Abstract
Objective
To elucidate the mechanism by which tyrosine phosphatase SHP2 protects CRC through modulation of TFEB-mediated ferritinophagy, thereby suppressing ROS and ferroptosis.
Methods
SW480 and SW620 cells, in the logarithmic growth phase, were treated with or without the SHP2 inhibitor PHPS1, the activator Trichomide A, EGF, or MMP inhibitors, and randomly assigned to four groups. Additionally, SW480 cells in the logarithmic phase underwent treatments with EGF, the ferroptosis inducer erastin, Trichomide A, or the curcumin analog C1, forming seven groups. Cell migration assessment in these groups employed scratch and Transwell assays. Protein expression analysis of total SHP2, total PI3K, p-SHP2, p-PI3K, p-TFEB, TFEB, SQSTM1, LC3, LAMP2, NCOA4, FTH1, GPX4, NOX4, and ACSL4 in the seven SW480 groups was conducted through Western blot and immunofluorescence. Apoptosis analysis was performed on these seven groups, while gene co-expression analysis utilized bioinformatics. SW480 and CCD-841CoN cells were categorized into four groups, undergoing treatment with saline, EGFR-OE lentivirus, SHP2-KD lentivirus, or SHP2-OE lentivirus. Western blot analysis in SW480 cells detected EGFR, total SHP2, p-SHP2, GPX4, and ACSL4 proteins, and tumor volume observations were conducted in a nude mouse xenograft model. Western blot also evaluated total SHP2, p-SHP2, GPX4, and ACSL4 protein expression in CCD-841CoN cells.
Results
Bioinformatics analysis revealed correlations between EGFR and SHP2, SHP2 and PIK3CA, SHP2 and MAPK1, BRK4 and HIF1A, HIF1A and NCOA4, as well as TFEB and FTH1. Scratch and Transwell assays showed that SHP2 diminishes the migratory capacity of SW480 and SW620 cells. Western blot and immunofluorescence demonstrated that EGFR activation of SHP2 markedly elevated p-TFEB levels while reducing TFEB protein expression. EGF stimulation enhanced the expression of FTH1, GPX4, NOX4, and ACSL4. Combined stimulation with EGF and SHP2 further amplified the expression of p-SHP2, p-TFEB, and NCOA4 while reducing TFEB, SQSTM1, LC3, and LAMP2. Erastin augmented FTH1, GPX4, NOX4, and ACSL4 expression while decreasing p-SHP2, p-TFEB, TFEB, SQSTM1, LC3, LAMP2, and NCOA4. TFEB activation suppressed p-SHP2, p-TFEB, NCOA4, FTH1, and GPX4 expression, while promoting TFEB, SQSTM1, LC3, LAMP2, NOX4, and ACSL4 expression. Apoptosis assays indicated that SHP2 activation decelerated apoptosis in SW480 cells, whereas erastin under EGF stimulation accelerated apoptosis, as did TFEB activation. Western blot results in SW480 cells displayed that overexpression of EGFR or SHP2 significantly increased total SHP2, p-SHP2, and GPX4 expression while decreasing ACSL4 levels. SHP2 knockdown decreased total SHP2, p-SHP2, and GPX4 expression, with an increase in ACSL4 expression. In CCD-841CoN cells, overexpression of EGFR or SHP2 resulted in a decrease in p-SHP2 and an increase in total SHP2, more pronounced with SHP2 overexpression, while GPX4 and ACSL4 levels remained stable. SHP2 knockdown led to reduced EGFR, total SHP2, p-SHP2, and GPX4 expression, without a significant impact on ACSL4 levels. The nude mouse xenograft model demonstrated that EGFR overexpression significantly increased tumor size, whereas SHP2 overexpression markedly decreased tumor volume. SHP2 knockdown resulted in significantly larger tumors.
Conclusion
SHP2 advances CRC progression by modulating TFEB-mediated ferritinophagy, suppressing ROS and ferroptosis. Targeting SHP2 presents a promising therapeutic strategy for CRC.
Keywords: CRC, SHP2, TFEB, Ferritinophagy, Ferroptosis
Introduction
CRC ranks among the most prevalent cancers and presently constitutes the third leading cause of cancer-related mortality globally [1]. In its incipient stages, CRC frequently manifests without overt symptoms, and approximately 50% of patients are diagnosed at an advanced stage or with potential distant metastasis upon initial symptom presentation, with tumor metastasis persisting as a major contributor to cancer-related mortality [2]. EGFR resides on the cell membrane and influences tumor glucose metabolism via the Erk1/2 pathway [3]. In the context of cancer metastasis, cancer-associated EGFR mutants activate the PI3K/AKT pathway, thereby facilitating cell proliferation, migration, and invasion [4]. SHP2 participates in the activation of Ras/Erk under the influence of EGF and other growth factors [3]. Activating mutations in PI3K confer resistance to Ferroptosis in cancer cells, and inhibition of the PI3K-AKT-mTOR signaling axis sensitizes cancer cells to Ferroptosis induction [5]. TFEB is a critical regulator of lysosomal biogenesis, ALP, and catabolism, with the majority of TFEB protein localized on the lysosomal membrane [6]. During lysosomal stress, mTORC1 becomes inactive, ceasing phosphorylation of TFEB, while TFEB is dephosphorylated by calcineurin, enabling its translocation to the nucleus [7]. Phosphorylated p38 MAPK can translocate into the cell nucleus [8].
Iron can generate ROS through redox cycling, exerting potential cytotoxic effects such as inducing Ferroptosis [9]. Ferroptosis is an iron-dependent oxidative process resulting in significant cellular damage and diminished cell viability. Its occurrence involves the production of upstream ROS, followed by the activation of downstream Ferroptosis pathways [9]. Enzymes involved in lipid metabolism, such as ACSL4 and LPCATs, integrate free polyunsaturated fatty acids into phospholipids, leading to lipid peroxidation. The accumulation of lipid peroxides is a critical indicator of Ferroptosis [10]. Ferroptosis is initiated by three primary regulatory pathways: the GPX4 pathway, iron metabolism pathway, and lipid metabolism pathway. Inhibition of GPX4 results in the accumulation of lipid peroxides, thereby inducing Ferroptosis [10]. BRD4 is widely expressed in various cancerous and fibrotic cells [11]. Iron metabolism is crucial for Ferroptosis. Hypoxia induces the transcription of the TFR1 gene through the binding of HIF to specific promoter elements [12]. Iron enters cells by binding to TFR1 on the cell membrane, where Fe3 + is reduced to Fe2 + , participating in the generation of hydroxyl radicals (OH∙) that promote lipid peroxidation. Dysregulation of iron metabolism leads to the accumulation of free iron, thereby driving Ferroptosis [13].
TF and TFR1 serve as pivotal positive regulators of Ferroptosis [14]. Upon binding, TF and TFR1 mediate cellular iron uptake via endocytosis, facilitating the synthesis of heme, iron-sulfur clusters, or storage within ferritin [15]. Excessive activation of NCOA4-mediated Ferritinophagy has been shown to incite Ferroptosis [16]. In serum, TF binds two Fe3 + ions, and TFR1 on the cell membrane recognizes this complex [17]. Research has found that NCOA4 interacts with FTH1, mediating its autophagic degradation to release Fe2 + , a process known as ferritinophagy [18]. Excessive ferritinophagy has been confirmed to induce Ferroptosis by increasing intracellular free iron levels, thereby promoting lipid peroxidation [19]. Ferritinophagy is intimately linked to the pathogenesis of various cancers, cardiovascular diseases, and liver diseases, rendering its regulation a promising therapeutic strategy and target [20]. Altogether, the occurrence of Ferroptosis is the result of the interplay among these three pathways. Dysregulation or abnormalities in one pathway can affect the others, thereby fostering Ferroptosis. GPX4 can avert Ferroptosis by inhibiting lipid peroxidation. Iron metabolism provides the essential iron required for the lipid peroxidation process. Lipid metabolism determines whether sufficient peroxidation substrates accumulate for Ferroptosis. Current research remains inadequate, warranting further inquiry.
Methods
Bioinformatics analysis
Gene expression data were procured from the GEO database on NCBI (http://www.ncbi.nlm.nih.gov/geo/), concentrating on CRC-related datasets. Data were normalized utilizing R software (version 4.1.3) and examined for differential gene expression employing the Limma package. Volcano plots and heatmaps were constructed with a |logFC|> 1 and p < 0.05 threshold. DEGs were further scrutinized using DAVID for GO and KEGG enrichment, emphasizing biological distinctions between CRC and normal tissues. Co-expression scatter plots were generated for genes such as EGFR, PTPN11, PIK3CA, MAPK1, BRD4, HIF1α, NCOA4, TFEB, and FTH1.
Materials
Human CRC cell lines SW480 and SW620 were procured from Shanghai Biyuntian Biotechnology Co., Ltd., while the CCD-841CoN cell line was acquired from Hefei Wanwu Biotech Co., Ltd. PHPS1 was obtained from Omics Bio (Shanghai), whereas MMP, erastin, and Curcumin analog C1 were from Selleck China. Trichomide A was purchased from Bide Pharmatech (Shanghai). SPF-grade C57BL/6 J male mice, weighing 20 ± 4 g, were sourced from Henan Sikebase Biotech. This study was reviewed and approved by the Medical Ethics Committee of the 980 Hospital of the Joint Service Support Force of the Chinese People's Liberation Army (2024-KY-87).
Cell culture and grouping
SW480 and CCD-841CoN cells were routinely cultured in modified glucose-free RPMI-1640 medium supplemented with 10% FBS and 1% penicillin/streptomycin at 37 °C in a 5% CO₂ atmosphere. The medium was replaced every three days. When cell confluence reached approximately 80%, cells were digested with 0.25% trypsin and passaged. Cells in the logarithmic growth phase were utilized for subsequent experiments. SW620 cells were cultured under conditions similar to those of SW480 cells.
Logarithmic phase SW480 and SW620 cells
Logarithmic phase SW480 and SW620 cells were randomly divided into four equal groups and treated as follows:
• DMSO Group (Group I): Treated with physiological saline equivalent in volume to the other groups as a control. The final DMSO concentration remained below 0.1%.
• EGF-MMP-DMSO Group (Group II): Treated with EGF and MMP.
• EGF-MMP-PHPS1 Group (Group III): Treated with EGF, MMP, and the SHP2 inhibitor PHPS1.
• EGF-MMP-Trichomide A Group (Group IV): Treated with EGF, MMP, and the SHP2 activator Trichomide A.
EGF was applied at a concentration of 50 ng/mL, MMP at 5 μg/mL, PHPS1 at 20 μM, and Trichomide A at 20 μM. Cells were co-cultured in serum-free medium for 48 h following treatment.
Logarithmic phase SW480 cells were randomly divided into the following seven groups:
• NC Group (Group a): Treated with physiological saline as a control.
• EGF-NC Group (Group b): Treated with EGF (50 ng/mL) under hypoxic conditions (1% O₂) for 48 h.
• EGF-Trichomide A Group (Group c): Treated with EGF and Trichomide A.
• EGF-Fe-NC Group (Group d): Treated with EGF and the ferroptosis inducer erastin.
• EGF-Fe-Trichomide A Group (Group e): Treated with EGF, erastin, and Trichomide A.
• EGF-Fe-Curcumin analog C1 Group (Group f): Treated with EGF, erastin, and the TFEB activator Curcumin analog C1.
• EGF-Fe-Trichomide A-Curcumin analog C1 Group (Group g): Treated with EGF, erastin, Trichomide A, and Curcumin analog C1.
For Groups b to g, EGF was used at 50 ng/mL, erastin at 10 μM, Trichomide A at 20 μM, and Curcumin analog C1 at 20 μM under hypoxic conditions (1% O₂) for 48 h. All cells were treated under different conditions for 48 h.
Logarithmic phase SW480 colon cancer cells and CCD-841CoN normal colon cells
SW480 and CCD-841CoN cells were allocated into four groups and subjected to treatment with or without physiological saline, EGFR-OE lentivirus, SHP2-KD lentivirus, or SHP2-OE lentivirus under hypoxic conditions (1% O₂) for 48 h.
SW480 cells were treated as follows:
• SW480-NC Group (Group ①): Treated with physiological saline.
• SW480-EGFR-OE Group (Group ②): Treated with EGFR-OE lentivirus.
• SW480-SHP2-KD Group (Group ③): Treated with SHP2-KD lentivirus.
• SW480-SHP2-OE Group (Group ④): Treated with SHP2-OE lentivirus.
CCD-841CoN cells were treated as follows:
• CCD-841CoN-NC Group (Group ⑤): Treated with physiological saline.
• CCD-841CoN-EGFR-OE Group (Group ⑥): Treated with EGFR-OE lentivirus.
• CCD-841CoN-SHP2-KD Group (Group ⑦): Treated with SHP2-KD lentivirus.
• CCD-841CoN-SHP2-OE Group (Group ⑧): Treated with SHP2-OE lentivirus.
Lentivirus construction for SW480 cells treated with EGFR-OE lentivirus:
EGFR-OE lentivirus was constructed by co-transfecting EGFR-OE plasmids with packaging plasmids into 293 T cells. Culture supernatant was collected 48–72 h later, centrifuged to remove cell debris, and concentrated to obtain the virus. SW480 cells were plated and infected with diluted lentivirus (40 μL of viral stock and 400 μL of medium containing 5 μg/mL Polybrene), with an MOI of 10–20, for 24 h. After infection, the medium was replaced for subsequent screening and detection.
Lentivirus construction for SW480 cells treated with SHP2-KD lentivirus:
The SHP2-KD lentivirus was created by co-transfecting SHP2-KD plasmids with packaging plasmids into 293 T cells. After viral particle collection and concentration, SW480 cells were plated and infected as described above, with an MOI of 10–20. Upon overnight infection, the virus was removed, and the cells were subjected to puromycin selection and KD effect analysis.
Lentivirus construction for SW480 cells treated with SHP2-OE lentivirus:
SHP2-OE lentivirus particles were prepared using the same packaging and infection protocol. Post-infection, stable SHP2-OE cell lines were obtained and verified by fluorescence protein expression, mRNA, and protein levels.
Lentivirus construction for CCD-841CoN and SW480 cells:
The lentivirus construction method was analogous for CCD-841CoN and SW480 cells. However, as CCD-841CoN cells are non-cancerous, they may exhibit lower infection efficiency or necessitate higher Polybrene concentrations (5–10 μg/mL) for successful infection. Infection efficiency and cell health were meticulously monitored to ensure successful viral infection and stable cell conditions.
Wound healing assay
The wound healing assay was employed to evaluate the migratory capacity of SW480 and SW620 cells in Groups I–IV. Lines were meticulously drawn across the back of 6-well plates using a marker, intersecting each well. Approximately 5 × 105 cells were seeded per well, ensuring full confluence overnight. A scratch was made in the cell layer the following day using a pipette tip or sterile toothpick. After washing with sterile PBS three times to remove detached cells, fresh serum-free medium was added. Cells were incubated at 37 °C with 5% CO₂ for 48 h. The scratch width was measured via microscopy, and images were analyzed with ImageJ software.
Transwell assay
The Transwell assay was utilized to assess cell migration in SW480 and SW620 cells from each group. Matrigel-coated Transwells were used, and 2 × 104 cells per well were introduced to the upper chamber. The lower chamber was filled with 600 μL of medium containing FBS, and cells were incubated at 37 °C with 5% CO₂ for 12 h. Cells in the upper chamber were removed, and cells in the lower chamber were fixed with 4% paraformaldehyde and stained with 0.1% crystal violet. Migrated cells were enumerated under an inverted fluorescence microscope, and the assay was repeated three times.
Xenograft tumor formation assay in nude mice
SW480 cells from Groups ①-④ were adjusted to a concentration of 1 × 10⁶ to 5 × 10⁶ cells/mL and resuspended in sterile PBS. Nude mice were randomly divided into two cages, with six mice per group. Each mouse was injected with 100 μL of cell suspension (1 × 10⁶ cells) into the groin area. Post-injection, mice were acclimatized to laboratory conditions for 1 week, with regular monitoring of health and tumor growth. Tumor volume was measured every 2 days using a caliper and calculated as (volume = 1/2 × length × width2). Mice were euthanized using 3% isoflurane, and tumors were collected for subsequent Western blot analysis.
Apoptosis detection using annexin V-FITC staining
Apoptosis in Groups a-g was evaluated using Annexin V-FITC staining. SW480 cells were gently dissociated into single cells using 0.25% trypsin–EDTA, washed with PBS, fixed, and incubated with antibodies before analysis using CytoFLEX. Data were analyzed with CytExpert software.
Immunofluorescence staining
Immunofluorescence staining was employed to detect NOX4 and GPX4 protein expression in SW480 cells from Groups a-g. Cells were fixed with 4% paraformaldehyde, blocked, and incubated with primary and secondary antibodies. Images were captured using an Olympus 1X73 fluorescence microscope, and quantitative analysis was carried out using ImageJ.
Western blot analysis
Western blot analysis was performed on SW480 cells from Groups I–IV to assess protein expression of p-SHP2, p-PI3K, BRD4, p-TFEB, TFEB, Cathepsin L, and Cathepsin B. Additionally, cells from Groups a-g were analyzed for p-SHP2, p-TFEB, TFEB, SQSTM1, LC3, LAMP2, NCOA4, FTH1, GPX4, NOX4, and ACSL4 expression. Protein analysis was similarly conducted on SW480 cells from Groups ①-④ and CCD-841CoN cells from Groups ⑤-⑧.
Statistical analysis
Statistical analysis was conducted using GraphPad Prism 5.0 software, with all data expressed as mean ± SD. Group comparisons were analyzed using ANOVA. Further validation of statistical significance was performed using SPSS 26.0, with P < 0.05 considered statistically significant and P < 0.01 considered highly significant.
Results
Bioinformatics analysis of pathways and differential gene correlations in colorectal cancer
We explored the GEO database for datasets pertinent to Colorectal Cancer and acquired the CRC-related mRNA expression dataset GPL17586 (based on [HTA-2_0] Affymetrix Human Transcriptome Array 2.0 [transcript (gene) version]). RNA-seq data from GSE100179 were quantile normalized using the R package limma, both pre- and post-normalization. The dataset, comprising 20 normal samples and 20 cancerous samples, was selected for differential gene analysis (|logFC|> 1, p-value < 0.05). A volcano plot illustrating DEGs was constructed using ggplot2 in R, and a heatmap of the top 100 DEGs was generated using the pheatmap package in R.
GOplot and ggplot2 packages in R were employed to visualize the top 10 upregulated DEGs in BP, CC, and MF categories in GO pathway enrichment plots and chord diagrams, demonstrating enrichment in biological processes such as apoptotic process, innate immune response, and inflammatory response. The top 10 downregulated DEGs in BP, CC, and MF categories were enriched in immune response, adaptive immune response, and immunoglobulin-mediated immune response. KEGG enrichment analysis revealed significant enrichment in pathways including the PI3K-Akt signaling pathway and Cytokine-cytokine receptor interaction.
Correlation scatter plots were generated for dataset GSE100179, and Pearson correlation coefficients were calculated to evaluate gene correlations. We observed that EGFR suppresses PTPN1 expression, PTPN11 promotes PIK3CA and MAPK1 expression, BRK4 inhibits HIF1Α, HIF1Α inhibits NCOA4, and TBEB inhibits FTH1 expression (Fig. 1).
Fig. 1.
Bioinformatics analysis of pathways and differential gene correlations in colorectal cancer. A Boxplot of raw data from the GSE100179 dataset. B Boxplot of normalized data from the GSE100179 dataset. C Volcano plot of differential genes in the GSE100179 dataset. D Heatmap of differential genes in the GSE100179 dataset. E Bar plot of GO enrichment analysis for upregulated genes. F Chord diagram of GO enrichment analysis for upregulated genes. G Bar plot of GO enrichment analysis for downregulated genes. H Chord diagram of GO enrichment analysis for downregulated genes. I Bubble chart of KEGG enrichment analysis for differential genes. J Correlation scatter plot between EGFR and PTPN11. K Correlation scatter plot between PTPN11 and PIK3CA. L Correlation scatter plot between PTPN11 and MAPK1. M Correlation scatter plot between BRK4 and HIF1A. N Correlation scatter plot between HIF1A and NCOA4. O Correlation scatter plot between TFEB and FTH1
EGFR activates SHP2 leading to increased p-TFEB expression and suppression of TFEB protein expression
Western blot analysis of Groups I–IV revealed that, upon EGF stimulation and MMP broad-spectrum inhibitor treatment, protein levels in Group II were elevated compared to Group I. This suggests that EGF stimulation and MMP inhibition activate EGFR in SW480 cells, recruiting p-SHP2, and resulting in increased expression of p-PI3K, BRD4, p-TFEB, TFEB, Cathepsin L, and Cathepsin B proteins, with significant differences noted. In Group III, the addition of the SHP2 inhibitor PHPS1 under EGF stimulation and MMP inhibition led to decreased p-SHP2, p-PI3K, BRD4, and p-TFEB activities compared to Group II, while TFEB, Cathepsin L, and Cathepsin B activities increased, demonstrating notable differences. Group IV, treated with the SHP2 activator Trichomide A under EGF stimulation and MMP inhibition, exhibited increased p-SHP2, p-PI3K, BRD4, and p-TFEB activities and decreased TFEB, Cathepsin L, and Cathepsin B activities compared to Group II, indicating that EGFR activation of SHP2 promotes p-TFEB expression and inhibits TFEB protein expression (Fig. 2A and B).
Fig. 2.
EGFR activation results in elevated p-TFEB expression and suppression of the TFEB protein. A Protein expression levels of p-SHP2, p-PI3K, BRD4, p-TFEB, TFEB, Cathepsin L, and Cathepsin B in each group. B Compared to Group I, all proteins in Group II are upregulated. In Group III, p-SHP2, p-PI3K, BRD4, and p-TFEB activities are decreased compared to Group II, whereas TFEB, Cathepsin L, and Cathepsin B activities are increased, showing significant differences. In Group IV, p-SHP2, p-PI3K, BRD4, and p-TFEB activities are elevated compared to Group II, while TFEB, Cathepsin L, and Cathepsin B activities are reduced, denoting significant differences. (*P < 0.05; **P < 0.01)
Wound healing assay demonstrates that SHP2 modulates migration of SW480 and SW620 cells
At 0 h, there were no differences between cells. After 48 h of culturing SW480 and SW620 cells, Group I displayed minimal migration. Group II, treated with an MMP broad-spectrum inhibitor, exhibited narrower wound gaps compared to Group I, indicating enhanced migration. In Group III, the addition of an SHP2 inhibitor to the MMP broad-spectrum inhibitor resulted in even narrower wound gaps compared to Group II, indicating stronger migration. In Group IV, addition of an SHP2 activator to the MMP inhibitor led to wider wound gaps compared to Group II, indicating reduced migration. Comparisons between Groups III, IV, and II suggest that SHP2 downregulates the migration ability of SW480 and SW620 cells (Fig. 3A).
Fig. 3.
A Scratch assay indicates that SHP2 downregulates the migratory capacity of SW480 and SW620 cells. B Transwell assay demonstrates that SHP2 attenuates the migratory capacity of SW480 and SW620 cells. (*P < 0.05; **P < 0.01)
Transwell assay shows that SHP2 reduces the migration of SW480 and SW620 cells
After culturing SW480 and SW620 cells in the upper chamber, Transwell results indicated that Group I had few cells. Group II, treated with an MMP broad-spectrum inhibitor, exhibited increased cell numbers compared to Group I, suggesting enhanced migration. In Group III, the addition of an SHP2 inhibitor to the MMP inhibitor further increased cell numbers compared to Group II, indicating that SHP2 inhibition augments the migration of SW480 and SW620 cells. In Group IV, the addition of an SHP2 activator to the MMP inhibitor reduced cell numbers compared to Group II, suggesting that SHP2 activation diminishes cell migration. Comparisons among Groups III, IV, and II confirm that SHP2 diminishes the migratory capacity of SW480 and SW620 cells (Fig. 3B).
Combined EGF and SHP2 activation promotes p-SHP2, p-TFEB, and NCOA4 protein expression while suppressing TFEB, SQSTM1, LC3, and LAMP2 protein expression; erastin inhibits p-SHP2, p-TFEB, TFEB, SQSTM1, LC3, LAMP2, and NCOA4 protein expression; TFEB activation inhibits p-SHP2, p-TFEB, and NCOA4 protein expression while promoting TFEB, SQSTM1, LC3, and LAMP2 protein expression
Western blot analysis of groups a-g under 1% hypoxia and low serum conditions showed that EGF stimulation in group b compared to group a led to the activation of EGFR, resulting in increased expression of p-SHP2, p-TFEB, TFEB, SQSTM1, LC3, LAMP2, and NCOA4, with significant differences. Group c, treated with the SHP2 activator Trichomide A, exhibited increased expression of p-SHP2, p-TFEB, and NCOA4, and decreased TFEB, SQSTM1, LC3, and LAMP2, indicating that SHP2 activation following EGF stimulation promotes p-SHP2, p-TFEB, and NCOA4 expression while inhibiting TFEB, SQSTM1, LC3, and LAMP2 expression. In Group d, the addition of erastin following EGF stimulation in group b reduced expression of p-SHP2, p-TFEB, TFEB, SQSTM1, LC3, LAMP2, and NCOA4. Similarly, group e, compared to group c, also exhibited decreased expression of these proteins upon erastin treatment, suggesting that erastin inhibits these proteins under EGF stimulation. In Group e, the addition of SHP2 activator Trichomide A following EGF and erastin stimulation increased p-SHP2, p-TFEB, and NCOA4 expression while decreasing TFEB, SQSTM1, LC3, and LAMP2 expression. Group f, compared to group d, showed decreased p-SHP2, p-TFEB, and NCOA4 and increased TFEB, SQSTM1, LC3, and LAMP2 upon TFEB activation with Curcumin analog C1. In Group g, the addition of TFEB activator to group e significantly decreased p-SHP2, p-TFEB, and NCOA4 expression while significantly increasing TFEB, SQSTM1, LC3, and LAMP2 expression. Comparisons between groups f-d and g-e indicate that under EGF and erastin stimulation, TFEB activation suppresses p-SHP2, p-TFEB, and NCOA4 while promoting TFEB, SQSTM1, LC3, and LAMP2 expression (Fig. 4A).
Fig. 4.
A Protein expression levels of p-SHP2, p-TFEB, TFEB, SQSTM1, LC3, LAMP2, and NCOA4 in each group. Combined EGF and SHP2 stimulation augments the expression of p-SHP2, p-TFEB, and NCOA4 while inhibiting TFEB, SQSTM1, LC3, and LAMP2 expression. Erastin inhibits the expression of p-SHP2, p-TFEB, TFEB, SQSTM1, LC3, LAMP2, and NCOA4. TFEB activation suppresses p-SHP2, p-TFEB, and NCOA4 expression while promoting TFEB, SQSTM1, LC3, and LAMP2 expression. B SW480 cell apoptosis assay reveals that SHP2 activation decelerates apoptosis, whereas erastin under EGF stimulation accelerates apoptosis, and TFEB activation further accelerates apoptosis. (*P < 0.05; **P < 0.01)
SW480 cell apoptosis assay shows that SHP2 activation slows SW480 cell apoptosis; EGF stimulation with erastin accelerates SW480 cell apoptosis; TFEB activation accelerates SW480 cell apoptosis
In group a, cells exhibited a high level of apoptosis. Compared to group a, group b demonstrated reduced apoptosis in SW480 cells upon EGF stimulation, indicating that EGF mitigates apoptosis in SW480 cells. Group c, with SHP2 activation relative to group b, showed even fewer apoptotic cells, suggesting that SHP2 activation decelerates apoptosis. Group d, compared to group b with erastin treatment, exhibited increased apoptosis, and group e, relative to group c with erastin treatment, also demonstrated increased apoptosis. Comparisons between groups b-d and c-e imply that erastin treatment under EGF stimulation accelerates apoptosis in SW480 cells. Group f, compared to group d with TFEB activation, showed increased apoptosis, as did group g compared to group e, indicating that TFEB activation expedites SW480 cell apoptosis (Fig. 4B).
Western blot and immunofluorescence results show that EGF and/or erastin stimulate ferroptosis-related protein expression; TFEB activation inhibits FTH1 and GPX4 while promoting NOX4 and ACSL4 expression
In group a, without any stimulation, the expression of FTH1, GPX4, NOX4, and ACSL4 proteins was weak. In group b, EGF stimulation enhanced the expression of all these proteins, indicating that EGF promotes Ferroptosis-related protein expression. Compared to group b, group c with SHP2 activation showed increased expression of FTH1 and GPX4 and decreased expression of NOX4 and ACSL4, suggesting that SHP2 activation augments FTH1 and GPX4 while inhibiting NOX4 and ACSL4. Both groups d and e showed increased expression of these proteins with erastin stimulation, confirming that erastin enhances Ferroptosis-related protein expression. Group e, compared to group d under EGF and erastin stimulation with SHP2 activation, exhibited increased expression of FTH1 and GPX4 and decreased expression of NOX4 and ACSL4. In group f, TFEB activation compared to group d reduced the expression of FTH1 and GPX4 and increased the expression of NOX4 and ACSL4, and group g, compared to group e, showed significant reduction in FTH1 and GPX4 and a significant increase in NOX4 and ACSL4 expression, indicating that TFEB activation inhibits FTH1 and GPX4 and promotes NOX4 and ACSL4 (Fig. 5A and B).
Fig. 5.
A Protein expression levels of FTH1, GPX4, NOX4, and ACSL4 in each group. Western blot results indicate that EGF and erastin stimulation enhance ferroptosis-related protein expression. TFEB activation inhibits FTH1 and GPX4 expression while promoting NOX4 and ACSL4 expression. B Immunofluorescence results demonstrate that EGF and erastin stimulation augment GPX4 and NOX4 expression, while TFEB activation inhibits GPX4 expression and enhances NOX4 expression. (*P < 0.05; **P < 0.01)
Western blot results show that overexpression of EGFR or SHP2 in SW480 cells significantly increases total SHP2, p-SHP2, and GPX4 expression while decreasing ACSL4 expression; SHP2 knockdown reduces total SHP2, p-SHP2, and GPX4 expression while increasing ACSL4 expression
In SW480 cells, group ① exhibited weak expression of total SHP2, p-SHP2, and GPX4 and moderate ACSL4 expression without any stimulation. In group ②, EGFR overexpression compared to group ① enhanced total SHP2 and p-SHP2 self-phosphorylation, resulting in increased total SHP2, p-SHP2, and GPX4 expression and reduced ACSL4 expression, with significant differences. In group ③, SHP2 knockdown compared to group ① resulted in reduced total SHP2, p-SHP2, and GPX4 expression and increased ACSL4 expression, with significant differences. Group ④, with SHP2 overexpression in comparison to group ①, showed significantly increased total SHP2, p-SHP2, and GPX4 expression and significantly decreased ACSL4 expression (Fig. 6A).
Fig. 6.
A Western blot results in SW480 cells show that in Group ①, without stimulation, total SHP2, p-SHP2, and GPX4 expression levels are low, while ACSL4 expression is moderate. In Group ②, EGFR overexpression significantly elevates total SHP2, p-SHP2, and GPX4 expression, whereas ACSL4 expression decreases. In Group ③, SHP2 knockdown markedly decreases total SHP2, p-SHP2, and GPX4 expression, while ACSL4 expression increases. In Group ④, SHP2 overexpression significantly enhances total SHP2, p-SHP2, and GPX4 expression while reducing ACSL4 expression. (*P < 0.05; **P < 0.01) B Western blot results in CCD-841CoN cells show that in Group ⑤, without stimulation, EGFR, total SHP2, p-SHP2, GPX4, and ACSL4 expression levels are low. In Group ⑥, EGFR overexpression decreases EGFR and p-SHP2 expression but increases total SHP2 expression, with no significant changes in GPX4 and ACSL4 levels. In Group ⑦, SHP2 knockdown increases EGFR and p-SHP2 expression, decreases total SHP2 expression, and significantly lowers GPX4 expression, with no significant changes in ACSL4. In Group ⑧, SHP2 overexpression reduces EGFR and p-SHP2 expression, increases total SHP2 expression, but causes no significant changes in GPX4 and ACSL4 expression. (*P < 0.05; **P < 0.01). C Tumor growth experiments in nude mice demonstrate that EGFR overexpression significantly increases tumor volume, while SHP2 knockdown also enlarges tumor size. SHP2 overexpression, however, reduces tumor volume. Compared to Group ①, tumor volumes in Groups ②, ③, and ④ are significantly larger. In contrast, Group ③ (with SHP2 inhibition) exhibits larger tumors than Group ② (with EGFR overexpression), while Group ④ (with SHP2 overexpression) shows a significant reduction in tumor size. (*P < 0.05; **P < 0.01)
Western blot results show that EGFR or SHP2 overexpression in CCD-841CoN cells downregulates p-SHP2 and EGFR expression while upregulating total SHP2 expression; SHP2 overexpression shows greater differences than EGFR overexpression; GPX4 and ACSL4 show no significant changes; SHP2 knockdown reduces EGFR, Total SHP2, p-SHP2, and GPX4 expression while ACSL4 shows no significant change
In group ⑤, epithelial cells of human normal colon tissue CCD-841CoN demonstrated low expression of EGFR, total SHP2, p-SHP2, GPX4, and ACSL4 without any stimulation. In group ⑥, EGFR overexpression compared to group ⑤ led to reduced EGFR and p-SHP2 expression, with significant differences; total SHP2 expression increased without significant difference; GPX4 and ACSL4 showed no significant change. In group ⑦, SHP2 knockdown compared to group ⑤ enhanced EGFR and p-SHP2 expression, with significant differences; total SHP2 expression decreased with significant differences; GPX4 expression decreased with significant differences; and ACSL4 expression showed no significant change. In group ⑧, SHP2 overexpression downregulated EGFR and p-SHP2 expression with significant differences; total SHP2 expression increased with significant differences; and GPX4 and ACSL4 exhibited no significant change (Fig. 6B).
Tumor formation assay in nude mice shows that EGFR overexpression increases tumor volume, SHP2 knockdown significantly increases tumor volume, and SHP2 overexpression reduces tumor volume
In group ②, the tumor volume was larger compared to group ①; in group ③, the tumor volume was significantly larger than in group ①; and in group ④, tumor volume increased compared to group ①. Group ③, with SHP2 inhibition relative to group ② with EGFR overexpression, exhibited increased tumor volume; group ④, with SHP2 overexpression compared to group ② with EGFR overexpression, demonstrated reduced tumor volume (Fig. 6C).
Discussion
Ferroptosis prominently induces cell death through iron-dependent lipid peroxidation, distinct from traditional apoptotic mechanisms. It involves ROS accumulation and the collapse of antioxidant defense systems, particularly disrupting GPX4 and lipid metabolism pathways [21, 22]. Tumor suppressor genes such as p53 can trigger ferroptosis, thereby inhibiting tumor progression. Conversely, cancer cells upregulate antioxidant systems like SLC7A11 and GPX4 to counteract oxidative stress [23, 24]. As an emerging cell death mechanism, ferroptosis is intricately linked with alterations in intracellular iron metabolism and signaling pathways [25, 26].
EGF is the primary ligand for EGFR and plays a pivotal role in various aspects of cancer cell metastasis [27]. SHP2 functions in mammals by facilitating signal transduction through its phosphatase activity and also acts as an adaptor protein for signal mediation [28]. In this study, upon EGF binding to EGFR, SHP2 is recruited [29]. The GTPase Ras serves as a key mediator of signal transduction for receptor tyrosine kinases [30]. Evidence suggests that Gab1 is a substrate for SHP2, implying this interaction may participate in Ras activation [3]. In this study, SHP2 activates Ras. Ras can stimulate several effectors, notably the downstream mitogen-activated protein kinases Erk1/2, which are major regulators of cell proliferation, differentiation, and survival [31]. Upon EGF stimulation, SHP2 is swiftly recruited to the complex with activated EGFR and undergoes phosphorylation [32]. SHP2 is essential for the full activation of the RAS-ERK pathway in receptor tyrosine kinase and cytokine receptor signaling, acting as a positive regulator of RAS signaling [33]. In this study, SHP2 facilitates RAS activation of p-P38 and Erk1/2. The ERK1/2 and p38 pathways are canonical MAPK activation pathways [34]. The combination of EGFR and p38 MAPK may optimize the therapeutic potential of EGFR in cancer [8]. HIF binds specific promoter elements to induce TFR1 gene transcription [12]. HIF1α promotes high expression of NCOA4 [35], while BRD4 inhibits TFEB [36]. Activated Ras further promotes HIF binding to specific promoters, inducing TFR1 transcription and elevating NCOA4 expression. PI3K, as the regulatory subunit p85, can bind the EGFR growth factor receptor, modifying the structure of the Akt protein [37]. The PI3K-mTORC1 signaling network orchestrates cellular metabolism and growth [38]. Lysosomal mTORC1 integrates upstream PI3K and amino acid-dependent signals [39]. The classic mode of TFEB activation is: under nutrient-rich conditions, mTORC1 enhances phosphorylation of TFEB, preventing its nuclear translocation, while under nutrient-deprived conditions, mTORC1 phosphorylation activity diminishes, allowing dephosphorylated TFEB to translocate into the nucleus [40, 41]. SHP2 activates the PI3K/AKT signaling pathway, subsequently inhibiting TSC2/mTORC1, promoting TFEB phosphorylation, and indirectly enhancing p-TFEB.
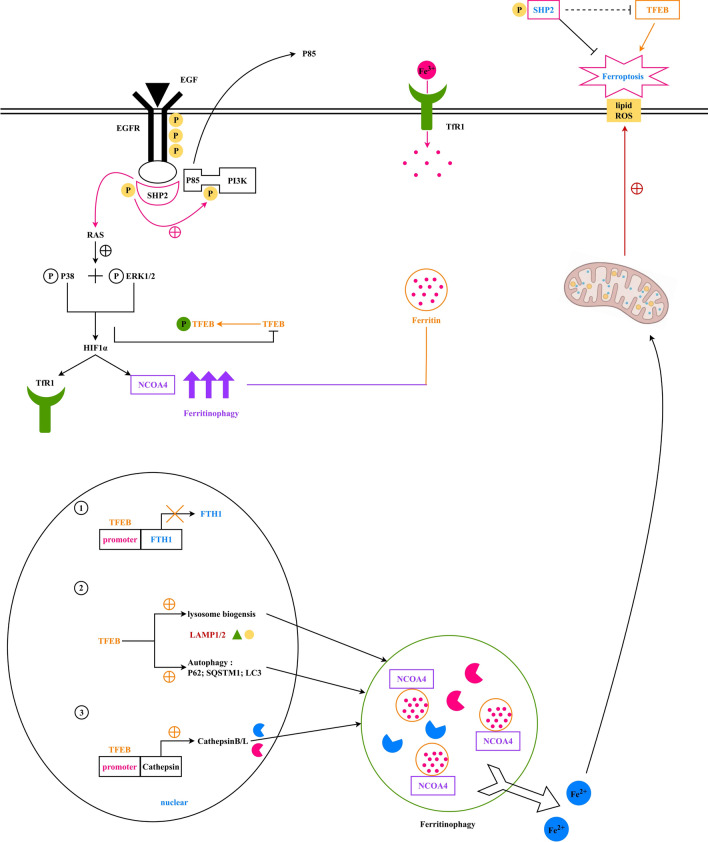
Ferritin is a pivotal intracellular iron storage protein composed of FTL and FTH1, playing an essential role in host defense against oxidative stress [18]. When TFR1 expression on the cell membrane is enhanced, more TF is recognized and binds, increasing Fe3 + within cellular endosomes [42]. NCOA4 efficiently regulates ferritin levels via ferritinophagy to maintain systemic iron balance; when the body is iron-deficient, NCOA4 levels rise, promoting ferritin degradation and releasing Fe2 + [43]. Ferritinophagy facilitates the release of Fe2 + , leading to increased intracellular free iron concentration, causing iron overload and ultimately promoting ferroptosis [44]. Common ferroptosis biomarkers include GPX4, FSP1, COX2, ACSL4, and FTH1 [45]. LIP in lysosomes actively sequesters free cytosolic iron, rendering lysosomes the primary organelles inducing ferroptosis [46]. Our study indicates that TFEB promotes ferritinophagy and lysosome biogenesis. Ferritin is delivered to lysosomes for degradation through ferritinophagy, with autophagy receptors including SQSTM1, p62, and LC3, among which LC3 is a classical autophagy marker [47]. Research has shown that NCOA4 specifically interacts with LC3 only when SQSTM1 is present [47]. In this study, erastin stimulation promotes NCOA4 expression and its binding to ferritin, leading to the release of Fe2 + , which triggers mitochondrial oxidative stress, generates a substantial amount of ROS, and promotes ferritinophagy (Fig. 7).
Fig. 7.
Upon EGFR stimulation, SHP2 promotes p-TFEB via PI3K/BRD4, indirectly inhibiting TFEB. Subsequently, TFEB promotes LAMP2 and LC3 in lysosome synthesis and SQSTM1 in ferritinophagy, enhancing NCOA4’s interaction with FTH1, GPX4, NOX4, and ACSL4, which ultimately leads to iron overload and triggers ferroptosis
Conclusion
In summary, upon EGFR stimulation, SHP2 promotes p-TFEB via PI3K/BRD4 pathways, indirectly inhibiting TFEB. Consequently, TFEB enhances LAMP2 and LC3 for lysosomal synthesis and SQSTM1 for ferritinophagy, promoting NCOA4 interaction with FTH1, GPX4, NOX4, and ACSL4, leading to iron overload and ultimately ferroptosis. This study demonstrates that SHP2 regulates TFEB-mediated ferritinophagy to inhibit ROS and ferroptosis, providing protection against CRC invasion and metastasis, offering significant potential for clinical applications.
Acknowledgements
None.
Abbreviations
- ACSL4
Acyl-CoA Synthetase Long-Chain Family Member 4
- AKT
Protein Kinase B
- BRD4
Bromodomain-containing protein 4
- CRC
Colorectal cancer
- DAVID
Database for Annotation, Visualization and Integrated Discovery Database
- EGF
Epidermal growth factor
- EGFR
Epidermal growth factor receptor
- Erk
Extracellular signal-regulated kinase
- FTH1
Ferritin Heavy Chain
- FTL
Ferritin Light Chain
- Gab1
Grb2-associated binding protein 1
- GO
Gene Ontology
- GPX4
Glutathione Peroxidase 4
- HIF
Hypoxia-Inducible Factor
- HIF1α
Hypoxia-Inducible Factor 1 alpha
- KEGG
Kyoto Encyclopedia of Genes and Genomes
- LAMP2
Lysosome-associated membrane protein 2
- LC3
Microtubule-associated protein 1 light chain 3
- LIP
Labile ironpool
- MAPK
Mitogen-Activated Protein Kinase
- MAPK1
Mitogen-Activated Protein Kinase 1
- MMP
Matrix Metalloproteinase
- mTORC1
Mechanistic Target of Rapamycin Complex 1
- NCBI
National Center for Biotechnology Information
- NCOA4
Nuclear Receptor Coactivator 4
- NOX4
NADPH Oxidase 4
- p38
P38 Mitogen-Activated Protein Kinase
- p85
Phosphoinositide 3-kinase regulatory subunit 1
- PHPS1
PH domain and leucine-rich repeat-containing protein 1
- PI3K
Phosphoinositide 3-Kinase
- PIK3CA
Phosphatidylinositol-4,5-Bisphosphate 3-Kinase Catalytic Subunit Alpha
- PTPN11
Protein Tyrosine Phosphatase Non-Receptor Type 11
- Ras
Rat Sarcoma
- ROS
Reactive oxygen species
- SHP2
SH2-containing Protein Tyrosine Phosphatase 2
- SQSTM1
Sequestosome 1
- TF
Transferrin
- TFEB
Transcription Factor EB
- TFR1
Transferrin receptor 1
Author contributions
J. C. And W. Captain. The Experiment Protocol was designed, SW480 and SW620 cells were treated, and the effects of different drugs and stimuli on cell migration, protein expression and apoptosis were observed. C. Noah. And D. H. Captain. Cell Scratch Test, Transwell Test, Western blot test and immunofluorescence test were carried out to analyze the changes of cell migration, protein expression and apoptosis. J. C. ANDC. J. Bioinformatics analysis revealed correlations between EGFR and SHP2, Shp2 and PIK3CA, SHP2 and MAPK1, and improved understanding of signaling pathways. J. C. The results showed that EGFR activated SHP2 to promote p-TFEB expression, and EGF stimulated FTH1, GPX4, NOX4, ACSL4 expression. J. C. It was concluded that the phosphatase Shp2 induces iron death by regulating the PI3KBRD4 TFEB signaling pathway, thereby promoting the development of colorectal cancer. In summary, the five authors revealed the mechanism of the role of SHP2 in CRC progression through design experiments, experimental manipulation, data analysis, and conclusion summary, it provides an important reference for the treatment and prevention of tumor.
Funding
This study was supported by the 2022 Hebei Medical Science Research Project (20220231): the effect of ileocolectomy on blood glucose level in cardiac cancer patients with type II diabetes mellitus. This project is supported by the Incubation Program of the 980th Hospital of the Joint Logistics Support Force (2024FYMSXM06, 2024FYQNXM01).
Data availability
The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
Declarations
Ethics approval and consent to participate
This study has been People’s Liberation Army by the Medical Ethics Committee of the 980 Hospital of the Joint Logistics Support Unit. Medical Ethics Committee of the 98O Hospital of the Joint Logistics Support Force of the People's Liberation Army of China (2024-KY-87).
Competing interests
The authors declare no competing interests.
Footnotes
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
- 1.Sazin Y, et al. Association between adherence to a dietary approach to stop hypertension and the Mediterranean diets and risk of colorectal cancer: a matched case-control study. Clinical Nutrition ESPEN. 2024;60:195–202. [DOI] [PubMed] [Google Scholar]
- 2.Ciardiello F, et al. Clinical management of metastatic colorectal cancer in the era of precision medicine. CA Cancer J Clin. 2022;72(4):372–401. [DOI] [PubMed] [Google Scholar]
- 3.Herbst R, et al. Recruitment of the protein tyrosine phosphatase CSW by DOS is an essential step during signaling by the sevenless receptor tyrosine kinase. Embo j. 1999;18(24):6950–61. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Tong D, et al. LAMC2 promotes EGFR cell membrane localization and acts as a novel biomarker for tyrosine kinase inhibitors (TKIs) sensitivity in lung cancer. Cancer Gene Ther. 2023;30(11):1498–512. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Yi J, et al. Oncogenic activation of PI3K-AKT-mTOR signaling suppresses ferroptosis via SREBP-mediated lipogenesis. Proc Natl Acad Sci U S A. 2020;117(49):31189–97. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Sarkar C, et al. PLA2G4A/cPLA2-mediated lysosomal membrane damage leads to inhibition of autophagy and neurodegeneration after brain trauma. Autophagy. 2020;16(3):466–85. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Pan B, et al. The calcineurin-TFEB-p62 pathway mediates the activation of cardiac macroautophagy by proteasomal malfunction. Circ Res. 2020;127(4):502–18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Mak VC, et al. p85β alters response to EGFR inhibitor in ovarian cancer through p38 MAPK-mediated regulation of DNA repair. Neoplasia. 2021;23(7):718–30. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Battaglia AM, et al. Ferroptosis and cancer: mitochondria meet the “iron maiden” cell death. Cells. 2020;9(6):1505. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Wang Y, et al. Overcoming cancer chemotherapy resistance by the induction of ferroptosis. Drug Resist Updat. 2023;66:100916. [DOI] [PubMed] [Google Scholar]
- 11.Chen X, et al. Cellular degradation systems in ferroptosis. Cell Death Differ. 2021;28(4):1135–48. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Shen Y, et al. Transferrin receptor 1 in cancer: a new sight for cancer therapy. Am J Cancer Res. 2018;8(6):916–31. [PMC free article] [PubMed] [Google Scholar]
- 13.Hu C, et al. Polyphyllin B Suppresses gastric tumor growth by modulating iron metabolism and inducing ferroptosis. Int J Biol Sci. 2023;19(4):1063–79. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Wu Y, et al. Identification of a novel inhibitor of TfR1 from designed and synthesized muriceidine A derivatives. Antioxidants. 2022;11(5):834. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Chang X, et al. Identification and characterization of elevated expression of transferrin and its receptor TfR1 in mouse models of depression. Brain Sci. 2022;12(10):1267. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Wu H, et al. ATM orchestrates ferritinophagy and ferroptosis by phosphorylating NCOA4. Autophagy. 2023;19(7):2062–77. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Xiao Y, et al. Fe(3+)-binding transferrin nanovesicles encapsulating sorafenib induce ferroptosis in hepatocellular carcinoma. Biomater Res. 2023;27(1):63. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Xiu Z, et al. Caryophyllene oxide induces ferritinophagy by regulating the NCOA4/FTH1/LC3 pathway in hepatocellular carcinoma. Front Pharmacol. 2022;13:930958. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Tian Y, et al. FTH1 inhibits ferroptosis through ferritinophagy in the 6-OHDA model of Parkinson’s disease. Neurotherapeutics. 2020;17(4):1796–812. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.He J, et al. Ferroptosis and ferritinophagy in diabetes complications. Mol Metab. 2022;60:101470. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Xue Q, et al. Copper-dependent autophagic degradation of GPX4 drives ferroptosis. Autophagy. 2023;19(7):1982–96. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Shen F, et al. Metal ions and nanometallic materials in antitumor immunity: function, application, and perspective. J Nanobiotechnology. 2023;21(1):20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Liu Y, Gu W. p53 in ferroptosis regulation: the new weapon for the old guardian. Cell Death Differ. 2022;29(5):895–910. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Carver PL. Metals in medicine: the therapeutic use of metal ions in the clinic. Met Ions Life Sci. 2019. 10.1515/9783110527872. [DOI] [PubMed] [Google Scholar]
- 25.Yang P, et al. Recent progress in drug delivery and cancer theranostic built from metal-organic framework. Biomed Mater. 2021;16(4):042011. [DOI] [PubMed] [Google Scholar]
- 26.Zhou Q, et al. Ferroptosis in cancer: from molecular mechanisms to therapeutic strategies. Signal Transduct Target Ther. 2024;9(1):55. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Grahovac J, Wells A. Matrikine and matricellular regulators of EGF receptor signaling on cancer cell migration and invasion. Lab Invest. 2014;94(1):31–40. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Sodir NM, et al. SHP2: a pleiotropic target at the interface of cancer and its microenvironment. Cancer Discov. 2023;13(11):2339–55. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Cicek E, et al. EGF-SNX3-EGFR axis drives tumor progression and metastasis in triple-negative breast cancers. Oncogene. 2022;41(2):220–32. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Tang KH, et al. Combined Inhibition of SHP2 and CXCR1/2 promotes antitumor T-cell response in NSCLC. Cancer Discov. 2022;12(1):47–61. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Montagner A, et al. A novel role for Gab1 and SHP2 in epidermal growth factor-induced Ras activation. J Biol Chem. 2005;280(7):5350–60. [DOI] [PubMed] [Google Scholar]
- 32.Kong M, et al. Gab2 tyrosine phosphorylation by a pleckstrin homology domain-independent mechanism: role in epidermal growth factor-induced mitogenesis. Mol Endocrinol. 2003;17(5):935–44. [DOI] [PubMed] [Google Scholar]
- 33.Neel BG, Gu H, Pao L. The ’Shp’ing news: SH2 domain-containing tyrosine phosphatases in cell signaling. Trends Biochem Sci. 2003;28(6):284–93. [DOI] [PubMed] [Google Scholar]
- 34.Papanicolaou KN, et al. Inhibiting O-GlcNAcylation impacts p38 and Erk1/2 signaling and perturbs cardiomyocyte hypertrophy. J Biol Chem. 2023;299(3):102907. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Tian P, et al. Hypoxia causes trophoblast cell ferroptosis to induce miscarriage through lnc-HZ06/HIF1α-SUMO/NCOA4 axis. Redox Biol. 2024;70:103073. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Li X, et al. Autophagy enhanced by curcumin ameliorates inflammation in atherogenesis via the TFEB-P300-BRD4 axis. Acta Pharm Sin B. 2022;12(5):2280–99. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Wang JF, et al. Upregulated PD-L1 delays human neutrophil apoptosis and promotes lung injury in an experimental mouse model of sepsis. Blood. 2021;138(9):806–10. [DOI] [PubMed] [Google Scholar]
- 38.Wang J, et al. Targeting PI3K/AKT signaling for treatment of idiopathic pulmonary fibrosis. Acta Pharm Sin B. 2022;12(1):18–32. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Prossomariti A, et al. Are Wnt/β-catenin and PI3K/AKT/mTORC1 distinct pathways in colorectal cancer? Cell Mol Gastroenterol Hepatol. 2020;10(3):491–506. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Alesi N, et al. TFEB drives mTORC1 hyperactivation and kidney disease in tuberous sclerosis complex. Nat Commun. 2024;15(1):406. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Cui Z, et al. Structure of the lysosomal mTORC1-TFEB-Rag-ragulator megacomplex. Nature. 2023;614(7948):572–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Li N, et al. Ferritinophagy-mediated ferroptosis is involved in sepsis-induced cardiac injury. Free Radic Biol Med. 2020;160:303–18. [DOI] [PubMed] [Google Scholar]
- 43.Xiu Z, et al. Inhibitory effects of esculetin on liver cancer through triggering NCOA4 pathway-mediation ferritinophagy in vivo and in vitro. J Hepatocell Carcinoma. 2023;10:611–29. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Jin Y, et al. LncRNA CACNA1G-AS1 up-regulates FTH1 to inhibit ferroptosis and promote malignant phenotypes in ovarian cancer cells. Oncol Res. 2023;31(2):169–79. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Bersuker K, et al. The CoQ oxidoreductase FSP1 acts parallel to GPX4 to inhibit ferroptosis. Nature. 2019;575(7784):688–92. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Hider RC, et al. Optical imaging opportunities to inspect the nature of cytosolic iron pools. Molecules. 2023;28(18):6467. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Liu L, et al. Suppression of USP8 sensitizes cells to ferroptosis via SQSTM1/p62-mediated ferritinophagy. Protein Cell. 2023;14(3):230–4. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.