its relationship with Runx3, Pim1 inhibitor (SGI-1776) and Runx3 siRNA (siRunx3) were used in this study. The cells were divided into five groups: control, PMA, PMA+SGI-1776, PMA+SGI-1776+si NC and PMA+SGI-1776+siRunx3. Briefly, the cells in the PMA group were stimulated with 50 nM PMA only for 48 hours. The cells in the PMA+SGI-1776 group were treated with 50 nM PMA and 5 nM SGI-1776 for 48 hours. Cells in the PMA+SGI-1776+siRunx3 group were transfected with 50 nM siRunx3 and then treated with 50 nM PMA, 5 nM SGI-1776 for 48 hours. Cells in the PMA+SGI-1776+si NC group were transfected with 50 nM empty vectors and then treated with 50 nM PMA, 5 nM SGI-1776 for 48 hours. The control group did not undergo any treatment. Runx3 siRNA and corresponding empty vectors were purchased from Shanghai Shengbo Biomedical Co. (Shengbo, China). Cell transfection was performed by using Lipofectamine 3000 (Invitrogen) according to the manufacturer’s instructions.
Flow cytometry
Apoptosis was analysed using flow cytometry. BEAS-2B cells (2×105 cells/well) were cultured in a 6-well plate for 72 hours. Next, the cells were digested using 0.25% trypsin without EDTA and resuspended in 500 µL of annexin binding buffer. Subsequently, the cells were stained with Annexin V-FITC and PI (P-CA-201, Pricella, China) in the dark. The apoptotic cells were analysed using a flow cytometer (BD Biosciences, San Jose, USA).
ELISA
The levels of IL-5, IL-13, monocyte chemotactic protein-1 (MCP-1) and human cysteinyl leukotrienes (Cys-LT) in the culture supernatant of cells and BALF were measured using an ELISA kit (ab181421, Abcam, UK) according to the manufacturer’s recommendations.
Immunofluorescence
Pim1/Runx3 double immunofluorescence and MUC5AC single immunofluorescence labelling were performed. The following primary antibodies were used: Pim1(ab308006, Abcam, UK), Runx3 (ab135248, Abcam, UK) and MUC5AC (ab198294, Abcam, UK). Cells were fixed with 4% paraformaldehyde solution for 10 min, permeabilised with 0.5% Triton X-100 for 15 min, blocked with 5% BSA for 1 hour and then incubated with primary antibodies at 4°C overnight. Fluorescent detection antibodies were added and the cells were incubated at room temperature in the dark for 1 hour. The nuclei were labelled with 4,6-diamidino-2-phenylindole for 5 min. Images were captured by using fluorescence microscope (IX7I Olympus, Tokyo, Japan). The results were analysed by Image J software.
Western blot assay
Proteins were isolated from cells or tissues using RIPA lysis buffer containing 20 mM Tris-HCl (pH 7.5), 150 mM NaCl, 1 mM EDTA and 1% Triton X-100. Bicinchoninic acid kit (Abcam, UK) was used to determine protein concentrations of proteins. Then, 50 µg protein samples were separated by 10% SDS-PAGE and then transferred onto 0.22 µm polyvinylidene difluoride (PVDF) membranes (Millipore, USA). PVDF membranes were blocked with 5% non-fat dried milk and incubated overnight at 4°C with primary antibodies as following: anti-Pim1, anti-Runx3, anti-p-PI3K, anti-PI3K, anti-p-AKT, anti-AKT, anti-p-PTEN, anti-PTEN, anti-p-NF-κB, anti-p-NF-κB and anti-actin. Following day, the membranes were incubated with horseradish peroxidase-conjugated IgG secondary antibodies (Santa Cruz Biotechnology) at room temperature for 2 hours. Finally, the bands were visualised using enhanced chemiluminescence reagents (Beyotime, China) and a Tanon 5200 multi-imager (Tanon Science & Technology, China). Protein bands were analysed using ImageJ software.
Animals
To further verify the effect of Pim1 on asthma and its relationship with Runx3, Runx3 knockout (Runx3-KO) and wild-type (WT) mice were used. Runx3-KO and WT mice were purchased from Shandong Experimental Animal Center. All the mice were housed under SPF conditions. All animal care protocols and experiments were performed in accordance with the ARRIVE guidelines and the China Council on Animal Care and Use and were approved by the ethics committee of Yantaishan Hospital, Yantai (No. 2022056).
Sensitisation, challenge and treatment
Runx3-KO mice (n=15) and WT mice (n=15) were randomly divided into three groups (5 mice each group): control, OVA and OVA+SGI-1776 groups. The OVA-induced (DEOVAE01, Demeditec, Germany) AA model was established in mice. On days 0, 7 and 14, except control group, all mice were sensitised by intraperitoneal injection of OVA (150 µg dissolved in phosphate buffer saline) and 2.25 mg aluminium hydroxide (21645-51-2 Invitrogen, Germany). On days 21–26, the OVA+SGI-1776 group was injected intraperitoneally with SGI-1776 (50 mg/kg) diluted in DMSO for 1 hour before challenged with 50 µg OVA via inhalation in a volume of PBS once a day. Mice in the control group were sensitised and challenged with only 0.95% saline. During the experiment, there were no dead mice in all groups. After all animal experimental testing and sampling was completed, the mice were euthanised by cervical dislocation.
Assessment of AHR
AHR was indirectly assessed using non-invasive single-chamber whole-body plethysmography system (WBP System, Buxco Electronics, USA) tracing 24 hours after the last OVA stimulation. Conscious and spontaneously breathing mice were placed in the main chamber of a non-invasive plethysmography system and subsequently exposed to increasing concentrations of methacholine (3.125 mg/mL, 6.25 mg/mL, 12.5 mg/mL, 25 mg/mL and 50 mg/mL) for 3 min. Recording time was 3 min and the recovery time was 1 min. Respiratory resistance values were calculated to evaluate changes in airway resistance in mice after each methacholine excitation.
Analysis of BALF
After anaesthesia, the thorax was clipped to expose the trachea, a small incision was made on the trachea, a lagging needle was inserted into the trachea and 500 µL PBS was used to perform bronchoalveolar lavage with a syringe, which was repeated three times before being withdrawn and the alveolar lavage fluid was collected. The BALF was centrifuged at 2000×g for 10 min at 4°C. The cell pellets were resuspended in 1 mL PBS. The total number of cells was counted using a haemocytometer and differential cell counts were calculated using Wright’s staining.
H&E staining and periodic acid-Schiff (PAS) staining
Bronchial biopsy tissues were fixed in 10% formalin and embedded in paraffin. The 3-µm sections were cut and stained using the H&E staining kit (ab245880, Abcam) and PAS staining kit (Beyotime, China). All images were acquired and observed using a microscope (Olympus BX51, Japan). Inflammation Scoring criteria: no inflammatory cells, 0 points; a small number of inflammatory cells that inflammatory cells form a ring, layer thickness of 1 cell, 2 points; inflammation inflammatory cells forming a ring, layer thickness of 2–4 cells, 3 points; inflammatory cells forming inflammatory cells forming a ring with a layer of 4–5 cells is worth 4 points. PAS-positive rate was analysed by Image J software.
Immunohistochemistry staining
Bronchial biopsy sections were sequentially deparaffinised and rehydrated. The sections were subjected to antigen retrieval using Tris-EDTA (pH 9.0) in a microwave for 15 min at 100°C, incubated with rabbit monoclonal MUC5AC (ab198294, Abcam, UK) overnight at 4°C, and then incubated with FITC goat anti-rabbit secondary antibody (ab6662, Abcam, UK). The sections were stained with DAB solution for 3 min and counterstained with hematoxylin for 3 min. Acid alcohol (95% ethanol+2.5 mL hydrochloric acid) was added for colour separation for approximately 1 min and rinsed with running water three times, followed by dehydration, mounting and sealing. Images were acquired using a microscope.
Statistical analysis
Statistical analyses were conducted using SPSS V.20.0 (IBM) for Windows. Statistical comparisons between the groups were performed using one-way analysis of variance. Test data were normally distributed by Shapiro-Wilk test results. Multiple comparisons were performed using Duncan’s test. All data are reported as mean±SD, and the level of statistical significance was set at p<0.05.
Results
Pim1 and Runx3 were negatively correlated in human airway epithelial cells BEAS-2B inflammation induced by PMA
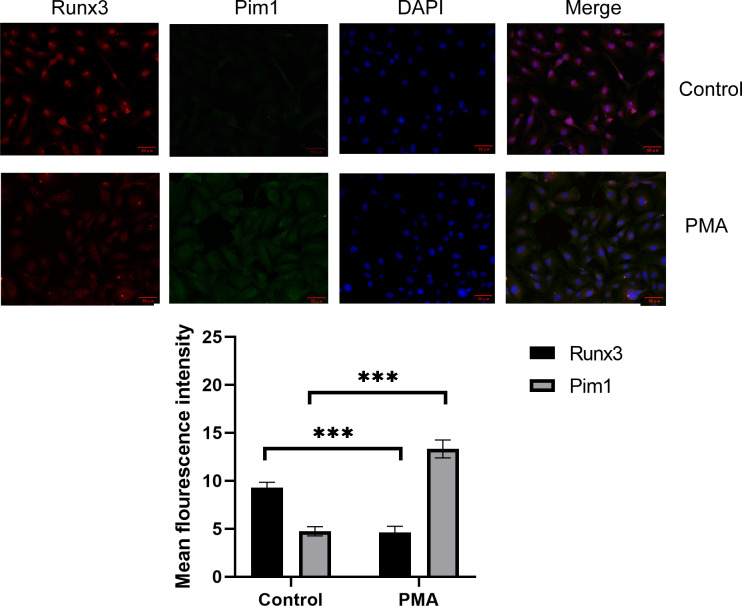
To confirm the differential expression of Pim1 and Runx3 in normal BEAS-2B cells and inflammatory BEAS-2B cells, human airway epithelial cells BEAS-2B were used to construct a PMA-stimulated cellular inflammation model. Double immunofluorescence labelling showed that Pim1 (green) and Runx3 (red) were coexpressed in BEAS-2B cells, as shown in figure 1. Interestingly, the expression of Pim1 was significantly increased, whereas that expression of Runx3 was significantly decreased compared with the control group. PMA stimulation upregulated the expression of Pim1 and downregulated expression of Runx3, suggesting a negative correlation between Pim1 and Runx3 with PMA-induced inflammation.
Figure 1. The localisation and expression of Pim1 and Runx3 in BEAS-2B cells after PMA stimulation. Dual immunofluorescence staining of Pim1/Runx3 was performed. Pim1 (green), RUNX3 (red) and nucleus (blue). The scale bar was 50 µm (the magnification of 200×).
Pim1 inactivation inhibited the epithelial cell apoptosis by Runx3
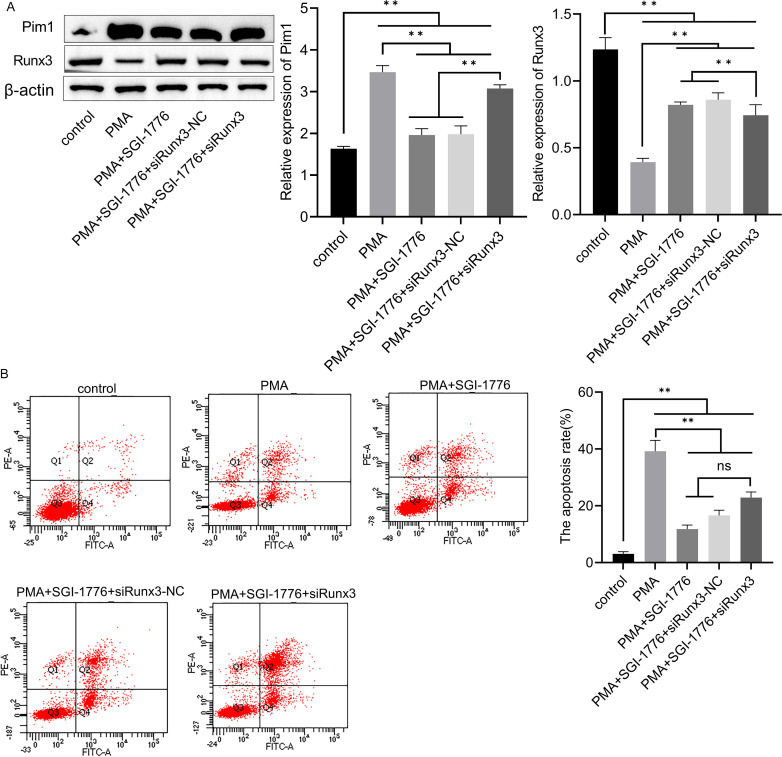
Numerous studies have shown that allergen stimulation-induced epithelial cell apoptosis is strongly associated with airway injury. Thus, we used flow cytometry to examine whether apoptosis of BEAS-2B cells is associated with PMA-induced inflammatory injury. As shown in figure 2A, compared with the control group, the apoptosis rate of BEAS-2B cells was markedly increased in PMA group, whereas SGI-1776 treatment significantly reduced apoptosis. Compared with the PMA+SGI-1776 group, the apoptosis rate of BEAS-2B cells was markedly increased in the PMA+SGI-1776+siRunx3 group. The results showed that Pim1 inactivation inhibits epithelial cell apoptosis via Runx3.
Figure 2. Pim1 inactivation inhibits epithelial cell apoptosis induced by RUNX3. (A) The expression of Pim1 and Runx3 proteins in BEAS-2B cells was detected using western blotting. (B) Cell apoptosis of BEAS-2B cells was detected using flow cytometry. **P <0.05.
Pim1 inactivation decreased the inflammatory factor levels and MUC5AC expression
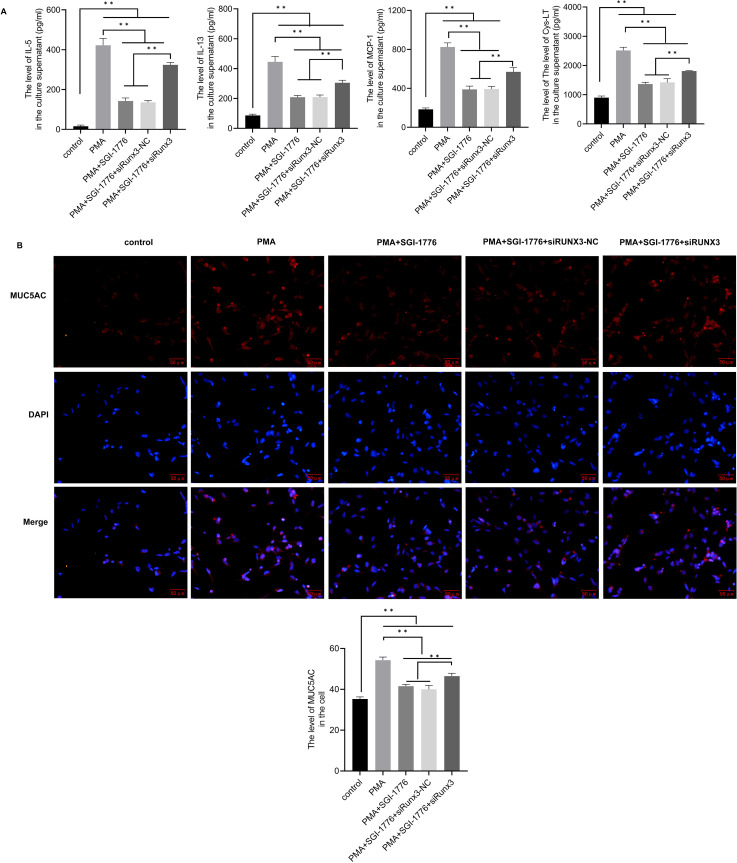
The levels of inflammatory cytokines, including IL-5, IL-13, MCP-1 and Cys-LT, were used as parameters to evaluate the role of Pim1 inactivation in the regulation of inflammation. As shown in figure 3A, the levels of inflammatory cytokines were significantly increased in the PMA group compared with the control group. However, the levels of these cytokines were significantly decreased in SGI-1776 treated group compared with the PMA group, and the change was reversed to a certain extent. Notably, the levels of these inflammatory cytokines were markedly elevated in the siRunx3 group compared with the siRunx3 control group; however, the ability of SGI-1774 to inhibit the release of inflammatory factors was significantly reduced.
Figure 3. Pim1 inactivation decreases inflammatory factor levels and MUC5AC expression in phorbol-12-myristate-13-acetate (PMA)-induced BEAS-2B cells. (A) The levels of interleukin 5 (IL-5), interleukin 13 (IL-13), monocyte chemotactic protein-1 (MCP-1) and human cysteinyl leukotrienes (Cys-LT) were detected by the ELISA method. (B) The expression of MUC5AC was detected using immunofluorescence. **P <0.05.
MUC5AC is a major mucus protein in respiratory diseases, and mucus secretion is increased in asthmatic states. Immunofluorescence assay was used to detect the MUC5AC level. The results in figure 3B showed that MUC5AC level was increased clearly after PMA stimulation. However, compared with the PMA group, MUC5AC level had markedly decreased after treatment with SGI-1776, while siRunx3 attenuated the effect of SGI-1776.
Pim1 regulated Runx3 by PI3K/AKT/NF-κB pathway
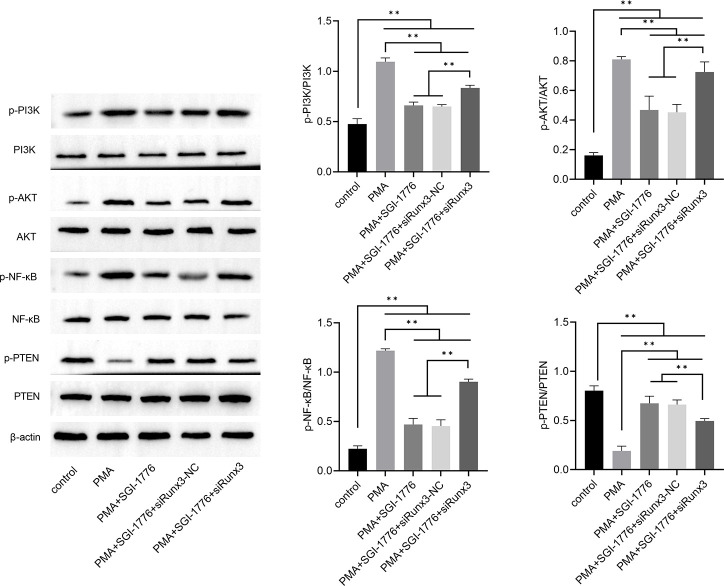
The PI3K/AKT pathway is a key pathway to regulate the release of various cytokines and inflammatory factors. PI3K/AKT was activated after PMA stimulation; the levels of p-PI3K/ PI3K and p-AKT/AKT were markedly increased as shown in figure 4 and online supplemental figure 1. PTEN is a critical negative regulator of PI3K/Akt activation. The results showed that PMA stimulation significantly decreased the levels of p-PTEN/PTEN. However, SGI-1776 significantly decreased the levels of p-PI3K/ PI3K and p-AKT/AKT, whereas increased the levels of p-PTEN/ PTEN; however, siRunx3 attenuated these changes. These findings suggest that PI3K/AKT is closely related to inflammation and Pim1 inactivation promotes PI3K/AKT activation via Runx3.
Figure 4. Pim1 regulated Runx3 via the PI3K/AKT/NF-κB pathway in phorbol-12-myristate-13-acetate (PMA)-induced BEAS-2B cells. The relative expressions of p-PI3K, PI3K, p-Akt, Akt, p-NF-κB, nuclear factor-κB (NF-κB), p-PTEN and PTEN were detected by western blotting. **P <0.05.
An increasing number of studies have shown that NF-κB is a nuclear transcriptional factor that was indirectly regulated by Akt, and its transcriptional activity modulated by inducing phosphorylation. NF-κB plays an important role in regulating diverse cellular activities related to inflammation and immune response. The expression of p-NF-κB and NF-κB was assessed by NF-κB, and the results showed that p-NF-κB/NF-κB significantly increased in the PMA group compared with the control group, while siRunx3 attenuated this change. Overall, Pim1 regulates Runx3 via the PI3K/AKT/NF-κB pathway.
Pim1 regulated methacoline-induced AHR and eosinophils by Runx3
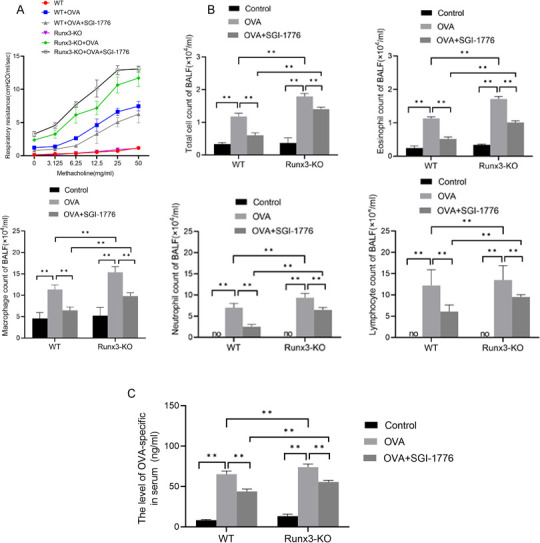
To further support the aforementioned correlation between Pim1, Runx3 and PI3K/AKT/NF-κB pathways, we established a mouse asthma model. The results showed that methacholine-induced AHR (figure 5A), as well as the numbers of the eosinophil, macrophage, neutrophil, lymphocyte (figure 5B) and IgE (figure 5C) was significantly elevated in WT and Runx3-KO mice in the model group compared with the control group, and SGI-1776 significantly attenuated methacholine-induced AHR (figure 5A). However, the levels of AHR, the numbers of the eosinophil, macrophage, neutrophil, lymphocyte and IgE in OVA+SGI-1776 group were higher in Runx3-KO mice than in WT mice. These findings showed that the AA model was successfully established and Pim1 could regulate methacholine-induced AHR and eosinophils via Runx3.
Figure 5. Pim1 regulated asthma progression through Runx3. (A) The respiratory resistance value was calculated to evaluate the changes in airway resistance in mice after each methylcholine excitation. (B) The number of total cells, eosinophils, macrophages, neutrophils and lymphocytes in BALF were counted. (C) IgE levels were measured by ELISA. **P <0.05.
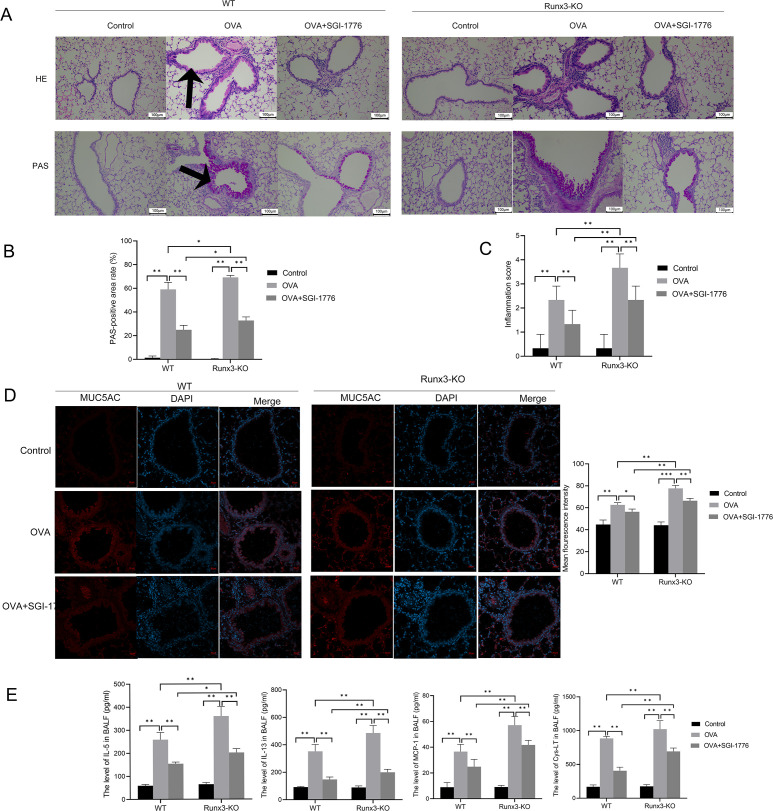
Pim1 regulated Runx3 to improve airway inflammation and inhibit mucus secretion
H&E and PAS staining was used to observe the inflammatory cell infiltration and mucus secretion in the airway. As shown in figure 6, no significant inflammatory cell infiltration and mucus secretion was observed in control group. However, in the OVA group, inflammatory cell infiltration was observed. PAS staining showed a large amount of mucus and mucus plugs. SGI-1776 ameliorated inflammatory cell infiltration and mucus secretion. In particular, inflammatory cell infiltration and mucus secretion were more severe in Runx3-KO asthmatic mice than in WT asthmatic mice. Biomarkers of inflammation and mucus secretion were also detected. The results showed that the levels of IL-5, IL-13, MCP-1 and Cys-LT (figure 6D) and the protein expression of MUC5AC (figure 6C) were significantly increased in the OVA group compared with the control group, whereas SGI-1776 significantly decreased these cytokines levels. In the OVA and OVA+SGI-1776 groups, the cytokines levels were higher in Runx3-KO mice than in WT mice. These results suggest that Pim1 regulates Runx3 to ameliorate airway inflammation and inhibit mucus secretion.
Figure 6. Pim1 regulates Runx3 to improve airway inflammation and inhibit mucus secretion via the PI3K/AKT/NF-κB pathway. (A) H&E and periodic acid-Schiff (PAS) staining was used to assess inflammatory cell infiltration and mucus secretion in the airway. Black arrows indicate positive PAS expression. (B) PAS-positive rate. (C) Inflammatory score. (D) The expression of MUC5AC was detected using immunofluorescence. (E) The levels of interleukin 5 (IL-5), interleukin 13 (IL-13), monocyte chemotactic protein-1 (MCP-1) and human cysteinyl leukotrienes (Cys-LT) in BALF were detected by ELISA. **P <0.05.
Pim1 regulated PI3K/AKT/NF-κB pathway by Runx3
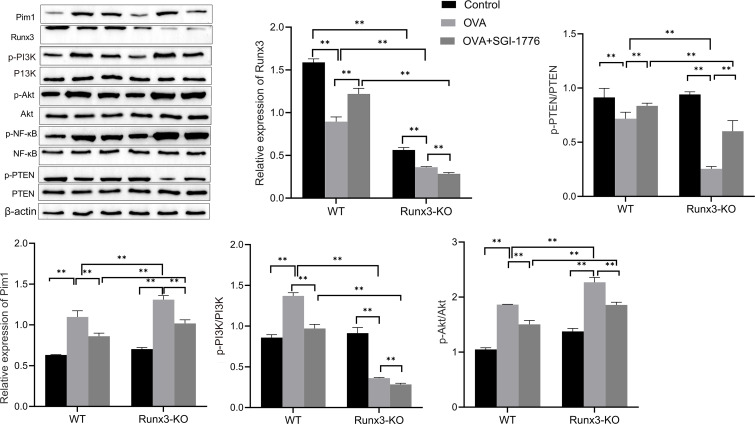
The results in figure 7 and online supplemental figure 2 showed that p-NFκB/NFκB, p-PI3K/ PI3K and p-AKT/AKT were increased in the OVA group compared with the control group, whereas the reduction of p-PTEN/PTEN and SGI-1776 significantly reversed these changes. In the OVA and OVA+SGI-1776 groups, the cytokines levels were higher in Runx3-KO mice than in WT mice. These findings further confirm that Pim1 regulates the PI3K/AKT/NF-κB pathway via Runx3.
Figure 7. Pim1 regulates the PI3K/AKT/NF-κB pathway via Runx3. The expression of Pim1, Runx3, p-PI3K, PI3K, p-Akt, Akt, p-NF-κB, nuclear factor-κB (NF-κB), p-PTEN and PTEN was detected by western blotting. **P <0.05.
Discussion
Asthma is a frequently occurring disease and is a severe health risk in children.15 The pathophysiological process of asthma is complex and involves airway structures including airway cells, cellular components and inflammatory substances.16 Numerous studies have showed that T cells excessive immune responses and the amounts of inflammatory cytokines (such as IL-4, IL-5 and IL-6) are closely related to the development and progression of asthma.17,19 Thus, identifying the involvement of T cells in asthma development and its underlying mechanisms had been repeatedly shown to be a possible new target and therapeutic strategy for the treatment of asthma. In the current study, we demonstrated that Pim1 inactivation ameliorates airway inflammation and mucus hypersecretion by upregulating Runx3 expression. Pim1 expressed in cell types associated with allergic diseases, such as T cells and eosinophils were upregulated in peripheral T cells after antigen activation.20 21 IL-5 and IL-13, mainly derived from activated T cell mast cells, are pivotal Th2 effector cytokines involved in the development of several allergic diseases.22 23 A study revealed that inactivation of Pim1 led to a reduction in T-cell and mast cell recruitment and activation, resulting in lower levels of IL-13 and improved prognosis of sensitised mice after the challenge.13 Th2 expansion and cytokine production are considered to be the main mechanisms by Pim1 kinase which regulates the initiation of an allergic response.13
To understand the consequences of Pim1 activation and explore new therapeutic targets for AA, we focused on the potential downstream transcriptional regulator Runx3. Increasing evidence had shown that Runx3 proteins are involved in multiple cellular functions, especially cell differentiation and apoptosis, and that deficiency of Runx3 results in the spontaneous development of inflammatory bowel diseases. Xu et al24 reported that Pim1 alters the subcellular localisation of Runx3 via phosphorylation in SACC cells. In our study, we found that PMA stimulation led to apoptosis in BEAS-2B cell and promoted MUC5AC expression. SGI-1776 treatment partially reversed these effects, but siRunx3 attenuated the effects of SGI-1776 on PMA-stimulated BEAS-2B cells. In vivo results showed that inhibition of Pim1 kinase had less effect on airway inflammation, pathological injury and mucus secretion in Runx3-KO mice, and the expression of Pim1 kinase was higher than in WT asthmatic mice. These results suggest that Pim1 inactivation ameliorates airway inflammation and mucus hypersecretion by upregulating Runx3 expression in asthma.
Small RNA profile analysis showed that dysregulation of the PTEN/PI3K/Akt pathway in bronchial smooth muscle cells of patients with asthma.25 PI3K inhibitors are considered good therapeutic options for inflammatory respiratory diseases. In the present study, we evaluated PI3K/Akt activity in vivo and in vitro asthma model, and the results showed that PMA-induced activation of the PI3K/Akt pathway was inhibited by SGI-1776, which in turn ameliorated inflammatory manifestations. However, this inhibitory effect of SGI-1776 was significantly attenuated in Runx3-KO mice and siRunx3 cell groups. In the current study, we assessed the activity of PI3K/Akt in vivo and in vitro asthma model26 27 and the results showed that PMA induced activation of the PI3K/Akt pathway, which was inhibited by SGI-1776, which in turn ameliorated inflammatory manifestations. However, this inhibitory effect of SGI-1776 was significantly attenuated in Runx3-KO mice and siRunx3 cell groups.
NF-κB is a key downstream factor of PI3K/Akt signalling pathway that induces overexpression of the MUC5AC.28 A previous study demonstrated that Alpinetin attenuated OVA-induced inflammatory response by inhibiting the activation of the PI3K/Akt /NF-κB and HO-1 pathways.29 Asthmatic mice showed significant upregulation of p-PI3K p85/p55 and p-AktSer473, downregulation of phospho-PTENTyr366, increase in nuclear NF-κB p65 and decrease in cytosolic IκBα.30 In the present study, the results of in vivo and in vitro experiments showed increased expression of p-NF-κB/NF-κB, and this change could be suppressed to a certain extent by SGI-1776. Notably, this suppression was significantly attenuated when Runx3 was silenced or knocked down. In conclusion, our experimental data suggest that PMA induces Pim1 activation, which in turn models asthma inflammation in vivo. Inactivation of Pim1 by the Pim1 inhibitor SGI-1776 induces upregulation of Runx3 that significantly reduces inflammatory factor release, mucus hypersecretion and improves airway inflammation, and we suggest that the effect can be mediated through the PI3K/AKT/NF-κB pathway. These findings provide strong evidence for the important role of Pim1 in the development of asthma and suggest that Pim1 inactivation can upregulate Runx3 expression by inhibiting the activation of the PI3K/AKT/NF-κB signalling pathway, which in turn improves the inflammatory manifestations of asthma. Therefore, we believed that Runx3 is likely to be a potential target for asthma therapy.
Our study also has some limitations: we used OVA stimulation induction to establish a mouse model of AA; we verified the therapeutic effect of Pim1 inhibitors after Runx3 knockdown, and the mechanism that Runx3 is a downstream protein of Pim1 and negatively regulated should be explored more closely.
supplementary material
Footnotes
Funding: This work was supported by Yantai Science and Technology Innovation Program (Grant No. 2023YD039).
Provenance and peer review: Not commissioned; externally peer reviewed.
Patient consent for publication: Not applicable.
Animal ethics approval: All animal care protocols and experiments were performed in accordance with the ARRIVE guidelines and the China Council on Animal Care and Use, and were approved by the ethics committee of Yantaishan Hospital, Yantai (No. 2022056).
Patient and public involvement: Patients and/or the public were not involved in the design, or conduct, or reporting, or dissemination plans of this research.
Contributor Information
Yanni Fang, Email: f5853567925@163.com.
Zhen Guo, Email: guozhen1101@163.com.
Lanzhi Zhou, Email: 912023336@qq.com.
Juan Zhang, Email: 626906921@qq.com.
Haiyan Li, Email: mhaiyanli@163.com.
Jumei Hao, Email: h1315695561@163.com.
Data availability statement
No data are available.
References
- 1.Chen J, Chan WM, Leung HY, et al. Anti-Inflammatory Effects of a Cordyceps sinensis Mycelium Culture Extract (Cs-4) on Rodent Models of Allergic Rhinitis and Asthma. Molecules. 2020;25:4051. doi: 10.3390/molecules25184051. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Murray CS, Woodcock A, Custovic A. The role of indoor allergen exposure in the development of sensitization and asthma. Curr Opin Allergy Clin Immunol. 2001;1:407–12. doi: 10.1097/01.all.0000011053.76412.14. [DOI] [PubMed] [Google Scholar]
- 3.Sheehan WJ, Rangsithienchai PA, Wood RA, et al. Pest and allergen exposure and abatement in inner-city asthma: a work group report of the American Academy of Allergy, Asthma & Immunology Indoor Allergy/Air Pollution Committee. J Allergy Clin Immunol. 2010;125:575–81. doi: 10.1016/j.jaci.2010.01.023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Brasó-Maristany F, Filosto S, Catchpole S, et al. PIM1 kinase regulates cell death, tumor growth and chemotherapy response in triple-negative breast cancer. Nat Med. 2016;22:1303–13. doi: 10.1038/nm.4198. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Li H, Xie L, Zhu L, et al. Multicellular immune dynamics implicate PIM1 as a potential therapeutic target for uveitis. Nat Commun. 2022;13:5866. doi: 10.1038/s41467-022-33502-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Linowes BA, Ligons DL, Nam AS, et al. Pim1 permits generation and survival of CD4+ T cells in the absence of γc cytokine receptor signaling. Eur J Immunol. 2013;43:2283–94. doi: 10.1002/eji.201242686. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Shin YS, Takeda K, Shiraishi Y, et al. Inhibition of Pim1 kinase activation attenuates allergen-induced airway hyperresponsiveness and inflammation. Am J Respir Cell Mol Biol. 2012;46:488–97. doi: 10.1165/rcmb.2011-0190OC. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Liu Z, Li X, Gao Y, et al. Epigenetic reprogramming of Runx3 reinforces CD8 + T-cell function and improves the clinical response to immunotherapy. Mol Cancer. 2023;22:84. doi: 10.1186/s12943-023-01768-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Milner JJ, Toma C, Yu B, et al. Runx3 programs CD8+ T cell residency in non-lymphoid tissues and tumours. Nature New Biol. 2017;552:253–7. doi: 10.1038/nature24993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Haider A, Steininger A, Ullmann R, et al. Inactivation of RUNX3/p46 Promotes Cutaneous T-Cell Lymphoma. J Invest Dermatol. 2016;136:2287–96. doi: 10.1016/j.jid.2016.05.126. [DOI] [PubMed] [Google Scholar]
- 11.Overgaard NH, Jung JW, Steptoe RJ, et al. CD4+/CD8+ double-positive T cells: more than just a developmental stage? J Leukoc Biol. 2015;97:31–8. doi: 10.1189/jlb.1RU0814-382. [DOI] [PubMed] [Google Scholar]
- 12.Zhu X, Li W, Gao J, et al. RUNX3 improves CAR-T cell phenotype and reduces cytokine release while maintaining CAR-T function. Med Oncol . 2023;40:89. doi: 10.1007/s12032-022-01913-7. [DOI] [PubMed] [Google Scholar]
- 13.Wang M, Okamoto M, Domenico J, et al. Inhibition of Pim1 kinase prevents peanut allergy by enhancing Runx3 expression and suppressing T(H)2 and T(H)17 T-cell differentiation. J Allergy Clin Immunol. 2012;130:932–44. doi: 10.1016/j.jaci.2012.07.032. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Liu H, Chen C, Ma D, et al. Inhibition of PIM1 attenuates the stem cell–like traits of breast cancer cells by promoting RUNX3 nuclear retention. J Cellular Molecular Medi. 2020;24:6308–23. doi: 10.1111/jcmm.15272. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Basharat S, Jabeen U, Zeeshan F, et al. Adherence to asthma treatment and their association with asthma control in children. J Pak Med Assoc. 2018;68:725–8. [PubMed] [Google Scholar]
- 16.Ohno I. Neuropsychiatry phenotype in asthma: Psychological stress-induced alterations of the neuroendocrine-immune system in allergic airway inflammation. Allergol Int. 2017;66S:S2–8. doi: 10.1016/j.alit.2017.06.005. [DOI] [PubMed] [Google Scholar]
- 17.Alobaidi AH, Alsamarai AM, Alsamarai MA. Inflammation in Asthma Pathogenesis: Role of T Cells, Macrophages, Epithelial Cells and Type 2 Inflammation. AIAAMC . 2021;20:317–32. doi: 10.2174/1871523020666210920100707. [DOI] [PubMed] [Google Scholar]
- 18.Harb H, Stephen-Victor E, Crestani E, et al. A regulatory T cell Notch4-GDF15 axis licenses tissue inflammation in asthma. Nat Immunol. 2020;21:1359–70. doi: 10.1038/s41590-020-0777-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Miller RL, Grayson MH, Strothman K. Advances in asthma: New understandings of asthma’s natural history, risk factors, underlying mechanisms, and clinical management. J Allergy Clin Immunol. 2021;148:1430–41. doi: 10.1016/j.jaci.2021.10.001. [DOI] [PubMed] [Google Scholar]
- 20.Ainge GD, Hudson J, Larsen DS, et al. Phosphatidylinositol mannosides: Synthesis and suppression of allergic airway disease. Bioorg Med Chem. 2006;14:5632–42. doi: 10.1016/j.bmc.2006.04.037. [DOI] [PubMed] [Google Scholar]
- 21.Didichenko SA, Spiegl N, Brunner T, et al. IL-3 induces a Pim1-dependent antiapoptotic pathway in primary human basophils. Blood. 2008;112:3949–58. doi: 10.1182/blood-2008-04-149419. [DOI] [PubMed] [Google Scholar]
- 22.Liu H, Li L, Hao Y, et al. Identification of two migratory colon ILC2 populations differentially expressing IL-17A and IL-5/IL-13. Sci China Life Sci. 2023;66:67–80. doi: 10.1007/s11427-022-2127-2. [DOI] [PubMed] [Google Scholar]
- 23.Olaguibel J, Sastre J, Rodríguez J, et al. Eosinophilia Induced by Blocking the IL-4/IL-13 Pathway: Potential Mechanisms and Clinical Outcomes. J Investig Allergol Clin Immunol . 2022;32:165–80. doi: 10.18176/jiaci.0823. [DOI] [PubMed] [Google Scholar]
- 24.Xu J, Zhu X, Li Q, et al. Loss of PIM1 correlates with progression and prognosis of salivary adenoid cystic carcinoma (SACC) Cancer Cell Int. 2018;18:22. doi: 10.1186/s12935-018-0518-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Alexandrova E, Miglino N, Hashim A, et al. Small RNA profiling reveals deregulated phosphatase and tensin homolog (PTEN)/phosphoinositide 3-kinase (PI3K)/Akt pathway in bronchial smooth muscle cells from asthmatic patients. J Allergy Clin Immunol. 2016;137:58–67. doi: 10.1016/j.jaci.2015.05.031. [DOI] [PubMed] [Google Scholar]
- 26.Xue JF, Shi ZM, Zou J, et al. Inhibition of PI3K/AKT/mTOR signaling pathway promotes autophagy of articular chondrocytes and attenuates inflammatory response in rats with osteoarthritis. Biomed Pharmacother. 2017;89:1252–61. doi: 10.1016/j.biopha.2017.01.130. [DOI] [PubMed] [Google Scholar]
- 27.Tian Q, Guo Y, Feng S, et al. Inhibition of CCR2 attenuates neuroinflammation and neuronal apoptosis after subarachnoid hemorrhage through the PI3K/Akt pathway. J Neuroinflamm. 2022;19:312. doi: 10.1186/s12974-022-02676-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Chen X, Yang J, Shen H, et al. Muc5ac Production Inhibited by Decreased lncRNA H19 via PI3K/Akt/NF-kB in Asthma. J Asthma Allergy . 2021;14:1033–43. doi: 10.2147/JAA.S316250. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Wu D, Li S, Liu X, et al. Alpinetin prevents inflammatory responses in OVA-induced allergic asthma through modulating PI3K/AKT/NF-κB and HO-1 signaling pathways in mice. Int Immunopharmacol. 2020;89:107073. doi: 10.1016/j.intimp.2020.107073. [DOI] [PubMed] [Google Scholar]
- 30.Yang N, Zhang H, Cai X, et al. Epigallocatechin-3-gallate inhibits inflammation and epithelial‑mesenchymal transition through the PI3K/AKT pathway via upregulation of PTEN in asthma. Int J Mol Med. 2018;41:818–28. doi: 10.3892/ijmm.2017.3292. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
No data are available.