Abstract
Oral cancer, the most prevalent cancer worldwide, is far more likely to occur after the age of forty-five, according to the World Health Organization. Although many biomarkers have been discovered over the years using non-invasive saliva samples, biopsies, and human blood, these biomarkers have not been incorporated into standard clinical practice. Investigating the function of microRNAs (miRNAs) in the diagnosis, aetiology, prognosis, and treatment of oral cancer has drawn more attention in recent years. Though salivary microRNA can act as a window into the molecular environment of the tumour, there are challenges due to the heterogeneity of oral squamous cell carcinoma (OSCC), diversity in sample collection, processing techniques, and storage conditions. The up and downregulation of miRNAs has been found to have a profound role in OSCC as it regulates tumour stages by targeting many genes. As a result, the regulatory functions of miRNAs in OSCC underscore their significance in the field of cancer biology. Salivary miRNAs are useful diagnostic and prognostic indicators because their abnormal expression profiles shed light on tumour behaviour and patient prognosis. In addition to their diagnostic and prognostic value, miRNAs hold promise as therapeutic targets for oral cancer intervention. The current review sheds light on the challenges and potentials of microRNA studies that could lead to a better understanding of oral cancer prognosis, diagnosis, and therapeutic intervention. Furthermore, the clinical translation of OSCC biomarkers requires cooperation between investigators, physicians, regulatory bodies, and business partners. There is much potential for improving early identification, tracking therapy response, and forecasting outcomes in OSCC patients by including saliva-based miRNAs as biomarkers.
Keywords: Oral squamous cell carcinoma (OSCC), microRNAs (miRNAs), Salivary biomarkers, Diagnosis and prognosis, Therapeutic targets
Introduction
A crucial global health concern is oral cancer, which is categorized under general oral cancer groups. It is expected to have an annual incidence of over 300,000 cases and causes over 170,000 deaths annually, making it the sixteenth most prevalent cancer globally. These malignancies affect the gums, tongue, lips, floor of the mouth, and other oral cavity components [1]. These malignancies affect the gums; as per the WHO, the risk of oral cancer increases significantly beyond the age of forty-five. Additionally, the five-year survival rates differ significantly based on the stage of cancer diagnosis. Eighty per cent of patients with a stage I survive for five years, while only twenty per cent of patients with a stage III or IV has about five years of survival period [2]. Although many biomarkers have been discovered over the years using non-invasive saliva samples, biopsies, and human blood, these biomarkers have not been incorporated into standard clinical practice.
Recently, increased interest in studying microRNAs (miRNAs) and their role in oral cancer’s aetiology, diagnosis, prognosis, and therapy. Small RNA molecules known as miRNAs are crucial for regulating post-transcriptional genes because they are non-coding. Their method is binding to the 3’ untranslated region of the target mRNAs, which results in mRNA destruction or translational repression. Oral cancer is among the cancers where miRNA dysregulation has been connected, as miRNAs can function as oncogenes or tumour suppressors [3].
Numerous bodily fluids contain these miRNAs, such as blood, urine, and saliva. Saliva is unique among these biological samples because of its practical and non-invasive collection technique with significant advantages. It is inexpensive, simple to gather, and provides numerous sample opportunities—all of which are especially helpful for tracking the course of the disease and how well a medication works. Furthermore, saliva collection improves healthcare personnel’s safety by decreasing the chance of infectious agent transmission. Salivary biomolecules have demonstrated potential as a reliable diagnostic agent for identifying hormonal imbalances, genetic diseases, and a variety of cancers. They have a great deal of promise for highly reliable monitoring of prognosis, illness development, and systemic health [4].
The direct interaction of saliva with oral cancers improves its detection capacity. Saliva can act as a window into the molecular environment of the tumour due to the presence of microRNA markers and other cellular and molecular changes associated with the tumour. Due to their critical involvement in gene regulation, miRNAs have emerged as significant biomarkers for several malignancies, including OSCC (oral squamous cell carcinoma). MiRNAs’ stability, endurance, and quantitative measurability in saliva make them reliable indicators of the onset and progression of cancer [3]. Recent technological advances have made it feasible to identify specific miRNA patterns in saliva that correspond with OSCC. This capability offers a promising new path for early prognosis, diagnosis, and therapy response monitoring in patients with oral cancer.
The use of saliva-based miRNA diagnostics in clinical settings could completely change how oral cancer is treated. These diagnostics can potentially improve patient outcomes by enabling earlier detection and more precise prognostication, thereby changing the paradigm in healthcare from reactive to proactive. It is imperative since early detection dramatically improves long-term survival and successful treatment prospects. Additionally, developing targeted therapeutics may be guided by understanding the molecular pathways behind oral cancer that saliva-based miRNA signatures can offer. There are still several obstacles to these encouraging advances. Standardization of saliva collection and miRNA detection techniques is crucial to guarantee accuracy and reproducibility in various therapeutic contexts.
Furthermore, more extensive research is required to determine the most reliable biomarkers for oral cancer and confirm the clinical usefulness of particular miRNA signatures. Effectively incorporating saliva-based miRNA diagnostics into standard clinical practice will depend on how these issues are resolved. The transformational promise of this burgeoning subject will be highlighted as this review delves into the scientific underpinning, detection methodologies, clinical research, advantages, limitations, and future potential of saliva-based miRNA diagnostics in oral cancer.
Overview of the current biomarkers for OSCC
One of the most prevalent types of oral cancer is oral squamous cell carcinoma (OSCC), for which early diagnosis is essential for better prognosis of the patient. Numerous biomarkers, such as DNA methylation indicators, non-coding RNAs like microRNAs (miRNAs), and protein biomarkers have been identified for diagnosis and prognosis of OSCC. Protein biomarkers, such as p53, a tumor suppressor protein that is often altered in OSCC and associated with a poor prognosis, have attracted a lot of attention in many studies [5–9]. Similarly, increased expression of EGFR (Epidermal Growth Factor Receptor) has been associated with poor survival rate and tumor development in OSCC [10]. Cyclin D1, protein that’s important for controlling the cell cycle, was observed to be often overexpressed and many literatures associated the overexpression of cyclin D1 to aggressiveness of the lesion [11]. DNA methylation indicators have demonstrated promise in OSCC diagnosis in addition to proteins. Genes like p16 and DAPK1 frequently exhibit aberrant methylation patterns. The prognosis of OSCC has been associated to hypermethylation of the p16 gene promoter region, indicating increased expression of DAPK1 (Death-Associated Protein Kinase 1) to poor prognosis [12, 13].
MicroRNAs, or miRNAs, have become a promising biomarker for OSCC diagnosis and prognosis in recent years. Small non-coding RNA molecules known as miRNAs control the expression of genes and are essential to many biological processes in carcinogenesis. miR-21 is one valid predictive and prognostic biomarker since its overexpression in OSCC has been demonstrated to stimulate tumor growth, invasion, and treatment resistance [13–15]. Similarly, miR-375 is usually downregulated in OSCC and has been associated to advanced tumor stages and poorer prognoses [15, 16] (Table 1). Number of studies has been conducted with saliva as sample for miRNA analysis in OSCC because of its non-invasive nature to collect and also that miRNAs are very stable in body fluids [13]. MiRNAs especially those detectable in saliva, offer a novel and promising strategy for non-invasive OSCC biomarker research, even if protein and DNA methylation markers also offer valuable diagnostic and prognostic information. By interdisciplinary research techniques, these biomarkers must be validated, confirmed and incorporated into clinical practice. The role of miRNAs in OSCC has grown in terms of pathogenesis, metastasis, and resistance to therapeutic agents [41].
Table 1.
MicroRNAs involved in oral squamous cell carcinoma (OSCC)
| S.no | miRNA | Role in OSCC | Mechanism | Regulation | References |
|---|---|---|---|---|---|
| 1. | miR-21 | Promotes cell proliferation and survival | Targets tumor suppressor genes (TSG) PTEN and PDCD4 | Upregulated | [17] |
| 2. | miR-155 | Inhibits apoptosis | Targets pro-apoptotic genes | Upregulated | [18] |
| 3. | miR-31 | Facilitates tumor invasion and metastasis | Targets genes involved in cell adhesion and extracellular matrix breakdown to regulate the epithelial-mesenchymal transition (EMT). | Upregulated | [19] |
| 4. | miR-10b | Promotes metastasis | Targets genes related to the breakdown of extracellular matrix and cell adhesion | Upregulated | [20] |
| 5. | miR-210 | Promotes angiogenesis | Upregulated under hypoxic conditions, targets angiogenic pathway genes | Upregulated | [21] |
| 6. | miR-375 | Correlates with advanced tumor stages and poorer outcomes | Targets PDK1, leading to suppressed tumor growth | Downregulated | [22] |
| 7. | miR-196a | Associated with increased tumor aggressiveness and poor survival rates | Mechanism not fully elucidated, but linked to tumor progression | Upregulated | [23] |
| 8. | miR-99a | Associated with tumor progression | Targets mTOR signaling pathway components | Downregulated | [24] |
| 9. | miR-100 | Linked to poor prognosis | Targets mTOR signaling pathway components | Downregulated | [24] |
| 10. | miR-34a | Inhibits OSCC cell growth | Mimics function as tumor-suppressive miRNAs | Downregulated | [25] |
| 11. | miR-200c | Inhibits EMT and metastasis | Targets ZEB1 and ZEB2, which are EMT regulators | Downregulated | [26] |
| 12. | miR-21-5p | Promotes tumor growth and invasion | Targets PTEN, which causes the PI3K/AKT pathway to be activated. | Upregulated | [27] |
| 13. | miR-146a | Modulates inflammatory response and cell proliferation | Targets IRAK1 and TRAF6, which are part of the NF-κB pathway. | Downregulated | [28] |
| 14. | miR-124 | Suppresses tumor growth and invasion | Targets CDK6, leading to cell cycle arrest | Downregulated | [29] |
| 15. | miR-204 | Inhibits proliferation and metastasis | Targets BCL2 and promotes apoptosis | Downregulated | [30] |
| 16. | miR-17-92 cluster | Promotes cell proliferation and survival | Targets several TSGs, including PTEN and TGFBR2 | Upregulated | [31] |
| 17. | miR-221/222 | Enhances proliferation and invasion | Targets p27Kip1 and TIMP3, which are involved in inhibiting invasion and regulating the cell cycle | Upregulated | [32] |
| 18. | miR-205 | Inhibits EMT and promotes epithelial phenotype | Targets ZEB1 and ZEB2 | Downregulated | [33] |
| 19. | miR-23b | Inhibits the growth and invasion of cells | Targets MET and induces cell death | Downregulated | [34] |
| 20. | miR-191 | Promotes cell proliferation and invasion | Targets TIMP3 and CDK6 | Upregulated | [35] |
| 21. | miR-203 | Suppresses tumor growth and EMT | Targets Src and promotes cell differentiation | Downregulated | [36] |
| 22. | miR-7 | Inhibits cell proliferation and metastasis | Targets EGFR and RAF1 | Downregulated | [37] |
| 23. | miR-26a | Inhibits cell proliferation | Targets EZH2 and promotes apoptosis | Downregulated | [38] |
| 24. | miR-29b | Inhibits EMT and metastasis | Targets MMP2 and MMP9, which are involved in matrix degradation | Downregulated | [39] |
| 25. | miR-125b | Prevents the invasion and proliferation of cells | Targets MMP13 and induces apoptosis | Downregulated | [40] |
Potential of MiRNA biomarkers in OSCC
MiRNA can interact with genes to change their expression, impacting essential biological processes such as cell cycle regulation, apoptosis, migration, invasion, and angiogenesis [42]. MiR-21, acts as a critical link in oncogenesis and inversely correlated with TSGs such as PTEN and PDCD4. MiR-155 exerted a pro-metastatic effect on ARID2 (AT-rich interaction domain 2) by significantly suppressing its protein level and controlling the biological conduct of OSCC. MiRNAs regulate the intrinsic and extrinsic pathways of apoptosis, and a faulty apoptotic pathway may be brought on by the dysregulation of miRNAs [43]. MiRNAs like miR-31 and miR-10b have been shown to modulate the EMT (Epithelial mesenchymal transition), involving extracellular matrix breakdown and cell adhesion, which is an essential stage in cancer metastasis. Angiogenesis is necessary for a tumour’s growth, and dissemination is also regulated by MiR-210, which targets genes linked to angiogenic pathways in hypoxic environments [44].
Advanced tumour stages and poor patient outcomes are correlated with miR-375, which is typically downregulated in OSCC and targets the 3’ UTR of PDK1 [45]. Similarly, low survival rates are associated with miR-196a, which is often upregulated and associated with increased tumour aggressiveness and may be a prognostic marker. miR-100 and miR-99a, which are both downregulated in OSCC and target elements of the mTOR signalling pathway, are linked to cancer growth and a poor prognosis [1].
Since miRNAs can be targets for innovative therapies. MiRNA mimics like miR-34a mimic are introduced to bring back the function of tumour-suppressive miRNAs, which have shown promise in preclinical trials by decreasing OSCC cell proliferation by modulating miRNA levels. Furthermore, oncogenic miRNAs can be silenced with miRNA inhibitors, such as antagomirs or locked nucleic acid (LNA) inhibitors [46]. Of these, inhibitors of miR-21 are effective in decreasing tumour development and increasing OSCC cell sensitivity to chemotherapy. Oncogenic miRNAs can also be reserved by miRNA sponges, RNA molecules engineered to trap particular miRNAs and stop them from interacting with their target mRNAs [47].
Salivary MicroRNA in OSCC
The regulatory functions of miRNAs in OSCC underscore their significance in the field of cancer biology. They are useful diagnostic and prognostic indicators because their abnormal expression profiles shed light on tumour behaviour and patient prognosis. Salivary microRNAs (miRNAs) have been found to originate from OSCC tissues, providing a non-invasive means of detecting these biomarkers in patients with oral cancer [48]. Significant promise exists for enhancing OSCC diagnosis and treatment outcomes by saliva-based miRNA profiling. Further investigation into the complex networks of miRNA regulatory profiles and their identification in saliva is expected to provide new insights into OSCC pathogenesis and lead to the development of innovative treatment approaches [49].
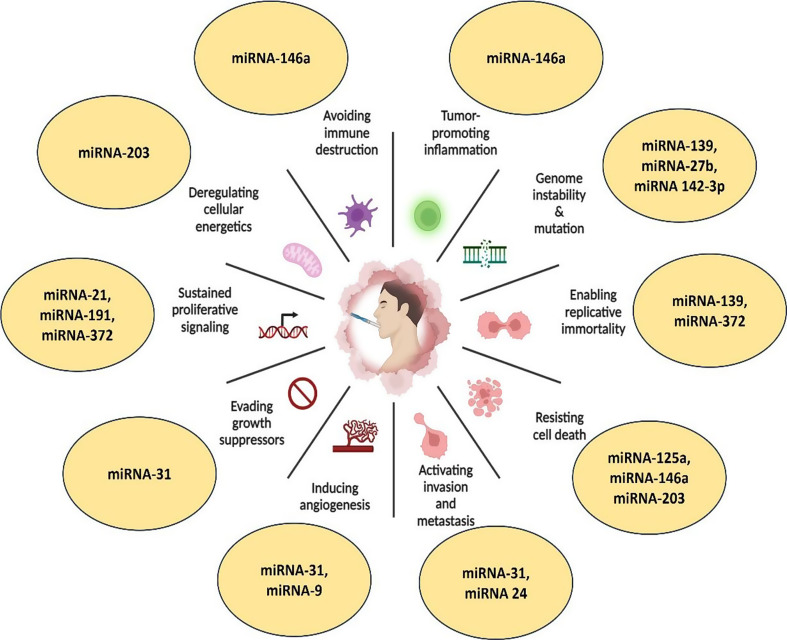
Studying salivary miRNAs has become increasingly attractive for oral squamous cell cancer (OSCC) (Table 2). The potential of miRNAs in saliva for non-invasive cancer diagnosis, especially for OSCC, has drawn interest in recent years (Fig. 1). The strategy takes advantage of the fact that miRNAs are stable in saliva and indicate the disease of the oral cavity, which qualifies them as good candidates for OSCC early identification and tracking [55].
Table 2.
Salivary microRNAs associated with oral squamous cell carcinoma (OSCC)
| miRNA | Sample type | Role in OSCC | Mechanism | Regulation | References |
|---|---|---|---|---|---|
| miR-21 | Whole saliva | Promotes cell proliferation and survival | Targets TSG, PTEN and PDCD4 | Upregulated | [50] |
| miR-31 | Whole saliva | Facilitates tumor invasion and metastasis | Regulates EMT by targeting genes involved in cell adhesion and ECM degradation | Upregulated | [51] |
| miR-184 | Saliva supernatant | Associated with tumor progression and aggressiveness | Targets AKT2, affecting cell proliferation and apoptosis | Upregulated | [52] |
| miR-125a | Saliva supernatant | Suppresses tumor growth | Targets ERBB2, affecting cell proliferation | Downregulated | [52] |
| miR-200a | Whole saliva | Inhibits EMT and metastasis | Targets ZEB1 and ZEB2, which are EMT regulators | Downregulated | [50] |
| miR-146a | Saliva supernatant | Modulates inflammatory response and cell proliferation | Targets TRAF6 and IRAK1, involved in NF-κB pathway | Downregulated | [53] |
| miR-139 | Whole saliva | Suppresses tumor growth and invasion | Targets MMP11, affecting cell migration | Downregulated | [54] |
| miR-372 | Saliva supernatant | Promotes cell proliferation | Targets LATS2, affecting Hippo signaling pathway | Upregulated | [54] |
| miR-155 | Whole saliva | Inhibits apoptosis | Targets pro-apoptotic genes | Upregulated | [50] |
| miR-9 | Saliva supernatant | Promotes cell migration and invasion | Targets E-cadherin, affecting cell adhesion | Upregulated | [53] |
| miR-24 | Whole saliva | Associated with tumor aggressiveness | Targets Bim, affecting apoptosis | Upregulated | [51] |
| miR-27b | Saliva supernatant | Suppresses cell proliferation | Targets MYC, affecting cell cycle progression | Downregulated | [54] |
| miR-142-3p | Whole saliva | Inhibits cell proliferation and migration | Targets CDK4, affecting cell cycle regulation | Downregulated | [52] |
| miR-486-5p | Saliva supernatant | Correlates with advanced tumor stages | Targets OLFM4, affecting cell adhesion and invasion | Downregulated | [51] |
| miR-191 | Whole saliva | Promotes cell proliferation and invasion | Targets TIMP3 and CDK6 | Upregulated | [50] |
| miR-203 | Saliva supernatant | Suppresses tumor growth and EMT | Targets Src, promoting cell differentiation | Downregulated | [53] |
Fig. 1.
Schematic representation of salivary miRNAs associated with the hallmarks of cancer in OSCC
One of the main reasons salivary miRNAs are considered potential biomarkers is their exceptional stability [56]. Although primarily produced by salivary gland cells, saliva can contain molecules from systemic circulation, making it a valuable biofluid for non-invasive diagnostics [57, 58]. Of all body fluids saliva/ whole mouth fluid, which is considered to be the most non-invasive and easy to collect. Since saliva is largely derived from the systemic circulation, its biochemical composition and properties are closely analogous to those of blood. Furthermore, miRNAs can bind to RNA-binding proteins like AGO2 (Argonaute 2) to form complexes that protect them from being broken down by nucleases. Due to these defense mechanisms, miRNAs in saliva are preserved and observable, which makes them trustworthy options for the identification of biomarkers. Their ability to withstand being exposed to a biological fluid that is easily accessible facilitates their application in the early identification, diagnosis, and prognostication of OSCC. This presents a viable substitute for more intrusive biopsy-based methods [59].
Differentially expressed salivary miRNAs between OSCC patients and healthy individuals have been found in multiple investigations. For instance, it has been repeatedly observed that OSCC has elevated levels of miR-21, one of the miRNAs investigated the most in cancer research, which targets TSGs like PTEN and PDCD4 and enhances cell survival and proliferation [60, 61]. MiR-31, on the other hand, is increased in OSCC and promotes tumour invasion and metastasis by controlling the process of the EMT, which includes modifications to cell adhesion and extracellular matrix breakdown [62].
The aggressiveness and tumour progression in OSCC have been associated with another miRNA, miR-184, which affects cell division and death by targeting the AKT2 gene. The saliva of OSCC patients likewise exhibits a substantial upregulation of this miRNA. On the other hand, the downregulated miR-125a in OSCC functions as a tumour suppressor by blocking the growth of cells via targeting the ERBB2 gene [63]. By focusing on ZEB1 and ZEB2, important EMT regulators, the downregulation of miR-200a in OSCC also aids in suppressing EMT and metastasis. The development of OSCC depends on controlling inflammation and the immunological response, and miR-146a is involved in both processes [64]. It is usually downregulated in OSCC and targets TRAF6 and IRAK1, components of the NF-κB pathway, to influence the inflammatory response and cell proliferation.
Similarly, OSCC has downregulated miR-139, which targets MMP11 and inhibits tumour development and invasion by influencing cell movement [65]. Another interesting miRNA is miR-372, which is increased in OSCC and targets LATS2, a part of the Hippo signalling system, to encourage cell proliferation. Conversely, miR-27b, suppressed in OSCC, targets the MYC gene to inhibit cell proliferation, affecting the cell cycle’s progression [66].
MiR-9, which targets E-cadherin and is elevated in OSCC, also influences the control of cell invasion and migration in this situation. Furthermore, by targeting the pro-apoptotic gene Bim, miR-24, which has been connected to tumour aggressiveness, is elevated and influences apoptosis. Other notable miRNAs in OSCC include miR-142-3p and miR-486-5p, which are also downregulated [67]. Targeting the critical cell cycle regulator CDK4, miR-142-3p prevents cell proliferation and migration, whereas miR-486-5p, which targets the gene OLFM4, which is involved in cell adhesion and invasion, is correlated with advanced tumour stages [68]. Moreover, OSCC has increased expression of miR-191, which targets TIMP3 and CDK6 to encourage cell invasion and proliferation. On the other hand, OSCC has downregulated miR-203, which inhibits tumour growth and EMT by targeting Src and promoting cell differentiation [69].
Saliva sample types
Cellular miRNAs found in saliva/whole mouth fluid are produced from immune cells, salivary gland secretions, and exfoliated oral mucosa cells. These miRNAs are pivotal for diseases like OSCC [70], as they can offer a broad picture of the molecular landscape within the oral cavity. When these miRNAs attach to the 3’ untranslated regions (3’ UTRs) of messenger RNAs (mRNAs), they cause translational repression or mRNA degradation. These miRNAs are released during cellular processes like death and secretion. In the oral cavity, this mechanism is essential for controlling oncogenic and tumor-suppressive pathways [71, 72].
Saliva, on the other hand, contains miRNAs, which represent localized and systemic cancer-related molecular alterations [73]. To prevent RNAse-mediated degradation, these extracellular miRNAs are frequently encapsulated in EVs or linked to proteins such as Argonaute-2 (Ago2). Saliva is a useful tool because of its abundance in extracellular miRNAs, which reduces the possibility of cellular miRNA contamination. The extracellular miRNAs in saliva supernatant, such as miR-21, miR-31, and miR-184, have been implicated in disease progression by targeting genes involved in cellular proliferation, apoptosis, and EMT [74]. miR-31 plays a dual role by enhancing invasion and metastasis through pathways like PI3K/AKT and modulating cellular migration via EMT regulatory genes [75, 76].
The encapsulation of extracellular miRNAs in vesicles, such as exosomes, facilitates their stability in saliva and permits efficient intercellular signaling, potentially serving as cancer biomarkers [77, 78]. By modifying gene expression in recipient cells, these miRNAs can contribute to the pathogenesis of OSCC. By modifying gene expression in recipient cells, these miRNAs can contribute to the pathogenesis of OSCC. As a result, determining the miRNA concentration in saliva supernatant provides a focused method for comprehending the molecular mechanisms underlying OSCC and makes it easier to identify particular miRNAs linked to the advancement of the illness and resistance to treatment.
MicroRNA quantification techniques
The highly sensitive technique of qRT-PCR makes it possible to precisely quantify the miRNAs implicated in the evolution of OSCC. A more comprehensive perspective is made possible by next-generation sequencing (NGS), which profiles the full miRNA transcriptome and finds novel miRNAs like miR-184 and miR-125a that contribute to the pathophysiology of OSCC. By blocking FIH-1, which raises HIF-1α activity and supports cancer cell viability under hypoxia, miR-184 supports OSCC. Tumor suppressor miR-125a controls apoptosis; its downregulation accelerates the growth of OSCC [79]. This method sheds light on the intricate regulation of biological processes like metastasis, apoptosis, and proliferation.
The assessment of therapy responses and OSCC progression can be aided by tracking the dynamics of miRNA throughout time. The EMT-inhibiting miR-200a is essential for preventing metastasis. The degree of its expression can be tracked during therapy to evaluate the effectiveness of the medication. On the other hand, miR-155 and miR-372, associated with more aggressive tumor characteristics, contribute to oncogenesis by their inhibition of apoptosis and targeting of tumor suppressor genes, respectively. Their expression functions as a possible therapeutic target and a predictor of a bad prognosis [18, 74, 76].
Low-abundance miRNAs can be detected using newly developed methods like digital droplet PCR (ddPCR), which divides material into thousands of droplets to increase sensitivity. In the event that standard approaches fail to identify important markers, this is critical for early identification of OSCC [80, 81]. Furthermore, real-time, label-free salivary miRNA detection is being achieved through the development of nanotechnology-based biosensors, which may boost specificity and sensitivity [77–83].
Clinical research on salivary miRNA markers in oral cancer
New research has illuminated the role of particular miRNAs in developing oral cancer and their possible therapeutic applications. Developing and validating a novel saliva-based miRNA for the early diagnosis and prediction of oral cancer was the goal of a recent study by Balakittnen et al. [61]. Six differentially expressed miRNAs (miR-7-5p, miR-10b-5p, miR-182-5p, miR-215-5p, miR-431-5p, miR-486-3p, miR-3614-5p, and miR-4707-3p) were found by researchers using saliva samples and data from The Cancer Genome Atlas (TCGA). Their results suggested that these novel salivary miRNAs might revolutionize oral cancer management by aiding diagnosis and prediction [61, 84]. A pilot study was carried out to investigate blood and salivary miRNAs associated with oral cancer, with a focus on smoking status stratification. To ensure that smokers and non-smokers were equally distributed in each group, researchers took serum and saliva samples from 23 oral cancer patients as well as 21 healthy volunteers. According to their research, miR-21 is especially significant in those who have smoked in the past, and blood and salivary levels of miR-136, miR-3928, and miR-29b may also be linked to oral cancer, even in its early stages. They stressed, however, that more validation in a larger cohort of participants with premalignant and early malignant lesions is necessary to validate these correlations [85].
Salivary miRNA-21 and miRNA-184 have also been studied for their potential as diagnostic tools for OSCC and oral potentially malignant diseases (OPMD). The 90 volunteers the researchers enrolled included healthy controls, OPMD patients, and OSCC patients. To determine the expression levels of miRNA-21 and miRNA-184, they extracted RNA from saliva samples and used qRT-PCR. MiRNA-21 and miRNA-184 expression levels in OSCC, OPMD, and healthy controls were found to differ significantly (p < 0.001) from one another. It is possible that these miRNAs could be used for early detection because they were able to distinguish OSCC and OPMD from the control group. Saline is a readily available, non-invasive method for identifying malignant alterations in the oral mucosa, according to the study [86].
Furthermore, a study evaluating the function, sensitivity, and specificity of salivary LINC00657 and miRNA-106a as diagnostic markers in OSCC patients was retrospectively registered on clinicaltrials.gov under the identifier NCT05821179. The study was first registered on Mar 26, 2023, with the official registration date being Apr 19, 2023. These markers were compared to healthy individuals and patients with oral lichen planus (OLP), a sign of potentially malignant oral diseases. Thirty-six volunteers were split into three groups for the study: Twelve patients with OSCC were in Group I, twelve patients with OLP were in Group II, and twelve healthy people without oral mucosal lesions were in Group III. Using quantitative real-time PCR, saliva samples from each participant were examined. According to the findings, LINC00657 was more accurate in diagnosing OSCC (83.3%) than miR-106a (80.4%).
On the other hand, LINC00657 (52.5%) and miR-106a had worse diagnostic accuracy (61%) when identifying OLP. Both markers showed similar diagnostic accuracy (75%) in differentiating between OSCC and OLP. According to the study, salivary LINC00657 has a high potential for differentiating OSCC from OLP, while low levels of miR-106a may be a sign of cancer. These findings suggest that both markers have promise as OSCC diagnostic tools.
In addition to miRNAs’ diagnostic and prognostic value, they also hold promise as therapeutic targets for oral cancer intervention. Preclinical studies utilizing miRNA-based therapeutics, such as miRNA mimics or antagomiRs, have shown promising results in suppressing tumour growth and enhancing chemosensitivity in oral cancer models. With technological advancements, point-of-care assays for salivary miRNA detection are on the horizon. Rapid and cost-effective diagnostic tests based on miRNA biomarkers could simplify the early detection of oral cancer in primary care settings, especially in resource-limited settings where access to specialized healthcare facilities is limited. Interestingly, salivary miRNAs have the potential to stratify individuals based on their risk of developing oral cancer. Integrating miRNA biomarkers into population-based screening programs may enable targeted surveillance and early intervention among high-risk populations, reducing disease burden and mortality. Integrating salivary miRNA analysis with further omics technologies (proteomics and metabolomics) holds promise for comprehensive biomarker discovery in oral cancer. Multi-omics approaches may uncover novel biomarker signatures with enhanced diagnostic and prognostic accuracy, providing a holistic understanding of disease pathogenesis. Beyond diagnostics, dysregulated miRNAs implicated in oral cancer pathogenesis represent potential therapeutic targets. Targeted modulation of aberrant miRNA expression using miRNA mimics or inhibitors could offer novel therapeutic avenues for precision medicine in oral cancer treatment. Clinical trials exploring the efficacy and safety of miRNA-based therapies are underway, and these findings are intended to be translated into clinical practice [70, 87].
Pitfalls of salivary miRNA in OSCC detection
Several concerns need to be considered when using salivary miRNAs as a reliable diagnostic method for OSCC. Intermittent findings might arise from a wide range of factors, including circadian rhythms, food, dental cleanliness, and sample-collecting techniques, all of which can affect the amounts of miRNAs in saliva. MiRNA analysis may be hampered by contamination from food particles, the oral microbiota, and other materials in the mouth cavity. The disparities across various research are made worse by the absence of established data collection, processing, and analysis processes. Furthermore, salivary miRNAs in the oral cavity are vulnerable to destruction by RNases, compromising the data’s stability and accuracy. Due to these challenges, alternative biofluids with distinct miRNA expression patterns and stability, like human peripheral blood and body fluids, are considered considerably more reliable for early cancer diagnosis and prognosis [88]. Recent studies have demonstrated that salivary exosomes, believed to contain thousands of miRNAs, have promise as non-invasive diagnostic biomarkers for OSCC and other cancers [79, 89].
Considering the pitfalls in using salivary miRNAs as diagnostic tools of OSCC, human peripheral blood and body fluids, due to their stability and unique expression pattern of miRNAs, make them perfect biomarkers for cancer prognosis and early diagnosis. Recent research indicates that thousands of miRNAs may be present in salivary exosomes, which may be utilized as non-invasive diagnostic indicators to find OSCC and other malignancies [90, 91].
Heterogeneity of microRNAs in OSCC
The fundamental heterogeneity of OSCC is reflected in its broad spectrum of tumours, each with a unique molecular profile, clinical characteristics, and responses to therapy. Because of this variability, miRNA expression patterns vary between patients, making it difficult to find universal biomarkers that work for all OSCC subtypes. This may affect salivary miRNA-based diagnostic accuracy, implying that tumour diversity must be carefully considered in the biomarker identification and validation process [48]. There is a need to focus on the study of the sub-specificity of OSCC because the aetiology and clinical features of SCC vary between oral mucosa. These studies can shed light on genetic variation and behavioural characteristics unique to oral cavity locations. Large-scale studies are needed to strengthen the rapidly growing field of miRNA research [92].
Non-specificity of miRNA expression
Salivary miRNAs exhibit dysregulation in various physiological and pathological circumstances, including stomatitis, infection, and systemic disorders, making them useful as biomarkers for identifying OSCC. Their specificity for detecting OSCC should be carefully compared with other oral and systemic diseases to reduce the risk of false positive results. A recent next-generation sequencing study shows that some miRNAs are expressed differently in oral cancer. The area under the curve (AUC) and receiver operating characteristic (ROC) analyses indicate that these miRNAs may help differentiate oral cancer patients from healthy persons [93–95]. Furthermore, supporting the clinical significance and prospective use of miRNAs as prognostic indicators, they have also been linked to survival in patients with oral cancer. However, careful validation and standardization of miRNA biomarkers are necessary to ensure their accuracy and reliability in OSCC diagnosis and prognostic assessment [96].
Influence of confounding factors
Environmental and lifestyle factors, including nutrition, oral hygiene practices, alcohol intake, and smoking, can significantly impact salivary miRNA expression levels, complicating their use as diagnostic biomarkers in OSCC. Dietary habits, such as high-fat diet, may alter saliva composition and miRNA content due to systemic inflammation could be affected [97–99]. To control for these influences, participants should follow standardized dietary guidelines or undergo fasting before saliva collection. Oral hygiene practices, including toothpaste and mouthwash use, can also affect saliva composition and miRNA levels. Antibacterial products may alter microbial flora and influence miRNA profiles. Therefore, participants should be instructed to avoid oral hygiene products for a specified period before saliva collection to minimize variability [100].
Moreover, inconsistent miRNA readings may result from differences in sample preparation and saliva collection techniques. Providing participants with instructions to abstain from eating, drinking, smoking, or engaging in oral hygiene activities before to collection, together with a regulated collection environment to minimize contamination, are essential components of standardized collection devices and procedures. MiRNA stability is also impacted by processing and storage circumstances, which calls for quick processing, a reduction in freeze-thaw cycles, and uniform procedures for RNA stabilizers and storage temperatures [100]. Age-related changes in miRNA levels should be carefully taken into account. A trustworthy control miRNA for normalization is miR-16-5p, which has been consistently abundant in a dataset of 922 healthy persons. By addressing these variables using established protocols, salivary miRNA biomarkers for OSCC diagnosis can be made more sensitive and accurate, and research reliability can be increased [102].
Ethical and regulatory considerations
Salivary miRNAs are being used in clinical settings as diagnostic tools for OSCC, but their responsible deployment requires careful consideration of ethical and regulatory considerations. One of the main ethical obligations is to get informed consent. It is imperative that patients receive comprehensive information regarding miRNA-based diagnostic tests, encompassing their intended use, possible advantages, and potential drawbacks. Maintaining patient autonomy and confidence requires open communication regarding test results intended use and how they may affect patient management [103]. To safeguard patient confidentiality, strict data privacy standards must also be followed when handling biological samples and patient data. To protect patient health information, researchers and healthcare providers must make sure that they are in accordance with rules and regulations like the Health Insurance Portability and Accountability Act (HIPAA) in the US and the General Data Protection Regulation (GDPR) in Europe [104].
The European Medicines Agency (EMA) or the U.S. Food and Drug Administration (FDA) must rigorously evaluate and approve salivary miRNA-based diagnostic assays before they can be incorporated into clinical practice [105]. As part of this procedure, the tests’ analytical and clinical performance are validated to guarantee their precision, dependability, and safety. In order to obtain clinical acceptance and verify that the tests fulfill the required quality and performance standards, adherence to regulatory standards is essential. By addressing these moral and legal issues, salivary miRNA diagnostics can be developed and integrated responsibly, guaranteeing their morally and effectively in clinical contexts [106]. The ICMR regulations, which place a strong emphasis on unambiguous communication in regional languages, regulate informed consent for salivary miRNA diagnostics in India. In accordance with international norms such as GDPR, patient confidentiality is safeguarded by the Personal Data Protection Bill (PDPB). The CDSCO is in charge of regulatory approval for diagnostics, and all clinical research must have ethics committee approval in accordance with ICMR guidelines. The Indian Council of Medical Research [107] states that these frameworks guarantee test safety, data confidentiality, and ethical compliance.
Sample collection and processing variability
Reliability and reproducibility of miRNA-based diagnostics can be negatively affected by biases and inconsistencies introduced into miRNA expression data due to differences in RNA extraction methods, storage conditions, and saliva sample collection methods. Accurate miRNA estimates are complicated by variables such as saliva viscosity, oral microbial contamination, and sample degradation over time. Standardized methods of saliva collection, processing and quality control are essential to overcome these problems as they reduce technical variation and ensure stable and reliable miRNA data [108, 109]. The development of repeatable techniques for manipulating saliva is required due to the increasing interest in using salivary cell-free (cf.) miRNA as a liquid biopsy indication in different forms of cancer. Controlling pre-analytical factors affecting miRNA stability is one way to successfully incorporate salivary cf-miRNA indicators into clinical settings. Researchers can improve the validity and feasibility of miRNA-based cancer detection and monitoring diagnostics by adopting standards used in saliva processing [110].
Limited understanding of miRNA function
Many salivary miRNAs have been concerned with the pathophysiology of OSCC, although their functional role in OSCC carcinogenesis remains unclear. To find valid biomarkers and treatment targets, it is critical to comprehend the molecular mechanisms behind miRNA-mediated regulatory networks in OSCC initiation, development, and metastasis. Integrative approaches combining miRNA profiling with functional genomics, pathway analysis and experimental validation can provide insight into the function of miRNAs and their clinical relevance in OSCC [1].
The dysregulation of specific miRNA expression profiles associated with various clinicopathological features has been frequently observed in mature miR expression profile studies in OSCC tumour samples. For instance, compared to normal keratinocytes in oral cancer cell lines, studies have shown upregulation of miR-10b, miR-196a, miR-196b, and others, while miR-503 and miR-31 were downregulated. Additionally, significant differences in the expression levels of certain miRNAs were observed in oropharyngeal malignancies, including let-7a, miR-200c, and miR-34a. Dysregulated miRNAs were also associated with regulating genes related to the PI3K/AKT and p53 signalling pathways involved in OSCC carcinogenesis. These miRNAs include let-7a, miR-16, miR-29b, and miR-144. However, the variability of miRNA expression profiles across studies underscores the need for patient selection, study design, and methodology standardization to ensure reliable and reproducible results. Differences in study results may result from differences in patient populations, experimental methods and the use of appropriate controls [48, 111]. Despite these obstacles, defining the unique regulatory networks and miRNA expression profiles of OSCC tumours may improve patient outcomes and understand the pathophysiology of the disease.
Normalization challenges
Eliminating systematic biases and variances during sample preparation and analysis necessitates normalizing miRNA expression data. However, due to the lack of globally stable miRNAs, finding suitable endogenous controls or reference miRNAs for normalization in saliva samples can be difficult. The importance of methodological optimization and validation in miRNA biomarker studies is further emphasized by the impact the normalization strategy (e.g., global mean normalization, spike-in controls) can have on data interpretation and diagnostic performance [112, 113].
A comprehensive approach is necessary to overcome the challenges of using salivary microRNA (miRNA) as a diagnostic tool for OSCC. Developing robust biomarker selection algorithms that consider the diversity of tumour subtypes and the comprehensive molecular profile of each is only one part of the multimodal strategy required to address the heterogeneity of OSCC [114]. To improve the specificity of salivary miRNA biomarkers, OSCC-specific miRNA panels should be identified and evaluated extensively against oral and systemic diseases. Minimizing variability and ensuring reproducibility of miRNA expression data requires careful consideration of confounding factors and standardization of sampling techniques. Improvements in sample handling methods and quality control methods are essential to reduce variability in saliva collection, RNA extraction, and end-stage analysis [115, 116].
Exploring new normalization techniques and finding reliable reference miRNAs with low variability across multiple sample populations is necessary to overcome normalization hurdles. Comprehensive methods integrating high-throughput profiling, functional genomics, pathway analysis and experimental validation in preclinical models and clinical data are needed to fully understand the functional significance of salivary miRNAs in the aetiology of OSCC. Determining the clinical validity and value of salivary miRNA biomarkers requires validation in large and well-defined cohorts of patients with diverse demographic and clinical characteristics. Improving the accessibility and cost-effectiveness of miRNA-based diagnostics requires the creation of affordable assay platforms, point-of-care testing techniques suitable for use in various healthcare settings, and optimized workflows. These challenges can be resolved collectively, advancing the field of salivary miRNAs as dependable, non-invasive markers for early diagnosis, prognosis, and personalized treatment of OSCC. This will benefit patients worldwide by reducing the disease burden and improving patient outcomes.
Conclusion
In conclusion, saliva-based miRNA signatures have considerable potential as non-invasive biomarkers for OSCC early identification and treatment. Saliva is a readily available and simple-to-collect diagnostic medium, which makes it an ideal choice for OSCC diagnosis. This review has discussed the possible applications of salivary miRNAs for OSCC diagnosis. A few issues must be resolved for these indicators to be therapeutically valuable. The considerations mentioned earlier encompass the variability of miRNA profiles, the non-specificity of miRNA expression, and the influence of confounding variables such as sample collection variability and the restricted comprehension of miRNA functions in OSCC.
Furthermore, using salivary miRNAs for accurate OSCC detection is difficult due to miRNA normalization and quantification difficulties. Transforming these biomarkers into proper diagnostic instruments will need overcoming these challenges via standardized procedures and additional studies into miRNA processes. By addressing these problems, more precise non-invasive diagnostic techniques will become available, improving early detection and patient outcomes for those with OSCC.
Author contributions
MP and MIK participated in the conception and design of the study; MP, RS, and MDP determined the workflow; MP, R, MDP and SRV conducted the data collection; MP, RS, and MDP wrote the manuscript; MIK reviewed and revised the manuscript. MIK supervised this work.
Funding
This research received no external funding.
Data availability
No datasets were generated or analysed during the current study.
Declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Footnotes
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
- 1.Wang J, Lv N, Lu X, Yuan R, Chen Z, Yu J. Diagnostic and therapeutic role of microRNAs in oral cancer (review). Oncol Rep. 2021;45(1):58–64. 10.3892/or.2020.7854. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Aghiorghiesei O, Zanoaga O, Nutu A, et al. The world of oral Cancer and its risk factors viewed from the aspect of MicroRNA expression patterns. Genes (Basel). 2022;13(4):594. 10.3390/genes13040594. Published 2022 Mar 26. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Rishabh K, Khadilkar S, Kumar A, Kalra I, Kumar AP, Kunnumakkara AB. MicroRNAs as modulators of oral Tumorigenesis-A focused review. Int J Mol Sci. 2021;22(5):2561. 10.3390/ijms22052561. PMID: 33806361; PMCID: PMC7961687. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Yoshizawa JM, Wong DT. Salivary microRNAs and oral cancer detection. Methods Mol Biol. 2013;936:313–24. 10.1007/978-1-62703-083-0_24. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Pekarek L, Garrido-Gil MJ, Sánchez-Cendra A, Cassinello J, Pekarek T, Fraile-Martinez O, García-Montero C, Lopez-Gonzalez L, Rios-Parra A, Álvarez-Mon M, Acero J, Diaz-Pedrero R, Ortega MA. Emerging histological and serological biomarkers in oral squamous cell carcinoma: applications in diagnosis, prognosis evaluation and personalized therapeutics (review). Oncol Rep. 2023;50(6):213. 10.3892/or.2023.8650. Epub 2023 Oct 20. PMID: 37859591; PMCID: PMC10620846. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Selvaraj J, Yasothkumar D, Vishnu Priya V, Raj AT, Babu SD, Patil S. Development and tumorigenic potential of TP53: a therapeutic target for head and neck squamous cell carcinoma. Oral Oncol. 2022;130:105922. 10.1016/j.oraloncology.2022.105922. Epub 2022 May 18. PMID: 35596977. [DOI] [PubMed] [Google Scholar]
- 7.Hyodo T, Kuribayashi N, Fukumoto C, et al. The mutational spectrum in whole exon of p53 in oral squamous cell carcinoma and its clinical implications. Sci Rep. 2022;12:21695. 10.1038/s41598-022-25744-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Liu R, Sun K, Wang Y, Jiang Y, Kang J, Ma H. The effects of proliferating cell nuclear antigen and p53 in patients with oral squamous cell carcinoma: a systematic review and meta-analysis. Ann Transl Med. 2021;9(23):1739. 10.21037/atm-21-6133. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Field JK, Pavelic ZP, Spandidos DA, Stambrook PJ, Jones AS, Gluckman JL. The role of the p53 tumor suppressor gene in squamous cell carcinoma of the Head and Neck. Arch Otolaryngol Head Neck Surg. 1993;119(10):1118–22. 10.1001/archotol.1993.01880220064009. [DOI] [PubMed] [Google Scholar]
- 10.Tan Y, Wang Z, Xu M, Li B, Huang Z, Qin S, Nice EC, Tang J, Huang C. Oral squamous cell carcinomas: state of the field and emerging directions. Int J Oral Sci. 2023;15(1):44. 10.1038/s41368-023-00249-w. PMID: 37736748; PMCID: PMC10517027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.De Paoli P. Bio-banking in microbiology: from sample collection to epidemiology, diagnosis and research. FEMS Microbiol Rev. 2005;29(5):897–910. 10.1016/j.femsre.2005.01.005. Epub 2005 Feb 26. PMID: 16219511; PMCID: PMC7110355. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Zhong L, Liu Y, Wang K, He Z, Gong Z, Zhao Z, Yang Y, Gao X, Li F, Wu H, Zhang S, Chen L. Biomarkers: paving stones on the road towards the personalized precision medicine for oral squamous cell carcinoma. BMC Cancer. 2018;18(1):911. 10.1186/s12885-018-4806-7. PMID: 30241505; PMCID: PMC6151070. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Prakash N, Pradeep GL, J Oral Maxillofac Pathol. Circulating biomarkers in oral cancer: Unravelling the mystery. 2022 Jul-Sep;26(3):300–306. doi: 10.4103/jomfp.jomfp_338_22. Epub 2022 Oct 17. PMID: 36588829; PMCID: PMC9802512. [DOI] [PMC free article] [PubMed]
- 14.Bodaghi A, Fattahi N, Ramazani A, Biomarkers. Promising and valuable tools towards diagnosis, prognosis and treatment of Covid-19 and other diseases. Heliyon. 2023;9(2):e13323. 10.1016/j.heliyon.2023.e13323. Epub 2023 Jan 30. PMID: 36744065; PMCID: PMC9884646. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Pitikultham P, Putnin T, Pimalai D, Sathirapongsasuti N, Kitiyakara C, Jiang Q, Ding B, Japrung D. Ultrasensitive Detection of MicroRNA in Human Saliva via Rolling Circle Amplification Using a DNA-Decorated Graphene Oxide Sensor. ACS Omega. 2023;8(17):15266–75. 10.1021/acsomega.3c00411. PMID: 37151566; PMCID: PMC10157686. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Bakker E, Starokozhko V, Kraaijvanger JWM, Heerspink HJL, Mol PGM. Precision medicine in regulatory decision making: biomarkers used for patient selection in European Public Assessment reports from 2018 to 2020. Clin Transl Sci. 2023;16(11):2394–412. 10.1111/cts.13641. Epub 2023 Oct 18. PMID: 37853917; PMCID: PMC10651650. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Volinia S, Calin GA, Liu CG, Ambs S, Cimmino A, Petrocca F, Croce CM. A microRNA expression signature of human solid tumors defines cancer gene targets. PNAS. 2006;103(7):2257–61. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Zhang T, Liu M, Wang C, Lin C, He L, Lei Y. miR-155 silencing induces apoptosis and enhances cisplatin chemosensitivity in OSCC. Biomed Pharmacother. 2013;67(4):387–94.23643356 [Google Scholar]
- 19.Lu YC, Chen YJ, Wang HM, Tsai CY, Chen WH, Huang YC, Chang KP. Oncogenic function and early detection potential of miRNA-31 in human squamous cell carcinoma. Oncogene. 2014;33(25):324–34. [Google Scholar]
- 20.Wang X, Liu S, Chen L, Fan Y. MicroRNA-10b promotes invasion by targeting Tiam1 in head and neck cancer. Cell Physiol Biochem. 2012;28(1):196–204. [Google Scholar]
- 21.Huang, X., Ding, L., Bennewith, K. L., Tong, R. T., Welford, S. M., Ang, K. K., Giaccia, A. J. (2009). Hypoxia-inducible mir-210 regulates normoxic gene expression involved in tumor initiation. Mol Cell, 35(6), 856–867. [DOI] [PMC free article] [PubMed]
- 22.Avissar M, Christensen BC, Kelsey KT, Marsit CJ. MicroRNA expression ratio is predictive of head and neck squamous cell carcinoma. Clin Cancer Res. 2009;15(8):2850–5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Gao W, Yu Y, Cao H, Shen H, Li X, Pan S. Deregulated expression of miR-196a and miR-196b in gastric cancer. Mol Carcinog. 2011;50(3):145–54. [Google Scholar]
- 24.Sun Y, Liu FZ, Liang X, Sun CC. miR-100 inhibits the proliferation and metastasis of oral squamous cell carcinoma by targeting mTOR. Biochem Biophys Res Commun. 2013;431(3):705–10. [Google Scholar]
- 25.Li N, Fu H, Tie Y, Hu Z, Kong W, Wu Y, Zheng X. miR-34a inhibits migration and invasion by down-regulation of c-Met. Oral Oncol. 2010;46(7):526–31. [DOI] [PubMed] [Google Scholar]
- 26.Park SM, Gaur AB, Lengyel E, Peter ME. The miR-200 family determines the epithelial phenotype of cancer cells by targeting the E-cadherin repressors ZEB1 and ZEB2. Genes Dev. 2008;22(7):894–907. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Siow ZR, De Boer B, Tilley WD. Should miRNA expression data be standardized? BMC Bioinformatics. 2014;15(1):387.25495206 [Google Scholar]
- 28.Bhaumik D, Scott GK, Schokrpur S, Patil CK, Campisi J, Benz CC, Laderoute KR. Expression of microRNA-146 suppresses NF-κB activity with reduction of metastatic potential in breast cancer cells. Oncogene. 2008;27(42):5643–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Xia H, Sun S, Wang B, Wang T, Liang C, Li G. miR-124 inhibits cell proliferation in gastric cancer through down-regulation of SPHK1. J Pathol. 2012;227(4):470–80. [DOI] [PubMed] [Google Scholar]
- 30.Li H, Li J, Yu B, Shi Y. MicroRNA-204 regulates epithelial-mesenchymal transition by targeting SLUG in oral squamous cell carcinoma. Biochim Biophys Acta. 2011;1816(1):50–7.21447370 [Google Scholar]
- 31.Hayashita, Y., Osada, H., Tatematsu, Y., Yamada, H., Yanagisawa, K., Tomida, S., Takahashi, T. (2005). A polycistronic microRNA cluster, miR-17-92, is overexpressed in human lung cancers and enhances cell proliferation. Cancer Res, 65(21), 9628–9632. [DOI] [PubMed]
- 32.Liu X, Chen Z, Yu J, Xia J, Zhou X. MicroRNA profiling and head and neck cancer. Comp Funct Genomics. 2009;2009:837514. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Gregory, P. A., Bert, A. G., Paterson, E. L., Barry, S. C., Tsykin, A., Farshid, G., Goodall, G. J. The miR-200 family and miR-205 regulate epithelial to mesenchymal transition by targeting ZEB1 and SIP1. Nat Cell Biol. 2008;10(5), 593–601. [DOI] [PubMed]
- 34.Zhang X, Xu J, Jiang T, Liu G. (2012). MicroRNA-23b targets the MET oncogene to inhibit cell proliferation and invasion in glioma. PLoS ONE, 7(7), e41502.
- 35.Wu W, Sun M, Zou GM, Chen J. MicroRNA and cancer: current status and prospective. Int J Cancer. 2011;130(6):1236–45. [DOI] [PubMed] [Google Scholar]
- 36.Sonkoly E, Loven J, Xu N, Meisgen F, Wei T, Brodin P, Stahle M. MicroRNA-203 functions as a tumor suppressor in basal cell carcinoma. Oncogenesis. 2010;2(1):e1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Jiang L, Liu X, Chen Z, Jin Y, Heidbreder CE, Kolokythas A, Zhou X. MicroRNA-7 targets IGF1R (insulin-like growth factor 1 receptor) in tongue squamous cell carcinoma cells. Biochem J. 2010;432(1):199–205. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Kota, J., Chivukula, R. R., O’Donnell, K. A., Wentzel, E. A., Montgomery, C. L., Hwang,H. W., Mendell, J. T. Therapeutic microRNA delivery suppresses tumorigenesis in a murine liver cancer model. Cell. 2009;137(6), 1005–1017. [DOI] [PMC free article] [PubMed]
- 39.Gebeshuber CA, Zatloukal K, Martinez J. miR-29a suppresses tristetraprolin, which is a regulator of epithelial polarity and metastasis. EMBO Rep. 2009;10(4):400–5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Shi B, Sepp-Lorenzino L, Prisco M, Linsley P, deAngelis T. MicroRNA 125b targets the 3’ UTR of multiple pro-apoptotic genes in ovarian cancer cells. Biochem Biophys Res Commun. 2011;413(1):3–6. [Google Scholar]
- 41.Ratti M, Lampis A, Ghidini M, Salati M, Mirchev MB, Valeri N, Hahne JC. MicroRNAs (miRNAs) and long non-coding RNAs (lncRNAs) as New Tools for Cancer Therapy: first steps from Bench to Bedside. Target Oncol. 2020;15(3):261–78. PMID: 32451752; PMCID: PMC7283209. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Ying SY, Chang DC, Lin SL. The microRNA (miRNA): overview of the RNA genes that modulate gene function. Mol Biotechnol. 2008;38(3):257 – 68. doi: 10.1007/s12033-007-9013-8. Epub 2007 Nov 13. PMID: 17999201; PMCID: PMC7091389. [DOI] [PMC free article] [PubMed]
- 43.Wang G, Wang JJ, Tang HM, To SS. Targeting strategies on miRNA-21 and PDCD4 for glioblastoma. Arch Biochem Biophys. 2015;580:64–74. 10.1016/j.abb.2015.07.001. Epub 2015 Jul 2. PMID: 26142886. [DOI] [PubMed] [Google Scholar]
- 44.Zhang J, Ma L. MicroRNA control of epithelial-mesenchymal transition and metastasis. Cancer Metastasis Rev. 2012;31(3–4):653–62. 10.1007/s10555-012-9368-6. PMID: 22684369; PMCID: PMC3686549. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Palaia G, Pippi R, Rocchetti F, Caputo M, Macali F, Mohsen A, Del Vecchio A, Tenore G, Romeo U. Liquid biopsy in the assessment of microRNAs in oral squamous cell carcinoma: a systematic review. J Clin Exp Dent. 2022;14(10):e875–84. 10.4317/jced.59736. PMID: 36320672; PMCID: PMC9617270. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Gambari R, Brognara E, Spandidos DA, Fabbri E. Targeting oncomiRNAs and mimicking tumor suppressor miRNAs: Νew trends in the development of miRNA therapeutic strategies in oncology (review). Int J Oncol. 2016;49(1):5–32. 10.3892/ijo.2016.3503. Epub 2016May 4. PMID: 27175518; PMCID: PMC4902075. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Si W, Shen J, Zheng H, Fan W. The role and mechanisms of action of microRNAs in cancer drug resistance. Clin Epigenetics. 2019;11(1):25. 10.1186/s13148-018-0587-8. PMID: 30744689; PMCID: PMC6371621. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Manikandan M, Deva Magendhra Rao AK, Arunkumar G, Manickavasagam M, Rajkumar KS, Rajaraman R, Munirajan AK. Oral squamous cell carcinoma: microRNA expression profiling and integrative analyses for elucidation of tumourigenesis mechanism. Mol Cancer. 2016;15:28. 10.1186/s12943-016-0512-8. PMID: 27056547; PMCID: PMC4823852. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Rapado-González Ó, Majem B, Muinelo-Romay L, Álvarez-Castro A, Santamaría A, Gil-Moreno A, López-López R, Suárez-Cunqueiro MM. Human salivary microRNAs in Cancer. J Cancer. 2018;9(4):638–49. 10.7150/jca.21180. PMID: 29556321; PMCID: PMC5858485. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Park NJ, Zhou H, Elashoff D, Henson BS, Kastratovic DA, Abemayor E, Wong DT. Salivary microRNA: discovery, characterization, and clinical utility for oral cancer detection. Clin Cancer Res. 2009;15(17):5473–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Matse JH, Yoshizawa J, Wang X, Elashoff D, Bolscher JG, Veerman EC, Wong DT. Discovery and prevalidation of salivary extracellular microRNA biomarkers panel for the non-invasive detection of benign and malignant parotid gland tumors. Clin Cancer Res. 2013;19(11):3032–8. [DOI] [PubMed] [Google Scholar]
- 52.Wong TS, Liu XB, Wong BY, Ng RW, Yuen AP, Wei WI. Mature miR-184 as potential oncogenic microRNA of squamous cell carcinoma of tongue. Clin Cancer Res. 2008;14(9):2588–92. [DOI] [PubMed] [Google Scholar]
- 53.He Q, Chen J, Lin L, Shao X, Li S, Zheng H, Pan Q. Differential expression of miRNAs in the saliva of patients with oral lichen planus and oral squamous cell carcinoma. Oncol Lett. 2014;8(4):1403–7.25202342 [Google Scholar]
- 54.Liu CJ, Tsai MM, Hung PS, Kao SY, Liu TY, Wu KY, Lin SC. miR-31 ablates expression of the HIF regulatory factor FIH to activate the HIF pathway in head and neck carcinoma. Cancer Res. 2012;70(4):1635–44. [DOI] [PubMed] [Google Scholar]
- 55.Zahran F, Ghalwash D, Shaker O, Al-Johani K, Scully C. Salivary microRNAs in oral cancer. Oral Dis. 2015;21(6):739–47. 10.1111/odi.12340. Epub 2015 Apr 22. PMID: 25784212. [DOI] [PubMed] [Google Scholar]
- 56.Bahn JH, Zhang Q, Li F, Chan TM, Lin X, Kim Y, Xiao X. The landscape of microRNA, Piwi-interacting RNA, and circular RNA in human saliva. Clin Chem. 2015;61(1):221–30. 10.1373/clinchem.2014.230433. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Lee YH, Wong DT. Saliva: an emerging biofluid for early detection of diseases. Am J Dent. 2009;22(4):241–8. [PMC free article] [PubMed] [Google Scholar]
- 58.Spielmann N, Wong DT. Saliva: diagnostics and therapeutic perspectives. Oral Dis. 2011;17(4):345–54. 10.1111/j.1601-0825.2010.01773.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Kinoshita T, Yip KW, Spence T, Liu FF. MicroRNAs in extracellular vesicles: potential cancer biomarkers. J Hum Genet. 2017;62(1):67–74. 10.1038/jhg.2016.87. Epub 2016 Jul 7. PMID: 27383658. [DOI] [PubMed] [Google Scholar]
- 60.Lin X, Wu W, Ying Y, Luo J, Xu X, Zheng L, Wu W, Yang S, Zhao S. MicroRNA-31: a pivotal oncogenic factor in oral squamous cell carcinoma. Cell Death Discov. 2022;8(1):140. 10.1038/s41420-022-00948-z. PMID: 35351880; PMCID: PMC8964740. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Balakittnen J, Ekanayake Weeramange C, Wallace DF, Duijf PHG, Cristino AS, Hartel G, Barrero RA, Taheri T, Kenny L, Vasani S, Batstone M, Breik O, Punyadeera C. A novel saliva-based miRNA profile to diagnose and predict oral cancer. Int J Oral Sci. 2024;16(1):14. 10.1038/s41368-023-00273-w. PMID: 38368395; PMCID: PMC10874410. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Eslami M, Khazeni S, Khanaghah XM, Asadi MH, Ansari MA, Garjan JH, Lotfalizadeh MH, Bayat M, Taghizadieh M, Taghavi SP, Hamblin MR, Nahand JS. MiRNA-related metastasis in oral cancer: moving and shaking. Cancer Cell Int. 2023;23(1):182. 10.1186/s12935-023-03022-5. PMID: 37635248; PMCID: PMC10463971. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Thomas P, Preethi KA, Selvakumar SC, Ramani P, Sekar D. Relevance of micro-RNAs and their targets as a diagnostic and prognostic marker in oral squamous cell carcinoma. J Oral Maxillofac Pathol. 2023 Apr-Jun;27(2):364–73. 10.4103/jomfp.jomfp_349_22. Epub 2023 Jul 13. PMID: 37854932; PMCID: PMC10581285. [DOI] [PMC free article] [PubMed]
- 64.Díaz-López A, Díaz-Martín J, Moreno-Bueno G, Cuevas EP, Santos V, Olmeda D, Portillo F, Palacios J, Cano A. Zeb1 and Snail1 engage miR-200f transcriptional and epigenetic regulation during EMT. Int J Cancer. 2015;136(4): E62-73. 10.1002/ijc.29177. Epub 2014 Sep 12. PMID: 25178837. [DOI] [PubMed]
- 65.Jiang Q, Cao Y, Qiu Y, Li C, Liu L, Xu G. Progression of squamous cell carcinoma is regulated by miR-139-5p/CXCR4. Front Biosci (Landmark Ed). 2020;25(9):1732–1745. 10.2741/4875. PMID: 32472755. [DOI] [PubMed]
- 66.Tajik F, Alian F, Yousefi M, Azadfallah A, Hoseini A, Mohammadi F, Karimi-Dehkordi M, Alizadeh-Fanalou S. MicroRNA-372 acts as a double-edged sword in human cancers. Heliyon. 2023;9(5):e15991. 10.1016/j.heliyon.2023.e15991. PMID: 37251909; PMCID: PMC10208947. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Ma L, Young J, Prabhala H, Pan E, Mestdagh P, Muth D, Teruya-Feldstein J, Reinhardt F, Onder TT, Valastyan S, Westermann F, Speleman F, Vandesompele J, Weinberg RA. miR-9, a MYC/MYCN-activated microRNA, regulates E-cadherin and cancer metastasis. Nat Cell Biol. 2010;12(3):247–56. Epub 2010 Feb 21. PMID: 20173740; PMCID: PMC2845545. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Zhu X, Ma SP, Yang D, Liu Y, Wang YP, Lin T, Li YX, Yang SH, Zhang WC, Wang XL. Mir-142-3p suppresses cell growth by targeting CDK4 in Colorectal Cancer. Cell Physiol Biochem. 2018;51(4):1969–81. Epub 2018 Dec 4. PMID: 30513513. [DOI] [PubMed] [Google Scholar]
- 69.Wang Z, Guan W, Ma Y, Zhou X, Song G, Wei J, Wang C. MicroRNA-191 regulates oral squamous cell carcinoma cells growth by targeting PLCD1 via the Wnt/β-catenin signaling pathway. BMC Cancer. 2023;23(1):668. 10.1186/s12885-023-11113-9. PMID: 37460940; PMCID: PMC10351167. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Cristaldi M, Mauceri R, Di Fede O, Giuliana G, Campisi G, Panzarella V. Salivary biomarkers for oral squamous cell carcinoma diagnosis and Follow-Up: current status and perspectives. Front Physiol. 2019;10:1476. 10.3389/fphys.2019.01476. PMID: 31920689; PMCID: PMC6914830. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Faur CI, Rotaru H, Osan C, Jurj A, Roman RC, Moldovan M, Chirila M, Hedesiu M. Salivary exosomal microRNAs as biomarkers for head and neck cancer detection-a literature review. Maxillofac Plast Reconstr Surg. 2021;43(1):19. 10.1186/s40902-021-00303-9. PMID: 34191144; PMCID: PMC8245637. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Coenen-Stass AML, Pauwels MJ, Hanson B, Martin Perez C, Conceição M, Wood MJA, Mäger I, Roberts TC. Extracellular microRNAs exhibit sequence-dependent stability and cellular release kinetics. RNA Biol. 2019;16(5):696–706. Epub 2019 Mar 5. PMID: 30836828; PMCID: PMC6546368. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Xie ZJ, Chen G, Zhang XC, Li DF, Huang J, Li ZJ. Saliva supernatant miR-21: a novel potential biomarker for esophageal cancer detection. Asian Pac J Cancer Prev. 2012;13(12):6145-9. 10.7314/apjcp.2012.13.12.6145. PMID: 23464420. [DOI] [PubMed]
- 74.Gao S, Chen LY, Wang P, Liu LM, Chen Z. MicroRNA expression in salivary supernatant of patients with pancreatic cancer and its relationship with ZHENG. Biomed Res Int. 2014;2014:756347. 10.1155/2014/756347. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Yang Y, Ma L, Qiao X, Zhang X, Dong SF, Wu MT, Zhai K, Shi HZ. Salivary microRNAs show potential as biomarkers for early diagnosis of malignant pleural effusion. Transl Lung Cancer Res. 2020;9(4):1247–57. 10.21037/tlcr-19-530. PMID: 32953502; PMCID: PMC7481620. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Cherchi R, Cusano R, Orrù S, Ferrari PA, Massidda M, Fotia G, De Matteis S, Cocco P. Next generation sequencing for miRNA detection on the exhaled breath condensate: a pilot study. Epigenet Insights. 2023;16:25168657231160985. 10.1177/25168657231160985. PMID: 37025420; PMCID: PMC10070752. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Maharaj PD, Widen SG, Huang J, Wood TG, Thangamani S. Discovery of mosquito saliva microRNAs during CHIKV infection. PLoS Negl Trop Dis. 2015;9(1):e0003386. 10.1371/journal.pntd.0003386. PMID: 25612225; PMCID: PMC4303268. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Ekanayake Weeramange C, Tang KD, Barrero RA, Hartel G, Liu Z, Ladwa R, Langton-Lockton J, Frazer I, Kenny L, Vasani S, Punyadeera C. Salivary micro RNAs as biomarkers for oropharyngeal cancer. Cancer Med. 2023;12(14):15128–40. 10.1002/cam4.6185. Epub 2023 Jun 6. PMID: 37278132; PMCID: PMC10417169. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Michael A, Bajracharya SD, Yuen PS, Zhou H, Star RA, Illei GG, Alevizos I. Exosomes from human saliva as a source of microRNA biomarkers. Oral Dis. 2010;16(1):34–8. 10.1111/j.1601-0825.2009.01604.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Bautista-Sánchez D, Arriaga-Canon C, Pedroza-Torres A, et al. The promising role of miR-21 as a Cancer Biomarker and its importance in RNA-Based therapeutics. Mol Ther Nucleic Acids. 2020;20:409–20. 10.1016/j.omtn.2020.03.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Stepicheva NA, Song JL. Function and regulation of microRNA-31 in development and disease. Mol Reprod Dev. 2016;83(8):654–74. 10.1002/mrd.22678. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Kumar MA, Baba SK, Sadida HQ et al. Extracellular vesicles as tools and targets in therapy for diseases. Signal Transduct Target Ther. 2024;9(1):27. Published 2024 Feb 5. 10.1038/s41392-024-01735-1 [DOI] [PMC free article] [PubMed]
- 83.Wang J, Jing J, Zhou C, Fan Y. Emerging roles of exosomes in oral diseases progression. Int J Oral Sci. 2024;16(1):4. 10.1038/s41368-023-00274-9. PMID: 38221571; PMCID: PMC10788352. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Dumache R. Early Diagnosis of Oral Squamous Cell Carcinoma by Salivary microRNAs. Clin Lab. 2017;63(11):1771–1776. 10.7754/Clin.Lab.2017.170607. PMID: 29226639. [DOI] [PubMed]
- 85.Vageli DP, Doukas PG, Shah R, Boyi T, Liu C, Judson BL. A novel saliva and serum miRNA panel as a potential useful index for oral Cancer and the Association of miR-21 with smoking history: a pilot study. Cancer Prev Res (Phila). 2023;16(12):653–9. 10.1158/1940-6207.CAPR-23-0219. [DOI] [PubMed] [Google Scholar]
- 86.Garg A, Urs AB, Koner BC, Augustine J, Guru SA. Evaluation of diagnostic significance of salivary miRNA-184 and miRNA-21 in oral squamous cell carcinoma and oral potentially malignant disorders. Head Neck Pathol. 2023;17(4):961–8. 10.1007/s12105-023-01600-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Tarrad NAF, Hassan S, Shaker OG, AbdelKawy M. Salivary LINC00657 and miRNA-106a as diagnostic biomarkers for oral squamous cell carcinoma, an observational diagnostic study. BMC Oral Health. 2023;23(1):994. Published 2023 Dec 12. 10.1186/s12903-023-03726-0 [DOI] [PMC free article] [PubMed]
- 88.Safari Z, Firouzi A, Rezaeikalantari N, et al. The salivary exosomal microRNA as a potential biomarker in patients with periodontitis and oral cancers. Chem Biol Drug Des. 2023;101(5):1204–15. 10.1111/cbdd.14159. [DOI] [PubMed] [Google Scholar]
- 89.Koh, W., Sheng, C. T., Tan, B., Lee, Q. Y., Kuznetsov, V. A., Kiang, L. S., Chia,K. S. (2014). Analysis of extracellular microRNAs in saliva and other biofluids for cancer diagnostics. Nature Protocols, 9(9), 2111–2128. doi: 10.1038/nprot.2014.143.
- 90.Yoshizawa JM, Schafer CA, Schafer JJ, Farrell JJ, Paster BJ, Wong DT. Salivary biomarkers: toward future clinical and diagnostic utilities. Clin Microbiol Rev. 2013;26(4):781–91. 10.1128/CMR.00021-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Chiabotto G, Ceccotti E, Tapparo M, Camussi G, Bruno S. Human liver stem cell-derived extracellular vesicles target hepatic stellate cells and attenuate their pro-fibrotic phenotype. Front Cell Dev Biol. 2021;9:777462. 10.3389/fcell.2021.777462. PMID: 34796180; PMCID: PMC8593217. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Romani P, Valcarcel-Jimenez L, Frezza C, Dupont S. Crosstalk between mechanotransduction and metabolism. Nat Rev Mol Cell Biol. 2021;22(1):22–38. 10.1038/s41580-020-00306-w. [DOI] [PubMed] [Google Scholar]
- 93.Bugshan A, Farooq I. Oral squamous cell carcinoma: metastasis, potentially associated malignant disorders, etiology and recent advancements in diagnosis. F1000Res. 2020;9:229. 10.12688/f1000research.22941.1. PMID: 32399208; PMCID: PMC7194458. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Mehterov N, Vladimirov B, Sacconi A, Pulito C, Rucinski M, Blandino G, Sarafian V. Salivary miR-30c-5p as potential biomarker for detection of oral squamous cell carcinoma. Biomedicines. 2021;9(9):1079. 10.3390/biomedicines9091079. PMID: 34572265; PMCID: PMC8465705. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Bastías D, Maturana A, Marín C, Martínez R, Niklander SE. Salivary biomarkers for oral Cancer detection: an exploratory systematic review. Int J Mol Sci. 2024;25(5):2634. 10.3390/ijms25052634. Published 2024 Feb 23. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Sinha A, Bhattacharjee R, Bhattacharya B, et al. The paradigm of miRNA and siRNA influence in oral-biome. Biomed Pharmacother. 2023;159:114269. 10.1016/j.biopha.2023.114269. [DOI] [PubMed] [Google Scholar]
- 97.Deutsch FT, Khoury SJ, Sunwoo JB, Elliott MS, Tran NT. Application of salivary non-coding microRNAs for the diagnosis of oral cancers. Head Neck. 2020;42(10):3072–83. 10.1002/hed.26348. [DOI] [PubMed] [Google Scholar]
- 98.DeLucas M, Sánchez J, Palou A, Serra F. The impact of Diet on miRNA Regulation and its implications for health: a systematic review. Nutrients. 2024;16(6):770. 10.3390/nu16060770. Published 2024 Mar 7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99.Bailey RL. Overview of dietary assessment methods for measuring intakes of foods, beverages, and dietary supplements in research studies. Curr Opin Biotechnol. 2021;70:91–6. 10.1016/j.copbio.2021.02.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.Anderson AC, Al-Ahmad A, Schlueter N, Frese C, Hellwig E, Binder N. Influence of the long-term use of oral hygiene products containing stannous ions on the salivary microbiome - a randomized controlled trial. Sci Rep. 2020;10(1):9546. Published 2020 Jun 12. 10.1038/s41598-020-66412-z [DOI] [PMC free article] [PubMed]
- 101.Bhattarai KR, Kim HR, Chae HJ. Compliance with Saliva Collection Protocol in healthy volunteers: strategies for managing risk and errors. Int J Med Sci. 2018;15(8):823–31. 10.7150/ijms.25146. Published 2018 May 22. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102.Romani C, Baronchelli M, Assoni C, et al. Stability of circulating miRNA in saliva: the influence of sample associated pre-analytical variables. Clin Chim Acta. 2024;553:117702. 10.1016/j.cca.2023.117702. [DOI] [PubMed] [Google Scholar]
- 103.Tan Y, Wang Z, Xu M, et al., et al. Oral squamous cell carcinomas: state of the field and emerging directions. Int J Oral Sci. 2023;15(1):44. 10.1038/s41368-023-00249-w. Published 2023 Sep 22. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104.McGraw D, Mandl KD. Privacy protections to encourage use of health-relevant digital data in a learning health system. NPJ Digit Med. 2021;4(1):2. 10.1038/s41746-020-00362-8. Published 2021 Jan 4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105.Koturbash I, Tolleson WH, Guo L, et al. microRNAs as pharmacogenomic biomarkers for drug efficacy and drug safety assessment. Biomark Med. 2015;9(11):1153–76. 10.2217/bmm.15.89. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 106.Goldsack JC, Coravos A, Bakker JP et al. Verification, analytical validation, and clinical validation (V3): the foundation of determining fit-for-purpose for Biometric Monitoring Technologies (BioMeTs). NPJ Digit Med. 2020;3:55. Published 2020 Apr 14. 10.1038/s41746-020-0260-4 [DOI] [PMC free article] [PubMed]
- 107.Indian Council of Medical Research. (2017). National ethical guidelines for biomedical and health research involving human participants. https://ethics.ncdirindia.org/asset/pdf/ICMR_National_Ethical_Guidelines.pdf [DOI] [PMC free article] [PubMed]
- 108.Mestry C, Ashavaid TF, Shah SA. Key methodological challenges in detecting circulating miRNAs in different biofluids. Ann Clin Biochem. 2023;60(1):14–26. 10.1177/00045632221129778. [DOI] [PubMed] [Google Scholar]
- 109.Sekaran S. Corrigendum to salivary microRNA- 375: A novel biomarker in the malignant transformation of oral potentially malignant disorders [Oral Oncol. 134 (2022) 106065]. Oral Oncol. 2023;138:106326. 10.1016/j.oraloncology.2023.106326. Epub 2023 Jan 28. PMID: 36716561. [DOI] [PubMed]
- 110.Shen Z, Silva RM. MicroRNAs: emerging players in apical periodontitis. J Appl Oral Sci. 2021;29:e20201058. 10.1590/1678-7757-2020-1058. PMID: 33886945; PMCID: PMC8054647. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111.Thomaidou AC, Batsaki P, Adamaki M, Goulielmaki M, Baxevanis CN, Zoumpourlis V, Fortis SP. Promising biomarkers in Head and Neck Cancer: the most clinically important miRNAs. Int J Mol Sci. 2022;23(15):8257. 10.3390/ijms23158257. PMID: 35897831; PMCID: PMC9367895. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 112.Kalfert D, Pesta M, Kulda V, Topolcan O, Ryska A, Celakovsky P, et al. MicroRNA profile in site-specific head and neck squamous cell cancer. Anticancer Res. 2015;35:2455–63. [PubMed] [Google Scholar]
- 113.Marabita F, de Candia P, Torri A, Tegnér J, Abrignani S, Rossi RL. Brief Bioinform. 2016;17(2):204–12. 10.1093/bib/bbv056. Epub 2015 Aug 3. PMID: 26238539; PMCID: PMC4793896. Normalization of circulating microRNA expression data obtained by quantitative real-time RT-PCR. [DOI] [PMC free article] [PubMed]
- 114.Pamedytyte D, Leipute E, Zilaitiene B, Sarauskas V, Dauksiene D, Dauksa A, Zvirbliene A. Different stability of miRNAs and endogenous control genes in archival specimens of papillary thyroid carcinoma. Mol Med. 2020;26(1):100. 10.1186/s10020-020-00218-7. PMID: 33153429; PMCID: PMC7643475. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 115.Kumar P, Gupta S, Das BC. Saliva as a potential non-invasive liquid biopsy for early and easy diagnosis/prognosis of head and neck cancer. Transl Oncol. 2024;40:101827. 10.1016/j.tranon.2023.101827. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 116.Ghosh S, Rajendran RL, Mahajan AA, Chowdhury A, Bera A, Guha S, Chakraborty K, Chowdhury R, Paul A, Jha S, Dey A, Dubey A, Gorai S, Das P, Hong CM, Krishnan A, Gangadaran P, Ahn BC. Harnessing exosomes as cancer biomarkers in clinical oncology. Cancer Cell Int. 2024;24(1):278. 10.1186/s12935-024-03464-5. PMID: 39113040; PMCID: PMC11308730. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Citations
- Marabita F, de Candia P, Torri A, Tegnér J, Abrignani S, Rossi RL. Brief Bioinform. 2016;17(2):204–12. 10.1093/bib/bbv056. Epub 2015 Aug 3. PMID: 26238539; PMCID: PMC4793896. Normalization of circulating microRNA expression data obtained by quantitative real-time RT-PCR. [DOI] [PMC free article] [PubMed]
Data Availability Statement
No datasets were generated or analysed during the current study.