Abstract
Long-term control of viral replication relies on the efficient differentiation of memory T cells into effector T cells during secondary immune responses. Recent findings have identified T cell precursors for both memory and exhausted T cells, suggesting the existence of progenitor-like effector T cells. These cells can persist without antigenic challenge but expand and acquire effector functions upon recall immune responses. In this study, we demonstrate that the combination of SLAMF7 with either CD27 or TCF-1 effectively identifies progenitor-like effector CD8 T cells, while SLAMF7 with GPR56 or TOX defines effector CD8 T cells. These markers allow for the clear segregation of these distinct cell subsets. SLAMF7+ CD8T cells are dynamically modulated during viral infections, including HIV, HCV, CMV, and SARS-CoV-2, as well as during aging. We further characterize the SLAMF7 signature at both phenotypic and transcriptional levels. Notably, during aging, the SLAMF7 pathway becomes dysregulated, resulting in persistent phosphorylation of STAT1. Additionally, SLAMF7 ligation in the presence of IL-15 induces TCF-1 expression, which promotes the homeostatic proliferation of progenitor-like effector CD8 T cells.
Subject terms: Viral infection, Lymphocyte differentiation
Introduction
T cells evolve both transcriptionally and functionally into diverse phenotypes with age. The burden of persistent infections, low-grade inflammation, and thymic involution independently contribute to immune cell differentiation, leading to the development of heterogeneous T-cell phenotypes. While much progress has been made in studying T-cell exhaustion in cancer and chronic infection, the development of “replicative senescence” in T cells remains poorly understood in these contexts1. Identifying markers that distinguish early effector (or progenitor-like effector) from terminal effector T-cells is crucial for optimizing immunotherapies, particularly those based on immune checkpoint blockade or therapies that promote the differentiation of T cell precursors into functional effector T cells.
The factors contributing to the terminal differentiation of effector T cells are multifaceted. Thymic atrophy, combined with persistent T-cell receptor (TCR) activation in chronic infections such as CMV, promotes memory inflation in T cells2. Persistent latent CMV infection drives the accumulation of CMV-specific T cells, exhibiting an “effector” phenotype in the peripheral blood3,4. Similar to the replicative senescence seen in fibroblasts, terminally differentiated T cells exhibit specific features, such as shortened telomeres, reduced proliferative capacity, altered mitochondrial functions, elevated expression of p53 and p38 MAPK, and increased production of reactive oxygen species. (ROS)5. However, these hallmarks of T-cell senescence have not been consistently linked to reliable phenotypic markers. Several studies have explored potential signatures for T-cell senescence, including the detection of cell cycle regulators p16, p21, and p27. Surface markers associated with terminal differentiation in T cells include the loss of CD27 and CD28, along with the acquisition of CD57, and KLRG1 expression6–8. Recently, GPR56 has emerged as a marker for identifying functional memory9, effector, and terminally differentiated T cells10,11. Despite these findings, further validation is required for the reliable identification of progenitor-like effector CD8 T cells, which we investigate here.
Terminally differentiated CD8 T cells and NK cells share some common transcriptional signatures, including the expression of T-bet, Eomes, ZEB2, RUNX3, and HOBIT12–14. These transcription factors govern not only the differentiation of these cells but also their cytotoxic properties and preferential metabolic pathways15–17. Here, we investigate hallmark transcriptional signature in effector CD8 T cells and antigen-specific T cells across aging process.
Using single-cell proteomics and RNA sequencing, we identified SLAMF7 (also known as CRACC, CD319, CS1) as an early marker of effector CD8 T cells. SLAMF7 was initially shown to regulate NK cells18,19, as well as myeloid20 and plasma cell activity21,22. In patients with Systemic Lupus Erythematous (SLE), lower SLAMF7 expression levels were observed in memory and effector CD8 T cells. This defect was reversible through SLAMF7 engagement, which enhanced the cytotoxicity of antigen-specific CD8 T cells23. In aged donors, CMV-specific CD8 T cells expansion may lead to TCR repertoire restriction, resulting in poor vaccination responses to and diminished tumour immunity. Therefore, we investigated whether CMV-specific CD8 T cells characteristics evolve with age24 and whether SLAMF7 influences the functionality of these cells. Impaired CD8 T-cell cytotoxicity may lead to the accumulation of terminally differentiated cells in tissues, contributing to increased “inflammaging”. We evaluated the SLAMF7 signalling pathway to identify mechanisms involved in its regulation.
The identification of SLAMF7 as an early effector marker may help in developing strategies to reverse the detrimental effects of terminal differentiation, which may impede responsiveness to immunotherapy, while enhancing immune cell effector functions. Here, we describe SLAMF7 expression changes in aging and individuals with chronic infections such as CMV or people living with HIV (PLWH). Although SLAMF7+CD8 T cells exhibit heterogeneity, we identified two main subsets characterized by differential CD27/TCF-1 (co-expressed with CD5/CD127) and GPR56/TOX (co-expressed with CX3CR1/SLAMF6) expression. The frequencies of these subsets may predict patient responsiveness to checkpoint blockade therapy and the functional capacities of effector T cells.
Results
Heterogeneity of SLAMF7 expressing T cells
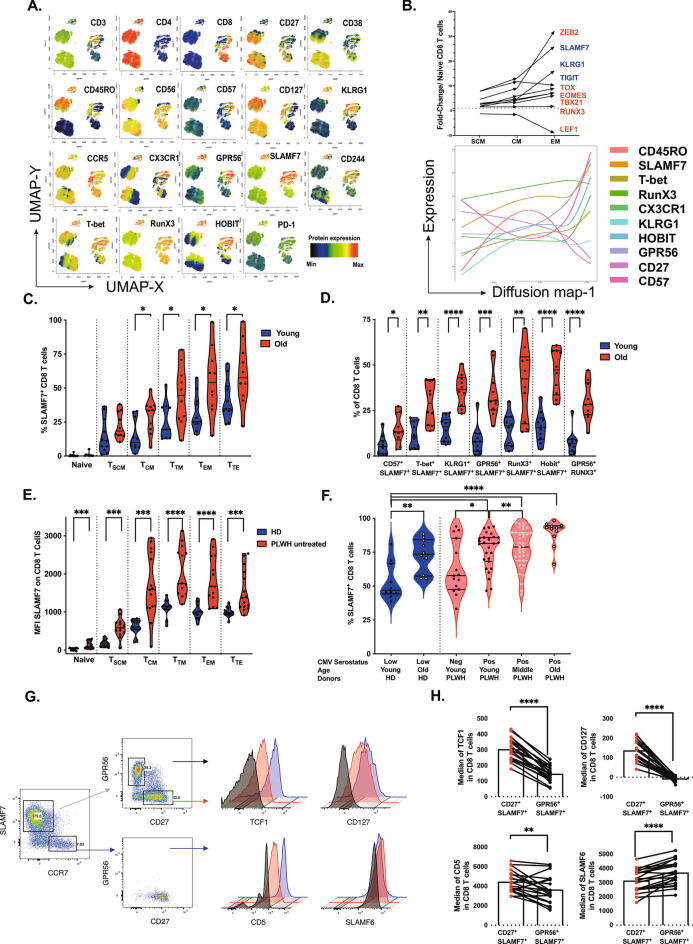
We began by investigating the expression patterns of SLAMF7 across the main T-cell subsets using a high-dimensional flow cytometry panel of 26 antibodies, Figure S1. Visualized by UMAP, unsupervised clustering of T cells based on combinatorial expression of 19 surface markers identified clusters enriched for SLAMF7. These SLAMF7+ clusters were found in both CD4 and CD8 T cells and were closely associated with high expression of transcription factors (TFs) such as T-bet, RUNX3, and HOBIT, as well as chemokine receptors like CX3CR1 and CCR5, and other surface markers including GPR56, CD57, KLRG1, and CD244 (Fig. 1A). This finding aligns with prior RNAseq data25,26, which we now extend to protein-level insights in CD8 T cell subsets, including Stem Cell Memory (SCM), Central Memory (CM), and Effector Memory (TEM) populations (Fig. 1B, top). To assess the relationship between SLAMF7 expression and T-cell differentiation, we developed a linear protein model that traced marker expression kinetics. SLAMF7 expression increased in tandem with T-bet, RUNX3, HOBIT, KLRG1, CD57, CX3CR1, and GPR56, and coincided with the loss of CD27 and CD45RO expression, confirming SLAMF7 as a marker of progressive T-cell differentiation (Fig. 1B, bottom).
Fig. 1.
Heterogeneity of SLAMF7 expressing T cells. (A) Heterogeneity of T cells by high-dimensional single-cell flow cytometry staining. Frozen PBMCs were thawed and stained for flow cytometry. T cells of 20 donors (n = 10 each for young and older donors, all CMV seropositive) were concatenated. T cells were visualized by UMAP and a cold to hot heat map was used to represent the intensity of each marker. (B) Characterization of human CD8 T cell maturation. The mRNA fold change expression in comparison to naïve CD8 T cells was calculated in sorted populations of T Stem Cell Memory (TSCM), Central Memory (TCM), and Effector Memory (TEM), top. The mRNA coding for transcription factors is labelled in red and surface molecules in black. The cellular progression according to the diffusion map was visualized by the expression profile of CD45RO, SLAMF7, CX3CR1, KLRG1, GPR56, CD27, CD57, T-bet, RunX3, and HOBIT in CMV seropositive donors, bottom. (C) SLAMF7 expression in CD8 T cell subsets during aging. The frequency of SLAMF7 was evaluated in naïve (CD45RO-CCR7+CD27+CD95-), TSCM (CD45RO-CCR7+CD27+CD95+), TCM (CD45RO+CCR7+CD27+CD95+), TTM (CD45RO+CCR7-CD27+CD95+), TEM (CD45RO+CCR7-CD27-CD95+), and TTE (CD45RO-CCR7-CD27-CD95+) of frozen PBMCs from young and older donors (n = 10 and n = 11 respectively). The statistical analysis was performed on unpaired samples (U Mann–Whitney test) (** and **** for p < 0.01 and p < 0.0001, respectively). (D) Heterogeneity of SLAMF7 expressing T cells during aging. Staining was performed on frozen PBMCs of young (n = 10) and elderly donors (n = 10). The statistical analysis was performed on unpaired samples (U Mann–Whitney test; *, **, ***, and **** for p < 0.05, p < 0.01, p < 0.001, and p < 0.0001, respectively). (E) SLAMF7 expression in CD8 T cell subsets during untreated HIV infection. The median intensity of SLAMF7 expression was evaluated in subsets (as defined in Fig. 1C) of frozen PBMCs from PLWH and healthy donors (n = 15 and n = 12 respectively). The statistical analysis was performed on unpaired samples (U Mann–Whitney test) (** and **** for p < 0.01 and p < 0.0001, respectively). (F) SLAMF7 expression in CD8 T cell subsets during chronic viral infections and aging. The frequencies of SLAMF7 expression were evaluated in total CD8 T cells from frozen PBMCs of PLWH under anti-retroviral therapy and co-infected or not with CMV. Patients have been stratified by age (< 40, 40–65, > 65 years old as young, middle age, and old with n = 42, n = 43, and n = 11 respectively) and compared to healthy donors (n = 20). The statistical analysis was performed on unpaired samples (U Mann–Whitney test) (** and **** for p < 0.01 and p < 0.0001, respectively). (G) Identification of progenitor-like and terminal effector CD8 T cells based on SLAMF7 expression. Description of the gating strategy used to identify SLAMF7+-progenitor-like (CD27+GPR56-) and terminal effector (CD27-GPR56+) population in total CD8 T cells. The histograms represented the overlaid intensity of TCF-1, CD127, SLAMF6, and CD5 in SLAMF7 subsets in comparison to Naïve/ Central Memory (CD27+CCR7+) CD8 T cells. (H) Characterization of progenitor-like and terminal effector CD8 T cells. The median intensity of fluorescence was compared between progenitor-like and terminal effector CD8 T cell populations. The statistical analysis was performed on paired samples (n = 20, Wilcoxon signed-rank test), (** and **** for p < 0.01 and p < 0.0001, respectively).
Using conventional gating approaches and classifying CD8 T cells into subsets (Naïve, TSCM, TCM, TTM, TEM, TTE) based on markers such as CD27, CD31, CD45RO, CCR7, and CD95, we observed a clear increase in the frequency of SLAMF7+ cells with progressive differentiation (Fig. 1C). Moreover, the percentage of SLAMF7+ CD8 memory T cells was significantly higher in elderly donors compared to younger individuals across all differentiated subsets (p < 0.05), except for Naïve and TSCM cells (Fig. 1C). Combining SLAMF7 with additional markers revealed pronounced differences between young and older donors (T-bet (p < 0.01), HOBIT (p < 0.0001), KLRG1 (p < 0.0001), CD57 (p < 0.05), RUNX3 (p < 0.01) and GPR56 (p < 0.001)), highlighting the complex regulation of TF and surface markers during aging (Fig. 1D).
To further explore SLAMF7’s role in T cell differentiation, we examined its expression in the context of chronic HIV infection, a model of accelerated T-cell aging (due to chronic immune activation caused by microbial translocation and chronic TCR engagement by persistent viral antigens). SLAMF7 expression was significantly higher in CD8 T cells from people living with HIV (PLWH) compared to age-matched healthy donors (HD) (p < 0.0001 for TTM or TEM and p < 0.001 for other subsets), with progressive increases observed along the differentiation spectrum (Fig. 1E). Additionally, the absolute count of SLAMF7-expressing CD8 T cells in the peripheral blood of ART-treated PLWH was increased (p < 0.01), Figure S2A. We wondered whether SLAMF7 expression might be regulated by the control of HIV replication in successfully treated PLWH (Viral load < 50 copies per mL and CD4 > 500 cells per mm3), Figure S2B. ART did not affect SLAMF7-expression but simultaneously increased the long-term memory marker CD12727 (p < 0.05) and decreased exhaustion markers such as Tim-3 (p < 0.05), Ceacam-1 (p < 0.001), and TFs involved in T cell differentiation and exhaustion such as T-bet (p < 0,05), and Eomes (p < 0.05). Given the role of CMV co-infection in exacerbating T-cell differentiation, we stratified PLWH based on CMV serostatus and age, Fig. 1F. We found that aging, CMV, and HIV infection all contributed to the accumulation of SLAMF7+ CD8 T cells, with elderly CMV-seropositive PLWH showing the highest frequencies of these cells (Fig. 1F).
Considering that the loss of CD127 expression and proliferation capacity are hallmarks of exhaustion, we examined SLAMF7 expression alongside CD127 and various inhibitory receptors (PD-1, TIGIT, Tim-3, Caecam-1, and CD160) in PLWH co-infected with CMV or with CMV and HCV during aging (Fig. S2C–E). We found that SLAMF7+ CD8 T cells were more prevalent in terminally differentiated subsets (T-bethighEomesdimCD57+) rather than in exhausted cells (T-betdimEomeshighCD160+TIGIT+). PD-1 expression was continuous and not exclusive to exhaustion, Fig. S2E.
To characterize the heterogeneity within SLAMF7+ CD8 T cells, we defined two main subsets based on effector functions: SLAMF7+CD27+ (Progenitor-like Effector) and SLAMF7+GPR56+ (Terminal Effector) (Fig. 1G). These subsets differed in their expression of key markers, including TCF-1, CD127, CD5 and SLAMF6 (Fig. 1G,H) as well as CD28, CD45RO and CD57 (or by using CX3CR128 instead of GPR56) (Fig. S2F). Interestingly, SLAMF6 expression increased globally with age and the frequency of the GPR56+ subset co-expressing SLAMF7 was significantly higher in elderly individuals (p < 0.01 in comparison to young donors with similar CMV serostatus), Fig. S2G.
Multi-omic signature of SLAMF7 expressing T cells
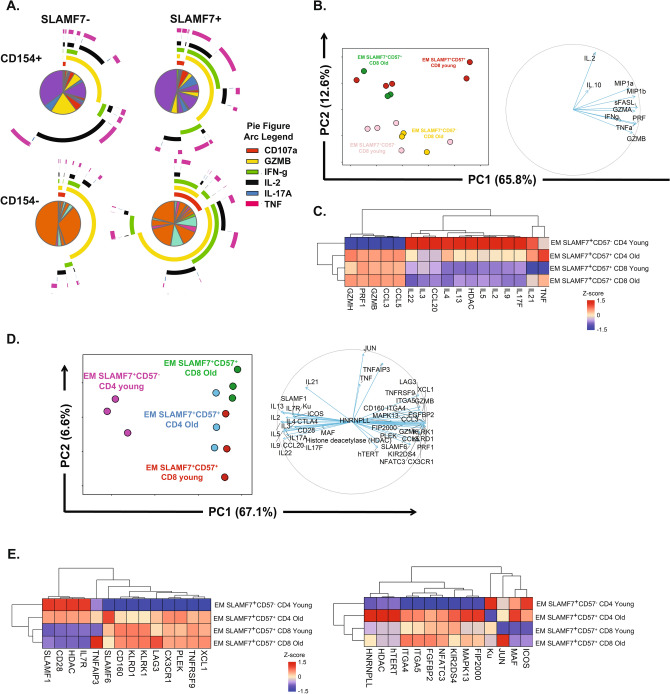
To further investigate the functional implications of SLAMF7 expression, we analyzed its association with effector functions in T cells. Polyclonally stimulated SLAMF7+ T cells demonstrated high expression of IFN-γ, TNF-α, CD107a, Granzyme B, and Eomes (Fig. 2A and Figure S3A), but reduced levels of CD154, IL-2 and IL-17A (Fig. S3B). Using SPICE analysis, we categorized the data based on CD154 expression to distinguish between CD4 and CD8 T cells (Fig. 2A). SLAMF7+ CD154- T cells (predominantly CD8) displayed greater poly-functionality compared to SLAMF7- T cells (Fig. 2A and Fig. S3B). Notably, SLAMF7+ T cells exhibited higher frequencies of IFN-γ (p < 0.001), CD107a (p < 0.01), Granzyme B (p < 0.01), and Eomes (p < 0.01), compared to their SLAMF7- counterparts (Fig. S3C).
Fig. 2.
Multiomic signature of SLAMF7 expressing T cells. (A) Poly-functionality of SLAMF7+ T cells. Functional heterogeneity of T cells revealed by Flow Cytometry analysis after PMA/Ionomycin stimulation. Poly-functionality of T cells was analyzed by SPICE and compared SLAMF7+ with SLAMF7- CD4 (CD154+) and CD8 (CD154-) T cells. The pies represented the frequencies of T cells with a defined combination of effector molecules. The arcs above the pie indicate which cytokines or cytolytic molecules are expressed by each slice of the pie. (B) Effector signature of SLAMF7+ T cells during aging. Effector Memory (CD45RO+CD27-) T cells from young (n = 5) and older donors (n = 3) were sorted and stimulated overnight with PMA/Ionomycin. Supernatants were analyzed by Luminex and visualized by PCA. The contribution of individual cytokines to the main components was also indicated. (C) Effector signature of SLAMF7+ T cells during aging. Sorted SLAMF7+ T cell subsets were directly lysed ex-vivo. Gene expression was quantified by customized senescence Nanostring, normalized by Z-score calculation, and represented by a cold-to-hot heat map. (D) Transcriptional signature of SLAMF7+ T cells during aging. Top50 of differentially expressed genes between SLAMF7-expressing EM T cells was visualized by PCA. The contribution of individual genes to the main principal components was also indicated. (E) Exhaustion and transcription factors profile of SLAMF7+ T cells during aging. The SLAMF7-specific gene expression assessed as in (D), was represented by a cold to hot heat map for genes associated with activation/inhibitory molecules (left) and TF/signaling pathways (right).
Subsequently, we used CD57, as an established marker of terminal T-cell differentiation, to isolate and stimulate CD57+SLAMF7+ (as Terminal effector) and CD57-SLAMF7+ (as Progenitor-like) CD8 T cells in order to assess the production of cytokines, chemokines and cytotoxic molecules, including IFN-γ, MIP-1α, MIP-1β, IL-2, Granzyme B, and Perforin. The main difference was observed between CD57+SLAMF7+ and CD57-SLAMF7+ CD8 T cells but was not significantly influenced by aging, Fig. 2B. While aging did affect CD57-SLAMF7+ CD4 T cells, no significant differences in the effector functions were noted between both SLAMF7+ CD4 T cell subsets from old patients, Fig. S3D.
Next, we examined the transcriptional profiles of SLAMF7+CD57+ T cells during aging, Fig. 2C. Due to the limited number of SLAMF7+CD57+ CD4 T cells in young donors, we focused on SLAMF7+CD57- effector memory (EM) CD4 T cells from young donors. Our analysis revealed a shift from a helper (IL-4, IL-5, IL-9, IL-13, IL-17A, IL-17F, IL-22, IL-2, IL-3) to cytotoxic phenotype (Perforin, Granzyme B, Granzyme H) in older individuals. For SLAMF7+CD57+ CD8 T cells, cytotoxic signature was largely maintained during aging, with the exception of TNF expression.
We also identified the top 50 differentially expressed genes in SLAMF7+ EM T cells during aging, Fig. 2D. PCA revealed significant changes in SLAMF7+CD57+ CD8 T cells during aging, associated with the expression of genes such as TNF, TNFRSF9, TNFAIP3, JUN, LAG3, CD160, SLAMF6, hTERT, CX3CR1 and ITGA4/5 (Fig. 2D-E).
SLAMF7 expression in virus-specific T cells
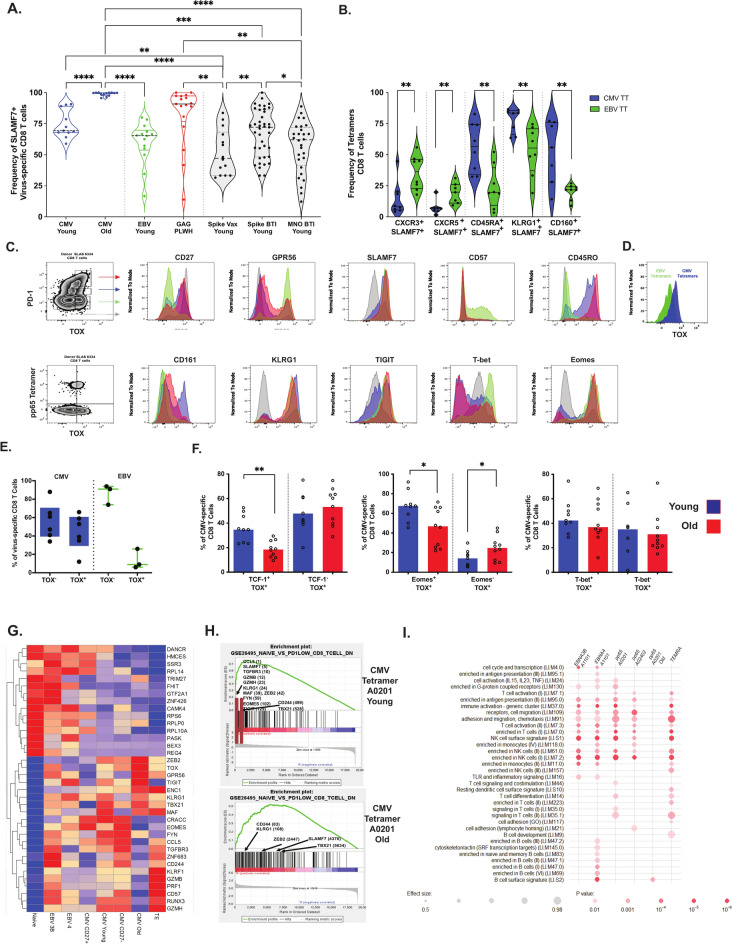
We then explored the role of SLAMF7 in virus-specific T cells during aging, vaccination, and viral infections (HIV, SARS-CoV-2), Fig. 3A. SLAMF7 was predominantly expressed on CMV-, GAG-, Spike- or MNO-specific CD8 T cells in HD, PLWH, and individuals after breakthrough infection respectively. Given the heterogeneity of SLAMF7+ T cells, we combined SLAMF7 with other markers to further define these virus-specific T cells, Fig. 3B. For EBV-specific CD8 T cells, SLAMF7 was associated with CXCR3 or CXCR5, while CMV-specific CD8 T cells expressed higher levels of CD45RA, KLRG1, or CD160 (p < 0.01), Fig. 3B.
Fig. 3.
SLAMF7 expressions in virus-specific T cells during aging. (A) SLAMF7 expression in virus-specific CD8 T cells. The expression of SLAMF7 was compared between virus-specific CD8 T cells in HD during aging (n = 8 for young and n = 11 for elderly), in PLWH (n = 17, including 10 non-treated and 7 under HAART), after SARS-CoV-2 vaccination (n = 14, 5 months post dose 2) and after SARS-CoV-2 breakthrough infection (n = 37, 3 weeks post-infection). The statistical analysis was performed on unpaired samples (U Mann–Whitney test) (*, **, ***, **** for p < 0.05, p < 0.01, p < 0.001, and p < 0.0001 respectively). (B) Specific SLAMF7 signature in virus-specific CD8 T cells. CMV- and EBV-specific CD8 T cells were statistically analyzed on unpaired samples (U Mann–Whitney test) (** for p < 0.01). (C) TOX expression in CD8 T cells. A zebra plot according to TOX and PD-1 or pp65-specific tetramer expressions represented CD8 T cells from an old donor. Gates were defined according to PD-1 and TOX expression as PD-1-TOX- (grey), PD-1-TOX+ (green), PD-1dimTOX+ (blue), and PD-1+TOX+ (red). The respective expressions of surface and intracellular markers were overlaid on histograms. (D) Specific TOX expression in CMV-specific CD8 T cells. The representative expression of TOX in CMV- and EBV-specific CD8 T cells from the same donor was overlaid on the histogram and the frequencies of TOX expressing were represented. (E) Quantification of TOX expression in virus-specific CD8 T cells. (F) Transcription factors signature of CMV-specific CD8 T cells during aging. The frequencies of CMV-specific CD8 T cells expressing TOX with and without TCF-1/7, Eomes, and T-bet were evaluated and compared in young (n = 9) and older donors (n = 10) (Mann–Whitney test, with * and ** for p < 0.05 and p < 0.01 respectively). (G) Transcriptional signature of CMV-specific CD8 T cells during aging. Total and subsets of CMV-specific CD8 T cells were sorted from 10 young and 3 old healthy donors. The extracted transcripts were analyzed by RNA sequencing. The differentially expressed genes were clustered and visualized by a cold-to-hot heat map representing the gene expression intensity. Naïve and Terminal effector T cells were used as internal controls. (H) Non-exhaustion T cell signature of CMV-specific CD8 T cells. The enrichment of gene expression detected in CMV-specific CD8 T cells was calculated in comparison to the naïve T cell signature. Enrichment plot of the gene set reported by GSEA as most enriched among PD-1low gene sets (GO: 26,495). The profile shows the enrichment score (green curve) and positions of gene set members (black vertical bars) rank-ordered list of differential gene expression. (I) Transcriptional pathway analysis of virus-specific CD8 T cells during aging. Ingenuity pathway analysis of bulk RNASeq data from virus-specific CD8 T cells. The enrichment of a specific pathway is proportional to the bubble size and the significance is associated with the different shades of red (from p < 10–2 to p < 10–6).
To determine if the heterogeneity of SLAMF7 expression was regulated by specific master genes, we focused on transcription factor signatures in virus-specific CD8 T cells. We analyzed the expression of Eomes, T-bet, TCF-1, BATF29, and TOX, which have been previously implicated in T cell exhaustion in both animal models and human viral infections30–34, Figure S4A. Segregating CD8 T cells by TOX and PD-1 expression (Fig. 3C), we defined TOX+PD-1Bright, TOX+PD-1Dim, and TOX+PD-1Neg CD8 T cells. Interestingly, TOX-negative CD8 T cells lacked SLAMF7 expression, while TOX+PD-1Neg CD8 T cells were characterized by GPR56, CD57, and T-bet expressions. Both exhausted and terminal effector CD8 T cells shared TOX expression. Extending this analysis to antigen-specific T cells (Fig. 3C-D), we found that CMV- specific CD8 T cells expressed higher levels of TOX compared to EBV-specific CD8 T cells (Fig. 3E).
Next, we wondered whether aging can impact on transcription factor expression in virus -specific CD8 T cells. In CMV-specific CD8 T cells, we observed age-related decreases in TCF-1/ TOX (p < 0.01) and TCF-1/ Eomes (p < 0.05) co-expression, Fig. 3F and Figures S4A-B. Moreover, CMV-specific CD8 T cells expressing CD27 and GPR56 were inversely modulated during aging, preferentially expressing TCF-1 and TOX respectively (Figure S4C). Due to the regulation of other genes either at a transcriptional or epigenetic level by these transcription factors, we used RNA sequencing to provide a comprehensive signature of antigen-specific T cells associated with SLAMF7 expression during aging.
To better distinguish Progenitor-like and Terminal effector CD8 T cells, we used CD27 as a marker to sort CMV-specific CD8 T cells into defined subsets. Naïve (CD27+CD45RO-) and terminally differentiated (CD27-CD45RO-CD57+) CD8 T cells were included as controls (Fig. 3G). Our analysis revealed SLAMF7 mRNA expression in CMV-specific CD8 T cells from young donors, but its presence was scarce barely in young EBV-specific CD8 T cells and CMV-specific CD8 T cells from older donors. The CMV-specific CD8 T cells in older donors exhibited greater differentiation, characterized by the expression of transcription factors such as ZEB2 and TBX21, which drive terminal differentiation and TOX, indicative of exhaustion and senescence. Interestingly, although TOX mRNA was also detected in CMV-specific CD8 T cells from younger donors, its expression was notably lower compared to older donors. Additionally, in older donors, there was an absence of key markers like RUNX3, ZNF683 (Hobit), EOMES, or CD57, suggesting an alternative differentiation pathway associated with aging. To further explore these findings, we applied Gene Set Enrichment Analysis (GSEA) to compare the transcriptional signature of CMV-specific CD8 T cells with public immune signatures. The transcriptional profile of CMV-specific CD8 T cells from young donors was significantly enriched for PD-1low CD8 T cells (vs Naïve, GSE26495) whereas in older individuals, only limited enrichment was observed (Fig. 3H). SLAMF7 emerged as one of the top 40 genes in the PD-1low CD8 T cells signature, corroborating our observations of TOX+PD-1Neg CD8 T cells. Further analysis of Blood Transcription Modules (BTM) revealed functional gene modules significantly expressed in CMV-specific CD8 T cells, highlighting pathways related to T cell activation and NK cell signatures (Fig. 3I). Notably, these pathways were diminished in older donors. An analysis of cellular functions associated with genes expressed in CMV-specific CD8 T cells (compared with naïve and terminal effector controls) showed a stronger activation of apoptosis pathways in older donors (Figure S4D).
Finally, we hypothesized that the loss of “SLAMF7 signature” in CMV-specific CD8 T cells may result from the disappearance of “younger” clones over time. To test this, we assessed the clonal diversity of TCR sequences in CMV-specific CD8 T cells. In comparison to naïve CD8 T cells, CMV-specific CD8 T cells displayed reduced TCR clonal diversity, particularly in older donors, Figure S4E. This trend was even more pronounced in the CD27- CMV-specific CD8 T cells, which exhibited increased oligo-clonality, Figure S4F. Thus, CMV-specific CD8 T cells undergo significant alterations in phenotype, transcriptional profiles, and TCR repertoire during aging.
SLAMF7 signaling during aging
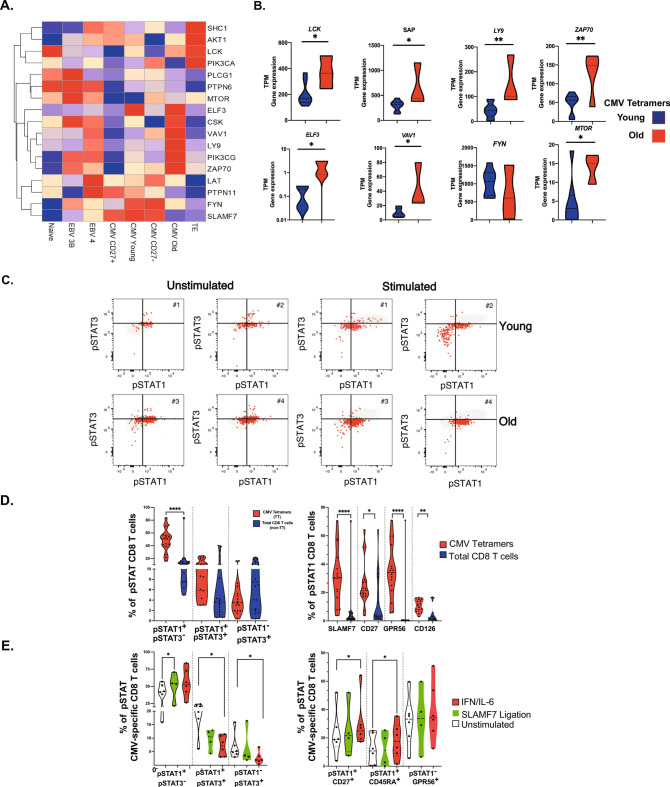
We focused on SLAMF7 signaling within the SLAM pathway to determine whether this pathway is altered during aging, Fig. 4A. ELF3, CSK, VAV1, LY9, PIK3CG, and ZAP70 mRNA were overexpressed in CMV-specific CD8 T cells from older donors. Significant up-regulation of LCK (p < 0.05), SAP (p < 0.05), ZAP70 (p < 0.01), or LY9 (p < 0.01) was observed, Fig. 4B. Interestingly, many of these transcripts did not correlate with other SLAM signaling components, such as SLAMF7, FYN, or PTPN11, suggesting a non-classical SLAM pathway in CMV-specific CD8 T cells during aging (Fig. 4A).
Fig. 4.
SLAMF7 signaling during aging. (A) SLAM signaling pathway in CMV-specific CD8 T cells. RNA Sequencing was used to evaluate the mRNA expression of constituent molecules of this pathway. The scaled gene expression intensity was visualized by a cold-to-hot heat map. (B) Modulation of the SLAM signaling pathway in CMV-specific CD8 T cells during aging. The gene expression was quantified by bulk RNA Sequencing in total or CD27 subsets of CMV-specific CD8 T cells from young or older donors (n = 10 and n = 3 respectively) and expressed as transcript per Million. (C) STAT1/3 signaling pathway in CMV-specific CD8 T cells. Flow cytometry was used to evaluate the constitutive (unstimulated) and IFNγ/IL-6-induced phosphorylation of STAT1 and STAT3 in CMV-specific CD8 T cells (red) and total CD8 T cells (grey) from young and older donors. Dot plots from four representative patients are described. (D) Quantification of STAT1/3 phosphorylation in CMV-specific CD8 T cells. The combination of phosphoSTAT1 (pSTAT1) and pSTAT3 was compared between total and CMV-specific CD8 T cells, left. The co-expression of pSTAT1 with phenotypic markers such as SLAMF7, CD27, GPR56, and CD126 was evaluated for total and CMV-specific CD8 T cells, right. The statistical analysis was performed on paired samples (n = 16, Wilcoxon signed-rank test), (*, **, and **** for p < 0.05, p < 0.01, and p < 0.0001, respectively). (E) SLAMF7 and STAT1 signaling in CMV-specific CD8 T cells. The pSTAT1 and pSTAT3 were evaluated after SLAMF7 ligation in CMV-specific CD8 T cells. The co-expression of pSTAT1 and pSTAT3 or pSTAT1 with phenotypic markers were analyzed as in (D).
Given our earlier findings regarding Wnt/β-catenin pathway disruption during aging35, we hypothesized that inflammation from chronic viral infection may affect the differentiation and gene expression of virus-specific T cells. Inflammatory cytokines such as IFN, TNF, or IL-6 regulated immune responses by activating the Janus kinase/signal transducers and activators of the transcription (JAK/STAT) pathway. Ex-vivo analysis of phosphorylation status revealed constitutive STAT1 and STAT3 activation in CMV-specific CD8 T cells and total CD8 T cells, Fig. 4C. The majority of CMV-specific CD8 T cells expressed pSTAT1 but not pSTAT3, in contrast to total CD8 T cells (p < 0.0001), irrespective of age, Fig. 4D. pSTAT1 was mainly detected in SLAMF7+ CMV-specific CD8 T cells, (p < 0.0001), as well as in GPR56+ (p < 0.0001), and IL-6R+ subsets (p < 0.01), Fig. 4D.
Next, we assessed the responsiveness of T cells to pro-inflammatory cytokines IFNγ and IL-6, Fig. 4C and Figures S5A-B. No significant increase in pSTAT1 and pSTAT3 was observed in CMV-specific CD8 T cells following stimulation, Fig. 4E. However, a significant reduction in pSTAT3 was noted after stimulation (p < 0.05). Interestingly, only CD27+CMV-specific CD8 T cells were responsive and expressed pSTAT1 after stimulation (p < 0.05), Fig. 4E. Finally, we also investigated whether SLAMF7 ligation could activate virus-specific T cells independently of TCR stimulation36, Figures S5A-B. SLAMF7 engagement led to increased pSTAT1 expression but not pSTAT3 in CMV-specific CD8 T cells (p < 0.05), Fig. 4E. Despite SLAMF7 engagement, there was a limited enhancement in the functional response of CMV-specific T cells when combined with TCR stimulation, Figure S5C.
Homeostatic regulation of SLAMF7+ CD8 T cells
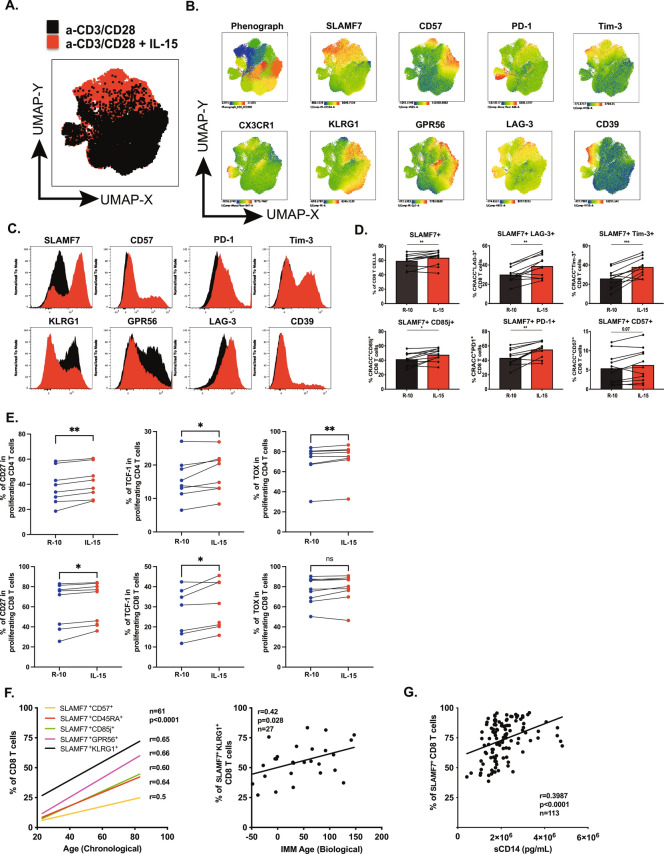
Next, we sought to decipher the mechanism regulating SLAMF7 expression. Since effector CD8 T cells lack surface IL-7R, their long-term survival depends on IL-15. We investigated whether IL-15 could regulate SLAMF7 expression during TCR stimulation (Fig. 5A). SLAMF7 up-regulation was associated with increased expression of inhibitory receptors, such as PD-1 (p < 0.01), Tim-3 (p < 0.001), LAG-3 (p < 0.01) and CD85j (p < 0.05) but not CD39, CD57, KLRG1, GPR56 or CX3CR1 (Fig. 5B–D). We hypothesized that IL-15 might favor the expansion of Progenitor-like effector CD8 T cells rather than Terminal effector CD8 T cells. To test this, we measured the proliferation of CellTrace™ Violet-stained CD8 T cells in response to TCR stimulation, with or without IL-15 (Fig. S6A,B). We confirmed that CD27+CD8 T cells proliferate, whereas CD57+ KLRG1+ CD8 T cells did not (Figure S6A). SLAMF7+CD57- showed higher proliferation than SLAMF7+CD57+ CD8 T cells (Figure S6C) (p < 0.0001) and IL-15 enhanced proliferation in all subsets (p < 0.05 for all subsets). Further, CD27+TCF-1+ CD8 T cells proliferated more after SLAMF7 ligation than IgG control (p < 0.05), Fig. 5E, suggesting that SLAMF7+CD57- CD27+TCF-1+ cells represent a Progenitor-like effector subset able to proliferate in response to TCR or SLAMF7 ligation and IL-15 stimulation.
Fig. 5.
Homeostatic regulation of SLAMF7+ CD8 T cells. (A) Distribution of IL-15/ TCR stimulated CD8 T cells. Total CD8 T cells from stimulated PBMCs (n = 12) were concatenated and visualized by UMAP. The phenotype was measured after 4 days of stimulation in the presence of anti-CD3 microbeads with and without IL-15 (10 ng/mL). TCR-stimulated and TCR/IL-15-stimulated CD8 T cells were overlaid on the UMAP representation in black and red respectively. (B) Phenotype of IL-15/ TCR stimulated CD8 T cells. UMAP visualized the phenotype of activated CD8 T cells as in (A). Clusters were automatically determined by phenograph. A cold-to-hot heatmap represented the scaled intensity of each marker. (C) SLAMF7-associated phenotype of IL-15/ TCR stimulated CD8 T cells. The fluorescence intensity expression of each marker was overlaid for both conditions. (D) Acquisition of SLAMF7 and inhibitory receptors in response to IL-15/ TCR stimulation. The frequency of total and SLAMF7 expressing CD8 T cells was measured after 4 days of in vitro stimulation. The statistical analysis was performed on paired samples (Wilcoxon signed-rank test) (n = 12 and *, **, *** for p < 0.05, p < 0.01 and p < 0.001 respectively). (E) SLAMF7 signaling and T cell proliferation. The proliferation was measured by the dilution of Cell Trace Violet (CTV) after 4 days of IL-15/ TCR stimulation with and without SLAMF7 ligation. The expression of CD27, TCF-1, and TOX were quantified on proliferating T cells, defined as CTVlow. The statistical analysis was performed on paired samples (Wilcoxon signed-rank test) (n = 8 and *, ** for p < 0.05, and p < 0.01 respectively). (F) Accumulation of SLAMF7+ CD8 T cells during aging. The correlations between the frequencies of CD8 T cell populations and the chronological age were evaluated by the Spearman test, left. The IMM Age was calculated according to the expression of 57 pre-determined genes detected by RNA-Seq and correlated to the frequencies of SLAMF7 expressing CD8 T cells in the ATTRACT cohort (n = 27, r = 0.42 and p < 0.05). (G) Accumulation of SLAMF7+ CD8 T cells during inflammation. The quantification of sCD14 was performed by Elisa to measure the systemic inflammation in the plasma of PLWH. The correlation between the frequency of SLAMF7+CD8 T cells and the concentration of sCD14 was evaluated by a Spearman test.
Whether IL-15 can regulate CD8 T cells in vivo as observed in centenarians37, is still an unanswered question. To unravel the dynamic of cellular differentiation through the lifespan, we assessed whether SLAMF7 expression could serve as a marker for immune aging. In two independent cohorts, the decline of Naïve CD8 T cells was associated with an accumulation of Terminally effector differentiated cells re-expressing CD45RA (TEMRA), Fig. 5F, and Figure S6D. SLAMF7+ GPR56+ and SLAMF7+ KLRG-1+ CD8 T cells correlated with chronological age in both healthy and non-healthy donors (with heart failure-related co-morbidities). The frequency of SLAMF7+ KLRG-1+ CD8 T cells was identified as a robust predictor of biological age, termed immune age38 (IMM-Age) (Fig. 5F). Notably, inflammation is a hallmark of co-morbidities during aging (Figure S6E). An increase of sCD14 is observed during aging and correlated with inflammatory markers (TNF, IL-12p70) or IDO activity (Figure S6F). The concentration of sCD14 was positively associated with the frequency of SLAMF7+ CD8 T cells in PLWH (Fig. 5G). Further validation is required, but SLAMF7 expression holds potential as a biomarker to guide personalized therapy in aged populations.
Discussion
The primary objective of this study was to deepen our understanding of the ontogeny of terminally differentiated CD8 T cells by delineating their heterogeneity using well-established markers such as CD27, CD57, GPR56, KLRG1, and the less explored SLAMF7. CD8 T cell diversity is closely associated with their immune history, and we analyzed their differentiation dynamics across diverse cohorts, including healthy young and aged donors, individuals with chronic HIV, or latent infections (EBV and CMV), and those with comorbidities.
Our findings revealed an early acquisition of SLAMF7 during CD8 T cell differentiation, challenging the classification of some CD8 T cells as “senescent” based solely on CD27 or CD28 expression loss. To refine the characterization of early versus late stage CD8 T cell differentiation, we propose SLAMF7, CD27, and GPR56 as key markers to distinguish progenitor-like from terminal effector T cells, showing near-exclusive expressions and enabling more precise classification.
A unique aspect of this study involved documenting both surface markers (CD5, CD28, CD127, KLRG1, CX3CR1, and CD57) and intracellular transcription factors (TCF-1, TOX, Eomes, T-bet, RUNX3, and HOBIT) to provide a comprehensive phenotypic characterization of CD8 T cell subsets. For the first time in the context of aging, we offer a detailed signature of TFs in human T cells. TCF-1 expression was linked to CD27+ subset, while TOX was more prominent in GPR56+ subset, indicating possible roles in initiating and maintaining effector differentiation. TF expression was tightly regulated in antigen-specific CD8 T cells, with RUNX3 and Eomes emerging early and T-bet and HOBIT appeared later. This suggest that RUNX3 and Eomes may initiate differentiation39 while T-bet and HOBIT sustain and lock it down40,41. This hypothesis warrants validation in animal models to identify therapeutic windows for inducing a robust and long-lasting effector immune response. Second, there was preferential expression of TFs in CD8 T cell subsets with different functions. Terminal Effector CD8 T cells were indeed characterized as T-bethighEomeslow, while exhausted CD8 T cells were characterized as T-betlowEomeshigh, consistent with previous findings42,43. Additionally, the co-expression of TOX30,31,34,44 and TCF-1 indicated specialized functions, as seen in CD8 T cells, which act as precursors of exhausted (TPEX) T cells during chronic infections45 and cancer46–48. The stemness properties of TPEX enabled them to proliferate and restore robust T-cell immunity after immune checkpoint blockade. Similarly, during chronic viral infection (including HCV and HIV)30,49 and cancer, naïve and central memory CD8 T cells expressed TCF-150,51 but not TOX, while effector and exhausted CD8 T cells were TCF-1-TOX+, implying that the continuous TCR activation due to the persistence of viral or tumor antigens enhanced T cell differentiation and TOX acquisition. In previous reports, the co-expression of TCF-1 and TOX was differentially detected in human CMV, EBV, FLU, and HIV-specific CD8 T cells49. We confirmed earlier observation that the frequency of TCF-1 expressing CMV-specific CD8 T cells decreases with age, which may compromise immunity against latent infections.
In CMV-specific CD8 T cells from older donors, TOX protein expression remained elevated, correlating with terminal effector genes (ZEB2, TBX21, GPR56, and KLRG1) but not with effector functions genes (GZMB, PRF1, GZMH) or effector TFs such as RUNX3 or ZNF683 (HOBIT). It is known that TOX auto-regulates its transcription and also controls other TFs, such as Tcf7 or Nr4a2. However, the regulation of epigenetic and transcriptional activity during aging is still poorly understood, and further studies are needed to clarify TOX’s role in immune function during aging.
SLAMF7 isoforms, originally described in NK cells52, were found to play a significant role in T cells as well. The ratio between long and short SLAMF7 variants, which impacts signaling, has been described in autoimmune disease like SLE53, but its role in aging is yet to be determined. Further research is required to reconcile the observed in vivo systemic activation of monocytes (sCD14), which correlates with SLAMF7 expression on CD8 T cells during aging, with the inhibitory effects of SLAMF7 ligation on activated monocytes proinflammatory cytokines secretion in vitro54.
In PLWH undergoing anti-retroviral therapy, we observed an increase of CD127 (a marker of long-term immune memory), and a decrease in inhibitory receptors (Tim-3, Caecam-1)55 as well as TFs associated with terminal differentiation and exhaustion (T-bet/ Eomes). However, SLAMF7 expression on CD8 T cells, including HIV-specific CD8 T cells, remained elevated. The increased TOX expression during HIV infection suggests these molecules as potential therapeutic targets to rejuvenate immunity in PLWH. Chronic infection histories and persistent inflammation during aging leave deep imprints on immune signaling pathways, potentially disrupting Wnt/β-Catenin family signaling, including TCF1 and therefore stemness of T cells35. We also detected the modulation of genes associated with the SLAM family in CMV-specific CD8 T cells from older individuals. Therefore, SLAMF7 or TOX blockade alone might not be sufficient to restore full pathways functionality.
We also explored whether directly targeting SLAMF7 might enhance immune function, particularly by expanding or boosting progenitor-like effector CD8 T cells to differentiate into competent effector T cells. SLAMF7 engagement may enhance the immune surveillance for persistent antigens such as in cancer or chronic infections, as observed in autoimmune diseases like SLE23, polymicrobial sepsis56 and chronic inflammation (Crohn’s disease, severe COVID-19)57. Additionally, novel therapies like CAR T cells or Elotuzumab58 have shown success in targeting SLAMF7 in Multiple Myeloma (MM). Interestingly MM patients with high frequencies of SLAMF7+CD8 T cells have diminished anti-tumor responses59, suggesting a broader potential application for SLAMF7-targeting therapies. The proof of concept of vaccination with SLAMF7-derived peptides to directly stimulate anti-tumoral CD8 T cells and therefore to induce the lysis of MM cells has been performed only in vitro60. The agonist engagement of SLAMF7 can directly boost the cytotoxicity of tumor-specific CD4 T cells61, but we only observed a slight increase in the cytolytic potential of CMV-specific CD8 T cells from HDs after SLAMF7 engagement.
IL-15 stimulation has been shown to drive the proliferation of SLAMF7+CD27+ CD8 T cells, with TCF-1 expression indicating a progenitor-like state. Given IL-15’s role in supporting naïve and memory CD8 T cells in animal models62,63, it represents another therapeutic avenue, alongside SLAMF7 engagement, to promote robust T cell immunity in humans. The anti-tumor activity of TSCM CD8 was also enhanced by IL-1564. IL-15 could also drive the differentiation of tissue-resident CD8 T cells65, expand CX3CR1+CD57+ CD8 T cells or Granzyme B+ CD8 T cells in PLWH, and maintain terminally differentiated CD8 T cells in the bone marrow66.
In summary, we identified SLAMF7 as a surrogate marker for CD57, which along with CD27, GPR56, and transcription factors such as TOX, TCF-1 and Eomes, serves as a robust tool for identifying effector T cell subsets. The fine regulation of these markers and their interactions across different immune histories—aging, persistent viral infection, and chronic inflammation—may represent promising therapeutic targets for restoring or enhancing immune function in aging populations or individuals with chronic infections.
Methods
Donors and sample preparation
Blood was collected from participants of the Singapore Longitudinal Aging Study (SLAS) cohort. Characteristics of the SLAS cohort are detailed in our previous publication, related to physical frailty67. Blood was collected into BD Vacutainer CPT Cell Preparation tubes with Sodium Citrate (BD Biosciences, San Jose, CA, USA). After centrifugation at 1600 rcf for 20 min at room temperature, plasma and peripheral blood mononuclear cells (PBMCs) were isolated. Plasma was stored at − 80 °C before use. PBMCs were frozen in 90% foetal bovine serum (FBS) containing 10% DMSO and stored in liquid nitrogen.
The study has been approved by the National University of Singapore-Institutional Review Board 04–140 and all participants gave informed consent. Young donors were recruited at the National University of Singapore. The study has been approved by the Ethics Committee of the NUS-IRB 09–256. All study participants provided informed written consent.
Blood was collected from HIV-infected individuals attending the University Malaya Medical Centre (UMMC), Malaysia,35 and the service of infectious disease from Virgen del Rocio Hospital, Sevilla, Spain. Data on HIV-specific characteristics including HIV RNA, CD4 T cell counts, antiretroviral drug history, and history of co-infections were obtained from patient medical records (Table 1). The study was approved by the hospital institutional review board for Malaysian HIV-infected patients (MEC 975.6) and the hospital institutional review board for Spanish HIV-infected patients (2012PI/240). All study participants provided informed written consent.
Table 1.
Clinical characteristics of HIV-infected donors (Spain).
| # patient | Patient ID |
Age | CD4 | CD8 | HIV Viral Load | CD4 Nadir |
Gender | HCV coinfection |
|---|---|---|---|---|---|---|---|---|
| No Anti-Retroviral Therapy | ||||||||
| 1 | 20385 | 45 | 204 | 1334 | 75900 | 204 | Male | Yes |
| 2 | 22369 | 41 | 358 | 594 | 172000 | 291 | Female | No |
| 3 | 23612 | 45 | 501 | 438 | 9850 | 390 | Female | No |
| 4 | 19263 | 38 | 319 | 1444 | 116011 | 319 | Male | No |
| 5 | 23060 | 45 | 404 | 1937 | 64000 | 404 | Male | No |
| 6 | 22468 | 41 | 249 | 458 | 18400 | 249 | Female | Yes |
| 7 | 24856 | 51 | 485 | 1056 | 23200 | 485 | Male | No |
| 8 | 19978 | 46 | 224 | 306 | 201000 | 168 | Male | No |
| 9 | 22786 | 42 | 280 | 646 | 166000 | 268 | Male | No |
| 10 | 27109 | 54 | 20 | 256 | 1200000 | 20 | Female | Yes |
| 11 | 23418 | 45 | 262 | 912 | 6900 | 262 | Male | No |
| 12 | 21772 | 45 | 375 | 1410 | 41200 | 364 | Male | Yes |
| 13 | 27232 | 46 | 326 | 1039 | 68800 | 326 | Male | No |
| 14 | 26172 | 40 | 565 | 2203 | 24300 | 522 | Male | No |
| 15 | 22759 | 40 | 600 | 884 | 3650 | 542 | Male | No |
| 16 | 25991 | 70 | 454 | 1403 | 81600 | 396 | Male | No |
| 17 | 25884 | 52 | 549 | 722 | 55200 | 549 | Female | No |
| 18 | 26451 | 40 | 609 | 944 | 2710 | 606 | Male | No |
| 19 | 26228 | 39 | 383 | 606 | 123000 | 383 | Male | No |
| 20 | 26003 | 37 | 633 | 908 | 13200 | 468 | Male | No |
| 21 | 25989 | 35 | 350 | 574 | 8460 | 350 | Male | No |
| Anti-Retroviral Therapy | ||||||||
| 1 | 24011 | 47 | 386 | 1075 | 20 | |||
| 2 | 27251 | 45 | 697 | 576 | 24 | |||
| 3 | 26147 | 47 | 695 | 364 | 20 | |||
| 4 | 22118 | 40 | 578 | 1382 | 20 | |||
| 5 | 27654 | 48 | 621 | 943 | 20 | |||
| 6 | 26497 | 44 | 278 | 566 | 20 | |||
| 7 | 27200 | 54 | 748 | 1144 | 20 | |||
| 8 | 22385 | 48 | 590 | 306 | 20 | |||
| 9 | 27939 | 46 | 330 | 284 | 20 | |||
| 10 | 29258 | 56 | 199 | 638 | 20 | |||
| 11 | 26911 | 48 | 605 | 557 | 139 (blip) | |||
| 12 | 28648 | 50 | 410 | 1160 | 71 | |||
| 13 | 29898 | 48 | 761 | 739 | 50 | |||
| 14 | 28328 | 42 | 754 | 1236 | 20 | |||
| 15 | 25907 | 42 | 647 | 805 | 20 | |||
| 16 | 27560 | 72 | 817 | 1940 | 20 | |||
| 17 | 26792 | 53 | 812 | 501 | 20 | |||
| 18 | 28733 | 42 | 737 | 601 | 20 | |||
| 19 | 28216 | 41 | 547 | 732 | 20 | |||
| 20 | 28443 | 39 | 563 | 516 | 20 | |||
| 21 | 28068 | 36 | 644 | 804 | 20 | |||
Cryo-preserved PBMCs of patients included in the seasonal Influenza vaccination (Table 2) or the ATTRACT cohort (Table 3) were used at baseline, randomized, and analyzed simultaneously. All study participants provided informed written consent.
Table 2.
Clinical characteristics of aging healthy donors (Singapore, SLAS cohort).
| Patient ID | Age | Gender | Frailty Status |
|---|---|---|---|
| SNF001 | 67 | Male | Robust |
| SNF020 | 75 | Female | Pre-frail |
| SNF029 | 71 | Male | Pre-frail |
| SNF031 | 73 | Female | Robust |
| SNF039 | 73 | Female | Robust |
| SNF047 | 67 | Female | Robust |
| SNF049 | 73 | Male | Pre-frail |
| SNF050 | 26 | Female | Young |
| SNF052 | 33 | Female | Young |
| SNF054 | 31 | Female | Young |
| SNF055 | 26 | Male | Young |
| SNF060 | 34 | Female | Young |
| SNF061 | 26 | Female | Young |
| SNF062 | 27 | Female | Young |
| SNF064 | 66 | Male | Robust |
| SNF068 | 25 | Male | Young |
| SNF070 | 30 | Female | Young |
| SNF071 | 24 | Male | Young |
| SNF074 | 27 | Male | Young |
| SNF075 | 29 | Female | Young |
| SNF076 | 25 | Female | Young |
| SNF077 | 23 | Female | Young |
| SNF079 | 33 | Male | Young |
| SNF080 | 27 | Female | Young |
| SNF081 | 23 | Male | Young |
| SNF082 | 27 | Male | Young |
| SNF084 | 31 | Male | Young |
| SNF086 | 26 | Male | Young |
| SNF099 | 73 | Male | Pre-frail |
| SNF106 | 71 | Female | Robust |
| SNF113 | 81 | Female | Pre-frail |
| SNF143 | 70 | Female | Pre-frail |
| SNF185 | 76 | Male | Robust |
| SNF187 | 73 | Female | Pre-frail |
| SNF205 | 74 | Female | Pre-frail |
| SNF211 | 74 | Male | Robust |
| SNF215 | 66 | Female | Pre-frail |
| SNF223 | 69 | Male | Pre-frail |
| SNF227 | 76 | Male | Robust |
| SNF231 | 72 | Female | Pre-frail |
Table 3.
Clinical characteristics of aging nonhealthy donors (Singapore, ATTRACT cohort).
| Patient ID | Age | Diabetes | Hypertension | Hyperlipidemia | BMI | Gender | Ethnicit | Status |
|---|---|---|---|---|---|---|---|---|
| CSHOP0083A | 53 | N | N | ? | 23.2 | Female | Chinese | Control |
| CSHOP0776A | 38 | N | N | ? | 25.8 | Male | Chinese | Control |
| CSHOP0809A | 42 | N | N | ? | 22 | Male | Indian | Control |
| CSHOP0850A | 69 | N | Y | ? | 23.6 | Male | Chinese | Control |
| CSHOP0934A | 46 | N | N | ? | 26 | Male | Chinese | Control |
| CSHOP0950A | 51 | N | N | ? | 30.8 | Female | Indian | Control |
| CSHOP0999A | 36 | N | N | ? | 32.2 | Male | Indian | Control |
| SHOPNH0163 | 64 | N | N | N | 21.4 | Male | Chinese | HFrEF |
| SHOPNH0173 | 76 | Y | Y | N | 23.4 | Female | Indian | HFrEF |
| SHOPNH0174 | 62 | N | N | N | 33.5 | Female | Chinese | HFrEF |
| SHOPNH0185 | 68 | Y | Y | Y | 23.2 | Male | Chinese | HFrEF |
| SHOPNH0187 | 39 | Y | Y | Y | 33.8 | Male | Chinese | HFrEF |
| SHOPNH0188 | 37 | N | N | Y | 29.3 | Female | Malay | HFrEF |
| SHOPNH0193 | 51 | Y | Y | Y | 25.5 | Male | Chinese | HFrEF |
| SHOPNU0129 | 63 | Y | N | N | 44 | Female | Indian | HFrEF |
| SHOPNU0137 | 63 | N | Y | N | 29.6 | Male | Indian | HFrEF |
| SHOPNU0252 | 63 | Y | Y | N | 27.1 | Female | Chinese | HFrEF |
| SHOPNU0334 | 66 | N | Y | N | 33.7 | Female | Indian | HFrEF |
| SHOPSG0156 | 74 | N | Y | N | 23.3 | Male | Chinese | HFrEF |
| SHOPSG0165 | 50 | Y | Y | N | 25.1 | Male | Chinese | HFrEF |
| SHOPSG0172 | 49 | Y | N | N | 43.3 | Female | Indian | HFrEF |
| SHOPTT0050 | 66 | N | Y | N | 24 | Male | Chinese | HFrEF |
| SHOPTT0171 | 38 | N | Y | Y | 36 | Male | Chinese | HFrEF |
| SHOPTT0309 | 43 | Y | Y | Y | 29 | Male | Malay | HFrEF |
| SHOPKT0094 | 83 | N | N | N | 24.9 | Male | Chinese | HFrEF |
| SHOPNH0208 | 50 | N | Y | N | 28.6 | Female | Indian | HFrEF |
| SHOPNU0168 | 54 | Y | N | N | 27.8 | Female | Malay | HFrEF |
| SHOPNH0219 | 57 | N | Y | N | 19 | Female | Chinese | HFrEF |
| SHOPNH0006 | 49 | Y | Y | Y | 26.9 | Female | Chinese | HFrEF |
The demographic data of acute COVID-19 BTI have been published previously68. Blood sampling was processed 10 days after SARS-CoV-2 infection in double dose vaccinated patients (4 months since the last vaccine administration). Ethical approvals were obtained from regulatory committee authority (Approval numbers: REK 2021.233704; REK 2020.135924; REK 2021.229359). All study participants provided informed written consent.
All methods were performed in accordance with the relevant guidelines and regulations.
Phenotyping of PBMCs
Cell phenotyping was performed by flow cytometry on frozen PBMC samples from young and older donors. For each staining, 2 × 106 PBMCs were used. Lymphocytes were gated based on FSC/SSC profile and doublets/dead cell exclusion.
Absolute cell count was performed by flow cytometry on freshly collected blood of 10 healthy donors and 6 HAART HIV-infected patients. For each staining, 100 μL of blood was used in Trucount tubes (BD Biosciences). After doublets/dead cell exclusion, lymphocytes were gated based on FSC/SSC profile and CD45 expression.
The antibodies are listed in the Sup Table. Flow cytometry was performed on a BDSymphony A5 (BD Biosciences) and automatic compensation was applied.
Flow cytometry of HIV participants was performed on a BD FACS Celesta (BD Biosciences) at the University of Malaya and LSR Fortessa Cell Analyzer (BD Biosciences) at IBIS. Automatic compensations were applied.
Peptides
Pools of peptides corresponding to known optimal epitopes derived from CMV, EBV, and Flu, were purchased from Miltenyi Biotec, and HIV was purchased from Mimotopes. All peptides were synthesized at a purity of > 95%. Lyophilized peptides were reconstituted at 1 mg/ml in dimethyl sulfoxide (DMSO) and further diluted to 100 μg/ml in PBS.
Tetramers
MHC class I tetramer conjugated to FITC, BV421 or PE were used to detect CD8 + T cells with the following specificities: CMV NLVPMVATV (NV9/HLA-A*0201), EBV GLCTLVAML (GL9/HLA-A*0201), Flu GILGFVFTL (GL9/HLA-A*0201), HIV SLYNTVATL (SL9/HLA-A*0201). All tetramers were purchased from Immudex or Biolegend.
Spike-Specific CD8 T cells were detected using PE-conjugated Dextramers (Immudex) targeting Spike and restricted to HLA-A*0101 (LTDEMIAQY), HLA-A*0201 (YLQPRTFLL), HLA-A*2402 (QYIKWPWYI), and HLA-B*0702 (SPRRARSVA). The panel was expanded using Flex-T tetramer according to the manufacturer´s instructions (BioLegend). We UV-exchanged peptides for Spike epitopes restricted to HLA-A*0101 (YTNSFTRGVY), HLA-A*0201 (LITGRLQSL and RLNEVAKNL), HLA-A*2402 (NYNYLYRLF), and HLA-B*0702 (APHGVVFL) and tetramerized with Streptavidin-PE (Biolegend). A similar approach was performed for non-Spike derived epitopes, including HLA-A*0101 (ORF3a, FTSDYYQLY, and ORF1ab, TTDPSFLGRY), HLA-A*0201 (ORF3a, LLYDANYFL), HLA-A*2402 (ORF3a, VYFLQSINF), and HLA-B*0702 (Nucleoprotein, SPRWYFYYL) and tetramerized with Streptavidin-APC (Biolegend).
Flow cytometry functional assay
Proliferation
Freshly purified PBMCs were isolated. Cell Trace Violet-stained PBMC were stimulated with anti-CD3/CD28 micro-beads and IL-15 for 4 days. Proliferation of T cells was measured through the dilution of CTV.
ICS
Freshly purified PBMCs were stimulated with PMA/Ionomycin (1 µg/ml and 100 ng/ml respectively) or Cytostim or overlapping peptides (Pepmix, Myltenyi) for 4 h to detect the secretion of cytokines by CD8 T cells. Brefeldin A and Monensin (eBioscience) were added during the final 2 h incubation. For the list of antibodies used, refer to Sup Table 4. Flow cytometry was performed on a BD Symphony A5 (BD Biosciences).
Table 4.
List of antibodies.
| Name | Clone | Company | Dilution |
|---|---|---|---|
| Surface Staining | |||
| CCR5 | J418F1 | Biolegend | 1/25 |
| CCR7 | G043H7 | Biolegend | 1/25 |
| CD127 | HIL-7R-M21 | BD Biosciences | 1/50 |
| CD15s | CSLEX1 | BD Biosciences | 1/50 |
| CD160 | BY55 | BD Biosciences | 1/50 |
| CD161 | HP-3G10 | Biolegend | 1/50 |
| CD244 | C1.7 | Biolegend | 1/25 |
| CD27 | L128 | BD Biosciences | 1/50 |
| CD28 | CD28.2 | BD Biosciences | 1/100 |
| CD3 | UCHT1 | BD Biosciences | 1/50 |
| CD352 | REA 339 | Miltenyi | 1/25 |
| CD38 | HIT2 | BD Biosciences | 1/50 |
| CD39 | A1 | Biolegend | 1/50 |
| CD4 | OKT4 | BD Biosciences | 1/50 |
| CD45RA | HI100 | BD Biosciences | 1/100 |
| CD45RO | UCHL1 | Biolegend | 1/50 |
| CD5 | UCHT2 | BD Biosciences | 1/50 |
| CD56 | HCD56 | BD Biosciences | 1/50 |
| CD57 | HNK-1 | Biolegend | 1/100 |
| CD62L | DREG-56 | Biolegend | 1/50 |
| CD8 | OKT8 | BD Biosciences | 1/50 |
| CD85j | GHI/75 | Biolegend | 1/25 |
| CD95 | DX2 | BD Biosciences | 1/50 |
| CRACC | 162.1 | Biolegend | 1/50 |
| CX3CR1 | 2A9-1 | Biolegend | 1/50 |
| CXCR3 | G025H7 | Biolegend | 1/25 |
| CXCR5 | J252D4 | Biolegend | 1/50 |
| GPR56 | CG4 | Biolegend | 1/50 |
| HLA-DR | L243 | BD Biosciences | 1/100 |
| KLRG1 | 2F12/KLRG1 | eBioscience | 1/25 |
| LAG-3 | REA351 | Miltenyi | 1/10 |
| NKG2C | REA205 | Miltenyi | 1/10 |
| PAN GD | 11F2 | BD Biosciences | 1/25 |
| PD-1 | EH12.1 | BD Biosciences | 1/50 |
| TIGIT | 741182 | BD Biosciences | 1/50 |
| TIM-3 | F38-2E2 | Biolegend | 1/25 |
| Intra-cellular Intra-nuclear Staining | |||
| CD107a | H4A3 | Biolegend | 1/25 |
| CD154 | TRAP-1 | BD Biosciences | 1/25 |
| CTLA-4 | 14D3 | BD Biosciences | 1/25 |
| Eomes | WD1928 | eBioscience | 1/50 |
| Granulysin | DH2 | Biolegend | 1/25 |
| GZM A | CB9 | Biolegend | 1/25 |
| GZMB | GB11 | BD Biosciences | 1/100 |
| Hobit | ZNF683 | BD Biosciences | 1/25 |
| IFN-g | 4S.B3 | Biolegend | 1/50 |
| IL-17A | 64DEC17 | eBioscience | 1/25 |
| IL-2 | MQ1-17H12 | Biolegend | 1/50 |
| KI-67 | Ki-67 | BD Biosciences | 1/25 |
| Perforin | B-D48 | Biolegend | 1/100 |
| RunX3 | R3-5G4 | BD Biosciences | 1/25 |
| Sestrin-2 | D1B6 | Cell Signaling | 1/25 |
| T-bet | 4B10 | eBioscience | 1/50 |
| TCF-1 | 7F11A10 | Biolegend | 1/25 |
| TNF | MAb11 | BD Biosciences | 1/50 |
| TOX | TXRX10 | eBioscience | 1/10 |
Multiplex analytes screening
Cell sorting was performed with a FACS Aria III (BD Biosciences) on 10 samples according to CD27, CD45RO, CD57, and SLAMF7 expression in T cells. For the list of antibodies used for sorting, refer to Supplementary Table 4. After 18-h incubation with PMA/Ionomycin, supernatants were collected and tested by Luminex assay. The Milliplex HCD8MAG-15 K (Millipore) was used according to the manufacturer’s instructions and signal detected by Luminex™ FLEXMAP 3D®.
ELISA
Soluble CD14 (R&D Systems), was measured according to the manufacturer’s instructions. Plasma samples of HIV-infected patients and corresponding controls were Triton-inactivated and diluted in the appropriate buffer and assayed in duplicate.
nCounter human senescence v2
Direct mRNA expression levels of the samples were measured using the NanoString nCounter gene expression system. – 20,000 sorted CD8 T cell subsets in 5μL of RLT buffer from Qiagen RNeasy Mini kit (Qiagen, Hilden, Germany) were hybridized with probes from the nCounter Human Senescence v2 panel (Nanostring, Seattle, USA) at 65 °C for 16–19 h according to the nCounter™ Gene Expression Assay Manual. Excess probes were washed away using a two-step magnetic bead-based purification on the nCounter™ Prep Station (GEN1). The nCounter™ Digital Analyzer (GEN1) was used to count individual fluorescent barcodes and quantify target molecules present in each sample. For each assay, a high-density scan (600 fields of view) was performed.
RNAseq
Total RNA was extracted following the double extraction protocol: RNA isolation by acid guanidinium thiocyanate-phenol–chloroform extraction (TRIzol, Thermo Fisher Scientific, Waltham, MA, USA) followed by a Qiagen RNeasy Micro clean-up procedure (Qiagen, Hilden, Germany). All human RNAs were analyzed on Agilent Bioanalyzer for quality assessment with RNA Integrity Number (RIN) ranging from 6.2 to 9.6 and median RIN 8.9 (Agilent, Santa Clara, CA, USA). cDNA libraries were prepared using 1 ng of total RNA and 0.5ul of a 1:50,000 dilution of ERCC RNA Spike in Controls (Ambion Thermo Fisher Scientific, Waltham, MA, USA) using SMARTSeq v2 protocol except for the following modifications: 1, Use of 20 µM TSO; 2, Use of 250 pg of cDNA with 1/5 reaction of Illumina Nextera XT kit (Illumina, San Diego, CA, USA). The length distribution of the cDNA libraries was monitored using a DNA High Sensitivity Reagent Kit on the Perkin Elmer Labchip (Perkin Elmer, Waltham, MA, USA). All samples were subjected to an indexed PE sequencing run of 2 × 51 cycles on an Illumina HiSeq 2000 (16 samples/lane).
Seasonal influenza vaccination
The trivalent inactivated (split virion) Vaxigrip® (Sanofi Pasteur) 2013–14 seasonal influenza vaccine was used in this trial. The vaccine contained the three strains A/California/07/2009 (H1N1), A/Texas/50/2012 (H3N2), and B/Massachusetts/02/2012. Viruses were grown in embryonated chicken eggs, inactivated with formaldehyde, and split with anionic detergent. The vaccine was administered to the 205 elderly study participants between January and August 2014. Venous blood specimens were collected from the participants immediately before the vaccination (day 0) and on days 2, 7, and 28 after the vaccination.
Data analysis
Flow cytometry data were analyzed using FlowJo (Treestar) and FACSDiva (BD Biosciences). Samples were compared using GraphPad Prism software (v.8.0c). t-SNE analysis of flow cytometry data: unbiased representations of multi-parameter flow cytometry data were obtained using the uniform Manifold approximation and projection (UMAP) for dimension reduction. UMAP analysis was performed using FlowJo, custom R scripts, and Cytofkit software.
Gene expression
Paired-end 2 × 150 bp reads from Illumina sequencing were quality-checked using fastqc. Abundances of human transcripts obtained from gencode version 27 were estimated from the sequencing reads using Salmon version 0.8.2. Transcript abundances were summarized into gene counts using the tximport R/Bioconductor package. The gene abundances were normalized to transcript per million (TPM) and then log-transformed as log 2 (TPM + 1.0) with a pseudo-count of 1.0 added to prevent negative values. The normalized log transformed abundances were displayed in heatmaps where they were scaled to a Z-score in each row for coloring. Gene set enrichment analysis was performed using Broad Institute’s GSEA tool version 3.0 with immunologic signature gene sets (C7) collection of the MSIG database version 6.2 as the reference. The connectivity map (CMAP) algorithm was implemented using custom R scripts. Gene signatures used in CMAP analysis were defined by performing differential gene expression analysis using DESeq2 (for RNA-seq data) or limma (for microarray data) R/Bioconductor packages and then selecting differentially expressed genes based on a false discovery rate of 0.05. TCR repertoire analysis was performed with MiXCR where the same number of initial reads (8 million) was taken for analysis from each sample to normalize the results.
Statistical analysis
Groups of young and elderly donors were analyzed by the Mann–Whitney U test to compare values. The Wilcoxon matched-pairs signed rank test was used for paired testing of median values of different subsets from the same donor. Analysis with p < 0.05 (*), p < 0.01 (**), p < 0.001 (***) and p < 0.0001 (****) were considered significantly different between the groups.
Supplementary Information
Acknowledgements
We thank, Ivy Low (Flow Cytometry Platform), Esther Mok (Multiplex Analysis Platform), and Foo Shihui (Immunogenomics Platform) from SIgN Immunomonitoring Platform (BMRC IAF 311006 grant and BMRC transition funds #H16/99/b0/011). We thank, Nicolas Ang, from the Computational Immunology Platform for the kind assistance to RNAseq data deposition in GEO. We thank BD Biosciences and BioMarketing Services (BMS) for the generous loan of a BD FACS Celesta at the University of Malaya. We would like to acknowledge Dr. Robert Balderas (BD Biosciences) for his input in antibody panel design and high-dimension flow cytometry experiments. We thank Siri Mjaaland for the previous collaboration on COVID-19 breakthrough infection68.
Abbreviations
- ART
Anti-Retroviral Therapy
- BTM
Blood Transcription Modules
- CM
Central Memory
- CMV
Cytomegalovirus
- CTL
Cytotoxic T lymphocyte
- DEG
Differentially Expressed Gene
- EBV
Epstein-Barr virus
- EM
Effector Memory
- Flu
Influenza virus
- GSEA
Gene Set Enrichment Analysis
- HCV
Hepatitis C virus
- HD
Healthy Donors
- HIV
Human Immunodeficiency Virus
- ICS
Intra-Cellular Staining
- RNAseq
RNA Sequencing
- ROS
Reactive Oxygen Species
- SCM
Stem Cell Memory
- SLAM
Signaling Lymphocytic Activation Molecule
- SLE
Systemic Lupus Erythematous
- TCF7
Transcription factor 7
- TE
Terminal Effector
- TIL
Tumor Infiltrating Lymphocytes
- TM
Transitional memory
- TOX
Thymocyte selection-associated high mobility group box protein
- UMAP
Uniform Manifold Approximation and Projection
Author contributions
Conceptualization, H.K. and A.L; Experimental work, H.K., S.W.T., C.T, A.T.S.W., M.S, C.H.X. Samples acquisition, R.R., A.K., E.R.M., T.P.N.; Data analysis, H.K., V.N., M.S.; J.L.; Writing – original draft, H.K.; Writing – review and editing, A.L., V.N.; Supervision, H.K. and A.L.; Funding acquisition, A.L., E.R.M., T.P.N, R.R.
Funding
The study is supported by a research grant from the Agency for Science, Technology and Research (No. 10-036), by the Singapore Immunology Network A.L. is a scholar of the International Society for Advancement of Cytometry (ISAC). RR and AK are funded by the High Impact Research/Ministry of Higher Education Research Grant, Malaysia (HIR/MOHE; H-20001-E000001) and the RU grant (UMRG RP029-14HTM). E.R.M. was supported by Consejeria de Salud y Bienestar Social of Junta de Andalucıa through the Nicolas Monardes program (C-0032/17) and by Consejo Superior de Investigaciones Científicas (CSIC) and by the grants PI16/00684, PI19/01127 by Instituto de Salud Carlos III-FEDER. V.N. is supported by SIgN Immunomonitoring platform grants BMRC IAF 311006, BMRC transition funds#H16/99/b0/011, BMRC IAF-PP H1901a0024 and NRF SIS NRF2017_SISFP09.
Data availability
The authors declare that the data supporting the findings of this study are available within the paper [and its supplementary information files]. RNA sequencing data are accessible from NCBI’s Gene Expression Omnibus repository using the dataset ID GSE 282172. The data that support the other findings of this study are available from the corresponding author upon reasonable request.
Declarations
Competing interests
The authors declare no competing interests.
Footnotes
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
The online version contains supplementary material available at 10.1038/s41598-024-80971-5.
References
- 1.Brenchley, J. M. et al. Expression of CD57 defines replicative senescence and antigen-induced apoptotic death of CD8+ T cells. Blood101(7), 2711–2720 (2003). [DOI] [PubMed] [Google Scholar]
- 2.Kared, H., Martelli, S., Ng, T. P., Pender, S. L. & Larbi, A. CD57 in human natural killer cells and T-lymphocytes. Cancer Immunol. Immunother.65(4), 441–452 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Ouyang, Q. et al. Age-associated accumulation of CMV-specific CD8+ T cells expressing the inhibitory killer cell lectin-like receptor G1 (KLRG1). Exp. Gerontol.38(8), 911–920 (2003). [DOI] [PubMed] [Google Scholar]
- 4.Voehringer, D. et al. Viral infections induce abundant numbers of senescent CD8 T cells. J. Immunol.167(9), 4838–4843 (2001). [DOI] [PubMed] [Google Scholar]
- 5.Henson, S. M., Macaulay, R., Riddell, N. E., Nunn, C. J. & Akbar, A. N. Blockade of PD-1 or p38 MAP kinase signaling enhances senescent human CD8(+) T-cell proliferation by distinct pathways. Eur. J. Immunol.45(5), 1441–1451 (2015). [DOI] [PubMed] [Google Scholar]
- 6.Ibegbu, C. C. et al. Expression of killer cell lectin-like receptor G1 on antigen-specific human CD8+ T lymphocytes during active, latent, and resolved infection and its relation with CD57. J. Immunol.174(10), 6088–6094 (2005). [DOI] [PubMed] [Google Scholar]
- 7.McMahon, C. W. et al. Viral and bacterial infections induce expression of multiple NK cell receptors in responding CD8(+) T cells. J. Immunol.169(3), 1444–1452 (2002). [DOI] [PubMed] [Google Scholar]
- 8.Voehringer, D., Koschella, M. & Pircher, H. Lack of proliferative capacity of human effector and memory T cells expressing killer cell lectinlike receptor G1 (KLRG1). Blood100(10), 3698–3702 (2002). [DOI] [PubMed] [Google Scholar]
- 9.Truong, K. L. et al. Killer-like receptors and GPR56 progressive expression defines cytokine production of human CD4(+) memory T cells. Nat. Commun.10(1), 2263 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Peng, Y. M. et al. Specific expression of GPR56 by human cytotoxic lymphocytes. J. Leukoc. Biol.90(4), 735–740 (2011). [DOI] [PubMed] [Google Scholar]
- 11.Tian, Y. et al. Unique phenotypes and clonal expansions of human CD4 effector memory T cells re-expressing CD45RA. Nat. Commun.8(1), 1473 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Dominguez, C. X. et al. The transcription factors ZEB2 and T-bet cooperate to program cytotoxic T cell terminal differentiation in response to LCMV viral infection. J. Exp. Med.212(12), 2041–2056 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Omilusik, K. D. et al. Transcriptional repressor ZEB2 promotes terminal differentiation of CD8+ effector and memory T cell populations during infection. J. Exp. Med.212(12), 2027–2039 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.van Helden, M. J. et al. Terminal NK cell maturation is controlled by concerted actions of T-bet and Zeb2 and is essential for melanoma rejection. J. Exp. Med.212(12), 2015–2025 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Henson, S. M. et al. p38 signaling inhibits mTORC1-independent autophagy in senescent human CD8(+) T cells. J. Clin. Invest.124(9), 4004–4016 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Jeng, M. Y. et al. Metabolic reprogramming of human CD8(+) memory T cells through loss of SIRT1. J. Exp. Med.215(1), 51–62 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Schurich, A. & Henson, S. M. The many unknowns concerning the bioenergetics of exhaustion and senescence during chronic viral infection. Front. Immunol.5, 468 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Cruz-Munoz, M. E., Dong, Z., Shi, X., Zhang, S. & Veillette, A. Influence of CRACC, a SLAM family receptor coupled to the adaptor EAT-2, on natural killer cell function. Nat. Immunol.10(3), 297–305 (2009). [DOI] [PubMed] [Google Scholar]
- 19.Tassi, I. & Colonna, M. The cytotoxicity receptor CRACC (CS-1) recruits EAT-2 and activates the PI3K and phospholipase Cgamma signaling pathways in human NK cells. J. Immunol.175(12), 7996–8002 (2005). [DOI] [PubMed] [Google Scholar]
- 20.Hagberg, N. et al. Systemic lupus erythematosus immune complexes increase the expression of SLAM family members CD319 (CRACC) and CD229 (LY-9) on plasmacytoid dendritic cells and CD319 on CD56(dim) NK cells. J. Immunol.191(6), 2989–2998 (2013). [DOI] [PubMed] [Google Scholar]
- 21.Frigyesi, I. et al. Robust isolation of malignant plasma cells in multiple myeloma. Blood123(9), 1336–1340 (2014). [DOI] [PubMed] [Google Scholar]
- 22.Lee, J. K., Mathew, S. O., Vaidya, S. V., Kumaresan, P. R. & Mathew, P. A. CS1 (CRACC, CD319) induces proliferation and autocrine cytokine expression on human B lymphocytes. J. Immunol.179(7), 4672–4678 (2007). [DOI] [PubMed] [Google Scholar]
- 23.Comte, D. et al. Signaling lymphocytic activation molecule family member 7 engagement restores defective effector CD8+ T cell function in systemic lupus erythematosus. Arthritis Rheumatol.69(5), 1035–1044 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Wertheimer, A. M. et al. Aging and cytomegalovirus infection differentially and jointly affect distinct circulating T cell subsets in humans. J. Immunol.192(5), 2143–2155 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Galletti, G. et al. Two subsets of stem-like CD8(+) memory T cell progenitors with distinct fate commitments in humans. Nat. Immunol.21(12), 1552–1562 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Gattinoni, L. et al. A human memory T cell subset with stem cell-like properties. Nat. Med.17(10), 1290–1297 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Huster, K. M. et al. Selective expression of IL-7 receptor on memory T cells identifies early CD40L-dependent generation of distinct CD8+ memory T cell subsets. Proc. Natl. Acad. Sci. U. S. A.101(15), 5610–5615 (2004). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Zwijnenburg, A. J. et al. Graded expression of the chemokine receptor CX3CR1 marks differentiation states of human and murine T cells and enables cross-species interpretation. Immunity56(8), 1955–7410 (2023). [DOI] [PubMed] [Google Scholar]
- 29.Quigley, M. et al. Transcriptional analysis of HIV-specific CD8+ T cells shows that PD-1 inhibits T cell function by upregulating BATF. Nat. Med.16(10), 1147–1151 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Alfei, F. et al. TOX reinforces the phenotype and longevity of exhausted T cells in chronic viral infection. Nature571(7764), 265–269 (2019). [DOI] [PubMed] [Google Scholar]
- 31.Khan, O. et al. TOX transcriptionally and epigenetically programs CD8(+) T cell exhaustion. Nature571(7764), 211–218 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Scott, A. C. et al. TOX is a critical regulator of tumour-specific T cell differentiation. Nature571(7764), 270–274 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Seo, H. et al. TOX and TOX2 transcription factors cooperate with NR4A transcription factors to impose CD8(+) T cell exhaustion. Proc. Natl. Acad. Sci. U. S. A.116(25), 12410–12415 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Yao, C. et al. Single-cell RNA-seq reveals TOX as a key regulator of CD8(+) T cell persistence in chronic infection. Nat. Immunol.20(7), 890–901 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Kared, H. et al. Immunological history governs human stem cell memory CD4 heterogeneity via the Wnt signaling pathway. Nat. Commun.11(1), 821 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.O’Connell, P. et al. SLAMF7 signaling reprograms T cells toward exhaustion in the tumor microenvironment. J. Immunol.206(1), 193–205 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Gangemi, S. et al. Age-related modifications in circulating IL-15 levels in humans. Mediators Inflamm.2005(4), 245–247 (2005). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Alpert, A. et al. A clinically meaningful metric of immune age derived from high-dimensional longitudinal monitoring. Nat. Med.25(3), 487–495 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Cruz-Guilloty, F. et al. Runx3 and T-box proteins cooperate to establish the transcriptional program of effector CTLs. J. Exp. Med.206(1), 51–59 (2009). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Intlekofer, A. M. et al. Effector and memory CD8+ T cell fate coupled by T-bet and eomesodermin. Nat. Immunol.6(12), 1236–1244 (2005). [DOI] [PubMed] [Google Scholar]
- 41.Oja, A. E. et al. The transcription factor hobit identifies human cytotoxic CD4(+) T cells. Front. Immunol.8, 325 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Buggert, M. et al. T-bet and Eomes are differentially linked to the exhausted phenotype of CD8+ T cells in HIV infection. PLoS Pathog.10(7), e1004251 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Dolfi, D. V. et al. Increased T-bet is associated with senescence of influenza virus-specific CD8 T cells in aged humans. J. Leukoc. Biol.93(6), 825–836 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Page, N. et al. Expression of the DNA-binding factor TOX promotes the encephalitogenic potential of microbe-induced autoreactive CD8(+) T cells. Immunity48(5), 937–50 e8 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Utzschneider, D. T. et al. T cell factor 1-expressing memory-like CD8(+) T cells sustain the immune response to chronic viral infections. Immunity45(2), 415–427 (2016). [DOI] [PubMed] [Google Scholar]
- 46.Kurtulus, S. et al. Checkpoint blockade immunotherapy induces dynamic changes in PD-1(-)CD8(+) tumor-infiltrating T cells. Immunity50(1), 181–94 e6 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Sade-Feldman, M. et al. Defining T cell states associated with response to checkpoint immunotherapy in melanoma. Cell175(4), 998-1013 e20 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Siddiqui, I. et al. Intratumoral Tcf1(+)PD-1(+)CD8(+) T cells with stem-like properties promote tumor control in response to vaccination and checkpoint blockade immunotherapy. Immunity50(1), 195-211 e10 (2019). [DOI] [PubMed] [Google Scholar]
- 49.Sekine, T. et al. TOX is expressed by exhausted and polyfunctional human effector memory CD8(+) T cells. Sci. Immunol.10.1126/sciimmunol.aba7918 (2020). [DOI] [PubMed] [Google Scholar]
- 50.Jeannet, G. et al. Essential role of the Wnt pathway effector Tcf-1 for the establishment of functional CD8 T cell memory. Proc. Natl. Acad. Sci. U. S. A.107(21), 9777–9782 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Zhou, X. et al. Differentiation and persistence of memory CD8(+) T cells depend on T cell factor 1. Immunity33(2), 229–240 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Lee, J. K., Boles, K. S. & Mathew, P. A. Molecular and functional characterization of a CS1 (CRACC) splice variant expressed in human NK cells that does not contain immunoreceptor tyrosine-based switch motifs. Eur. J. Immunol.34(10), 2791–2799 (2004). [DOI] [PubMed] [Google Scholar]
- 53.Kim, J. R., Mathew, S. O., Patel, R. K., Pertusi, R. M. & Mathew, P. A. Altered expression of signalling lymphocyte activation molecule (SLAM) family receptors CS1 (CD319) and 2B4 (CD244) in patients with systemic lupus erythematosus. Clin. Exp. Immunol.160(3), 348–358 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Kim, J. R., Horton, N. C., Mathew, S. O. & Mathew, P. A. CS1 (SLAMF7) inhibits production of proinflammatory cytokines by activated monocytes. Inflamm. Res.62(8), 765–772 (2013). [DOI] [PubMed] [Google Scholar]
- 55.Khairnar, V. et al. CEACAM1 promotes CD8(+) T cell responses and improves control of a chronic viral infection. Nat. Commun.9(1), 2561 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Wu, Y. et al. SLAMF7 regulates the inflammatory response in macrophages during polymicrobial sepsis. J. Clin. Invest.10.1172/JCI150224 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Simmons, D. P. et al. SLAMF7 engagement superactivates macrophages in acute and chronic inflammation. Sci. Immunol.7(68), eabf2846 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Chu, E., Wu, J., Kang, S. S. & Kang, Y. SLAMF7 as a promising immunotherapeutic target in multiple myeloma treatments. Curr. Oncol.30(9), 7891–7903 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Awwad, M. H. S. et al. Selective elimination of immunosuppressive T cells in patients with multiple myeloma. Leukemia35(9), 2602–2615 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Bae, J. et al. Identification and characterization of HLA-A24-specific XBP1, CD138 (Syndecan-1) and CS1 (SLAMF7) peptides inducing antigens-specific memory cytotoxic T lymphocytes targeting multiple myeloma. Leukemia32(3), 752–764 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Cachot, A. et al. Tumor-specific cytolytic CD4 T cells mediate immunity against human cancer. Sci. Adv.10.1126/sciadv.abe3348 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Berard, M., Brandt, K., Bulfone-Paus, S. & Tough, D. F. IL-15 promotes the survival of naive and memory phenotype CD8+ T cells. J. Immunol.170(10), 5018–5026 (2003). [DOI] [PubMed] [Google Scholar]
- 63.Richer, M. J. et al. Inflammatory IL-15 is required for optimal memory T cell responses. J. Clin. Invest.125(9), 3477–3490 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Hurton, L. V. et al. Tethered IL-15 augments antitumor activity and promotes a stem-cell memory subset in tumor-specific T cells. Proc. Natl. Acad. Sci. U. S. A.113(48), E7788–E7797 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Strutt, T. M. et al. IL-15 supports the generation of protective lung-resident memory CD4 T cells. Mucosal Immunol.11(3), 668–680 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Pangrazzi, L. et al. Increased IL-15 production and accumulation of highly differentiated CD8(+) effector/memory T cells in the bone marrow of persons with cytomegalovirus. Front. Immunol.8, 715 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Lu, Y. et al. Inflammatory and immune markers associated with physical frailty syndrome: Findings from Singapore longitudinal aging studies. Oncotarget7(20), 28783–28795 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Kared, H. et al. Immune responses in Omicron SARS-CoV-2 breakthrough infection in vaccinated adults. Nat. Commun.13(1), 4165 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The authors declare that the data supporting the findings of this study are available within the paper [and its supplementary information files]. RNA sequencing data are accessible from NCBI’s Gene Expression Omnibus repository using the dataset ID GSE 282172. The data that support the other findings of this study are available from the corresponding author upon reasonable request.