Abstract
Helicobacter pylori infection is a major risk factor for gastric adenocarcinomas. In the case of the intestinal subtype, chronic gastritis and intestinal metaplasia are well-known sequential steps in carcinogenesis. H. pylori has high genetic diversity that can modulate virulence and pathogenicity in the human host as a cag Pathogenicity Island (cagPAI). However, bacterial gene combinations do not always explain the clinical presentation of the disease, indicating that other factors associated with H. pylori may play a role in the development of gastric disease. In this context, we characterized the microbial composition of patients with chronic gastritis (inactive and active), intestinal metaplasia, and gastric cancer as well as their potential association with H. pylori. To this end, 16 S rRNA metagenomic analysis was performed on gastric mucosa samples from patients with different types of lesions and normal gastric tissues. Our main finding was that H. pylori virulence status can contribute to significant differences in the constitution of the gastric microbiota between the sequential steps of the carcinogenesis cascade. Differential microbiota was observed in inactive and active gastritis dependent of the H. pylori presence and status (p = 0.000575). Pseudomonades, the most abundant order in the gastritis, was associated the presence of non-virulent H. pylori in the active gastritis. Notably, there are indicator genera according to H. pylori status that are poorly associated with diseases and provide additional evidence that the microbiota, in addition to H. pylori, is relevant to gastric carcinogenesis.
Keywords: Helicobacter pylori, Gastric cancer, Metagenomics
Subject terms: Gastrointestinal cancer, Gastroenterology, Gastrointestinal diseases
Introduction
Gastric cancer (GC) is a global public health problem owing to its high incidence and mortality rates. It is the fifth most frequently diagnosed cancer and the fourth leading cause of cancer-related death according to the Globocan estimative1. Approximately 95% of gastric tumors are adenocarcinomas that can histologically be classified as diffuse or intestinal type2.
Helicobacter pylori infection is the major risk factor for gastric adenocarcinomas in which, chronic active gastritis and intestinal metaplasia are part of bacterium-related intestinal subtype3. Notably, H. pylori has co-evolved with humans up to approximately l00 years ago4,5, indicating a bacterium-host equilibrium, and this symbiotic relationship must be altered for the GC development. H. pylori has high genetic diversity due to rearrangements and horizontal gene transfer, which reflect historical human demographic events. Among the horizontally acquired gene regions, cag pathogenicity island (cagPAI), a cluster of genes encoding the Type IV secretion system, contains a virulent factor that mediates colonization and pathogenicity in the human host6,7. However, the island is not a uniformly conserved entity, and the correlation between different cagPAI rearrangements and clinical presentation is not always found8,9, indicating that other factors, in association with H. pylori, play a role in the development of gastric disease.
Currently, research using 16 S rRNA sequence analysis, and more recently, high-throughput sequencing technology, has revealed that the stomach harbors a distinct and complex ecosystem in addition to H. pylori. Under physiological conditions, the microbiota plays an essential role in human health, and alterations in the microbiome have been hypothesized to be linked to gastric disease development, as they can stimulate the inflammatory process10,11.
Previous studies have investigated the characteristics of gastric microbiota in different gastric diseases; however, few studies have addressed bacterial genotypes, which are relevant factors for GC development. In addition, considering the multifactorial nature of gastric lesions, diverse populations with different habits should be studied to clarify this association. To address this issue, we characterized the microbial composition of patients with chronic gastritis, intestinal metaplasia, and GC. Given the role of reactive oxygen species derived from neutrophils, which are characteristic of ACG and linked to DNA damage, the gastritis was categorized as inactive (ICG) or active (ACG), In addition, we explored potential associations with H. pylori strains, considering the cagPAI genes, cagA and cagE, which have not yet been considered in gastric microbiota studies.
Patients and methods
Sixty-six patients were included in this study after selection based on histological classification, gastric region, and H. pylori genotype. Fifteen intestinal GC samples were selected from 172 patients who underwent gastrectomy after GC diagnosis at the Hospital Universitario Walter Cantídeo (HUWC) and Santa Casa de Misericordia Hospital in Fortaleza, Ceará, Brazil. Forty-eight non-cancerous gastric lesions (NGL), including 20 intestinal metaplasia, 12 active chronic gastritis, 16 inactive chronic gastritis, and 3 normal gastric tissues, were selected from 202 patients with dyspeptic symptoms who underwent endoscopy at the HUWC and Hospital Geral de Fortaleza, Ceará, Brazil. All patients were from a non-cardiac region. Exclusion criteria for NGL were the presence of duodenal gastric or esophageal masses, Barrett’s esophagus lesions, previous gastric or duodenal surgery, treatment with any anti-inflammatory, proton-pump inhibitor drugs, or antibiotics during the last three months and previous therapy for H. pylori. The histopathological diagnosis of GC was made using Lauren12 and NGL according to the update Sydney system13. ICG was defined as diffuse lymphocytic inflammatory infiltration or organization in the follicular/nodular structures. ACG was defined by the presence of neutrophilic infiltrates permeating the glands and/or lamina propria. Normal cases included patients who underwent endoscopy with negative results and were H. pylori negative. Table 1 presents the demographic data of the study participants grouped according to the type of gastric lesion.
Table 1.
Demographic characteristics of study participants, distributed by type of gastric lesion.
| Variables | Groups | |||
|---|---|---|---|---|
| Control | Gastritis | Metaplasia | Cancer | |
| Age (years) | 34.33 ± 11.85 | 55.2 ± 17.74 | 63.7 ± 18.9 | 65.3 ± 15.40 |
| Sex | ||||
| Female | 2 (33.3) | 15 (53.6) | 14 (70) | 5 (33.3) |
| Male | 1 (66.7) | 13 (46.4) | 6 (30) | 10 (66.7) |
DNA extraction, H. pyloridetection and genotyping
Metagenomic DNA was extracted from frozen tumor tissue samples consisting mainly of tumor cells (> 80%) and single frozen stomach fragments in which the NGL was representative, using the cetyltrimethylammonium bromide method adapted from Foster and Twell14. Concentration and quality of metagenomic DNA were determinate with NanoDrop ND-1000®.
H. pylori infection was detected using amplification of the ureC gene as described by Lage et al.15. cagPAI genes (cagA and cagE) were identified using specific primers and conditions described by Lima et al.16. Negative (water) and positive controls were assayed in each run. The PCR products were visualized using 1% agarose gel electrophoresis with ethidium bromide.
16SrRNA amplicon for library preparation
For 16 S rRNA gene amplification, DNA from all samples selected for this study was purified using DNA clean-up kit (Qiagen, Germany). The amplicon library of the V4 variable region of the 16 S rRNA gene was prepared according to the protocol suggested by Illumina (San Diego, CA, USA), using primers 515 F/806R.17 PCR mixtures (25 µL) were composed of Taq HotStart HiFi buffer plus, 1U of Taq HotStar High Fidelity (Kapa Biosystems, Boston, MA, EUA), 0.75mM dNTPs, and 0.75mM of each primer and 30 ng of DNA template of each sample. PCR amplification was carried out under the following conditions: 94°C for 4 min of initial heating followed 25 cycles of 94°C for 45 s, 60°C for 60 s and 72°C for 60 s, after which a final elongation step at 72°C for 10 min. Each sample was put in triplet with one negative control. The success of the amplification, with expected bands around 250 bp, was assessed on a 1% agarose gel.
Library preparation was done using Illumina Nextera XT (Illumina, San Diego, CA, USA), according to the manufacturer’s instructions. Briefly, 5 µL of the each confirmed amplified sample was submitted to a second reaction in a 50 µL PCR mixture using 1x Taq HotStart HiFi buffer, 1U of Taq DNA polimerase HotStart High fidelity, (Kapa Biosystems, Boston, MA, EUA), 0.75mM dNTPs and 5 µL of each Illumina Nextera XT index. PCR conditions consisted of initial heating of 94°C for 4 min followed 8 cycles of 94°C for 30 s, 55°C for 30 s and 72°C for 30 s and final elongation steps at 72°C for 5 min.
The PCR products were purified using Agencourt AMPure XP magnetic beads (Beckman Coulter, Brea, CA, USA) according to the manufacturer’s instructions. Amplicons from each sample were pooled and quantified by spectrophotometry using a dsDNA BR Assay Kit (Invitrogen). DNA libraries were multiplexed using equal amount of the pooled amplicon, diluted to a final concentration of 8.0 pM and it was spiked with the PhiX sequencing positive control (Illumina), in a final ratio of 80:20. Sequencing was performed on an Illumina MiSeq platform (Illumina, San Diego, CA, USA) in a 2 × 300 paired-end configuration.
Sequencing data processing
After sequencing, Illumina adapter sequences were trimmed from already-demultiplexed raw fastq files using Cutadapt v1.818 in paired-end mode, and the reads quality was assessed using FastQC v.0.11.819 and vsearch v2.10.420. Subsequent analyses were performed in the R v3.5.3 environment21, following the DADA2 v1.11.1 package22 pipeline for obtaining a table of non-chimeric amplicon sequence variants (ASVs) free of low-quality and non-prokaryotic sequences (sequences differing by as little as one nucleotide)23. Taxonomy assignment and removal of non-prokaryotic sequences was performed against the SILVA reference database (release 132)24.
Data analyses
Observed, Chao1, Shannon and Simpson were calculated for alpha diversity evaluation. Alpha diversity metrics summarize the structure of an ecological community with respect to its richness (number of taxonomic groups) and evenness (distribution of abundances of the groups). Shapiro-Wilk test was used to assess whether the data set was normally distributed. As Shannon diversity was parametric, one-way analysis of variance and Tukey’s honest significant difference (HSD) post-hoc tests were used for multiple comparisons of means at a 95% confidence interval. For Chao1 and observed ASVs, we used the Kruskal–Wallis non-parametric test25.
To analyze beta diversity, microbial communities were compared using distance-based comparisons and a comparison of composition profiles related to each of the studied lesions. For the distance-based approach, the relative abundances of ASVs were square-root transformed, and the Bray-Curtis similarity coefficient was calculated for each sample pair. The similarity matrix was visualized using nonmetric multidimensional scaling. Indicator analysis was performed using the package indicspecies26 to identify the main genera associated with each type of lesion, clinical status, and virulence.
The relationship between clinical variables and the bacterial community was evaluated through canonical correspondence analysis (CCA), using the Vegan package26 in the R environment21. The ‘decorana’ function was used to choose the best ordering model, which indicated the use of CCA. Preliminary CCA was performed using all available variables (age, lesion type, cagA, cagE, virb11, and other genotypic markers). The variables that demonstrated significant influence (p < 0.05) were selected using the ‘ordistep’ function and confirmed using the ‘anova.cca’ function for the final CCA. The inflation factor of the variation (function ‘vif.cca’) showed that the retained explanatory variables did not present collinearity among themselves (FIV < 10).
In addition, the samples were classified into 11 groups, considering gastric lesions and H. pylori genes cagA/cagE, as described in Table 2.
Table 2.
Distribution of samples according to gastric lesions and Helicobacter pylori genes.
| Lesion | HP presence | HP virulence status | cagA | cagE | N |
|---|---|---|---|---|---|
| 1- cancer | |||||
| 1a | pos | Virulent | pos | pos | 8 |
| 1b | pos | Non virulent | neg | pos | 3 |
| 1c | pos | Non virulent | neg | neg | 4 |
| 2-metaplasia | |||||
| 2a | pos | Virulent | pos | pos | 9 |
| 2b | pos | Virulent | pos | neg | 2 |
| 2c | neg | - | neg | neg | 9 |
| 3-active gastritis | |||||
| 3a | pos | Virulent | pos | pos | 7 |
| 3b | pos | Non virulent | neg | neg | 5 |
| 4-inactive gastritis | |||||
| 4a | pos | Virulent | pos | neg | 6 |
| 4b | pos | Non virulent | neg | neg | 6 |
| 4c | neg | - | neg | neg | 4 |
| 5-reference | neg | - | neg | neg | 3 |
HP = Helicobacter pylori; pos = presence; neg = absence; N = number of cases; Reference = normal cases (endoscopy with negative results)
Results
Sequencing and coverage data
Metagenomic data collected from 66 samples were grouped according to histopathological classification, H. pylori, and virulence status. From these, 1,957,697 high-quality reads, with a minimum of 125 sequences (CG150 sample) and a maximum of 72,056 sequences (LG76 sample), were obtained. Three samples had fewer than 10,000 sequences (CG150, LG171, and TG10) and were excluded from subsequent analyses. The Good’s coverage estimator was > 97% for the overall sequence set.
Alpha diversity
Alpha diversity was evaluated using Chao1-richness, Simpson-diversity index, and Shannon diversity index (combining richness and diversity).
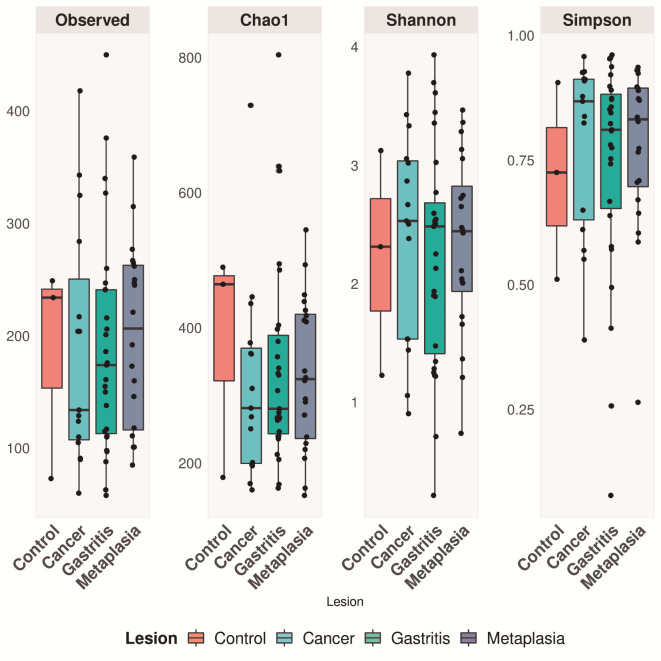
The Shapiro-Wilk test showed that all diversity metrics for this dataset were non-parametric, except for the Shannon test. ANOVA followed by Tukey’s test for Shannon and Kruskal–Wallis were applied to the other estimators, but no statistically significant difference was observed between them for the alpha diversity metric (Fig. 1). Nevertheless, it was possible to observe from Fig. 2, a microbial signature composition in each lesion group.
Fig. 1.
Alpha Diversity by Observed, Chao1, Shannon and Simpson analysis. The samples were separated by type of lesion (Control, Cancer, Gastritis and Metaplasia).
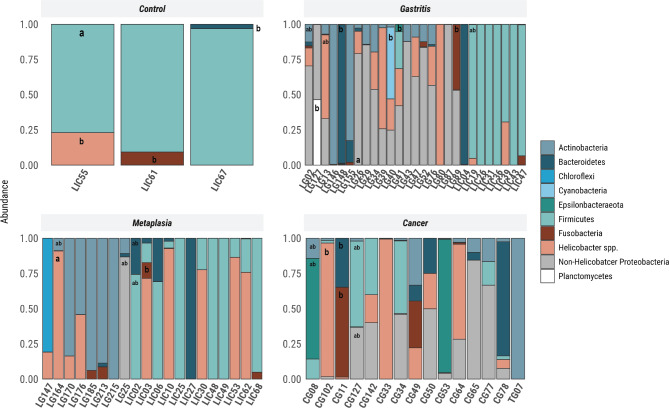
Fig. 2.
Distribution of files with an abundance above 5%, separated by lesion (Gastritis, Metaplasia and Cancer). Control refers to patients without gastric lesions. The different colors in the vertical bars indicate the main phyla observed in each sample. Lowercase letters show the statistical importance of that phylum in the sample group.
Global taxonomic composition
Phylum
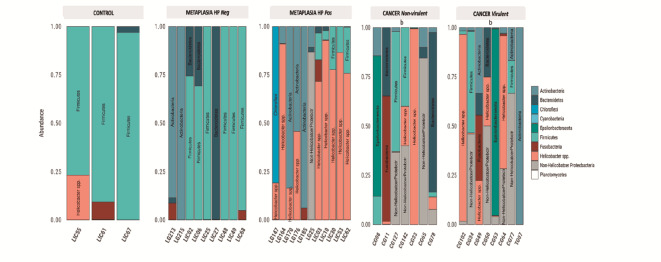
Operational taxonomic units (OTUs) with > 5% abundance were affiliated with 10 bacterial phyla. The phylum composition varied among the different gastric lesions (p = 0.0292, two-way ANOVA and Tukey’s HSD test) (Fig. 2). A statistically significant difference in microbial composition was observed in each group according to the inflammation status (p = 1.26e-11), H. pylori presence (p = 0.000133), and virulence (p = 0.044126). In the metaplasia group, the microbial composition differed from that of the control (p = 6.73e-09) and cancer groups (p = 0.049779) (Fig. 3). The abundance of the phylum Firmicutes in the control cluster decreased among the gastric lesions; conversely, Actinobacteria were present in all gastric lesions but not in normal tissue. Notably, in addition to Firmicutes being the most abundant phylum in the control group, this was also the case for inactive gastritis without H. pylori and non-virulent H. pylori. (p = 2.11 e−12). The Proteobacteria phylum was the most frequently detected group in cancer, metaplasia, and active gastritis. Notably, we observed a negative association between the presence of Firmicutes and Helicobacter spp. (p = 1.45e-07) in this group (Fig. 3), with Helicobacter spp. being statistically more abundant in the metaplasia samples than in the other lesions (Fig. 2). No significant differences in H. pylori virulence were observed in the cancer group (Fig. 3).
Fig. 3.
Distribution of phyla with abundance above 5% for Metaplasia and Cancer. Control refers to patients without gastric lesions. Samples derived from patients with metaplasia are separated into “HP Pos” and “HP Neg” and, referring to the presence or absence of H. pylori. Samples derived from cancer patients are separated into “Non-virulent” and “Virulent”, referring to the virulence profile of the H. pylori.
Order
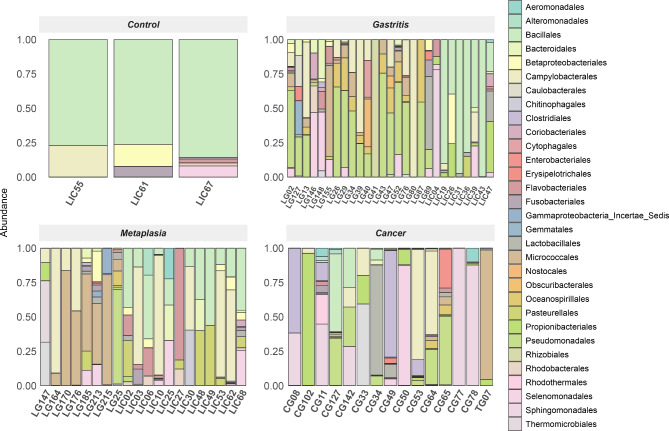
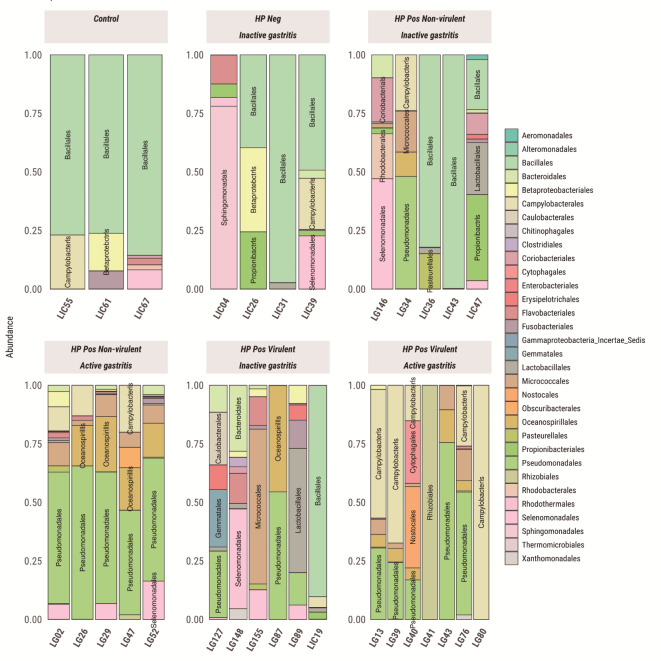
At the order level, the abundance significantly varied among lesions (p = 3.3e-09). The most frequent order in the control group was Bacillales, while that in gastritis was Pseudomonadales and Campylobacterales and, in metaplasia was Campylobacterales (Fig. 4). Between gastritis, the H. pylori presence and status significantly changed the microbiota composition (p = 0.000575), with Pseudomonadales being more abundant in active gastritis carrying non-virulent H. pylori strains (Fig. 5). In metaplasia, the microbial composition differed significantly from that of the control (p = 2.15e-08) and between the presence and absence of H. pylori (6.392 3.4e-16). In addition, a negative association was observed between Bacillales and Campylobacterales (Fig. 6). Considering the cancer group, Sphingomonadales was more abundant than that in the control group (p = 0.00391) (Fig. 4).
Fig. 4.
Distribution of orders with abundance above 5% separated by lesion. Control refers to patients without gastric lesions.
Fig. 5.
Distribution of orders with abundance above 5% for Gastritis. Control refers to patients without gastric lesions. Gastritis samples are separated according to the presence and virulence of H. pylori as well as active or inactive gastritis status.
Fig. 6.
Distribution of orders with abundance above 5% for Metaplasia. Control refers to patients without gastric lesions. Samples derived from patients with metaplasia are separated into “HP Pos” and “HP Neg”, referring to the presence or absence of H. pylori.
Genus
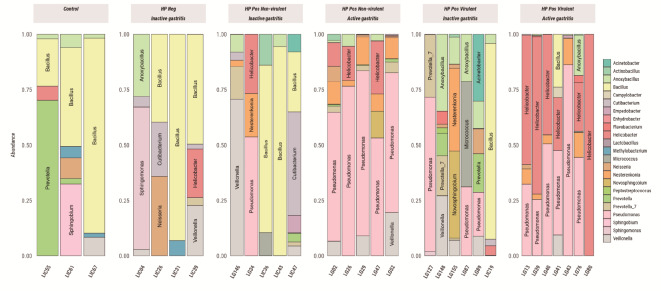
A diverse community of 270 genera was detected in this study. Among these, 79, 205, 176, and 177 genera were in controls, gastritis, metaplasia, and cancer patients, respectively. Overall, Helicobacter was the most common genus, followed by Pseudomonas and Bacillus. Analyses at the genus level (abundance > 20%) are presented in Figs. 7, 8, 9 and 10. A significant difference in microbiome abundance was observed in the general comparison between lesions (p = 2e-16), Fig. 7). Gastritis status (active and non-active) still showed differences in microbiome composition, but there was a notable positive association between the presence of Pseudomonas and Helicobacter, especially in active gastritis (Fig. 8). A higher frequency of Bacillus was observed in the control and inactive gastritis groups (Fig. 8). Similarly, among the metaplasia clusters, the differential microbiomes were influenced by H. pylori infection (p = 2.67e-09; Fig. 9).
Fig. 7.
Distribution of genus with abundance above 20%, separated by lesion. Control refers to patients without gastric lesions.
Fig. 8.
Distribution of genera with abundance above 20% for the Gastritis group. Control refers to patients without gastric lesions. Samples derived from patients with gastritis are separated according to the presence and virulence of H. pylori as well as the active or inactive state of the gastritis.
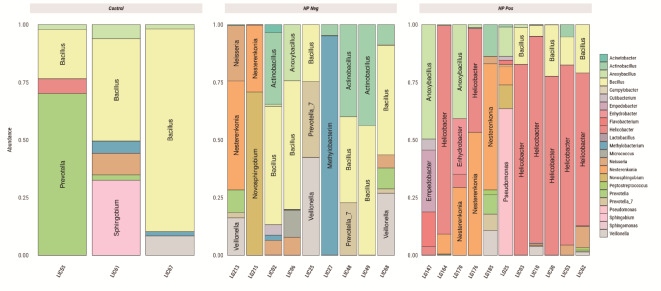
Fig. 9.
Distribution of genera with abundance above 20% for Metaplasia. Control refers to patients without gastric lesions. Samples derived from patients with metaplasia are separated into “HPneg” and “HPpos”, referring to the absence or presence of H. pylori, respectively.
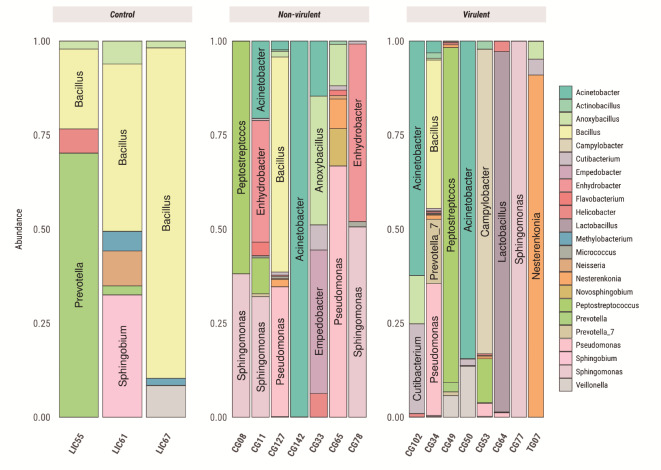
Fig. 10.
Genus’s distribution with abundance above 20% for Cancer. Control refers to patients without gastric lesions. Samples derived from cancer patients are separated into “Non-virulent” and “Virulent”, referring to the virulence profile of the H. pylori.
The microbial composition differed between the control and cancer groups (p = 0.0045), with Sphingomonas being more frequent in the cancer group. Although there was no significant difference in virulence, Fig. 10 shows a greater abundance of this genus in the non-virulent H. pylori cluster. Finally, Fig. 11 summarizes the main genera involved in each lesion group classified into families.
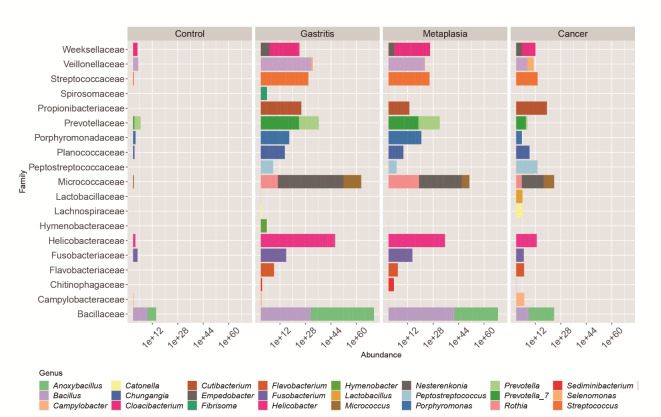
Fig. 11.
Main genera involved in each group of gastric lesions, classified in families.
Indicator genus
Indicator analysis was performed to identify the main genera associated with lesion type, clinical status, and virulence. This analysis revealed that 10 of 270 genera were responsible for the statistically significant differences. Most indicator genera were part of a rare biosphere, with abundances below 0.01%. Four of them (Microvirga, Massilia, Mesorhizobium, and Aquincola) were associated with the control group, Salinicola was associated with active gastritis, Lactobacillus was associated with the cancer group, Raoultella and Leuconostoc were associated with cancer non-virulent H. pylori and Sarcina and Centipeda were associated with cancer virulent HP (Table 3; Figs. 11 and 12).
Table 3.
Indicator Genus statistically associated with gastric lesions.
| Lesion group | Indicator | p-value* |
|---|---|---|
| Control | Microvirga | 0.020 |
| Massilia | 0.040 | |
| Mesorhizobium | 0.045 | |
| Aquincola | 0.045 | |
| Active gastritis | Salinicola | 0.001 |
| Cancer | Lactobacillus | 0.002 |
| Cancer non-virulent HP | Raoultella | 0.020 |
| Leuconostoc | 0.021 | |
| Cancer virulent HP | Sarcina | 0.032 |
| Centipeda | 0.044 |
*P-value < 0.05 was considered statistically significant
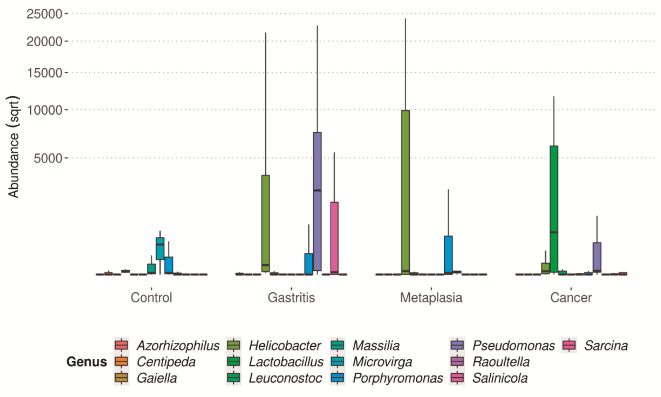
Fig. 12.
Most abundant genera in each group of gastric lesions.
Discussion
In this study, we investigated the gastric microbiota by considering sequential histological events in intestinal GC development. Gastritis was grouped according to neutrophilic infiltration as ACG or ICG, reflecting differences in host response to infections. In addition, H. pylori was categorized as virulent mainly by the presence of the cagA or cagE genes. This aspect differentiates the present study from the gastric metagenomics currently published, in which the association with H. pylori genotype, when performed, was only with the presence of cagA.
Consistent with previous studies, the phylum composition differed among the gastric lesion’s studies27–30; however, Dicksved31 and Wang et al.32 did not find any differences in the diversity indices of bacterial phyla. Nonetheless, both studies had a small number of samples; the study by Dicksved included 10 patients with GC and 5 controls with dyspepsia, while in Wang’s study, only 12 out of 315 patients with gastritis and GC were not analyzed using 16 S rRNA NGS.
Firmicutes seem to be associated with more benign lesions since this phylum was statistically more abundant in the control group and with inactive gastritis in the absence or presence of less virulent H. pylori. In a study by Gantuya et al.33 with gastritis patients from Mongolia, an area with high GC incidence and mortality, Firmicutes was the most frequent phylum in the control group and was negative for H. pylori. Thus, a negative association between the presence of Firmicutes and Helicobacter spp. was observed. Consistent with this result, Maldonado-Contreras et al.34 in a study aiming to characterize the structure of the human gastric bacterial community in relation to H. pylori status, observed a decreased abundance of Firmicutes, Actinobacteria and Bacteroidetes in H. pylori positive cases. In contrast to Firmicutes, the Actinobacteria phylum was found only in the lesions (not in the control samples); however, there were no differences among the groups with respect to H. pylori status. The Proteobacteria phylum, which includes Helicobacter, was the most frequent group associated with cancer, metaplasia, and active gastritis.
At order level, Bacillales was the Firmicutes phylum in the control group. Bacillales play an active role in the production of bioproducts, such as organic acids, chemicals, surfactants, enzymes, and insecticides35. Also, Bacillales are found in milk and cheese36, and have been used in the food industry35. Staphylococci, a representative of Bacillales, are abundant in the human skin microbiome, with Staphylococcus epidermidis being the most frequently isolated species37,38. This explains the predominance of the order in the control group.
Protobacteria were represented by Pseudomonadales, the most abundant order in gastritis; however, their abundance was associated with the presence of non-virulent H. pylori in active gastritis. The association Proteobacteria, including Pseudomonadales, with the presence of inflammation has been observed in pediatric Crohn disease patients39 which may explain gastric activity in the presence of less virulent H. pylori. In contrast, Campylobacterales, the order in which H. pylori belongs, was the most abundant order in metaplasia, which is in accordance with the Correia cascade for intestinal cancer progression, associated with H. pylori as the main etiologic factor.
A diverse community of 270 genera was detected in this study. Among these, 79, 205, 176 and 177 genera were in controls, gastritis, metaplasia and cancer patients, respectively. Overall, Helicobacter was the most common genus, followed by Pseudomonas and Bacillus. In this study, ten genera were highlighted because they were related to the groups of lesions. Most of these indicator genera were part of the rare microbiota with abundances below 0.01%. Lactobacillus, which was associated with cancer samples is in accordance with other studies40–43.
The higher relative Lactobacillus abundance in cancer could be a consequence of its overgrowth as a result of the metabolic changes that occur during the carcinogenesis process, in which the pH increases due to the gradual reduction in gastric secretion. This gastric acid reduction can lead to colonization of the stomach by species from ingestion, migration from the oropharyngeal cavity or by enterogastric reflux44–46. In this scenario, Lactobacillus, being lactic acid bacteria, could tolerate this new acid level and proliferate, increasing their relative abundance47,48. In addition, the metabolism of oral Lactobacillus generates volatile sulphur compounds, short-chain fatty acids (SCFAs), reactive oxygen species, hydrogen peroxide and lactic acid, all of which have already been implicated in chronic inflammation, genomic instability and carcinogenesis49. Although species of this genus are widely used as probiotic microorganisms, it is important to highlight that biological behaviors substantially vary between different species. Therefore, studies evaluating species that are effectively associated with GC are important for developing a therapeutic approach.
In this series of patients, it was also possible to identify the indicator genera according to H. pylori status. An interesting association was found between Leuconostoc and Raoultella with cancer-associated non-virulent H. pylori. Leuconostoc spp are used in the food industry for their fermentation properties and production and contribute to the flavor of fermented products50. They are not part of the human microbiota51. However, studies have reported infection by this microorganism in the blood, parenteral nutrition catheters, cerebrospinal fluid, and urine, especially in immunocompromised patients52–55. Recently, Nouri et al.56 compared the oral microbiota of healthy individuals to patients with several cancer types (oral, gastric, head and neck, and pancreatic) and observed that a higher abundance of Leuconostoc was associated with a higher risk of all cancers. In addition, the authors suggested that fluctuations in certain genera of oral bacteria, such as Streptococcus, Haemophilus, Neisseria, and Leuconostoc, could induce the production of SCFAs, causing the production of cytokines and inflammation, potentially leading to cancer.
Reports of human infections by the genus Raoultella have increased over the last decade56. Raoultella spp. are very similar to Klebsiella, sharing ecological, biochemical, clinical, and microbiological characteristics and making differential diagnosis very difficult with traditional microbiological techniques57. These bacteria are found in plant ecosystems, water and soil and can colonize humans and animals57. Raoultella spp. can cause a wide range of clinical syndromes, such as bacteremia, pneumonia, and urinary and biliary tract infections, especially in immunocompetent patients58. Except for a case report of a patient with early GC and Raoultella planticola bacteremia59, our study is the first to report Raoultella spp. as an indicator of GC. Further studies are needed to clarify its relationship with nonvirulent H. pylori cancer samples.
Sarcina and Centipeda were associated with cancer-virulent H. pylori. Sarcina spp. are present in soil and cereal grains60 and are often associated with delayed gastric emptying and movement of food to the intestine in addition to dyspepsia, gastritis, ulcers, and gastric perforation61–63. Studies indicate Sarcina ventriculi is a relevant species in gastric pathologies, as extensively reviewed by Tartaglia et al.64. However, it should be noted that Sarcina spp. have also been detected in healthy individuals65. In our study, we observed an interesting association of Sarcina with the cancer-virulent H. pylori group. The coexistence between Sarcina and H. pylori has already been reported by Sauter et al. in patients with Gastritis/Duodenitis66. Only one study, from Ugarde et al. (2022), reported infection by Sarcina ventriculi in the biopsy of two gastric tumors67. These data encouraged us to conduct more in-depth analyses of this notable relationship between virulent H. pylori and Sarcina ventriculi. Regarding Centipeda, the most relevant findings were its association with chronic periodontitis68, dental caries69 and, as a driver for colorectal carcinoma70 and its prognostic value for patients with oral cancer71. Our study is the first to report this genus in gastric tumor samples, and this relationship should be studied in greater depth.
The genus Salinicola, classified in the Halomonadaceae family, currently includes 12 gram-negative, aerobic or facultative anaerobes, which are halophilic. It is widely distributed in saline aquatic and terrestrial habitats and is associated with halophyte plants and sea animals. Salinicola species or strains have never been isolated from unequivocally pathological material from humans, animals, or plants72. This species was identified in our study as an indicator of active gastritis (p = 0.001), therefore, studies that clarify this relationship will be relevant.
Overall, this series of patients demonstrated that H. pylori virulence status can contribute to significant differences in the composition of the gastric microbiota between the sequential steps of the carcinogenesis cascade. The identification of indicative bacterial genera is emphasized, which are still poorly associated with diseases, and provides additional evidence that the microbiota is relevant for gastric carcinogenesis in addition H. pylori. Further studies with functional analysis are needed to clarify the clinical implications of these findings.
Acknowledgements
The authors would like to thank the Genomics and Bioinformatics Center (CeGenBio) of Drug Research and Development Center of Federal University of Ceara for technical support.
Author contributions
Conceptualization: SHBR, ACF, VMMM; Data curation: SHBR, ACF, VMMM, MMOB; Project administration: SHBR; Manuscript draft: SHBR, ACF; Review and editing: SHBR, ACF, VMMM; Approval of final version: all.**SHBR and ACF have contributed equally to this work and share first authorship.
Funding
This work was supported by Fundação Cearense de Apoio ao Desenvolvimento Científico e Tecnológico (FUNCAP), process no. 07939716/2020; SHBR and VMMM thank Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq) for their fellowship of research.
Data availability
The datasets generated and/or analyzed during the current study are available in the National Center for Biotechnology Information repository at [https://www.ncbi.nlm.nih.gov/bioproject/PRJNA1145484] (accession number PRJNA1145484). Additional information is available from the corresponding author upon request.
Declarations
Competing interests
The authors declare no competing interests.
Ethics approval
The study was approved by Ethics Committee (COMEP) of the Federal University of Ceará (protocol 071002/10) and conducted in accordance with the Declaration of Helsinki. All participants of the study signed individual informed consent forms.
Footnotes
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Silvia Helena Barem Rabenhorst and Adriana Camargo Ferrasi contributed equally to this work.
References
- 1.Sung, H. et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin.71(3), 209–249. 10.3322/caac.21660 (2021). [DOI] [PubMed] [Google Scholar]
- 2.Oue, N., Sentani, K., Sakamoto, N., Uraoka, N. & Yasui, W. Molecular carcinogenesis of gastric cancer: Lauren classification, mucin phenotype expression, and cancer stem cells. Int J Clin Oncol.24(7), 771–778. 10.1007/s10147-019-01443-9 (2019). [DOI] [PubMed] [Google Scholar]
- 3.Conteduca, V. et al. pylori infection and gastric cancer: state of the art (review). Int J Oncol.42(1), 5–18. 10.3892/ijo.2012.1701 (2013). [DOI] [PubMed] [Google Scholar]
- 4.Moodley, Y. et al. Age of the association between Helicobacter pylori and man. PLoS Pathog.8(5), e1002693. 10.1371/journal.ppat.1002693 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Maixner, F. et al. The 5300-year-old Helicobacter pylori genome of the Iceman. Science.351(6269), 162–165. 10.1126/science.aad2545 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Olbermann, P. et al. A global overview of the genetic and functional diversity in the Helicobacter pylori cag pathogenicity island. PLoS Genet.6(8), e1001069. 10.1371/journal.pgen.1001069 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Kennemann, L. et al. Helicobacter pylori genome evolution during human infection. Proc Natl Acad Sci U S A.108(12), 5033–5038. 10.1073/pnas.1018444108 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Nishiya, D. et al. Genes inside the cagPAI of Helicobacter pylori are not associated with gastric cancer in Japan. Hepatogastroenterology.51(57), 891–894 (2004) (PMID: 15143941). [PubMed] [Google Scholar]
- 9.Matteo, M. J. et al. Helicobacter pylori cag pathogenicity island genotype diversity within the gastric niche of a single host. J Med Microbiol.56(Pt 5), 664–669. 10.1099/jmm.0.46885-0 (2007). [DOI] [PubMed] [Google Scholar]
- 10.Panebianco, C., Potenza, A., Andriulli, A. & Pazienza, V. Exploring the microbiota to better understand gastrointestinal cancers physiology. Clin Chem Lab Med.56(9), 1400–1412. 10.1515/cclm-2017-1163 (2018). [DOI] [PubMed] [Google Scholar]
- 11.Ortigão, R., Pimentel-Nunes, P., Dinis-Ribeiro, M. & Libânio, D. Gastrointestinal microbiome - What we need to know in clinical practice. GE Port J Gastroenterol.27(5), 336–351. 10.1159/000505036 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Lauren, P. THE two histological main types of gastric carcinoma: Diffuse and so-called intestinal-type carcinoma. An attempt at a histo-clinical classification. Acta Pathol Microbiol Scand.64, 31–49 (1965). [DOI] [PubMed] [Google Scholar]
- 13.Dixon, M. F., Genta, R. M., Yardley, J. H. & Correa, P. Classification and grading of gastritis. The updated Sydney System. International workshop on the histopathology of gastritis, Houston 1994. Am J Surg Pathol.20(10), 1161–1181 (1996). [DOI] [PubMed] [Google Scholar]
- 14.Foster, G. D. & Twell, D. Plant Gene Isolation: Principles and Practice 426 (Wiley, 1996). [Google Scholar]
- 15.Lage, A. P. et al. Diagnosis of Helicobacter pylori infection by PCR: comparison with other invasive techniques and detection of cagA gene in gastric biopsy specimens. J Clin Microbiol.33(10), 2752–2756. 10.1128/jcm.33.10.2752-2756.1995 (1995). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Lima, V. P., Silva-Fernandes, I. J., Alves, M. K. & Rabenhorst, S. H. Prevalence of Helicobacter pylori genotypes (vacA, cagA, cagE and virB11) in gastric cancer in Brazilian’s patients: An association with histopathological parameters. Cancer Epidemiol.35(5), e32–e37. 10.1016/j.canep.2011.02.017 (2011). [DOI] [PubMed] [Google Scholar]
- 17.Caporaso, J. G. et al. Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc Natl Acad Sci USA108(Suppl 1), 4516–4522. 10.1073/pnas.1000080107 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet. J.17, 10. 10.14806/ej.17.1.200 (2011). [Google Scholar]
- 19.Andrews, S. FastQC A quality control tool for high throughput sequence data. Babraham Bioinformatics. http://www.bioinformatics.babraham.ac.uk/projects/fastqc (2010).
- 20.Rognes, T., Flouri, T., Nichols, B., Quince, C. & Mahé, F. VSEARCH: A versatile open source tool for metagenomics. PeerJ.4, e2584. 10.7717/peerj.2584.PMID:27781170;PMCID:PMC5075697 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.R Core Team. R: The R Project for Statistical Computing. (R core team, 2016).
- 22.Callahan, B. J. et al. DADA2: High-resolution sample inference from Illumina amplicon data. Nat Methods.13(7), 581–3. 10.1038/nmeth.3869 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Yilmaz, P. et al. The SILVA and “All-species Living Tree Project (LTP)” taxonomic frameworks. Nucleic Acids Res.42, D643-8. 10.1093/nar/gkt1209 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.de Cáceres, M., Sol, D., Lapiedra, O. & Legendre, P. A framework for estimating niche metrics using the resemblance between qualitative resources. Oikos.120, 1341–1350. 10.1111/j.1600-0706.2011.19679.x (2011). [Google Scholar]
- 25.Tavares, T. C. L., Bezerra, W. M., Normando, L. R. O., Rosado, A. S. & Melo, V. M. M. Brazilian semi-arid mangroves-associated microbiome as pools of richness and complexity in a changing world. Front Microbiol.12, 715991. 10.3389/fmicb.2021.715991 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Oksanen, J. et al. Package “vegan” - Community Ecology Package. 296 (2019).
- 27.Nardone, G. & Compare, D. The human gastric microbiota: Is it time to rethink the pathogenesis of stomach diseases. United Eur. Gastroenterol. J.3, 255–260. 10.1177/2050640614566846 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Ferreira, R. M. et al. Gastric microbial community profiling reveals a dysbiotic cancer-associated microbiota. Gut.67, 226–236. 10.1136/gutjnl-2017-314205 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Stewart, O. A., Wu, F. & Chen, Y. The role of gastric microbiota in gastric cancer. Gut Microbes.11, 1220–1230 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Eun, C. S. et al. Differences in gastric mucosal microbiota profiling in patients with chronic gastritis, intestinal metaplasia, and gastric cancer using pyrosequencing methods. Helicobacter.19(6), 407–416. 10.1111/hel.12145 (2014). [DOI] [PubMed] [Google Scholar]
- 31.Dicksved, J. et al. Characterization of the stomach microbiota in patients with gastric cancer and in controls. J Med Microbiol.58(4), 509–516. 10.1099/jmm.0.007302-0 (2009). [DOI] [PubMed] [Google Scholar]
- 32.Wang, L. et al. Bacterial overgrowth and diversification of microbiota in gastric cancer. Eur J Gastroenterol Hepatol.28(3), 261–266. 10.1097/MEG.0000000000000542 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Gantuya, B. et al. Gastric microbiota in helicobacter pylori-negative and -positive gastritis among high incidence of gastric cancer area. Cancers (Basel).11(4), 504. 10.3390/cancers11040504 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Maldonado-Contreras, A. et al. Structure of the human gastric bacterial community in relation to Helicobacter pylori status. ISME J.5(4), 574–579. 10.1038/ismej.2010.149 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Harirchi, S. et al. Bacillales: From taxonomy to biotechnological and industrial perspectives. Microorganisms.10(12), 2355. 10.3390/microorganisms10122355 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Delbès, C., Ali-Mandjee, L. & Montel, M. C. Monitoring bacterial communities in raw milk and cheese by culture-dependent and -independent 16S rRNA gene-based analyses. Appl Environ Microbiol.73(6), 1882–1891. 10.1128/AEM.01716-06 (2007). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Coates, R., Moran, J. & Horsburgh, M. J. Staphylococci: Colonizers and pathogens of human skin. Future Microbiol.9(1), 75–91. 10.2217/fmb.13.145 (2014). [DOI] [PubMed] [Google Scholar]
- 39.Alonzo, F. Toward uncovering the complexities of bacterial interspecies communication and competition on the skin. mBio.13(4), e0132022. 10.1128/mbio.01320-22 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Haberman, Y. et al. Pediatric Crohn disease patients exhibit specific ileal transcriptome and microbiome signature. J Clin Invest.124(8), 3617–3633. 10.1172/JCI75436 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Vinasco, K., Mitchell, H. M., Kaakoush, N. O. & Castaño-Rodríguez, N. Microbial carcinogenesis: Lactic acid bacteria in gastric cancer. Biochim Biophys Acta Rev Cancer.1872(2), 188309. 10.1016/j.bbcan.2019.07.004 (2019). [DOI] [PubMed] [Google Scholar]
- 41.Li, Z. P. et al. Overgrowth of Lactobacillus in gastric cancer. World J Gastrointest Oncol.13(9), 1099–1108. 10.4251/wjgo.v13.i9.1099 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Wang, L. et al. Gastric mucosa-associated microbial signatures of early gastric cancer. Front Microbiol.11, 1548. 10.3389/fmicb.2020.01548 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Gunathilake, M. et al. Alterations in gastric microbial communities are associated with risk of gastric cancer in a Korean population: A case-control study. Cancers (Basel).12(9), 2619. 10.3390/cancers12092619 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Beasley, D. E., Koltz, A. M., Lambert, J. E., Fierer, N. & Dunn, R. R. The evolution of stomach acidity and its relevance to the human microbiome. PLoS ONE.10(7), e0134116. 10.1371/journal.pone.0134116 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Del Piano, M. et al. The innovative potential of Lactobacillus rhamnosus LR06, Lactobacillus pentosus LPS01, Lactobacillus plantarum LP01, and Lactobacillus delbrueckii Subsp. delbrueckii LDD01 to restore the “gastric barrier effect” in patients chronically treated with PPI: A pilot study. J Clin Gastroenterol.46(Suppl), S18-26. 10.1097/MCG.0b013e318267b55d (2012). [DOI] [PubMed] [Google Scholar]
- 46.Gray, J. D. & Shiner, M. Influence of gastric pH on gastric and jejunal flora. Gut.8(6), 574–581. 10.1136/gut.8.6.574 (1967). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Azcarate-Peril, M. A., Altermann, E., Hoover-Fitzula, R. L., Cano, R. J. & Klaenhammer, T. R. Identification and inactivation of genetic loci involved with Lactobacillus acidophilus acid tolerance. Appl Environ Microbiol.70, 5315–5322. 10.1128/AEM.70.9.5315-5322.2004 (2004). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Shin, C. M. et al. Impact of long-term proton pump inhibitor therapy on gut microbiota in F344 rats: Pilot study. Gut Liver.10, 896–901 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Hussain, S. P., Hofseth, L. J. & Harris, C. C. Radical causes of cancer. Nat. Rev. Cancer.3, 276–285. 10.1038/nrc1046 (2003). [DOI] [PubMed] [Google Scholar]
- 50.Roșca, M. F. et al. Leuconostoc citreum: A promising sourdough fermenting starter for low-sugar-content baked goods. Foods.13(1), 96. 10.3390/foods13010096 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Cuervo, M. S. I., Cortés, L. J., Rodríguez, R. E., Hormaza, A. N. & Vargas, S. E. Leuconostoc sp en pacientes con cáncer: Estudio descriptivo [Leuconostoc sp in cancer patients: A descriptive study]. Rev Chilena Infectol.25(3), 184–8 (2008). [PubMed] [Google Scholar]
- 52.Dhodapkar, K. M. & Henry, N. K. Leuconostoc bacteriemia in an infant with short-gut syndrome: Case report and literature review. Mayo Clin Proc.71, 1171–1174. 10.4065/71.12.1171 (1996). [DOI] [PubMed] [Google Scholar]
- 53.Espinoza, R. et al. Leuconostoc bacteremia after liver transplantation: Another cause of vancomycin resistant gram-positive infection. Clin Transp.11, 322–324 (1997). [PubMed] [Google Scholar]
- 54.Jofré, L. et al. Infección por Leuconostoc en pacientes con síndrome de intestino corto, nutrición parenteral y alimentación enteral continua. Rev Chil Infect.23, 340–345 (2006). [DOI] [PubMed] [Google Scholar]
- 56.Fauchais, A. L. et al. Rare opportunistic infection due to Leuconostoc. Rev Med Interne.24, 268–75. 10.1016/s0248-8663(03)00032-8 (2003). [DOI] [PubMed] [Google Scholar]
- 56.Nouri, Z. et al. Exploring connections between oral microbiota, short-chain fatty acids, and specific cancer types: A study of oral cancer, head and neck cancer, pancreatic cancer, and gastric cancer. Cancers (Basel).15(11), 2898. 10.3390/cancers15112898 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Sękowska, A. Raoultella spp.-clinical significance, infections and susceptibility to antibiotics. Folia Microbiol.62, 221–7. 10.1007/s12223-016-0490-7 (2017). [DOI] [PubMed] [Google Scholar]
- 58.Appel, T. M., Quijano-Martínez, N., De La Cadena, E., Mojica, M. F. & Villegas, M. V. Microbiological and clinical aspects of Raoultella spp. Front Public Health.9, 686789. 10.3389/fpubh.2021.686789 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Yamamoto, S., Nagatani, K., Sato, T., Ajima, T. & Minota, S. Raoultella planticola bacteremia in a patient with early gastric cancer. Intern Med.57(10), 1469–1473. 10.2169/internalmedicine.9611-17 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Makovska, M. et al. Species and strain variability among Sarcina isolates from diverse mammalian hosts. Animals (Basel).13(9), 1529. 10.3390/ani13091529 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Dumitru, A. et al. Fatal outcome of gastric perforation due to infection with Sarcina spp. A case report. IDCases.19, e00711. 10.1016/j.idcr.2020.e00711 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Singh, K. Emphysematous gastritis associated with Sarcina ventriculi. Case Rep. Gastroenterol.13, 207–213. 10.1159/000499446 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.de Meij, T. G., van Wijk, M. P., Mookhoek, A. & Budding, A. E. Ulcerative gastritis and esophagitis in two children with Sarcina ventriculi infection. Front. Med.4, 145. 10.3389/fmed.2017.00145 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Tartaglia, D. et al. Sarcina Ventriculi infection: A rare but fearsome event. A systematic review of the literature. Int. J. Infect. Dis.115, 48–61. 10.1016/j.ijid.2021.11.027 (2022). [DOI] [PubMed] [Google Scholar]
- 66.Haroon Al Rasheed, M. R., Kim, G. J. & Senseng, C. A rare case of Sarcina ventriculi of the stomach in an asymptomatic patient. Int. J. Surg. Pathol.24(2), 142–145. 10.1177/1066896915610196 (2016). [DOI] [PubMed] [Google Scholar]
- 66.Sauter, J. L. et al. Co-existence of Sarcina organisms and helicobacter pylori gastritis/duodenitis in pediatric siblings. J Clin Anat Pathol.1(1), 103. 10.17303/jcap.2013.103 (2013). [PMC free article] [PubMed] [Google Scholar]
- 67.Ugarte Bilbao, A. et al. Sarcina ventriculi in gastric biopsies of two patients with an underlying neoplasia. Rev Esp Enferm Dig.114(9), 557. 10.17235/reed.2022.8667/2022 (2022). [DOI] [PubMed] [Google Scholar]
- 68.Rams, T. E. et al. Centipeda periodontii in human periodontitis. Odontology.103(3), 286–291. 10.1007/s10266-014-0166-1 (2015). [DOI] [PubMed] [Google Scholar]
- 69.Celik, Z. C. et al. The complex microbiome of caries-active and caries-free supragingival plaques in permanent dentition. Niger J Clin Pract.24(10), 1535–1540. 10.4103/njcp.njcp_49_21 (2021). [DOI] [PubMed] [Google Scholar]
- 70.Wang, Y. et al. Alterations in the oral and gut microbiome of colorectal cancer patients and association with host clinical factors. Int J Cancer10.1002/ijc.33596 (2021). [DOI] [PubMed] [Google Scholar]
- 71.Granato, D. C. et al. Meta-omics analysis indicates the saliva microbiome and its proteins associated with the prognosis of oral cancer patients. Biochim Biophys Acta Proteins Proteom.1869(8), 140659. 10.1016/j.bbapap.2021.140659 (2021). [DOI] [PubMed] [Google Scholar]
- 72.Plotnikova, E. G., Anan’ina, L. N., Ariskina, E. V. & Evtushenko, L. I. Salinicola. Bergey’s Manual of Systematics of Archaea and Bacteria10.1002/9781118960608.gbm01719 (2024). [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
The datasets generated and/or analyzed during the current study are available in the National Center for Biotechnology Information repository at [https://www.ncbi.nlm.nih.gov/bioproject/PRJNA1145484] (accession number PRJNA1145484). Additional information is available from the corresponding author upon request.