Abstract
Analysis of clones isolated from a cosmid DNA library indicates that the Serratia marcescens chromosome contains at least two genes, chiA and chiB, which encode distinct secreted forms of the enzyme chitinase. These genes have been characterized by inspection of chitinase activity and secreted proteins in Escherichia coli strains containing subclones of these cosmids. The two chitinase genes show no detectable homology to each other. DNA sequence analysis of one of the genes predicts an amino acid sequence with an N-terminal signal peptide typical of genes encoding secreted bacterial proteins. This gene was mutagenized by cloning a neomycin phosphotransferase gene within its coding region, and the insertion mutation was recombined into the parental S. marcescens strain. The resulting chiA mutant transconjugant showed reduced chitinase production, reduced inhibition of fungal spore germination and reduced biological control of a fungal plant pathogen.
Keywords: biological control, catabolite repression, chitinase, protein secretion, Serratia
Full text
PDF

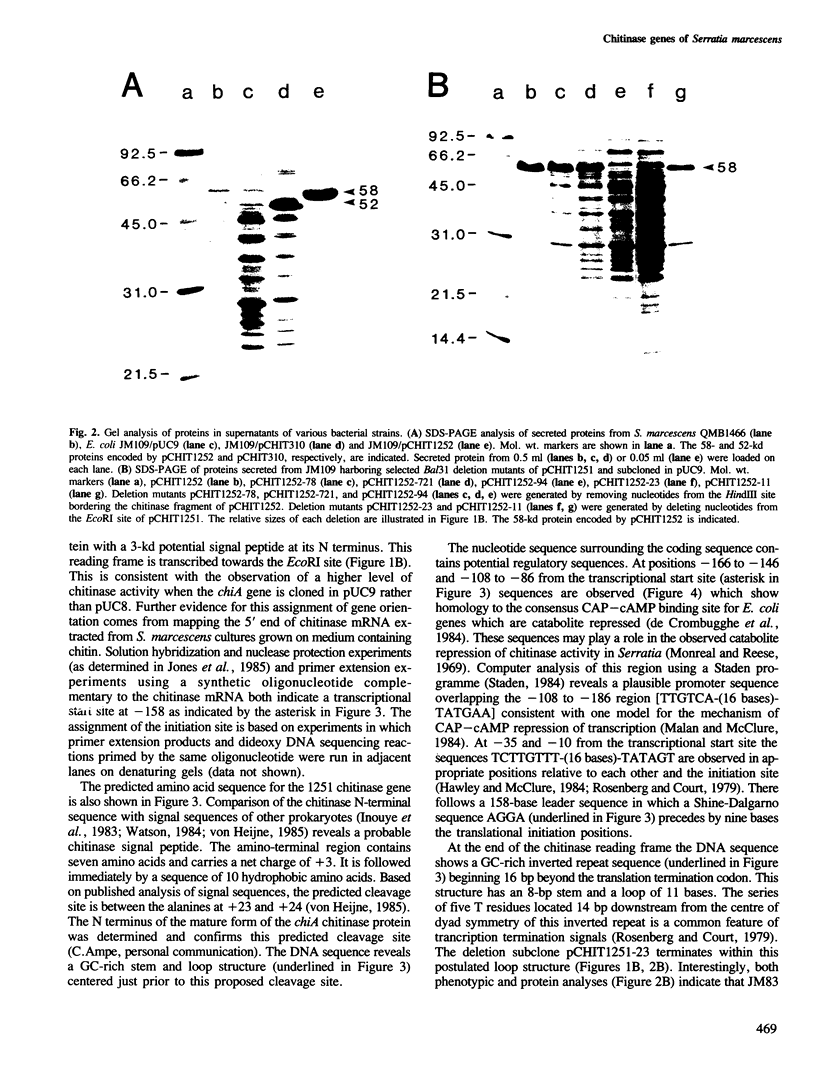

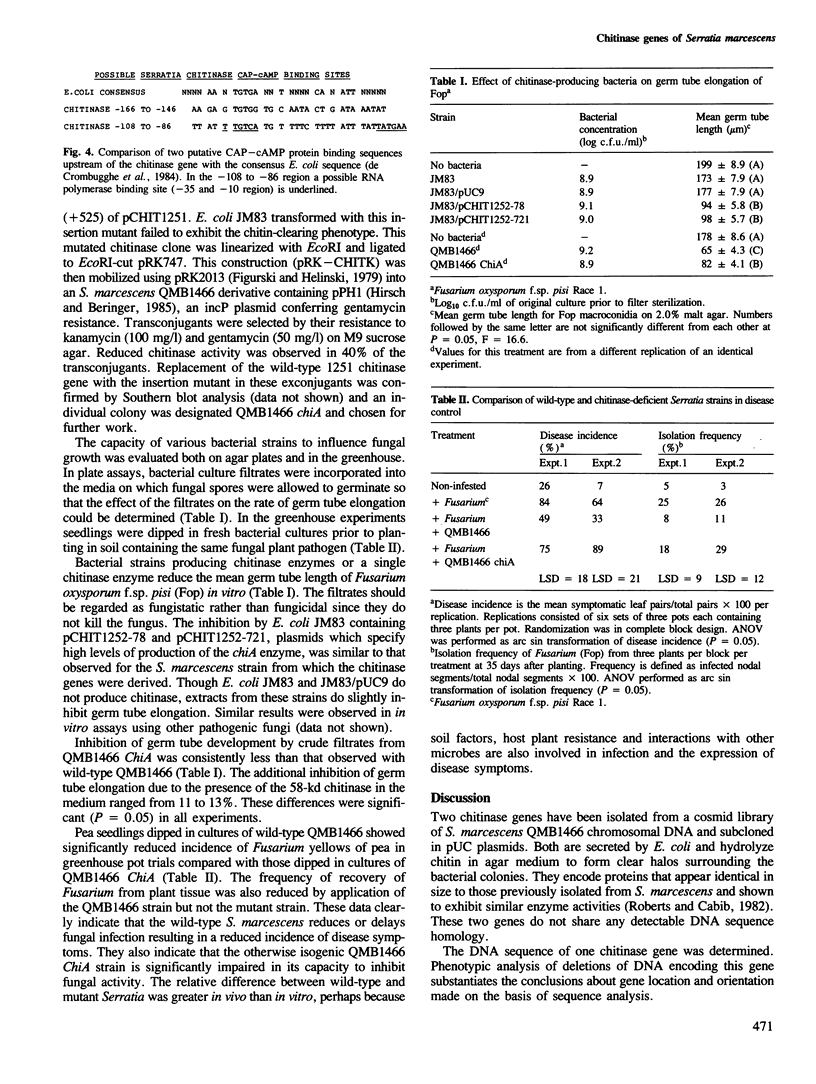


Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Abeles F. B., Bosshart R. P., Forrence L. E., Habig W. H. Preparation and purification of glucanase and chitinase from bean leaves. Plant Physiol. 1971 Jan;47(1):129–134. doi: 10.1104/pp.47.1.129. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bertrand K., Korn L. J., Lee F., Yanofsky C. The attenuator of the tryptophan operon of Escherichia coli. Heterogeneous 3'-OH termini in vivo and deletion mapping of functions. J Mol Biol. 1977 Nov 25;117(1):227–247. doi: 10.1016/0022-2836(77)90032-8. [DOI] [PubMed] [Google Scholar]
- Dean C., Elzen P., Tamaki S., Dunsmuir P., Bedbrook J. Differential expression of the eight genes of the petunia ribulose bisphosphate carboxylase small subunit multi-gene family. EMBO J. 1985 Dec 1;4(12):3055–3061. doi: 10.1002/j.1460-2075.1985.tb04045.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dhaese P., De Greve H., Decraemer H., Schell J., Van Montagu M. Rapid mapping of transposon insertion and deletion mutations in the large Ti-plasmids of Agrobacterium tumefaciens. Nucleic Acids Res. 1979 Dec 11;7(7):1837–1849. doi: 10.1093/nar/7.7.1837. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ditta G., Schmidhauser T., Yakobson E., Lu P., Liang X. W., Finlay D. R., Guiney D., Helinski D. R. Plasmids related to the broad host range vector, pRK290, useful for gene cloning and for monitoring gene expression. Plasmid. 1985 Mar;13(2):149–153. doi: 10.1016/0147-619x(85)90068-x. [DOI] [PubMed] [Google Scholar]
- Elango N., Correa J. U., Cabib E. Secretory character of yeast chitinase. J Biol Chem. 1982 Feb 10;257(3):1398–1400. [PubMed] [Google Scholar]
- Figurski D. H., Helinski D. R. Replication of an origin-containing derivative of plasmid RK2 dependent on a plasmid function provided in trans. Proc Natl Acad Sci U S A. 1979 Apr;76(4):1648–1652. doi: 10.1073/pnas.76.4.1648. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Friedman A. M., Long S. R., Brown S. E., Buikema W. J., Ausubel F. M. Construction of a broad host range cosmid cloning vector and its use in the genetic analysis of Rhizobium mutants. Gene. 1982 Jun;18(3):289–296. doi: 10.1016/0378-1119(82)90167-6. [DOI] [PubMed] [Google Scholar]
- Hawley D. K., McClure W. R. Compilation and analysis of Escherichia coli promoter DNA sequences. Nucleic Acids Res. 1983 Apr 25;11(8):2237–2255. doi: 10.1093/nar/11.8.2237. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hirsch P. R., Beringer J. E. A physical map of pPH1JI and pJB4JI. Plasmid. 1984 Sep;12(2):139–141. doi: 10.1016/0147-619x(84)90059-3. [DOI] [PubMed] [Google Scholar]
- Inouye S., Hsu C. P., Itakura K., Inouye M. Requirement for signal peptide cleavage of Escherichia coli prolipoprotein. Science. 1983 Jul 1;221(4605):59–61. doi: 10.1126/science.6344218. [DOI] [PubMed] [Google Scholar]
- Jones J. D., Dunsmuir P., Bedbrook J. High level expression of introduced chimaeric genes in regenerated transformed plants. EMBO J. 1985 Oct;4(10):2411–2418. doi: 10.1002/j.1460-2075.1985.tb03949.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Laemmli U. K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature. 1970 Aug 15;227(5259):680–685. doi: 10.1038/227680a0. [DOI] [PubMed] [Google Scholar]
- Malan T. P., McClure W. R. Dual promoter control of the Escherichia coli lactose operon. Cell. 1984 Nov;39(1):173–180. doi: 10.1016/0092-8674(84)90203-4. [DOI] [PubMed] [Google Scholar]
- Messing J. New M13 vectors for cloning. Methods Enzymol. 1983;101:20–78. doi: 10.1016/0076-6879(83)01005-8. [DOI] [PubMed] [Google Scholar]
- Monreal J., Reese E. T. The chitinase of Serratia marcescens. Can J Microbiol. 1969 Jul;15(7):689–696. doi: 10.1139/m69-122. [DOI] [PubMed] [Google Scholar]
- Roberts R. L., Cabib E. Serratia marcescens chitinase: one-step purification and use for the determination of chitin. Anal Biochem. 1982 Dec;127(2):402–412. doi: 10.1016/0003-2697(82)90194-4. [DOI] [PubMed] [Google Scholar]
- Rosenberg M., Court D. Regulatory sequences involved in the promotion and termination of RNA transcription. Annu Rev Genet. 1979;13:319–353. doi: 10.1146/annurev.ge.13.120179.001535. [DOI] [PubMed] [Google Scholar]
- Ruvkun G. B., Ausubel F. M. A general method for site-directed mutagenesis in prokaryotes. Nature. 1981 Jan 1;289(5793):85–88. doi: 10.1038/289085a0. [DOI] [PubMed] [Google Scholar]
- Sanger F., Coulson A. R., Barrell B. G., Smith A. J., Roe B. A. Cloning in single-stranded bacteriophage as an aid to rapid DNA sequencing. J Mol Biol. 1980 Oct 25;143(2):161–178. doi: 10.1016/0022-2836(80)90196-5. [DOI] [PubMed] [Google Scholar]
- Southern E. M. Detection of specific sequences among DNA fragments separated by gel electrophoresis. J Mol Biol. 1975 Nov 5;98(3):503–517. doi: 10.1016/s0022-2836(75)80083-0. [DOI] [PubMed] [Google Scholar]
- Staden R. Computer methods to locate signals in nucleic acid sequences. Nucleic Acids Res. 1984 Jan 11;12(1 Pt 2):505–519. doi: 10.1093/nar/12.1part2.505. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vieira J., Messing J. The pUC plasmids, an M13mp7-derived system for insertion mutagenesis and sequencing with synthetic universal primers. Gene. 1982 Oct;19(3):259–268. doi: 10.1016/0378-1119(82)90015-4. [DOI] [PubMed] [Google Scholar]
- Watson M. E. Compilation of published signal sequences. Nucleic Acids Res. 1984 Jul 11;12(13):5145–5164. doi: 10.1093/nar/12.13.5145. [DOI] [PMC free article] [PubMed] [Google Scholar]
- de Crombrugghe B., Busby S., Buc H. Cyclic AMP receptor protein: role in transcription activation. Science. 1984 May 25;224(4651):831–838. doi: 10.1126/science.6372090. [DOI] [PubMed] [Google Scholar]
- von Heijne G. Signal sequences. The limits of variation. J Mol Biol. 1985 Jul 5;184(1):99–105. doi: 10.1016/0022-2836(85)90046-4. [DOI] [PubMed] [Google Scholar]



