Abstract
The camelid single-domain antibody fragment, commonly referred to as a nanobody, achieves the targeting power of conventional monoclonal antibodies (mAbs) at only a fraction of their size. Isolated from camelid species (including llamas, alpacas, and camels), their small size at ∼15 kDa, low structural complexity, and high stability compared with conventional antibodies have propelled nanobody technology into the limelight of biologic development. Nanobodies are proving themselves to be a potent complement to traditional mAb therapies, showing success in the treatment of, for example, autoimmune diseases and cancer, and more recently as therapeutic options to treat infectious diseases caused by rapidly evolving biological targets such as the SARS-CoV-2 virus. This review highlights the benefits of applying a proteomic approach to identify diverse nanobody sequences against a single antigen. This proteomic approach coupled with conventional yeast/phage display methods enables the production of highly diverse repertoires of nanobodies able to bind the vast epitope landscape of an antigen, with epitope sampling surpassing that of mAbs. Additionally, we aim to highlight recent findings illuminating the structural attributes of nanobodies that make them particularly amenable to comprehensive antigen sampling and to synergistic activity—underscoring the powerful advantage of acquiring a large, diverse nanobody repertoire against a single antigen. Lastly, we highlight the efforts being made in the clinical development of nanobodies, which have great potential as powerful diagnostic reagents and treatment options, especially when targeting infectious disease agents.
Keywords: nanobody, single-domain antibody, monoclonal antibody, mass spectrometry, antibody display, antibody selection, antibody engineering, antibody therapeutics, VHH
Graphical Abstract

Highlights
-
•
Successful methods used to produce high-affinity nanobodies, with a focus on proteomic approaches.
-
•
Nanobody features enabling comprehensive sampling of an antigen’s surface.
-
•
Advantages of a large diverse repertoire of nanobodies against an antigen.
-
•
Nanobodies as promising therapeutics targeting infectious diseases, e.g.SARS-CoV-2.
-
•
Key challenges when creating a nanobody-based therapeutic.
In Brief
This review summarizes successful, widely used methods to produce high-affinity nanobodies against an antigen. We discuss the structural attributes of nanobodies that enable them to comprehensively sample their antigen surface and to engage in synergy pairs—underscoring the powerful advantages of acquiring a large, diverse nanobody repertoire against a single antigen. Lastly, we highlight the efforts being made in the development of nanobody therapeutics, which have great potential as diagnostic reagents and treatment options, especially when targeting infectious disease agents.
The precision targeting power of antibodies makes them some of the most widely utilized biologic reagents in both general scientific research and therapeutic development (1, 2, 3). Antibody-based therapeutics have provided a significant advance in the fight against numerous human diseases, especially cancer (4, 5, 6, 7). A compelling single domain antibody fragment, termed nanobody (a registered trademark of Ablynx NV, Sanofi) has emerged as an alternative to the precision targeting power of conventional monoclonal antibodies (mAbs), with several advantages over mAbs whilst filling complementary roles (8, 9). Generated from the variable domain of a unique subset of immunoglobulins found in camelids (e.g. llamas and alpacas), nanobodies consist only of heavy-chain homodimers, with no associated light chains (Fig. 1), and at only ∼15 kDa they are the smallest known antigen-binding single polypeptide chain occurring in any natural antibody (10, 11). Due to their small size, nanobodies can target epitopes inaccessible to conventional antibodies, and coupled with their ease of production (especially when compared with other mAbs), they can make ideal research tools with myriad applications from crystallographic chaperones to diagnostic development (8).
Fig. 1.
Structural overview of nanobodies.A, diagram of HCAb and IgG domain structures. B, organization of typical VHH (nanobody) and VH variable domains. CDR3 lengths vary substantially, but on average are higher in VHH domains from all species (averaging ∼14–18 vs. ∼11 residues in VH) (138, 139). In addition to the canonical disulfide (yellow) between cysteines at positions 23 and 104 in FR1 and FR3, camels commonly have a disulfide between CDR3 and position 35 to 38 in CDR1. At lower frequency (∼50%), alpacas have a cysteine at position 55 in FR2 that can form a disulfide with CDR3 (139, 140). Similar non-canonical cysteines and disulfides can also be found in llama VHH sequences, typically between CDR3 and CDR1 or FR2 residues, and more rarely in VH sequences. Positions are numbered by IMGT convention. (CDR = complementarity-determining region; FR = framework). C, the crystal structure of an anti-green fluorescent protein (GFP) nanobody (gray) bound to GFP (sand) (PDB ID: 8SG3), highlighting the three CDR and FR regions.
The application of nanobodies in scientific research is well-established and well-covered elsewhere (8, 12, 13, 14). In recent years, it is in therapeutic and diagnostic development that nanobodies have received considerable attention. To date, numerous nanobody therapeutics have entered the global therapeutic market, beginning with Caplacizumab, a bivalent nanobody drug that received global approval for use in 2019 for the treatment of the autoimmune disease aTTP (acquired thrombotic thrombocytopenic purpura) (15, 16). Following the success of Caplacizumab, other nanobody-based therapeutics have been approved for use in various countries, including Ozoralizumab in Japan, for treatment of rheumatoid arthritis (17, 18), the cancer immunotherapy Envafolimab in China (19, 20), and the nanobody-based CAR-T therapeutic Ciltacabtagene autoleucel, with both FDA and EU approval (21). Infectious disease research has also seen significant interest in the therapeutic potential of nanobodies, including efforts to combat multi-drug resistant bacteria (22, 23) and rapidly evolving human viruses (24, 25, 26). This review aims to summarize the benefits in generating large, diverse repertoires of nanobodies against a single antigen, the different approaches used to achieve this, with a focus (as befitting this journal) on more recent mass spectrometry (MS)-based proteomic techniques, and to highlight promising applications of nanobody technology, especially for the treatment and diagnosis of infectious diseases.
Techniques for Nanobody Discovery
A variety of alternative approaches to generating nanobodies have emerged over recent years; all underscore the relative ease of nanobody production over conventional mAbs. These techniques all start with a large, diverse collection of nanobody clones, ultimately derived from camelid immune cells (usually from llamas and alpacas, and less commonly, camels). Nanobodies against a target of interest are then selected by a range of methods: through expression display and panning of an intermediate clonal library (phage, yeast, or ribosome display), or by direct detection of relevant serum-derived antibodies (MS-based identification) (Fig. 2). The key step in all of these techniques is identifying sequences of nanobodies (the antigen-binding VHH variable domains) from target-specific heavy chain-only antibodies (HCAbs) (Fig. 1). Libraries used in these techniques are typically prepared from camelids immunized with the target antigen of interest, and for savings in cost and time, mixtures of at least 5 to 10 antigens can be used in immunizations with no loss in immune response strength (27, 28). As animal husbandry for camelids can be demanding and require outsourcing, mice have been engineered with germline camelid nanobody sequences that can mature HCAb antibodies in response to immunization as a more laboratory-accessible option, though the overall immune diversity of these mice is significantly lower than camelids (29).
Fig. 2.
Outline of methods used to identify antigen-specific nanobody candidates. Libraries of VHH (nanobody) DNA sequences can be produced from camelids (llamas, alpacas, camels) immunized with antigen(s) of interest, or mice genetically engineered with germline VHH. These can be directly sequenced or cloned into a display library, or libraries can be prepared from naïve or synthetic libraries independent of immunization. Libraries can then be screened in a variety of systems, including phage display, yeast display, or ribosome display. Antigen-binding clones can be enriched by affinity capture, or by FACS after incubating with fluorescently-labeled antigen. Binding antibodies can alternatively be identified directly from serum, with the HCAb fraction affinity-purified, and antigen-bound nanobody fragments analyzed by LC-MS/MS to identify nanobodies from an in silico sequence library.
There has been increasing interest in the use of naïve libraries for nanobody screening, in which sufficiently large and diverse libraries from non-immunized animals, expanded using synthetic techniques, or fully synthetic libraries, can be panned for antigen-specific HCAbs (30, 31, 32, 33, 34). Publicly or commercially available libraries have made this a particularly accessible option for investigators by eliminating the need for animal work. While this saves considerable time and resources on animal immunization, starting affinities of binders identified from naïve libraries tend to be lower, typically requiring affinity maturation or multimerization to approach the sub-nanomolar affinities routinely achieved by immune-raised nanobodies (31).
Display Library Approaches
Display library screening has been the most common technology for nanobody selection, providing the majority of nanobodies in use. Phage display was the first method applied to identifying nanobodies from camelids and continues to be used routinely, though yeast display and ribosome display are now in wide use as well. Typical display approaches rely on cloning libraries from nanobody sequences amplified from lymphocyte cDNA for surface expression in phage (35, 36, 37) or yeast (33, 38, 39), and panning against the antigen of interest. This is a widely available approach that has been successfully applied to a multitude of antigens, and remains the most accessible approach in nanobody development. While both yeast and phage libraries have proven effective, yeast display offers advantages by allowing for direct selection of clones by cell sorting, and even release of soluble protein from the cell surface for analysis (39). Recent improvements in this method have made it more robust, with optimizations to surface fusion protein design, library preparation, and panning parameters (33, 39, 40).
Ribosome display is an alternative and cell-free screening approach, in which library mRNA is translated in vitro but protein is prevented from leaving the ribosome, to preserve a complex with both a nanobody protein and its encoding transcript that can then be screened against antigens and sequenced (41, 42). This fully in vitro approach allows for larger libraries than cell-based methods (1010-1012), making it particularly useful for naïve libraries (see below) (34, 43). This technique has also been incorporated into Illumina high-throughput sequencing platforms, allowing in situ sequencing and measurement of antigen binding activity of up to ∼108 clones (44). While this technique is subject to the limitations of in vitro translation and naïve libraries (above), it provides a highly parallelized alternative to nanobody screening, with possibilities for additional functional assays on immobilized libraries.
Mass Spectrometry in Nanobody Identification
The introduction of proteomic, mass spectrometry-based methods for nanobody identification has provided a complementary approach to display procedures (45, 46, 47, 48, 49). With this technique, lymphocyte cDNA is again prepared from immunized animals, but rather than cloning into a display library, PCR-amplified nanobody sequences are directly analyzed by next-gen sequencing to generate an in silico library of HCAb variable domains. To identify antigen-specific sequences, HCAbs from sera are affinity-purified against the target antigen, optionally digested to remove Fc, and bound nanobody fragments analyzed by LC-MS/MS. Identified peptides are then matched to the in silico nanobody library to determine full-length sequences to be synthesized and expressed as nanobody candidates. This approach has the advantage of directly analyzing antibodies in serum, allowing correlation of nanobody sequences to the animal’s effective immune response. The use of protein rather than cell-based affinity isolation also allows significantly more flexibility in this stage of selection, as there is no need to maintain cell viability for a display library. This allows for significantly higher stringency in wash conditions (e.g. salt, detergent, pH), facilitating enrichment of ultra-high-affinity binders, or antibodies able to retain stability and binding under a given biochemical condition.
However, MS analysis of purified HCAb samples also presents unique challenges compared to typical proteomic samples. These concerns are broadly shared with antibody proteomics in general, though significantly simplified by the lack of a light chain (50). First, antibody repertoires are highly divergent between individuals, meaning that sequence libraries are needed for each immunized animal. This can be addressed by sampling B cells from animals post-immunization for HCAb sequencing from cDNA. The need for animal-specific libraries can be alleviated to some extent by combining many antigens for a single immunization round.
Second, the high sequence similarity of framework regions within the variable domain necessitates specialized processing to unambiguously identify unique nanobody sequences; the nanobody-defining complementarity-determining regions (CDRs), the essential antigen-binding loops of all antibodies, must therefore have adequate peptide coverage for sequence matching, in particular CDR3, generally the longest, most diverse, and most relevant to binding (51). Multiple groups have utilized similar approaches using bottom-up MS to facilitate such analyses (Fig. 3) (45, 46, 49). Key features of these pipelines include customized software used to search large databases of nanobody sequences, and matching and validating peptides covering CDRs to identify high-confidence nanobody candidates. High sequence conservation, as well as the potential for non-specific HCAb contaminants, makes false positives a particular concern in this analysis, and these pipelines are largely reliant on unique fingerprint peptides. One way to reduce these false positives is by using a decoy database to identify and filter out non-specific nanobody sequences, either from a separate non-immunized animal or if possible from a pre-immune library of the same target animal (46).
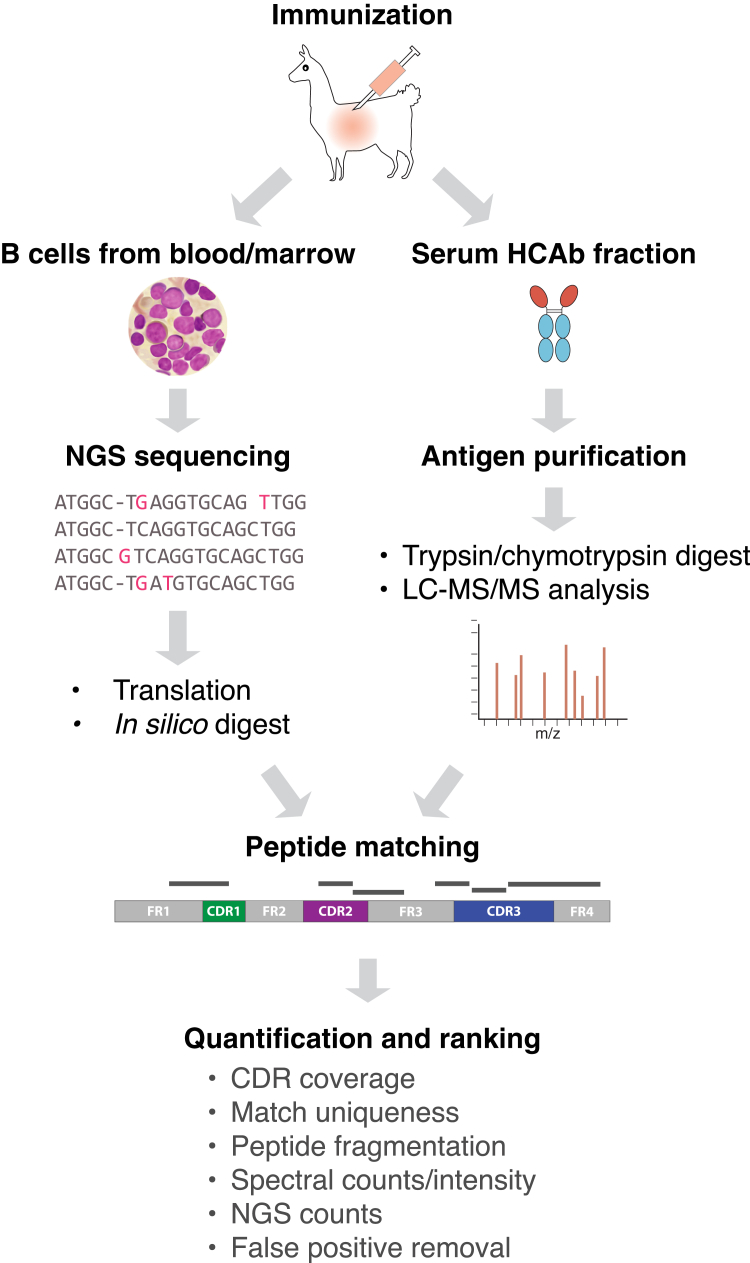
Fig. 3.
Overview of a typical mass spectrometry approach to nanobody identification. After camelid immunization, both sera and lymphocytes from blood or marrow are sampled from the animal. A sequence library of all nanobody-expressing transcripts is generated by next-gen DNA sequencing and prepared for peptide searching by in silico translation and protease digestion. In parallel, an HCAb fraction is prepared from serum and affinity-purified against the target antigen. Purified nanobody-encoding HCAb fragments are separated by SDS-PAGE and digested by trypsin and/or chymotrypsin before LC-MS/MS analysis. MS data is then searched against the corresponding animal-generated sequence library, and peptide matches used to identify high-confidence nanobody clones by algorithmic weighting of coverage and other parameters.
In addition to explicit filtering using decoy data, optimized searching and sequence ranking algorithms are key to the confident identification of antigen-specific nanobody sequences. Peptide coverage of variable CDR regions must be appropriately weighted, as these are the key determinant fingerprints of unique sequences. In particular, peptide fragmentation across CDRs must be assessed to confirm true matches, as framework sequence alignment alone is not sufficient for unique identification. Furthermore, the amino acid distribution in and around the key CDR3 region tends to limit coverage by tryptic peptides; chymotrypsin has been found to more reliably cover this area, and thus a combination of proteases is preferred for optimal sequence coverage (46, 49).
Comparative Advantages and Applications
MS-based nanobody identification is highly complementary to display library screening, each with unique advantages and disadvantages. MS can be biased against sequences that do not readily provide detectable or identifying peptides, while display approaches can select against rare clones in the library or those that are poor expressers or binders in the context of the surface display system. Differences in sensitivity to binding affinity may also affect results between these approaches, as display methods with multiple antibody copies will have a greater avidity effect (higher effective affinity due to multiple binding interactions) than single-molecule serum IgG isolation. Notwithstanding these differences, both approaches have been applied with great success against diverse antigens, and can even be used in combination to increase overall nanobody coverage and diversity (40, 47). While it is difficult to directly compare the results of these techniques due to the large number of variables in antigen immunogenicity, animal responses, and repertoire sizes, in one example targeting SARS-CoV-2 spike with identical source libraries using either yeast display or MS-based identification, similarly high hit rates and affinities were obtained up to at least 30 candidates, though greater overall diversity allowed larger numbers of unique candidates from MS identification in this case (40, 47).
The Benefits of Large Repertoires of Nanobodies
As discussed above, an MS approach enables the identification of high-confidence nanobody sequences from the immune response of the animal (47), where the uniqueness of the CDR3 region is first identified and further used to group the repertoire of potential nanobodies based on CDR3 sequence similarity. Mast et al. (47) revealed the successful application of this method when generating nanobodies against the full-length spike protein from ancestral SARS-CoV-2. They identified 374 high-confidence nanobody sequences, which were further categorized via their CDR3 sequences into 183 groups or “nodes.” They observed that nanobodies within each node displayed similar antigen binding behavior, to suggest more similar epitope coverage within each node than between each node. The authors then functionally characterized nanobodies from each major node, which enabled them to generate a final repertoire of 116 high-affinity nanobodies, importantly targeting all major domains of the spike protein. The repertoire of nanobodies identified by Mast et al. (47) revealed numerous nanobody unique (“hidden”) epitopes on spike, uncharted by mAbs targeting the same antigen (47, 52).
The small size of nanobodies is believed to be key to accessing these “hidden” epitopes on antigens. Additionally, numerous studies have collated structural information from the protein data bank (PDB) to demonstrate that nanobodies can bind their antigen in different orientations (often utilizing their framework (FR) regions to do so), greatly aiding in comprehensive sampling of their antigen surface to access and bind a variety of epitope shapes (53, 54, 55, 56). This is in stark contrast to conventional mAbs which interact primarily via their three CDRs, in what can be described as a “head-on” manner (10, 53). This comparative freedom of interaction orientation and ability to utilize large portions of the FR regions enables nanobodies to create unique compositions of epitope binding surfaces (paratopes) when bound to their antigen, such as seen with anti-GFP nanobodies (53, 57), anti-norovirus capsid nanobodies (58), and anti-SARS-CoV-2 RBD nanobodies (53, 59) (Fig. 4A). These different orientations have numerous benefits, as they potentially aid in: (i) comprehensive sampling of the antigen surface (Fig. 4B); (ii) attacking numerous distinct epitopes on an antigen and (iii) enabling multiple nanobodies to bind the same antigen simultaneously, an ability greatly aided by their size.
Fig. 4.
Nanobodies bind antigens in diverse orientations.A, the three different orientations (Binding mode 1, Binding mode 2 and Binding mode 3) a nanobody can bind its antigen as determined by the composition of its paratope. B, nanobodies which bind their antigen in one of the three binding modes as defined in (A) are shown to be able to bind simultaneously to their target antigen. Panel I – three different anti-RBD nanobodies bound to RBD (59); Panel II – four different nanobodies bound to P domain of norovirus (58); and Panel III – four different anti-GFP nanobodies bound to GFP (53).
The ability of multiple nanobodies to bind a single antigen simultaneously is especially useful when applying nanobody technology to the generation of therapeutics and diagnostics, particularly against antigens of rapidly evolving disease agents. This was exemplified by Mast et al. (47), who showed how neutralizing nanobodies binding distinct epitopes on the receptor binding domain (RBD) of SARS-CoV-2 spike readily engaged in synergy, in which nanobody pairs produced neutralization mixtures far more potent than the predicted additive behavior of the nanobodies’ separate activities (47). The authors showcased numerous synergistic relationships between different nanobody pairs selected from their nanobody repertoire, many of which remained effective against numerous SARS-CoV-2 strains (47, 52). In addition, factoring in the different orientations different nanobodies adopt when binding an antigen can greatly facilitate the rational design of multivalent formulations of nanobodies as research tools and therapeutics, where multimeric formatting is a common feature of the nanobody therapeutics currently licensed and in clinical trials (60).
Applying Nanobodies in Proteomic Research
Researchers have utilized the unique advantages of nanobodies in a wide variety of research and clinical applications, including microscopy and structural biology, which have been extensively surveyed (8, 9, 61, 62).
To highlight one expanding area of use, nanobodies’ size and ease of modification have made them particularly suited to various approaches in proteomic research. Several nanobody toolkits have been implemented for the purification of protein complexes for ex vivo proteomic and structural studies, by targeting widespread tags such as GFP or novel peptide tags (45, 57, 63, 64). While retaining the high affinity and specificity of traditional antibody immunoprecipitation, nanobodies offer further flexibility in ease and low cost of production. The resilience and small size of nanobodies also make them more amenable to on-bead structural analysis of captured complexes, for example by cross-linking MS or cryo-EM (65, 66). Nanobodies can also be compatible with in vivo expression in heterologous systems and have been extensively used in intracellular constructs or “intrabodies” to explore protein localization and for proteomic and interactomic studies. Nanobodies can be expressed either independently or as genetic fusions to selectively target a tagged or native antigen for functional regulation, visualization, or biosensing (67). Intracellular nanobodies can also be used for targeted protein modifications, such as biotin-labeling via fusions with biotin ligase (BioID or TurboID), enabling proximity-based mapping of protein-protein interactions. This adaptable technique has been applied with nanobodies targeting GFP-tagged proteins in zebrafish and Caenorhabditis elegans (68, 69). Developing nanobodies for intracellular use can be a challenge, as many nanobodies do not fold properly in the cytoplasm, primarily due to the lack of required disulfide formation in the reducing intracellular environment, demanding either selection or engineering of suitable clones. Many approaches have been developed to engineer suitable nanobodies (surveyed in (67)) including cloning CDRs into optimized scaffolds (32), directly selecting candidates for intracellular activity in two-hybrid or other functional screens (70, 71), or sequence engineering by mutations in key residues (72).
Several nanobody-tag combinations have also been characterized to facilitate in vivo use of nanobodies. These include 10 to 14 amino acid “NanoTag” peptide tags that were developed with corresponding nanobodies and validated in Drosophila (73), the 14 amino acid ALFA-tag and its high-affinity NbALFA nanobody partner (63), and others (74). These short peptide tags are more easily incorporated into target proteins without disrupting function, making them well-suited to studies using nanobodies against intracellular targets, and can be used in combination (64, 75).
Nanobodies Against SARS-CoV-2 and Infectious Diseases
Nanobodies are increasingly being applied as reagents for the treatment and diagnosis of infectious diseases. Currently, two nanobodies, ARP1 and LMN-101, are in clinical trials for the treatment of diarrheal disease caused by rotavirus (76) and Campylobacter jejuni (77) respectively. Additionally, nanobodies are being explored as a treatment option for placental malaria by targeting the VAR2CSA protein of Plasmidum falciparum (78); influenza by targeting the M2 surface protein of the virus (79) and also conserved regions on Hemagglutinin (HA) (80); human papilloma virus (HPV) (81, 82, 83) and human African trypanosomal disease (84). As mentioned previously, the ability of nanobodies to target non-immunodominant regions on proteins, which are often highly conserved, establishes the opportunity of generating broad-spectrum nanobody therapeutics against a pathogen. This is exemplified by the nanobodies targeting influenza virus, where nanobody M2-7A and several other nanobodies successfully targeted the highly conserved ion channel and regions of HA on the viral surface (79, 85, 86). This suggests the possibility of formulating an influenza treatment effective against multiple influenza strains.
In recent years, SARS-CoV-2 has been a prime testbed for therapeutic solutions to a rapidly evolving disease agent, with many efforts demonstrating the unique properties of nanobodies. The COVID-19 pandemic saw a dramatic rise in the development of nanobodies as both therapeutics and diagnostics against the SARS-CoV-2 virus, targeting the viral spike protein. Numerous SARS-CoV-2 spike-neutralizing nanobodies have been identified, binding spike epitopes distinct from those bound by mAbs (47, 59, 87, 88). In the work by Mast et al. (47), their repertoire of over 100 high-affinity nanobodies targeted all major regions of the SARS-CoV-2 spike, with many nanobodies binding highly conserved regions hidden from the selective pressure of the conventional mAb response. This included the highly glycosylated S2 domain and regions on the RBD resistant to mutational escape. Several of these regions showed strong conservation between SARS-CoV-2 variants, resulting in the identification of pan-variant nanobodies that remained effective at neutralizing emerging strains of the virus (52, 59, 87), in contrast to conventional mAbs that began to lose viral neutralizing efficacy as new strains of the virus rapidly emerged (89). Furthermore, Ketaren et al. (52), showed how nanobodies within their repertoire were specific for certain variants of SARS-CoV-2, which introduces the possibility of creating diagnostic agents able to discriminate between different viral strains, enabling a tailored approach to treatment upon infection.
Unsurprisingly, efforts are being made to develop broad-spectrum nanobody-based therapeutics against betacoronaviruses by targeting highly conserved regions on spike, where a single nanobody could potentially target multiple members of the betacoronaviral family. This is exemplified by Laroche et al. (90), where the authors used deep mutational scanning to design nanobodies able to target both SARS-CoV-1 and SARS-CoV-2. Furthermore, as mentioned earlier, the ability of two nanobodies to readily engage in synergy to create potent neutralization mixtures against the ancestral SARS-CoV-2 virus has recently been shown to extend the effectiveness of the nanobody repertoire established by Mast et al. (47) in neutralizing SARS-CoV-2 variants from successive new strains (52). Intriguingly, the authors revealed a unique synergy relationship between two nanobodies, where the combination of a neutralizing and non-neutralizing nanobody targeting the Delta variant of SARS-CoV-2 still exhibited synergy to create a potent synergistic cocktail. This flexible capacity for synergistic behavior opens up a potentially powerful property of nanobodies to consider when designing nanobody therapeutics, which is enabled by a repertoire of nanobodies large enough to contain binders with numerous non-overlapping epitopes. Aside from the synergy observed between nanobodies targeting spike of SARS-CoV-2 (47, 52), nanobody synergy mixtures have also been observed between nanobodies developed against the capsid of norovirus (91) and HPV nanobodies (82). With the increasing use of nanobody technology to target human disease agents, especially with pipelines (as described in this review) available that maximize nanobody sequence identification to generate highly diverse nanobody repertoires, the compelling property of synergy is sure to become a powerful feature when utilizing nanobodies in both a research and clinical setting.
Collectively, the large body of work accumulated since the onset of the COVID-19 pandemic in generating nanobodies against the spike protein of SARS-CoV-2 has exploited many unique facets of nanobodies, drawing particular attention to the power of their much smaller size to overcome many limitations of conventional mAbs when developing persistent therapeutic options against infectious disease agents. Additionally, the rapid response of researchers during the COVID-19 pandemic who engaged nanobody technology to provide both nanobody tools and strategies to combat the virus, reveals the enormous potential to apply this technology to other viruses, particularly those that may have a high pandemic potential.
Considerations When Developing Nanobodies as Therapeutics
Critical design elements when developing a nanobody therapeutic include (i) nanobody humanization to limit potential immunogenicity; (ii) the final drug structure and oligomeric state, which often requires optimization to prevent overly rapid systemic clearance of nanobodies and/or improve antigen binding (92), and (iii) the drug administration route, which coupled to (ii) influences the bioavailability of the drug (Fig. 5).
Fig. 5.
Humanization and therapeutic applications of nanobodies.A, linear schematic of a typical nanobody structure. Regions on the structure commonly humanized, yet that have been shown to participate in antigen binding, are represented as red circles with the corresponding sequence numbering (IMGT) (Adapted from a meta-analysis of ∼100 nanobody-antigen interactions analyzed from the protein data bank (PDB) (53)). The amino acid substitutions within FR2 for the corresponding drugs are annotated (determined from the sequences of each drug sourced from https://gsrs.ncats.nih.gov/ginas/app/beta/home or 7EOW for Caplacizumab). The potential effect on binding when targeting these four residues on nanobodies that adopt Binding mode 2 (PBD ID: 5my6) is shown. B, different strategies to deliver and create approved nanobody therapeutics. Figure created with BioRender.com. CN, China; JPN, Japan.
Humanization
Mechanisms to humanize nanobodies are covered extensively elsewhere (9, 93, 94) and largely involve mutating residues within the nanobody scaffold (FR region) to better mimic that of the human VH domain, particularly that of the FR2 region that is a main point of difference between camelid VHH and human VH (94). (Fig. 5A). Though nanobody humanization is commonly practiced, there are conflicting interpretations as to whether humanization in fact reduces immunogenicity or limits anti-drug antibodies (ADAs) targeting the nanobody therapeutic. A study performed by Ackaert et al. (95) tested non-humanized nanobodies targeting human epidermal growth factor receptor 2 (HER2) and macrophage mannose receptor (MMR) in an in vitro immunogenicity assay, and saw no significant induction of a T-cell response to suggest strong immunogenicity. Though many humanized nanobody drugs tested clinically have similarly shown minimal immunogenicity (93, 96), this is not always the case – for example, one humanized tetravalent nanobody targeting the TNF death receptor 5 (DR5), resulted in hepatotoxicity and detection of ADAs targeting the nanobody-drug, leading to termination of the study (97). These results suggest that immunogenicity is not limited to the presence of non-human sequences on the protein-based therapeutic, and the contribution of other factors such as pre-existing ADAs and instability of the therapeutic must also be considered (97, 98). Furthermore, it is important to note that the presence of ADAs has been shown to be a common characteristic of many approved protein-based therapeutics, including human monoclonal antibodies (99, 100). Ultimately, with humanization a feature of all approved nanobody-based drugs (Fig. 5A), it remains a measure commonly pursued when developing nanobodies for human use.
The strategies that are employed to humanize nanobodies largely involve engineering the FR regions, particularly FR2, which, as shown above, can impact antigen binding and/or the biophysical properties of nanobodies (53, 94). This is seen when mutating the FR2 region, which is shown in numerous structures to be involved in both antigen binding (53) and maintaining nanobody solubility (94). As a nanobody’s binding and biophysical properties can be impacted by humanization, one way to circumvent the impact of classical humanization strategies on nanobody binding would be to select for nanobodies within a repertoire that did not utilize key scaffold regions for binding—underscoring the value of having a large, diverse starting repertoire of nanobodies. With the growing number of high-resolution nanobody-antigen structures, it is increasingly likely that computational modeling can be used to predict what residues are involved in antigen binding (i.e. predict the nanobody paratope). For example, FR2 is often recruited to bind an antigen in nanobodies with a short CDR3 (53, 101) – this information could be used to predict nanobodies that use FR2 to bind their antigen, preventing issues with humanization. Furthermore, to aid in the rational design of humanized nanobody therapeutics, computational programs such as Llamanade (102) and AbNativ (103) have been developed to guide strategies to humanize nanobodies, potentially without compromising both solubility and binding, as recently reviewed in Gordon et al. (96).
Post-Translational Modifications
A major concern for protein therapeutics, and particularly antibodies, is enzymatic or chemical modification during production or storage. Post-translational modifications (PTMs) such as glycosylation, deamidation, and oxidation can affect therapeutic efficacy and bioavailability, and controlling and identifying such changes is a major challenge in biologic drug development (104, 105, 106). Nanobody-based therapeutics share many of these challenges as protein products, but the smaller and simpler structure relative to mAbs can make them more amenable to production, optimization and analysis. Modifications during cellular expression can be a significant challenge for production of molecularly homogeneous conventional antibodies, but are generally not a concern for nanobodies—they are typically produced recombinantly in bacteria, avoiding enzymatic glycosylation or modification otherwise seen with antibodies. When eukaryotic production is required, the nanobody’s single domain lacks canonical sites of antibody modification (i.e. the Fc region) (107), and can be engineered as needed to avoid spurious post-translational modification sites as these are not normally required for activity.
With their smaller size, nanobodies are also more amenable to structural analysis than standard antibodies, particularly by mass spectrometry. At ∼15 kDa, nanobodies are small enough for direct analysis in top-down mass spectrometry (TD-MS) (108, 109, 110), allowing for detection of modifications, and avoiding the need for middle-down methods (pre-cleavage of antibody fragments) or other customized techniques typically required for intact MS of 10× larger mAbs (111, 112). Effective direct fragmentation of nanobodies for sequence assessment has also been demonstrated with TD-MS, in one case showing sequence coverage of 87% from a combination of HCD, ETD, and ultraviolet photodissociation (UVPD) (110). As with standard antibodies, bottom-up MS can also be applied for detection of modifications, with a combination of trypsin and chymotrypsin particularly effective in increasing sequence coverage, as seen in nanobody discovery studies (46, 49).
Multimerization
As mentioned earlier, two major factors that determine the final oligomeric structure of a nanobody drug are the required systemic circulatory half-life and in vivo antigen binding kinetics. To address these variables, multimerization of the nanobody is often performed, enabling the nanobody drug to both (i) increase its systemic retention and (ii) aid in stronger antigen binding in vivo. The approved drugs Ozoralizumab (approved for use in Japan) (17, 18) and Sonelokimab (113) are both nanobody trimers that incorporate a human serum albumin-binding nanobody as their central domain to increase their circulatory half-life to 18 (114) and 11 (115) days respectively. Additionally, the FDA-approved Caplacizumab is a homodimer linked by a tri-alanine repeat linker composed of two nanobodies that binds the A1 subunit of the blood glycoprotein von Willebrand factor to prevent platelet aggregation (15, 16). Furthermore, for these therapeutics, two identical antigen binding domains are incorporated into the final drug formulation, as the monomer showed significantly reduced antigen binding in vivo compared to the multimerized versions (116, 117).
However, the drug formulation strategy shown to be effective for currently approved nanobody drugs is not universally applicable, as there are instances where nanobody multimerization affects drug bioavailability, such as in cancer therapy when targeting solid tumors. In their monomeric state, nanobodies have been shown to successfully penetrate the tumor microenvironment better than their multimerized counterparts (118). This was seen with the anti-HER2 nanobody 2Rb17c, which showed homogenous tumor distribution of the monomeric form within minutes of injection (119), in contrast to its dimeric version and the mAb, trastuzumab, which targets the same antigen. Furthermore, when multimerizing nanobodies, especially in the context of exploring the possibility of creating a hetero-bivalent nanobody therapeutic composed of synergistic nanobodies targeting distinct epitopes on an antigen, structural information (e.g. X-ray crystallography or cryo-EM structures) is essential to determine drug parameters such as linker length, nanobody orientation, and nanobody formulation. Instead of experimental structural information, numerous robust computational protein modeling tools (9, 120) are available to model nanobody and antigen structures, which coupled with molecular docking programs (121, 122, 123), can be used to facilitate the best multimerization design strategy to maintain the synergistic relationship between nanobodies.
Therapeutic Administration Routes
The structural robustness of nanobodies makes them amenable to different strategies of administration, making it possible to tailor the administration route depending on the location of the target. Of the successful nanobody drugs available, Caplacizumab (15, 16) and Ciltacabtagene autoleucel (21) are both administered intravenously, whereas Ozoralizumab (17, 18) and Envafolimab (19, 20, 124) are both administered subcutaneously (Fig. 5B).
More recently, orally administered nanobody drugs have been developed, which carry numerous advantages over other delivery systems including long-term stability, low production costs, and ease of administration for patients (125, 126). The primary site of drug absorption for these drugs is the gastrointestinal tract, making them particularly suitable for diseases of the gut as exemplified by the three monomeric nanobody-based drugs V565 (127, 128, 129), LMN-101 (77), and ARP1/VHH batch 203,027 (76), which have been developed as oral therapeutics to treat Crohn’s disease, traveler’s diarrhea, and rotavirus infection respectively. Notably, the delivery of LMN-101—packaged as capsules—incorporates the emerging technology of nanobody expression in the edible cyanobacterium spirulina (130), which releases the nanobody upon digestion of the cyanobacterium. The small size and low structural complexity of nanobodies readily enables the application of this technique, where orally available nanobody therapeutics could become a powerful tool in distributing drugs to developing nations, especially for the treatment of gastrointestinal diseases.
Finally, an emerging administration route that is being explored for the treatment of respiratory infections is via an intranasal or inhalable route, bypassing the circulatory system to actively target the lungs. These two administration routes require robust biophysical properties of the drug to withstand the formulation required for inhalable or intranasal use (131), which nanobodies can fulfill. This strategy aims to focus the nanobodies on the site of the respiratory infection to increase drug uptake and efficacy, as drugs destined for the lungs delivered systemically have shown decreased bioavailability (132, 133). In one study, a nanobody trimer treating mice infected with SARS-CoV-2 administered intranasally resulted in a faster recovery time for the animals over the same dosage administered intravenously (132). Even for classical mAbs, a mAb destined for the lung given intravenously for the treatment of asthma showed only 0.2% of the mAb reaching the lungs (134). Recently, inhalable formulations of nanobodies have been tested as a treatment option for respiratory syncytial virus (135) and SARS-CoV-2 (136, 137) infection, which could dramatically increase the accessibility of such treatments, due to their relatively easy mode of administration.
Perspectives
Advances in techniques for large-scale nanobody generation and characterization have further spurred the already rapid growth of this class of antibodies in diverse areas of research. With multiple nanobody-based drugs coming into active clinical use in recent years, interest in the therapeutic applications of nanobodies has become particularly robust. As dramatically more nanobodies are developed and methodologies gain wider adoption, the advantages of extensive nanobody repertoires have also become increasingly clear. Nanobodies can cover greater epitope space than typical mAbs, allowing them to target difficult antigen targets, and complementary binders are capable of potent synergistic effects. Nanobodies are thus ideal candidates for combinatorial use and multimerization, along with their general flexibility as single domains suitable for wider functionalization. Many investigators are taking advantage of these uniquely favorable properties to explore new protein-targeted approaches in proteomic and cell biology research, and to expand the territory of antibody-based clinical reagents. Furthermore, the high quality and quantity of work in developing nanobodies targeting SARS-CoV-2 virus and other infectious disease agents underscores the versatility and robustness of nanobodies in the context of diagnostic and therapeutic development. Consequently, nanobodies could prove especially powerful tools in the control, treatment, and prevention of future infectious disease outbreaks, especially viral—an important strategy for the global goal of pandemic preparedness.
Data Availability
All supporting data are provided within the manuscript, supplementary data and supplementary tables.
Conflict of interests
The authors declare that they have no conflicts of interest with the contents of this article.
Acknowledgments
We acknowledge the support of the National Institutes of Health grant P41GM109824, the G. Harold and Leila Y. Mathers Charitable Foundation, the Robertson Therapeutic Development Fund and the Stavros Niarchos Foundation (SNF) as part of its grant to the SNF Institute for Global Infectious Disease Research at The Rockefeller University. Nanobody is a registered trademark of Ablynx NV, Sanofi. All atomic resolution structure representations were created using ChimeraX (https://www.rbvi.ucsf.edu/chimerax).
Author contributions
P. C F., N. E. K., and M. P. R. writing–review & editing, P. C F. and N. E. K. writing–original draft, P. C F. and N. E. K. conceptualization. M. P. R. supervision; M. P. R. funding acquisition.
References
- 1.Singh S., Kumar N.K., Dwiwedi P., Charan J., Kaur R., Sidhu P., et al. Monoclonal antibodies: a review. Curr. Clin. Pharmacol. 2018;13:85–99. doi: 10.2174/1574884712666170809124728. [DOI] [PubMed] [Google Scholar]
- 2.Sharma P., Joshi R.V., Pritchard R., Xu K., Eicher M.A. Therapeutic antibodies in medicine. Molecules. 2023;28:6438. doi: 10.3390/molecules28186438. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Nelson P.N., Reynolds G.M., Waldron E.E., Ward E., Giannopoulos K., Murray P.G. Monoclonal antibodies. Mol. Pathol. 2000;53:111–117. doi: 10.1136/mp.53.3.111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Riccardi F., Dal Bo M., Macor P., Toffoli G. A comprehensive overview on antibody-drug conjugates: from the conceptualization to cancer therapy. Front. Pharmacol. 2023;14 doi: 10.3389/fphar.2023.1274088. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Kaplon H., Chenoweth A., Crescioli S., Reichert J.M. Antibodies to watch in 2022. MAbs. 2022;14 doi: 10.1080/19420862.2021.2014296. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Delgado M., Garcia-Sanz J.A. Therapeutic monoclonal antibodies against cancer: present and future. Cells. 2023;12:2837. doi: 10.3390/cells12242837. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Zahavi D., Weiner L. Monoclonal antibodies in cancer therapy. Antibodies (Basel) 2020;9:34. doi: 10.3390/antib9030034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Muyldermans S. Applications of nanobodies. Annu. Rev. Anim. Biosci. 2021;9:401–421. doi: 10.1146/annurev-animal-021419-083831. [DOI] [PubMed] [Google Scholar]
- 9.Yong Joon Kim J., Sang Z., Xiang Y., Shen Z., Shi Y. Nanobodies: robust miniprotein binders in biomedicine. Adv. Drug Deliv. Rev. 2023;195 doi: 10.1016/j.addr.2023.114726. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Muyldermans S. Nanobodies: natural single-domain antibodies. Annu. Rev. Biochem. 2013;82:775–797. doi: 10.1146/annurev-biochem-063011-092449. [DOI] [PubMed] [Google Scholar]
- 11.Hamers-Casterman C., Atarhouch T., Muyldermans S., Robinson G., Hamers C., Songa E.B., et al. Naturally occurring antibodies devoid of light chains. Nature. 1993;363:446–448. doi: 10.1038/363446a0. [DOI] [PubMed] [Google Scholar]
- 12.Muyldermans S. A guide to: generation and design of nanobodies. FEBS J. 2021;288:2084–2102. doi: 10.1111/febs.15515. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Bao G., Tang M., Zhao J., Zhu X. Nanobody: a promising toolkit for molecular imaging and disease therapy. EJNMMI Res. 2021;11:6. doi: 10.1186/s13550-021-00750-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Jin B.K., Odongo S., Radwanska M., Magez S. Nanobodies: a review of generation, diagnostics and therapeutics. Int. J. Mol. Sci. 2023;24:5994. doi: 10.3390/ijms24065994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Duggan S. Caplacizumab: first global approval. Drugs. 2018;78:1639–1642. doi: 10.1007/s40265-018-0989-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Scully M., Cataland S.R., Peyvandi F., Coppo P., Knobl P., Kremer Hovinga J.A., et al. Caplacizumab treatment for acquired thrombotic thrombocytopenic purpura. N. Engl. J. Med. 2019;380:335–346. doi: 10.1056/NEJMoa1806311. [DOI] [PubMed] [Google Scholar]
- 17.Keam S.J. Ozoralizumab: first approval. Drugs. 2023;83:87–92. doi: 10.1007/s40265-022-01821-0. [DOI] [PubMed] [Google Scholar]
- 18.Tanaka Y. Ozoralizumab: first Nanobody(R) therapeutic for rheumatoid arthritis. Expert Opin. Biol. Ther. 2023;23:579–587. doi: 10.1080/14712598.2023.2231344. [DOI] [PubMed] [Google Scholar]
- 19.Markham A. Envafolimab: first approval. Drugs. 2022;82:235–240. doi: 10.1007/s40265-022-01671-w. [DOI] [PubMed] [Google Scholar]
- 20.Zhang F., Wei H., Wang X., Bai Y., Wang P., Wu J., et al. Structural basis of a novel PD-L1 nanobody for immune checkpoint blockade. Cell Discov. 2017;3 doi: 10.1038/celldisc.2017.4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Mullard A. FDA approves second BCMA-targeted CAR-T cell therapy. Nat. Rev. Drug Discov. 2022;21:249. doi: 10.1038/d41573-022-00048-8. [DOI] [PubMed] [Google Scholar]
- 22.Yu S., Zhang L., Wang A., Jin Y., Zhou D. Nanobodies: the potential application in bacterial treatment and diagnosis. Biochem. Pharmacol. 2023;214 doi: 10.1016/j.bcp.2023.115640. [DOI] [PubMed] [Google Scholar]
- 23.Qin Q., Liu H., He W., Guo Y., Zhang J., She J., et al. Single Domain Antibody application in bacterial infection diagnosis and neutralization. Front. Immunol. 2022;13 doi: 10.3389/fimmu.2022.1014377. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Moliner-Morro A., McInerney G.M., Hanke L. Nanobodies in the limelight: multifunctional tools in the fight against viruses. J. Gen. Virol. 2022;103 doi: 10.1099/jgv.0.001731. [DOI] [PubMed] [Google Scholar]
- 25.Vanlandschoot P., Stortelers C., Beirnaert E., Ibanez L.I., Schepens B., Depla E., et al. Nanobodies(R): new ammunition to battle viruses. Antivir. Res. 2011;92:389–407. doi: 10.1016/j.antiviral.2011.09.002. [DOI] [PubMed] [Google Scholar]
- 26.Bhattacharya M., Chatterjee S., Lee S.S., Chakraborty C. Therapeutic applications of nanobodies against SARS-CoV-2 and other viral infections: current update. Int. J. Biol. Macromol. 2023;229:70–80. doi: 10.1016/j.ijbiomac.2022.12.284. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Chow K.M., Whiteheart S.W., Smiley J.R., Sharma S., Boaz K., Coleman M.J., et al. Immunization of alpacas (Lama pacos) with protein antigens and production of antigen-specific single domain antibodies. J. Vis. Exp. 2019 doi: 10.3791/58471. [DOI] [PubMed] [Google Scholar]
- 28.Pardon E., Laeremans T., Triest S., Rasmussen S.G., Wohlkonig A., Ruf A., et al. A general protocol for the generation of Nanobodies for structural biology. Nat. Protoc. 2014;9:674–693. doi: 10.1038/nprot.2014.039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Xu J., Xu K., Jung S., Conte A., Lieberman J., Muecksch F., et al. Nanobodies from camelid mice and llamas neutralize SARS-CoV-2 variants. Nature. 2021;595:278–282. doi: 10.1038/s41586-021-03676-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Yan J., Wang P., Zhu M., Li G., Romao E., Xiong S., et al. Characterization and applications of Nanobodies against human procalcitonin selected from a novel naive Nanobody phage display library. J. Nanobiotechnol. 2015;13:33. doi: 10.1186/s12951-015-0091-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Huo J., Le Bas A., Ruza R.R., Duyvesteyn H.M.E., Mikolajek H., Malinauskas T., et al. Neutralizing nanobodies bind SARS-CoV-2 spike RBD and block interaction with ACE2. Nat. Struct. Mol. Biol. 2020;27:846–854. doi: 10.1038/s41594-020-0469-6. [DOI] [PubMed] [Google Scholar]
- 32.Moutel S., Bery N., Bernard V., Keller L., Lemesre E., de Marco A., et al. NaLi-H1: a universal synthetic library of humanized nanobodies providing highly functional antibodies and intrabodies. Elife. 2016;5:e16228. doi: 10.7554/eLife.16228. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.McMahon C., Baier A.S., Pascolutti R., Wegrecki M., Zheng S., Ong J.X., et al. Yeast surface display platform for rapid discovery of conformationally selective nanobodies. Nat. Struct. Mol. Biol. 2018;25:289–296. doi: 10.1038/s41594-018-0028-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Zimmermann I., Egloff P., Hutter C.A.J., Kuhn B.T., Brauer P., Newstead S., et al. Generation of synthetic nanobodies against delicate proteins. Nat. Protoc. 2020;15:1707–1741. doi: 10.1038/s41596-020-0304-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Romao E., Morales-Yanez F., Hu Y., Crauwels M., De Pauw P., Hassanzadeh G.G., et al. Identification of useful nanobodies by phage display of immune single domain libraries derived from camelid heavy chain antibodies. Curr. Pharm. Des. 2016;22:6500–6518. doi: 10.2174/1381612822666160923114417. [DOI] [PubMed] [Google Scholar]
- 36.Romao E., Poignavent V., Vincke C., Ritzenthaler C., Muyldermans S., Monsion B. Construction of high-quality camel immune antibody libraries. Methods Mol. Biol. 2018;1701:169–187. doi: 10.1007/978-1-4939-7447-4_9. [DOI] [PubMed] [Google Scholar]
- 37.Arbabi Ghahroudi M., Desmyter A., Wyns L., Hamers R., Muyldermans S. Selection and identification of single domain antibody fragments from camel heavy-chain antibodies. FEBS Lett. 1997;414:521–526. doi: 10.1016/s0014-5793(97)01062-4. [DOI] [PubMed] [Google Scholar]
- 38.Roth L., Krah S., Klemm J., Gunther R., Toleikis L., Busch M., et al. Isolation of antigen-specific VHH single-domain antibodies by combining animal immunization with yeast surface display. Methods Mol. Biol. 2020;2070:173–189. doi: 10.1007/978-1-4939-9853-1_10. [DOI] [PubMed] [Google Scholar]
- 39.Uchanski T., Zogg T., Yin J., Yuan D., Wohlkonig A., Fischer B., et al. An improved yeast surface display platform for the screening of nanobody immune libraries. Sci. Rep. 2019;9:382. doi: 10.1038/s41598-018-37212-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Cross F.R., Fridy P.C., Ketaren N.E., Mast F.D., Li S., Olivier J.P., et al. Expanding and improving nanobody repertoires using a yeast display method: targeting SARS-CoV-2. J. Biol. Chem. 2023;299 doi: 10.1016/j.jbc.2023.102954. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Pluckthun A. Ribosome display: a perspective. Methods Mol. Biol. 2012;805:3–28. doi: 10.1007/978-1-61779-379-0_1. [DOI] [PubMed] [Google Scholar]
- 42.Ferrari D., Garrapa V., Locatelli M., Bolchi A. A novel nanobody scaffold optimized for bacterial expression and suitable for the construction of ribosome display libraries. Mol. Biotechnol. 2020;62:43–55. doi: 10.1007/s12033-019-00224-z. [DOI] [PubMed] [Google Scholar]
- 43.Chen X., Gentili M., Hacohen N., Regev A. A cell-free nanobody engineering platform rapidly generates SARS-CoV-2 neutralizing nanobodies. Nat. Commun. 2021;12:5506. doi: 10.1038/s41467-021-25777-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Porebski B.T., Balmforth M., Browne G., Riley A., Jamali K., Furst M., et al. Rapid discovery of high-affinity antibodies via massively parallel sequencing, ribosome display and affinity screening. Nat. Biomed. Eng. 2023;8:214–232. doi: 10.1038/s41551-023-01093-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Fridy P.C., Li Y., Keegan S., Thompson M.K., Nudelman I., Scheid J.F., et al. A robust pipeline for rapid production of versatile nanobody repertoires. Nat. Methods. 2014;11:1253–1260. doi: 10.1038/nmeth.3170. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Xiang Y., Sang Z., Bitton L., Xu J., Liu Y., Schneidman-Duhovny D., et al. Integrative proteomics identifies thousands of distinct, multi-epitope, and high-affinity nanobodies. Cell Syst. 2021;12:220–234.e229. doi: 10.1016/j.cels.2021.01.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Mast F.D., Fridy P.C., Ketaren N.E., Wang J., Jacobs E.Y., Olivier J.P., et al. Highly synergistic combinations of nanobodies that target SARS-CoV-2 and are resistant to escape. Elife. 2021;10 doi: 10.7554/eLife.73027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Shen Z., Xiang Y., Vergara S., Chen A., Xiao Z., Santiago U., et al. A resource of high-quality and versatile nanobodies for drug delivery. iScience. 2021;24 doi: 10.1016/j.isci.2021.103014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Fridy P.C., Farrell R.J., Molloy K.R., Keegan S., Wang J., Jacobs E.Y., et al. A new generation of nanobody research tools using improved mass spectrometry-based discovery methods. J. Biol. Chem. 2024;300 doi: 10.1016/j.jbc.2024.107623. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Snapkov I., Chernigovskaya M., Sinitcyn P., Le Quy K., Nyman T.A., Greiff V. Progress and challenges in mass spectrometry-based analysis of antibody repertoires. Trends Biotechnol. 2022;40:463–481. doi: 10.1016/j.tibtech.2021.08.006. [DOI] [PubMed] [Google Scholar]
- 51.Safarzadeh Kozani P., Naseri A., Mirarefin S.M.J., Salem F., Nikbakht M., Evazi Bakhshi S., et al. Nanobody-based CAR-T cells for cancer immunotherapy. Biomark Res. 2022;10:24. doi: 10.1186/s40364-022-00371-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Ketaren N.E., Mast F.D., Fridy P.C., Olivier J.P., Sanyal T., Sali A., et al. Nanobody repertoire generated against the spike protein of ancestral SARS-CoV-2 remains efficacious against the rapidly evolving virus. bioRxiv. 2023 doi: 10.1101/2023.07.14.549041. [preprint] [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Ketaren N.E., Fridy P.C., Malashkevich V., Sanyal T., Brillantes M., Thompson M.K., et al. Unique binding and stabilization mechanisms employed by and engineered into nanobodies. bioRxiv. 2023 doi: 10.1101/2023.10.22.563475. [preprint] [DOI] [Google Scholar]
- 54.Mitchell L.S., Colwell L.J. Comparative analysis of nanobody sequence and structure data. Proteins. 2018;86:697–706. doi: 10.1002/prot.25497. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Mitchell L.S., Colwell L.J. Analysis of nanobody paratopes reveals greater diversity than classical antibodies. Protein Eng. Des. Sel. 2018;31:267–275. doi: 10.1093/protein/gzy017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Zavrtanik U., Lukan J., Loris R., Lah J., Hadzi S. Structural basis of epitope recognition by heavy-chain camelid antibodies. J. Mol. Biol. 2018;430:4369–4386. doi: 10.1016/j.jmb.2018.09.002. [DOI] [PubMed] [Google Scholar]
- 57.Rothbauer U., Zolghadr K., Muyldermans S., Schepers A., Cardoso M.C., Leonhardt H. A versatile nanotrap for biochemical and functional studies with fluorescent fusion proteins. Mol. Cell Proteomics. 2008;7:282–289. doi: 10.1074/mcp.M700342-MCP200. [DOI] [PubMed] [Google Scholar]
- 58.Koromyslova A.D., Hansman G.S. Nanobodies targeting norovirus capsid reveal functional epitopes and potential mechanisms of neutralization. PLoS Pathog. 2017;13 doi: 10.1371/journal.ppat.1006636. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Xiang Y., Huang W., Liu H., Sang Z., Nambulli S., Tubiana J., et al. Superimmunity by pan-sarbecovirus nanobodies. Cell Rep. 2022;39 doi: 10.1016/j.celrep.2022.111004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Silva-Pilipich N., Smerdou C., Vanrell L. A small virus to deliver small antibodies: new targeted therapies based on AAV delivery of nanobodies. Microorganisms. 2021;9 doi: 10.3390/microorganisms9091956. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Frecot D.I., Froehlich T., Rothbauer U. 30 years of nanobodies - an ongoing success story of small binders in biological research. J. Cell Sci. 2023;136 doi: 10.1242/jcs.261395. [DOI] [PubMed] [Google Scholar]
- 62.Chanier T., Chames P. Nanobody engineering: toward next generation immunotherapies and immunoimaging of cancer. Antibodies (Basel) 2019;8:13. doi: 10.3390/antib8010013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Gotzke H., Kilisch M., Martinez-Carranza M., Sograte-Idrissi S., Rajavel A., Schlichthaerle T., et al. The ALFA-tag is a highly versatile tool for nanobody-based bioscience applications. Nat. Commun. 2019;10:4403. doi: 10.1038/s41467-019-12301-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Stevens T.A., Tomaleri G.P., Hazu M., Wei S., Nguyen V.N., DeKalb C., et al. A nanobody-based strategy for rapid and scalable purification of human protein complexes. Nat. Protoc. 2024;19:127–158. doi: 10.1038/s41596-023-00904-w. [DOI] [PubMed] [Google Scholar]
- 65.Shi Y., Pellarin R., Fridy P.C., Fernandez-Martinez J., Thompson M.K., Li Y., et al. A strategy for dissecting the architectures of native macromolecular assemblies. Nat. Methods. 2015;12:1135–1138. doi: 10.1038/nmeth.3617. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Arimura Y., Konishi H.A., Funabiki H. MagIC-Cryo-EM: structural determination on magnetic beads for scarce macromolecules in heterogeneous samples. bioRxiv. 2024 doi: 10.1101/2024.01.21.576499. [preprint] [DOI] [Google Scholar]
- 67.Wagner T.R., Rothbauer U. Nanobodies right in the middle: intrabodies as toolbox to visualize and modulate antigens in the living cell. Biomolecules. 2020;10:1701. doi: 10.3390/biom10121701. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Xiong Z., Lo H.P., McMahon K.A., Martel N., Jones A., Hill M.M., et al. In vivo proteomic mapping through GFP-directed proximity-dependent biotin labelling in zebrafish. Elife. 2021;10:e64631. doi: 10.7554/eLife.64631. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Holzer E., Rumpf-Kienzl C., Falk S., Dammermann A. A modified TurboID approach identifies tissue-specific centriolar components in C. elegans. PLoS Genet. 2022;18 doi: 10.1371/journal.pgen.1010150. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Pellis M., Muyldermans S., Vincke C. Bacterial two hybrid: a versatile one-step intracellular selection method. Methods Mol. Biol. 2012;911:135–150. doi: 10.1007/978-1-61779-968-6_9. [DOI] [PubMed] [Google Scholar]
- 71.Schmidt F.I., Hanke L., Morin B., Brewer R., Brusic V., Whelan S.P., et al. Phenotypic lentivirus screens to identify functional single domain antibodies. Nat. Microbiol. 2016;1 doi: 10.1038/nmicrobiol.2016.80. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Dingus J.G., Tang J.C.Y., Amamoto R., Wallick G.K., Cepko C.L. A general approach for stabilizing nanobodies for intracellular expression. Elife. 2022;11:e68253. doi: 10.7554/eLife.68253. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Xu J., Kim A.R., Cheloha R.W., Fischer F.A., Li J.S.S., Feng Y., et al. Protein visualization and manipulation in Drosophila through the use of epitope tags recognized by nanobodies. Elife. 2022;11:e74326. doi: 10.7554/eLife.74326. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Traenkle B., Emele F., Anton R., Poetz O., Haeussler R.S., Maier J., et al. Monitoring interactions and dynamics of endogenous beta-catenin with intracellular nanobodies in living cells. Mol. Cell Proteomics. 2015;14:707–723. doi: 10.1074/mcp.M114.044016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Igreja C., Loschko T., Schafer A., Sharma R., Quiobe S.P., Aloshy E., et al. Application of ALFA-tagging in the nematode model organisms Caenorhabditis elegans and pristionchus pacificus. Cells. 2022;11:3875. doi: 10.3390/cells11233875. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Sarker S.A., Jakel M., Sultana S., Alam N.H., Bardhan P.K., Chisti M.J., et al. Anti-rotavirus protein reduces stool output in infants with diarrhea: a randomized placebo-controlled trial. Gastroenterology. 2013;145:740–748.e748. doi: 10.1053/j.gastro.2013.06.053. [DOI] [PubMed] [Google Scholar]
- 77.Al-Ibrahim M. LMN-101 in a Campylobacter Human Challenge Model. ClinicalTrials.gov, National Library of Medicine; Bethesda, MD: 2022. [Google Scholar]
- 78.Ditlev S.B., Florea R., Nielsen M.A., Theander T.G., Magez S., Boeuf P., et al. Utilizing nanobody technology to target non-immunodominant domains of VAR2CSA. PLoS One. 2014;9 doi: 10.1371/journal.pone.0084981. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Wei G., Meng W., Guo H., Pan W., Liu J., Peng T., et al. Potent neutralization of influenza A virus by a single-domain antibody blocking M2 ion channel protein. PLoS One. 2011;6 doi: 10.1371/journal.pone.0028309. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Gaiotto T., Hufton S.E. Cross-Neutralising nanobodies bind to a conserved pocket in the hemagglutinin stem region identified using yeast display and deep mutational scanning. PLoS One. 2016;11 doi: 10.1371/journal.pone.0164296. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Woodham A.W., Cheloha R.W., Ling J., Rashidian M., Kolifrath S.C., Mesyngier M., et al. Nanobody-antigen conjugates elicit HPV-specific antitumor immune responses. Cancer Immunol. Res. 2018;6:870–880. doi: 10.1158/2326-6066.CIR-17-0661. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Zhang W., Shan H., Jiang K., Huang W., Li S. A novel intracellular nanobody against HPV16 E6 oncoprotein. Clin. Immunol. 2021;225 doi: 10.1016/j.clim.2021.108684. [DOI] [PubMed] [Google Scholar]
- 83.Li S., Zhang W., Jiang K., Shan H., Shi M., Chen B., et al. Nanobody against the E7 oncoprotein of human papillomavirus 16. Mol. Immunol. 2019;109:12–19. doi: 10.1016/j.molimm.2019.02.022. [DOI] [PubMed] [Google Scholar]
- 84.Arias J.L., Unciti-Broceta J.D., Maceira J., Del Castillo T., Hernandez-Quero J., Magez S., et al. Nanobody conjugated PLGA nanoparticles for active targeting of African Trypanosomiasis. J. Control Release. 2015;197:190–198. doi: 10.1016/j.jconrel.2014.11.002. [DOI] [PubMed] [Google Scholar]
- 85.Voronina D.V., Shcheblyakov D.V., Esmagambetov I.B., Derkaev A.A., Popova O., Shcherbinin D.N. Development of neutralizing nanobodies to the hemagglutinin stem domain of influenza A viruses. Acta Naturae. 2021;13:33–41. doi: 10.32607/actanaturae.11495. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Gaiotto T., Ramage W., Ball C., Risley P., Carnell G.W., Temperton N., et al. Nanobodies mapped to cross-reactive and divergent epitopes on A(H7N9) influenza hemagglutinin using yeast display. Sci. Rep. 2021;11:3126. doi: 10.1038/s41598-021-82356-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Sun D., Sang Z., Kim Y.J., Xiang Y., Cohen T., Belford A.K., et al. Potent neutralizing nanobodies resist convergent circulating variants of SARS-CoV-2 by targeting diverse and conserved epitopes. Nat. Commun. 2021;12:4676. doi: 10.1038/s41467-021-24963-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Chen F., Liu Z., Jiang F. Prospects of neutralizing nanobodies against SARS-CoV-2. Front. Immunol. 2021;12 doi: 10.3389/fimmu.2021.690742. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Widyasari K., Kim J. A review of the currently available antibody therapy for the treatment of Coronavirus Disease 2019 (COVID-19) Antibodies (Basel) 2023;12:5. doi: 10.3390/antib12010005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Laroche A., Orsini Delgado M.L., Chalopin B., Cuniasse P., Dubois S., Sierocki R., et al. Deep mutational engineering of broadly-neutralizing nanobodies accommodating SARS-CoV-1 and 2 antigenic drift. MAbs. 2022;14 doi: 10.1080/19420862.2022.2076775. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Ruoff K., Kilic T., Devant J., Koromyslova A., Ringel A., Hempelmann A., et al. Structural basis of nanobodies targeting the prototype norovirus. J. Virol. 2019;93 doi: 10.1128/JVI.02005-18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Jovcevska I., Muyldermans S. The therapeutic potential of nanobodies. BioDrugs. 2020;34:11–26. doi: 10.1007/s40259-019-00392-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Rossotti M.A., Belanger K., Henry K.A., Tanha J. Immunogenicity and humanization of single-domain antibodies. FEBS J. 2022;289:4304–4327. doi: 10.1111/febs.15809. [DOI] [PubMed] [Google Scholar]
- 94.Vincke C., Loris R., Saerens D., Martinez-Rodriguez S., Muyldermans S., Conrath K. General strategy to humanize a camelid single-domain antibody and identification of a universal humanized nanobody scaffold. J. Biol. Chem. 2009;284:3273–3284. doi: 10.1074/jbc.M806889200. [DOI] [PubMed] [Google Scholar]
- 95.Ackaert C., Smiejkowska N., Xavier C., Sterckx Y.G.J., Denies S., Stijlemans B., et al. Immunogenicity risk profile of nanobodies. Front. Immunol. 2021;12 doi: 10.3389/fimmu.2021.632687. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Gordon G.L., Raybould M.I.J., Wong A., Deane C.M. Prospects for the computational humanization of antibodies and nanobodies. Front. Immunol. 2024;15 doi: 10.3389/fimmu.2024.1399438. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97.Papadopoulos K.P., Isaacs R., Bilic S., Kentsch K., Huet H.A., Hofmann M., et al. Unexpected hepatotoxicity in a phase I study of TAS266, a novel tetravalent agonistic Nanobody(R) targeting the DR5 receptor. Cancer Chemother. Pharmacol. 2015;75:887–895. doi: 10.1007/s00280-015-2712-0. [DOI] [PubMed] [Google Scholar]
- 98.Holland M.C., Wurthner J.U., Morley P.J., Birchler M.A., Lambert J., Albayaty M., et al. Autoantibodies to variable heavy (VH) chain Ig sequences in humans impact the safety and clinical pharmacology of a VH domain antibody antagonist of TNF-alpha receptor 1. J. Clin. Immunol. 2013;33:1192–1203. doi: 10.1007/s10875-013-9915-0. [DOI] [PubMed] [Google Scholar]
- 99.Pizano-Martinez O., Mendieta-Condado E., Vazquez-Del Mercado M., Martinez-Garcia E.A., Chavarria-Avila E., Ortuno-Sahagun D., et al. Anti-drug antibodies in the biological therapy of autoimmune rheumatic diseases. J. Clin. Med. 2023;12:3271. doi: 10.3390/jcm12093271. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.Strand V., Balsa A., Al-Saleh J., Barile-Fabris L., Horiuchi T., Takeuchi T., et al. Immunogenicity of biologics in chronic inflammatory diseases: a systematic review. BioDrugs. 2017;31:299–316. doi: 10.1007/s40259-017-0231-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.Kirchhofer A., Helma J., Schmidthals K., Frauer C., Cui S., Karcher A., et al. Modulation of protein properties in living cells using nanobodies. Nat. Struct. Mol. Biol. 2010;17:133–138. doi: 10.1038/nsmb.1727. [DOI] [PubMed] [Google Scholar]
- 102.Sang Z., Xiang Y., Bahar I., Shi Y. Llamanade: an open-source computational pipeline for robust nanobody humanization. Structure. 2022;30:418–429.e413. doi: 10.1016/j.str.2021.11.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Ramon A., Ali M., Atkinson M., Saturnino A., Didi K., Visentin C., et al. Assessing antibody and nanobody nativeness for hit selection and humanization with AbNatiV. Nat. Machine Intelligence. 2024;6:74–91. [Google Scholar]
- 104.Le Basle Y., Chennell P., Tokhadze N., Astier A., Sautou V. Physicochemical stability of monoclonal antibodies: a review. J. Pharm. Sci. 2020;109:169–190. doi: 10.1016/j.xphs.2019.08.009. [DOI] [PubMed] [Google Scholar]
- 105.Batra J., Rathore A.S. Glycosylation of monoclonal antibody products: current status and future prospects. Biotechnol. Prog. 2016;32:1091–1102. doi: 10.1002/btpr.2366. [DOI] [PubMed] [Google Scholar]
- 106.Jefferis R. Posttranslational modifications and the immunogenicity of biotherapeutics. J. Immunol. Res. 2016;2016 doi: 10.1155/2016/5358272. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107.Liu L. Antibody glycosylation and its impact on the pharmacokinetics and pharmacodynamics of monoclonal antibodies and Fc-fusion proteins. J. Pharm. Sci. 2015;104:1866–1884. doi: 10.1002/jps.24444. [DOI] [PubMed] [Google Scholar]
- 108.Haselberg R., De Vijlder T., Heukers R., Smit M.J., Romijn E.P., Somsen G.W., et al. Heterogeneity assessment of antibody-derived therapeutics at the intact and middle-up level by low-flow sheathless capillary electrophoresis-mass spectrometry. Anal. Chim. Acta. 2018;1044:181–190. doi: 10.1016/j.aca.2018.08.024. [DOI] [PubMed] [Google Scholar]
- 109.Macias L.A., Wang X., Davies B.W., Brodbelt J.S. Mapping paratopes of nanobodies using native mass spectrometry and ultraviolet photodissociation. Chem. Sci. 2022;13:6610–6618. doi: 10.1039/d2sc01536f. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110.Benazza R., Papadakos G., Thom J., Diemer H., Cotton G., Cianférani S., et al. Development of fine-tuned top-down mass spectrometry strategies in the chromatographic time scale (LC-TD-MS) for the complete characterization of an anti EGFR single domain antibody-drug conjugate (sdADC) ChemRxiv. 2024 doi: 10.26434/chemrxiv-2024-wnx5w. [preprint] [DOI] [Google Scholar]
- 111.Srzentic K., Fornelli L., Tsybin Y.O., Loo J.A., Seckler H., Agar J.N., et al. Interlaboratory study for characterizing monoclonal antibodies by top-down and middle-down mass spectrometry. J. Am. Soc. Mass Spectrom. 2020;31:1783–1802. doi: 10.1021/jasms.0c00036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 112.Wei B., Lantz C., Liu W., Viner R., Ogorzalek Loo R.R., Campuzano I.D.G., et al. Added value of internal fragments for top-down mass spectrometry of intact monoclonal antibodies and antibody-drug conjugates. Anal. Chem. 2023;95:9347–9356. doi: 10.1021/acs.analchem.3c01426. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113.Papp K.A., Weinberg M.A., Morris A., Reich K. IL17A/F nanobody sonelokimab in patients with plaque psoriasis: a multicentre, randomised, placebo-controlled, phase 2b study. Lancet. 2021;397:1564–1575. doi: 10.1016/S0140-6736(21)00440-2. [DOI] [PubMed] [Google Scholar]
- 114.Takeuchi T., Chino Y., Kawanishi M., Nakanishi M., Watase H., Mano Y., et al. Efficacy and pharmacokinetics of ozoralizumab, an anti-TNFalpha NANOBODY((R)) compound, in patients with rheumatoid arthritis: 52-week results from the OHZORA and NATSUZORA trials. Arthritis Res. Ther. 2023;25:60. doi: 10.1186/s13075-023-03036-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 115.Svecova D., Lubell M.W., Casset-Semanaz F., Mackenzie H., Grenningloh R., Krueger J.G. A randomized, double-blind, placebo-controlled phase 1 study of multiple ascending doses of subcutaneous M1095, an anti-interleukin 17A/F nanobody, in moderate-to-severe psoriasis. J. Am. Acad. Dermatol. 2019;81:196–203. doi: 10.1016/j.jaad.2019.03.056. [DOI] [PubMed] [Google Scholar]
- 116.Coppieters K., Dreier T., Silence K., de Haard H., Lauwereys M., Casteels P., et al. Formatted anti-tumor necrosis factor alpha VHH proteins derived from camelids show superior potency and targeting to inflamed joints in a murine model of collagen-induced arthritis. Arthritis Rheum. 2006;54:1856–1866. doi: 10.1002/art.21827. [DOI] [PubMed] [Google Scholar]
- 117.Roovers R.C., Laeremans T., Huang L., De Taeye S., Verkleij A.J., Revets H., et al. Efficient inhibition of EGFR signaling and of tumour growth by antagonistic anti-EFGR Nanobodies. Cancer Immunol. Immunother. 2007;56:303–317. doi: 10.1007/s00262-006-0180-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 118.Jumapili N.A., Zivalj M., Barthelmess R.M., Raes G., De Groof T.W.M., Devoogdt N., et al. A few good reasons to use nanobodies for cancer treatment. Eur. J. Immunol. 2023;53 doi: 10.1002/eji.202250024. [DOI] [PubMed] [Google Scholar]
- 119.Debie P., Lafont C., Defrise M., Hansen I., van Willigen D.M., van Leeuwen F.W.B., et al. Size and affinity kinetics of nanobodies influence targeting and penetration of solid tumours. J. Control Release. 2020;317:34–42. doi: 10.1016/j.jconrel.2019.11.014. [DOI] [PubMed] [Google Scholar]
- 120.Jumper J., Evans R., Pritzel A., Green T., Figurnov M., Ronneberger O., et al. Highly accurate protein structure prediction with AlphaFold. Nature. 2021;596:583–589. doi: 10.1038/s41586-021-03819-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121.Longsompurana P., Rungrotmongkol T., Plongthongkum N., Wangkanont K., Wolschann P., Poo-Arporn R.P. Computational design of novel nanobodies targeting the receptor binding domain of variants of concern of SARS-CoV-2. PLoS One. 2023;18 doi: 10.1371/journal.pone.0293263. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 122.Weng G., Gao J., Wang Z., Wang E., Hu X., Yao X., et al. Comprehensive evaluation of fourteen docking programs on protein-peptide complexes. J. Chem. Theor. Comput. 2020;16:3959–3969. doi: 10.1021/acs.jctc.9b01208. [DOI] [PubMed] [Google Scholar]
- 123.Schneidman-Duhovny D., Inbar Y., Nussinov R., Wolfson H.J. PatchDock and SymmDock: servers for rigid and symmetric docking. Nucleic Acids Res. 2005;33:W363–W367. doi: 10.1093/nar/gki481. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 124.Papadopoulos K.P., Harb W., Peer C.J., Hua Q., Xu S., Lu H., et al. First-in-Human phase I study of Envafolimab, a novel subcutaneous single-domain anti-PD-L1 antibody, in patients with advanced solid tumors. Oncologist. 2021;26:e1514–e1525. doi: 10.1002/onco.13817. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 125.Alqahtani M.S., Kazi M., Alsenaidy M.A., Ahmad M.Z. Advances in oral drug delivery. Front. Pharmacol. 2021;12 doi: 10.3389/fphar.2021.618411. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 126.Homayun B., Lin X., Choi H.J. Challenges and recent progress in oral drug delivery systems for biopharmaceuticals. Pharmaceutics. 2019;11 doi: 10.3390/pharmaceutics11030129. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 127.Nurbhai S., Roberts K.J., Carlton T.M., Maggiore L., Cubitt M.F., Ray K.P., et al. Oral anti-tumour necrosis factor domain antibody V565 provides high intestinal concentrations, and reduces markers of inflammation in ulcerative colitis patients. Sci. Rep. 2019;9 doi: 10.1038/s41598-019-50545-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 128.Nurbhai S. A Six Week Efficacy, Safety and Tolerability Study of V565 in Crohn’s Disease (HarbOR) ClinicalTrials.gov, National Library of Medicine; Bethesda; Bethesda, MD: 2021. [Google Scholar]
- 129.Crowe J.S., Roberts K.J., Carlton T.M., Maggiore L., Cubitt M.F., Clare S., et al. Preclinical development of a novel, orally-administered anti-tumour necrosis factor domain antibody for the treatment of inflammatory bowel disease. Sci. Rep. 2018;8:4941. doi: 10.1038/s41598-018-23277-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 130.Jester B.W., Zhao H., Gewe M., Adame T., Perruzza L., Bolick D.T., et al. Development of spirulina for the manufacture and oral delivery of protein therapeutics. Nat. Biotechnol. 2022;40:956–964. doi: 10.1038/s41587-022-01249-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 131.Van Heeke G., Allosery K., De Brabandere V., De Smedt T., Detalle L., de Fougerolles A. Nanobodies(R) as inhaled biotherapeutics for lung diseases. Pharmacol. Ther. 2017;169:47–56. doi: 10.1016/j.pharmthera.2016.06.012. [DOI] [PubMed] [Google Scholar]
- 132.Huo J., Mikolajek H., Le Bas A., Clark J.J., Sharma P., Kipar A., et al. A potent SARS-CoV-2 neutralising nanobody shows therapeutic efficacy in the Syrian golden hamster model of COVID-19. Nat. Commun. 2021;12:5469. doi: 10.1038/s41467-021-25480-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 133.Wu X., Cheng L., Fu M., Huang B., Zhu L., Xu S., et al. A potent bispecific nanobody protects hACE2 mice against SARS-CoV-2 infection via intranasal administration. Cell Rep. 2021;37 doi: 10.1016/j.celrep.2021.109869. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 134.Hart T.K., Cook R.M., Zia-Amirhosseini P., Minthorn E., Sellers T.S., Maleeff B.E., et al. Preclinical efficacy and safety of mepolizumab (SB-240563), a humanized monoclonal antibody to IL-5, in cynomolgus monkeys. J. Allergy Clin. Immunol. 2001;108:250–257. doi: 10.1067/mai.2001.116576. [DOI] [PubMed] [Google Scholar]
- 135.Cunningham S., Piedra P.A., Martinon-Torres F., Szymanski H., Brackeva B., Dombrecht E., et al. Nebulised ALX-0171 for respiratory syncytial virus lower respiratory tract infection in hospitalised children: a double-blind, randomised, placebo-controlled, phase 2b trial. Lancet Respir. Med. 2021;9:21–32. doi: 10.1016/S2213-2600(20)30320-9. [DOI] [PubMed] [Google Scholar]
- 136.Aksu M., Kumar P., Guttler T., Taxer W., Gregor K., Mussil B., et al. Nanobodies to multiple spike variants and inhalation of nanobody-containing aerosols neutralize SARS-CoV-2 in cell culture and hamsters. Antivir. Res. 2024;221 doi: 10.1016/j.antiviral.2023.105778. [DOI] [PubMed] [Google Scholar]
- 137.Nambulli S., Xiang Y., Tilston-Lunel N.L., Rennick L.J., Sang Z., Klimstra W.B., et al. Inhalable Nanobody (PiN-21) prevents and treats SARS-CoV-2 infections in Syrian hamsters at ultra-low doses. Sci. Adv. 2021;7 doi: 10.1126/sciadv.abh0319. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 138.Vu K.B., Ghahroudi M.A., Wyns L., Muyldermans S. Comparison of llama VH sequences from conventional and heavy chain antibodies. Mol. Immunol. 1997;34:1121–1131. doi: 10.1016/s0161-5890(97)00146-6. [DOI] [PubMed] [Google Scholar]
- 139.Maass D.R., Sepulveda J., Pernthaner A., Shoemaker C.B. Alpaca (Lama pacos) as a convenient source of recombinant camelid heavy chain antibodies (VHHs) J. Immunol. Methods. 2007;324:13–25. doi: 10.1016/j.jim.2007.04.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 140.Liu Y., Yi L., Li Y., Wang Z., Jirimutu Characterization of heavy-chain antibody gene repertoires in Bactrian camels. J. Genet. Genomics. 2023;50:38–45. doi: 10.1016/j.jgg.2022.04.010. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
All supporting data are provided within the manuscript, supplementary data and supplementary tables.