Abstract
B lymphopoiesis is orchestrated by lineage-specific transcription factors. In B cell progenitors, lineage commitment is mediated by Pax5, which is commonly mutated in B cell acute lymphoblastic leukemia. Despite its essential role in immunity, the mechanisms regulating Pax5 function remain largely unknown. Here, we found that the NAD+-dependent enzyme SIRT7 coordinates B cell development through deacetylation of Pax5 at K198, which promotes Pax5 protein stability and transcriptional activity. Neither Pax5K198 deacetylated nor acetylated mimics rescued B cell differentiation in Pax5−/− pro-B cells, suggesting that B cell development requires Pax5 dynamic deacetylation. The Pax5K198 deacetylation mimic restored lineage commitment in Pax5−/− pro-B cells and B cell differentiation in Sirt7−/− pro-B cells, suggesting the uncoupling of differentiation from lineage commitment. The SIRT7–Pax5 interplay was conserved in B cell acute lymphoblastic leukemia, where SIRT7 expression correlated with good prognosis. Our findings reveal a crucial mechanism for B lymphopoiesis and highlight the relevance of sirtuins in immune function.
Subject terms: B cells, Acetylation, Leukaemia
Gámez-García et al. show that the deacetylase SIRT7 modulates the acetylation of Pax5 and its ability to repress alternate lineage programs and promote B cell differentiation and commitment in B cell progenitor cells.
Main
Lineage-instructive transcription factors establish the gene regulatory networks that drive differentiation of hematopoietic progenitors toward all mature immune cell types. Among these, the transcription factor Pax5 regulates B cell identity1. Following specification, Pax5 orchestrates downstream differentiation and drives lineage commitment by promoting simultaneous activation of B cell-specific programs and repression of alternative-lineage genes2–4. Pax5 deficiency in mice results in a complete block in B cell differentiation at the pro-B cell stage, leading to the accumulation of uncommitted cells5–7. In human B cell acute lymphoblastic leukemia (B-ALL), which is the most common cancer in children, Pax5 is a haploinsufficient tumor suppressor, and the deletion of one Pax5 allele in mice cooperates with oncogenic mutations and with heterozygosity of the transcription factor Ebf1 to drive malignant transformation8–11.
In B cells, Pax5 controls the global genome architecture12 while simultaneously limiting pro-B cell proliferation through repression of Myc13, restricting cellular metabolism14 and facilitating loop extrusion of the whole immunoglobulin heavy chain (Igh) locus15–17. Pax5 collaborates with chromatin remodeling coactivator and co-repressor complexes and with other B lineage transcription factors to establish a self-reinforcing network that drives differentiation18,19. Furthermore, Pax5 dynamically binds to various genomic regions and regulates distinct target genes in developing and mature B cells20. Although the importance of Pax5 for B lymphopoiesis and human leukemia is well established, the mechanisms that control Pax5 biology and dynamics are still unclear.
Sirtuins are a family of conserved NAD+-dependent deacetylases implicated in mammalian immunity21 that regulate inflammation, T cell polarization and responses and hematopoietic stem cell exhaustion through different mechanisms, including the deacetylation of histone and nonhistone substrates22. Here, we show that SIRT7, a sirtuin with central roles in genome stability and stress response23, controls Pax5 activity during B cell lymphopoiesis and human B-ALL. SIRT7 directly promotes B cell development and commitment by establishing an acetylation switch at a single Pax5 residue located within a putative intrinsically disordered region. Our results indicate that post-translational modifications of lineage-instructive transcription factors may be sufficient to disrupt lineage commitment and suggest key roles for sirtuins in the physiology and malignancy of hematopoietic progenitors.
Results
SIRT7 is required for normal B cell development
To understand the role of sirtuins in hematopoiesis, we analyzed the expression levels of sirtuins Sirt1, Sirt2, Sirt6 and Sirt7 in Lin−(Ter119−CD11b−B220−Gr-1−) progenitors using publicly available single-cell RNA-sequencing (scRNA-seq) data from mouse bone marrow (BM) cells (Broad Institute, study SCP978). Among them, Sirt7 displayed the highest expression, particularly in the cluster annotated as B cells (Fig. 1a and Extended Data Fig. 1a). In publicly available data from human BM cells (Broad Institute, study SCP101), SIRT7 was highly expressed in Cd79a+Ighm+ B lineage cells, Nkg7+Klrf1+ natural killer (NK) cells and Trbc2+Cd3d+ T cells (Extended Data Fig. 1b,c). SIRT7 protein was highly expressed in BM CD19+ B cells purified by magnetic-activated cell separation compared to in the CD19− fraction (Fig. 1b). In mouse B cell progenitors, Sirt7 expression peaked at the committed B220+CD19+IgM−CD43− pre-B cell stage (Extended Data Fig. 1d,e). Sirt2 expression increased to a lesser extent in the same stage, whereas the expression of other sirtuin transcripts did not change during B cell differentiation (Extended Data Fig. 1d,e). Intracellular flow cytometry of mouse BM B220+ B cell progenitor cells showed a gradual upregulation of SIRT7 expression from B220+CD19− pre-pro-B cells to B220+CD19+IgM−CD43− pre-B cells and was maintained in B220+CD19+IgM+ immature and mature B220hiCD19+ B cells (Fig. 1c,d and Extended Data Fig. 1f). These observations indicated that SIRT7 expression was upregulated during B lymphopoiesis.
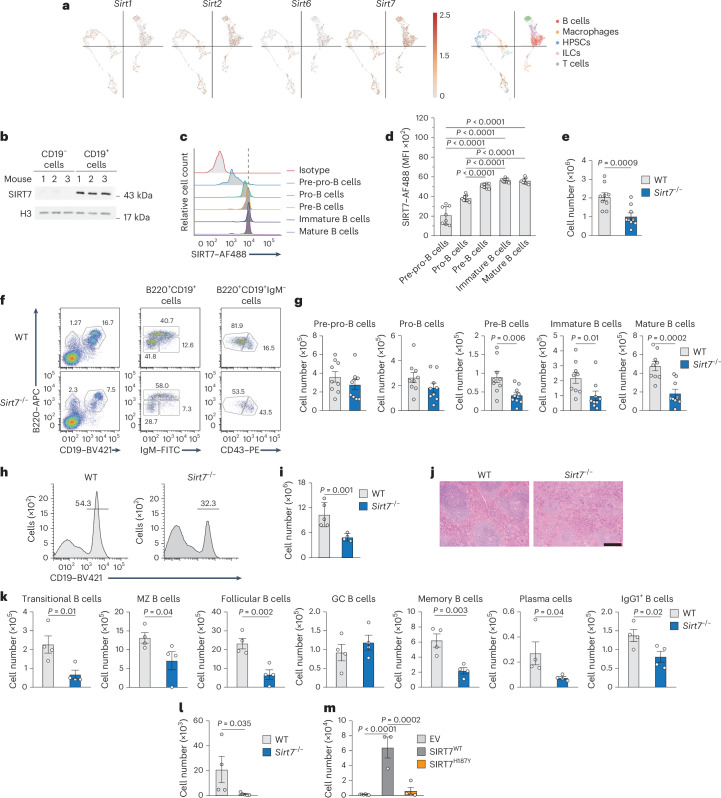
Fig. 1. SIRT7 is required for normal B cell development.
a, t-Distributed stochastic neighbor embedding plots displaying the single-cell expression profiles of nuclear sirtuins in purified mouse BM Lin– cells (left) and scRNA-seq feature plot (right) identifying B cells, macrophages, hematopoietic and progenitor stem cells (HPSCs), innate lymphoid cells (ILCs) and T cells. Data were obtained from singlecell.broadinstitute.org (study SCP978). b, Immunoblot of SIRT7 expression in mouse BM CD19− cells and CD19+ B cells (n = 3). c,d, Representative histograms (c) and quantification of SIRT7 median fluorescence intensity (MFI; d) in B220+CD19– pre-pro-B cells, B220+CD19+IgM−CD43+ pro-B cells, B220+CD19+IgM−CD43− pre-B cells, B220+CD19+IgM+ immature B cells and B220hiCD19+ mature B cells measured by intracellular flow cytometry (n = 8 mice). e, Total number of B220+CD19+ B cells in the BM of wild-type and Sirt7Δ4–10 129Sv mice (n = 9). Data were pooled from four independent experiments. f,g, Representative histograms (f) and numbers (g) of pre-pro-B cell, pro-B cell, pre-B cell, immature B cell and mature B cell populations identified as in c in the BM of 129Sv wild-type and Sirt7Δ4–10 mice (n = 9). Data were pooled from four independent experiments. h,i, Representative histograms (h) and numbers (i) of splenic CD19+ B cells from wild-type (n = 5) and Sirt7−/− (n = 4) mice. j, Hematoxylin and eosin staining of histological sections from the spleens of wild-type and Sirt7−/− mice (n = 5); scale bar, 500 µm. k, Number of splenic B220+CD19+CD21+CD23+CD93+ transitional B cells, B220+CD19+CD21hiCD23− marginal zone (MZ) B cells, B220+CD19+CD21+CD23+CD93− follicular B cells, B220+CD19+IgM+Gl7+Fas+ germinal center (GC) B cells, B220+IgG1+ class-switched B cells and BM CD19+CD38+CD138−Gl7− memory B cells and B220loCD138+ plasma cells from wild-type and Sirt7−/− mice (n = 4). l,m, Numbers of donor-derived CD45.1–CD45.2+CD19+ B cells in the spleens of recipient CD45.1/CD45.2 mice 4 weeks after congenic transplantation of either CD45.2 wild-type (n = 4) or Sirt7−/− (n = 5) Lin−B220+CD19+IgM− pro-B cells expanded ex vivo with OP9 cells and 10 ng ml–1 IL-7, SCF and FLT3-L (l) or Sirt7−/− Lin−B220+CD19+IgM− pro-B cells retrovirally expressing empty vector (EV; n = 6), SIRT7WT (n = 3) or SIRT7H187Y (n = 4; m). Data are presented as mean ± s.d. (d, i, and k–m) or mean ± s.e.m. (e, g, l and m) and were analyzed by one-tailed t-test (e, g, i, k and l) or one-way analysis of variance (ANOVA) with Sidak multiple comparisons (d and m).
Extended Data Fig. 1. SIRT7 is upregulated in B cell progenitors.
a, Heatmap displaying the single-cell expression of Sirt1, Sirt2, Sirt6 and Sirt7 in the populations (B cells, innate lymphoid cells (ILCs), macrophages (Mac), hematopoietic and progenitor stem cells (HPSC) and T cells) annotated by sc-RNA-seq from purified murine BM Lin− cells. Data was obtained from singlecell.broadinstitute.org (study SCP978). b,c, sc-RNA-seq feature plot (b) and single-cell expression of Sirt7 (c) in B cells, T cells, NK cells, plasma cells, dendritic cells, plasmacytoid dendritic cells, platelets and erythroid cells of human BM. Data was obtained from singlecell.broadinstitute.org (study SCP101). In panel c, only those cells with detectable counts are plotted, and the dashed lines indicate the median and the first and fourth quartiles. Statistical significance was assessed by one-way ANOVA with Dunnet comparison. d, Microarray expression of Sirt1 (top left), Sirt2 (top right), Sirt6 (bottom left) and Sirt7 (bottom right) in B lymphopoiesis stages (n = 3; common lymphoid progenitors (CLP), n = 2). Data are presented as mean ± s.d. A two-tailed t-test was performed comparing the expression in pooled CLP–pro-B and in pooled pre-B–plasma cells. Data were obtained from the Immgen consortium (GSE15907). e, RNA-Seq expression patterns of Sirt1, Sirt2, Sirt6 and Sirt7 in pro-B cells and pre-B cells. Data are presented as the mean of the two replicates (n = 2). f, Gating strategy defining B cell subsets (B220+CD19− pre-pro-B cells, B220+CD19+IgM−CD43+ pro-B cells, B220+CD19+IgM−CD43− pre-B cells, B220+CD19+IgM+ immature B cells and B220hiCD19+ mature B cells) for intracellular flow cytometry measurement of SIRT7 protein levels in fixed mouse BM.
To determine the role of SIRT7 in B cell development, we measured the number of B220+CD19+ B cells in the BM of two reported models of Sirt7−/− mice, Sirt7Δ4−10 129Sv and Sirt7Δ4–9 C57BL/6 mice (Methods)24,25. In both strains, Sirt7−/− mice displayed a significant reduction in the number of B220+CD19+ B cells compared to wild-type littermates (used throughout unless otherwise specified; Fig. 1e and Extended Data Fig. 2a). We also found decreased numbers of B220+CD19+IgM−CD43− pre-B cells, B220+CD19+IgM+ immature B cells and B220hiCD19+ recirculating mature B cells (Fig. 1f,g and Extended Data Fig. 2b), indicative of an impaired pro-B-to-pre-B cell transition. We used the Sirt7Δ4–10 129Sv model, hereafter referred to as Sirt7−/−, throughout for all experiments. The spleens of Sirt7−/− mice exhibited a contraction of the CD19+ cell compartment (Fig. 1h,i) and compromised structural architecture, with fewer and less-organized follicles than spleens in the wild-type mice (Fig. 1j). There was a significant reduction in the number of B220+CD19+CD21+CD23+CD93+ transitional, B220+CD19+CD21hiCD23− marginal zone and B220+CD19+CD21+CD23+CD93− follicular B cells in Sirt7−/− mice compared to in wild-type mice (Fig. 1k and Extended Data Fig. 2c). The number of B220+CD19+IgM+Gl7+Fas+ germinal center B cells in Sirt7−/− mice was normal, whereas the number of CD19+CD38+CD138−Gl7− memory B cells, B220loCD138+ plasma cells and B220+ class-switched IgG1+ B cells was reduced compared to wild-type mice (Fig. 1k and Extended Data Fig. 2d–g), suggesting a defective germinal center response in Sirt7−/− mice. The levels of hen egg lysozyme (HEL)-specific IgM, IgG1 and IgG3 antibodies in serum at day 14 after immunization with HEL antigen coupled to NPP hapten (NP–HEL) were significantly reduced in Sirt7−/− mice compared to wild-type mice (Extended Data Fig. 2h), suggesting that SIRT7 deficiency leads to impaired B cell development and immunity.
Extended Data Fig. 2. SIRT7 is required for normal B cell development.
a,b, Number of B220+CD19+ B cells (a) and percentages of B220+CD19−pre-pro-B cells, B220+CD19+IgM−CD43+ pro-B cells, B220+CD19+IgM−CD43− pre-B cells and B220+CD19+IgM+ immature B cells (b) determined by flow cytometry in the BM of wild-type and Sirt7 Δ4-9 C57BL/6 mice. Data are presented as mean ± s.d. and significance was assessed by two-tailed t-test (wild-type, n = 5 (a) and 7 (b); Sirt7 Δ4-9, n = 9 (a) and 7 (b)). Pooled from two independent experiments. c-g, Representative FACS plots of splenic B220+CD19+CD21highCD23− marginal zone B cells, B220+CD19+CD21+CD23+CD93−follicular B cells and B220+CD19+CD21+CD23+CD93+ transitional B cells. (c), B220+CD19+IgM+Gl7+Fas+ germinal center B cells (d), BM CD19+CD38+CD138−Gl7+memory B cells (e), BM B220loCD138+plasma cells (f) and class-switched splenic B220+IgG1+ B cells (g) from wild-type and Sirt7−/− mice (n = 4). h, Levels of anti-HEL antibody isotypes in the sera of wild-type and Sirt7−/− mice 14 days after NP-HEL immunization, relative to naive mice, as determined by ELISA. Data are shown as in a (wild-type, n = 6; Sirt7−/−, n = 5). One of two separate experiments is shown. i, Gating strategy used to identify donor-derived CD45.1-CD45.2+CD19+ mature B cells in the spleen of recipient CD45.1/CD45.2 mice four weeks after transplantation of CD45.2 wild-type or Sirt7−/− Lin−B220+CD19+IgM− pro-B cells.
To test the contribution of the BM stroma in Sirt7−/− mice, purified Lin−B220+CD19+IgM− pro-B cells from the BM of CD45.2 wild-type and Sirt7−/− mice were expanded in the presence of OP9 stromal cells, interleukin-7 (IL-7), stem cell factor (SCF) and FLT3-L for 4 days and injected into sublethally irradiated CD45.1/CD45.2 mice to generate separate wild-type and Sirt7−/− BM chimeras. Wild-type pro-B cells repopulated the splenic CD45.1–CD45.2+CD19+ B cell compartment of recipient mice, whereas Sirt7−/− pro-B cells did not (Fig. 1l and Extended Data Fig. 2i), indicating that SIRT7 had a B cell-autonomous role. Retroviral expression of wild-type SIRT7 into Sirt7−/− pro-B cells injected into sublethally irradiated CD45.1/CD45.2 mice reversed the B cell differentiation block, whereas expression of a catalytically inactive SIRT7H187Y mutant did not (Fig. 1m). These findings indicate that SIRT7 promotes pro-B-to-pre-B cell transition through a mechanism dependent on its deacetylase activity.
SIRT7 promotes B cell development independently of V(D)J
SIRT7 facilitates DNA damage repair by controlling nonhomologous end joining24. Sirt7−/− BM pre-B cells, but not Sirt7−/− pro-B cells, showed a twofold increase in 7AAD+Annexin-V+ apoptotic cells (Extended Data Fig. 3a). Sirt7−/− B220+CD19+IgM−CD43−FSChi large pre-B cells were partially arrested at G1 (Extended Data Fig. 3b) and displayed reduced phosphorylation of STAT5 at Y694, the major driver of pre-B cell proliferation26, without altering the expression of the upstream IL-7 receptor subunit CD127 (Extended Data Fig. 3c,d). SIRT7 recruits 53BP1, a key player in distal Vh-to-DJh recombination, to double strand breaks24,27. Splenic IgM+ Sirt7−/− B cells exhibited reduced usage of distal Vh588 and Vh7183 segments, whereas the proximal segments DhL-to-Jh3 and VhQ52-DJh remained unaffected (Extended Data Fig. 3e,f). However, crossing wild-type and Sirt7−/− mice with mice expressing a transgenic HEL-specific immunoglobulin (IgHEL)28, which bypasses the V(D)J-related checkpoints, did not restore B cell differentiation in Sirt7−/− mice (Extended Data Fig. 3g). These observations indicate that SIRT7 promotes B cell development through a mechanism independent of V(D)J recombination.
Extended Data Fig. 3. SIRT7 promotes B cell development independently of V(D)J.
a, Representative apoptosis FACS plots (top) and relative numbers (bottom) of 7AAD+Annexin V+ cells (gated on B220+CD19+IgM−CD43+ pro-B and B220+CD19+IgM−CD43− pre-B cells) in the BM of wild-type (n = 3) and Sirt7−/− (n = 4) mice. Data was normalized to wild-type values. b, Representative histograms (left) and quantification (right) of the percentages of cells in G1, S and G2/M cell cycle stages of gated wild-type and Sirt7−/− B220+CD19+IgM−CD43−FSChi large and B220+CD19+IgM−CD43−FSClo small pre-B cells, determined by intracellular flow cytometry (n = 4 mice). Pooled from two separate experiments. c, Representative histogram (left) and p-STAT5Y694 MFI (right) in gated wild-type and Sirt7−/− B220+CD19+IgM−CD43− pre-B cells, measured by intracellular flow cytometry (n = 3 mice; p = 0.005). Pooled from three separate experiments d, Flow cytometric quantification of CD127 expression in gated B220+CD19+IgM−CD43− pre-B cells from the BM of wild-type and Sirt7−/− mice (n = 5). Pooled from three separate experiments. e, Schematic representation of the mouse IgH locus, showing constant (C), joining (J), diversity (D) and variable (V) gene segment organization (top) and semi-quantitative genomic DNA PCR using degenerate primers targeting DHL-JH3 and VH-DJH gene segments in IgM+ B cells sorted from the BM of wild-type and Sirt7−/− mice. Mouse embryonic fibroblasts (MEFs) and H2O were used as negative controls of PCR amplification. The independent locus Kat8 was used as a loading control (n = 5 mice). f, Densitometric quantification of DHL-JH3, VHJ588-DJH3, VH7183-DJH3 and VHQ52-DJH3 gene segment amplification in the mouse IgH locus normalized to Kat8 in sorted splenic wild-type and Sirt7−/− IgM+ B cells (n = 5 mice). g, Representative flow cytometry plots (left) and number of B220+CD19+ B cells (right) in the BM of IgHEL−/− (n = 5) IgHEL−/−Sirt7−/− (n = 5), IgHEL+/- (n = 6) and IgHEL+/-Sirt7−/− (n = 6) mice. Data in a, b, c, d, f and g are shown as mean ± s.d. Data in a, c and f were analyzed using one-tailed t-test, and data in b and g were analyzed using two-way ANOVA with Sidak multiple comparisons.
SIRT7 regulates Pax5 in B cell progenitors
To investigate the mechanisms through which SIRT7 regulates B cell development, we performed RNA-seq in sorted wild-type and Sirt7−/− B220+CD19+IgM−CD43+ pro-B cells and B220+CD19+IgM−CD43− pre-B cells. In a principal component analysis (PCA), wild-type and Sirt7−/− pre-B cells clustered at a greater distance than wild-type and Sirt7−/− pro-B cells (Fig. 2a). SIRT7 regulated the expression of 220 genes in pro-B cells and 429 genes in pre-B cells (false discovery rate (FDR) < 0.05), including genes important for B cell development (pro-B: Vpreb2, Ikzf3 and Irf4; pre-B: Vpreb1, Runx2, Erg, Mef2c, Rag1 and Bcl2; Supplementary Table 1). Gene set enrichment analysis (GSEA) indicated that SIRT7 mainly repressed lineage-inappropriate (Thy1 and Il2ra) and cell cycle-related genes (Myc and Grb7), such as ‘lung NK and T cell’ and ‘E2F targets’ gene sets (Fig. 2b,c and Supplementary Table 1). Accordingly, T cell-related genes, such as Trbc1, Trbc2, Zap70, Thy1, Cd3e and Tigit, were derepressed in ex vivo-expanded Sirt7−/− B cell progenitors (Fig. 2d).
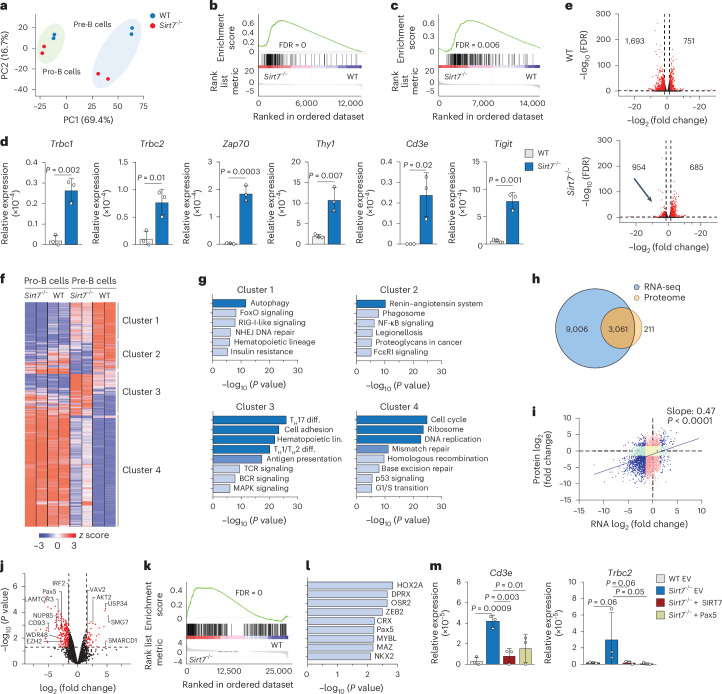
Fig. 2. SIRT7 regulates Pax5 in B cell progenitors.
a, PCA clustering of B220+CD19+IgM−CD43+ pro-B cells and B220+CD19+IgM−CD43− pre-B cells sorted from the BM of wild-type and Sirt7−/− 129Sv mice based on the top 2,000 differentially expressed genes (n = 2); PC, principal component. b,c, GSEA of the ‘Travaglini lung proliferating NK T cell’ (b) and ‘Hallmark E2F targets’ (c) gene sets in Sirt7−/− versus in wild-type B220+CD19+IgM−CD43− pre-B cells. d, RT–qPCR of Trbc1, Trbc2, Zap70, Thy1, Cd3e and Tigit expression relative to Hprt (n = 3). e, Volcano plot of differentially expressed genes in pre-B cells versus in pro-B cells (defined as in a) from wild-type (top) and Sirt7−/− (bottom) mice. Numbers on the right and left represent significantly induced or repressed genes, respectively. The black arrow indicates the lack of downregulated genes. Red dots represent significantly regulated genes (| log2 (fold change) | > 1.5, FDR < 0.05). f, Unsupervised clustering of the top 2,000 differentially expressed genes (P < 0.03) in wild-type and Sirt7−/− pro-B and pre-B cells defined as in a. g, Gene Ontology analyses of genes in clusters 1–4. Only terms with a –log10 (P value) of >5 are reported; NHEJ, nonhomologous end joining; TH17 diff., differentiation of IL-17-producing helper T cells; lin., lineage; TH1/TH2 diff., differentiation of type 1 helper T/type 2 helper T cells; TCR, T cell antigen receptor; BCR, B cell receptor. h,i, Venn diagram displaying the number of detected proteins and transcripts (h) and scatter plot (i) showing the linear correlation between the mean log2 (fold change) in protein expression and their corresponding transcript levels in wild-type B220+CD19+IgM−CD43− pre-B cells, as determined by proteomics and RNA-seq, respectively. i, Changes in protein and RNA levels with a | log2 (fold change) | of ≥1.5 (blue), changes in protein levels only (light red), changes in RNA levels only (light green) and changes of | log2 (fold change) | ≤1.5 (yellow) are shown. Fold change values were calculated relative to Nup50 expression. The Spearman correlation slope = 0.47; P < 0.0001 between protein and transcript pairs. j, Volcano plot of differentially expressed proteins in sorted Sirt7−/− versus wild-type B220+CD19+IgM−CD43− pre-B cells. Red dots indicate | log2 (fold change) | of ≥1.5 and a P value of <0.05. k, GSEA of Sirt7−/− versus wild-type B220+CD19+IgM−CD43− pre-B cells compared to Pax5 target genes20. l, HOMER de novo motif enrichment analysis showing nine of the top transcription factors enriched in cluster 3 genes. Motif enrichment was performed on promoter regions (transcription start site to 2,000 bp upstream). m, RT–qPCR of Cd3e and Trcb2 relative to Hprt in wild-type or Sirt7−/− B220+CD19+IgM− pro-B cells retrovirally expressing empty vector, SIRT7 or Pax5 (n = 3). Data are shown as mean ± s.d. (d and m) and were analyzed by one-tailed t-test (i), two-tailed t-test (d) or one-way ANOVA with Fisher’s least significant different (LSD) test (m).
Volcano plots indicated that Sirt7−/− pre-B cells induced gene expression normally but repressed fewer genes than wild-type pre-B cells (Fig. 2e and Extended Data Fig. 4a). We identified four gene clusters regulated by SIRT7 (Fig. 2f). Genes within clusters 1 and 2 showed lower expression in Sirt7−/− pro-B and pre-B cells than in their wild-type counterparts and were involved in autophagy, nonhomologous end joining DNA repair and cellular signaling. Cluster 3 genes were increased in Sirt7−/− pre-B cells relative to wild-type pre-B cells and were enriched for genes involved in T cell differentiation, suggesting compromised lineage commitment in Sirt7−/− pre-B cells. Cluster 4 comprised genes similarly expressed in wild-type and Sirt7−/− pro-B cells that were downregulated in wild-type pro-B cells but were only partially reduced in Sirt7−/− pre-B cells, including genes implicated in DNA repair and the cell cycle (Fig. 2g).
Extended Data Fig. 4. SIRT7 regulates Pax5 in B cell progenitors.
a, Pie charts showing the number and percentage of genes significantly induced and repressed (|log2(fold-change)| ≥ 1.5, FDR < 0.05) during the pre-B-to-pro-B cell transition in wild-type (left) and Sirt7−/− (right) mice (n = 2). b, Percentages of significantly regulated (|log2(fold change)| ≥ 1, FDR < 0.1) transcripts and proteins in Sirt7−/− vs wild-type B220+CD19+IgM−CD43− pre-B cells, relative to all the transcripts and proteins detected by RNA-Seq and proteomics, respectively. c, Gene ontology analysis of significantly regulated proteins in Sirt7−/− vs wild-type B220+CD19+IgM−CD43− pre-B cells. d, Percentage of Cluster 3 genes from wild-type and Sirt7−/− B220+CD19+IgM−CD43− pre-B cell and B220+CD19+IgM−CD43+ pro-B cell bulk RNA-Seq data with significant (q < 0.05) Pax5 peaks17 in their promoters only (±2Kb from TSS) (left panel) or in the whole gene (including ± 2Kb from TSS, gene bodies and 3’UTR, right panel).
Quantitative mass spectrometry (MS) in sorted wild-type and Sirt7−/− B220+CD19+IgM−CD43− pre-B cells identified 2,917 proteins (Supplementary Table 2), most of which were also identified by bulk RNA-seq analysis of the same cells (Fig. 2h). mRNA and protein expression levels positively correlated for most transcript–protein pairs (Fig. 2i), as previously reported for B-ALL29. Although the RNA-seq analysis identified differential expression for 1.6% of all detected transcripts between wild-type and Sirt7−/− pre-B cells, 5.2% of the proteome (Extended Data Fig. 4b), including proteins involved in DNA repair, cell cycle and intracellular signaling (Extended Data Fig. 4c), was differentially expressed.
Among the top differentially expressed proteins between wild-type and Sirt7−/− pre-B cells, we found several transcriptional and chromatin regulators (Pax5, IRF2, EZH2 and SMARCD1; Fig. 2j). Pax5 was downregulated 2.8-fold in Sirt7−/− pre-B cells compared to wild-type cells (Fig. 2j and Supplementary Table 2). GSEA of the Sirt7−/− pre-B cell bulk RNA-seq dataset indicated that SIRT7 and Pax5 controlled a similar group of genes, including signaling genes, transcriptional regulators and surface receptors, such as Vpreb1, Runx2, Bach2, Il2ra and Thy1 (Fig. 2k), whereas unsupervised de novo motif discovery analysis identified enrichment of the Pax5 motif in cluster 3 genes (Fig. 2l). Analysis of publicly available chromatin immunoprecipitation with sequencing (ChIP–seq) datasets17 revealed that Pax5 bound to 62% of cluster 3 genes and to 43% of their promoters (Extended Data Fig. 4d). Retroviral expression of either SIRT7 or Pax5 in wild-type or Sirt7−/− BM CD19+ cells restored Cd3e and Trcb2 repression in Sirt7−/− CD19+ cells (Fig. 2m), indicating that Pax5 expression bypassed SIRT7 deficiency to repress lineage-inappropriate genes. According to bulk RNA-seq datasets of wild-type and Sirt7−/− pro-B and pre-B cells, SIRT7 also regulated transcriptional programs not associated with Pax5, such as DNA replication and repair and ribosome biogenesis (Fig. 2g). These observations suggest that SIRT7 collaborates with Pax5 to repress transcription of lineage-inappropriate genes during early B cell development.
SIRT7 regulates Pax5 stability by deacetylating K198
Intracellular flow cytometry of B220+CD19+IgM−CD43+ pro-B cells and B220+CD19+IgM−CD43− pre-B cells and immunoblotting of B220+CD19+IgM− pro-B cells expanded in vitro indicated reduced amounts of Pax5 protein in Sirt7−/− pro-B and pre-B cells compared to their wild-type counterparts (Fig. 3a,b and Extended Data Fig. 5a). By contrast, RNA-seq and quantitative PCR with reverse transcription (RT–qPCR) on the same cells showed normal Pax5 mRNA expression (Fig. 3c and Extended Data Fig. 5b), indicating that SIRT7 directly regulates Pax5 protein levels. SIRT7 retroviral expression in Sirt7−/− pro-B cells rescued Pax5 protein expression (Fig. 3d and Extended Data Fig. 5c). To test whether SIRT7 controlled Pax5 turnover, we used CRISPR–Cas9 to knock out Sirt7 in the mouse pre-B cell line HAFTL (HAFTLSIRT7KO cells). Pax5 was downregulated in HAFTLSIRT7KO cells compared to in HAFTL cells, which was fully reversed by treatment with the proteasome inhibitor lactacystin (Fig. 3e and Extended Data Fig. 5d). Cycloheximide protein stability assays in HAFTLSIRT7KO compared to in HAFTL cells indicated that Pax5 half-life was reduced by twofold in the former (Fig. 3f,g), indicating that SIRT7 prevents Pax5 proteasomal degradation. SIRT7 interacted with endogenous Pax5 in HAFTL cells and with transiently expressed PAX5 in HEK293F cells (Fig. 3h and Extended Data Fig. 5e). In gel filtration chromatography of HAFTL nuclear extracts, a subpopulation of SIRT7 and Pax5 coeluted in high-molecular-weight (≈1 MDa) fractions (Fig. 3i), in line with previous observations that Pax5 works in concert with other transcription factors and epigenetic regulators18.
Fig. 3. SIRT7 regulates Pax5 stability by deacetylating K198.
a, Pax5 MFI measured by intracellular flow cytometry of B220+CD19+IgM−CD43+ pro-B and B220+CD19+IgM−CD43− pre-B cells from the BM of wild-type (n = 6) and Sirt7−/− (n = 8) mice. Data were pooled from three independent experiments. b, Immunoblot of Pax5 protein in wild-type and Sirt7−/− B220+CD19+IgM− pro-B cells expanded ex vivo for 4 days with OP9 cells and 10 ng ml–1 IL-7, SCF and FLT3-L. c, RT–qPCR analysis of Pax5 gene expression in wild-type and Sirt7−/− pro-B cells expanded as in b (n = 3). d, Immunoblot of Pax5 and SIRT7 protein in wild-type and Sirt7−/− B220+CD19+IgM− pro-B cells retrovirally transduced with empty vector and SIRT7. e, Quantification of Pax5 protein expression in wild-type and CRISPR–Cas9-generated HAFTLSIRT7KO cells treated with vehicle (Ct) or 2 µM lactacystin for 8 h (n = 3 biological replicates). f,g, Time course immunoblot (f) and Pax5/H3 ratio (g) in HAFTL (n = 6 independent time courses) and HAFTLSIRT7KO (n = 8 independent time courses) cells treated with 100 μg ml–1 cycloheximide for 3, 6, 9 and 24 h. Nonlinear fits with variable slope (four parameters) are depicted. h, Flag-specific coimmunoprecipitation from HAFTL cells retrovirally transduced with empty vector or SIRT7–Flag, followed by Pax5 and SIRT7 immunoblotting; IP, immunoprecipitate. i, Immunoblot of Pax5 and SIRT7 in the input and gel filtration chromatography fractions in HAFTL cells. Approximate molecular weights are shown. Red brackets indicate fractions in which Pax5 and SIRT7 coelute. j, Immunoblot of Pax5 protein in HAFTL and KOPN-8 cells treated with vehicle (–) or 5 mM nicotinamide (NAM) for 48 h. k, Pan-acetyl-lysine (AcK) immunoblotting of in vitro deacetylation assays with purified Pax5 alone, with SIRT7 or SIRT7 and NAD+. l, Fragmentation MS/MS spectra of the KacRDEGIQ(+.98)ESPVPNGHSLPGR peptide, as determined by proteomic analysis of Pax5 protein immunoprecipitated from SIRT7−/− HEK293F cells transiently expressing Pax5 and empty vector (left) or Pax5 and SIRT7 (right) by polyethylenimine transfection. m, Schematic of mouse Pax5 functional domains; PRD, paired box domain; OP, octapeptide domain; HD, partial homeodomain; TAD, transactivation domain; ID, inhibitory domain. The start and end of each domain and the detected acetyl-lysine residues are indicated. n, Conservation of Pax5 186–229 peptide in Mus musculus, Homo sapiens, Gallus gallus, Danio rerio and Xenopus laevis. K198 is shown in red. o, Expression of Pax5WT and Pax5K198Q and Pax5K198R mutants expressed in HEK293F cells as in l. p, Ubiquitination of Flag-purified Pax5WT and Pax5K198Q and Pax5K198R mutants in HEK293F cells coexpressing HA–ubiquitin (HA–Ub) as in l. q, Expression of Pax5WT, Pax5K198Q and Pax5K198R in HEK293F cells treated as in g (n = 4). r, Scatter plot of the correlation between Pax5 and PCAF protein in individuals with B-ALL29, as determined by proteomics. Linear regression and 95% confidence intervals (dashed lines) are shown (n = 27, P = 0.02). s, Pax5 expression (left; pooled from three independent experiments) and Pax5 acetylation (right; Ack/Pax5, pooled from four (Pax5WT) and three (Pax5K198R) independent experiments) in HEK293F cells expressing Pax5WT or Pax5K198R together with empty vector, p300, PCAF, NCOA3 or GTF3C4. Data are shown as mean ± s.d. (a, c, e, g, q and s) and were analyzed by two-tailed t-tests with a Holm–Sidak comparison (a), one-tailed t-test (e and r), two-way ANOVA with Sidak comparisons (g) or one-way ANOVA with Fisher’s LSD test (q and s).
Extended Data Fig. 5. SIRT7 regulates Pax5 stability by deacetylating K198.
a, Representative histograms of Pax5 expression in gated B220+CD19+IgM−CD43+ pro-B and B220+CD19+IgM−CD43− pre-B cells from the BM of wild-type and Sirt7−/− mice (wild-type, n = 6; Sirt7−/−, n = 8). Pooled from three separate experiments b, RNA-Seq analysis of Pax5 expression in sorted pro-B and pre-B cells defined as in a. Data are shown as mean of two replicates (n = 2). c, Pax5/GAPDH ratio in B220+CD19+IgM−CD43+ pro-B cells retrovirally expressing empty vector (EV) and SIRT7 and expanded ex vivo for four days with OP9 cells and 10 ng/ml IL-7, SCF and FLT3-L (n = 3). Pooled from 3 separate experiments. d, Immunoblots of Pax5 and SIRT7 protein from wild-type and CRISPR-Cas9-generated HAFTLSIRT7KO pre-B cells treated with vehicle (Ct) or 2 µM lactacystin for 8 h. e, FLAG-specific immunoprecipitation from HEK293F cells transiently co-expressing SIRT7-HA with EV or Pax5-FLAG using polyethyleneimine transfection, followed by FLAG and HA immunoblotting. f, RT-qPCR analysis of Pax5 expression relative to Hprt in HAFTL cells treated with vehicle (Ct) or 5 mM nicotinamide (NAM) for 48 h (n = 3). g, Pax5/H3 ratio in HAFTL (n = 4 biological replicates; p < 0.0001) and KOPN-8 (n = 2 biological replicates) cells treated as in f. h,i, FLAG-tagged Pax5WT, Pax5K198Q and Pax5K198R levels (h, n = 3) and ubiquitination (i, n = 3). Expressed in HEK293F cells as in e. Pooled from three separate experiments. j,k, Immunoblots (j) and quantification (k) of Pax5WT, Pax5K198A and Pax5K198R forms expressed as in e. Data in k are pooled from three separate experiments. l, Relative levels of Pax5WT, Pax5K198A and Pax5K198R forms in HEK293F cells treated with vehicle (Ct) or 100 μg/mL cycloheximide (CHX) for 8 h (n = 3, pooled from three separate experiments). m, Subcellular fractionation of HEK293F cells expressing Pax5WT, Pax5K198Q and Pax5K198R forms as in e. GAPDH, Fibrillarin and H3; cytoplasmic, nuclear and chromatin controls. n, Venn diagram depicting the overlap between Pax5 protein−protein interactions (PPIs) and mammalian histone acetyltransferases31 (HATs) (see methods). o, Scatter plots showing the correlation between the protein levels of Pax5, p300, NCOA3 and GTF3C4, as determined by proteomics in human B-ALL patients samples29 (n = 27). Each point corresponds to one sample. Linear regression and 95% confidence intervals (dashed lines) are shown. p-r, Immunoblots depicting the expression (p,q) and acetylation levels (r) of Pax5WT and Pax5K198R proteins from HEK293F transiently co-expressing Pax5WT or Pax5K198R together with p300, PCAF, NCOA3 or GTF3C4. Data in c, f, g, h, i and k are shown as mean ± sd. Data in c, h, I, k were analyzed using one-way ANOVA with Fisher’s LSD test, and data in g (HAFTL) were analyzed using two-tailed t-test.
We next investigated whether the catalytic activity of SIRT7 was required for Pax5 protein stabilization. Treatment with the pan-sirtuin inhibitor nicotinamide induced Pax5 degradation in HAFTL cells and B-ALL KOPN-8 cells, whereas the expression of Pax5 mRNA was unaffected (Fig. 3j and Extended Data Fig. 5f,g). To test whether SIRT7 deacetylated Pax5 directly, we expressed and purified them in HEK293F cells and performed in vitro deacetylation assays. Incubation of Pax5 with SIRT7 in the presence of the cofactor NAD+ partially reduced Pax5 global acetylation (Fig. 3k). Ectopic expression of Pax5 alone or together with SIRT7 in Sirt7−/− HEK293F cells followed by Pax5 purification and MS detected several acetylated residues, including K79, K87, K98, K198 and K257 (Fig. 3l,m). Of them, only K198 was lost in Pax5 isolated from SIRT7−/− HEK293F cells overexpressing SIRT7 (as evidenced by the loss of the cluster of acetylated peptide peaks spanning from 554.79 to 555.54 m/z; Fig. 3l), indicating that SIRT7 specifically deacetylated Pax5 at K198. K198 is located within a putative Pax5 intrinsically disordered region of unknown function, located between the conserved octapeptide and the partial homeodomain30 and conserved across Pax5 orthologs in chordates (Fig. 3n). A lysine-to-glutamine Pax5 mutant at K198 (Pax5K198Q) that mimicked acetylated Pax5 was expressed at lower levels than wild-type Pax5 protein (Pax5WT) in HEK293F cells, whereas a lysine-to-arginine mutant (Pax5K198R) that mimicked the deacetylated form of Pax5 was expressed at higher levels than Pax5WT (Fig. 3o and Extended Data Fig. 5h). When Pax5WT, Pax5K198Q and Pax5K198R were expressed together with hemagglutinin–ubiquitin (HA–ubiquitin) in HEK293F cells, ubiquitin was preferentially loaded into Pax5K198Q (Fig. 3p and Extended Data Fig. 5i), suggesting increased proteasomal degradation. The half-life of Pax5K198Q was diminished 2.5-fold compared to Pax5WT, whereas the half-life of Pax5K198R increased 1.7-fold (Fig. 3q). A K198 lysine-to-alanine mutant (Pax5K198A) showed similar protein levels (Extended Data Fig. 5j) and half-life as Pax5K198R in untreated and cycloheximide-treated HEK293F cells, respectively (Extended Data Fig. 5k,l), ruling out the possibility that the increased stability of Pax5K198R was due to potential modifications on arginine. Finally, cellular fractionation experiments showed that Pax5WT, Pax5K198Q and Pax5K198R were similarly distributed in the nucleoplasm and chromatin and were excluded from the cytoplasm in HEK293F cells (Extended Data Fig. 5m). Thus, SIRT7-mediated deacetylation of Pax5 at K198 prevented the proteasomal degradation of Pax5 and enhanced its stability.
To identify the acetyltransferase(s) that catalyze Pax5 K198 acetylation, we searched for proteins with acetyltransferase activity31 among reported interactors of Pax5 (Methods) and identified four potential candidates (p300, PCAF, GTF3C4 and NCOA3; Extended Data Fig. 5n and Supplementary Table 3). Analysis of the expression of Pax5 and the four candidates in publicly available proteomics data from 27 individuals with B-ALL29 showed that only PCAF levels correlated negatively with Pax5 protein expression (Fig. 3r and Extended Data Fig. 5o). Following coexpression of Pax5 with all four enzymes in HEK293F cells, Pax5-WT expression was strongly reduced when coexpressed with PCAF, but not with the other acetyltransferases, compared to the levels of Pax5 expressed alone (Fig. 3s and Extended Data Fig. 5p). Pax5-K198R expression was similar whether coexpressed with PCAF or not (Fig. 3s and Extended Data Fig. 5q), suggesting that PCAF modulates Pax5 expression through K198 acetylation. To measure the ability of all four enzymes to acetylate Pax5, we coexpressed them with Pax5WT or Pax5K198R, purified Pax5 by immunoprecipitation and measured global Pax5 acetylation by pan-acetyl-lysine immunoblotting. Only PCAF significantly increased Pax5WT, but not Pax5K198R acetylation (Fig. 3s and Extended Data Fig. 5r), indicating that PCAF specifically acetylates Pax5 at K198. Thus, SIRT7 and PCAF regulate Pax5 protein stability by controlling Pax5 K198 acetylation.
Pax5 K198 acetylation regulates gene expression
To investigate the impact of Pax5 K198 acetylation on Pax5 genome distribution and transcriptional activity, we performed ChIP–seq and RNA-seq in sorted Lin–B220+IgM− pro-B cells from Pax5−/− mice retrovirally infected with Pax5WT, Pax5K198Q and Pax5K198R. ChIP–seq indicated that Pax5K198Q had markedly reduced binding to chromatin, whereas Pax5K198R occupancy at Pax5 target loci was strongly increased compared to Pax5WT (Fig. 4a). Motif enrichment analysis indicated that Pax5K198Q and Pax5K198R bound to the same Pax5 binding motif (Extended Data Fig. 6a). Differences in the binding to other transcription factor motifs were observed, including a reduced binding to the PU.1 motif and increased binding to the FLI1 motif by Pax5K198R compared to Pax5WT (Extended Data Fig. 6b).
Fig. 4. Pax5 K198 acetylation regulates gene expression.
a, ChIP–seq analysis showing the genomic occupancy of Pax5-WT, Pax5-K198Q and Pax5-K198R in Pax5−/− Lin–B220+IgM− pro-B cells retrovirally transduced with Pax5WT, Pax5K198Q and Pax5K198R and expanded ex vivo for 7 days in OP9 cells and in the presence of 10 ng ml–1 IL-7, SCF and FLT3-L. One thousand random significant peaks are displayed. Top, read coverage profiles. b, Venn diagram of the overlap between the significant peaks (q < 0.05) detected by ChIP–seq of Pax5WT, Pax5K198Q and Pax5K198R forms. c, Binding of Pax5WT, Pax5K198Q and Pax5K198R in the Actr6 (top) and Nus1 regions (bottom). The y axis represents read coverage. d, PCA clustering of RNA-seq data from Pax5−/− Lin−B220+IgM− pro-B cells retrovirally transduced with empty vector, Pax5WT, Pax5K198Q and Pax5K198R. e, Unsupervised clustering of differentially expressed genes (FDR < 0.05) in Pax5−/− Lin−B220+IgM− pro-B cells retrovirally transduced with empty vector, Pax5WT, Pax5K198Q and Pax5K198R as in d. Significant Gene Ontology terms for clusters C1 and C2 are shown. f, Volcano plots of differentially expressed genes in Pax5−/− Lin−B220+IgM− pro-B cells retrovirally transduced with Pax5WT, Pax5K198Q and Pax5K198R versus empty vector. The black arrow indicates the absence of downregulated genes; significantly regulated genes (| log2 (fold change) | ≥ 1.5, FDR < 0.05) are shown in red. Numbers on the right and left show the number of genes significantly induced and repressed, respectively; FC, fold change; Fr. A, fraction A. g, ChIP–seq of Pax5WT, Pax5K198Q and Pax5K198R forms and RNA-seq of Pax5−/− Lin−B220+IgM− pro-B cells retrovirally expressing empty vector, Pax5WT, Pax5K198Q or Pax5K198R in Mycn (top) and Rag1 (bottom) genes. The y axis represents read coverage.
Extended Data Fig. 6. Pax5K198 acetylation regulates gene expression.
a, DNA binding motifs of Pax5WT, Pax5K198Q and Pax5K198R in Pax5−/− Lin-B220+IgM−pro-B cells retrovirally transduced with Pax5WT, Pax5K198Q and Pax5K198R forms and expanded ex vivo for seven days with OP9 cells and 10 ng/ml IL-7, SCF and FLT3-L. b, Motif enrichment analysis of the significant peaks (q < 0.05) detected by ChIP-Seq of Pax5WT, Pax5K198Q and Pax5K198R as in a. c, Comparison of the significant peaks bound by Pax5K198R as in a and those reported in Ref. 17. d, Genomic occupancies of Pax5WT, Pax5K198Q and Pax5K198R forms in the unique Pax5K198R peaks. Top panels, read coverage profiles. e, Gene ontology terms of the unique Pax5K198R peaks. Only terms with -log10(p-value)>3 are reported. f, RNA-Seq analysis of the expression of Blnk, Irf4, Rag1, Il7r, Dusp4 and Mycn the indicated genes in Lin−B220+IgM− Pax5−/− pro-B cells expressing an empty vector (EV), Pax5WT, Pax5K198Q and Pax5K198R and expanded as in a. Data are presented as mean (n = 2). g, ‘HALLMARK_E2F_TARGETS’ GSEA of RNA-seq data from Pax5WT versus EV (upper) pro-B cells and PAX5K198R versus EV (lower) pro-B cells. FDR, false discovery rate.
Comparison of the Pax5K198Q, Pax5K198R and Pax5WT peaks revealed that Pax5K198Q lost occupancy in nearly 80% of the peaks occupied by Pax5WT and Pax5K198R (for example, peaks in Klf6, Myc and Bcl6 genes; Fig. 4b and Supplementary Table 4). Notably, Pax5K198R bound 3,093 new regions (28.2% of all detected peaks), such as Csf1 and Actr6, and lost 1,659 binding sites (15.1%; for example, Nus1 and Fundc1), compared to Pax5WT, whereas 6,616 peaks (55.8%; for example Cd19 and Vpreb1) were shared with Pax5WT (Fig. 4b). More than 90% of the regions bound by Pax5K198R were previously reported17, such as those in the Actr6 and Foxo1 genes (Extended Data Fig. 6c), indicating that K198 deacetylation stabilized Pax5 binding at regions where it binds weakly rather than causing Pax5 redistribution. Consistently, Pax5WT and Pax5K198Q exhibited mild but clear binding to regions that were only bound significantly by Pax5K198R (Fig. 4c and Extended Data Fig. 6d). Similarly, we observed mild binding of Pax5K198R to unique Pax5WT sites, such as the Nus1 gene (Fig. 4c). Gene ontology analysis of the peaks uniquely bound by Pax5K198R indicated enrichment in lineage-inappropriate genes (Csf1, Lif and Itga6) as well as in genes involved in B cell differentiation (Ikzf1 and Cxcr4), the cell cycle (Rb1 and Cdk4) and metabolism (Pten and Cox20; Extended Data Fig. 6e). These data suggest that K198 deacetylation promotes Pax5 binding to regions linked to B cell development and commitment.
In a PCA of RNA-seq data, Pax5−/− pro-B cells transduced with empty vector clustered closer to Pax5−/− pro-B cells expressing Pax5K198Q than to those expressing Pax5WT (Fig. 4d), whereas Pax5−/− pro-B cells expressing Pax5K198R clustered away from the rest (Fig. 4d). Unsupervised clustering unveiled two clusters: cluster C1, which comprised genes repressed by Pax5, such as lineage-inappropriate genes (Zap70 and Nkg7) and cytokine signaling genes (Stat3 and Il2rg; Fig. 4e), and cluster C2, which comprised genes induced by Pax5, such as genes related to pre-B cell receptor (Vpreb1a/b and Cd79a/b), IL-7 receptor-mediated signaling (Il7r and Stat5b) and V(D)J recombination (Foxo1 and Rag1/Rag2; Fig. 4e). Pax5K198R induced genes related to V(D)J recombination (Blnk, Irf4 and Rag1) more strongly than Pax5WT and Pax5K198Q, whereas genes linked to proliferation (Il7r, Dusp4 and Mycn) were similarly regulated by Pax5WT and Pax5K198Q, but not by Pax5K198R (Extended Data Fig. 6f,g). All three Pax5 proteins significantly increased the expression of a similar number of genes compared to Pax5−/− pro-B cells (for example, Cd19 and Vpreb3), with Pax5K198R inducing the expression of some targets (Cd55 and Bcar3) stronger than Pax5WT (Fig. 4f). By contrast, introduction of Pax5K198Q significantly reduced the expression of only 13 genes (for example, Cyp2d13 and Bcl2a1a), whereas Pax5K198R stringently repressed 375 genes (including Gata3 and Il2rb; Fig. 4f and Supplementary Table 5). These observations indicate that Pax5 K198 acetylation affects the extent to which Pax5 regulates its canonical targets rather than controlling different subsets of genes. Binding of Pax5K198Q and Pax5K198R to the promoter or to putative proximal enhancer regions correlated with the induction or silencing of the bound genes (Fig. 4g). Therefore, modulation of acetylation at Pax5 K198 creates a regulatory switch whereby K198-deacetylated Pax5 potently represses gene expression of lineage-inappropriate and cytokine signaling genes in pro-B cells, whereas K198-acetylated Pax5 fails to repress lineage-inappropriate genes but efficiently regulates genes linked to V(D)J and proliferation.
Pax5 K198 deacetylation regulates B cell identity
To confirm that Pax5 deacetylation is required for B cell development and lineage restriction, we injected CD45.2 Pax5−/− Lin−B220+IgM− pro-B cells retrovirally expressing either empty vector or vectors encoding Pax5WT, Pax5K198Q or Pax5K198R into sublethally irradiated CD45.1 mice. Four weeks after transplantation, CD45.2+B220+CD19+ B cells were detected in the BM of mice injected with Pax5−/− pro-B cells expressing Pax5WT, but not in mice transferred with Pax5−/− pro-B cells expressing Pax5K198Q or Pax5K198R (Fig. 5a and Extended Data Fig. 7a), indicating that Pax5 K198 dynamic acetylation/deacetylation is required for B cell development. Pax5-deficient pro-B cells differentiate into alternative lineages, including T cells7,32. Donor-derived CD45.2+ CD4+ and CD8+ T cells were detected in the thymus of mice injected with Pax5−/− pro-B cells expressing empty vector or Pax5K198Q, but not in those injected with Pax5−/− pro-B cells expressing Pax5WT or Pax5K198R (Fig. 5b and Extended Data Fig. 7b), suggesting that SIRT7-mediated K198 deacetylation is required for pro-B cell commitment. Transfer of sorted CD45.2 wild-type or Sirt7−/− Lin–B220+CD19+IgM− pro-B cells into sublethally irradiated CD45.1/CD45.2 mice revealed that 4 weeks after transplantation, donor-derived CD45.1–CD45.2+TCRβ+ T cells and CD45.1–CD45.2+NKP46+ NK cells were present in the spleens of mice injected with Sirt7−/− pro-B cells (Fig. 5c–e), indicating that Sirt7−/− pro-B cells differentiated into T cells and NK cells. Lin−IgM+IgD+ mature B cells sorted from the spleens of CD45.2 wild-type or Sirt7−/− mice and transferred into sublethally irradiated CD45.1/CD45.2 mice did not lose CD19 expression, and no CD45.1–CD45.2+TCRβ+ T cells were detectable in the spleens of mice injected with wild-type or Sirt7−/− mature B cells 6 weeks after transplantation (Extended Data Fig. 7c), suggesting that SIRT7 is required for the establishment, but not maintenance, of B cell commitment.
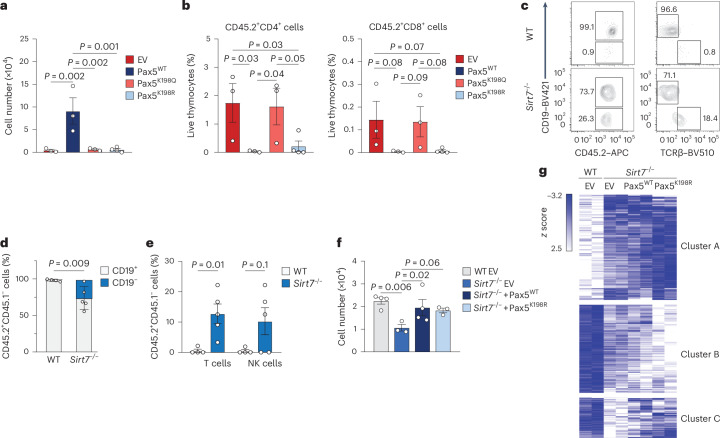
Fig. 5. Pax5 K198 deacetylation regulates B cell identity.
a,b, Number of donor-derived CD45.2+CD19+B220+ cells in the BM (a) and percentages of donor-derived CD45.2+CD4+ cells (left) and CD45.2+CD8+ (right) cells in the thymus (b) of CD45.1 wild-type recipient mice 4 weeks after transplantation of Lin−B220+IgM− Pax5−/− pro-B cells retrovirally transduced with empty vector, Pax5WT, Pax5K198Q or Pax5K198R (empty vector, n = 3; Pax5WT, n = 3; Pax5K198Q, n = 3; Pax5K198R, n = 4). Data are presented as mean ± s.e.m. and were analyzed by one-way ANOVA with Fisher’s LSD test. c–e, Representative flow cytometry plots (c) and percentages of donor-derived CD45.1–CD45.2+CD19+ and CD45.1–CD45.2+CD19− cells (d) and donor-derived CD45.1–CD45.2+TCRβ+ and CD45.1–CD45.2+NKP46+ cells (e) in the spleens of CD45.1/CD45.2 recipient mice injected with CD45.2 wild-type (n = 4) or Sirt7−/− (n = 5) Lin−B220+CD19+IgM− pro-B cells. Data are presented as mean ± s.d. (d) or mean ± s.e.m. (e) and were analyzed by two-way ANOVA with Fisher’s LSD test (d) or multiple t-tests with Holm–Sidak comparisons (e). f, Number of donor-derived B220+CD19+GFP+ B cells in the spleens of CD45.1/CD45.2 recipient mice injected with wild-type and Sirt7−/− Lin−B220+CD19+IgM− pro-B cells expressing empty vector, Pax5WT and Pax5K198R retroviruses (wild-type + empty vector, n = 4; Sirt7−/− + empty vector, n = 3; Sirt7−/− + Pax5WT, n = 3; Sirt7−/− + Pax5K198R, n = 3). Data are presented as mean ± s.d. and were analyzed by one-way ANOVA with Fisher’s LSD test. g, Unsupervised clustering of the differentially expressed genes (P < 0.05) between wild-type and Sirt7−/− Lin−B220+CD19+IgM− pro-B cells retrovirally transduced with empty vector, Pax5WT or Pax5K198R.
Extended Data Fig. 7. Pax5K198 deacetylation regulates B cell identity.
a,b, Gating strategies used to identify CD45.2+B220+CD19+ B cells (a) or CD45.2+CD4+ and CD45.2+CD8+ T cells (b) derived from donor CD45.2 Pax5−/− Lin−B220+IgM−pro-B cell expressing empty vector (EV), Pax5WT, Pax5K198Q and Pax5K198R by retroviral transduction in the bone marrow (a) or thymus (b) of CD45.1 recipient mice 4 weeks after transplantation. c, Representative histograms of the donor derived CD45.1-CD45.2+CD19+ B cells in the spleens of recipient CD45.1/CD45.2 mice 6 weeks after injection of wild-type or Sirt7−/− Lin−IgM+IgD+ splenic B cells. d) Gating strategy used to identify donor-derived B220+CD19+GFP+ B cells in the spleens of CD45.1/CD45.2 recipient mice four weeks after transplantation of ex vivo expanded (with OP9 cells and 10 ng/ml IL-7, SCF and FLT3-L, for 7 days) wild-type and Sirt7−/− Lin−B220+CD19+GFP+ pro-B cells retrovirally transduced with empty vector (EV, pMIG), Pax5WT or Pax5K198R. e, Gene ontology analysis of the genes in Clusters A, B and C generated by unsupervised clustering analysis of bulk RNA-seq data from the same cells as in d. Only terms with -log10(p-value) > 1.5 are reported.
Conditional deletion of PCAF and its homolog Gcn5 in B cell progenitors impairs B cell development at the pro-B cell stage33. To test whether acetylation of Pax5 K198 is required for B lymphopoiesis, we retrovirally expressed Pax5WT or Pax5K198R (together with green fluorescent protein (GFP)) in CD45.2 Sirt7−/− pro-B cells. After 4 weeks of transplantation into sublethally irradiated CD45.1/CD45.2 mice, we detected reduced numbers of B220+CD19+GFP+ B cells in the spleens of mice injected with Sirt7−/− pro-B cells compared to mice injected with wild-type cells. This effect was reversed in Sirt7−/− pro-B cells expressing Pax5WT or Pax5K189R (Fig. 5f and Extended Data Fig. 7d), indicating that PAX5 K198 deacetylation rescues defective B cell development in Sirt7−/− pro-B cells. To understand how Pax5 rescues B cell development in Sirt7−/− progenitors at the gene expression level, we performed RNA-seq in these pro-B cells. Unsupervised clustering analysis revealed that expression of Pax5K198R and, to a lesser extent, Pax5WT restored the repression of 93 of the 297 genes (31.3%) upregulated in Sirt7−/− pro-B cells, including Cxcr5 and Tet1 (cluster C), and did not affect the expression of genes downregulated in Sirt7−/− pro-B cells (cluster A; Fig. 5g and Supplementary Table 6). Pax5WT expression in Sirt7−/− pro-B cells mostly repressed lineage-inappropriate genes (for example, Nfatc2 and Cyyr1; Extended Data Fig. 7e), consistent with its role in lineage commitment. These results indicated that B cell development required the dynamic acetylation and deacetylation of Pax5, with Pax5 deacetylation being necessary to restrict lineage plasticity.
The Pax5–SIRT7 interplay is conserved in human B-ALL
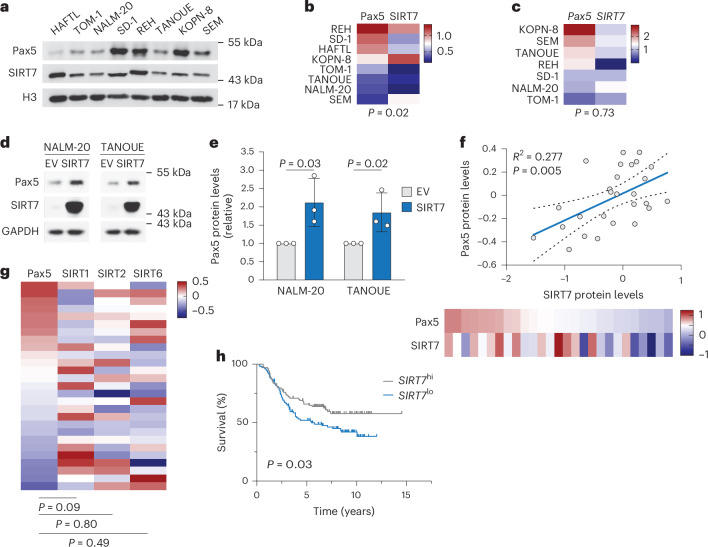
Because Pax5 is a haploinsufficient tumor suppressor in human B-ALL8,10, we investigated whether the SIRT7–PAX5 interplay was functionally relevant in this disease. In a panel of B-ALL cell lines, Pax5 and SIRT7 protein, but not mRNA, levels strongly correlated, despite their diverse genetic backgrounds (Fig. 6a–c). Retroviral overexpression of SIRT7 in NALM-20 and TANOUE cells (two B-ALL cell lines with reduced Pax5 levels) significantly increased Pax5 protein expression (Fig. 6d,e). Analysis of publicly available proteomics data from a cohort of ETV6-RUNX1+ and high hyperploidy (HeH+) B-ALL cases29 found a highly significant correlation between Pax5 and SIRT7 protein levels in these individuals (P = 0.005; Fig. 6f). PAX5 and SIRT7 mRNA correlated weakly in HeH+ B-ALL (Extended Data Fig. 8a), although the correlation was stronger for the protein (protein, P = 0.019; RNA, P = 0.026). A comparable link at the protein level was found in a panel of B cell chronic lymphoblastic leukemia (B-CLL) cell lines34 (Extended Data Fig. 8b), indicating that the SIRT7–Pax5 expression correlation was independent of genetic background and differentiation stage. Among the tested sirtuins, only SIRT7 associated with Pax5 protein levels in individuals with B-ALL29 (Fig. 6g and Extended Data Fig. 8c). Stratification of pediatric individuals with B-ALL from the COG9906 study35,36 according to their SIRT7 mRNA expression indicated that those with higher than median SIRT7 mRNA levels had a better prognosis than those expressing lower than median SIRT7 mRNA (Fig. 6h), indicating that SIRT7 could be an independent prognostic factor in human B-ALL. PAX5 inactivating mutations in B-ALL are usually monoallelic8. Classification of individuals with B-ALL35–37 into those with or without PAX5 deletions and further into SIRT7hi and SIRT7lo subgroups indicated that higher than median SIRT7 expression tended to be protective only in individuals with B-ALL with PAX5 deletions (Extended Data Fig. 8d), suggesting a tumor-suppressive function of SIRT7 by increasing PAX5 dosage in individuals with B-ALL with haploinsufficient PAX5.
Fig. 6. The Pax5–SIRT7 interplay is conserved in human B-ALL.
a, Immunoblot of Pax5, SIRT7 and H3 in HAFTL, TOM-1, NALM-20, SD-1, REH, TANOUE, KOPN-8 and SEM cell lines. b,c, Heat map of the correlation between Pax5 and SIRT7 protein expression normalized to H3 expression (b) and Pax5 and Sirt7 RNA expression normalized to Hprt RNA expression (c) in B-ALL cell lines as in a. Mean z scores for the normalized protein (b) and mRNA (c) are shown. The P values were determined by Spearman’s rank correlation. Data were pooled from three (b) or two (c) experiments. d,e, Immunoblots of the levels of Pax5 and SIRT7 in NALM-20 and TANOUE B-ALL cells expressing empty vector and SIRT7 retroviruses (d) and quantification of the Pax5/GAPDH ratio (e). Data are shown as mean ± s.d. and were analyzed by one-tailed t-tests (n = 3 biological replicates). f, Scatter plot (top) and heat map (bottom, z score) of Pax5 and SIRT7 protein determined by proteomics in individuals with B-ALL29 (n = 27). Each point corresponds to one sample. Linear regression, 95% confidence intervals (dashed lines) and Spearman’s rank correlation coefficient (R2) are shown (P = 0.005, one-tailed t-test). g, Heat map of the correlation between the expression of Pax5, SIRT1, SIRT2 and SIRT6 in the same human B-ALL samples as in f. h, Kaplan–Meier survival curves for children with high-risk B-ALL from the COG-P9906 study35,36 (n = 209) stratified as higher (SIRT7hi) or lower (SIRT7lo) than median SIRT7 RNA expression. Statistical significance was determined by a long-rank test (P value < 0.03).
Extended Data Fig. 8. The PAX5/SIRT7 interplay is conserved in human B-ALL.
a, Scatter plots of Pax5 and SIRT7 levels derived from proteomics (left panels) and RNA-Seq (right panels) of human B-ALL HeH (n = 18) and ETV6-RUNX1 (n = 9) patient samples29. Each point corresponds to one sample. Linear regression and 95% confidence intervals (dashed lines) are shown in red. The p-values were calculated by one-tailed t-test. b, Heatmap showing the correlation between the protein levels of Pax5 and SIRT7 in B-CLL cell lines versus healthy donor B cells, as determined by proteomics34. The p-values were determined by one-tailed t-test (p = 0.0016). c, Correlation between the protein levels of Pax5 and SIRT1, SIRT2 and SIRT6 in human B-ALL patient samples29, as determined by proteomics (n = 27). Data are presented and analyzed as in a. d, Kaplan-Meier survival curves of children with high-risk B-ALL (COG-P9906 study)35,36 stratified into four groups based on higher- or lower-than-median SIRT7 RNA expression levels and the occurrence of PAX5 deletions37. Statistical significance was determined by log-rank test (PAX5WTSIRT7hi, n = 14; PAX5WTSIRT7lo, n = 15; PAX5delSIRT7hi, n = 11; PAX5delSIRT7lo, n = 12).
Discussion
Here, we present evidence implicating SIRT7 in B cell lymphopoiesis by establishing an acetylation switch on Pax5 that regulates its essential role in B cell development and identity. SIRT7-dependent deacetylation of Pax5 K198 promoted its protein stability, resulting in increased occupancy in a wide range of target genes, and enhanced its repressive activity. Conversely, PCAF-mediated Pax5 K198 acetylation reduced Pax5 protein levels and was associated with a global reduction of Pax5 occupancy and impaired repression of lineage-inappropriate genes. Transplantation experiments of Pax5 K198-acetylated and K198-deacetylated mimics demonstrated that, although a deacetylated mimic Pax5-K198R was enough to drive lineage commitment, both forms were required for B cell differentiation.
SIRT7 mRNA and protein levels were upregulated during early B cell development, coinciding with the activation of Pax5 expression in pro-B cells38. SIRT7 participated in repression of lineage-potential genes and balanced the expression of DNA repair and cell cycle factors to preserve pre-B cell proliferation and survival. SIRT7 also enhanced distal Igh recombination in pro-B cells and behaved as a tumor suppressor in B-ALL, suggesting that SIRT7 may contribute to the prevention of malignant transformation in hematopoietic progenitors. The identification of PCAF as the major Pax5 K198 acetyltransferase provided a mechanistic explanation for the impairment of pro-B cell development described following conditional deletion of Pcaf and its homolog Gcn5 in mice33. Furthermore, the observed deacetylation of PCAF by SIRT7 in colon cancer cells39 suggests that both enzymes may have an interdependent relationship during B cell lymphopoiesis.
Expression of the constitutively deacetylated Pax5K198R mutant in Pax5−/− pro-B cells abrogated their T cell potential but failed to restore B cell development, suggesting that PAX5 promotes these processes through independent mechanisms. Because Pax5-acetylated or Pax5-deacetylated mimics could not rescue B cell differentiation, both forms seem to be required for lymphopoiesis, raising the question of how Pax5 acetylation is regulated during B development. One possibility is that pools of acetylated and deacetylated Pax5 coexist in B cell progenitors and that their balanced actions lead to optimal B cell differentiation. Despite its ability to efficiently repress lineage plasticity in pro-B cells, the deacetylated mimic Pax5K198R overactivated Igh recombination genes and failed to induce normal expression of clonal expansion programs, consistent with its roles in V(D)J recombination15–17 and as a negative regulator of pro-B cell proliferation13. Thus, it is also possible that a pool of constitutively deacetylated Pax5 ensures lineage commitment throughout B cell development, whereas a pool of Pax5 that undergoes dynamic acetylation and deacetylation controls the B cell developmental programs.
K198 is located within a putative intrinsically disordered region of Pax5, between the octapeptide and the partial homeodomain30. Although K198 acetylation strongly decreased Pax5 protein stability, it may also regulate Pax5 in other ways. Pax5 shapes chromatin organization and transcription by recruiting chromatin remodelers and histone modifiers to its target loci18, so the observed cofractionation of Pax5 and SIRT7 in high-molecular-weight complexes suggests that K198 deacetylation may also influence the ability of Pax5 to mediate chromatin modification. Additionally, acetylation of Pax5 K198 might also affect global nuclear organization12, as a substantial number of Pax5 binding sites lie in distal intergenic regions18,20, and Pax5 binding to DNA plays a key organizational role in genome architecture, independent of transcription12.
Finally, our findings provide a potential avenue to target leukemia pharmacologically. Inactivating mutations of the PAX5 gene are present in 30% of B-ALL cases8, and restoration of Pax5 levels in human and mouse B-ALL cells leads to leukemia regression40,41. Because PAX5 mutations in B-ALL are usually monoallelic, enhancing wild-type PAX5 allele function to overcome its haploinsufficiency represents a promising strategy for B-ALL treatment. Because SIRT7 activity strongly increased Pax5 levels in B cell progenitors, human B-ALL samples and cell lines, we propose that developing SIRT7-activating compounds to stimulate Pax5 functions may provide a basis for leukemia therapies. Further research should establish the relevance of the SIRT7–Pax5 regulatory axis in B cell malignancies.
Methods
Mice
Two previously reported germline CD45.2 Sirt7−/− mouse models that contained exon 4 to 10 (ref. 24; Sirt7Δ4–10) or exon 4 to 9 deletions25 (Sirt7Δ4–9) and that were maintained on the 129Sv or C57BL/6 genetic background, respectively, were used. 129Sv Sirt7−/− mice were used for most experiments. IgHEL28 mice (CD45.2) were on the 129Sv background, whereas Pax5−/−5 and CD45.1 mice were on the C57BL/6 background. Heterozygous CD45.1/CD45.2 mice were generated by crossing CD45.1 C57BL/6 and wild-type CD45.2 129Sv mice for one generation to avoid rejection following transplantation of 129Sv cells. All mice were bred at the Comparative Medicine and Bioimage Centre of Catalonia animal facility of the Germans Trias i Pujol Research Institute. Animal studies were conducted at Josep Carreras Leukemia Research Institute (IJC) (Spain) according to national authorities and institutional ethics committees (Germans Trias i Pujol Reserach Institute Ethics Committee). The collection of BM samples from C57BL/6 wild-type and Sirt7−/− mice and the generation of Pax5−/− mouse B cell progenitors were conducted according to national authorities and institutional ethics committees at Max Plank Institute for Heart and Lung Research (MPI-HLR) (Germany) and Lund University (Sweden), respectively. Mouse housing conditions included a temperature of 21–25 °C, 40–70% humidity and a light cycle from 0800 to 2000 h with a 15-min intensity ramp simulating sunrise and sunset.
Determination of anti-HEL isotypes
To analyze antigen-specific responses, mice were intraperitoneally injected with 100 μl of an emulsion containing 50 μg of NP–HEL along with complete Freund adjuvant. After 14 days, mice were bled, and serum samples were collected. For antibody isotype detection, enzyme-linked immunosorbent assay plates were coated overnight at 4 °C with HEL peptide (6 μg ml–1). Following removal of excess peptide, serum dilutions from control or immunized mice were added and incubated for 1 h at room temperature. After three washes with a 0.05% Tween–PBS solution, rat anti-mouse immunoglobulin subclasses (IgM, IgG1 and IgG3; 1:250) were added for 1 h at room temperature. After 3 washes, an anti-rat immunoglobulin (1:5,000) was added for 1 h at room temperature, followed by three additional washes. Finally, TMB developing solution was added for 15 min and stopped with 1 N H2SO4. Color production was measured at 450 nm and 570 nm (for background subtraction) using a Multiskan Sky (Thermo Fisher) plate reader.
Transplantation experiments
Two million short-term expanded pro-B cells or 1.5 million splenic Lin–IgM+IgD+ B cells were washed and resuspended in 200 µl of PBS supplemented with 1% heat-inactivated fetal calf serum. Cells were injected via the tail vein into sublethally irradiated (5 Gy) CD45.1/CD45.2 or CD45.2 6- to 10-week-old randomized recipient mice. Four weeks after transplantation, the spleen (CD45.1/CD45.2 mice) or BM and thymus (CD45.2 mice) were collected and analyzed by fluorescence-activated cell sorting (FACS).
Flow cytometry and cell sorting
BM, spleen and thymus samples were collected from wild-type, Sirt7−/−, IgHEL+/–, Sirt7−/−IgHEL+/–, CD45.1/CD45.2 and CD45.2 mice. BM samples were crushed in staining buffer (3% fetal bovine serum (FBS) and 2 mM EDTA in PBS) to obtain single-cell suspensions. Spleen and thymus samples were similarly processed. Red blood cells were lysed in ACK buffer (Gibco), and the reaction was stopped by adding 5 volumes of staining buffer. Cells were filtered through 40-μm sterile strainers and incubated with Fc-block (eBioscience) before staining for 30 min (4 °C) with anti-B220 (RA3-6B2, eBioscience, 1:400), anti-CD19 (HIB19, eBioscience, 1:400), anti-CD43 (eBioR2/60 or 1G10, BD Biosciences, 1:800), anti-IgM (II/41, eBioscience, 1:800), anti-IgD (11-26c, eBioscience, 1:400), anti-CD21 (7G6, BD Biosciences, 1:800), anti-CD23 (B3B4, BD Biosciences, 1:200), anti-CD93 (AA4.1, eBioscience, 1:400), anti-Gl7 (Gl7, eBioscience, 1:200), anti-CD38 (90, eBioscience, 1:400), anti-CD138 (300506, Invitrogen, 1:400), anti-Fas (SA367H8, Biolegend, 1:600), anti-IgG1 (A85-1, BD Pharmigen, 1:600), anti-CD127 (eBioSD/199, eBioscience, 1:200), anti-CD45.1 (A20, eBioscience, 1:400), anti-CD45.2 (104, eBioscience, 1:200), anti-TCRβ (H57-597, BD Biosciences, 1:200), anti-NKP46 (29A1.4, Biolegend, 1:100), anti-CD4 (GSK1.5, eBioscience, 1:800), anti-CD8 (53-6.7, eBioscience, 1:800), anti-hCD4 (RPA-T4, Biolegend, 1:400), anti-CD3e (145-2C11, BD Biosciences, 1:400), anti-Ly76 (TER-119, BD Biosciences, 1:400), anti-CD11b (M1/70, BD Biosciences, 1:400) or anti-GR1 (RB6-8C5, BD Biosciences, 1:400). After staining, cells were washed twice and resuspended in staining buffer before analysis with a FACS Canto II (BD Biosciences) or sorting with a FACSAria II (BD Biosciences). B cell subsets are defined as pre-pro-B (B220+CD19−), pro-B (B220+CD19+IgM−CD43+), pre-B (B220+CD19+IgM−CD43−), large pre-B (B220+CD19+IgM−CD43−FSChi), small pre-B (B220+CD19+IgM−CD43−FSClo), immature B (B220+CD19+IgM+), BM mature B (B220hiCD19+), marginal zone B (B220+CD19+CD21hiCD23−), transitional B (B220+CD19+CD21+CD23+CD93+), follicular B (B220+CD19+CD21+CD23+CD93−), germinal center B (B220+CD19+IgM+Gl7+Fas+), memory B (CD19+CD38+CD138−Gl7−), class-switched IgG1+ B cells (B220+IgG1+) and plasma cells (B220loCD138+).
Apoptosis was measured in BM pro-B and pre-B cells stained with 7AAD (BD Biosciences) and Annexin V-FITC (Abcam), following the manufacturer’s instructions. To measure cell cycle distribution, BM pre-B cells were stained with surface markers and fixed and incubated with a solution containing 1 µg ml–1 DAPI in permeabilization buffer for at least 1 h before FACS analysis. All flow cytometry experiments were analyzed with FlowJo.
For SIRT7 and Pax5 intracellular staining, BM single-cell suspensions were stained and washed before fixation for 30 min and permeabilization with Foxp3/Transcription Factor Fixation/Permeabilization buffers (eBioscience) according to manufacturer’s instructions. Permeabilized cells were blocked with 2% FBS for 10 min, stained with anti-SIRT7 (D3K5A, Cell Signaling, 0.25 µg per sample) or anti-Pax5 (1H9, eBioscience, 0.2 µg per sample) for 1 h at room temperature and washed twice with permeabilization buffer containing 2% FBS. For SIRT7 staining, cells were subsequently incubated with a polyclonal anti-IgG (H + L) secondary antibody (Invitrogen, 0.25 µg per sample) for 1 h at room temperature before FACS analysis.
For phospho-STAT5-Y694 staining (47/Stat5(pY694), BD Biosciences, 20 µl per sample) in pre-B cells, BM cells were incubated in RPMI (Gibco) for 30 min at 37 °C and stimulated with IL-7 (5 ng ml–1) in RPMI for an additional 30 min at 37 °C. Cells were fixed with Foxp3/Transcription Factor Fixation buffer, followed by washing and incubation with ice-cold 100% methanol for 1 h. After Fc blocking, cells were stained and analyzed with a FACS Canto II.
Isolation of primary B cell progenitors
BM wild-type and Sirt7−/− pro-B cells were isolated by magnetic-activated cell separation enrichment of CD19+ cells, followed by cell sorting. Briefly, BM single-cell suspensions were prepared in staining buffer. Cells were stained with Fc-block for 20 min and an additional 30 min with a biotinylated antibody to CD19 (1D3, BD Biosciences, 0.1 µg per 10 million cells). After washing, stained cells were incubated with Streptavidin MicroBeads (Miltenyi) and separated magnetically. CD19+ cells were cultured overnight in Opti-MEM supplemented with 10% heat-inactivated FBS, 25 mM HEPES, 50 µg ml–1 gentamicin and 50 µM β-mercaptoethanol in the presence of 10 ng ml–1 IL-7, 10 ng ml–1 SCF and 10 ng ml–1 FTL3-L before Lin−CD19+B220+IgM− cell sorting. Fetal liver Pax5−/− B cell progenitors were obtained as described previously13.
Cells and reagents
HAFTL, TANOUE, NALM-20, REH, KOPN-8, SD-1, SEM and TOM-1 cells were cultured in RPMI supplemented with 10% heat-inactivated FBS and 100 U ml–1 penicillin/streptomycin (Gibco), whereas HEK293F, SIRT7−/− HEK293F (described in Simonet et al. 42) and Platinum E cells were grown in DMEM (Gibco) supplemented with 10% FBS and 100 U ml–1 penicillin/streptomycin. OP9 cells were maintained in MEM-α (Gibco) supplemented with 20% FBS and 100 U ml–1 penicillin/streptomycin. KOPN-8, NALM-20, REH, TANOUE, SD-1, TOM-1 and SEM cells were kindly provided by M. Parra (IJC; purchased from the DSMZ German Collection of Microorganisms and Cell Cultures). OP9 and HEK293F cells were purchased from ATCC, and Platinum E cells were purchased from Cell Biolabs. None of the cell lines used were found in the Commonly misidentified lines database. Primary pro-B cells from wild-type, Sirt7−/− or Pax5−/− mice were plated onto a layer of mitomycin C-inactivated OP9 feeder cells and grown on Opti-MEM supplemented with 10% heat-inactivated FBS, 25 mM HEPES, 50 µg ml–1 gentamicin and 50 µM β-mercaptoethanol in the presence of 10 ng ml–1 IL-7, 10 ng ml–1 SCF and 10 ng ml–1 FTL3-L. All cells were cultured at 37 °C in a humidified atmosphere containing 5% CO2. For transient transfections, HEK293F or SIRT7−/− HEK293F cells were transfected using polyethylenimine and the corresponding plasmids. For retroviral transduction of pro-B cells and B-ALL cell lines, Platinum E cells were transiently transfected with polyethylenimine, a pVSV-G vector encoding the viral envelop and pMIG bicistronic vectors encoding either the selection marker (hCD4 or GFP) alone or together with the SIRT7WT, SIRT7H187Y, Pax5WT, Pax5K198Q or Pax5K198R coding sequences. Pro-B cells were resuspended in retroviral supernatants, centrifuged for 1.5 h at 1,000g (32 °C) and selected by hCD4+ cell sorting 96 h after infection. Treatments were performed with 5 mM nicotinamide (Sigma) for 48 h, 100 µg ml–1 cycloheximide (Sigma-Aldrich) for the indicated times, 2 µM lactacystin (Santa Cruz Biotechnology) for 8 h and 1 µM trichostatin A (Sigma-Aldrich) for 3 h.
Histology
Spleens from wild-type and Sirt7−/− mice were collected, fixed in 10% formalin for 24 h, embedded in paraffin and sectioned at 4 µm before staining with hematoxylin and eosin. Histological sections were visualized with an Olympus BX53 microscope.
Immunoprecipitation, gel filtration high-performance liquid chromatography and immunoblotting
For immunoprecipitation, cell pellets were lysed in RIPA buffer (50 mM Tris-HCl (pH 8.0), 150 mM NaCl, 0.5% sodium deoxycholate, 0.1% SDS, 1% NP-40 and 2 mM MgCl2) containing cOmplete Protease Inhibitor (Roche) and incubated for 8 h with benzonase nuclease (Millipore) at 4 °C. Cell lysates were clarified by centrifugation (17,000g for 10 min at 4 °C) and incubated overnight with anti-Flag beads (Millipore) at 4 °C with gentle rotation. The immunoprecipitated protein complexes were washed five times with lysis buffer (20 mM Tris-HCl (pH 8.0), 500 mM NaCl, 10% glycerol and 1 mM EDTA) and eluted with Laemmli buffer supplemented with 10% β-mercaptoethanol. Samples were then boiled at 95 °C for 5 min and analyzed by immunoblotting. Densitometric quantification of immunoblotting experiments was performed with ImageJ software.
Cellular fractionation experiments were performed using the Dignam method, as described in Simonet et al.42. For size-exclusion chromatography analysis, nuclei from HAFTL cells were purified and lysed under native conditions according to the Dignam method, as previously described42. Nuclear lysates were incubated overnight with benzonase nuclease before clarification and concentration with Amicon Ultra centrifugal filters (Millipore). Concentrated nuclear lysates were then fractionated by molecular weight on the gel filtration column Superose 6 (Cytiva) with a fractionation range of 5 × 103–5 × 106 Da. The eluted fractions containing size-excluded proteins and protein complexes were denatured in Laemmli buffer supplemented with 10% β-mercaptoethanol and analyzed by immunoblotting. The following antibodies were used for immunoblotting: anti-SIRT7 (D3K5A, Cell Signaling, 1:1,000), anti-Pax5 (D19F8, Cell Signaling, 1:1,000), anti-H3 (ab1791, Abcam, 1:1,000), anti-fibrillarin (B1, Santa Cruz Biotechnology, 1:1,000), anti-acetyl-lysine (9814, Cell Signaling, 1:200), anti-Flag (M2, Sigma-Aldrich, 1:10,000), anti-HA (6908, Sigma), anti-Myc (9B11, Cell Signaling, 1:1,000), anti-V5 (ab9116, Abcam, 1:1,000) and anti-actin (A1978, Sigma, 1:5,000).
SIRT7 and Pax5 purification and in vitro deacetylation assay
SIRT7−/− HEK293F cells were transiently transfected with vectors encoding Pax5-Myc–Flag or SIRT7–Flag for 48 h. Before collection, cells expressing these constructs were treated overnight with 5 mM nicotinamide and for 3 h with 1 µM trichostatin A to hyperacetylate Pax5. Cell pellets were lysed, incubated with benzonase nuclease for 8 h, clarified and incubated overnight with anti-Flag beads (Millipore). The immunoprecipitated protein complexes were washed five times with BC500 buffer (20 mM Tris-HCl (pH 8.0), 500 mM NaCl, 10% glycerol and 1 mM EDTA), eluted with synthetic Flag peptide (0.6 µg ml–1; GenScript) and dialyzed in BC100 buffer (20 mM Tris-HCl (pH 8.0), 100 mM NaCl, 10% glycerol and 1 mM EDTA). Purified PAX5 and SIRT7 proteins were incubated for 1 h at 37 °C, with or without 1.25 mM NAD+, in deacetylation buffer (10 mM Tris-HCl (pH 8.0), 150 mM NaCl, 1 mM DTT and 10% glycerol), and the reaction was stopped with 5× Laemmli buffer containing 10% β-mercaptoethanol. PAX5 acetylation was determined by immunoblotting using anti-pan-acetyl-lysine.
Pre-B cell proteome and Pax5 acetylation analysis
For determination of the pre-B cell proteome, pre-B cells were sorted from the BM of wild-type and Sirt7−/− mice. Proteins were extracted with 6 M urea, 100 mM Tris (pH 8.0) and the help of a bioruptor, quantified using a NanoDrop at 280 nm and precipitated with trichloroacetic acid/acetone. Samples were then reduced and alkylated with 10 mM DTT and 55 mM chloroacetamide, respectively. Proteins were then resuspended in 6 M urea and 100 mM Tris (pH 8.0) and digested with LysC/trypsin. LysC digestion was performed for 16 h, while trypsin was added for 8 h, and both reactions were performed at 30 °C. The reactions were stopped with 10% formic acid. The peptides were then desalted with a C18 reverse-phase ultramicrospin column and desiccated in a speedvac. Total proteome samples were separated using a C18 analytical column (nanoEaseTM M/Z HSS C18 T3; 75 µm × 25 cm, 100 Å; Waters) with a 180-min run comprising three consecutive steps with linear gradients from 3% to 35% B in 150 min, from 35% to 50% B in 5 min and from 50% to 85% B in 2 min, followed by isocratic elution at 85% B in 5 min and stabilization to initial. The mass spectrometer was operated in data-dependent acquisition mode, and the data were acquired with Xcalibur software 4.0.27.10 (Thermo Scientific).
For identification of acetylated Pax5 residues, PAX5 was purified from SIRT7−/− HEK293F cells transiently expressing Pax5-Myc–Flag together with an empty vector or a vector encoding SIRT7–Flag. PAX5-containing beads were washed three times with 100 mM Tris (pH 8.0) and then resuspended in 6 M urea and 100 mM Tris (pH 8.0). Reduction and alkylation were then performed by using 10 mM and 55 mM chloroacetic acid. The digestion was performed by adding 1 μg of trypsin for 16 h at 30 °C. Finally, the digestion was stopped with 10% formic acid, and the peptides were desalted with a polyLC C18 pipette tip and dried in a speedvac. The acetylomes were separated using an Evosep EV1000 column (150 μm × 150 mm, 1.9 μm; Evosep) with an 88-min run. The spectrometer was working in positive polarity mode, and single-charge state precursors were rejected for fragmentation. The data were acquired with Xcalibur software 4.2.28.14 (Thermo Scientific).
For both total proteome and acetylome analyses, the peptides were reconstituted with 3% acetonitrile and 0.1% formic acid aqueous solution at 100 ng µl–1, and 800 ng was injected into the mass spectrometer.
Semiquantitative PCR, RT–qPCR and RNA-seq
Semiquantitative PCR of Igh segments was performed as described in Ng et al.43 using genomic DNA extracted from sorted splenic IgM+ cells and degenerate primers. RNA for RNA-seq and RT–qPCR was extracted from frozen pellets using a Maxwell RSC simplyRNA Tissue kit (Promega). For RT–qPCR, cDNA was synthesized using a Transcriptor First Strand cDNA Synthesis kit (Roche) according to manufacturer’s instructions. RT–qPCR reactions were performed in a QuantStudio 5 Real-Time PCR System. Primer sequences (Integrated DNA Technologies) are shown in Extended Data Table 1.
Extended Data Table 1.
List of primer sequences used for RT-qPCR and IgH recombination semi-quantitative PCR

RNA for RNA-seq was extracted from sorted wild-type and Sirt7−/− BM pro-B cells and pre-B cells or Pax5−/− ex vivo-expanded B cell progenitors expressing an empty vector, PAX5-WT, PAX5-K198Q or PAX5-K198R. After library construction, 150-bp paired-end sequencing was performed on a DNBSEQ-G400 (MGI Tech).
ChIP–seq
Chromatin was prepared using a truChIP Chromatin Shearing kit (Covaris) with modifications. Ten million pro-B cells were collected and washed once in PBS before fixation in 1 mg ml–1 DSG (Thermo Fisher) for 30 min, followed by an additional 10-min incubation with 1% formaldehyde. The cross-linking reaction was stopped by adding 1/20 quenching buffer, and fixed cells were washed twice with ice-cold 0.5% bovine serum albumin (wt/vol) in PBS. Nuclei were purified and sonicated following the manufacturer’s instructions. One volume of 2× dilution buffer supplemented with 0.1% SDS and protease inhibitors was added before clarifying the lysate (10,000g, 5 min, 4 °C). ChIP was performed overnight at 4 °C with 10 µg of anti-Pax5 (ab183575, Abcam) previously conjugated with 20 µl of Pierce ChIP-grade Protein A/G Magnetic Beads (Thermo Fisher) for 4 h at 4 °C. After immunoprecipitation, samples were serially washed once with low-salt wash buffer (0.1% SDS, 1% Triton, 2 mM EDTA, 20 mM Tris-HCl (pH 8.1) and 150 mM NaCl), high-salt wash buffer (0.1% SDS, 1% Triton, 2 mM EDTA, 20 mM Tris-HCl (pH 8.1) and 500 mM NaCl) and LiCl immune complex wash buffer (10 mM Tris-HCl (pH 8), 250 mM LiCl, 1% NP-40, 1% sodium deoxycholate and 1 mM EDTA) supplemented with protease inhibitors and twice with modified TE buffer (0.1 mM EDTA and 10 mM Tris-HCl (pH 8)). Cross-linking was reversed in elution buffer (0.1 M NaHCO3 and 1% SDS) with 10 µg of RNase A (Thermo Fisher) for 30 min at 37 °C, followed by an additional 6-h incubation at 65 °C with 50 µg of Proteinase K (Apollo Scientific). DNA was subsequently cleaned up using NucleoSpin Gel and PCR clean-up columns (Macherey-Nagel) before library construction and 100-bp paired-end sequencing on a DNBSEQ-G400.
Public data analysis
Mouse and human BM 10x scRNA-seq data were derived from the Broad Institute Single Cell Portal (https://singlecell.broadinstitute.org; projects ‘Bone Marrow from B6 Mice, 10x’ and ‘A Census of Immune Cells’, respectively). For human scRNA-seq analysis, the associated Loom file was downloaded and reanalyzed using Scanpy44. Cell-type annotation was performed using DecoupleR (v1.34)45 and PanglaoDB46. Mouse scRNA-seq feature and t-distributed stochastic neighbor embedding plots were directly downloaded from the Single Cell Portal. Microarray normalized data from mouse B cell progenitors were from the Immgen Consortium datasets47 (accession number GSE15907). Pax5-regulated genes were retrieved from accession number GSE38046 (ref. 20). Pax5 peaks were obtained from a publicly available PAX5 ChIP–seq dataset17. The list of mammalian proteins with acetyltransferase activity was from a previous report31, whereas Pax5 protein-protein interactions (PPIs) combined the data from another study19 with the curated interactions compiled in the BioGRID repository. Normalized proteomics and RNA-seq data for individuals with B-ALL were from Yang et al.29. Proteomics and RNA-seq raw data from this project are available at the ProteomeXchange Consortium (identifier PXD010175) and at the European Genome–Phenome Archive (accession EGAS00001003079), respectively. Relative Pax5 and SIRT7 protein levels in B-CLL samples were obtained from Johnston et al.34, and raw data are deposited at the PRIDE archive (PXD002004). Children’s Oncology Group clinical trial P9906 RNA expression and outcome data are available at the Genomic Data Commons (https://portal.gdc.cancer.gov) and were generated by the Therapeutically Applicable Research to Generate Effective Treatments (https://www.cancer.gov/ccg/research/genome-sequencing/target; GSE11877) initiative, whereas PAX5 deletion data in these individuals were retrieved from Roberts et al.37 (EGAS00001000654).
Proteomics data analysis
The raw thermo files were processed with MaxQuant 1.6.7.0 using a mouse database downloaded from https://www.uniprot.org/ (December 2018) or with PEAKS X+ software using a mouse database downloaded from https://www.uniprot.org/ (November 2019) for the total proteome and acetylome, respectively. In both cases, only reviewed entries were included (16,997 or 17,024 entries, respectively). The search was performed using the following parameters: trypsin was selected as the enzyme, and a maximum of two or three missed cleavages was allowed for the total proteome and acetylome. For the total proteome, the modifications were carbamidomethylation as a fixed modification, whereas oxidation in methionines and acetylation of protein N termini were used as variable modifications. The iBAQ intensity was used to quantify the proteins. Alternatively, for the acetylome, carbamidomethylation was set as a fixed modification, whereas oxidation in methionines; acetylation at both lysines and the protein N terminus; deamidation at asparagine and glutamine; phosphorylation at serines, threonines and tyrosines and dehydration at aspartic acid, tyrosines, threonines, serines, glutamine and asparagine were used as variable modifications. The mass tolerance for the parental ion and MS/MS fragments were set to 10 ppm and 0.5 Da. In both analyses, the results were filtered at 1% FDR at the peptide and protein levels.
Data processing and statistical analyses were performed using R (https://cran.r-project.org/) and RStudio (https://www.rstudio.com/) software. Total proteome statistical analysis was performed with the ‘limma’ package. For the PAX5 acetylome, the peptide spectrum matches were calculated by using a homemade algorithm and the PEAKS’ output files ‘peptide.csv’ and ‘DB search psm.csv’.
Bulk RNA-seq bioinformatic analysis
Bulk RNA-seq raw FastQ files were quality checked using FastQC, and raw counts were obtained by performing Salmon48 ‘quant’ pseudoalignment (mm10 reference mouse genome), with fragment-level GC bias correction (–gcBias), eight threads (-p 8) and selective alignment enabled (–validateMappings). The quant.sf files were used to import transcript-level quantification data into a Summarized Experiment using Tximeta49. The Summarized Experiment was imported into an R environment, and differential gene expression analysis was performed using DESeq2 (ref. 50). Counts were normalized by DESeq2’s median of ratios method51, and normalized counts were transformed to z score for data visualization (ggplot2). For generating BigWig files, raw FastQ files were mapped onto a reference mouse genome (mm10) using Bowtie2 (ref. 52) to generate SAM files. SAM files were converted to BAM files using SAMTools53, skipping alignments with MAPQ smaller than 37 (samtools view -bS -q 37). BAM files were sorted and filtered using Sambamba54, eliminating unmapped and duplicated reads (-F ‘[XS] == null and not unmapped and not duplicate’). Index BAI files were generated using SAMTools ‘index’, and BigWig files were created using deepTools55 ‘bamCoverage’ (–binSize 20–normalizeUsing BPM–ignoreForNormalization chrX–extendReads 150–centerReads–smoothLength 60). Visualization of BigWig files was performed using IGV56.
GSEA was performed using the software57 provided by the Broad Institute at https://www.gsea-msigdb.org/ with RNA-seq normalized count values and default parameters. Gene Ontology terms were obtained with the Enrichr tool58.
ChIP–seq bioinformatic analysis
ChIP–seq raw FastQ files were quality checked using FastQC and aligned onto a reference mouse genome (mm10) using Bowtie2 (ref. 52) to generate SAM files. SAM files were converted to BAM files using SAMTools53, skipping alignments with MAPQ smaller than 37 (samtools view -bS -q 37). BAM files were sorted and filtered using Sambamba54, eliminating unmapped and duplicated reads (-F ‘[XS] == null and not unmapped and not duplicate’). Index BAI files were generated using SAMTools ‘index’, and BigWig files were created using deepTools55 ‘bamCoverage’ (–binSize 20–normalizeUsing BPM–ignoreForNormalization chrX–extendReads 150–centerReads–smoothLength 60). Visualization of BigWig files was performed using IGV56. Peak calling was performed using MACS2 (ref. 59) ‘callpeak’ (-f BAM–nomodel–extsize 20 -g mm -B) taking into account both immunoprecipitation (-t) and input (-c) samples for each peak calling. Only significant peaks (q < 0.05) were included in downstream analyses. Differential peak analysis was performed with ChIPpeakAnno, and heat maps were generated with DiffBind or plotHeatmap.
Statistics and reproducibility
All statistical analyses were performed using GraphPad Prism 8.0.1. The statistical tests performed for data analysis are indicated in the corresponding figure legends. Individual P values of significant data are indicated in the figures. Results were similarly replicated in at least two independent experiments, and results pooled from independent experiments are indicated in the corresponding figure legends. For experiments involving mice, each replicate corresponds to an individual mouse, and no animals or data points were excluded. RNA-seq, ChIP–seq and proteomics replicates were generated on different days. No statistical methods were used to predetermine sample sizes. Data distribution was assumed to be normal, but this was not formally tested. Only recipient mice for transplantation experiments were randomized because the rest of the experiments required mouse genotyping and were performed on genetically identical mice. Data collection and analysis were not performed blind to the conditions of the experiments.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Online content
Any methods, additional references, Nature Portfolio reporting summaries, source data, extended data, supplementary information, acknowledgements, peer review information; details of author contributions and competing interests; and statements of data and code availability are available at 10.1038/s41590-024-01995-7.
Supplementary information
RNA-seq results from wild-type and Sirt7−/− B220+CD19+IgM–CD43+ pro-B cells and B220+CD19+IgM–CD43– pre-B cells sorted from mouse BM.
Quantitative MS results from sorted wild-type and Sirt7−/− B220+CD19+IgM–CD43– pre-B cells.
List of Pax5 protein–protein interactions and mammalian acetyltransferases.
Genes bound by Pax5WT, Pax5K198Q and Pax5K198R proteins from ChIP–seq data.
RNA-seq results from ex vivo-expanded Pax5−/− pro-B cells expressing empty vector, Pax5WT, Pax5K198Q or Pax5K198R.
RNA-seq results from ex vivo-expanded Sirt7−/− pro-B cells expressing empty vector, Pax5WT or Pax5K198R.
Source data
Unprocessed western blots and/or gels.
Statistical source data.
Statistical source data.
Unprocessed western blots and/or gels.
Statistical source data.
Statistical source data.
Unprocessed western blots and/or gels.
Statistical source data.
Statistical source data.
Statistical source data.
Unprocessed western blots and/or gels.
Statistical source data.
Unprocessed western blots and/or gels.
Statistical source data.
Statistical source data.
Statistical source data.
Acknowledgements
We are grateful to M. Parra and O. de Barrios (IJC) for providing B-ALL cell lines. We also thank the members of the laboratory of A.V. for stimulating discussions. This work was supported by a grant from the Scientific Foundation of the Spanish Association Against Cancer (AECC; grant number PROYE20042VAQU), the Spanish Ministry of Economy and Competitiveness (MINECO; SAF2017-88975R) and a grant PID2020-117284RB-I00 funded by MICIU/AEI/10.13039/501100011033 to A.V., the Spanish National Institute of Health Carlos III-ISCIII (ProteoRed) to A.V. and C.D.L.T., CP19/00176 to J.L.S. (cofunded by the European Social Fund ‘Investing in your future’), FI-AGAUR FI_B 00293 to A.G.-G., the European Commission’s Horizon research and innovation programme Marie Skłodowska-Curie (2020, 895979 to B.N.V.; 2021, 101065013 to A.I.), Beatriu de Pinos fellowship (2016-BP-00250 to B.N.V.), the Catalan Government Agency AGAUR (2017-SGR-148, 2021-SGR-01378 to A.V.; 2021-SGR-01494 to M.E.), the Human Genetics Institute of New Jersey (to L.S. and J.A.T.) and the Swedish Cancer Foundation (20-1153), the Childhood Cancer Foundation (2022-0019) and the Swedish Research Council (2021-02379) to M.S. Research at the laboratory of J.L.S. was supported by grant PID2019-111243RA-I00 funded by MICIU/AEI/10.13039/501100011033. The IJC Proteomics Unit is part of the Spanish Platform of Molecular and Bioinformatics Resources (ProteoRed), Instituto de Salud Carlos III (PT13/0001) and Proyecto Proteoma del Cáncer Participación Española en el CPTAC Internacional, The National Cancer Institute’s Clinical Proteomic Tumour Analysis Consortium. We also thank the CERCA Programme/Generalitat de Catalunya for institutional support.
Extended data
Author contributions
A.G.-G., B.N.V. and A.V. planned the project, designed the experiments and wrote the manuscript. A.V. and B.N.V. supervised the work, and M.S. provided critical reagents and key discussions on the experimental design and the manuscript. A.G.-G., M.E.-A., B.N.V., M.S., A.M.-D., E.A.-P., C.R., C.B., P.K., A.I. and L.S. performed experiments and/or analyzed data. A.G.-G., A.B.-C., J.L.S. and J.K.T. analyzed RNA-seq and ChIP–seq experiments. J.J.B. and C.D.L.T. performed and analyzed the proteomics experiments. M.E., L.S., J.L.S., A.I., T.B. and J.A.T. provided reagents and their expertise and participated in the discussion.
Peer review
Peer review information
Nature Immunology thanks Rhys Allan and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Ioana Staicu, in collaboration with the Nature Immunology team. Peer reviewer reports are available.
Data availability
All Pax5WT, Pax5K198Q and Pax5K198R ChIP–seq data and RNA-seq data from wild-type and Sirt7−/− BM pro-B and pre-B cells; Pax5−/− B cell progenitors retrovirally expressing empty vector, Pax5WT, Pax5K198Q or Pax5K198R and wild-type or Sirt7−/− B cell progenitors expressing empty vector, Pax5WT or Pax5K198R have been deposited at the National Center for Biotechnology Information Gene Expression Omnibus and are available under the accession number GSE246370. All sequencing data were aligned to the mm10 reference mouse genome. Proteomics data from wild-type and Sirt7−/− BM pre-B cells and Pax5 acetylation experiments have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifier PXD046457. Source data are provided with this paper.
Competing interests
The authors declare no competing interests.
Footnotes
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
These authors contributed equally: Maria Espinosa-Alcantud, Alberto Bueno-Costa.
Contributor Information
Berta N. Vazquez, Email: bvazquez@carrerasresearch.org
Alejandro Vaquero, Email: avaquero@carrerasresearch.org.
Extended data
is available for this paper at 10.1038/s41590-024-01995-7.
Supplementary information
The online version contains supplementary material available at 10.1038/s41590-024-01995-7.
References
- 1.Medvedovic, J., Ebert, A., Tagoh, H. & Busslinger, M. PAX5: a master regulator of B cell development and leukemogenesis. Adv. Immunol.111, 179–206 (2011). [DOI] [PubMed] [Google Scholar]
- 2.Delogu, A. et al. Gene repression by PAX5 in B cells is essential for blood cell homeostasis and is reversed in plasma cells. Immunity24, 269–281 (2006). [DOI] [PubMed] [Google Scholar]
- 3.Cobaleda, C., Schebesta, A., Delogu, A. & Busslinger, M. PAX5: the guardian of B cell identity and function. Nat. Immunol.8, 463–470 (2007). [DOI] [PubMed] [Google Scholar]
- 4.Schebesta, A. et al. Transcription factor PAX5 activates the chromatin of key genes involved in B cell signaling, adhesion, migration, and immune function. Immunity27, 49–63 (2007). [DOI] [PubMed] [Google Scholar]
- 5.Urbanek, P., Wang, Z. Q., Fetka, I., Wagner, E. F. & Busslinger, M. Complete block of early B cell differentiation and altered patterning of the posterior midbrain in mice lacking PAX5/BSAP. Cell79, 901–912 (1994). [DOI] [PubMed] [Google Scholar]
- 6.Nutt, S. L., Thevenin, C. & Busslinger, M. Essential functions of PAX-5 (BSAP) in pro-B cell development. Immunobiology198, 227–235 (1997). [DOI] [PubMed] [Google Scholar]
- 7.Nutt, S. L., Heavey, B., Rolink, A. G. & Busslinger, M. Commitment to the B-lymphoid lineage depends on the transcription factor PAX5. Nature402, 556–562 (1999). [DOI] [PubMed] [Google Scholar]
- 8.Mullighan, C. G. et al. Genome-wide analysis of genetic alterations in acute lymphoblastic leukaemia. Nature446, 758–764 (2007). [DOI] [PubMed] [Google Scholar]
- 9.Heltemes-Harris, L. M. et al. Ebf1 or Pax5 haploinsufficiency synergizes with STAT5 activation to initiate acute lymphoblastic leukemia. J. Exp. Med.208, 1135–1149 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Dang, J. et al. PAX5 is a tumor suppressor in mouse mutagenesis models of acute lymphoblastic leukemia. Blood125, 3609–3617 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Prasad, M. A. et al. Ebf1 heterozygosity results in increased DNA damage in pro-B cells and their synergistic transformation by Pax5 haploinsufficiency. Blood125, 4052–4059 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Johanson, T. M. et al. Transcription-factor-mediated supervision of global genome architecture maintains B cell identity. Nat. Immunol.19, 1257–1264 (2018). [DOI] [PubMed] [Google Scholar]
- 13.Somasundaram, R. et al. EBF1 and PAX5 control pro-B cell expansion via opposing regulation of the Myc gene. Blood137, 3037–3049 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Ramamoorthy, S. et al. EBF1 and PAX5 safeguard leukemic transformation by limiting IL-7 signaling, Myc expression, and folate metabolism. Genes Dev.34, 1503–1519 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Medvedovic, J. et al. Flexible long-range loops in the Vh gene region of the Igh locus facilitate the generation of a diverse antibody repertoire. Immunity39, 229–244 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Fuxa, M. et al. PAX5 induces V-to-DJ rearrangements and locus contraction of the immunoglobulin heavy-chain gene. Genes Dev.18, 411–422 (2004). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Hill, L. et al. Wapl repression by PAX5 promotes V gene recombination by Igh loop extrusion. Nature584, 142–147 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.McManus, S. et al. The transcription factor PAX5 regulates its target genes by recruiting chromatin-modifying proteins in committed B cells. EMBO J.30, 2388–2404 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Okuyama, K. et al. PAX5 is part of a functional transcription factor network targeted in lymphoid leukemia. PLoS Genet.15, e1008280 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Revilla, I. D. R. et al. The B-cell identity factor PAX5 regulates distinct transcriptional programmes in early and late B lymphopoiesis. EMBO J.31, 3130–3146 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Bosch-Presegue, L. & Vaquero, A. Sirtuins in stress response: guardians of the genome. Oncogene33, 3764–3775 (2014). [DOI] [PubMed] [Google Scholar]
- 22.Gamez-Garcia, A. & Vazquez, B. N. Nuclear sirtuins and the aging of the immune system. Genes12, 1856 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Kumari, P., Tarighi, S., Braun, T. & Ianni, A. SIRT7 acts as a guardian of cellular integrity by controlling nucleolar and extra-nucleolar functions. Genes12, 1361 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Vazquez, B. N. et al. SIRT7 promotes genome integrity and modulates non-homologous end joining DNA repair. EMBO J.35, 1488–1503 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Vakhrusheva, O. et al. SIRT7 increases stress resistance of cardiomyocytes and prevents apoptosis and inflammatory cardiomyopathy in mice. Circ. Res.102, 703–710 (2008). [DOI] [PubMed] [Google Scholar]
- 26.Goetz, C. A., Harmon, I. R., O’Neil, J. J., Burchill, M. A. & Farrar, M. A. STAT5 activation underlies IL7 receptor-dependent B cell development. J. Immunol.172, 4770–4778 (2004). [DOI] [PubMed] [Google Scholar]
- 27.Difilippantonio, S. et al. 53BP1 facilitates long-range DNA end-joining during V(D)J recombination. Nature456, 529–533 (2008). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Mason, D. Y., Jones, M. & Goodnow, C. C. Development and follicular localization of tolerant B lymphocytes in lysozyme/anti-lysozyme IgM/IgD transgenic mice. Int. Immunol.4, 163–175 (1992). [DOI] [PubMed] [Google Scholar]
- 29.Yang, M. et al. Proteogenomics and Hi-C reveal transcriptional dysregulation in high hyperdiploid childhood acute lymphoblastic leukemia. Nat. Commun.10, 1519 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Gruenbacher, S., Jaritz, M., Hill, L., Schafer, M. & Busslinger, M. Essential role of the PAX5 C-terminal domain in controlling B cell commitment and development. J. Exp. Med.220, e20230260 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Hyndman, K. A. & Knepper, M. A. Dynamic regulation of lysine acetylation: the balance between acetyltransferase and deacetylase activities. Am. J. Physiol. Ren. Physiol.313, F842–F846 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Ungerback, J., Ahsberg, J., Strid, T., Somasundaram, R. & Sigvardsson, M. Combined heterozygous loss of Ebf1 and Pax5 allows for T-lineage conversion of B cell progenitors. J. Exp. Med.212, 1109–1123 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Oksenych, V., Su, D. & Daniel, J. A. Acetyltransferases GCN5 and PCAF are required for B lymphocyte maturation in mice. Biomolecules12, 61 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Johnston, H. E. et al. Proteomics profiling of CLL versus healthy B-cells identifies putative therapeutic targets and a subtype-independent signature of spliceosome dysregulation. Mol. Cell Proteom.17, 776–791 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Harvey, R. C. et al. Identification of novel cluster groups in pediatric high-risk B-precursor acute lymphoblastic leukemia with gene expression profiling: correlation with genome-wide DNA copy number alterations, clinical characteristics, and outcome. Blood116, 4874–4884 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Kang, H. et al. Gene expression classifiers for relapse-free survival and minimal residual disease improve risk classification and outcome prediction in pediatric B-precursor acute lymphoblastic leukemia. Blood115, 1394–1405 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Roberts, K. G. et al. Targetable kinase-activating lesions in Ph-like acute lymphoblastic leukemia. N. Engl. J. Med.371, 1005–1015 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Fuxa, M. & Busslinger, M. Reporter gene insertions reveal a strictly B lymphoid-specific expression pattern of PAX5 in support of its B cell identity function. J. Immunol.178, 3031–3037 (2007). [DOI] [PubMed] [Google Scholar]
- 39.Lu, Y. F. et al. SIRT7 activates p53 by enhancing PCAF-mediated MDM2 degradation to arrest the cell cycle. Oncogene39, 4650–4665 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Liu, G. J. et al. PAX5 loss imposes a reversible differentiation block in B-progenitor acute lymphoblastic leukemia. Genes Dev.28, 1337–1350 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Chan, L. N. et al. Metabolic gatekeeper function of B-lymphoid transcription factors. Nature542, 479–483 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Simonet, N. G. et al. SIRT7 auto-ADP-ribosylation regulates glucose starvation response through mH2A1. Sci. Adv.6, 2590 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Ng, A. P. et al. An Erg-driven transcriptional program controls B cell lymphopoiesis. Nat. Commun.11, 3013 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol.19, 15 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Badia, I. M. P. et al. decoupleR: ensemble of computational methods to infer biological activities from omics data. Bioinform. Adv.2, vbac016 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Franzen, O., Gan, L. M. & Bjorkegren, J. L. M. PanglaoDB: a web server for exploration of mouse and human single-cell RNA sequencing data. Database2019, baz046 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Heng, T. S. & Painter, M. W.; Immunological Genome Project Consortium. The Immunological Genome Project: networks of gene expression in immune cells. Nat. Immunol.9, 1091–1094 (2008). [DOI] [PubMed] [Google Scholar]
- 48.Patro, R., Duggal, G., Love, M. I., Irizarry, R. A. & Kingsford, C. Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods14, 417–419 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Love, M. I. et al. Tximeta: reference sequence checksums for provenance identification in RNA-seq. PLoS Comput. Biol.16, e1007664 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol.15, 550 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol.11, R106 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods9, 357–359 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics25, 2078–2079 (2009). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Tarasov, A., Vilella, A. J., Cuppen, E., Nijman, I. J. & Prins, P. Sambamba: fast processing of NGS alignment formats. Bioinformatics31, 2032–2034 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Ramirez, F., Dundar, F., Diehl, S., Gruning, B. A. & Manke, T. deepTools: a flexible platform for exploring deep-sequencing data. Nucleic Acids Res.42, W187–W191 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Thorvaldsdottir, H., Robinson, J. T. & Mesirov, J. P. Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief. Bioinform.14, 178–192 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA102, 15545–15550 (2005). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Kuleshov, M. V. et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res.44, W90–W97 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Zhang, Y. et al. Model-based analysis of ChIP–seq (MACS). Genome Biol.9, 137 (2008). [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
RNA-seq results from wild-type and Sirt7−/− B220+CD19+IgM–CD43+ pro-B cells and B220+CD19+IgM–CD43– pre-B cells sorted from mouse BM.
Quantitative MS results from sorted wild-type and Sirt7−/− B220+CD19+IgM–CD43– pre-B cells.
List of Pax5 protein–protein interactions and mammalian acetyltransferases.
Genes bound by Pax5WT, Pax5K198Q and Pax5K198R proteins from ChIP–seq data.
RNA-seq results from ex vivo-expanded Pax5−/− pro-B cells expressing empty vector, Pax5WT, Pax5K198Q or Pax5K198R.
RNA-seq results from ex vivo-expanded Sirt7−/− pro-B cells expressing empty vector, Pax5WT or Pax5K198R.
Unprocessed western blots and/or gels.
Statistical source data.
Statistical source data.
Unprocessed western blots and/or gels.
Statistical source data.
Statistical source data.
Unprocessed western blots and/or gels.
Statistical source data.
Statistical source data.
Statistical source data.
Unprocessed western blots and/or gels.
Statistical source data.
Unprocessed western blots and/or gels.
Statistical source data.
Statistical source data.
Statistical source data.
Data Availability Statement
All Pax5WT, Pax5K198Q and Pax5K198R ChIP–seq data and RNA-seq data from wild-type and Sirt7−/− BM pro-B and pre-B cells; Pax5−/− B cell progenitors retrovirally expressing empty vector, Pax5WT, Pax5K198Q or Pax5K198R and wild-type or Sirt7−/− B cell progenitors expressing empty vector, Pax5WT or Pax5K198R have been deposited at the National Center for Biotechnology Information Gene Expression Omnibus and are available under the accession number GSE246370. All sequencing data were aligned to the mm10 reference mouse genome. Proteomics data from wild-type and Sirt7−/− BM pre-B cells and Pax5 acetylation experiments have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifier PXD046457. Source data are provided with this paper.