ABSTRACT
Klebsiella pneumoniae is a Gram-negative bacterium associated with a wide range of community- and hospital-acquired infections. The emergence of clonal hypervirulent strains resistant to last-resort antimicrobial agents has become a global concern. The Kingdom of Saudi Arabia (KSA), with its diverse population and high tourism traffic, serves as a platform where the spread of multidrug-resistant (MDR) strains are facilitated. However, the knowledge of epidemiology and population diversity of MDR K. pneumoniae in KSA is scarce. We conducted a comprehensive genomic survey on 352 MDR K. pneumoniae isolates systematically collected from bloodstream and urinary tract infections in 34 hospitals across 15 major cities in KSA during 2022 and 2023. Whole-genome sequencing on the isolates was performed, followed by genomic epidemiology and phylodynamic analysis. Our study revealed a dynamic population characterized by the rapid expansion of several dominant clones, including, ST2096, ST147, and ST231, which were estimated to have emerged within the past decade. These clones exhibited widespread dissemination across hospitals and were genetically linked to global strains, particularly from the Middle East and South Asia. All major clones harboured plasmid-borne ESBLs and carbapenemase genes, with plasmidome analysis identifying multiple IncH, IncA/C and IncL plasmids underlying the MDR-hypervirulent phenotype. These plasmids were shared between major clones and became acquired on the same time scales as the expansion of the dominant clones. Our results report ST2096 as an emerging MDR-hypervirulent clone, emphasizing the need for monitoring of the circulating clones and their plasmid content in the KSA and broader West Asia.
KEYWORDS: Klebsiella pneumoniae, precision epidemiology, whole genome sequencing, antimicrobial resistance, hypervirulence and multidrug resistance (MDR)
Introduction
K. pneumoniae is a major Gram-negative bacterial species implicated in several hospital-acquired infections, including urinary tract and bloodstream infections [1]. Pathogenic strains of K. pneumoniae colonize a broad range of niches. These strains have a dynamic genome in which key clinically significant genes for resistance and virulence can be acquired, leading to the emergence of high-risk strains from non-pathogenic strains [2]. The emergence and global spread of K. pneumoniae resistant to last-resort carbapenem antimicrobials over recent decades have been recognized as a health priority by the World Health Organization, which places this species on the priority list of pathogens for which imminent therapeutic solutions are required [3].
Elevated virulence and resistance levels in K. pneumoniae are associated with increased mortality and poor clinical outcomes. Clones of K. pneumoniae often emerge locally within hospitals and then spread regionally and globally [4]. These clones are frequently recovered from various distant geographical locations, can colonize or infect multiple hosts, and are rapidly transmitted. The best-characterized international clones belong to Clonal Group (CG) 258, which includes Sequence Types (ST) 258 and ST11, as well as clonal groups CG14/15, CG17/20, CG43, CG147, and ST101 [4–7]. These clades display a broad phylogenetic distribution and have rapidly disseminated across countries and healthcare systems. In recent decades, these clades have shown a concerning convergence of carbapenem resistance and virulence [8]. This is particularly alarming in clinical settings, where strains labelled as both multidrug-resistant (MDR) and hypervirulent (hv) are becoming more prevalent, as non-overlapping genomic signatures for resistance and virulence co-occur within the same genomic context [8,9]. These clones can cause severe infections not only within hospital settings but also in healthy individuals in the community, with very limited treatment options [10].
The Kingdom of Saudi Arabia (KSA) is strategically located, has an expansive geographical area, and a complex and dynamic demography. The country is also an international hub for travel, trade, and religious tourism, hosting mass gathering events. These factors have turned the country into a potential melting pot for the import and transmission of antimicrobial-resistant strains worldwide, as well as an environment for the horizontal transfer of resistance and virulence genes. High carbapenem usage in the kingdom is associated with a rapid increase in the incidence of carbapenem-resistant Enterobacterales (including K. pneumoniae). Previously, multiple small-scale studies on individual or small networks of hospitals have reported the incidence of MDR clones in the country [11–15]. However, these studies are limited, using only classical molecular typing methods with poor resolution or using heterogeneous and inconsistent datasets from different time points. Therefore, the dynamics of resistant clones and their mechanisms remain largely understudied within this region.
To fill this gap, we conducted a large-scale precision epidemiology analysis to decipher the evolution of multidrug-resistant K. pneumoniae clones within KSA. We curated a collection of samples systematically recovered from 34 hospitals across the country from bloodstream and urinary tract infections. The depth and breadth of the sampling provided us with the most comprehensive picture of clones circulating in the country during a period of 16 months. We fully characterized the diverse population of K. pneumoniae, including the identification of prevalent clones and their plasmid content, with a focus on resistance, virulence, and the convergence of these traits in these isolates. Our study revealed the dissemination of both globally and regionally endemic clones and the extensive sharing of plasmids associated with virulence and resistance between the circulating clones.
Methods
Sampling and collection
This study was approved by Institutional Review Board (IRB) of King Abdullah University for Science and Technology (approval number 23IBEC027) and IRB of Saudi Ministry of Health (approval number: 23–23 M). As part of a nationwide surveillance programme, the reference lab at the Saudi Ministry of Health received 63,688 Gram-negative ESKAPE samples from hospitals affiliated with the Ministry between January 2022 and April 2023. Isolates were deduplicated as they were retrieved from different patients. In total, 21,345 samples were identified as Klebsiella pneumoniae. Of these, 7,290 were classified as MDR, i.e. resistant to at least two classes of antimicrobials [16]. From the MDR subpopulation, 200 samples from bloodstream infections and 152 samples from urinary tract infections were randomly selected to ensure broad geographical and temporal representation throughout the study period.
For samples from bloodstream infections, we obtained clinical data for date and cause of death. We recovered isolates from patients who were infected with MDR K. pneumoniae via analysis in molecular microbiology laboratories in the hospital. A combination of phenotypic and genotypic methods was used to identify the MDR K. pneumoniae isolates. We used a PCR for identifying key resistance genes for carbapenemase and ESBL. Specifically, primers were designed for the ESBL gene of blaCTX-M (F (forward): 5′-ATGTGCAGCACCAGTAAAGT-3′, R (reverse): 5′-ACCGCGATATCGTTGGTGG-3′), and for the carbapenemase genes of blaKPC (F: 5′-CGTCTAGTTCTGCTGTCTT-3′, R: 5′-CTTGTCATCCTTGTTAGGCG-3′), blaNDM-1 (F: 5′-GGTTTGGCGATCTGGTTTTC-3′, R: 5′-CGGAATGGCTCATCACGATC-3′), and blaOXA-48 (F: 5′-TTGGTGGCATCGATTATCGG-3′, R: 5′-GAGCACTTCTTTTGTGATGGC-3′). All results from this method were confirmed with whole genome sequencing data. To explore the links with nonclinical reservoirs, we included nine strains recovered from retail food (lettuce) in Jeddah [17]. We also integrated our collection with the genomic collection from a single hospital study in Jeddah for phylodynamic analysis [18].
Strain identification and susceptibility testing
The Vitek®2 machine was used for identification and antimicrobial susceptibility testing (BioMerieux, Marcy-l’Etoile, France). ID-GN (reference: 22226) and AST-N 291 (reference: 415062) cards were used for identification and susceptibility testing, respectively. The isolates were cultured overnight on agar plates at 37°C. A 0.45% sodium chloride (NaCl) solution (CareFusion, USA) was used to prepare a bacterial suspension with an optimal density of 0.5–0.63 McFarland units. The cards were inoculated and loaded in the Vitek®2 compact according to the manufacturer’s instructions. The system output was generated automatically. AST-N 291 provides results for the following antimicrobials: Extended Spectrum β-Lactamase (ESBL), ampicillin (AMP), amoxicillin/clavulanic acid (AMC), piperacillin/tazobactam (PTZ), cefalotin (CEF), cefoxitin (FOX), ceftazidime (CAZ), ceftriaxone (CRO), cefepime (FEP), imipenem (IMP), meropenem (MER), amikacin (AMK), gentamicin (GEN), ciprofloxacin (CIP), tigecycline (TGC), nitrofurantoin (NIT) and trimethoprim/sulfamethoxazole (TMP/SMX). The Clinical & Laboratory Standards Institute (CLSI) breakpoints were used to define phenotypic resistance based on minimum inhibitory concentration (MIC) values [19]. For the subset of samples that were non-susceptible to all antimicrobials, we repeated the susceptibility testing using the AST-XN20 card. This card included two combinations of β-lactam/β-lactamase inhibitors: ceftazidime/avibactam and meropenem/vaborbactam This allowed us to assess the efficiency of combination therapy. In addition, results obtained from Vitek®2 were validated using broth dilution method (Table S2, Supplemental Methods).
Sequencing and genomics analysis
We conducted DNA extraction and subjected the isolates to short read sequencing. Multiple bioinformatics analyses including, variant calling, phylogenetic and phylodynamic, transmission analysis, contextualization of the genomes and identification of resistance and virulence genes were performed using the methods detailed in the Supplemental Material. Moreover, we integrated genomics data with data on in-hospital mortality and employed a hybrid long- and short-read sequencing approach for the characterization of plasmids on selected samples, as detailed in the Supplemental Material.
Statistical significance tests
We used a one-way proportion test in R to assess the significance of the differences in the ratios. The one-way Wilcoxon signed-rank test was used to assess the significance of differences between the means. To compare the significance of different parameters inferred from the Bayesian analysis, we considered the 95% lower and upper bounds of the credible intervals (highest posterior density [HPD]), which is the shortest interval that includes 95% of the probability density.
Data availability
Genomic data collected in this study were deposited in the European Nucleotide Archive (ENA) under the study accession number PRJEB66182. Moreover, the assemblies were uploaded to the NCBI GenBank database under the accession number PRJNA1018815. Detailed metadata associated with the genomes are provided in Supplemental Table S1. Sample IDs in the table were not known to anyone outside the research group.
Results
K. pneumoniae infections and resistance level in the KSA.
Klebsiella pneumoniae isolates were collected over 16 months from 34 hospitals across the Kingdom of Saudi Arabia (KSA). In total, 34% of all Gram-negative ESKAPE isolates collected were classified as multidrug resistant (MDR). K. pneumoniae accounted for 33.51% of all Gram negative ESKAPE pathogens, with it being the second most prevalent pathogenic species within the country, following E. coli (34% of all Gram negative ESKAPE pathogens). Among MDR K. pneumoniae isolates, we selected 200 isolates from bloodstream infections and 152 isolates from urinary tract infections as representatives of our population. Overall, this sampling, represents 1.5% (352/21,345) of the total K. pneumoniae population collected and 5% (352/7,290) of the MDR K. pneumoniae subpopulation. In addition, we included nine samples isolated from food sources (lettuce).
Dominance of clones of non-KL1/KL2 capsule types in KSA.
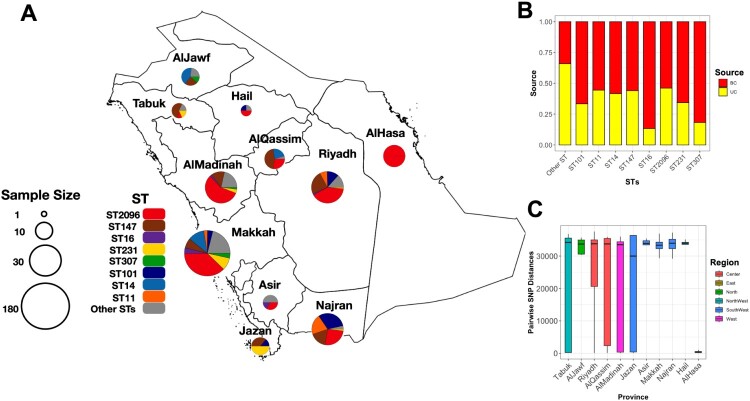
A high diversity was seen amongst our K. pneumoniae isolates, with 40 different sequence types (STs) observed. Among these, seven STs each represented more than 3% of the population: ST2096 (n = 115), ST147 (n = 75), ST231 (n = 35), ST101 (n = 32), ST11 (n = 18), ST16 (n = 15), and ST14 (n = 12). Collectively, these seven STs accounted for 87% of the total population (Figure 1A, Supplementary Table S1). When comparing our results with a single hospital study conducted in Jeddah [19], STs found in this study overlapped considerably with our collection, with ST2096, ST14, ST101, ST11, and ST231 being the most frequent STs in clinical settings in a single hospital as well as nationwide (Table 1, Supplementary Table S1), suggesting that the diversity observed in a single hospital study reflects the broader national diversity. Isolates retrieved from food showed a distinctive ST pattern, with only two of the nine samples recovered from food sources being ST101, potentially suggesting epidemiological links between food and human reservoirs. The capsular (K-locus) diversity within the major STs revealed the dominance of types distinct from the well-known globally circulating KL1/KL2 types, typically associated with pathogenicity. In our study, only ST14 isolates had the KL2 capsule type, while the other major STs were associated with KL64, KL51, KL17, and KL10. Within the ST147 isolates, three types (KL64, KL51, and KL10), were observed, pointing to multiple sub-lineages of ST147 circulating in KSA hospitals (Table 1).
Phylogenetic and geographical expansion of CG14 ST2096 clade and weak association between STs and infection type.
Figure 1.
The distribution of K. pneumoniae clones in the study cohort. (A) The frequency of STs across provinces. The size of the pie charts corresponds to the number of isolates. (B) Proportion of samples originating from different sources of isolation. The terms “BC” and “UC” stand for blood and urine culture, respectively. (C) SNP diversity of isolates in each province. Diversity is measured as the pairwise SNP distance between isolates originating from the same province. The colours represent the regions in the country.
Table 1.
Overview of the major clones. This table details the specific characteristics of the major clones. The SNP count column refers to the number of SNP sites after mapping the short reads to a local reference genome and removing hypervariable sites. The SNP count per sample column indicates the per-sample SNP within each clone. For both the SNP count and SNP count per sample columns, we included isolates from a single hospital study in Jeddah [18] in the collection. Key resistance genes include ESBL and carbapenemase genes identified by AMRfinderPlus. The Province column shows the number of provinces, out of a total of 11, where isolates from each ST clone were detected. The Hv-MDR column indicates the ratio of hypervirulence and multidrug resistant isolates, identified by Kleborate.
| Clone | SNP Count | GenomeCount | SNP Count Per Sample* | Province | K Locus | Hv Genes | Key resistance Genes | Ratio with Carbapenemase genes | Ratio with Colistin | Hv-MDR Ratio |
|---|---|---|---|---|---|---|---|---|---|---|
| ST14 | 1923 | 12 | 36.98 | 3/11 | KL2 | ybt 14; ICEKp5, ybt 9; ICEKp3 |
blaCTX-M-15, blaCTX-M-14, blaNDM-1, blaOXA-48 |
0.46 | 0.33 | 0 |
| ST231 | 579 | 35 | 13.46 | 6/11 | KL51 | ybt 14; ICEKp5, iuc 5 |
blaCTX-M-15, blaNDM-5, blaOXA-48, blaOXA-232 | 0.97 | 0.03 | 0.06 |
| ST101 | 1125 | 32 | 26.78 | 5/11 | KL17 | ybt 9; ICEKp3, iuc 1,rmpA2 |
blaOXA-48, blaOXA-232, blaCTX-M-15, blaCTX-M-27 | 0.87 | 0.69 | 0.16 |
| ST147 | 1837 | 75 | 22.4 | 6/11 | KL64/ KL10/ KL51 |
ybt 9; ICEKp3, ybt 16; ICEKp12, iuc 1,rmpA, rmpA2 |
mcr-1.1, blaCTX-M-15, blaCTX-M-14, blaNDM-1,NDM-5, blaOXA-232, blaOXA-48, blaOXA-181 | 0.96 | 0.16 | 0.55 |
| ST2096 | 2625 | 115 | 11.61 | 9/11 | KL64 |
ybt 14; ICEKp5, iuc 1,rmpA2 |
blaCTX-M-15, blaCTX-M-14, blaOXA-232, blaOXA-48, blaNDM-5 |
0.86 | 0.21 | 0.85 |
The phylogenetic analysis of the isolates revealed a diverse population composed of distinct lineages and expanding sequence types (STs) (Figure 2). The most prominent clones were linked to the CG14 clade, particularly the ST14 and ST2096 strains, aligning with recent findings from hospitals in the Middle East [20,21] and a single-hospital study in Jeddah [19]. The distribution of dominant clones showed no significant association with the site of isolation, whether from blood or urine (p > 0.05, proportion test), as isolates from various sources were dispersed throughout the phylogenetic tree (Figure 1B and Figure 2). The ST2096 clone also exhibited a broad geographical distribution, occurring in 9 out of 11 provinces (Table 1), a range greater than that of other major STs (Figure 1A, Table 1, Figure 2). Furthermore, K. pneumoniae population diversity was observed across all provinces except for the Eastern Province (Al Hasa), where all genomes belonged to a single clone, i.e. ST2096 (Figure 1A, Figure 2). The diversity of K. pneumoniae populations across provinces, measured by average SNP distance and number of STs after accounting for sample size, was comparable between provinces and did not correlate with human population size or density, which is highest in Riyadh and Makkah provinces (Figure 1C). This suggests the widespread dissemination of different clones across provinces.
ST231, ST147 and ST2096 have recently expanded across the country and show signatures of between-city transmissions.
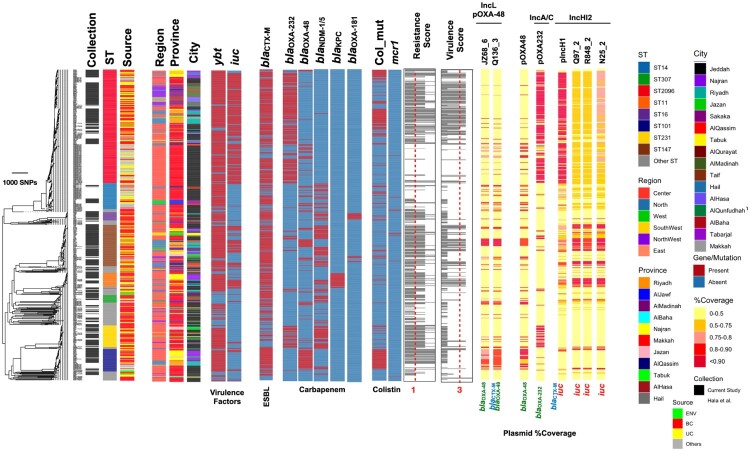
Figure 2.
Phylogenetic tree K. pneumoniae in the KSA. The tree included 361 K. pneumoniae genomes from the current study and 328 genomes from a single-hospital study in Jeddah [18]. The tree is an approximate maximum-likelihood phylogenetic tree from the alignment of 196,300 SNPs. Strains for STs with more than ten representative strains are shown. The resistance and virulence scores were computed with Kleborate. The dotted lines show the cut-off used for defining multidrug resistance and virulence factor genes. The plasmid bands correspond to the mapping coverage for the plasmids containing carbapenemase and ESBLs from long-read sequencing. “BC” and “UC” correspond to blood and urine culture samples, respectively.
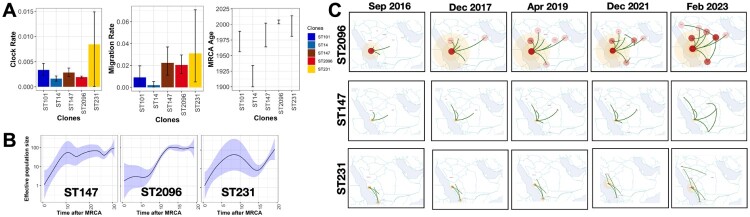
We analyzed the population dynamics of the major clones within across the country. Among the five most frequent STs, ST2096 exhibited the lowest SNP count per genome (Table 1), suggesting it has expanded more rapidly than the other clones. Using the temporal signals in the genomes of the five main clones (ST101, ST14, ST231, ST2096, and ST147), we examined their population dynamics. The clones had comparable clock rates (Figure 3A). The formation of ST2096, ST147, and ST231 clones were estimated to have occurred in 2004, 1992, and 1998, respectively, which are more recent than the formation of ST101 and ST14 clones (Figure 3A). The estimated ages for ST2096 and ST147 closely align with findings from other studies, suggesting that the population diversity of these clones in Saudi Arabia mirrors global patterns (Figure 3A) [20,21]. The skyline growth plot for the clones indicated a rapid initial expansion over 20 to 30 years for ST2096, ST147, and ST231 (Figure 3B). For ST2096 and ST147, the population stabilized following this initial increase, whereas for ST231, the population rebounded after an initial decline (Figure 3B). The average migration rates for ST2096, ST147, and ST231 were higher than those for the ST101 and ST14 clones (Figure 3A), indicating rapid dissemination across the country over seven years, including western, northern, and central regions of Saudi Arabia (Figure 3C). Reconstructed transmission networks for the three most frequent clones (ST2096, ST147, and ST231) also revealed multiple transmission subnetworks: 26 for ST2096, 13 for ST147, and 8 for ST231. Among these, 25/26 for ST2096, 13/13 for ST147, and 6/8 for ST231 included isolates from different cities (Figure S1), underscoring the nationwide dissemination of these clones on epidemiological time scales.
K. pneumoniae population is primarily linked with clones circulating in the Middle East and South Asia.
Figure 3.
Population dynamics of major clones. (A) The estimated population dynamics parameters from the phytogeography analysis. Clock rates show the rates based on the number of substitutions per site per year. The migration rates show the parameters for all the cities and the rates. The unit for migration is the number of migration events per year. The most recent common ancestor (MRCA) age indicates the time at which the clone formed. The error bars show the 95% confidence intervals. (B) The skyline growth plot. The shaded plot shows the 95% confidence intervals. (C) The recent spread of ST147, ST231 and ST2096 clones across the country. The trajectories show the inferred dissemination (migration routes) across the country. The circle sizes and colour intensity and rate show the relative marginal migration rate per city.
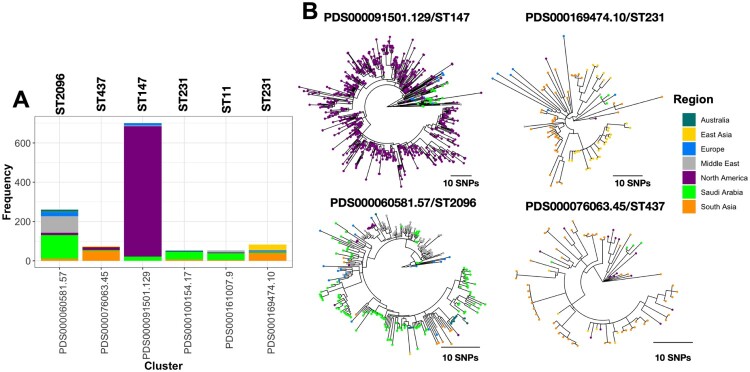
We contextualized the collection of isolates with a global dataset from the Pathogen Detection Database and identified epidemiological clusters, defined as SNP clusters with a maximum SNP distance of 50 (see Methods). In total, eleven clusters, including genomes from both the studied collection and the global dataset, were identified (Figure 4A). Among these, six SNP clusters (representing ST2096, ST437, ST11, ST147, and ST231) contained more than 50 genomes. A closer examination of the population structure for clusters containing at least ten Saudi strains revealed distinct patterns of how Saudi strains mixed with the global population (Figure 4B). Specifically, ST147 genomes from Saudi Arabia formed a unique clade, contrasting with the expanding clade observed in North America, Europe, and Australia. In contrast, ST2096, ST437, and ST11 genomes were primarily mixed with Middle Eastern and South Asian samples in the phylogenetic trees. Sensitivity analysis also identified potential cross-country transmissions within these clusters (highlighted in the yellow-shaded zone in Figure S2). For ST147, isolates from the Middle East were introduced into the cluster of Saudi isolates at SNP distances as low as 10. In the case of ST2096, isolates from the Middle East, Australia, and North America were found within the transmission threshold (i.e. 20 SNPs, see Methods). For ST231 and ST437, transmissions involving European and South Asian isolates (Figure S2) were detected within the transmission threshold. Since all the global isolates were of clinical origin, these findings suggest transmission of these isolates across healthcare settings at both regional and global levels (see Discussion).
K. pneumoniae population contains multiple clones with high resistance levels.
Figure 4.
Mixing of Saudi clones with the global collection. (A) The SNP clusters (clusters of genomes fewer than 50 SNPs apart) as derived from the pathogen detection database (see Methods for the definition of the clusters). (B) Neighbor-joining tree for the clusters that contained more than two isolates from Saudi Arabia and the global collection in the database. The colour tips represent strains from Saudi Arabia and other regions.
To investigate the level of antimicrobial resistance in our isolates, we performed antimicrobial susceptibility testing (See methods). The results demonstrated the collection was extensively resistant to multiple classes of antimicrobials, including β-lactams (cephalosporins (83%) and carbapenems (85%)), aminoglycosides (67%), fluoroquinolones (ciprofloxacin) (84%), trimethoprim (82%), and nitrofurantoin (62%) (Figure S3). Considering the high rate of resistance, we identified 56 strains which were non-susceptible to all drugs tested to undergo additional testing using β-lactam/ β-lactamase inhibitor combinations. This further testing revealed that of the 56 non susceptible strains, 44% (25/56) became susceptible under ceftazidime/avibactam and 21% (12/56) under meropenem/vaborbactam, pointing to the effectiveness of combination therapy (Table S1). Overall, the high resistance levels are evidenced by the high frequencies of genes or mutations conferring carbapenemase, ESBL, and colistin resistance, observed at 77%, 72%, and 27% frequencies, respectively. These determinants were distributed across the phylogenetic tree (Figure 2), with major STs having a greater content of these key genes and hence a greater resistance score, which was significantly greater than that of the minor STs combined (p-value from Wilcoxon signed rank test < 0.05) (see Methods) (Figure 2A, Figure S4A, S4B). ST2096 and ST147 clones had a significantly higher total number of resistance genes, including carbapenemase genes, than the rest of the population (one-sided p value from Wilcoxon test < 0.01) (Table 1, Figure 5A). When compared to the remaining population, the major clones exhibited a higher frequency of strains which genotypes were ESBL-positive or harboured non-ESBL β-lactamases, ciprofloxacin resistance mutations and carbapenemases (one-sided p value from proportion test < 0.05) (Figure S4A and S4B). The major clones also appeared to have distinctive set of resistance genes, in that in ST101 (12 genes), ST14 (11 genes), ST147 (31 genes), ST2096 (19 genes) and ST231 (15 genes) contained resistance genes from different antimicrobials classes that were at a higher frequency than the background populations (Table S3). Notably, major clones harboured distinctive overrepresented carbapenemase genes, i.e. blaOXA-48 in ST101, ST147 and ST14, blaOXA-232 in ST231 and ST2096, blaNDM-1 in ST14, ST147 and ST307 and blaNDM-5 in ST147 and ST231 (Figure S4, Table S3). A similar pattern was observed for ESBL blaCTX-M variants, which point to different evolutionary trajectories of resistance mediated by plasmid-borne resistance genes in different clones. The acquired mcr-1 gene against colistin, another last resort antimicrobial, was detected in ST147 clone (Figure S4A, Table S3). Colistin resistance truncation mutations in the mgrB and pmrB genes were found in 158 and 3 genomes across the phylogenetic tree, respectively, with the highest prevalence in strains ST2096, ST101 and ST14 (Figure S4B, Table S3). The colistin resistance mutations and genes was found also linked with the phenotypic resistance (MIC values greater than ≥ 4 µg/mL for isolates with colistin resistance determinants). The integration of the genotypic data with phenotypic data indicated strong link between resistance phenotypes and the presence of determinants for ESBLs, carbapenemase, aminoglycosides and ciprofloxacin (Figure S5), confirming strong phenotypic effect of these resistance genes.
K. pneumoniae population contains multiple clones with high resistance and virulence levels, linked with elevated mortality and sepsis rates.
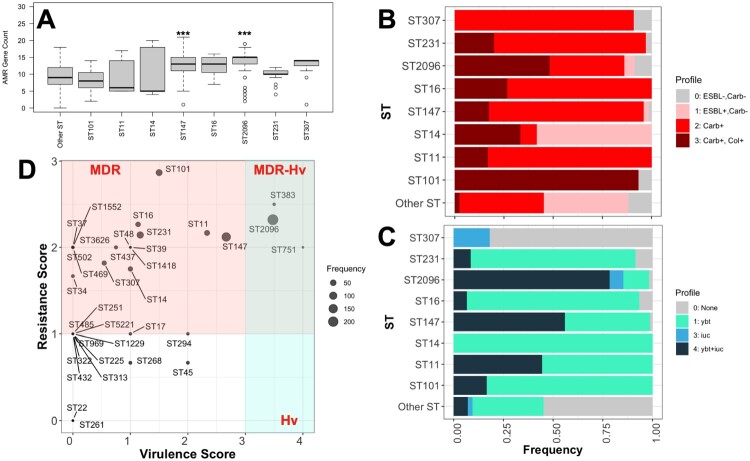
Figure 5.
Frequency of clinically significant resistance and virulence genes across major clones of K. pneumoniae from blood stream infections in the KSA. (A) The frequency of resistance genes across the major STs. The “***” symbol indicates the significance level of the higher number of resistance genes from the Wilcoxon rank-sum test (p-value < 0.001). (B) ESBL, colistin resistance and carbapenemase genes across STs. The numbers show the resistance scores (see methods for more information). (C) The distribution of aerobactin (iuc) and yersiniabactin (ybt) across major clones (STs) and other STs. The numbers show the virulence scores (see methods for more information). (D) The integration of the virulence score and resistance score for the STs in the population. The shaded areas define the hypervirulent, multidrug resistant and hypervirulent-MDR resistant areas. The scores and definitions of hypervirulence and MDR were adopted from Kleborate, and details are provided in Supplemental Table S1. The x- and y-axes correspond to the average virulence and resistance scores for isolates within each ST, respectively.
The patterns observed for key virulence factor genes revealed that major clones exhibited higher virulence scores, as determined by the presence of key virulence genes (see Methods), compared to the rest of the population (Figure 2, Figure 5B). All major STs, harboured the siderophore-encoding ybt gene. Most major STs possessed a single type of ybt locus, with the exception of ST147 strains, which contained ybt 9/ICEKp3 and ybt 16/ICEKp12 (Table 1). The genotoxin colibactin gene (clb) was absent in the major sequence types (STs) but was present in two ST294-KL30 isolates. Other key genes associated with the hypervirulence phenotype, i.e. the inner membrane transporter peg-344 and the mucoid phenotype regulators rmpA and rmpA2, were identified in 34, 37, and 4 isolates, respectively. These included isolates from major clones, such as ST101, ST147, and ST2096. A total of 163 strains contained the siderophore aerobactin iuc locus, which is known to be directly associated with sepsis and the promotion of bacterial growth in the bloodstream [10,22] (Figure 2, Figure 4B). By integrating resistance and virulence profiles (Figure 5B, 5C) of the sequenced strains, we identified strains from ST101, ST11, ST147, ST16, ST2096, ST231, ST307, ST751, ST45, and ST383 as carrying a dual resistance/virulence risk (MDR-hv), characterized by a resistance score ≥1 and a virulence score ≥3 (Figure 2, Figure 5D). The largest MDR-hv clone was ST2096, previously reported in a single-hospital study in Jeddah [21] (Table 1, Figure 5D). We further examined the clinical outcomes of infections caused by the strains harbouring both resistance and virulence genes, focusing on in-hospital mortality and sepsis incidence in a subset of patients. The results showed a significantly higher mortality and sepsis rate among patients carrying ST231 (Supplementary Results, Figure S6A, S6B). Mortality rates associated with the carriage of the ybt gene, and both iuc and ybt loci were higher compared to the rest of the population (Supplementary Results, Figure S6A, S6B). Moreover, the presence of carbapenemase genes was associated with a significantly higher mortality rate compared to isolates lacking these genes (Figure S6C), highlighting the clinical significance of these genes.
Sharing of IncL, IncA/C and IncH plasmids encoding virulence and resistance genes between major clones.
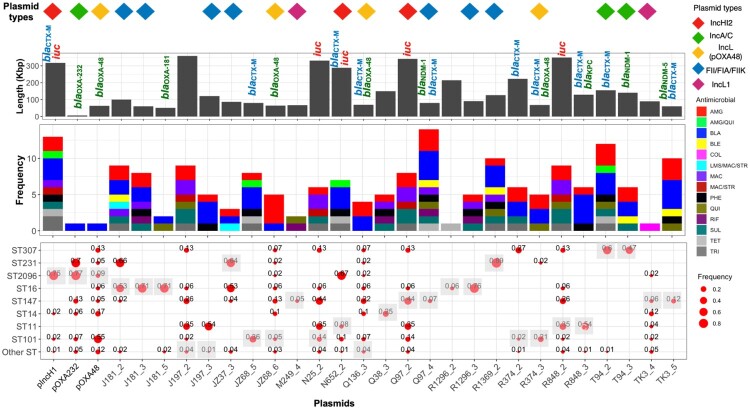
We analyzed the resistance-associated plasmids in the population using a hybrid sequencing approach on selected strains (see Methods). A total of 26 plasmid fragments with unique resistance gene profiles were identified from the isolates selected for third-generation sequencing (see Methods) (Table S4, Figure 6). We examined these plasmids, along with three resistance plasmids from ST2096 (IncHI, IncA/C (pOXA232) and IncL (pOXA-48)) that were isolated from ST2096 isolates in a single-hospital study [18] (Figure 6). Among the 26 plasmids, 19 were found in more than one clone, as indicated by higher coverage (i.e. >90% mapping of short reads to the reference plasmid sequence) (Figure 6). The screening of shared plasmids identified three types associated with carbapenemase, ESBL, or virulence genes: a non-conjugative IncA/C plasmid (6 kb) carrying the blaOXA-232 gene, and two conjugative plasmids (i.e. containing transfer (tra) genes) (Figure 2). These included the IncL (pOXA-48) plasmid with the blaOXA-48 gene [23], and the IncHI2 conjugative mosaic plasmid, which has a broad phylogenetic distribution and harbours both resistance genes and a pathogenicity island with the iuc locus and iutA genes (Figure 6, Table S4 and Figure S7). The virulence cassette in these plasmids was flanked by rmpA and rmpA2 genes, which regulate the hypervirulent mucoid phenotype. Both plasmids also contained the ESBL blaCTX-M genes in some isolates (Figure S7), though they exhibited variation in the configuration of resistance genes. These differences were evident in the gene-level alignment of the IncL (pOXA-48) plasmids from the ST101, ST1418 and ST2096 isolates. Although, all IncL plasmids had a resistance region with blaOXA-48 genes, the blaCTX-M-27 and aminoglycoside phosphorylase genes were variably present in the plasmids (Figure S7 and S8). The IncL plasmid from ST2096 isolates lacked a second resistance island. For the mosaic IncHI plasmid, which was previously characterized [18], gene arrangements varied among plasmids from ST101, ST11, ST147 and ST2096 clones, leading to different combinations of resistance genes (Figure S8). However, the order of virulence cassette genes remained consistent across plasmids from different clones (Figure S8). For ST2096, the IncHI plasmid is a distinguishing genomic element and underlies the higher virulence and resistance score of this clone (Figure 2), setting it apart from its sister clade, ST14. These findings suggest the backbones of the IncHI and IncL plasmids serve as a platform for the rapid horizontal transfer of resistance genes within and across clones. Beyond the recombination of resistance genes, our reconstructed phylodynamic tree for the major clones reveals that the horizontal transfer of the plasmids – harbouring both resistance and virulence factors – occur concurrently with clonal expansion, underscoring the complex evolutionary dynamics driving the spread of antimicrobial resistance (Figure S9).
Different patterns of the plasmid-mediated spread of the resistance plasmids.
Figure 6.
The retrieved plasmid fragments that contained a resistance gene. The top panel shows the length of the plasmid in base pairs. The bubble plot shows the frequency of plasmids across major clones. We mapped the short reads to the plasmid fragments and reported presence of plasmid if the mapping coverage was above 90%. The shaded boxes show the STs of the isolate from which the plasmid fragment was retrieved. The pIncH1, pOXA232 and pOXA48 fragments were from the ST2096 isolates, previously studied [18].
Coevolution analysis of the three plasmids, i.e. IncL (pOXA-48), IncA/C (pOXA232) and IncHI2, indicated distinctive modes of plasmid/core genome evolution. For the IncL plasmids, significant coevolution between plasmids and core genomes was observed (Mantel test p-value < 0.05), indicating that clones retained the same plasmids since their formation and the plasmid did not undergo genomic alterations during the clonal expansions. A tanglegram that connects plasmid-based, and core genome-based phylogenies confirmed the overall congruence between the trees but also showed evidence of plasmid horizontal transfer (Figure S10). In contrast, for IncHI plasmid, plasmid-core genome coevolution was not significant (Mantel test p-value > 0.05), with similar plasmids frequently found in distantly related isolates. This likely results from frequent recombination of resistance genes occurring concurrently with clonal expansion, conferring plasmids distinctive population dynamics than the core genome. The tanglegram for the plasmid-based and core-genome based also confirms the pervasive plasmid horizontal transfer for the IncHI plasmid (Figure S10). For the IncA/C (pOXA232) plasmid, highly similar plasmid variants (with a maximum SNP distance of five) were identified across different clones, indicating the horizontal transfer of a monomorphic plasmid between them.
Discussion
This study presents a large-scale genomic analysis of the epidemiology and evolution of MDR K. pneumoniae clones in the Kingdom of Saudi Arabia. Our systematic collection included a snapshot of clinical isolates from hospitalized patients for a large hospital network. Our results indicate the concurrent expansion of multiple clones across hospitals, with the dominance of a few clones, i.e. ST2096, ST147, and ST231, which commonly harbour Inc plasmids with multiple carbapenemase, ESBL, and virulence factor genes. Although previous studies reported the prevalence of some of these STs across the Middle East [13,21,24], no major study examined the dynamics of MDR clones at high resolution within a consistent framework. Our findings on the mixing of Saudi isolates with those of Middle Eastern and South Asian origins align with Saudi Arabia's role as a major religious tourism hub, hosting one of the largest annual human gatherings, with a significant number of visitors from these regions [25]. Moreover, Saudi Arabia is home to a substantial immigrant population from South Asia. This underscores the importance of human migration and movement in the dissemination of high-risk K. pneumoniae clones.
The evolution of MDR K. pneumoniae is characterized by the spread of clones at the hospital level through nosocomial transmission routes, followed by regional and global expansion. The epidemiological and clinical characteristics of carbapenem resistant K. pneumoniae clones have been reported in multicenter studies in different countries [6,26–29], revealing expansions of both endemic clones and imported clones. Although ST11 and ST258 with KL1/KL2 capsule types are commonly reported as the dominant multidrug-resistant hypervirulent K. pneumoniae strains globally, only 5% of the strains in our collection were ST11, and no ST258 cases were observed in our study cohort. The results indicate a distinctive pattern of K. pneumoniae epidemiology in the KSA, in which non-CG258 (non-KL1/KL2) strains are predominant. The ST147 and ST231 clones with KL10/KL64/KL51 capsule types are recognized as circulating clones in South and East Asia [9], but the KL64 harbouring ST2096 clone has not been widely reported as a global clone and mainly reported from the Middle East and South Asia [21,30–32]. However, the capsule switching and prevalence of KL64 capsule types were reported in a major ST11 clone in China [7,33,34], underscoring the increasing the clinical significance of the capsule type. Previously, an expanding ST2096 clone was reported in a single-hospital study over an 8-year period in Jeddah and was proposed to be an emerging clone, with pervasive transmission within the hospital wards and links to sepsis and adverse clinical outcomes [18]. Our study extends the epidemiology of the ST2096 beyond a single hospital and shows the nationwide dissemination of the clone. The success of ST2096 clone, compared with that of ancestral ST14, is attributable to the enhanced pathogenicity and resistance through the acquisition of MDR/hvKp IncH1 plasmid [18]. The acquisition of hybrid conjugative virulence plasmids played a similar role in the emergence of novel ST11 hv-MDR strains in K. pneumoniae from carbapenem resistant ancestral strains in China [7], and our study highlights the importance of this plasmid as a major risk factor in K. pneumoniae epidemiology.
Hybrid assembly has been increasingly employed to explore the coevolution of AMR-linked plasmids and bacterial core genomes. A recent multicenter study on clinical K. pneumoniae isolates demonstrated a correlation between carbapenemase-carrying plasmids and high-risk clones, with these plasmids accumulating through clonal expansion and spreading across hospital networks in Europe [35,36]. Our findings on the IncH1 and IncL plasmid types in a different geographical setting, align with these patterns and exhibit similar trends. The backbones of these plasmids, which act as platforms for resistance gene insertion, have been identified as facilitators of antimicrobial resistance gene spread in a study conducted on blood stream infections in a single geographical region in the UK [37]. Our study indicates that these findings may be generalized to the dissemination of resistance genes, occurring simultaneously with the expansion of high-risk clones, across hospitals in a broader geographical region. Extending the scope of plasmidome analysis in future studies to include larger numbers of isolates, including susceptible ones, could shed more light on the dynamics of resistance gene acquisition by plasmids during clonal expansion. This expansion will allow for the direct estimation of key epidemiological parameters, such as the rate of resistance gene acquisition and plasmid transmission, in multicenter regional studies.
The emergence of multidrug-resistant K. pneumoniae strains has been reported in clinical [8], urban [9], agricultural settings [10] and as a worldwide phenomenon [11–13]. To fully characterize the reservoirs and transmission dynamics of these endemic clones, unbiased and comprehensive surveillance is warranted. Despite the depth and breadth of our sampling, several limitations should be noted. First, our collection included only a few environmental samples from food. The limited sampling from food retail outlets prevented us from accurately assessing population diversity in non-human sources and identifying potential epidemiological links to human sources. Such links with environmental sources have been reported for K. pneumoniae strains in a previous study [38], suggesting spill-over of samples from hospitals into linked environmental sites [39]. Although the sample size from food was small in our study, we reported the sharing of ST101 strains between humans and food. This finding points to epidemiological transmission routes, which may become increasingly detectable with broader and more in-depth sampling. Furthermore, our collection predominantly consisted of multidrug-resistant strains and therefore did not include nonresistant or nonpathogenic strains. These strains might become pathogenic and multidrug resistant through plasmid acquisition. While we integrated phenotypic and genotypic data to elucidate the impact of known resistance genes on observed phenotypes, the ability to identify novel resistance determinants through GWAS was constrained by the prevalence of MDR isolates in the collection, and not all phenotypic variations could be attributed to known resistance genes. The unexplained resistance phenotype may originate from other factors, including non-genetic mechanisms such as regulatory mechanisms, epistatic interactions between resistance genes, the presence of pseudo-resistance genes, and testing errors [40–42]. Finally, our study covered 16 months and therefore did not reveal long-term clonal dynamics. In future studies, sampling could be extended to include not only environmental strains but also susceptible nonpathogenic strains in a purpose-designed sampling framework over an extended period to fully reveal the details of K. pneumoniae transmission routes in the KSA.
This study demonstrates that precision epidemiological analysis through whole-genome sequencing effectively elucidates transmission and population dynamics of K. pneumoniae at the regional level. By extending this study to encompass collections from geographically diverse healthcare centres across the Middle East and North Africa (MENA) regions – areas that are currently underrepresented in genomic databases – targeted interventional strategies based on the specificities of the region can be designed to control K. pneumoniae infections.
Supplementary Material
Acknowledgment
We would like to thank Liliane Okdah for the help with the experimental protocols.
.
Supplementary material (Methods)
Short-read sequencing
K. pneumoniae isolates were cultured overnight at 37°C in Luria–Bertani (LB) broth. Genomic DNA (gDNA) was extracted using a DNeasy Blood and Tissue Kit according to the manufacturer’s instructions (QIAGEN, Hilden, Germany). The quality of the gDNA was determined using a DS-11 DNA quantification spectrophotometer from Denovix (Denovix, US). The gDNA quantity was determined using a fluorometric method with analysis performed on a Qubit 4.0 fluorometer using a high-sensitivity double-stranded DNA assay kit (Thermo Fisher Scientific, US). We then prepared genomic libraries for the 361 isolates using the MGIEasy Fast FS DNA Library Prep in accordance with the instructions in the user manual provided by BGI Technology (BGI Technology, China). An enzymatic fragmentation protocol was used. The libraries were then denatured and circularized. Whole genome sequencing (WGS) was performed on the DNBSEQ-G400 platform (BGI Technology, China) using the 2150 bp paired-end read protocol.
The short reads were subjected to quality control with the fastqc package in R (v0.1.3). We assembled the genomes with a de novo assembly pipeline using Unicycler (v0.5.0) (https://github.com/rrwick/Unicycler#install-from-source) with default parameters [43]. Assemblies were processed, and contigs shorter than 200 bp were excluded from the analysis with Unicycler. We profiled and characterized the strains with Kleborate (v2.3.2) to determine the sequence and capsule types and the resistome and virulence gene profile of the collection [44]. The risk assessment scores for resistance and virulence levels were also obtained for Kleborate. The definition of hypervirulence is based on the presence of highly diagnostic gene markers, i.e. ybt, iuc, iro, rmpA, rmpA2 and peg-344 [10]. Virulence scores, graded on a scale of 0 to 5, are computed by the detection of a subset of these genetic markers, with yersiniabactin (ybt) having the smallest risk, followed by colibactin (clb), and then aerobactin (iuc). The presence of the loci ybt, ybt+clb, iuc, ybt+iuc and ybt+clb+iuc corresponded to virulence scores of 1 to 5. The absence of all these loci corresponded to virulence score of zero. Hypervirulent clones, i.e. ST23, ST86, ST65 and ST231 were shown to have a virulence score ≥ 3, owing to the prevalence of the iuc locus [44]. Resistance scores, ranging from 0 to 3, were determined by the presence of key antimicrobial resistance genes, with ESBL being the least resistant, followed by carbapenemase, and then carbapenemase combined with colistin resistance determinants. The convergence of resistance and virulence occurs when both virulence and resistance scores were equal to or greater than 3 and 1, respectively [44].
To complement the virulence genes that were detected by Kleborate, AMRFinderPlus [45] pipeline was used, with identity and coverage cut-off values of 50% and 70%, respectively. A one-sided proportion test in R with a significance level of 0.05 was used to identify genes differentially distributed in major clones and the background population.
We annotated de novo assemblies with Prokka (v1.14.5) [8] and fed the annotated assemblies into Panaroo to reconstruct the pangenome [9]. To generate the phylogenetic tree for our dataset, we initially aligned the short-read sequences against the genome of the reference strain K. pneumoniae Ecl8, an ST375/K2 hypervirulent clone of significant epidemiological importance in causing severe community-acquired infections (accession number: PRJEB401) [46]. This alignment was conducted using the Snippy pipeline (accessible at https://github.com/tseemann/snippy) with default settings. We then calculated the pairwise SNP distances among the core genome alignments and used the FastTree package (2.1) [47] with default parameters to construct an approximately maximum likelihood phylogenetic tree. To measure the genetic diversity within each province, we computed the average SNP distance for each province by taking the average of the number of SNP differences between core genomes of all possible pairs of isolates from the same province.
Transmission and phylodynamic analysis
We combined our genomes with those sequenced in a single hospital study in Jeddah [21], to attain a larger collection size. For the five prevalent sequence types (STs), i.e. STs with more than 1% of the population – ST14, ST101, ST231, ST147, and ST2096 – within the two collections combined, we conducted phylodynamic analysis to compute key epidemiological parameters of each clone and reconstruct dated phylodynamic trees.
We selected the strain with the best assembly statistics, namely, the highest N50, for each clone. We merged the contigs of these selected strains to form local reference genomes. By aligning the short reads of each strain in the clone to this reference genome, we derived a core genome SNP alignment. This alignment was fed into Gubbins (v.3.3.1) with five iterations to eliminate hypervariable regions. The SNP counts for the major clones are detailed in Table 1.
To estimate the ancestral locations of the major clones within the SNP clusters, we performed phylogeographic diffusion analysis in discrete space using BEAST (v2) [48]. We used the geographical location (city of isolation) as a discrete state of each taxon and employed a constant population size model with uniform priors on the clock rate. We also utilized a symmetric model with a uniform prior distribution for the discrete trait substitution model for the diffusion of the clone in space. Convergence of the Markov chain Monte Carlo (MCMC) chains was assessed by ensuring that the effective sample size (ESS) for key parameters exceeded 150. The spatial trajectories derived from the dated Bayesian tree were visualized using www.spreadviz.org website.
For a thorough examination of population size changes over time, we employed a nonparametric growth model to sample population sizes on the dated phylogenetic tree for the ST2096, ST231 and ST147 clones. We therefore used the skygrowth.mcmc function and visualized the results with the plot function within the skygrowth package (v0.3.1) [49].
To confirm the findings from the phylodynamic analysis, we reconstructed genealogies of sampled sequences assuming that ancestors and descendants were sampled together for three major clones, i.e. ST147, ST231 and ST2096. To this end, we used R package adegenet (V.1.3-1) [50], which maximizes the likelihood of the networks based on pairwise SNP distances in their core genomes and the isolation dates. To calculate the distances, we took the SNP alignment from the BEAST analysis, in which hypervariable sites were removed, for each clone and fed the distance information along with the collection dates of isolates into the seqTrack function in the adegenet package. To define transmission networks, we used the SNP threshold of 20 SNPs, which is based on a recent large-scale analysis of K. pneumoniae within and across hospitals [36,51]. The output networks were visualized with the igraph library in R.
Contextualization of the samples
We contextualized our strains using the Pathogen Detection database available at https://www.ncbi.nlm.nih.gov/pathogens/. We first retrieved epidemiological SNP clusters including the strains from the Pathogen Detection database on 06/08/2024. These SNP clusters contained all strains with pairwise SNP distances of up to 50 SNPs. The clustering process was conducted by the pipeline within the Pathogen Detection portal (https://www.ncbi.nlm.nih.gov/pathogens/pathogens_help/#references). This high-throughput automated pipeline employs a combined kmer approach and alignment to define clusters, with a fixed cut-off of 50 SNPs that cannot be customized. Subsequently, we extracted the trees for selected clusters from the portal and visualized them with the ggtree package (v3.8.2) in R [52].
To ensure the generalizability of the findings on SNP clusters, we also conducted sensitivity analysis. We, therefore, screened SNP distance thresholds for defining the clusters from 0 to 50 and counted the global strains that fell within the clusters containing Saudi genomes at different SNP thresholds. We used the SNP threshold of 20 SNPs, as the threshold for between hospital transmissions, as detailed above.
Plasmidome analysis and long-read sequencing
We performed third-generation sequencing on representative samples from the population to resolve the content of AMR-linked plasmids in the collection. Since the plasmid content of the ST2096 strains had been previously characterized [18], we selected samples from other sequence types (STs). To curate a representative collection, we selected twenty bloodstream isolates with the highest resistance and virulence scores, representing different clades and exhibiting distinctive patterns of resistance genes. These isolates were mostly from major ST clones, defined as those with more than ten representative genomes in the collection (one isolate was excluded during sequencing). The selected isolates included 3 from ST101, 2 from ST11, 2 from ST14, 3 from ST147, 2 from ST16, 2 from ST231, 1 from ST307, and the remaining from other STs.
To prepare sequenced libraries for long-read sequencing, we employed 96-plex Rapid Barcoding Kits for multiplexing. These libraries were then loaded into MinION flow cells (Oxford Nanopore Technologies) and subjected to a 48-hour run following the manufacturer's protocol. Hybrid assembly was conducted using Unicycler, employing the conservative option. Subsequently, the assembled contigs were screened for the presence of full copies of the origin of replications, virulence factor genes, and antimicrobial resistance genes using BLAST in conjunction with the abovementioned databases. The assembled genomes were visualized and checked with Bandage (v0.9.0) [53]. We extracted plasmid fragments containing virulence factors and resistance genes. Plasmids were visualized and annotated with the built-in tools in the Proksee portal (www.proksee.ca) [54]. We identified plasmid replicons and resistance genes via BLAST searches of the extracted plasmid fragments on PlasmidFinder [55] databases and AMRFinderPlus [45], respectively. We also typed the plasmid fragments with PlasmidMLST (https://pubmlst.org/organisms/plasmid-mlst) database [56]. We mapped the short reads of the other strains against these extracted plasmid fragments and used the mapping coverage of >90% to confirm the presence of these plasmids in the rest of the population.
Three types of plasmids were found recurring in plasmid fragments from isolates belonging to different STs: the IncH mosaic plasmid with ESBL blaCTX-M-15 and virulence genes, the IncA/C plasmid with blaOXA-232, and IncL (pOXA-48) plasmids, which carried blaOXA-48 gene. To understand the genomic rearrangement within plasmid structures, we conducted gene-level alignment and visualized the alignment using the pyGenomeViz (v1.0.0) package in Python (https://github.com/moshi4/pyGenomeViz) for IncH and IncL plasmids, which showed variations in gene content between the clones. To decipher the coevolution between the plasmid and the core genome, we first extracted distance matrices for the plasmids isolated in which the plasmid was detected (i.e. mapping > 90%) and the core genomes for the same isolates for the three plasmids: IncL, and IncH (the IncA/C plasmid with blaOXA-232 were highly similar in the isolates and therefore phylogenetic tree could not be reconstructed). We then conducted a Mantel test in R on the distance matrices for plasmids and core genomes to assess the significance of coevolution. We generated neighbor-joining trees for the plasmids and the core genomes from the distance matrices and visualized the tanglegram (i.e. a diagram for comparing two phylogenetic trees) using Dendroscope (v3.8.10) [57].
Integration of resistome and virulome data with phenotypic and clinical metadata
We integrated the genetic profile of resistance determinants obtained from Kleborate with the phenotypic resistance data using the odds ratio function from the Epitools package (v0.5.10.1) in R. Moreover, we examined the significance of associations between the accessory genes and SNPs and the resistance phenotype while considering population structure by employing Scoary (1.6.16) [58] on the Panaroo output and the SNPs identified via post-read mapping to the reference genome, respectively.. Specifically, we focused on pairwise p values (both worst and best p values) that adjusted for the confounding effect of population structure (lineage effect). The associations were assessed using Scoary (accessible at www.github.com/AdmiralenOla/Scoary). To visualize the tree and associated metadata, we utilized the ggtree package (v3.8.2) in R [52].
We obtained clinical metadata on the reported mortality, cause of death, and date of in-hospital death for all patients with bloodstream infections and those with urinary tract infections that subsequently led to bacteremia. We computed the mortality rates at days 7, 14, 21 and 30, considering the period between sample isolation and reported death, for patients infected by each major ST. We also computed the odds ratio of death for infection by the major STs and strains with different profiles of resistance determinants using the Epitools package.
Broth microdilution assays
Broth microdilution (BMD) assays were employed as a reference antimicrobial susceptibility testing (AST) method to confirm the results from the automated Vitek®2 platform. This was also performed to verify colistin resistance in isolates with known resistance determinants. The validation process included a subset of 58 isolates identified as non-susceptible (i.e. intermediate or resistant) to all tested drugs by the Vitek®2 platform. Three susceptible strains were used as controls. To maximize resource efficiency and ensure the generalizability of the findings, we tested antimicrobials from three key classes: cephalosporins (ceftazidime), fluoroquinolones (ciprofloxacin), and aminoglycosides (gentamicin). For colistin resistance testing, we included 52 isolates carrying mcr genes and those with truncated resistance genes (mgrB and pmr) (truncations greater than 50% as determined by Kleborate) along with three control strains.
BMD was performed according to Clinical & Laboratory Standards Institute (CLSI) guidelines using 96-well plates (Corning®) [59]. Stock concentrations of gentamicin, ciprofloxacin, ceftazidime, and colistin were prepared at 5 mg/mL using recommended solvents as described in CLSI guidelines [42]. For each antimicrobial, a twofold serial dilution was carried out by transferring equal volumes of antimicrobial into Muller Hinton Broth (MHB). The dilutions tested were as follows: gentamicin, ceftazidime, and colistin: 128 µg/mL – 0.25 µg/mL; ciprofloxacin: 32 µg/mL – 0.06 µg/mL. Isolates were streaked onto agar plates and incubated overnight at 37°C to achieve single colonies. Single colonies were then resuspended in saline solution (0.45% NaCl) to a turbidity of 0.5 McFarland units, followed by dilution in MHB to achieve 510⁵ CFU/mL per well. The plates were incubated at 37°C for 16 h. Results were interpreted visually according to CLSI guidelines [59].
The following metrics were measured for the agreement between the two methods:
Categorical Agreement (CA): The percentage of times Vitek®2 and BMD agreed on the susceptibility category (e.g. susceptible, intermediate, resistant).
Very Major Errors (VME): Instances where Vitek®2 incorrectly categorized a resistant strain as susceptible.
Major Errors (ME): Instances where Vitek®2 incorrectly categorized a susceptible strain as resistant.
Minor Errors (MinE): Instances where Vitek®2 and BMD differed between susceptible and intermediate or between intermediate and resistant.
The results, detailed in Table S2, confirm the congruence between the methods. For the isolates with colistin truncation mutations and mcr genes, all were found to have MIC values greater than the CLSI breakpoint for resistance (i.e. ≥ 4 µg/mL) (Supplemental Table S1).
Funding Statement
DM, MM, JH, DA, UA, and GZ were supported by KAUST faculty baseline fund (BAS/1/1108-01-01). AP is supported by KAUST baseline (BAS/1/1020-01-01). AP, DM, MM, JH, DA, UA, and GZ were also supported by FCC/1/5932-01-03 from KAUST Center of Excellence for Smart Health. The authors extend their appreciation to the Deputyship for Research and Innovation, “Ministry of Education” in Saudi Arabia for funding this research (IFKSUOR3-478).
Author contribution
J.H. and A.Y.A. contributed equally to this work. J.H. and A.Y.A., U.H.A., M.M. and G.Z. conducted the experimental and computational research. D.A., M.B., A.A.A., A.A.R., S.B., D.B., A.N.A., Z.A.A., S.M.A., P.-Y.H., M.A., and W.A.S. provided resources. A.P., W.A.S. and D.M. conceived and supervised the study. All authors have read and approved the manuscript.
Disclosure statement
No potential conflict of interest was reported by the author(s).
References
- 1.Dong N, Yang X, Chan EW, et al. Klebsiella species: taxonomy, hypervirulence and multidrug resistance. EBioMedicine. May 2022;79:103998. doi: 10.1016/j.ebiom.2022.103998 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Yang X, Dong N, Chan EW, et al. Carbapenem resistance-encoding and virulence-encoding conjugative plasmids in Klebsiella pneumoniae. Trends Microbiol. Jan 2021;29(1):65–83. doi: 10.1016/j.tim.2020.04.012 [DOI] [PubMed] [Google Scholar]
- 3.Tacconelli E, Carrara E, Savoldi A, et al. Discovery, research, and development of new antibiotics: the WHO priority list of antibiotic-resistant bacteria and tuberculosis. Lancet Infect Dis. Mar 2018;18(3):318–327. doi: 10.1016/S1473-3099(17)30753-3 [DOI] [PubMed] [Google Scholar]
- 4.Holt KE, Wertheim H, Zadoks RN, et al. Genomic analysis of diversity, population structure, virulence, and antimicrobial resistance in Klebsiella pneumoniae, an urgent threat to public health. Proc Natl Acad Sci U S A. Jul 7 2015;112(27):E3574–E3581. doi: 10.1073/pnas.1501049112 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Wyres KL, Lam MMC, Holt KE.. Population genomics of Klebsiella pneumoniae. Nat Rev Microbiol. Jun 2020;18(6):344–359. doi: 10.1038/s41579-019-0315-1 [DOI] [PubMed] [Google Scholar]
- 6.Teo JQ, Tang CY, Tan SH, et al. Genomic surveillance of carbapenem-resistant Klebsiella pneumoniae from a major public health hospital in Singapore. Microbiol Spectr. Oct 26 2022;10(5):e0095722. doi: 10.1128/spectrum.00957-22 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Liao WJ, Liu Y, Zhang W.. “Virulence evolution, molecular mechanisms of resistance and prevalence of ST11 carbapenem-resistant in China: A review over the last 10 years,” (in English). J Glob Antimicrob Re. Dec 2020;23:174–180. doi: 10.1016/j.jgar.2020.09.004 [DOI] [PubMed] [Google Scholar]
- 8.Lan P, Jiang Y, Zhou J, et al. A global perspective on the convergence of hypervirulence and carbapenem resistance in Klebsiella pneumoniae. J Glob Antimicrob Resist. Jun 2021;25:26–34. doi: 10.1016/j.jgar.2021.02.020 [DOI] [PubMed] [Google Scholar]
- 9.Heng H, Yang X, Ye L, et al. Global genomic profiling of Klebsiella pneumoniae: a spatio-temporal population structure analysis. Int J Antimicrob Agents. Feb 2024;63(2):107055. doi: 10.1016/j.ijantimicag.2023.107055 [DOI] [PubMed] [Google Scholar]
- 10.Russo TA, Olson R, Fang CT, et al. Identification of biomarkers for differentiation of hypervirulent Klebsiella pneumoniae from classical K. pneumoniae. J Clin Microbiol. Sep 2018;56(9). doi: 10.1128/JCM.00776-18 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Abd El Ghany M, Sharaf H, Al-Agamy MH, et al. Genomic characterization of NDM-1 and 5, and OXA-181 carbapenemases in uropathogenic Escherichia coli isolates from Riyadh, Saudi arabia. PLoS One. 2018;13(8):e0201613. doi: 10.1371/journal.pone.0201613 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Alotaibi FE, Bukhari EE, Al-Mohizea MM, et al. Emergence of carbapenem-resistant enterobacteriaceae isolated from patients in a university hospital in Saudi Arabia. Epidemiology, clinical profiles and outcomes. J Infect Public Health. Sep-Oct 2017;10(5):667–673. doi: 10.1016/j.jiph.2017.05.004 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Mouftah SF, Pal T, Higgins PG, et al. Diversity of carbapenem-resistant Klebsiella pneumoniae ST14 and emergence of a subgroup with KL64 capsular locus in the Arabian Peninsula. Eur J Clin Microbiol Infect Dis. Dec 2 2021. doi: 10.1007/s10096-021-04384-2 [DOI] [PubMed] [Google Scholar]
- 14.Zaman TU, Alrodayyan M, Albladi M, et al. Clonal diversity and genetic profiling of antibiotic resistance among multidrug/carbapenem-resistant Klebsiella pneumoniae isolates from a tertiary care hospital in Saudi Arabia. BMC Infect Dis. May 3 2018;18(1):205. doi: 10.1186/s12879-018-3114-9 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Memish ZA, Assiri A, Almasri M, et al. Molecular characterization of carbapenemase production among gram-negative bacteria in Saudi Arabia. Microb Drug Resist. Jun 2015;21(3):307–314. doi: 10.1089/mdr.2014.0121 [DOI] [PubMed] [Google Scholar]
- 16.Magiorakos AP, Srinivasan A, Carey RB, et al. Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: an international expert proposal for interim standard definitions for acquired resistance. Clin Microbiol Infect. Mar 2012;18(3):268–281. doi: 10.1111/j.1469-0691.2011.03570.x [DOI] [PubMed] [Google Scholar]
- 17.Toh BEW, Bokhari O, Kutbi A, Haroon MF, et al. Varying occurrence of extended-spectrum beta-lactamase bacteria among three produce types. J Food Safety. Feb 2018;38(1):e12373. doi:ARTN e12373 10.1111/jfs.12373. [DOI] [Google Scholar]
- 18.Hala S, Malaikah M, Huang J, et al. The emergence of highly resistant and hypervirulent Klebsiella pneumoniae CC14 clone in a tertiary hospital over 8 years. Genome Med. Apr 18 2024;16(1):58. doi: 10.1186/s13073-024-01332-5 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.James S. Lewis II P, FIDSA. CLSI M100 performance standards for antimicrobial susceptibility testing. 34th ed. USA: Clinical and Laboratory Standards Institute; 2024. [Google Scholar]
- 20.Peirano G, Chen L, Kreiswirth BN, et al. Emerging antimicrobial-resistant high-risk Klebsiella pneumoniae clones ST307 and ST147. Antimicrob Agents Ch. Sep 21 2020;64(10). doi: 10.1128/AAC.01148-20 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Hala S, Malaikah M, Huang J, et al. The emergence of highly resistant and hypervirulent Klebsiella pneumoniae CC14 clone in a tertiary hospital over 8 years. 2024. doi: 10.2139/ssrn.4574760 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Russo TA, Olson R, MacDonald U, et al. Aerobactin, but not yersiniabactin, salmochelin, or enterobactin, enables the growth/survival of hypervirulent (hypermucoviscous) Klebsiella pneumoniae ex vivo and in vivo. Infect Immun. Aug 2015;83(8):3325–3333. doi: 10.1128/IAI.00430-15 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Poirel L, Bonnin RA, Nordmann P.. Genetic features of the widespread plasmid coding for the carbapenemase OXA-48. Antimicrob Agents Ch. Jan 2012;56(1):559–562. doi: 10.1128/AAC.05289-11 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Sonnevend A, Abdulrazzaq N, Ghazawi A, et al. The first nationwide surveillance of carbapenem-resistant Enterobacterales in the United Arab Emirates – increased association of Klebsiella pneumoniae CC14 clone with Emirati patients. Int J Infect Dis. Jul 2022;120:103–112. doi: 10.1016/j.ijid.2022.04.034 [DOI] [PubMed] [Google Scholar]
- 25.Haseeb A, Saleem Z, Faidah HS, et al. Threat of Antimicrobial Resistance among Pilgrims with Infectious Diseases during Hajj: Lessons Learnt from COVID-19 Pandemic. Antibiotics. Antibiot-Basel. Aug 2023;12(8):1299. doi: 10.3390/antibiotics12081299. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Moradigaravand D, Martin V, Peacock SJ, et al. Evolution and epidemiology of multidrug-resistant Klebsiella pneumoniae in the United Kingdom and Ireland. mBio. Feb 21 2017;8(1). doi: 10.1128/mBio.01976-16 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Li C, Jiang X, Yang T, et al. Genomic epidemiology of carbapenemase-producing Klebsiella pneumoniae in China. Genom Proteomic Bioin. Dec 2022;20(6):1154–1167. doi: 10.1016/j.gpb.2022.02.005 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.David S, Reuter S, Harris SR, et al. Epidemic of carbapenem-resistant Klebsiella pneumoniae in Europe is driven by nosocomial spread. Nat Microbiol. Nov 2019;4(11):1919–1929. doi: 10.1038/s41564-019-0492-8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Wyres KL, Nguyen TNT, Lam MMC, et al. Genomic surveillance for hypervirulence and multi-drug resistance in invasive Klebsiella pneumoniae from South and Southeast Asia. Genome Med. Jan 16 2020;12(1):11. doi: 10.1186/s13073-019-0706-y [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Shankar C, Vasudevan K, Jacob JJ, et al. Hybrid plasmids encoding antimicrobial resistance and virulence traits among hypervirulent Klebsiella pneumoniae ST2096 in India. Front Cell Infect Microbiol. 2022;12:875116. doi: 10.3389/fcimb.2022.875116 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Hosbul T, Guney-Kaya K, Guney M, et al. Carbapenem and colistin resistant Klebsiella pneumoniae ST14 and ST2096 dominated in two hospitals in Turkey. Clin Lab. Sep 1 2021;67(9). doi: 10.7754/Clin.Lab.2021.201226 [DOI] [PubMed] [Google Scholar]
- 32.Urooj M, Shoukat M, Imran M, et al. Emergence of hypermucoviscous colistin-resistant high-risk convergent Klebsiella pneumoniae ST-2096 clone from Pakistan. Future Microbiol. Sep 2022;17:989–1000. doi: 10.2217/fmb-2021-0292 [DOI] [PubMed] [Google Scholar]
- 33.Zhou K, Xue C-X, Xu T, et al. A point mutation in recC associated with subclonal replacement of carbapenem-resistant Klebsiella pneumoniae ST11 in China. Nat Commun. Apr 28 2023;14(1):2464. doi: 10.1038/s41467-023-38061-z [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Zhou K, Xiao TT, David S, et al. “Novel subclone of carbapenem-resistant sequence type 11 with enhanced virulence and transmissibility, China,” (in English). Emerg Infect Dis. Feb 2020;26(2):289–297. doi: 10.3201/eid2602.190594 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.David S, Cohen V, Reuter S, et al. Integrated chromosomal and plasmid sequence analyses reveal diverse modes of carbapenemase gene spread among Klebsiella pneumoniae. Proc Natl Acad Sci U S A. Oct 6 2020;117(40):25043–25054. doi: 10.1073/pnas.2003407117 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.David S, Reuter S, Harris SR, et al. “Epidemic of carbapenem-resistant in Europe is driven by nosocomial spread,” (in English). Nat Microbiol. Nov 2019;4(11):1919–1929. doi: 10.1038/s41564-019-0492-8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Lipworth S, Matlock W, Shaw L, et al. Author correction: the plasmidome associated with gram-negative bloodstream infections: a large-scale observational study using complete plasmid assemblies. Nat Commun. Apr 9 2024;15(1):3060. doi: 10.1038/s41467-024-47494-z [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Runcharoen C, Moradigaravand D, Blane B, et al. Whole genome sequencing reveals high-resolution epidemiological links between clinical and environmental Klebsiella pneumoniae. Genome Med. Jan 24 2017;9(1):6. doi: 10.1186/s13073-017-0397-1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Ludden C, Moradigaravand D, Jamrozy D, et al. A one health study of the genetic relatedness of Klebsiella pneumoniae and their mobile elements in the East of England. Clin Infect Dis. Jan 2 2020;70(2):219–226. doi: 10.1093/cid/ciz174 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Davis MA, Besser TE, Orfe LH, et al. Genotypic-phenotypic discrepancies between antibiotic resistance characteristics of Escherichia coli isolates from calves in management settings with high and low antibiotic use. Appl Environ Microbiol. May 2011;77(10):3293–3299. doi: 10.1128/AEM.02588-10 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Deekshit VK, Srikumar S.. ‘To be, or not to be'-The dilemma of ‘silent’ antimicrobial resistance genes in bacteria. J Appl Microbiol. Nov 2022;133(5):2902–2914. doi: 10.1111/jam.15738 [DOI] [PubMed] [Google Scholar]
- 42.Chen Z, Qlu S, Wang Y, et al. Coexistence of blaNDM-1 with the prevalent blaOXA23 and blaIMP in pan-drug resistant Acinetobacter baumannii isolates in China. Clin Infect Dis. Mar 1 2011;52(5):692–693. doi: 10.1093/cid/ciq231 [DOI] [PubMed] [Google Scholar]
- 43.Wick RR, Judd LM, Gorrie CL, et al. Unicycler: resolving bacterial genome assemblies from short and long sequencing reads. PLoS Comput Biol. Jun 2017;13(6):e1005595. doi: 10.1371/journal.pcbi.1005595 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Lam MMC, Wick RR, Watts SC, et al. A genomic surveillance framework and genotyping tool for Klebsiella pneumoniae and its related species complex. Nat Commun. Jul 7 2021;12(1):4188. doi: 10.1038/s41467-021-24448-3 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Feldgarden M, Brover V, Gonzalez-Escalona N, et al. AMRFinderplus and the reference gene catalog facilitate examination of the genomic links among antimicrobial resistance, stress response, and virulence. Sci Rep. Jun 16 2021;11(1):12728. doi: 10.1038/s41598-021-91456-0 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Fookes M, Yu J, De Majumdar S, et al. Genome sequence of Klebsiella pneumoniae Ecl8, a reference strain for targeted genetic manipulation. Genome Announc. Jan 2013;1(1). doi: 10.1128/genomeA.00027-12 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Price MN, Dehal PS, Arkin AP.. Fasttree: computing large minimum evolution trees with profiles instead of a distance matrix. Mol Biol Evol. Jul 2009;26(7):1641–1650. doi: 10.1093/molbev/msp077 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Bouckaert R, Vaughan TG, Barido-Sottani J, et al. BEAST 2.5: An advanced software platform for Bayesian evolutionary analysis. PLoS Comput Biol. Apr 2019;15(4):e1006650. doi: 10.1371/journal.pcbi.1006650 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Volz EM, Didelot X.. Modeling the growth and decline of pathogen effective population size provides insight into epidemic dynamics and drivers of antimicrobial resistance. Syst Biol. Jul 1 2018;67(4):719–728. doi: 10.1093/sysbio/syy007 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Jombart T, Ahmed I.. Adegenet 1.3-1: new tools for the analysis of genome-wide SNP data. Bioinformatics. Nov 1 2011;27(21):3070–3071. doi: 10.1093/bioinformatics/btr521 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Thorpe HA, Booton R, Kallonen T, et al. “A large-scale genomic snapshot of Klebsiella spp. isolates in Northern Italy reveals limited transmission between clinical and non-clinical settings,” (in English). Nat Microbiol. Dec 2022;7(12):2054–20-+. doi: 10.1038/s41564-022-01263-0 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Yu G, Lam TT, Zhu H, et al. Two methods for mapping and visualizing associated data on phylogeny using ggtree. Mol Biol Evol. Dec 1 2018;35(12):3041–3043. doi: 10.1093/molbev/msy194 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Wick RR, Schultz MB, Zobel J, et al. Bandage: interactive visualization of de novo genome assemblies. Bioinformatics. Oct 15 2015;31(20):3350–3352. doi: 10.1093/bioinformatics/btv383 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Grant JR, Enns E, Marinier E, et al. Proksee: in-depth characterization and visualization of bacterial genomes. Nucleic Acids Res. Jul 5 2023;51(W1):W484–W492. doi: 10.1093/nar/gkad326 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Carattoli A, Hasman H.. Plasmidfinder and in silico pMLST: identification and typing of plasmid replicons in Whole-Genome Sequencing (WGS). Methods Mol Biol. 2020;2075:285–294. doi: 10.1007/978-1-4939-9877-7_20 [DOI] [PubMed] [Google Scholar]
- 56.Jolley KA, Bray JE, Maiden MCJ.. Open-access bacterial population genomics: BIGSdb software, the PubMLST.org website and their applications. Wellcome Open Res. 2018;3:124. doi: 10.12688/wellcomeopenres.14826.1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Huson DH, Richter DC, Rausch C, et al. Dendroscope: An interactive viewer for large phylogenetic trees. BMC Bioinformatics. Nov 22 2007;8:460. doi: 10.1186/1471-2105-8-460 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Brynildsrud O, Bohlin J, Scheffer L, et al. Rapid scoring of genes in microbial pan-genome-wide association studies with Scoary. Genome Biol. Nov 25 2016;17(1):238. doi: 10.1186/s13059-016-1108-8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.“CLSI. Performance Standards for Antimicrobial Susceptibility Testing” . CLSI supplement M100. Clinical and Laboratory Standards Institute, 2024.
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
Genomic data collected in this study were deposited in the European Nucleotide Archive (ENA) under the study accession number PRJEB66182. Moreover, the assemblies were uploaded to the NCBI GenBank database under the accession number PRJNA1018815. Detailed metadata associated with the genomes are provided in Supplemental Table S1. Sample IDs in the table were not known to anyone outside the research group.