Abstract
The impact of the tissue context on tumor growth and drug response in medulloblastoma (MB) is poorly understood. To gain insights into the growth and dissemination behavior of the MB tumor cells under treatment, we combined three-dimensional cell culture screening with ex vivo organotypic cerebellum slice co-culture (OCSC), which allowed the assessment of tumor cell behavior in the tissue context.
To identify druggable kinase pathways involved in invasion, we screened a panel of 274 kinase inhibitors and identified aurora kinase B (AURKB) as a potential anti-invasion drug target in MB. We validated tumor suppressive activities of the AURKB inhibitor (AURKBi) Barasertib (AZD1152-HQPA) and the structurally unrelated compound GSK-1070916 in cerebellum slice culture models for SHH, and Grp3 MB. Importantly, AURKBi are tumor suppressive in the tissue context, also in MB tumor cells that are in vitro resistant to the same treatment. We confirmed the requirement of AURKB for tumor growth and expansion in the tissue context through genetic suppression of AURKB by siRNA. We revealed that the combination of AURKBi with the SRC/BCR-ABL inhibitor Dasatinib acts synergistically to repress tumor growth and expansion in the highly invasive MB cell model ONS-76, but not in Grp3 MB cells. We demonstrate that tumor growth in the tissue context is suppressed by pharmacological inhibition of AURKB, comparable to the growth reduction observed after X-ray irradiation, which was used as the positive control. Finally, we show that exposure to µM concentrations of Barasertib does not cause developmental toxicity in fish larvae.
In conclusion, we demonstrate that AURKB is essential for MB tumor growth and expansion in the tissue context and the inhibition of AURKB is equally efficient as irradiation in repressing tumor cell growth. In patients younger than three years, pharmacological targeting of AURKB may thus constitute a novel means to overcome radiotherapy limitations.
Key words: Medulloblastoma, aurora kinase B, organotypic cerebellum slice culture, radiation therapy replacement, zebrafish larval model
Key points.
AURKB is essential for in tissue growth of MB.
Inactivation of AURKB causes p53 upregulation and represses in tissue growth and dissemination.
AURKBi Barasertib does not cause developmental toxicity
Alt-text: Unlabelled box
Introduction
Brain and nervous system tumors represent the second most common cancer entity in terms of incidence and mortality in children worldwide [Cancer Today, WHO], within which medulloblastoma (MB) is the most common malignant tumor. Current MB treatments consist of surgery complemented with chemotherapy and radiotherapy for children older than three years. Survivors often suffer from long-term side effects associated with radiotherapy, and approximately 30% of patients still die from the disease [1]. Four molecular subgroups (WNT, SHH, Group 3 (Grp3), and Group 4 (Grp4)) sub-divided into twelve subtypes have been molecularly characterized [2]. Although Grp3 and Grp4 MB tumors present a higher incidence of distal metastasis, metastasis is also a poor prognosis marker for SHH subgroup tumors [[1], [2], [3]]. MB metastasis occurs locally in the cerebellum midline and hemispheres and distally on the leptomeninges of the brain and the spinal cord. Because of tissue invasion, local recurrence is observed for SHH MBs, whereas Grp3 and Grp4 tumors tend to spread more distally [1]. Growth factor signaling pathways have been implicated in cerebellar tissue invasion [[4], [5], [6]] and chemokine signaling in distal spreading [7]. Druggable driver kinases of MB dissemination and distal growth remain to be identified.
Aurora Kinase A (AURKA), Aurora Kinase B (AURKB), and Aurora Kinase C (AURKC) are highly conserved proteins with homologous structures involved in mitosis [8]. AURKA controls centrosome maturation, bipolar spindle assembly, and cytokinesis, while AURKB and AURKC allow chromosome condensation, attachment to kinetochores and their alignment during metaphase and cytokinesis [9]. AURKA was identified as a critical gene involved in MB progression [10] and AURKA inhibition in MB cell lines was found to increase chemosensitivity [11]. Exposure of SHH MB models to pan-AURK inhibitors was reported to enhance the sensitivity to conventional chemotherapies in vitro and in vivo [12]. AURKB expression correlates positively with MYC in MB and specific inhibition of AURKB blocks MYC overexpressing MB tumors in vivo [13].
This study aimed to identify druggable kinases involved in invasion and dissemination control in MB. We used the spheroid invasion assay (SIA) [14] with a panel of 274 pharmacological kinase inhibitors to identify kinases involved in invasion control in MB. Pan AURKA/B inhibitors identified AURKB as a potential druggable target for invasion control. To further explore this finding and to systematically address the consequence of pharmacological targeting of AURKB in MB in the tissue context, we determined the impact of AURKB inhibition ex vivo using the organotypic cerebellum slice co-culture (OCSC) model for MB tumors [15]. We further compared the tumor-suppressive efficacy of genetic interference with AURKB, pharmacological AURKBi and X-ray radiation, and tested AURKBi for developmental toxicity in a zebrafish larval model, ultimately to explore AURKBi as a potential treatment for infant MB patients who cannot be treated with radiotherapy.
Materials and Methods
Tumor cells
Culture, origin, and authentication of ONS-76, HD-MB03 (Grp3), D425-Med (D425, Grp3) and D283-Med (D283, Grp3/4) cell lines are described in the Supplementary table 1. The ONS-76 cells were derived from a 2-year-old female with a cerebellar tumor [16] and later classified as a SHH cell line [17,18]. Unlike DAOY cells, where we could demonstrate GLI1 induction after smoothened activation [6], SHH status could not be confirmed in ONS-76 cells. Hence, we refer to this cell line as a spontaneously invasive MB cell line. The generation of LA-EGFP-lentivirus transduced descendants is described here: [5].
Patient-derived cells
Patient-derived tumor tissue samples were obtained from in-house patients with informed patient consent (Approval by: Cantonal Ethic Commission Zurich, Switzerland), and tumor tissue samples were processed as described [19]. 10’000 patient-derived cells (PDCs)/well were seeded in 96-well clear round bottom low adhesion plate to obtain spheroids, which were then implanted on cerebellum slices. 300′000 PDCs were seeded in 6-well low adhesion plates for 48 h to extract RNA and perform RT-qPCR. Patient samples: MB-PDC: Primary MB, group 3, subtype VII, male patient 16 years; ATRT-PDC: Primary ATRT, SHH subgroup, posterior fossa, female patient 1.5 years; EP-PDC, Primary Ependymoma, RELA-fusion, male patient 2 years.
Spheroid Invasion Assay (SIA)
SIA was performed as described in [14] with 2’500 ONS-76 cells seeded in 100 µl per well of complete RPMI medium in 96-well low adhesion plate. Cell invasion was expressed as the sum of invasion distances of the cells from the center of the spheroid after 24 h.
Kinase inhibitor screen, quantification of sum of distance of invasion
1’200 individual samples containing 3’000 ONS-76 mCherry-nuc cells/well were seeded in 96-well low adhesion plates for 24 h. Spheroids were embedded in collagen I as described in Ref. [14] and treated individually with a total of 275 kinase inhibitors at 3 µM concentration for 24 h. The full list of the 274 kinase inhibitors is shown in Supplementary table 2. Dasatinib treatment was used as a positive control and DMSO as negative control. mCherry-nuc fluorescence was acquired on a GE INCell2200 automated microscope using a Nikon 4X/0.2, Plan Apo, CFI/60 objective. For each well, a single field-of-view was acquired, composed of the maximum-intensity projection of a stack of 18 images with a step of 50 μm, an exposure time of 90 ms. No bining was applied, generating images of 2048 × 2048 pixels, with a pixel size of 1.6252 μm. Excitation and emission wavelength were 542/27 and 597/45, respectively. CellProfiler software [20] (v3.1.5) was used for image quantification as follows: A centroid Cmin around the spheroid (encompassing the optical halo of the spheroid) and a second centroid encompassing maximal migration distances of cells, Cmax, was created. Each individual nucleus was then outlined, and in each image, counts of cells in the area between Cmin and Cmax per image and sum of distances between each cell and spheroid were determined. Each screen plate was validated through a proper Z’-factor calculated on positive- and negative-control replicates on respective plates, and a z-score for each sample was used to establish a hit list of effective compounds [21]. Kinase inhibitors used for validation are listed in Supplementary table 3.
In vitro viability and cytotoxicity assay
CellTiter Glo® and CellToxTM Green assays were used with 750 cells seeded in 25 µl per well of complete medium in 96-well low adhesion plates for 24 h. Kinase inhibitors or DMSO for normalization were deposited on plates using a HP D300 Digital Dispenser (Hewlett-Packard Development Company). The relative luminescence (CellTiterGlow) or fluorescence (CellToxGreen, λex=485nm and λem=520nm) per well were measured after 24, 48 and 72 h of compound incubation at 37°C using a Cytation 3 imaging reader (BioTek®). ZIP (zero interaction potency) synergy scores of drug combinations were obtained using SynergyFinder® (https://synergyfinder.fimm.fi/). Kits and specific reagents used are listed in Supplementary table 4.
Irradiation (IR)
X-ray irradiation of cells in suspension cultures or OCSCs was performed using the CellRad X-ray irradiator PXI (Precision, North Branford, CT, USA) with the following settings: 1,8 Gy, 0,6 Gy/min, 5 mA, 105 kV.
Long-term viability assay
15’000 cells in 60 µl cell culture medium were seeded on Millipore inserts. The inserts were then transferred to 24-well plate containing 500 µl of OCSC medium per well. medium was changed daily, and cells were treated with 10, 100, or 500 nM Barasertib for five consecutive days. Cell viability was determined by CellTiter Glo 3D assay.
Immunoblot (IB)
200’000 cells were seeded in 6-well low adhesion plates for 48 h in complete medium and all treatments were performed in suspension. Cells were lysed with Laemmli buffer. HRP-linked secondary antibodies were used with either Pierce™ ECL Substrate or SuperSignal™ West Femto Maximum Sensitivity Substrate. Integrated densities of immuno-reactive bands were quantified using Image Lab (Version 5.2.1, Bio-Rad Laboratories). Antibodies used for IB and IF analysis are listed in Supplementary table 5.
Organotypic cerebellum slice co-culture (OCSC)
OCSC cultures were authorized by the Cantonal Veterinary Service of Zürich (ZH116/20) and performed as described in [15]. A maximum of three slices were placed per insert, and media were changed daily for two weeks. Tumor spheroids from ONS-76 LA-EGFP, HD-MB03 LA-EGFP, D425, D283, or from primary tumor-derived cells were then placed on the slices and incubated for 24 h. The tumor spheroid-slice co-cultures were subsequently drug-treated for five days.
Click-iT® EdU (C10340, Invitrogen) was used for detecting proliferating cells. 20 min EdU incubation was used for established tumor cell lines and 2 h for patient-derived cells. Antibodies used are listed in Supplementary table 5. The inserts were flat mounted onto glass slides in glycergel mounting medium.
Images were acquired on an SP8 Leica confocal microscope (Leica Microsystems, Mannheim, Germany). Tumor volume (TV) and proliferation volume (PV) were determined by quantifying the volume of the green fluorescence (human nucleoli) and the volume of the red fluorescence (EdU staining), respectively, from z-stacks using Imaris software (volume thresholding: smooth surfaces detail 15 µm [human nucleoli] and 5 µm [EdU], background subtraction [local contrast] 8.52 µm). Two measurements were acquired per slice, which were normalized to the negative control (DMSO-treated) condition (CTRL). Relative proliferation volume (rPV) = PV/TV.
siRNA-mediated depletion
10 nM siAURKB (AURKB Stealth siRNA, ThermoFisher Scientific, HSS190048) was used for AURKB depletion using lipofectamine RNAiMAX transfection reagent with reverse transfection. 300 µl of the siRNA transfection mix were deposited in a 6-well plate onto which 300’000 MB tumor cells were seeded. After 6 h of incubation, 5’000 transfected cells/well were seeded in a 96-well round bottom low adhesion plate for 24 h to form spheroids, which were then deposited on cerebellum slices. Co-cultures were then maintained in culture for 48 h (72 h after transfection). Silencing of AURKB was validated 72 h after transfection by IB from 250’000 cells seeded in low adhesion plates.
RT-qPCR: mRNA expression level after treatments
300’000 HD-MB03 or D425 were seeded per well in 6-well low adhesion plates. After 48 h, medium was replaced, cells were irradiated and incubated for another 8 or 24 h ± 50 nM Barasertib. At endpoint, cells were collected and RNA was purified using QIAGEN RNeasy Mini Kit. 1 µg of mRNA was reverse transcribed in cDNA using the high-capacity cDNA Reverse Transcription Kit. RT-qPCR was performed using TaqMan Master mix under conditions optimized for the ABI7900HT instrument. The ∆CT method was used to calculate the relative gene expression of each gene of interest relative to GAPDH. RT-qPCR primers used are listed in Supplementary table 6.
Zebrafish maintenance and treatment
Adult zebrafish were housed and maintained under standard procedures at the Fish Facility of the Department of Molecular Life Sciences, University of Zurich. Animals housing and husbandry are approved by the Veterinary Office of the Kanton of Zürich, Switzerland. This work did not require ethical approval from the animal welfare committee. Adult fish were kept in recirculating systems in 14:10 h light/dark cycle at 28°C. Embryos were obtained by natural spawning and raised at 28.5°C in embryo medium (E3). At 2 days post fertilization (dpf), embryos were manually dechorionated and treated with Barasertib at concentrations of 2,4,6,8,10 µM and with DMSO (0.01% v/v) as a vehicle control. The treatments were renewed daily, and larvae were monitored for survival until 5 dpf. Wild-type AB strain was used in this study.
Zebrafish image analysis
Treated zebrafish larvae were screened using a VAST BioImager™ platform (Union Biometrica, USA). A deep-learning method was developed to segment different morphological regions. We used 2250 images with annotations and a resnet50 model to train the segmentation algorithm [22]. From the segmented regions, we measured 6 features: body area, eye size, yolk area, body length, pericardiac area, notochord area, and inflated swim bladder.
Statistical analysis
Statistical differences were calculated with Kruskal-Wallis’ test followed by multiple comparison test with a two-stage linear step-up procedure of Benjamini, Krieger and Yekutieli or with Mann-Whitney test with p-value correction (cell and tissue culture experiments) or Kruskal-Wallis’ test followed by Dunn's multiple comparisons test (zebrafish toxicity) using Prism GraphPad 8 software. P-values of p ≤ 0.05 were considered statistically significant.
Results
3D invasion assay identifies Aurora and SRC kinase inhibitors as invasion inhibitors
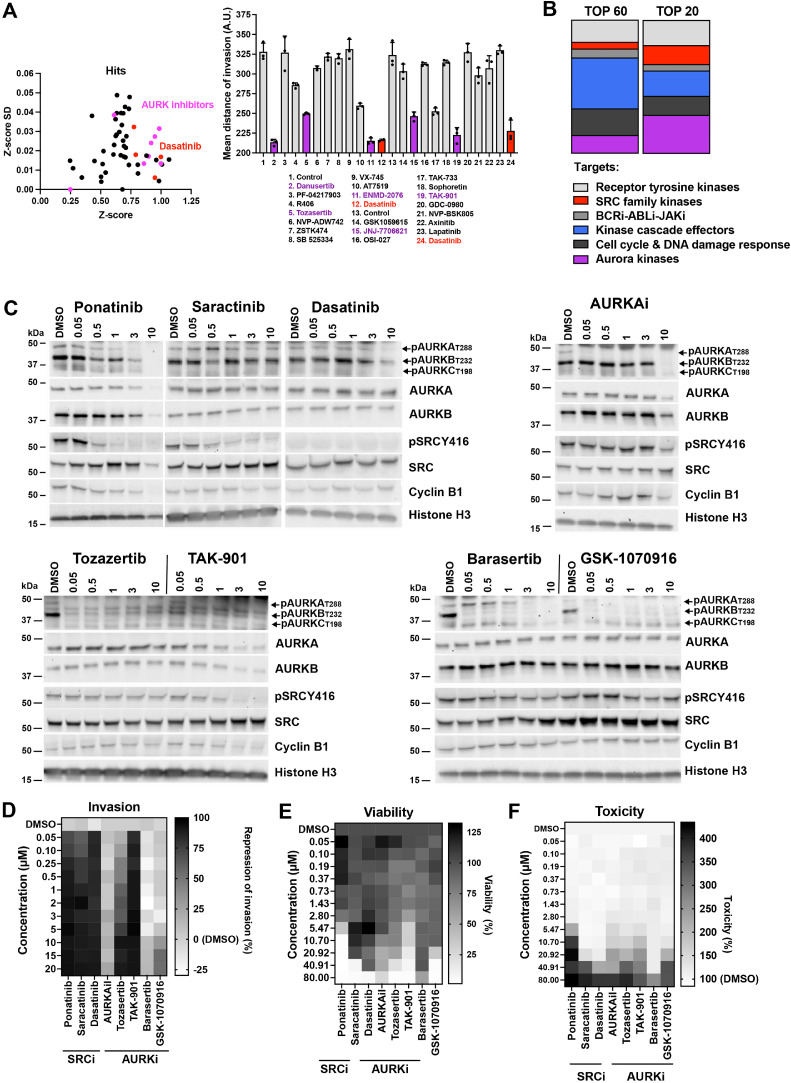
We tested 274 different kinase inhibitors in the ONS-76 MB cancer cell line [23] to identify inhibitors of pro-invasive kinases in MB. 60 inhibitors including Dasatinib [24] used as positive control reduced ONS-76 cell invasion (Fig. 1A, supplementary table 1). The top 20 compounds included several aurora kinase inhibitors (AURKi) (Fig. 1A, B). To validate the potential implication of SRC and AURK, we selected SRCi Ponatinib, Saracatinib, and Dasatinib and AURKi Aurora kinase A inhibitor I (AURKAi I), tozasertib (VX-680), and TAK-901. We first confirmed the specific inhibition of SRC and AURKs by immunoblot (IB) using phosphosite-specific antibodies [25,26] (Fig. 1C). The SRCi inhibited SRC kinase activity at 1 µM concentration without affecting AURK activities. We also confirmed AURKAi I specificity for AURKA. Tozasertib and TAK-901 reduced both AURKA and AURKB phosphorylation (Fig. 1C). All three SRCi effectively repressed collagen I invasion of ONS-76 cells at nanomolar concentrations (Fig. 1D, Supplementary Fig. 1A), consistent with the EC50 of Dasatinib in this line [24]. TAK-901 suppressed invasion of ONS-76 cells at 50 nM concentration. Tozasertib reduced collagen cell invasion in a dose-dependent manner with an EC50 of approximately 4 µM, whereas AURKAi I did not repress invasion up to 20 µM concentration (Fig. 1D). The two AURKB inhibitors [27] Barasertib (AZD1152-HQPA) and GSK-1070916, which both effectively reduced AURKB phosphorylation (Fig. 1C), were also ineffective in reducing collagen I invasion (Fig. 1D, Supplementary Fig. 1A). We, therefore, combined AURKAi I with either Barasertib or GSK-1070916 to test whether the inhibition of both AURKA and AURKB is necessary for blocking invasiveness (Supplementary Fig. 1B-C). As neither combination showed a reduction in ONS-76 cell invasion, we concluded that TAK-901 and Tozasertib effects are likely pan-kinase off-target activities of these inhibitors. None of the tested inhibitors at effective concentration reduced cell viability (Fig. 1E) or induced cell death (Fig. 1F)
Fig. 1.
Blockade of collagen I invasion by Aurora and BCR-ABL/SRC kinase inhibitors.
A) Left: X/Y plot of Z-score and Z-score SD of hit list. Right: Validation of selected drugs using SIA and aCDc software[14] showing mean distances of invasion from center of spheroids and SD (n=3 biological replicas). Purple: AURKBi, red: BCR-ABL/SRC inhibitors, all tested at 3 µM. B) Representation of top 60 and top 20 candidates across kinase inhibitor families. C) IB using antibodies specific for the phosphorylation of the active sites of SRC, and aurora kinase A and B proteins. Cyclin B1 protein expression confirms cell proliferation. D) Heat-map of mean values of anti-invasion efficacy of selected kinase inhibitors of SIA shown in A). E) Heat-map of mean cell viability in ONS-76 cells. F) Heat-map of cytotoxicity in ONS-76 cells. (D – E: n=3 biological replicas).
In conclusion, we found that Ponatinib, Saracatinib, Dasatinib, Tozasertib and TAK-901 possess excellent anti-invasion efficacy at concentrations below 1 µM. In contrast, specific inhibition of AURKA and AURKB by AURKAi I and Barasertib or GSK-1070916, respectively, is insufficient to block collagen I invasion in ONS-76 cells in vitro.
AURKB inhibition reduces growth and dissemination in the tissue context
We next tested a 5-day treatment regimen at 500 nM concentration on tumor cell growth and dissemination in the tissue context using OCSCs [15] (Supplementary Fig. 2). We calculated the DMSO-normalized tumor volume (TV) and relative proliferation volume (rPV) for each compound. Both Saracatinib and Dasatinib, reduced TV and rPV of ONS-76 cells at day 6 by 45% (p=0.0338) and 60% (p=0.0003), respectively (Fig. 2A, Supplementary Fig. 3A,4A, B). We observed no inhibitory effect of Ponatinib. Barasertib and GSK-1070916 reduced ONS-76 TV and rPV significantly in OCSCs by around 40-50% (p0.0338) (Fig. 2A, Supplementary Fig. 3A, 4A, B, E). The TV of the Grp3 line HD-MB03 increased 5-fold between day 1 and day 6 under control (DMSO) conditions (Supplementary Fig. 3B, 4C, E). The SRCi showed little effect on TV and rPV in HD-MB03 cells (Fig. 2A, Supplementary Fig. 3B, 4C-E). In contrast, both Barasertib and GSK-1070916 significantly reduced TV and rPV by >50% (p0.0051) and 80%-90% (p0.0002), respectively (Fig. 2A, Supplementary Fig. 3B, 4C-E). Importantly, the combination of SRCi and AURKBi led to the near-complete eradication of ONS-76 cells in the tissue context (Supplementary Fig. 3A, 4A, B, E). This additive effect of the compound combination was less pronounced in HD-MB03 cells, as AURKBi alone was already highly effective (Supplementary Fig. 3B, 4C-E).
Fig. 2.
AURKBi reduces medulloblastoma cell dissemination ex vivo.
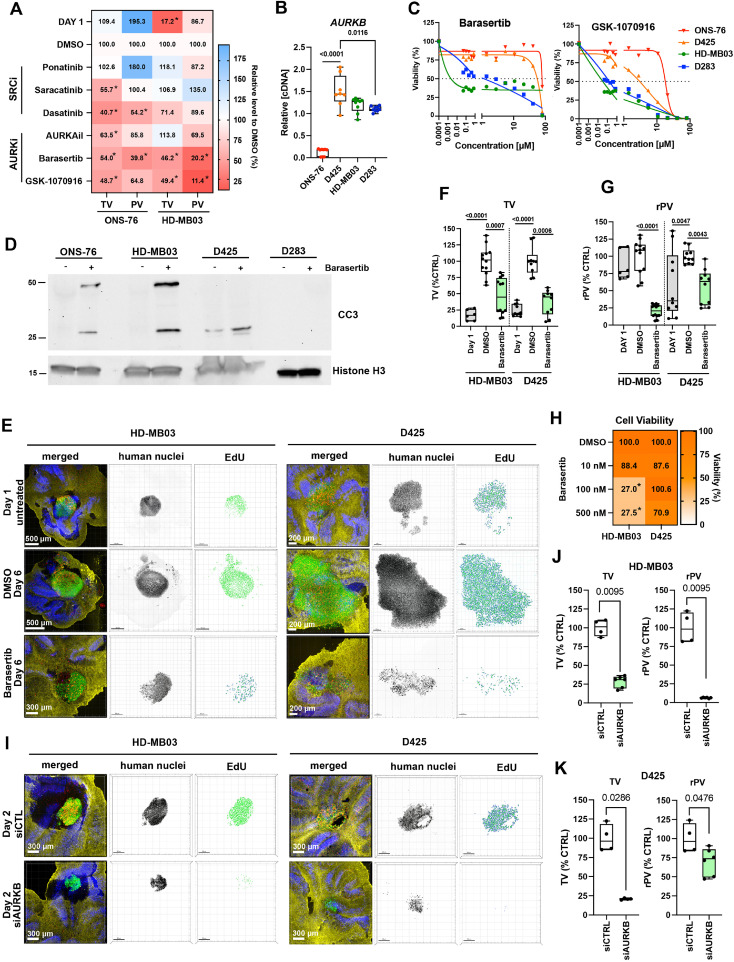
A) Heat-map of tumor volume (TV) and relative proliferation volume (rPV = PV/TV) of ONS-76 and HD-MB03 cells in OCSCs after 5 days of treatment (*p0.0338). B) RT-qPCR analysis of AURKB expression (n=3 biological replicas). C) Relative cell viability in response to Barasertib and GSK-1070916 (n=3 biological replicas). D) IB analyses of anti-cleaved caspase 3 (CC3) and anti-histone H3 of cells treated with 0.05 (HD-MB03) or 3 (ONS-76, D425, D283) µM Barasertib. E) Confocal microscopy IFA of OCSCs with implanted HD-MB03 and D425 cells (green). OCSCs with implanted tumor cells were treated with Barasertib at either 500 nM (HD-MB03) or 100 nM (D425) concentrations. Red: Click-iT® EdU, blue: anti-calbindin, and yellow: anti-GFAP. Human nuclei are displayed in inverted greyscale and EdU incorporation as volume renderings from Z-stack images. F) TV of n=5 individually implanted spheroids. G) rPV of n=5 individually implanted spheroids. H) Heat map of in vitro cell viability of HD-MB03 and D425 cells after 5-days of treatments (n=3 biological replicas, *p0.0018). I) Confocal microscopy IFA of OCSCs with implanted HD-MB03 and D425 cells (green) with siRNA-mediated depletion of AURKB in tumor cells. Human nuclei are displayed in inverted greyscale and EdU incorporation as volume renderings from Z-stack images. J) TV and rPV of HD-MB03 of experiment shown in I. Data of n=3 individually implanted spheroids are shown. K) TV rPV of D425 of experiment shown in I. Data of n=3 individually implanted spheroids are shown.
The Grp3 cell lines D425 and HD-MB03 as well as the Grp3/4 D283 cell line display considerably higher AURKB expression than ONS-76 cells (Fig. 2B). Similar relative expression of AURKB is also observed at protein levels, although D283 cells express less AURKB than HD-MB03 and D425 (Supplementary Fig. 5A). Barasertib effectively reduced viability of HD-MB03 and D283 cells in vitro, with an IC50 of 2 nM and 1 µM, respectively, unlike the D425 and ONS-76 cells, where we observed no inhibition up to 40 µM (Fig. 2C, Supplementary Fig. 5B). GSK-1070916 also effectively reduced HD-MB03 and D283 cell viabilities with IC50 of 0.033 and 0.2 µM, respectively. Barasertib treatment increased cleaved caspase 3 (CC3) signals in HD-MB03, D425 and ONS-76 cells (Fig. 2D). No increase in CC3 was detected in D283 cells, suggesting that decreased viability of this line (Fig. 2C, Supplementary Fig. 5B) is rather the consequence of reduced proliferation and not of apoptosis.
To explore the therapeutic potential of AURKBi for Grp3 MB, we compared the effect of AURKBi in HD-MB03 and D425 cell lines in OCSCs (Fig. 2E-G). We selected these two cell lines because (i) they highly express MYC (Supplementary Fig. 5A), a known marker of MB aggressiveness [3], and (ii) they represent an in vitro Barasertib-sensitive (HD-MB03) and -resistant cell line (D425). D425 cells display a similar propensity to grow and invade the cerebellum tissue as HD-MB03 (Fig. 2E). Barasertib treatment significantly reduced TV in HD-MB03 and D425 by 64% (p=0.0007) and 73% (p=0.0006), respectively. Similarly, the rPV in both cell lines decreased with a reduction of more than 80% (p<0.0001) for HD-MB03 and by 45% (p=0.0043) for D425. To assess slice impact on treatment efficacy, we cultured HD-MB03 and D425 cells in the absence of the cerebellum slices on inserts for five days using the OCSC medium (Fig. 2H). Under these conditions, Barasertib treatment reduced HD-MB03 viability by 70% (p=0.0018), similar to what we observed in the tissue context (Fig. 2B). No significant reduction of cell viability was observed for D425 cells. In contrast, the same concentration of Barasertib efficiently reduced the TV and rPV of D425 cells in OCSCs (Fig. 2E-H).
These results indicate that the cerebellar microenvironment can impact the functional activity of Barasertib. AURKB-depletion in tumor cells by siRNA phenocopied Barasertib effects and led to a near complete eradication of the tumor cells in OCSCs (Fig. 2I, 2J, K, Supplementary Fig. 6A-C), thus confirming the relevance of AURKB function in the tumor cells for growth and dissemination of MB in the tissue context.
Barasertib treatment increased p53 levels and induced p53 target gene expression
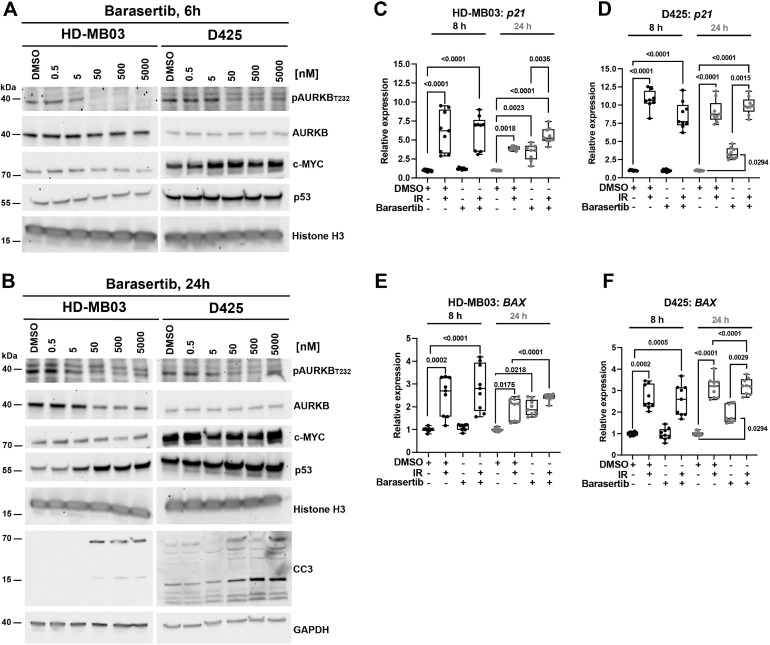
AURKB phosphorylation of MYC on S67 can stabilize MYC protein and control its transcriptional activity [28]. We did not observe a clear effect of Barasertib treatment on MYC protein levels (Fig 3A), neither did we observe a reduction of MYC or of the MYC target genes GLUT-1 [29], ITGA1 [30] and CDC42[31] at mRNA level (Supplementary Fig. 7A, B). AURKB can also phosphorylate p53 at S183, T211, and S215 to accelerate its degradation and suppresses target genes such as p21 [32]. 50 nM Barasertib is sufficient to repress AURKB Thr232 phosphorylation within 6 h in HD-MB03 and D425 cells (Fig. 3A) and to increase p53 protein and CC3 levels within 24 h (Fig. 3B). Consistently, we also observed the induction of the p53 target genes p21 and BAX (Fig. 3C-F). We used X-ray irradiation (IR) as control for p53 induction [33], which caused p53 target gene expression within 8 h. We did not observe a comparable induction of p21 and BAX after 8h of Barasertib treatment, suggesting that p53-induction is rather a long-term effect of AURKB inhibition.
Fig. 3.
AURKBi induces p53 expression and cleaved caspase 3 in Grp 3 MB.
A) IB of Barasertib treatment effects on AURKA,B,C phosphorylation (pThr232) and MYC and p53 protein levels after 6 h of treatment. B) IB of Barasertib treatment effects on AURKA,B,C phosphorylation and MYC, p53 protein levels and CC3 after 24 h of treatment. C-F) RT-qPCR quantifications of p21 and BAX mRNA levels in HD-MB03 (C,E) and D425 (D,F) cells. Cells were irradiated (IR, 1,8 Gy) and/or treated with Barasertib (50 nM) for 8 and 24 h (n=3 biological replicas).
Thus, reduced tumor expansion and proliferation after AURKB inhibition is not a consequence of impaired MYC function in MB but may rather involve the induction of p53 target genes.
Comparable efficacy of Barasertib and IR in the tissue context
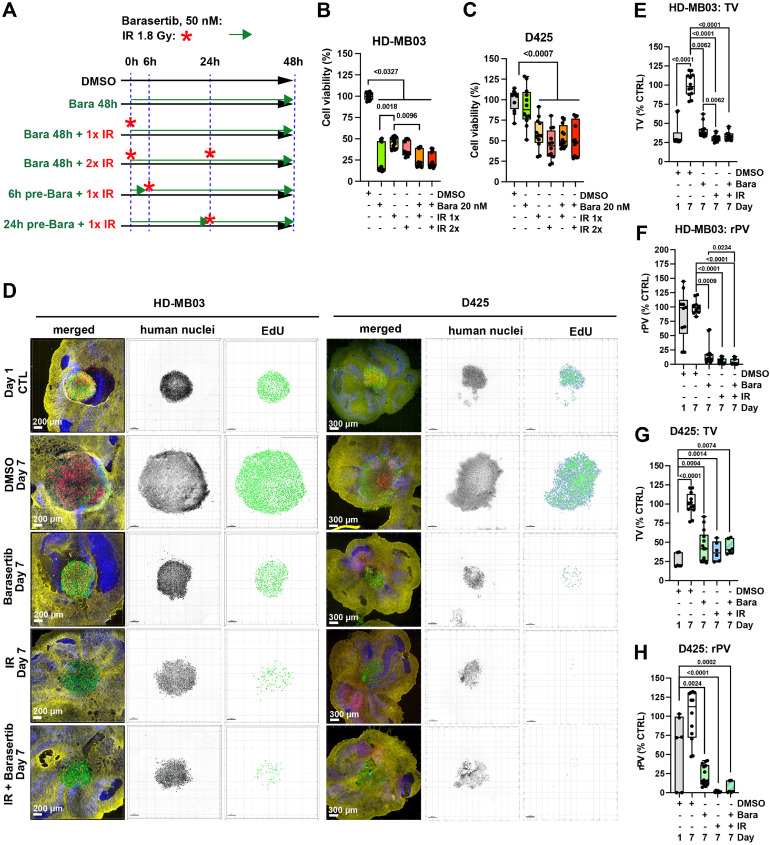
To explore Barasertib effects in comparison to IR, our positive control, we assessed Barasertib and IR treatment effects on cell viability. We first treated HD-MB03 and D425 cells in vitro with Barasertib and/or IR (1,8 Gy) as depicted in Fig. 4A. In HD-MB03, as little as 20 nM Barasertib reduced viability by more than 75% (Fig. 4B, Supplementary Fig. 8A, C). The same concentration had no effect in D425 (Fig. 4C, Supplementary Fig. 8B, C). IR reduced cell viability in both HD-MB03 (minus 55-63%) and D425 (minus 42-55%). Adding a second cycle of IR did not significantly increase treatment efficacy (Fig. 4A, B). Barasertib plus IR did not further reduce cell viability compared to Barasertib or IR alone (Fig. 4A, B). In HD-MB03, Barasertib treatment alone displayed a 25% (p=0.0018) increased inhibitory effect compared to IR treatment alone.
Fig. 4.
AURKBi and IR display comparable tumor suppressive activity in vitro and ex vivo.
A) Experimental procedure for the in vitro cell viability assays shown in B and C. B) Viability of HD-MB03 and C) D425 cells in response to IR (1,8 Gy) and Barasertib (20 nM) treatment alone or in combination after 48 h (n=3 biological replicas). D) Confocal microscopy IFA of OCSCs 6 days after treatments with Barasertib 100 nM and IR 1.8 Gy. Green: Anti-human nucleoli antibody, red: Click-iT® EdU, blue: anti-Cabindin Click-iT® EdU, yellow: anti-GFAP. Human nuclei are displayed in inverted grey scale and EdU incorporation as volume renderings from Z-stack images. E-H) Box plots of TV and rPV of OCSCs from D. (n≥3 biological replicas).
Ex vivo in OCSCs using HD-MB03 cells, Barasertib treatment for six days alone reduced TV by 60% (p=0.0062) and rPV by 83% (p=0.0009). In the same experiment, IR alone reduced TV of HD-MB03 cells by 70% (p0.0001), and rPV by 95% (p0.0001) (Fig. 4D-F). Barasertib plus IR reduced TV by 66% (p0.0001) and rPV by 96% (p0.0001) (Fig. 4D-F). In D425 cells, Barasertib alone reduced TV by 66% (p=0.0004) and rPV by 88% (p=0.0024). IR alone reduced the TV by 63% (p=0.0014) and the rPV by 99% (p=0.0033) (Fig. 4D, G, H). Barasertib plus IR reduced TV in D425 by 57% (p=0.0074) and rPV by 94% (p=0.0002). In both cell lines, Barasertib plus IR did not significantly increase efficacy compared to each treatment alone. Together, our results demonstrate that AURKBi-mediated growth repression is comparable to growth repression induced by IR.
Subgroup-specific sensitivities to combinatorial treatments in the tissue context
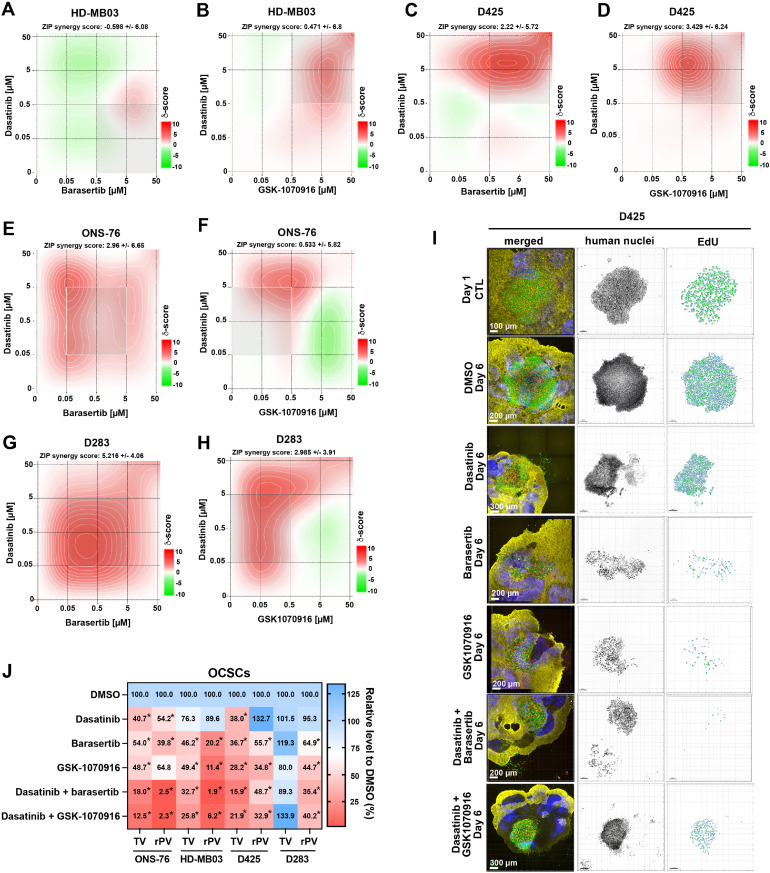
We next explored whether the combination of the invasion inhibitor Dasatinib and Barasertib or GSK-1070916 could exert an additive or synergistic effect in HD-MB03 (Fig. 5A, B), D425 (Fig. 5C-D), ONS-76 (Fig. 5E, F) and D283 (Fig. 5G, H) cells. The responses differed considerably between the cell lines. In HD-MB03 cells, the Barasertib-Dasatinib combination elicits antagonistic (Fig. 5A, B), and in ONS-76 (Fig. 5E, F) or D283 cells (Fig. 5 G, H) synergistic effects. We also noted some differences when comparing Barasertib-Dasatinib and GSK-1070916-Dasatinib combinations, indicating that the difference in chemical structure of the two AURKBi results in a differential drug response.
Fig. 5.
Synergistic activity of AURKBi and Dasatinib in vitro and ex vivo.
A-H) Synergy scores of drug combinations after 48 h of treatment with 0.005, 0.05, 0.5, 5 and 50 µM of Barasertib and Dasatinib in HD-MB03 (A), D425 (C), ONS-76 (E) and in D283 (G) cells. Synergy score of drug combination of GSK-1070916 and Dasatinib in HD-MB03 (B), D425 (D), ONS-76 (F) and in D283 (H) cells. I) Confocal microscopy IFA of OCSC-embedded D425 cells treated for 5 days with 100 nM Barasertib, GSK-1070916, or Dasatinib, and combinations of Barasertib and Dasatinib or GSK-1070916 and Dasatinib. Green: Anti-human nucleoli antibody, red: Click-iT® EdU, blue: anti-Cabindin, yellow: anti-GFAP. Human nuclei are displayed in inverted grey scale and EdU incorporation as volume renderings from Z-stack images. J) Heat map of TV and rPV of ONS-76, HD-MB03, D425 and D283 OCSC co-cultures after 5-days of treatments (n≥3 biological replicas, *p0.0338).
We also evaluated the combinatorial effect of AURKBi and Dasatinib, both at 100 nM concentration, in OCSCs with D425 (Fig. 5I, Supplementary Fig. 9D, E) and D283 (Supplementary Fig. 9A-C) cells. AURKBi strongly reduced TV and rPV in D425 (Fig. 5J), but not in D283 cells, where only rPV was reduced. Combining AURKBi with SRCi decreased TV of the ONS-76 cells by a factor of >-3 (p0.0478) and rPV by a factor >-17 (p=0.0300) compared to AURKBi single treatment alone. No significant additive effect of the compound combinations compared to AURKBi single treatment alone was noted in HD-MB03 cells. In D425 cells, we observed an additive effect of the Barasertib-Dasatinib combination on TV only (factor -2.3, p=0.0112), and in D283, we observed an additive effect of the Barasertib-Dasatinib combination on rPV only (factor -1.8, p=0.0139). Thus, AURKB plus BCR-ABL/SRC inhibition is additive for the ONS-76 cell line only.
Barasertib-Dasatinib combination treatment blocks growth of primary tumor samples in the tissue context
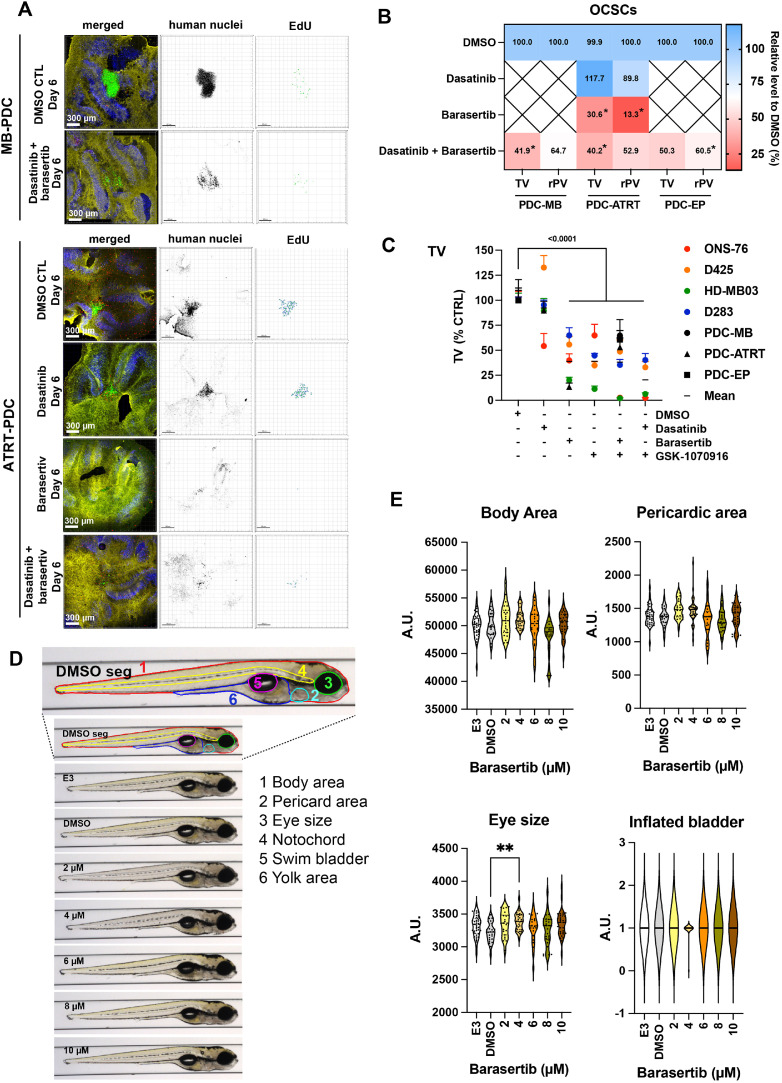
We then evaluated the efficacy of Barasertib alone or in combination with Dasatinib in patient-derived cells (PDC) from primary medulloblastoma (MB), atypical teratoid rhabdoid tumor (ATRT), and ependymoma (EPD) in OCSCs (Fig. 6A-B, Supplementary Fig. 10A). In MB PDCs, the combination of Barasertib and Dasatinib reduced TV by nearly 59% and rPV by 56%. With this treatment, a similar level of repression of TV and rPV was also observed in PDCs of ATRT (-59%, Fig. 6A, B) and EPD (-49%, Supplementary Fig. 10A). In ATRT, we additionally tested single Barasertib and Dasatinib treatments. Barasertib decreased TV by 70% and rPV by 86%. No additive or synergistic effect was observed when Barasertib was combined with Dasatinib.
Fig. 6.
Barasertib effectively blocks patient-derived cell growth in the tissue context as is not toxic.
Confocal microscopy IFA of OCSC implanted MB-PDC and ATART-PDC 5 days after treatments with 500 nM Barasertib or Dasatinib or the combination of Barasertib and Dasatinib. Green: Anti-human nucleoli antibody (PDCs), red: Click-iT® EdU, blue: anti-Cabindin, yellow: anti-GFAP. MB: medulloblastoma, ATRT: atypical teratoid rhabdoid tumor, EP: ependymoma. Human nuclei are displayed in inverted greyscale and EdU incorporation as volume renderings from Z-stacks. B) Heat map of TV and rPV of PDC-OCSC co-cultures after 5-days of treatments (n≥3 biological replicas, *p0.05). C-D) TV and the rPV of all cell lines and PDCs used (Mean + SEM). C) Compilation of all TVs from the OCSC experiments. D) Representative VAST images of lateral view of zebrafish in the different treatment groups. The first image corresponds to a control animal with segmentation of anatomical regions studied: body area (red), eye size (green), pericardiac area (light blue), inflated bladder (pink), yolk area (blue), and notochord (yellow). E) Quantification of a subset of morphological areas shown in D.
We also analyzed the drug response data of all OCSC experiments combined (Fig. 6C, Supplementary Fig. 10B). AURKB inhibition by either Barasertib or GSK-1070916 caused a significant reduction in TV and rPV in all cell lines and PDCs with the exception of D283 cells, where TV was not reduced. These data further corroborate the therapeutic efficacy of AURKB inhibition in restriction growth and tissue invasion in MB and potentially in other high-grade pediatric brain tumors.
Barasertib does not cause developmental toxicity in zebrafish larvae
Larval zebrafish treated with Barasertib at concentrations over 2 orders of magnitude higher than those used in vitro, show 100% survival and no signs of toxicity to any of the regions studied (Fig. 6D). The morphometric analysis from VAST imaging revealed no major differences between the control groups and the Barasertib-treated groups across all concentrations (Fig. 6E, Supplementary Fig. 10C). Some statistically significant differences were observed in the eye-size of the 4 µM-treated group and the yolk-area of the 8 µM-treated group. These data confirm the absence of developmental toxicity of Barasertib treatment.
Discussion
Drug exposure, drug import and drug extrusion mechanisms of tumor cells growing under 2D or 3D culture conditions differ from cells cultured as organoids or as xenografts in vivo [34,35] and the cellular, chemical, and biophysical context of the cerebellar microenvironment present in vivo is absent in current in vitro models. Some recent approaches addressed this problem and considerably improved 3D culture systems to better mimic MB tumor biology for high-throughput screenings [36,37]. Herein, we could demonstrate that despite the lack of in vitro efficacy in two cell models, both Barasertib and GSK-1070916 effectively reduced tumor volume and proliferation in the tissue context in all cell models tested. This highlights the relevance of AURKB activity and indicates a specific impact of the cerebellar microenvironment on the functional activities of drugs, analogous to the difference in drug sensitivity described in osteosarcoma [35]. Importantly, depletion of AURKB phenocopied the tumor-inhibiting effect of Barasertib and GSK-1070916, arguing against an indirect effect of Barasertib on the cerebellar slice leading to tumor suppression.
AURKBi have been tested in numerous clinical studies, whereby Barasertib is the most extensively tested molecule with clinical trials in leukemia, lymphoma, and diverse advanced solid malignancies [38]. Despite relatively well-manageable toxicities and promising objective complete response rates with acute myeloid leukemia for example [39], none of the so far tested 14 AURKBi made it to the clinics. Barasertib displays a low clearance rate in humans, but distribution kinetics govern the time to steady-state concentrations and the drug is eliminated before distribution equilibrium in the tissue can be achieved, possibly due to the increased association with the plasma protein fraction [40]. The lack of convincing in vivo efficacy in solid malignancies was also ascribed to the discrepancy between the proliferation rates in vitro and in vivo [41]. Indeed, higher proliferation rates increase the sensitivity of tumor cells to agents that target cell cycle progression. Barasertib targets mitotic chromatid positioning, leading to chromosomal segregation dysfunction [38]. To be effective, tumor cells may thus need to undergo several rounds of mitosis before apoptosis is induced. Our data show an effective reduction in tumor cell volume in the tissue context within five to six days at nanomolar concentrations, thus arguing for prolonged continuous, lower-dose treatment schemes. Reduced viability in response to Barasertib treatment promotes Caspase 3 cleavage in a dose and time-dependent manner in ONS-76, HD-MB03, and D425 but not D283 cells. The concomitant induction of BAX under these conditions could point towards AURKB implication in the repression of mitochondrial outer membrane permeabilization and the release of cytochrome C. Analogous antiapoptotic functions of AURKB via the repression of the pro-apoptotic BAX and BAK proteins were recently shown in the context of resistance to tyrosine kinase receptor inhibition in EGFR mutated lung cancer [42], and leukemia [43].
Anti-tumor efficacy of Barasertib has been demonstrated for pediatric acute lymphoblastic leukemia and neuroblastoma in vitro [[44], [45], [46]], and Barasertib efficacy was also shown in a MYC-overexpressing D425 MB flank model in vivo [13]. In this study, 12 days after the start of a four-day treatment, a considerable tumor re-growth was observed. This suggests that AURKB inhibition transiently repressed tumor growth without eradicating the tumor cells. A delay in tumor growth was also observed in a D425 intracranial tumor model [13], but all animals finally succumbed. The peak Barasertib levels measured in the brain of mice in this study were 0.7 ng/mg brain tissue [13], which is far above the minimal effective concentration of 100 nM we observed in the cerebellar slices. As the therapeutic efficacy of Barasertib in vivo is compromised by short exposure times and the onset of toxicity as a result of systemic exposure, alternative routes of drug administration into the brain must be considered. One possible route is intrathecal delivery via an Ommaya reservoir, which is widely used for the administration of chemotherapeutic drugs in pediatric patients [47,48]. Direct CNS delivery via an Ommaya reservoir would allow treatments with lower but still effective concentrations for a prolonged period. The toxicity analysis in the zebrafish larval model did not reveal any signs of overt developmental toxicity, although a statistical difference in the eye-size of the 4 µM-treated group and the yolk-area of the 8 µM-treated group was observed. However, these differences are unlikely to have biological relevance since no trend in the dose-dependent toxicity curve was observed; thus, we consider Barasertib, at the concentrations tested, to be non-toxic for larval development. Collectively, the data of the field and our data demonstrating in tissue efficacy of Barasertib at nanomolar concentrations, suggest that continuous, lower-dose drug exposure may be superior to high-dose short-term treatments.
MYC/MYCN and p53 are relevant transcription factors that could interact with AURKB in aggressive MB [[49], [50], [51]]. We confirmed that Barasertib effectively reduced viability defined as the capability of tumor cells to grow and proliferate in MYC overexpressing Grp3 MB cells [13] in the cerebellar slice culture model and additionally demonstrated AURKBi efficacy in MYC-low cell lines such as ONS-76. We also observed no marked change in MYC expression after Barasertib treatment. Our data thus indicate that AURKBi is effective independent of MYC-overexpression and that the previously described AURKB phosphorylation and stabilization of MYC on S67 and S373 [52,53] is likely not a tumor-driving mechanism in the models investigated. AURKB inhibition has both p53-dependent and p53-independent effects in cancer cells [54]. However, the increased p53 protein levels after 24 h treatment with inhibitory concentrations of Barasertib would support a model where repression of AURKB leads to p53 stabilization. Inhibiting AURKB could thus stabilize p53 and promote protective cell cycle arrests in p53 wt cells (ONS-76, HD-MB03, D283) [55]. Such Barasertib-induced p53 stabilization in MB cells could be induced by the loss of AURKB-dependent phosphorylation of p53, which accelerates its proteasomal degradation through an MDM2-dependent mechanism [32]. Consistent with AURKB-dependent destabilization of p53, we observed upregulation of the p53 target genes p21 and BAX after 24 h of Barasertib treatment, which also correlated with increased cleaved caspase 3. In D425 cells, the p53 protein is mutated [55]. p53 deficiencies sensitize cells to AURKB inhibition, leading to enhanced mitotic arrest and slippage [56]. However, whether a particular mutational alteration of D425 p53 could explain the differential in vitro sensitivity of D425 to Barasertib and GSK1070916 remains to be determined.
We expected increased p53 levels after Barasertib treatment to sensitize the cells to IR [57]. However, both in HD-MB03 and D425 cells, AURKBi plus IR did not significantly increase anti-proliferative or pro-apoptotic efficacy compared to each treatment alone. One possible explanation is that treatment-induced cell cycle arrest renders the cells less sensitive to either AURKBi or IR, which both target rapidly cycling cells. MB cell lines, primary tumors, and xenografts express different isoforms of p53 [58], and depending on p53 basal expression levels, opposite roles of p53 on the chemosensitivity of MB cells were described [59]. Determining the impact of AURKBi on differential p53 expression and function in MB will be necessary for better predicting patient's responses.
AURKB was recently found to promote tumor progression and invasion in hepatocellular carcinoma through interaction and regulation of the DHX9 helicase [60]. Interestingly, a strong positive correlation exists between AURKB and DHX9 expression in MB and other pediatric high-grade brain tumors (Supplementary Fig. 11), indicating that functional interaction of AURKB and DHX9 could contribute to the oncogenic functions of AURKB in MB and other high-grade brain malignancies in children.
Collectively our data demonstrate that genetic interference with or pharmacological inhibition of AURKB is comparably effective in repressing in tissue tumor growth as IR treatment. The lack of overt toxicity in the zebrafish larval model and on cerebellar slices indicates that AURKBi could represent a potential replacement treatment strategy for patients younger than three years, who cannot be treated by IR [61]. Finally, the increased treatment efficacy ex vivo in ONS-76 cells when AURKBi and SRCi were combined implies a beneficial effect of co-repression of tyrosine kinase receptor signaling in a subset of MB tumors [4,6].
Conclusion
We identified AURKB as a druggable vulnerability in medulloblastoma and demonstrated that two independent inhibitors of AURKB effectively repress MB tumor growth and dissemination in the cerebellar tissue. The beneficial effect of the combination of AURKBi with Dasatinib in ONS-76 cells indicates the relevance of the co-repression of tyrosine kinase signaling in a subset of MB tumors.
Funding
Swiss National Science Foundation (grants SNF_310030_188793 to MB and Sinergia_CRSII5_202245/1 to MB and SN), Swiss Cancer Research Foundation (grant KFS-4853-08-2019 to MB), and Childhood Cancer Research to MAG.
Data availability
The data will be made available upon reasonable request.
CRediT authorship contribution statement
Alexandre Gries: Writing – original draft, Visualization, Validation, Methodology, Investigation, Formal analysis, Conceptualization. Karthiga Santhana Kumar: Methodology, Investigation, Conceptualization. Fabien Kuttler: Methodology, Investigation, Formal analysis, Data curation. Özgün Özalp: Visualization, Validation, Methodology, Investigation. Veronica Akle: Writing – review & editing, Visualization, Validation, Supervision, Methodology, Investigation. Hanqing Zhang: Software, Methodology. Michael A. Grotzer: Writing – review & editing, Resources. Stephan C.F. Neuhauss: Writing – review & editing, Supervision, Resources, Methodology. Amin Allalou: Writing – review & editing, Visualization, Validation, Supervision, Software, Methodology, Investigation, Formal analysis, Data curation. Martin Baumgartner: Writing – review & editing, Writing – original draft, Visualization, Supervision, Resources, Project administration, Methodology, Funding acquisition, Formal analysis, Data curation, Conceptualization.
Declaration of competing interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Acknowledgments
The authors thank the Biomolecular Screening Core Facility of the EPFL and Jonathan Vesin and Julien Bortoli Chapalay for their competent technical support. We thank Ana Sofia Guerreiro Stücklin (University Children's Hospital Zürich) for providing patient samples, Sandra Laternser (University Children's Hospital Zürich) for technical assistance with patient samples, and Anastasia Emmanouilidou, Eugenia Mazzaferro, Endrina Mujica, Christoph Metzendorf and Marcel den Hoed, Uppsala University, for generating and sharing data used in training of the morphological analysis. Imaging was performed with equipment maintained by the Center for Microscopy and Image Analysis, University of Zurich. Animals were kept in the infrastructure of the Laboratory Animal Services Center (LASC), University of Zurich.
Footnotes
Supplementary material associated with this article can be found, in the online version, at doi:10.1016/j.neo.2024.101078.
Appendix. Supplementary materials
References
- 1.Wang J., Garancher A., Ramaswamy V., Wechsler-Reya R.J.Medulloblastoma. From molecular subgroups to molecular targeted therapies. Annu. Rev. Neurosci. 2018;41:207–232. doi: 10.1146/annurev-neuro-070815-013838. [DOI] [PubMed] [Google Scholar]
- 2.Cavalli FMG, Remke M, Rampasek L, Peacock J, Shih DJH, Luu B, Garzia L, Torchia J, Nor C, Morrissy AS, Agnihotri S, Thompson YY, Kuzan-Fischer CM, Farooq H, Isaev K, Daniels C, Cho BK, Kim SK, Wang KC, Lee JY, Grajkowska WA, Perek-Polnik M, Vasiljevic A, MD T. Intertumoral heterogeneity within medulloblastoma subgroups. Cancer Cell. 2017;31:737–754. doi: 10.1016/j.ccell.2017.05.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Schwalbe E.C., et al. Novel molecular subgroups for clinical classification and outcome prediction in childhood medulloblastoma: a cohort study. Lancet Oncol. 2017;18:958–971. doi: 10.1016/S1470-2045(17)30243-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Santhana Kumar K., et al. TGF-β Determines the Pro-migratory Potential of bFGF Signaling in Medulloblastoma. Cell. Rep. 2018;23:3798–3812. doi: 10.1016/j.celrep.2018.05.083. .e8. [DOI] [PubMed] [Google Scholar]
- 5.Migliavacca J, Züllig B, Capdeville C, Grotzer MA, M B. Cooperation of Striatin 3 and MAP4K4 promotes growth and tissue invasion. Commun. Bio. 2022;5:795. doi: 10.1038/s42003-022-03708-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Neve A., et al. Crosstalk between SHH and FGFR signaling pathways controls tissue invasion in medulloblastoma. Cancers (Basel) 2019;11:1–17. doi: 10.3390/cancers11121985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Garzia L, Kijima N, Morrissy AS, De Antonellis P, Guerreiro-Stucklin A, Holgado BL, Wu X, Wang X, Parsons M, Zayne K, Manno A, Kuzan-Fischer C, Nor C, Donovan LK, Liu J, Qin L, Garancher A, Liu KW, Mansouri S, Luu B, Thompson YY, Ramaswamy V, Peacock J, F T.MD. A Hematogenous Route for Medulloblastoma Leptomeningeal Metastases. Cell. 2018:172. doi: 10.1016/j.cell.2018.01.038. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Tang A., et al. Aurora kinases: Novel therapy targets in cancers. Oncotarget. 2017;8:23937–23954. doi: 10.18632/oncotarget.14893. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Willems E., et al. The functional diversity of Aurora kinases: A comprehensive review. Cell Div. 2018;13:1–17. doi: 10.1186/s13008-018-0040-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Liang B., et al. Integrated analysis of transcriptome data revealed AURKA and KIF20A as critical genes in medulloblastoma progression. Front Oncol. 2022;12:1–16. doi: 10.3389/fonc.2022.875521. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.El-Sheik A., et al. Inhibition of aurora kinase a enhances chemosensitivity of medulloblastoma cell lines. Pediatr. Blood Cancer. 2010;55:35–41. doi: 10.1002/pbc.22465. [DOI] [PubMed] [Google Scholar]
- 12.Markant S.L., et al. Targeting sonic hedgehog-associated medulloblastoma through inhibition of aurora and polo-like kinases. Cancer Res. 2013;73:6310–6322. doi: 10.1158/0008-5472.CAN-12-4258. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Diaz R.J., et al. Mechanism of action and therapeutic efficacy of Aurora kinase B inhibition in MYC overexpressing medulloblastoma. Oncotarget. 2015;6:3359–3374. doi: 10.18632/oncotarget.3245. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Kumar K.S., et al. Computer-assisted quantification of motile and invasive capabilities of cancer cells. Sci. Rep. 2015;5:15338. doi: 10.1038/srep15338. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Neve A., Kumar K.S., Tripolitsioti D., Grotzer M.A., Baumgartner M. Investigation of brain tissue infiltration by medulloblastoma cells in an ex vivo model. Sci. Rep. 2017;7:1–12. doi: 10.1038/s41598-017-05573-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Tamura K., et al. Expression of major histocompatibility complex on human medulloblastoma cells with neuronal differentiation. Cancer Res. 1989;49:5380–5384. [PubMed] [Google Scholar]
- 17.Triscott J., et al. Personalizing the treatment of pediatric medulloblastoma: Polo-like kinase 1 as a molecular target in high-risk children. Cancer Res. 2013;73:6734–6744. doi: 10.1158/0008-5472.CAN-12-4331. [DOI] [PubMed] [Google Scholar]
- 18.Cruzeiro G.A.V., et al. A simplified approach using Taqman low-density array for medulloblastoma subgrouping. Acta Neuropathol. Commun. 2019;7 doi: 10.1186/s40478-019-0681-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Mueller T, Laternser S, Guerreiro Stücklin AS, Gerber NU, Mourabit S, Rizo M, Rushing EJ, Kottke R, Grotzer M, Krayenbühl N, Nazarian J, S M. Real-time drug testing of paediatric diffuse midline glioma to support clinical decision making: The Zurich DIPG/DMG centre experience. Eur. J.Cancer. 2023;178:171–179. doi: 10.1016/j.ejca.2022.10.014. [DOI] [PubMed] [Google Scholar]
- 20.Mcquin C., et al. CellProfiler 3.0: Next-generation image processing for biology. PLoS Biol. 2018;16:1–17. doi: 10.1371/journal.pbio.2005970. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Huang W., et al. A high-throughput assay to identify allosteric inhibitors of the PLC-γ isozymes operating at membranes. Biochemistry. 2020;59:4029–4038. doi: 10.1021/acs.biochem.0c00511. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.He K., Zhang X., Ren S., Sun J. 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR) IEEE Computer Society Digital Library; Las Vegas, NV, USA: 2016. Deep residual learning for image recognition; pp. 770–778. [DOI] [Google Scholar]
- 23.Tamura K, Shimizu K, Yamada M, Okamoto Y, Matsui Y, Park KC, Mabuchi E, Moriuchi S, H M. Expression of major histocompatibility complex on human medulloblastoma cells with neuronal differentiation. Cancer Res. 1989;49:5380–5384. [PubMed] [Google Scholar]
- 24.Schönholzer M.T., et al. Real-time sensing of MAPK signaling in medulloblastoma cells reveals cellular evasion mechanism counteracting dasatinib blockade of ERK activation during invasion. Neoplasia (United States) 2020;22:470–483. doi: 10.1016/j.neo.2020.07.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.de Groot C.O., et al. A cell biologist's field guide to aurora kinase inhibitors. Front. Oncol. 2015;5:1–26. doi: 10.3389/fonc.2015.00285. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Roskoski R. Src protein-tyrosine kinase structure, mechanism, and small molecule inhibitors. Pharmacol. Res. 2015;94:9–25. doi: 10.1016/j.phrs.2015.01.003. [DOI] [PubMed] [Google Scholar]
- 27.Borah N.A., Reddy M.M. Aurora kinase B inhibition: A potential therapeutic strategy for cancer. Molecules. 2021;26:1–30. doi: 10.3390/molecules26071981. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Jiang J., et al. Direct phosphorylation and stabilization of MYC by Aurora B kinase promote T-cell Leukemogenesis. Cancer Cell. 2020;37:200–215. doi: 10.1016/j.ccell.2020.01.001. .e5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Osthus R.C., et al. Deregulation of glucose transporter 1 and glycolytic gene expression by c-Myc. J. Biol. Chem. 2000;275:21797–21800. doi: 10.1074/jbc.C000023200. [DOI] [PubMed] [Google Scholar]
- 30.Boudjadi S., Carrier J.C., Groulx J.F., Beaulieu J.F. Integrin α1β1 expression is controlled by c-MYC in colorectal cancer cells. Oncogene. 2016;35:1671–1678. doi: 10.1038/onc.2015.231. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Anderson S., et al. MYC-nick promotes cell migration by inducing fascin expression and Cdc42 activation. Proc. Natl. Acad. Sci. U S A. 2016;113:E5481–E5490. doi: 10.1073/pnas.1610994113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Gully C.P., et al. Aurora B kinase phosphorylates and instigates degradation of p53. Proc. Natl. Acad. Sci. U S A. 2012;109:1513–1522. doi: 10.1073/pnas.1110287109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Okazaki R. Role of p53 in regulating radiation responses. Life. 2022;12 doi: 10.3390/life12071099. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Hum N.R., et al. Comparative molecular analysis of cancer behavior cultured in vitro, in vivo, and ex vivo. Cancers (Basel) 2020;12 doi: 10.3390/cancers12030690. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Brulin B., et al. Evaluation of the chemotherapy drug response using organotypic cultures of osteosarcoma tumours from mice models and canine patients. Cancers (Basel) 2021;13 doi: 10.3390/cancers13194890. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Roper S.J., Coyle B. Establishing an In Vitro 3D spheroid model to study Medulloblastoma drug response and tumor dissemination. Curr. Protoc. 2022;2 doi: 10.1002/cpz1.357. [DOI] [PubMed] [Google Scholar]
- 37.Roper S.J., Linke F., Scotting P.J., Coyle B. 3D spheroid models of paediatric SHH medulloblastoma mimic tumour biology, drug response and metastatic dissemination. Sci. Rep. 2021;11:1–17. doi: 10.1038/s41598-021-83809-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Kovacs A.H., Zhao D., Hou J. Aurora B inhibitors as cancer therapeutics. Molecules. 2023;28 doi: 10.3390/molecules28083385. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Kantarjian H.M., et al. Stage i of a phase 2 study assessing the efficacy, safety, and tolerability of barasertib (AZD1152) versus low-dose cytosine arabinoside in elderly patients with acute myeloid leukemia. Cancer. 2013;119:2611–2619. doi: 10.1002/cncr.28113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Dennis M., et al. Phase i study of the Aurora B kinase inhibitor barasertib (AZD1152) to assess the pharmacokinetics, metabolism and excretion in patients with acute myeloid leukemia. Cancer Chemother. Pharmacol. 2012;70:461–469. doi: 10.1007/s00280-012-1939-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Boss D.S., et al. Clinical evaluation of AZD1152, an i.v. inhibitor of Aurora B kinase, in patients with solid malignant tumors. Ann. Oncol. 2011;22:431–437. doi: 10.1093/annonc/mdq344. [DOI] [PubMed] [Google Scholar]
- 42.Tanaka K., et al. Targeting Aurora B kinase prevents and overcomes resistance to EGFR inhibitors in lung cancer by enhancing BIM- and PUMA-mediated apoptosis. Cancer Cell. 2021;39:1245–1261. doi: 10.1016/j.ccell.2021.07.006. .e6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.He S.J., et al. Inhibition of Aurora kinases induces apoptosis and autophagy via AURKB/p70S6K/RPL15 axis in human leukemia cells. Cancer Lett. 2016;382:215–230. doi: 10.1016/j.canlet.2016.08.016. [DOI] [PubMed] [Google Scholar]
- 44.Hartsink-Segers S.A., et al. Aurora kinases in childhood acute leukemia: The promise of aurora B as therapeutic target. Leukemia. 2013;27:560–568. doi: 10.1038/leu.2012.256. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Goto H., et al. Aurora B kinase as a therapeutic target in acute lymphoblastic leukemia. Cancer Chemother. Pharmacol. 2020;85:773–783. doi: 10.1007/s00280-020-04045-9. [DOI] [PubMed] [Google Scholar]
- 46.Zekri A., Mesbahi Y., Boustanipour E., Sadr Z., Ghaffari S.H. The potential contribution of microRNAs in anti-cancer effects of aurora kinase inhibitor (AZD1152-HQPA) J. Mol. Neurosci. 2018;65:444–455. doi: 10.1007/s12031-018-1118-y. [DOI] [PubMed] [Google Scholar]
- 47.Peyrl A., et al. Safety of Ommaya reservoirs in children with brain tumors: A 20-year experience with 5472 intraventricular drug administrations in 98 patients. J. Neurooncol. 2014;120:139–145. doi: 10.1007/s11060-014-1531-1. [DOI] [PubMed] [Google Scholar]
- 48.Palmisciano P., et al. Intrathecal therapy for the management of leptomeningeal metastatic disease: a scoping review of the current literature and ongoing clinical trials. J. Neurooncol. 2022;160:79–100. doi: 10.1007/s11060-022-04118-0. [DOI] [PubMed] [Google Scholar]
- 49.Ramaswamy V., Nör C., Taylor M.D. P53 and Meduloblastoma. Cold Spring Harb. Perspect. Med. 2016;6:1–9. doi: 10.1101/cshperspect.a026278. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Shrestha S., Morcavallo A., Gorrini C., Chesler L. Biological role of MYCN in Medulloblastoma: Novel therapeutic opportunities and challenges ahead. Front. Oncol. 2021;11:1–19. doi: 10.3389/fonc.2021.694320. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Roussel M.F., Robinson G.W. Role of MYC in medulloblastoma. Cold Spring Harb. Perspect. Med. 2013;3:1–15. doi: 10.1101/cshperspect.a014308. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Jiang J., et al. Direct phosphorylation and stabilization of MYC by Aurora B Kinase promote T-cell Leukemogenesis. Cancer Cell. 2020;37:200–215. doi: 10.1016/j.ccell.2020.01.001. .e5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Li F., et al. AURKB/CDC37 complex promotes clear cell renal cell carcinoma progression via phosphorylating MYC and constituting an AURKB/E2F1-positive feedforward loop. Cell Death Dis. 2024;15 doi: 10.1038/s41419-024-06827-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Duan L., Maki C.G. Determinants of Aurora kinase B inhibitor sensitivity in small cell lung cancer. Transl. Lung Cancer Res. 2024;13:223–228. doi: 10.21037/tlcr-23-732. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Ivanov D.P., Coyle B., Walker D.A., Grabowska A.M. In vitro models of medulloblastoma: Choosing the right tool for the job. J. Biotechnol. 2016;236:10–25. doi: 10.1016/j.jbiotec.2016.07.028. [DOI] [PubMed] [Google Scholar]
- 56.Marxer M., Ma H.T., Man W.Y., Poon R.Y.C. P53 deficiency enhances mitotic arrest and slippage induced by pharmacological inhibition of Aurora kinases. Oncogene. 2014;33:3550–3560. doi: 10.1038/onc.2013.325. [DOI] [PubMed] [Google Scholar]
- 57.Tao Y., et al. Enhancement of radiation response by inhibition of Aurora-A kinase using siRNA or a selective Aurora kinase inhibitor PHA680632 in p53-deficient cancer cells. Br. J. Cancer. 2007;97:1664–1672. doi: 10.1038/sj.bjc.6604083. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Philipova T., et al. Differential forms of p53 in medulloblastoma primary tumors, cell lines and xenografts. Int. J. Oncol. 2011;38:843–849. doi: 10.3892/ijo.2010.884. [DOI] [PubMed] [Google Scholar]
- 59.Naeem A., et al. Regulation of chemosensitivity in human Medulloblastoma Cells by p53 and the PI3 Kinase signaling pathway. Mol. Cancer Res. 2022;20:114–126. doi: 10.1158/1541-7786.MCR-21-0277. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Zhu G., et al. AURKB targets DHX9 to promote hepatocellular carcinoma progression via PI3K/AKT/mTOR pathway. Mol. Carcinog. 2024:1814–1826. doi: 10.1002/mc.23775. [DOI] [PubMed] [Google Scholar]
- 61.Lafay-Cousin L., Dufour C. High-Dose Chemotherapy in Children with Newly Diagnosed Medulloblastoma. Cancers (Basel) 2022;14:1–14. doi: 10.3390/cancers14030837. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The data will be made available upon reasonable request.