Abstract
The p53 tumor suppressor protein, found mutated in over 50% of all human tumors, is a sequence-specific transcriptional activator. Recent studies have identified a p53 relative, termed p73. We were interested in determining the relative abilities of wild-type and mutant forms of p53 and p73α and -β isoforms to transactivate various p53-responsive promoters. We show that both p73α and p73β activate the transcription of reporters containing a number of p53-responsive promoters in the p53-null cell line H1299. However, a number of significant differences were observed between p53 and p73 and even between p73α and p73β. Additionally, a Saccharomyces cerevisiae-based reporter assay revealed a broad array of transcriptional transactivation abilities by both p73 isoforms at 37°C. Recent data have shown that p73 can associate with p53 by the yeast two-hybrid assay. When we examined complex formation in transfected mammalian cells, we found that p73α coprecipitates with mutant but not wild-type p53. Since many tumor-derived p53 mutants are capable of inhibiting transactivation by wild-type p53, we tested the effects of two representative hot-spot mutants (R175H and R248W) on p73. By cotransfecting p73α along with either p53 mutant and a p53-responsive reporter, we found that both R175H and R248W reduces the transcriptional activity of p73α. This decrease in transcriptional activity is correlated with the reduced ability of p73α to promote apoptosis in the presence of tumor-derived p53 mutants. Our data suggest the possibility that in some tumor cells, an outcome of the expression of mutant p53 protein may be to interfere with the endogenous p73 protein.
A new gene family whose encoded products show significant sequence similarity to the tumor suppressor protein p53 have been identified (32, 33, 50, 59, 60, 72). KET, the first to be identified, was cloned from a rat circumvallate taste papilla cDNA library (59). p73, the second identified from a COS cell cDNA library, encodes for at least two splicing variants, p73α and p73β (32, 33). Finally, the human homolog of KET, referred to as either p51 or p63, encodes at least six isoforms (p63α/p51B/p73L, p63β, p63γ/p51A, ΔNp63α, ΔNp63β, and ΔNp63γ) that are expressed in a tissue-specific manner and harbor different transactivation potentials (50, 60, 72). It has been proposed that this family of proteins is ancestral to human p53, in that all show significant amino acid similarity in their C-terminal p53-unrelated extensions to the squid p53 protein (33, 59, 72).
The p53 protein is modular and can be divided into at least four distinct domains: (i) the amino-terminal transcriptional transactivation domain (residues ∼1 to 70) (6, 7, 15, 55, 68), (ii) the PXXP domain (residues ∼61 to 94) (70), (iii) the sequence-specific DNA binding domain (residues ∼102 to 292) (1, 27, 51, 71), and (iv) the carboxy-terminal regulatory and tetramerization domains (residues ∼320 to 393 and ∼320 to 360, respectively) (3, 4, 57, 71). The various isoforms of p73 and p51/p63 display a modular structure similar to that of p53, having extensive homology to p53 within their DNA binding domains (63 and 60%, respectively), as well as possessing homologous amino-terminal transcriptional transactivation domains (29 and 22% [except ΔNp63 isoforms], respectively) and tetramerization domains (38 and 37%, respectively).
Given this degree of amino acid similarity between p53, p51/p63, and p73, it is not surprising that ectopic expression of p73 and p51/p63 can transactivate endogenous targets of p53, such as the cell cycle inhibitor gene p21 (32, 33, 50) as well as p21 and RGC (ribosomal gene cluster) promoter-containing reporters in p53-null cell lines (32, 50, 72). However, it is of interest to ascertain whether the various isoforms of p73 and p51/p63 are also capable of transactivating additional physiologically relevant p53 targets, such as the proapoptotic genes Bax (49) and IGF-BP3 (insulin-like growth factor binding protein 3) (5), and others (reviewed in references 22 and 37). The percent homology between the tetramerization domains of p53, p51/p63, and p73 suggests that these protein families may form heterotetramers. Indeed, Kaghad et al. have shown that p73β, but not p73α, can interact modestly with p53 in a yeast-two hybrid assay (33). It remains to be determined if these complexes can form in mammalian cells and whether they function during the developmental and/or pathological process.
One of the cellular functions of p53 is to induce apoptosis in response to genotoxic stress, such as damaged DNA (reviewed in references 22, 37, and 41). Similarly, it has been found that overexpression of both p73 and p51/p63 can inhibit cell growth by inducing apoptosis (32, 50, 72). However, despite these similarities to p53, p73 is not induced by exposure of cells to DNA-damaging agents such as UV irradiation (33), suggesting that p73 may have cellular functions distinct from those of p53. Supporting this notion is the fact that in contrast to the ubiquitous expression of p53, p51/p63 and KET have restricted tissue expression patterns (50, 59, 72). Nevertheless, mutations in p51 have been identified in some human epidermal tumors, whereas p73 is monoallelically expressed in cancers, including neuroblastoma, suggesting a potential tumor suppressor role for both p51 and p73 (33, 50).
In this study, we examined the ability of ectopically expressed p73α and p73β to transactivate p53-responsive reporters in the p53-null cell line H1299 as well as in a yeast-based reporter assay. Additionally, we determined if p73 can associate with wild-type and tumor-derived mutant p53 in mammalian cells. Based on these coimmunoprecipitation studies, we analyzed the effects of tumor-derived p53 mutants on p73 function in transient cotransfection assays. Finally, we discuss the role that tumor-derived p53 mutants may play in cellular transformation by their ability to selectively inactive other p53 family members.
MATERIALS AND METHODS
Yeast strains, media, and transformation.
All yeast strains were isogenic with S288C except that they were wild type at GAL2. Prior to the introduction of the wild-type or mutant p53, p73α, and p73β constructs, CUY5 (trp1-1 ura3-52 his3,200 leu2-3-112 lys2-801) was transformed with one of the HIS3 reporter plasmids (see below) or control plasmid. Rich (YP) and synthetic complete (SC) media were constituted as described elsewhere (58) except that the YP medium also contained 0.1 g of tryptophan per liter and 0.1 g of adenine per liter and the SC medium contained 0.2 g of leucine, 0.1 g of all other amino acids, 0.1 g of uracil, and 0.1 g of adenine per liter. All strains were grown in glucose to a final concentration of 2%. Yeast strains were transformed by a modified version of the lithium acetate method (24) as described elsewhere (12).
Yeast and mammalian expression plasmids.
pLS76 (pCUB7) and pFW601 (pCUB6) express full-length p53 and p53R273H cDNAs from the ADH1 (alcohol dehydrogenase) promoter with the CYC1 terminator downstream of the p53 cDNA, CEN6, and ARSH4 for stable, low-copy-number replication and the LEU2 gene for plasmid maintenance (29). The plasmid expressing full-length mutant p53R273H was isolated by FASAY (29) and sequenced to confirm that the p53 cDNA contained the missense mutation. pCUB274 expresses full-length hemagglutin epitope (HA) tagged p53 (HA:p53) cDNA from the ADH1 promoter, with the CYC1 terminator downstream of the p53 cDNA, the yeast 2 μm origin of replication, and the LEU2 gene for plasmid maintenance. pC53-SN3, pC53-175, and pC53-248 express full-length p53, p53R175H, and p53R248W cDNAs, respectively, from the cytomegalovirus (CMV) promoter (35). pCUB263 expresses full-length HA:p53 cDNA from the CMV promoter in pCDNA3. pCUB215, pCUB217, pCUB219, and pCUB221 express full-length HA:p73α, HA:p73αR292H, HA:p73β, and HA:p73βR292H cDNAs, respectively, from the ADH1 promoter, with the CYC1 terminator downstream of the p53 cDNA, the yeast 2 μm origin of replication, and the LEU2 gene for plasmid maintenance. Expression of simian HA:p73α, HA:p73αR292H, HA:p73β, and HA:p73βR292H cDNAs from the CMV promoter are as described elsewhere (32).
Yeast and mammalian reporter plasmids.
Yeast p53-responsive reporter plasmids are as described elsewhere (12). Briefly, a duplex oligonucleotide encoding one of the p53-responsive cis-acting elements was cloned upstream of the inactive GAL1 promoter which drives the HIS3 coding sequence on a TRP1/CEN plasmid. The RGC-containing p53-responsive reporter plasmid pSS1 (29) and the mammalian p53-responsive reporter plasmids (19) are as described elsewhere. p21min-luc (pCUB230) contains a duplex oligonucleotide encoding the p53-responsive cis-acting element from p21, cloned upstream of the minimal c-fos promoter (−53 to +42) in pGL3-OFLUC (kindly provided by N. Clarke). The sequences of the synthesized oligonucleotides (CUO3 and CUO4; Operon Technologies, Inc.) encoding the p21 cis-acting p53-responsive element were 5′GATCCTCGAGGAACATGTCCCAACATGTTGCTCGAG3′ and 5′GATCCTCGAGCAACATGTTGGGACATGTTCCTCGAG3′. The resulting plasmid was sequenced for the orientation and insert number of the oligoduplex.
Preparation of yeast whole-cell extracts and detection of p53, p73α, and p73β by immunoprecipitation and Western immunoblot analysis.
Procedures for preparation of yeast whole-cell extracts and Western immunoblotting were as described elsewhere (12). Immunoprecipitations of p53 proteins were performed by incubating 5 mg of whole-cell extract with 4 μl of anti-p53 monoclonal antibodies (MAbs; at 50 ng/μl) and rocking at 4°C for 1 h. After the primary incubation, 30 μl of protein A-Sepharose beads (Pharmacia) was added, and the samples were rocked at 4°C for 1 h. The samples were washed four times with 1 ml of lysis buffer (100 mM Tris-HCl [pH 7.5], 200 mM NaCl, 1 mM EDTA, 5% glycerol, 0.5 mM dithiothreitol [DTT], 1 mM phenylmethylsulfonyl fluoride), excess liquid was aspirated with a 1-ml syringe, and 30 μl of 2× sample buffer (62) was added. Samples were heated to 95°C for 5 min, centrifuged for 3 min at 13,000 × g and electrophoresed through a sodium dodecyl sulfate (SDS)–10% polyacrylamide gel. Protein gels were transferred to polyvinylidene fluoride membranes (Millipore). For p53 detection, we used a mixture of purified p53 MAbs (MAb 421, MAb 1801, MAb 240, and DO-1), each at a 1/3,000 dilution of a 50-ng/ml stock; for HA:p53, HA:p73α, and HA:p73β detection, the primary MAb was 12CA5 at a 1/3,000 dilution of a 50-ng/ml stock. Proteins were visualized with an enhanced chemiluminescence detection system (Amersham).
Cell culture, transfection, and luciferase assays.
H1299 cells (American Type Culture Collection) were maintained in Dulbecco’s modified Eagle medium (DMEM) supplemented with 10% fetal bovine serum (FBS) in 5% CO2 at 37°C. Cells were transfected by a lipopolyamine-based (Transfectam) protocol as described previously (21). Briefly, cells were grown in DMEM–10% FBS and transfected with various amounts of DNA. The precipitate was left on the cells for 6 h, after which fresh DMEM–10% FBS was added for the periods indicated. For luciferase assays, cells were seeded in 12-well, 3.8-cm2 plates and transfected with one of the expression vectors (200 ng of each) and two different reporter constructs (250 ng of each), a CMV-expressed luciferase cDNA from renilla and a p53-responsive luciferase cDNA from firefly. Luciferase activity was measured in each well 24 h later by a dual-luciferase reporter gene assay (Promega).
Preparation of mammalian whole-cell extracts and immunoprecipitation analysis.
H1299 cells were transfected in 10-cm plates with 20 μg of DNA and harvested at 48 h posttransfection. Cells were lysed in 300 μl of lysis buffer (10 mM Tris-HCl [pH 7.5], 1 mM EDTA, 0.5% NP-40, 150 mM NaCl, 1 mM DTT, 10% glycerol, 0.5 mM phenylmethylsulfonyl fluoride, protease inhibitor), and the extracts were centrifuged at 8,000 × g for 15 min to remove cell debris. Protein concentrations were determined by the Bio-Rad Laboratories (Hercules, Calif.) assay. Immunoprecipitations of p53 proteins were performed by incubating 1.5 mg of whole-cell extract with 100 μg each of anti-p53 MAbs 240, 421, and 1801 and rocking at 4°C for 1 h. After the primary incubation, 20 μl of protein A-Sepharose beads (Pharmacia) was added, and the samples were rocked at 4°C for 1 h. The samples were washed four times with 1 ml of wash buffer (10 mM Tris-HCl [pH 7.5], 1 mM EDTA, 0.5% NP-40, 1 mM DTT, 10% glycerol), the excess liquid was aspirated, and 30 μl of 2× sample buffer (62) was added. Samples were heated to 95°C for 5 min, centrifuged for 3 min at 13,000 × g, and electrophoresed through an SDS–10% polyacrylamide gel. Protein gels were transferred to nitrocellulose membranes (Schleicher & Schuell). For p53 detection, a mixture of p53 MAb (MAb 421, 1801, or 240) containing supernatants was used, each at a 1/4 dilution; for HA:p73α detection (Fig. 5C), a supernatant containing MAb 12CA5 was used at a 1/3 dilution; for HA:p73α, HA:p73β, and HA:p53 detection (Fig. 1C), MAb 16B12 (1 mg/ml; BAbCo) was used at 1/1,000. Proteins were visualized with an enhanced chemiluminescence detection system (Amersham).
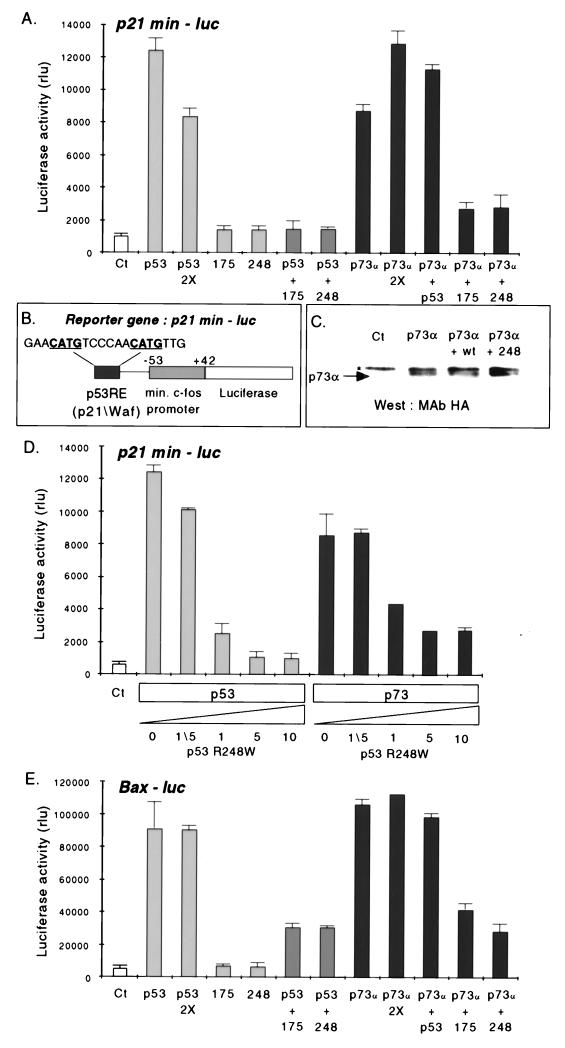
FIG. 5.
Tumor-derived p53 mutants reduce the transcriptional activity of p73α. (A) H1299 cells were transiently cotransfected with a reporter plasmid (p21min-luc) containing a derived p53-responsive human target gene cis-acting element from the p21 promoter (see panel B and Materials and Methods), and a plasmid expressing either p73α, wild-type p53 (p53), p53R175H (175), or p53R248W (248). In each transfection, equivalent amounts of DNA were used for all expression plasmids except where indicated (“2×” refers to a twofold increase in the amount of expression plasmid DNA transfected). (C) Representative Western blot loaded with 100 μg of extract and probed with anti-HA antibody 12CA5 to detect p73α in cells expressing either p73α alone, p73α plus wild-type p53 (wt), or p73α plus p53R248W (248). Ct, control plasmid; ∗, a nonspecific protein band that migrates more slowly than p73α and cross-reacts with the anti-HA antibody. (D) H1299 cells were cotransfected with increasing amounts of the p53R248W expression plasmid and a constant amount of either wild-type p53 or p73α and the p21min-luc reporter as for panel B. Ct, control plasmid; 0, 1/5, 1, 5, and 10, fold excess of p53R248W DNA used in the cotransfection. (E) H1299 cells were cotransfected as described for panel A except that the luciferase reporter construct used (Bax-luc) contained the full length Bax promoter. For the luciferase assays in panels A, D, and E, histograms represent relative luciferase units (rlu) and diagrams show the mean of a typical experiment of three performed in triplicate (bars indicate standard deviations).
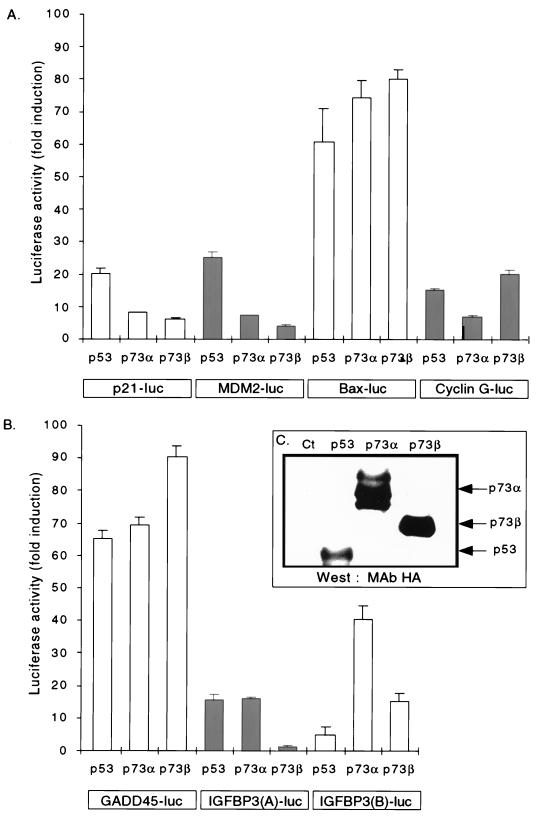
FIG. 1.
Wild-type p53, p73α, and p73β transactivate p53-responsive reporters in mammalian cells (A and B). H1299 cells (grown in DMEM supplemented with 10% FBS) were transiently cotransfected with the CMV-HA:p53, CMV-HA:p73α, or CMV-HA:p73β expression plasmid along with the luciferase (luc) reporter constructs indicated in boxes at the bottom. Cells were harvested for luciferase assay 18 h after cotransfection. Results are represented as fold induction of luciferase activity compared to control cells transfected with an empty CMV expression plasmid. Histograms show the mean of a typical experiment of three performed in triplicate; bars indicate the standard deviation of the mean. (C) Representative Western blot analysis. A blot loaded with 100 μg of extract was probed with the anti-HA antibody 16B12 to detect the expression levels of HA:p53, HA:p73α, and HA:p73β in transfected cells. Ct, control plasmid.
Apoptosis assays.
H1299 cells were cotransfected in 10-cm-diameter plates with 7 μg of expression vector containing either p73α, wild-type p53, or mutant p53 and 7 μg of a green fluorescent protein (GFP)-containing internal ribosomal entry site construct (Clontech). When appropriate, 7 μg of pRC-CMV vector (Invitrogen) was included to keep the total amount of transfected DNA constant in each sample. At 72 h after transfection, cells were observed under epifluorescence (Nikon Diaphot 300) and images were photographed with an Optronics 3CCD video camera linked to a Macintosh. For each condition, three plates were used and 500 GFP-stained cells were counted in randomly selected fields from each plate. Among the GFP-stained populations, apoptotic cells were identified by the presence of apoptotic bodies or membrane blebbing.
To confirm the apoptotic phenotype, cells were also subjected to fluorescence-activated cell sorting (FACS) analysis as described elsewhere (38). Briefly, 72 h posttransfection, cells were fixed in paraformaldehyde 2% [100 mM NaCl, 10 mM piperazine-N,N′-bis(2-ethanesulfonic acid) (PIPES; pH 6.8), 300 mM sucrose, 3 mM MgCl2, 1 mM EGTA] for 20 min and then in 95% methanol for 1 h. Fixed cells were washed three times with phosphate-buffered saline and exposed to propidium iodide (PI; 60 μg/ml) and RNase A (50 μg/ml) for 30 min before counting by FACS (FACScalibure; Becton Dickinson). Harvesting of cells and FACS analysis were performed on the same day to avoid loss of GFP staining. Cells (100,000) were gated for GFP staining with a 530/20-nm bandpass filter and then analyzed for DNA content (PI) with a 610-nm longpass filter. An excitation wavelength of 488 nm was used for GFP and PI. Data were analyzed with CELLQuest software (Becton Dickinson). From the DNA content profile, the sub-G1 fraction was gated and counted.
RESULTS
p73α and p73β differentially transactivate p53-regulated promoters in mammalian cells.
To analyze the transactivation potential of the two naturally occurring p73 isoforms for p53 target genes, a p53-null cell line (H1299) was transiently cotransfected with mammalian expression plasmids encoding HA:p53, HA:p73α, or HA:p73β under the control of the CMV promoter, and reporter plasmids containing either full-length promoters (p21, mdm2, cyclin G, and Bax) or p53 binding sites (GADD45, IGF-BP3 box A, and IGF-BP3 box B) from p53 target genes placed upstream of a luciferase cDNA (see Materials and Methods and reference 19). The simian p73 isoforms that we used are 97.6% identical (98.3% similar) to their human counterparts. As previously shown (19), wild-type p53 transactivated all promoters (full length or partial) in H1299 cells to various degrees (Fig. 1A and B). In addition, as previously demonstrated (32), p73α and p73β transactivated the p21 promoter-luciferase reporter, albeit to a lesser degree than p53 (Fig. 1A). Both p73α and p73β transactivated the remaining p53-responsive promoters to various degrees (Fig. 1), with one exception (IGF-BP3 box A, [Fig. 1B]; see below). While we observed a 20- to 25-fold induction of the p21 and mdm2 promoters by wild-type p53, both p73 isoforms activated transcription of these same promoters to a lesser degree (seven- to ninefold induction [Fig. 1A]). In contrast, the Bax and GADD45 promoters were transactivated by both p73α and p73β as well as, or somewhat better than, wild-type p53 (Fig. 1A and B, respectively). It should be noted that in these cells these two promoters are transactivated by wild-type p53 to a greater degree than any other. Interestingly, we observed significant differences between p73α and p73β in the ability to transactivate the cyclin G, IGF-BP3 box A, and IGF-BP3 box B promoters (Fig. 1A and B). Whereas p73β activated the cyclin G promoter 20-fold, p73α activated the same promoter less than 8-fold. Conversely, whereas p73α activated the IGF-BP3 box B promoter 40-fold, p73β activated this same promoter less than 16-fold. Finally, whereas both p53 and p73α activated the IGF-BP3 box A promoter 16 to 17-fold, p73β only activated this same promoter less than 2-fold. It should be noted as well that under these conditions, IGF-BP3 box B appears to a better target for p73α than for wild-type p53. As detected by Western blot analysis with MAb 16B12, the protein levels of ectopically expressed wild-type p73α and p73β were readily detected (Fig. 1C).
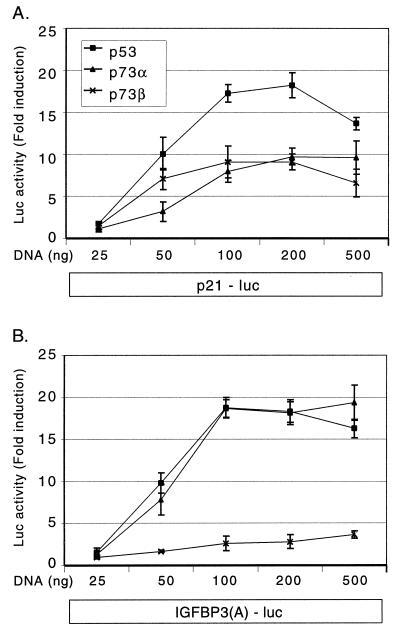
The ability of p53 to drive transcription from a reporter gene is dependent on the expression level of p53 (14, 40). The levels of p53/p73 expression constructs used for Fig. 1 were well within the range used in other such transient transfection studies (30, 32, 49, 54). However, it is frequently observed that a dose-response assay shows progressively increasing amounts of reporter activity as a function of p53-expressing constructs introduced into cells (10 to 200 ng) until a plateau is reached, while at still higher concentrations (e.g., 0.5 to 1 μg of DNA) p53 can “self-squelch” (53), leading to reduced transcriptional activation. To ensure that we were in the plateau range of activity, we examined the effects of increasing amounts (25 ng to 500 ng) of either p53, p73α, or p73β expression vectors on two distinct reporter genes. We chose the p21 promoter-luciferase reporter, which is transactivated to a lesser degree by both p73 isoforms than by p53 (Fig. 1), and the IGF-BP3 box A promoter-luciferase reporter, which is transactivated by both p53 and p73α and, very weakly, by p73β (Fig. 1). At expression vector concentrations ranging from 25 to 100 ng, p53, p73α, and p73β progressively increased the activity of the p21 promoter-luciferase reporter (Fig. 2A). At vector concentrations of 100 ng and 200 ng, p21 reporter activity plateaued. Further increasing the p53 and p73β, but not p73α, expression vector concentrations to 500 ng reduced p21 reporter transactivation, somewhat suggesting a squelching effect. The IGF-BP3 box A promoter-luciferase reporter gene, while similarly induced by p53 and p73α, was not strongly induced by p73β (Fig. 1), even at the highest concentration. Importantly, at lower DNA concentrations (50 and 100 ng), we did not observe transactivation of the IGF-BP3 box A reporter by p73β, ruling out the possibility of squelching (Fig. 2B). In contrast, the transactivation profiles of p53 and p73α were very similar, reaching a maximum at 100 ng and leveling off at 200 ng (Fig. 2). These results suggest that the different isoforms of p73 can discern between these two distinct p53-responsive reporters.
FIG. 2.
Transactivation of the p21 promoter and IGF-BP3 box A sequence by increasing amounts of p53, p73α, or p73β. Increasing amounts (0 to 500 ng) of CMV-HA:p53, CMV-HA:p73α, or CMV-HA:p73β expression plasmid along with one of the luciferase (luc) reporter constructs indicated in boxes at the bottom were transiently cotransfected into H1299 cells. DNA concentration was kept constant with an empty CMV expression vector. Cells were harvested for luciferase assay 18 h after cotransfection. Results are represented as fold induction of luciferase activity compared to control cells transfected with an empty CMV expression vector. Data points represent the mean of a typical experiment of two performed in triplicate; bars indicate the standard deviation of the mean.
These results taken together suggest that both isoforms of p73 have the ability to transactivate p53-responsive genes (albeit at possibly higher than normal protein levels) in the absence of endogenous p53. They also demonstrate significant quantitative differences in the relative transactivation abilities of these different related gene products.
p73α and p73β differentially transactivate p53-regulated promoters in yeast.
To further characterize the transactivation potential of p73α and p73β for p53 target genes, we constructed yeast strains (12) which contained the HIS3 gene under the control of one of the following derived p53-responsive human or murine target gene cis-acting elements: p21, mdm2, GADD45, cyclin G, Bax, IGF-BP3 box A, IGF-BP3 box B, RGC (34), and an artificial high-affinity-binding p53 consensus element (termed SCS) (26). Each of the above-mentioned reporter strains was transformed with a plasmid expressing either human wild-type or mutant p53 under the control of the constitutive ADH1 minimal promoter (29), which does not express extremely high levels of p53 (12). The growth assay used for our phenotypic analysis relies on the fact that the HIS3 gene is under the control of an inactive GAL1 promoter. This promoter is activated, and HIS3 is expressed only when bound by a transcriptional activator, such as p53 (or p73 [see below]), at sites placed upstream of the minimal GAL1 promoter. We scored transactivation as growth or lack thereof on histidine-deficient medium.
As previously demonstrated, we observed wild-type p53-dependent HIS3 transcription of all reporters to various degrees, with the exception of those containing the IGF-BP3 box A and box B cis-acting elements (Table 1 and reference 12). As a control, isogenic strains expressing either (i) wild-type p53 and containing a HIS3 reporter with no p53-responsive cis-acting element or (ii) vector control and any one of the p53-responsive cis-acting element reporters did not grow on histidine-deficient media (Table 1). Whereas the RGC- and Bax-containing reporter strains grew slowly on histidine-deficient media (Table 1), the p21-, SCS-, mdm2-, GADD45-, and cyclin G-containing reporter strains grew at the wild-type rate (Table 1). While IGF-BP3 box A and box B have been shown to be p53-responsive cis-acting elements in mammalian cells (Fig. 1 and references 5, 19, and 43), we detected no such activation in yeast (Table 1 and reference 12).
TABLE 1.
Activation of reporters by p53 and the p73 isoforms a
| cis-acting element | p53 | p73α | p73β |
|---|---|---|---|
| None | − | − | − |
| RGC | ++ | + | ++ |
| SCS | +++ | +++ | +++ |
| p21 | +++ | +++ | +++ |
| mdm2 | +++ | +++ | +++ |
| cyclin G | +++ | ++ | ++ |
| GADD45 | +++ | ++ | ++ |
| Bax | + | −/+ | −/+ |
| IGF-BP3 box A | − | − | − |
| IGF-BP3 box B | − | − | − |
Strains expressing wild-type or mutant p53, p73α, and p73β (on a LEU2 plasmid) and containing (on a TRP1 plasmid) one of the p53-responsive reporters (RGC, SCS, p21, mdm2, cyclin G, GADD45, Bax, IGF-BP3 box A, IGF-BP3 box B) or control reporter (None) were (i) streaked out for single colonies onto SC-minus-leucine-minus-tryptophan-minus-histidine plates and grown for ∼2 days at 37°C and (ii) replica plated from SC-minus-leucine-minus-tryptophan plates grown at 30°C to SC-minus-leucine-minus-tryptophan-minus-histidine plates and grown for 1 to 2 days at 37°C. Growth of wild-type p73-containing strains on histidine-deficient medium was scored against growth of strains expressing wild-type p53 and the relevant reporter. The LEU2/CEN (no p53, p73α, and p73β expression), p53R273H-, p73αR292H-, and p73βR292H-containing strains were inactive for transactivation with all reporters, scored −, −, no growth; −/+, slow growth; +, growth; ++, moderate growth; +++, wild-type growth.
To examine the transactivation ability of p73 in yeast, each of the above reporter strains was transformed with plasmids expressing either wild-type or mutant p73α and p73β under the control of the constitutive ADH1 minimal promoter. We observed wild-type p73-dependent HIS3 transcription of all reporters to various degrees, with the exception of those containing the IGF-BP3 box A and box B cis-acting elements (Table 1). In contrast, the p73αR292H and p73βR292H mutants, homologs of the p53R273H mutant that have been shown to be unable to transactivate an RGC- and p21-containing reporter in transient cotransfection assays (32), were similarly inactive for transactivation in our yeast-based assay. A representative example comparing the transactivation activities of wild-type and mutant p73α and -β to those of wild-type p53 in the p21:HIS3 strain is shown in Fig. 3C. As a control, isogenic strains expressing wild-type p73α and p73β and containing a HIS3 reporter with no p53-responsive cis-acting element did not grow on histidine-deficient medium (Table 1). Whereas the RGC-containing reporter strains grew slowly on histidine-deficient media (Table 1, p73α and p73β) and the Bax-containing reporter strains grew extremely slowly (Table 1), the p21-, SCS-, mdm2-, GADD45-, and cyclin G-containing reporter strains grew at rates ranging from ++ to +++ (Table 1). While p73α and p73β transactivated the p21-, mdm2-, and SCS-containing reporters as well as p53, both transactivated the cyclin G- and GADD45-containing reporters to lesser extents than p53 (Table 1). Additionally, p73β transactivated the RGC-containing reporter better than its isoform, p73α (Table 1).
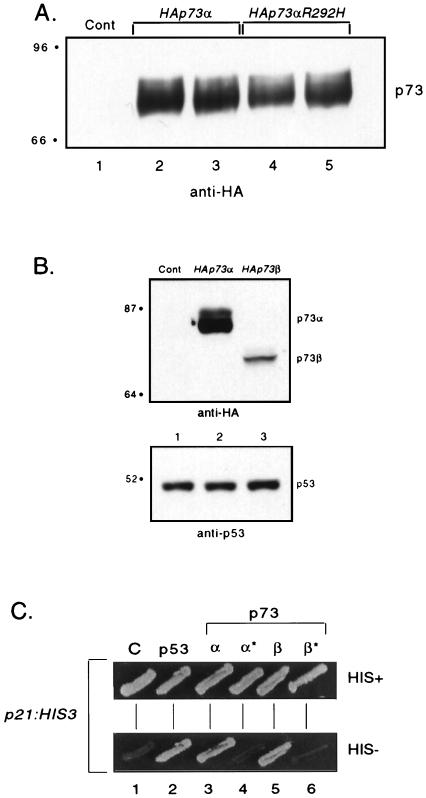
FIG. 3.
Expression levels of p73 in yeast. (A) Wild-type- and mutant p73α-expressing strains were grown to log phase; extracts were prepared and subjected to Western blot analysis in which 50 μg of total cell extract was loaded. The Western blot was probed with an anti-HA antibody at a 1/3,000 dilution. Shown is the expression from two independently isolated wild-type and mutant p73α clones. “Cont” (lane 1) refers to a strain containing a control vector (ADH1 promoter on a LEU2/2μm plasmid). (B) Wild-type p73α- and p73β-expressing strains were grown to log phase; extracts were prepared and subjected to Western blot analysis in which 100 μg of total cell extract was loaded. The Western blot was probed with an anti-HA antibody at a 1/3,000 dilution (top) and a mixture of anti-p53 antibodies at a 1/3,000 dilution (bottom). These strains also express wild-type untagged p53, which was used as a loading control. “Cont” (lane 1) refers to a strain containing a control vector (ADH1 promoter on a LEU2/2μm plasmid) and wild-type p53 (ADH1 promoter p53 on a URA3/CEN vector). (C) Strains expressing wild-type p53, p73α, or p73β, or mutant p73α or p73β, and containing a p53-responsive reporter (p21:HIS3 on a TRP/CEN plasmid) were patched onto SC-minus-leucine-minus-tryptophan plates and grown for 1 day at 37°C. Patches were replica plated onto SC-minus-leucine-minus-tryptophan-minus-histidine plates and grown for 1.5 days at 37°C. *, mutant p73αR292H or p73βR292H; C, strain containing a control vector (ADH1 promoter on a LEU2/CEN plasmid). Sizes in panels A and B are indicated in kilodaltons on the left.
As detected by Western blot analysis with MAb 12CA5, the protein levels of wild-type and mutant HA:p73 in their respective reporter yeast strains were readily detected and quite similar (Fig. 3A and data not shown). Therefore, the growth rate differences are not due to variations in the levels of wild-type and mutant p73 protein but are due to the ability of wild-type p73α and p73β to transactivate the cis-acting element present in each reporter. Interestingly, when we examined the expression levels of wild-type p73α and wild-type p73β by Western blot analysis and serial dilution of extracts from isogenic strains, we consistently observed ∼25-fold less p73β than p73α (Fig. 3B and data not shown). We also observed slower-migrating species for p73α (and p73β under certain gel conditions) which were not seen for p53 (Fig. 3A and B and data not shown). Despite being present at a lower concentration than p73α, p73β can activate transcription comparably to p73α. We do not know whether this represents a qualitative difference in the p73 gene products or the presence of both at levels greater than that required for transactivation in this yeast-based assay.
p73α coprecipitates with tumor-derived p53, but not wild-type p53, in mammalian cells.
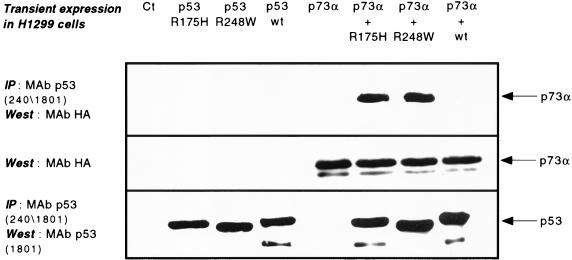
Utilizing the yeast two-hybrid system, Kaghad et al. demonstrated both homotypic and heterotypic interactions between p53 and p73β but not p73α (33). To test whether p73 associates with p53 in mammalian cells, we performed coimmunoprecipitation experiments from extracts of human H1299 cells ectopically expressing either wild-type or mutant forms of p53 and p73. Wild-type and mutant p53 were immunoprecipitated with a mixture of anti-p53 MAbs 240 and 1801, and HA:p73α was detected on Western blots of the immunoprecipitates by use of the anti-HA antibody 16B12. Surprisingly, p73α coprecipitated both with p53R248W and p53R175H but did not detectably coprecipitate with wild-type p53 (Fig. 4). By Western blot analysis of extracts, we estimated that 5 to 10% of p73α is stably associated with mutant p53 under these transient overexpression conditions. In contrast, despite repeated attempts using a variety of extraction and immunoprecipitation conditions, we were unable to detect protein-protein interactions between either wild-type or mutant p53 and either wild-type p73α or p73β by overexpression in yeast (data not shown).
FIG. 4.
Association of p73α with tumor-derived p53 mutants but not wild-type p53. H1299 cells were transfected with an expression plasmid for either wild-type p53 (p53 wt), p53R175H, p53R248W, or p73α, alone and in combination as indicated. p53 was precipitated from extracts (1.5 mg of protein) incubated with 100 ng each of anti-p53 MAbs 240 and 1801. Western blots of the immunoprecipitates (IP) were probed with either the anti-HA antibody 16B12 (upper panel) or the anti-p53 antibody 1801 (lower panel) (see Materials and Methods). The middle panel is a Western blot loaded with 100 μg of extract and probed with the anti-HA antibody to detect p73α.
Tumor-derived p53 mutants reduce p73α transcriptional activity in mammalian cells.
That we detected interactions between tumor-derived p53 mutants and p73 suggests that p53 mutants may affect p73 function. To test this, we cotransfected the p53-null cell line H1299 with p73α along with two tumor-derived p53 mutants and a reporter construct containing a p53-responsive human target gene cis-acting element derived either from the p21 promoter (Fig. 5B; see Materials and Methods) or from the Bax promoter (Fig. 5E). We confirmed that (i) wild-type p53 and p73α are transcriptionally active toward both reporters, while the two tumor-derived p53 mutants, R175H and R248W, are inactive, and (ii) that both p53 mutants reduced the transcriptional activity of wild-type p53 (Fig. 5A and E), as previously described (13, 17, 35); reviewed in reference 69). Importantly, we found that these same tumor-derived p53 mutants markedly reduced the transcriptional activity of p73α (Fig. 5A and E). To normalize for the reporter DNA utilized in each transfection, an additional reporter containing a CMV promoter upstream of the renilla luciferase cDNA (Promega) was employed. Under these experimental conditions, the ectopically expressed wild-type HA:p73α, as detected by Western blot analysis with MAb 12CA5, was readily detected and was not significantly affected by either wild-type or mutant p53 proteins (Fig. 5C). It is worth mentioning that in these experiments, we used a DNA transfection ratio of 1:1 for the various p53- and p73α-expressing vectors. Previous work has shown that the ratio of mutant to wild-type p53 DNA used for transfection determines the level of transcription measured (35, 68): more mutant p53 DNA transfected per wild-type p53 DNA leads to greater inhibition of transactivation (Fig. 5D). To further examine this dosage-dependent abrogation of transactivation with regard to p73α, increasing amounts of the expression plasmid p53R248W were cotransfected with constant amounts of either wild-type p53 or the HA:p73α expression plasmid. We found that a fivefold excess of transfected mutant p53 DNA decreased the transcriptional activity of p53 by a factor of 10 and that of p73α by a factor of 3 (Fig. 5D). Increasing the ratio to 10-fold did not further reduce activation in either case, although with p53 the level of repression approached that observed with the control vector. Thus, even this level of mutant p53 is not sufficient to completely abolish all transcriptional activity associated with p73α.
Expression of p53 mutants inhibits the ability of p73α to induce apoptosis.
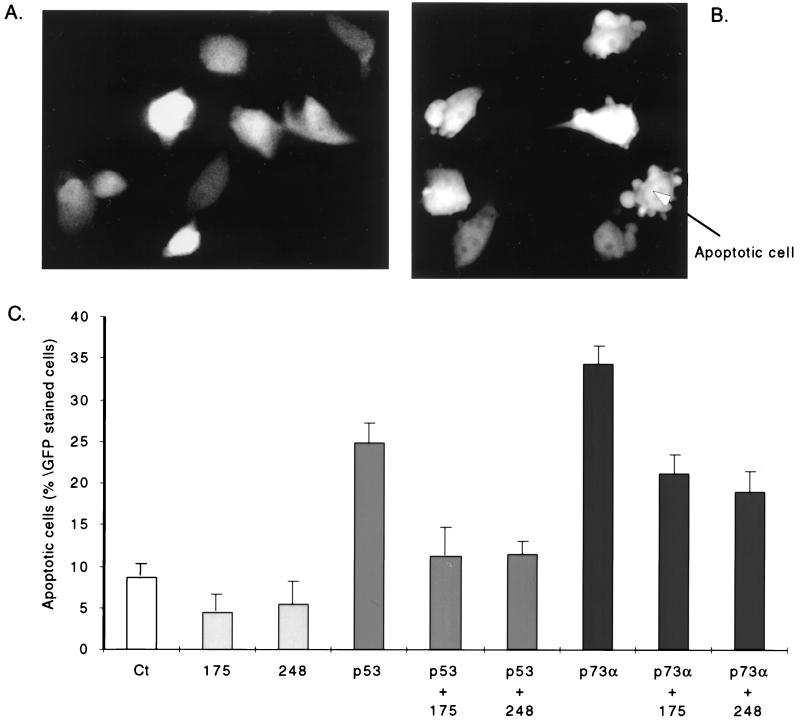
Both isoforms of p73, analogous to p53, can induce apoptosis (32). It was of interest to determine if the reduction in p73 transactivation activity by p53 mutants can be correlated with diminution of apoptosis as a result of overexpression of p73. To test this, we transfected H1299 cells with wild-type p73α, wild-type p53, and mutant p53 variants either alone or in combination along with a GFP-containing vector and analyzed the morphology of transfected cells. An example of the morphological changes induced in cells expressing p73α is shown in Fig. 6B, and an example of cells expressing a control plasmid is shown in Fig. 6A. As seen in Fig. 6B, p73 overexpression induced the apoptotic bodies and membrane blebbing characteristic of apoptotic cells (28). By enumerating such morphologically distinguishable cells as a function of the total GFP-expressing cells, we could quantitate the effects of the various forms of p53 and p73 separately and together (Fig. 6C). As expected, wild-type p53, but not the tumor-derived p53 mutants, induced apoptosis after transient overexpression (Fig. 6C). As previously demonstrated for other cell types (BHK and SaOs2 [32]), ectopically expressed p73α induced apoptosis in H1299 cells (Fig. 6B [representative profile] and C). It should be noted that the ability of p73α to induce apoptosis in H1299 cells is slightly better than that of wild-type p53. Moreover, coexpression of both tumor-derived p53 mutants with either wild-type p53 or p73α resulted in a reduction of apoptotic cells to a level approximately 50 and 44%, respectively, of that seen with either wild-type protein alone (Fig. 6C). Since the p53 mutants also caused a modest reduction in the number of apoptotic cells compared to the control vector, we cannot rule out the possibility that they have an additional counter apoptotic function. Nevertheless, our results show a good correlation between the ability of tumor-derived p53 mutants to inhibit the transactivation and proapoptotic functions of both wild-type p53 and of a closely related family member, p73α.
FIG. 6.
Tumor-derived p53 mutants reduce p73α-induced apoptosis. H1299 cells were transiently cotransfected with wild-type p53, p53R175H, p53R248W, and p73α expression plasmids, singly or in combination as indicated, and a GFP expression plasmid. After 72 h, the morphology of the cells was observed under an epifluorescence microscope. (B) Representative apoptotic morphology of cells overexpressing p73α compared to control cells in panel A transfected with an empty CMV expression plasmid. (C) Results represented as percentage of apoptotic cells over 500 GFP-stained cells in a 10-cm-diameter plate (mean of a typical experiment performed with triplicates cultures; bars indicate standard deviations).
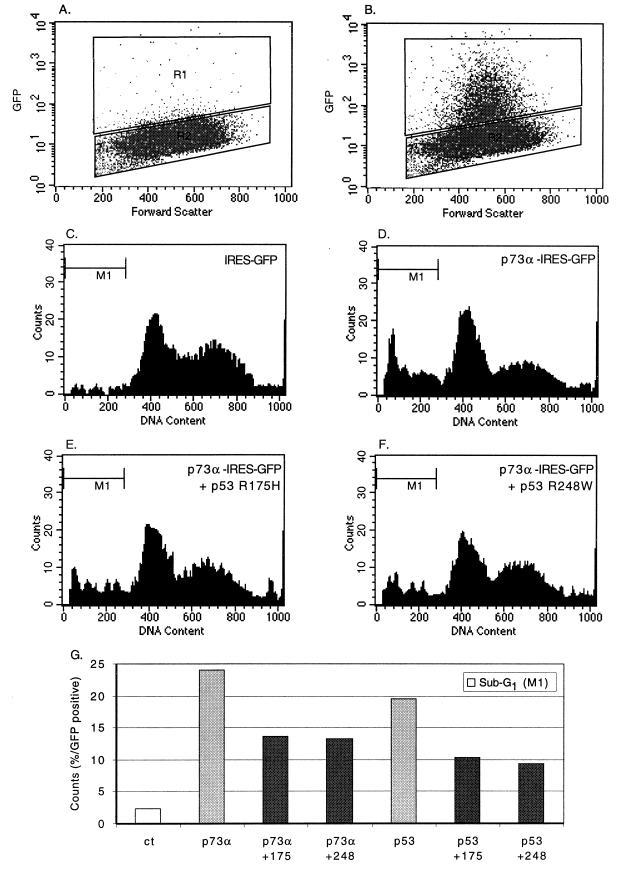
To confirm the reduction of apoptosis induced by the two p53 tumor-derived mutants, we used FACS analysis to quantify the amount of the sub-G1 fraction, which represents apoptotic cells. H1299 cells cotransfected with a p53 or p73α expression vector and a GFP expression vector as described in the legend to Fig. 6 and in Materials and Methods were fixed 72 h posttransfection, stained by PI, and analyzed by FACS. First, GFP-stained cells were gated by comparing cells transfected with an empty vector (Fig. 7A) to cells transfected with the GFP-expressing vector (Fig. 7B); then the GFP-positive cells were analyzed for DNA content to quantify the sub-G1 fraction when only GFP is expressed (Fig. 7C), when p73α is expressed alone (Fig. 7D), or when p73α is coexpressed with either p53R175H (Fig. 7E) or p53R248W (Fig. 7F). The results of the FACS analysis are summarized in Fig. 7G. Overexpression of p73α alone in H1299 cells induced a level of apoptotic cells (sub-G1 fraction) nearly the same as that induced by p53 alone. Importantly, both p53 tumor-derived mutants reduced the sub-G1 fraction (apoptotic cells) induced by both p53 and p73α. Thus, using two different assays, we demonstrated that expression of p53 mutants can lead to reduced apoptosis induced by p73α.
FIG. 7.
Tumor-derived p53 mutants reduce p73α-induced sub-G1 populations. H1299 cells were transiently cotransfected with a wild-type p53, p53R175H, p53R248W, or p73α expression plasmid singly or in combination as indicated, and a GFP expression plasmid. After 72 h, cells were fixed and analyzed by FACS analysis. (A and B) Representative gating of control cells and cells expressing GFP, respectively. Populations from gate R1 were analyzed for DNA content (C to F) and the sub-G1 population counted by the gate indicated M1. (G) Summary of results. Each bar represents the percentage of sub-G1 cells (gate M1) over the total number of GFP-stained cells.
DISCUSSION
In this study, we provide evidence that (i) p73α and p73β have the ability to transactivate several distinct p53-responsive reporters to various degrees, in both mammalian and yeast cells; (ii) two representative tumor-derived p53 mutants coimmunoprecipitate with p73 in vivo; and (iii) expression of these same tumor-derived p53 mutants can reduce the transcriptional and the proapoptotic activity of p73.
Both p73 isoforms differentially transactivate p53 targets.
We found that in transient cotransfection assays, p73α and p73β are both capable of inducing many p53 target gene promoters in mammalian cells and p53 binding sites in yeast cells but to variable extents. In mammalian cells, the GADD45 and Bax promoters are stimulated to the same extent by all three proteins, but p53 is a more potent transactivator of p21 and mdm2 promoters than p73. The two p73 isoforms differ in that while p73α is more active than p73β toward the A box of the IGF-BP3 promoter, p73β is more active toward the cyclin G promoter. An additional confirmation of the transcriptional activity associated with p73α and p73β was performed with a rapid and qualitative yeast-based assay. While both p73 isoforms are able to transactivate many of the p53-responsive reporters tested, we found lesser differences in the relative abilities of both isoforms to transactivate. Interestingly, in yeast p73α is expressed at higher levels than p73β. Whether this is due to stabilization or modification of p73α or enhanced degradation of p73β is not clear. What is clear is that this lower level of p73β than of p73α and p53 is sufficient to activate transcription as well as p53 (for p21, SCS, and mdm2) and better than p73α (for RGC).
While results for mammalian and yeast cells do not absolutely correlate with regard to p73’s transactivation ability, the yeast-based assay has proven useful for screening patients (29), cell lines, blood, and tumors (16) for somatic or germ line mutations in p53 as well as for monitoring adenovirus p53 transduction efficiency during gene therapy (64). Moreover, this approach allows one to distinguish among functionally silent mutations, inactivating mutations, and polymorphisms, which is not easily done by more widely used methods such as immunohistochemistry, single-stranded conformation polymorphism, anti-p53 antibody screening, and denaturing gradient gel electrophoresis. Our demonstration that wild-type, but not mutant, p73α and p73β are transcriptionally active in yeast should provide the ability to similarly assess the status of p73 (or p51/p63) in cell lines and in patient tumors.
Tumor-derived p53 mutants inhibit p73 function.
We have demonstrated that tumor-derived p53 mutants can be coimmunoprecipitated with other p53-related family members, specifically p73α, and that this interaction can lead to inhibition of p73α’s function. It is known that p53 missense mutations are expressed frequently at high levels in a wide variety of human tumors. Moreover, studies have shown that certain p53 mutants can abrogate wild-type p53 function in a dominant negative manner (13, 25, 35, 45, 47, 68). This dominant negative effect can also vary depending on the particular germ line mutation of p53 examined and the tumorigenic cell line assayed (17, 63). Overexpression of mutant p53 can also result in induction of a number of cellular genes, such as MDR-1, PCNA, VEGF, EGFR, IL-6, HSP70, and BFGF (8, 11, 36, 42, 44, 65, 66). More recently it has been demonstrated that certain tumor-derived p53 mutants, in addition to inhibiting wild-type p53 tumor suppressor activity, possess an activity that leads to the induction of endogenous cellular targets that promote tumorigenicity, such as c-myc (18), as well as promote genetic instability by disrupting spindle checkpoint control (23). Our observation that overexpression of tumor-derived p53 mutants inhibits transactivation by p73α and p73α-induced apoptosis suggests another possible mechanism by which tumor-derived p53 mutants may act, i.e., by physical association with related family members. Our data also lend support to the contention that tumor-derived p53 mutants may play an active role in cellular transformation, not only by activating genes involved in tumorigenicity, such as c-myc, but also by inactivating proapoptotic proteins such as p73. While the cellular function(s) of p73 and p51/p63 awaits further investigation, it is tempting to speculate that in cells which express both tumor-derived mutant p53 and p63 or p73 isoforms, the function of the latter may be abrogated or altered. Indeed, Prabhu et al. (52) found that p73α, but not p73β, lacks growth suppression activity in SW480 cells, a colon cancer cell line harboring mutant p53.
How do p53 and p73 proteins interact?
Wild-type p53 exists predominantly as tetramers in solution (20), and the structure of the tetramerization domain has been solved (10, 31, 39). Additionally, wild-type and tumor-derived mutant forms of p53 can readily form heterotetramers when cotranslationally expressed (2, 47, 48, 61, 63). Such wild-type/mutant p53 cotetramers are frequently inactive for DNA binding and transactivation (reviewed in reference 69). While it has not yet been determined whether wild-type p73 exists as tetramers, our observation that certain p53 mutants can associate with and inhibit p73 function suggests that cotetramers may exist in vivo. Alternatively, the association of mutant p53 with p73α may involve interactions distinct from those engendered by their oligomerization domains or even may be mediated by as yet unidentified cellular proteins. However, why were we not able to detect wild-type p73 complexed with wild-type p53 by coimmunoprecipitation? The crystal structure of the DNA binding domain of p53 bound to its cognate site (9) has helped to explain how mutations in this region interfere with DNA binding. It is plausible that these same mutations affect the overall structure of p53, including the transactivation and tetramerization domains, and not just the DNA binding domain. Perhaps only in this altered configuration does association by tumor-derived p53 mutants with p73α occur. It is noteworthy that a subset of tumor-derived mutations when fused to the GAL4 DNA binding domain are inert (46, 56, 67), suggesting propagation of the mutation within the p53 DNA binding domain to either the p53 activation region or the GAL4 DNA binding domain (or both). Clearly, determination of the domain(s) required for association and demonstration that other tumor-derived p53 mutants bind to p73 are required to better understand the nature of this potentially important interaction.
The experiments investigating p73 interactions shown in Figs. 4 to 7 were performed exclusively with p73α. Similar experiments with p73β are in progress; interestingly, we have preliminary indication that this isoform differs from p73α in that it can interact with both wild-type and mutant forms of p53 (data not shown). We are currently determining (i) which regions of the p53 and p73 proteins are required for their interactions and (ii) why p73α and p73β differ from each other.
There are a number of plausible explanations for the lack of detectable association between either wild-type or mutant p53 and p73 in yeast. If the interaction that we detect in mammalian cells requires an as yet unidentified adaptor protein, we would not see coimmunoprecipitation of both proteins if yeast cells lack such a factor. Suggestive of this are the results of Kaghad et al. (33), who detected no interaction between wild-type p53 and p73α in the yeast two-hybrid assay. Additionally, p73α and p73β may be posttranslationally modified in a yeast-specific manner, and it is this modification that may preclude any association with p53, wild type or mutant. Moreover, if an adaptor protein is required, this modification might also preclude association between p53 and p73 if the adaptor were modified. These possibilities, or others, remain to be tested.
Do p53 and p73 have overlapping functions?
The discovery that additional p53-related proteins exist requires a reexamination of the functions that we had previously attributed to p53. Not only do p73 and p51/p63 show structural similarities to p53, but the results of Jost et al. (32), Kaghad et al. (33), Osada et al. (50), and Yang et al. (72), as well as those presented here, demonstrate that p73 and p51/p63 show functional similarities to p53. That p73 and p51/p63 have activities overlapping those of p53 suggests that these proteins may be functionally interchangeable. However, the fact that p53−/− mice develop tumors argues against this assumption. Our results have demonstrated that both isoforms of p73, when overexpressed, can transactivate those target genes once thought of as being p53 specific, and they do so in a p53-independent context. What we have not shown is that these same target genes are transactivated by physiologically relevant levels of p73 in vivo. However, given the previous demonstration by Jost et al. (32) that p73 can indeed transactivate endogenous p21, it is possible that most, if not all, of these additional p53-responsive promoters can also be targets of p73 under the appropriate conditions in vivo. It is also of interest to examine whether in cells expressing both p53 and p73, the two factors compete or collaborate to activate the above-mentioned targets. Moreover, since p73α and p73β are not induced by DNA damage, it will be of interest to determine how p73 expression is regulated in cell types where it is found and what role(s) the two proteins play in p53-mediated growth arrest and/or apoptosis. Finally, we cannot exclude that one or both isoforms of p73 hetero-oligomerize with wild-type p53 under certain as yet unidentified physiological conditions and modulate some or all of their target genes. Elucidation of these questions, as well as the specific signals which regulate p73, will be necessary to uncover p73’s role in the cell. As more information about the normal expression of p73 is obtained, these and other questions can be approached.
ACKNOWLEDGMENTS
We gratefully thank W. Kaelin, Jr., for the simian p73α and p73β mammalian expression plasmids. We also thank R. Iggo for the human p53 yeast expression plasmids, the ΔUAS pGAL1:HIS3 construct, and RGC:HIS3 reporter. Critical comments and discussion on the manuscript were made by J. Ahn, G. Bond, and K. Okamoto.
C.J.D. is supported by the Cancer Research Fund of the Damon Runyon-Walter Winchell Foundation (fellowship DRG-1427). C.G. is supported by Human Frontier Foundation long-term fellowship LT0776/1997-M. This work was supported by grant {DAMD17-94}4275 from the U.S. Army.
The first two authors contributed equally to this work.
REFERENCES
- 1.Bargonetti J, Manfredi J J, Chen X, Marshak D R, Prives C. A proteolytic fragment from the central region of p53 has marked sequence-specific DNA-binding activity when generated from wild-type but not from oncogenic mutant p53 protein. Genes Dev. 1993;7:2565–2574. doi: 10.1101/gad.7.12b.2565. [DOI] [PubMed] [Google Scholar]
- 2.Bargonetti J, Reynisdottir I, Friedman P N, Prives C. Site-specific binding of wild-type p53 to cellular DNA is inhibited by SV40 T antigen and mutant p53. Genes Dev. 1992;6:1886–1898. doi: 10.1101/gad.6.10.1886. [DOI] [PubMed] [Google Scholar]
- 3.Bayle J H, Elenbaas B, Levine A J. The carboxyl-terminal domain of the p53 protein regulates sequence-specific DNA binding through its nonspecific nucleic acid-binding activity. Proc Natl Acad Sci USA. 1995;92:5729–5733. doi: 10.1073/pnas.92.12.5729. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Brain R, Jenkins J R. Human p53 directs DNA strand reassociation and is photolabelled by 8-azido ATP. Oncogene. 1994;9:1775–1780. [PubMed] [Google Scholar]
- 5.Buckbinder L, Talbott R, Velasco-Miguel S, Takenaka I, Faha B, Seizinger B R, Kley N. Induction of the growth inhibitor IGF-binding protein 3 by p53. Nature. 1995;377:646–649. doi: 10.1038/377646a0. [DOI] [PubMed] [Google Scholar]
- 6.Candau R, Scolnick D M, Darpino P, Ying C Y, Halazonetis T D, Berger S L. Two tandem and independent sub-activation domains in the amino terminus of p53 require the adaptor complex for activity. Oncogene. 1997;15:807–816. doi: 10.1038/sj.onc.1201244. [DOI] [PubMed] [Google Scholar]
- 7.Chang J, Kim D-H, Lee S W, Choi K Y, Sung Y C. Transactivation ability of p53 transcriptional activation domain is directly related to the binding affinity to TATA-binding protein. J Biol Chem. 1995;270:25014–25019. doi: 10.1074/jbc.270.42.25014. [DOI] [PubMed] [Google Scholar]
- 8.Chin K-V, Ueda K, Pastan I, Gottesman M M. Modulation of activity of the promoter of the human MDR1 gene by ras and p53. Science. 1992;255:459–462. doi: 10.1126/science.1346476. [DOI] [PubMed] [Google Scholar]
- 9.Cho Y, Gorina S, Jeffrey P D, Pavletich N P. Crystal structure of a p53 tumor suppressor-DNA complex: understanding tumorigenic mutations. Science. 1994;265:346–355. doi: 10.1126/science.8023157. [DOI] [PubMed] [Google Scholar]
- 10.Clore G M, Omichinski J G, Sakaguchi K, Zambrano N, Sakamoto H, Appella E, Gronenborn A M. High-resolution structure of the oligomerization domain of p53 by multidimensional NMR. Science. 1994;265:386–394. doi: 10.1126/science.8023159. [DOI] [PubMed] [Google Scholar]
- 11.Deb S, Jackson C T, Subler M A, Martin D W. Modulation of cellular and viral promoters by mutant human p53 proteins found in tumor cells. J Virol. 1992;66:6164–6170. doi: 10.1128/jvi.66.10.6164-6170.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Di Como C J, Prives C. Human tumor-derived p53 proteins exhibit binding site selectivity and temperature sensitivity for transactivation in a yeast-based assay. Oncogene. 1998;16:2627–2539. doi: 10.1038/sj.onc.1202041. [DOI] [PubMed] [Google Scholar]
- 13.Farmer G, Bargonetti J, Zhu H, Friedman P, Prywes R, Prives C. Wild-type p53 activates transcription in vitro. Nature. 1992;358:83–86. doi: 10.1038/358083a0. [DOI] [PubMed] [Google Scholar]
- 14.Farmer G, Friedlander P, Colgan J, Manley J L, Prives C. Transcriptional repression by p53 involves molecular interactions distinct from those with the TATA box binding protein. Nucleic Acids Res. 1996;24:4281–4288. doi: 10.1093/nar/24.21.4281. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Fields S, Jang S K. Presence of a potent transcription activating sequence in the p53 protein. Science. 1990;249:1046–1049. doi: 10.1126/science.2144363. [DOI] [PubMed] [Google Scholar]
- 16.Flaman J-M, Frebourg T, Moreau V, Charbonnier F, Martin C, Chappuis P, Sappino A-P, Limacher J-L, Bron L, Benhattar J, Tada M, Van Meir E G, Estreicher A, Iggo R D. A simple p53 functional assay for screening cell lines, blood, and tumors. Proc Natl Acad Sci USA. 1995;92:3963–3967. doi: 10.1073/pnas.92.9.3963. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Forrester K, Lupold S E, Ott V L, Chay C H, Band V, Wang X W, Harris C C. Effects of p53 mutants on wild-type p53-mediated transactivation are cell type dependent. Oncogene. 1995;10:2103–2111. [PubMed] [Google Scholar]
- 18.Frazier M W, He X, Wang J, Gu Z, Cleveland J L, Zambetti G P. Activation of c-myc gene expression by tumor-derived p53 mutants requires a discrete C-terminal domain. Mol Cell Biol. 1998;18:3735–3743. doi: 10.1128/mcb.18.7.3735. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Friedlander P, Haupt Y, Prives C, Oren M. A mutant p53 that discriminates between p53-responsive genes cannot induce apoptosis. Mol Cell Biol. 1996;16:4961–4971. doi: 10.1128/mcb.16.9.4961. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Friedman P N, Chen X, Bargonetti J, Prives C. The p53 protein is an unusually shaped tetramer that binds directly to DNA. Proc Natl Acad Sci USA. 1993;90:3319–3323. doi: 10.1073/pnas.90.8.3319. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Gaiddon C B, Monnier A-L, Mercken L, Loeffler J-P. Genomic effects of the putative oncogene G alpha s. Chronic transcriptional activation of the c-fos proto-oncogene in endocrine cells. J Biol Chem. 1994;269:22663–22671. [PubMed] [Google Scholar]
- 22.Gottlieb T, Oren M. p53 in growth control and neoplasia. Biochim Biophys Acta. 1996;1287:77–102. doi: 10.1016/0304-419x(95)00019-c. [DOI] [PubMed] [Google Scholar]
- 23.Gualberto A, Aldape K, Kozakiewicz K, Tlsty T. An oncogenic form of p53 confers a dominant, gain-of-function phenotype that disrupts spindle checkpoint control. Proc Natl Acad Sci USA. 1998;95:5166–5171. doi: 10.1073/pnas.95.9.5166. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Guthrie J A, Fink G R, editors. Methods in enzymology. Vol. 194. San Diego, Calif: Academic Press, Inc.; 1991. [Google Scholar]
- 25.Hachiya M, Chumakov A, Miller C W, Akashi M, Said J, Koeffler H P. Mutant p53 proteins behave in a dominant, negative fashion in vivo. Anticancer Res. 1994;14:1853–1860. [PubMed] [Google Scholar]
- 26.Halazonetis T D, Davis L J, Kandil A N. Wild-type p53 adopts a ‘mutant’-like conformation when bound to DNA. EMBO J. 1993;12:1021–1028. doi: 10.1002/j.1460-2075.1993.tb05743.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Halazonetis T D, Kandil A N. Conformational shifts propagate from the oligomerization domain of p53 to its tetrameric DNA binding domain and restore DNA binding to select p53 mutants. EMBO J. 1993;12:5057–5064. doi: 10.1002/j.1460-2075.1993.tb06199.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Hale A J, Smith C A, Sutherland L C, Stoneman V E A, Longthorne V L, Culhane A C, Williams G T. Apoptosis: molecular regulation of cell death. Eur J Biochem. 1996;236:1–26. doi: 10.1111/j.1432-1033.1996.00001.x. [DOI] [PubMed] [Google Scholar]
- 29.Ishioka C, Frebourg T, Yan Y-X, Vidal M, Friend S H, Schmidt S, Iggo R. Screening patients for heterozygous p53 mutations using a functional assay in yeast. Nat Genet. 1993;5:124–129. doi: 10.1038/ng1093-124. [DOI] [PubMed] [Google Scholar]
- 30.Jayaraman L, Murthy K G K, Zhu C, Curran T, Xanthoudakis S, Prives C. Identification of redox/repair protein Ref-1 as a potent activator of p53. Genes Dev. 1997;11:558–570. doi: 10.1101/gad.11.5.558. [DOI] [PubMed] [Google Scholar]
- 31.Jeffrey P D, Gorina S, Pavletich N P. Crystal structure of the tetramerization domain of the p53 tumor suppressor at 1.7 angstroms. Science. 1995;267:1498–1502. doi: 10.1126/science.7878469. [DOI] [PubMed] [Google Scholar]
- 32.Jost C A, Marin M C, Kaelin W G., Jr p73 is a human p53-related protein that can induce apoptosis. Nature. 1997;389:191–194. doi: 10.1038/38298. [DOI] [PubMed] [Google Scholar]
- 33.Kaghad M, Bonnet H, yang A, Creancier L, Biscan J-C, Valent A, Minty A, Chalon P, Lelias J-M, Dumont X, Ferrara P, McKeon F, Caput D. Monoallelically expressed gene related to p53 at 1p36, a region frequently deleted in neuroblastoma and other human cancers. Cell. 1997;90:809–819. doi: 10.1016/s0092-8674(00)80540-1. [DOI] [PubMed] [Google Scholar]
- 34.Kern S E, Kinzler K W, Bruskin A, Jarosz D, Friedman P, Prives C, Vogelstein B. Identification of p53 as a sequence-specific DNA-binding protein. Science. 1991;252:1708–1711. doi: 10.1126/science.2047879. [DOI] [PubMed] [Google Scholar]
- 35.Kern S E, Pientenpol J A, Thiagalingam S, Seymour A, Kinzler K W, Vogelstein B. Oncogenic forms of p53 inhibit p53-regulated gene expression. Science. 1992;256:827–830. doi: 10.1126/science.1589764. [DOI] [PubMed] [Google Scholar]
- 36.Kieser A, Weich H A, Brandner G, Marme D, Kolch W. Mutant p53 potentiates protein kinase C induction of vascular endothelial growth factor expression. Oncogene. 1994;9:963–969. [PubMed] [Google Scholar]
- 37.Ko L J, Prives C. p53: puzzle and paradigm. Genes Dev. 1996;10:1054–1072. doi: 10.1101/gad.10.9.1054. [DOI] [PubMed] [Google Scholar]
- 38.Lamm G M, Steinlein P, Cotten M, Christofori G. A rapid, quantitative and inexpensive method for detecting apoptosis by flow cytometry in transiently transfected cells. Nucleic Acids Res. 1997;25:4855–4857. doi: 10.1093/nar/25.23.4855. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Lee W, Harvey T S, Yin Y, Yau P, Litchfield D, Arrowsmith C H. Solution structure of the tetrameric minimum transforming domain of p53. Nat Struct Biol. 1994;1:877–890. doi: 10.1038/nsb1294-877. [DOI] [PubMed] [Google Scholar]
- 40.Lee Y, Chen Y, Chang L S, Johnson L F. Inhibition of mouse thymidylate synthase promoter activity by the wild-type p53 tumor suppressor protein. Exp Cell Res. 1997;234:270–276. doi: 10.1006/excr.1997.3605. [DOI] [PubMed] [Google Scholar]
- 41.Levine A J. p53, the cellular gatekeeper for growth and division. Cell. 1997;88:323–331. doi: 10.1016/s0092-8674(00)81871-1. [DOI] [PubMed] [Google Scholar]
- 42.Ludes-Meyers J H, Subler M A, Shivakumar C V, Munoz R M, Jiang P, Bigger J E, Brown D R, Deb S P, Deb S. Transcriptional activation of the human epidermal growth factor receptor promoter by human p53. Mol Cell Biol. 1996;16:6009–6019. doi: 10.1128/mcb.16.11.6009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Ludwig R L, Bates S, Vousden K H. Differential activation of target cellular promoters by p53 mutants with impaired apoptotic function. Mol Cell Biol. 1996;16:4952–4960. doi: 10.1128/mcb.16.9.4952. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Margulies L, Sehgal P B. Modulation of the human interleukin-6 promoter (IL-6) and transcription factor C/EBP beta (NF-IL6) activity by p53 species. J Biol Chem. 1993;268:15096–15100. [PubMed] [Google Scholar]
- 45.Michalovitz D, Halevy O, Oren M. Conditional inhibition of transformation and of cell proliferation by a temperature-sensitive mutant of p53. Cell. 1990;62:671–680. doi: 10.1016/0092-8674(90)90113-s. [DOI] [PubMed] [Google Scholar]
- 46.Miller C W, Chumakov A, Said J, Chen D L, Aslo A, Koeffler H P. Mutant p53 proteins have diverse intracellular abilities to oligomerize and activate transcription. Oncogene. 1993;8:1815–1824. [PubMed] [Google Scholar]
- 47.Milner J, Medcalf E A. Cotranslation of activated mutant p53 with wild type drives the wild-type p53 protein into the mutant conformation. Cell. 1991;65:765–774. doi: 10.1016/0092-8674(91)90384-b. [DOI] [PubMed] [Google Scholar]
- 48.Milner J, Medcalf E A, Cook A C. Tumor suppressor p53: analysis of wild-type and mutant p53 complexes. Mol Cell Biol. 1991;11:12–19. doi: 10.1128/mcb.11.1.12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Miyashita T, Reed J C. Tumor suppressor p53 is a direct transcriptional activator of the human bax gene. Cell. 1995;80:293–299. doi: 10.1016/0092-8674(95)90412-3. [DOI] [PubMed] [Google Scholar]
- 50.Osada M, Ohba M, Kawahara C, Ishioka C, Kanamaru R, Katoh I, Ikawa Y, Nimura Y, Nakagawara A, Obinata M, Ikawa S. Cloning and functional analysis of human p51, which structurally and functionally resembles p53. Nat Med. 1998;4:839–843. doi: 10.1038/nm0798-839. [DOI] [PubMed] [Google Scholar]
- 51.Pavletich N P, Chambers K A, Pabo C O. The DNA-binding domain of p53 contains the four conserved regions and the major mutation hot spots. Genes Dev. 1993;7:2556–2564. doi: 10.1101/gad.7.12b.2556. [DOI] [PubMed] [Google Scholar]
- 52.Prabhu N S, Somasundaram K, Satyamoorthy K, Herlyn M, El-Deiry W. P73β, unlike p53, suppresses growth and induces apoptosis of human papillomavirus E6-expressing cancer cells. Int J Oncol. 1998;13:5–9. doi: 10.3892/ijo.13.1.5. [DOI] [PubMed] [Google Scholar]
- 53.Ptashne M, Gann A A F. Activators and targets. Nature. 1990;346:329–331. doi: 10.1038/346329a0. [DOI] [PubMed] [Google Scholar]
- 54.Rainwater R, Parks D, Anderson M E, Tegtmeyer P, Mann K. Role of cysteine residues in regulation of p53 function. Mol Cell Biol. 1995;15:3892–3903. doi: 10.1128/mcb.15.7.3892. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Raycroft L, Wu H, Lozano G. Transcriptional activation by wild-type p53 but not transforming mutants of the p53 anti-oncogene. Science. 1990;249:1049–1051. doi: 10.1126/science.2144364. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Raycroft L, Schmidt J R, Yoas K, Hao M, Lozano G. Analysis of p53 mutants for transcriptional activity. Mol Cell Biol. 1991;11:6067–6074. doi: 10.1128/mcb.11.12.6067. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Reed M, Woelker B, Wang P, Wang Y, Anderson M E, Tegtmeyer P. The C-terminal domain of p53 recognizes DNA damaged by ionizing radiation. Proc Natl Acad Sci USA. 1995;92:9455–9459. doi: 10.1073/pnas.92.21.9455. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Rose M D, Winston F, Heiter P. Laboratory course manual for methods in yeast genetics. Cold Spring Harbor, N.Y: Cold Spring Harbor Laboratory; 1990. [Google Scholar]
- 59.Schmale H, Bamberger C. A novel protein with strong homology to the tumor suppressor p53. Oncogene. 1997;15:1363–1367. doi: 10.1038/sj.onc.1201500. [DOI] [PubMed] [Google Scholar]
- 60.Senoo M, Seki N, Ohira M, Sugano S, Watanabe M, Tachibana M, Tanaka T, Shinkai Y, Kato H. A second p53-related protein, p73L, with high homology to p73. Biochem Biophys Res Commun. 1998;248:603–607. doi: 10.1006/bbrc.1998.9013. [DOI] [PubMed] [Google Scholar]
- 61.Shaulian E, Zauberman A, Ginsberg D, Oren M. Identification of a minimal transforming domain of p53: negative dominance through abrogation of sequence-specific DNA binding. Mol Cell Biol. 1992;12:5581–5592. doi: 10.1128/mcb.12.12.5581. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Silhavy T J, Berman M L, Enquist L W. Experiments with gene fusions. Cold Spring Harbor, N.Y: Cold Spring Harbor Laboratory Press; 1984. [Google Scholar]
- 63.Srivastava S, Wang S, Tong Y A, Pirollo K, Chang E H. Several mutant p53 proteins detected in cancer-prone families with Li-Fraumeni syndrome exhibit transdominant effects on the biochemical properties of the wild-type p53. Oncogene. 1993;8:2449–2456. [PubMed] [Google Scholar]
- 64.Tada M, Sakuma S, Iggo R D, Saya H, Sawamura Y, Fujiwara T, Roth J A. Monitoring adenoviral p53 transduction efficiency by yeast functional assay. Gene Ther. 1998;5:339–344. doi: 10.1038/sj.gt.3300605. [DOI] [PubMed] [Google Scholar]
- 65.Tsutsumi-Ishi Y, Tadokoro A, Hanaoka F, Tsuchida N. Response of heat shock element within the human HSP70 promoter to mutated p53 genes. Cell Growth Differ. 1995;6:1–8. [PubMed] [Google Scholar]
- 66.Ueba T, Nosaka T, Takahashi J A, Shibata F, Florkiewicz R Z, Vogelstein B, Oda Y, Kikuchi H, Hatanaka M. Transcriptional regulation of basic fibroblast growth factor gene by p53 in human glioblastoma and hepatocellular carcinoma cells. Proc Natl Acad Sci USA. 1994;91:9009–9013. doi: 10.1073/pnas.91.19.9009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Unger T, Mietz J A, Scheffner M, Yee C L, Howley P M. Functional domains of wild-type and mutant p53 proteins involved in transcriptional regulation, transdominant inhibition, and transformation suppression. Mol Cell Biol. 1993;13:5186–5194. doi: 10.1128/mcb.13.9.5186. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Unger T, Nau M M, Segal S, Minna J D. p53: a transdominant regulator of transcription whose function is ablated by mutations occurring in human cancer. EMBO J. 1992;11:1383–1390. doi: 10.1002/j.1460-2075.1992.tb05183.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Vogelstein B, Kinzler K W. p53 function and dysfunction. Cell. 1992;70:523–526. doi: 10.1016/0092-8674(92)90421-8. [DOI] [PubMed] [Google Scholar]
- 70.Walker K K, Levine A J. Identification of a novel p53 functional domain that is necessary for efficient growth suppression. Proc Natl Acad Sci USA. 1996;93:15335–15340. doi: 10.1073/pnas.93.26.15335. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Wang Y, Reed M, Wang P, Stenger J E, Mayr G, Anderson M E, Schwedes J F, Tegtmeyer P. p53 domains: identification and characterization of two autonomous DNA-binding regions. Genes Dev. 1993;7:2575–2586. doi: 10.1101/gad.7.12b.2575. [DOI] [PubMed] [Google Scholar]
- 72.Yang A, Kaghad M, Wang Y, Gillett E, Fleming M D, Dotsch V, Andrews N C, Caput D, McKeon F. p63, a p53 homolog at 3q27-29, encodes multiple products with transactivating, death-inducing, and dominant-negative activities. Mol Cell. 1998;2:305–316. doi: 10.1016/s1097-2765(00)80275-0. [DOI] [PubMed] [Google Scholar]