Abstract
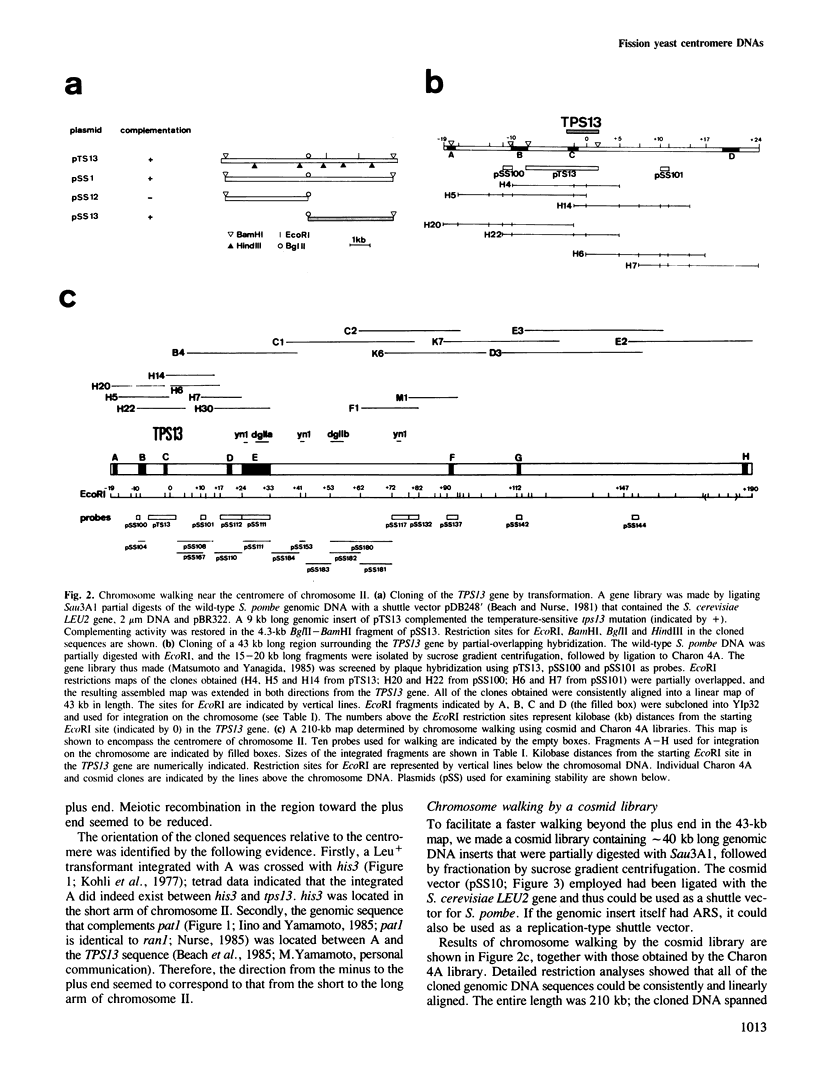
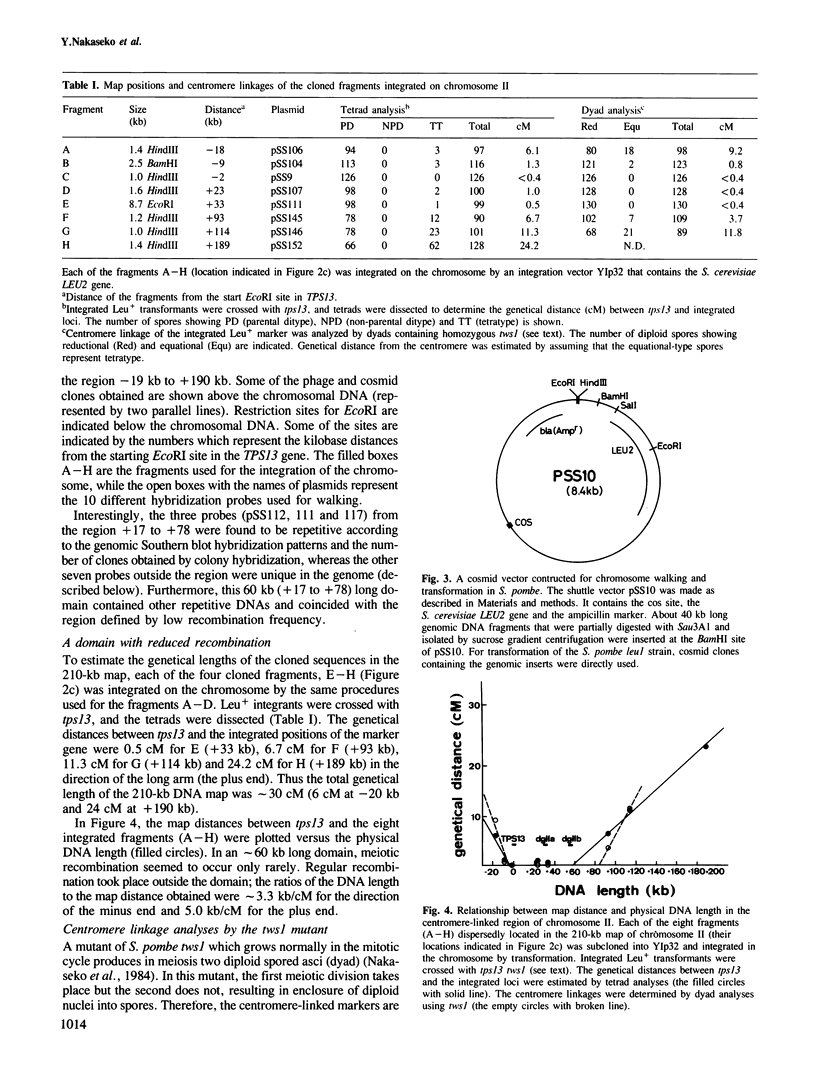
By cloning centromere-linked genes followed by partial overlapping hybridization, we constructed a 210-kb map encompassing the centromere in chromosome II and a 60-kb map near the centromere of chromosome I in the fission yeast Schizosaccharomyces pombe which has three chromosomes. Integration of the cloned sequences onto the chromosome and subsequent analyses of tetrads and dyads revealed an ∼50 kb long domain located in the middle of the 210-kb map, tightly linked to the centromere and greatly reduced in meiotic recombination. This domain contained at least two classes of repetitive sequences. One, designated yn1, was specifically present in a particular chromosome and repeated three times in the 210-kb map of chromosome II. The other, designated dg, was located in all the centromere regions of three chromosomes. One (dgI) and two (dgIIa, dgIIb) copies of the dg were found in the maps of chromosomes I and II, respectively. The dgIIa and dgIIb were arranged with a 20-kb interval within the repetitive domain. In the centric region of chromosome III, 3−4 copies of the dg appeared to exist. By determining the nucleotide sequences of dgI and dgIIa, the dg was identified to be 3.8 kb long. The sequence homology was 99% between dgI and dgIIa. These extraordinarily homologous sequences seemed not to be transcribed into RNA nor to be encoding any protein. The larger part of the dg sequence was internally non-repetitious, a 600-bp region existed which consisted of stretches of several short repeating units. The structures in or surrounding the centromeres of S. pombe appear to be much more complex than those of the budding yeast Saccharomyces cerevisiae.
Keywords: centromere, chromosome walking, domain structure, repetitive DNA, Schizosaccharomyces pombe
Full text
PDF



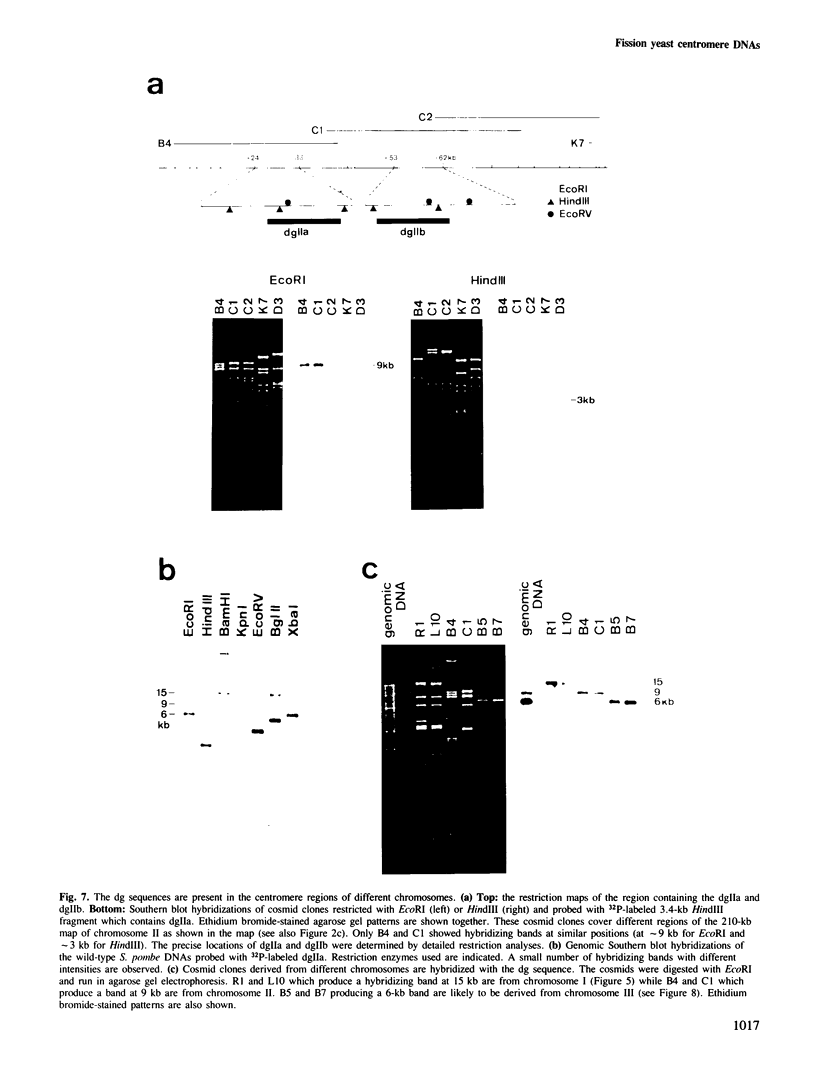




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Beach D., Piper M., Nurse P. Construction of a Schizosaccharomyces pombe gene bank in a yeast bacterial shuttle vector and its use to isolate genes by complementation. Mol Gen Genet. 1982;187(2):326–329. doi: 10.1007/BF00331138. [DOI] [PubMed] [Google Scholar]
- Beach D., Rodgers L., Gould J. ran1+ controls the transition from mitotic division to meiosis in fission yeast. Curr Genet. 1985;10(4):297–311. doi: 10.1007/BF00365626. [DOI] [PubMed] [Google Scholar]
- Blackburn E. H. The molecular structure of centromeres and telomeres. Annu Rev Biochem. 1984;53:163–194. doi: 10.1146/annurev.bi.53.070184.001115. [DOI] [PubMed] [Google Scholar]
- Bloom K. S., Carbon J. Yeast centromere DNA is in a unique and highly ordered structure in chromosomes and small circular minichromosomes. Cell. 1982 Jun;29(2):305–317. doi: 10.1016/0092-8674(82)90147-7. [DOI] [PubMed] [Google Scholar]
- Botstein D., Falco S. C., Stewart S. E., Brennan M., Scherer S., Stinchcomb D. T., Struhl K., Davis R. W. Sterile host yeasts (SHY): a eukaryotic system of biological containment for recombinant DNA experiments. Gene. 1979 Dec;8(1):17–24. doi: 10.1016/0378-1119(79)90004-0. [DOI] [PubMed] [Google Scholar]
- Carbon J. Yeast centromeres: structure and function. Cell. 1984 Jun;37(2):351–353. doi: 10.1016/0092-8674(84)90363-5. [DOI] [PubMed] [Google Scholar]
- Carle G. F., Olson M. V. Separation of chromosomal DNA molecules from yeast by orthogonal-field-alternation gel electrophoresis. Nucleic Acids Res. 1984 Jul 25;12(14):5647–5664. doi: 10.1093/nar/12.14.5647. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clarke L., Carbon J. Genomic substitutions of centromeres in Saccharomyces cerevisiae. Nature. 1983 Sep 1;305(5929):23–28. doi: 10.1038/305023a0. [DOI] [PubMed] [Google Scholar]
- Clarke L., Carbon J. Isolation of a yeast centromere and construction of functional small circular chromosomes. Nature. 1980 Oct 9;287(5782):504–509. doi: 10.1038/287504a0. [DOI] [PubMed] [Google Scholar]
- Fitzgerald-Hayes M., Buhler J. M., Cooper T. G., Carbon J. Isolation and subcloning analysis of functional centromere DNA (CEN11) from Saccharomyces cerevisiae chromosome XI. Mol Cell Biol. 1982 Jan;2(1):82–87. doi: 10.1128/mcb.2.1.82. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fitzgerald-Hayes M., Clarke L., Carbon J. Nucleotide sequence comparisons and functional analysis of yeast centromere DNAs. Cell. 1982 May;29(1):235–244. doi: 10.1016/0092-8674(82)90108-8. [DOI] [PubMed] [Google Scholar]
- Hieter P., Mann C., Snyder M., Davis R. W. Mitotic stability of yeast chromosomes: a colony color assay that measures nondisjunction and chromosome loss. Cell. 1985 Feb;40(2):381–392. doi: 10.1016/0092-8674(85)90152-7. [DOI] [PubMed] [Google Scholar]
- Hieter P., Pridmore D., Hegemann J. H., Thomas M., Davis R. W., Philippsen P. Functional selection and analysis of yeast centromeric DNA. Cell. 1985 Oct;42(3):913–921. doi: 10.1016/0092-8674(85)90287-9. [DOI] [PubMed] [Google Scholar]
- Hiraoka Y., Toda T., Yanagida M. The NDA3 gene of fission yeast encodes beta-tubulin: a cold-sensitive nda3 mutation reversibly blocks spindle formation and chromosome movement in mitosis. Cell. 1984 Dec;39(2 Pt 1):349–358. doi: 10.1016/0092-8674(84)90013-8. [DOI] [PubMed] [Google Scholar]
- Hohn B., Collins J. A small cosmid for efficient cloning of large DNA fragments. Gene. 1980 Nov;11(3-4):291–298. doi: 10.1016/0378-1119(80)90069-4. [DOI] [PubMed] [Google Scholar]
- Hsiao C. L., Carbon J. Direct selection procedure for the isolation of functional centromeric DNA. Proc Natl Acad Sci U S A. 1981 Jun;78(6):3760–3764. doi: 10.1073/pnas.78.6.3760. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ito H., Fukuda Y., Murata K., Kimura A. Transformation of intact yeast cells treated with alkali cations. J Bacteriol. 1983 Jan;153(1):163–168. doi: 10.1128/jb.153.1.163-168.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kohli J., Hottinger H., Munz P., Strauss A., Thuriaux P. Genetic Mapping in SCHIZOSACCHAROMYCES POMBE by Mitotic and Meiotic Analysis and Induced Haploidization. Genetics. 1977 Nov;87(3):471–489. doi: 10.1093/genetics/87.3.471. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Käufer N. F., Simanis V., Nurse P. Fission yeast Schizosaccharomyces pombe correctly excises a mammalian RNA transcript intervening sequence. Nature. 1985 Nov 7;318(6041):78–80. doi: 10.1038/318078a0. [DOI] [PubMed] [Google Scholar]
- Maine G. T., Surosky R. T., Tye B. K. Isolation and characterization of the centromere from chromosome V (CEN5) of Saccharomyces cerevisiae. Mol Cell Biol. 1984 Jan;4(1):86–91. doi: 10.1128/mcb.4.1.86. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Matsumoto S., Yanagida M. Histone gene organization of fission yeast: a common upstream sequence. EMBO J. 1985 Dec 16;4(13A):3531–3538. doi: 10.1002/j.1460-2075.1985.tb04113.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McCully E. K., Robinow C. F. Mitosis in the fission yeast Schizosaccharomyces pombe: a comparative study with light and electron microscopy. J Cell Sci. 1971 Sep;9(2):475–507. doi: 10.1242/jcs.9.2.475. [DOI] [PubMed] [Google Scholar]
- Messing J., Crea R., Seeburg P. H. A system for shotgun DNA sequencing. Nucleic Acids Res. 1981 Jan 24;9(2):309–321. doi: 10.1093/nar/9.2.309. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mitchison T. J., Kirschner M. W. Properties of the kinetochore in vitro. I. Microtubule nucleation and tubulin binding. J Cell Biol. 1985 Sep;101(3):755–765. doi: 10.1083/jcb.101.3.755. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mitchison T. J., Kirschner M. W. Properties of the kinetochore in vitro. II. Microtubule capture and ATP-dependent translocation. J Cell Biol. 1985 Sep;101(3):766–777. doi: 10.1083/jcb.101.3.766. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Murray A. W., Szostak J. W. Construction of artificial chromosomes in yeast. Nature. 1983 Sep 15;305(5931):189–193. doi: 10.1038/305189a0. [DOI] [PubMed] [Google Scholar]
- Nakaseko Y., Niwa O., Yanagida M. A meiotic mutant of the fission yeast Schizosaccharomyces pombe that produces mature asci containing two diploid spores. J Bacteriol. 1984 Jan;157(1):334–336. doi: 10.1128/jb.157.1.334-336.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nurse P., Thuriaux P., Nasmyth K. Genetic control of the cell division cycle in the fission yeast Schizosaccharomyces pombe. Mol Gen Genet. 1976 Jul 23;146(2):167–178. doi: 10.1007/BF00268085. [DOI] [PubMed] [Google Scholar]
- Panzeri L., Philippsen P. Centromeric DNA from chromosome VI in Saccharomyces cerevisiae strains. EMBO J. 1982;1(12):1605–1611. doi: 10.1002/j.1460-2075.1982.tb01362.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Peterson J. B., Ris H. Electron-microscopic study of the spindle and chromosome movement in the yeast Saccharomyces cerevisiae. J Cell Sci. 1976 Nov;22(2):219–242. doi: 10.1242/jcs.22.2.219. [DOI] [PubMed] [Google Scholar]
- Ris H., Witt P. L. Structure of the mammalian kinetochore. Chromosoma. 1981;82(2):153–170. doi: 10.1007/BF00286101. [DOI] [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schwartz D. C., Cantor C. R. Separation of yeast chromosome-sized DNAs by pulsed field gradient gel electrophoresis. Cell. 1984 May;37(1):67–75. doi: 10.1016/0092-8674(84)90301-5. [DOI] [PubMed] [Google Scholar]
- Stinchcomb D. T., Mann C., Davis R. W. Centromeric DNA from Saccharomyces cerevisiae. J Mol Biol. 1982 Jun 25;158(2):157–190. doi: 10.1016/0022-2836(82)90427-2. [DOI] [PubMed] [Google Scholar]
- Szostak J. W., Blackburn E. H. Cloning yeast telomeres on linear plasmid vectors. Cell. 1982 May;29(1):245–255. doi: 10.1016/0092-8674(82)90109-x. [DOI] [PubMed] [Google Scholar]
- Toda T., Yamamoto M., Yanagida M. Sequential alterations in the nuclear chromatin region during mitosis of the fission yeast Schizosaccharomyces pombe: video fluorescence microscopy of synchronously growing wild-type and cold-sensitive cdc mutants by using a DNA-binding fluorescent probe. J Cell Sci. 1981 Dec;52:271–287. doi: 10.1242/jcs.52.1.271. [DOI] [PubMed] [Google Scholar]
- Valdivia M. M., Brinkley B. R. Fractionation and initial characterization of the kinetochore from mammalian metaphase chromosomes. J Cell Biol. 1985 Sep;101(3):1124–1134. doi: 10.1083/jcb.101.3.1124. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yanisch-Perron C., Vieira J., Messing J. Improved M13 phage cloning vectors and host strains: nucleotide sequences of the M13mp18 and pUC19 vectors. Gene. 1985;33(1):103–119. doi: 10.1016/0378-1119(85)90120-9. [DOI] [PubMed] [Google Scholar]