Abstract
Rising cases of type 2 diabetes (T2D) in India, especially in metropolitan cities is an increasing concern. The individuals that were most affected are young professionals working in the corporate sector. However, the corporate sector has remained the least explored for T2D risk predisposition. Considering corporate employees’ lifestyles and the role of gene-environment interaction in T2D susceptibility, the study aims to find genetic variants associated with T2D predisposition. In this first kind of study, 680 young professionals (284 T2D cases, and 396 controls) were diagnosed and screened for 2658 variants on an array designed explicitly for the CoGsI study. The variant filtering was done at Bonferroni p-value of 0.000028. The genetic data was analysed using PLINK v1.09, SPSS, R programming, VEP tool, and FUMA GWAS tool. Interestingly, 42 variants were associated with the T2D risk. Out of 42, three missense variants (rs1402467, rs6050, and rs713598) in Sulfotransferase family 1 C member 4 (SULT1C4), Fibrinogen Alpha Chain (FGA), and Taste 2 Receptor Member 38 (TAS2R38) and two untranslated region (UTR) variants (rs1063320 and rs6296) in Major Histocompatibility Complex, Class I, G (HLA-G) and 5-Hydroxytryptamine Receptor 1B (HTR1B) were associated with the T2D risk. CoGsI identified potential genomic markers increasing susceptibility to the early onset of T2D. Present findings provide insights into mechanisms underlying T2D manifestation in corporate professionals due to genetics interacting with occupational stress and urban lifestyles.
Supplementary Information
The online version contains supplementary material available at 10.1038/s41598-024-84160-2.
Keywords: Type 2 diabetes, Genetics, Young professionals, SNPs, Genotyping
Subject terms: Genetics, Molecular biology, Diseases
Background
Type 2 diabetes (T2D) is a pervasive1 and consequential health concern characterised by chronic glycemia due to insulin resistance, impaired insulin production, and sometimes both2,3. The World Health Organization reported 422 million global diabetes cases, projected to reach 700 million by 20454.
T2D poses a significant burden, triggering complications like cardiovascular disorders, diabetic retinopathy, nephropathy, neuropathy, and many more, condensing quality of life5,6. The International Diabetes Federation estimated global healthcare expenditure related to diabetes to be over 825 billion USD by 2030 and 845 billion USD by 2045, constituting approximately 10% of total global healthcare spending7. To decrease such global burdens, it is crucial to decipher the factors affecting the severity of T2D8. It is a fact that genetics9, epigenetics10, and environmental factors play crucial roles in T2D progression11,12; mainly depending on the complex interaction of genetic variants and environmental factors13. Understanding the genetic and environmental contributors to T2D is vital, particularly in India, where it has a substantial impact14. Genetic risk variants across the human genome affect glucose metabolism, insulin signaling, and pancreatic function15. However, genetics operates within the context of environmental influences16, necessitating a comprehensive approach to address this global health challenge.
India, with a population exceeding 1.3 billion, faces a rapidly escalating T2D crisis, ranking second globally in the number of affected individuals17. Approximately 8.8% of India’s adult population, around 77 million individuals, have diabetes in 2023, with an alarming 44% being undiagnosed18,19. Diabetes-related healthcare expenditure accounted for about 10% of India’s total healthcare spending in 202320. Moreover, studies have indicated the association of chronic stress with the elevated risk of T2D21. A large sample sized based T2D study in the Indian population has shown the increased risk of T2D in young North Indians, with a prevalence of 14.4%, having higher Indian diabetes risk score compared to the other parts of India, suggesting that the youth of India is at high risk of T2D specifically the young North Indians due to increased stress, improper sleep, lifestyle habits, sedentary lifestyle, more screen time, smoking, alcohol consumption, and others22. This rise in the cases of youth onset T2D in India and globally has become a major health concern. Multi-level strategies need to be developed among young individuals to prevent the pathogenesis of T2D, including identification of the exposures, social determinants, individual and family level characteristics, disorders associated with the increased risk of T2D, and many more23. Thus, it is pertinent to understand the genetic factors contributing to T2D among Indian professionals to address this epidemic and improve the health outcomes of the Indian population.
The risk of developing T2D depends not only on genetic factors but also on environmental factors. This can be explained as the interplay of the genes and gene-environment interaction. The genetic predisposition to T2D may result from how an individual is exposed to a specific environmental factor. For example, those who adopt an unhealthy diet based on their preferences for taste and food, low daily activity level, exposure to pollutants, poor gut microbiota, psychological and work-related stress, and many more, are highly susceptible to T2D24–26.
Our research team developed the idea of Corporate Genome Screening India (CoGsI), an innovative genetic research approach that leverages genetic data to investigate T2D genetic impact in young working professionals. In CoGsI, we conducted a screening among young Indian professionals to assess their susceptibility towards T2D as corporate sector jobs often come with heavy workloads and stress, deskbound work with minimal physical activity, and unhealthy eating habits. These factors are considered significant risk factors for predisposition to T2D. This strategy utilises the genetic diversity often found among employees or members of such organisations, enhancing the statistical power of genetic analyses and enabling a deeper understanding of genetics interplay with shared environmental factors unique to the group. Advanced genomic techniques, like array-based genotyping are needed to be employed in the corporate genome screening to identify genetic risk variants and shed light on complex interactions between genetics, lifestyle, and the environment. The variants incorporated in the array (2658) were previously being reported to be associated with various phenotypes of T2D and its associated complexities, including lifestyle-related conditions, neurological disorders, and behavioural and stress-related disorders. By examining these variants, we aimed to gain insights into how genetic variants with established roles in T2D and its associated complexities influence T2D risk among young professionals. This approach opens avenues for personalised healthcare strategies based on individuals’ unique genetic profiles.
Materials and methods
Ethical clearance
The study was initiated after seeking ethical approval from the institutional ethics review board (IERB) of NMC Genetics India Pvt. Ltd. notification vide number NMC/IERB/2023/07, dated: 22-04-2023. All experimental protocols were conducted according to the guidelines and regulations set by the IERB.
Subjects
The present study is the first Corporate Genome Screening of young Indian professionals. The screening included 284 T2D (222 males and 62 females) individuals with age ranges between 24 and 50 years and 396 controls (304 males and 92 females) in the age group of 25 to 55 years. The post-hoc power of study calculation was performed using the Genetic Association Study (GAS) Power Calculator (https://csg.sph.umich.edu/abecasis/cats/gas_power_calculator/index.html)27. Further, all the cases were clinically diagnosed in accordance with the recommendations set by the American Diabetes Association28, and several clinical parameters were recorded, including Vitamin D3, Cholesterol levels, Liver profile, Protein, HbA1c, and many others that were evaluated in all the T2D affected individuals (Table 1).
Table 1.
Distribution of the clinical parameters observed in cases and controls.
| S No. | Blood parameters | Cases (N = 284) | Controls (N = 396) | p-value | ||
|---|---|---|---|---|---|---|
| Mean | SD | Mean | SD | |||
| Age (in years) | 38.98 | 7.62 | 40.07 | 12.19 | 0.2291 | |
| 1. | 25-OH VITAMIN D (TOTAL) (ng/mL) | 22.44 | 15.34 | 29.72 | 18.76 | 3.42E−06* |
| 2. | VITAMIN B-12 (pg/mL) | 309.94 | 203.18 | – | – | – |
| 3. | APOLIPOPROTEIN - A1 (APO-A1) (mg/dL) | 118.79 | 15.25 | 121.72 | 21.94 | 0.211 |
| 4. | APOLIPOPROTEIN - B (APO-B) (mg/dL) | 97.49 | 21.05 | 90.78 | 19.38 | 0.018* |
| 5. | APO B / APO A1 RATIO (APO B/A1) (Ratio) | 0.83 | 0.21 | 0.799 | 0.32 | 0.007* |
| 6. | TOTAL CHOLESTEROL (mg/dL) | 185.37 | 37.07 | 181.88 | 42.17 | 0.324 |
| 7. | HDL CHOLESTEROL - DIRECT (mg/dL) | 42.24 | 8.81 | 45.38 | 12.16 | 0.0009* |
| 8. | HDL / LDL RATIO | 0.38 | 0.14 | 0.42 | 0.28 | 0.042* |
| 9. | LDL CHOLESTEROL - DIRECT (mg/dL) | 120.48 | 30.44 | 121.66 | 36.12 | 0.692 |
| 10. | TRIGLYCERIDES (mg/dL) | 146.30 | 87.49 | 142.99 | 84.72 | 0.666 |
| 11. | TC/ HDL CHOLESTEROL RATIO | 5.07 | 8.54 | 4.2 | 1.24 | 0.111 |
| 12. | LDL / HDL RATIO | 2.96 | 0.90 | 2.84 | 1.07 | 0.173 |
| 13. | VLDL CHOLESTEROL (mg/dL) | 29.52 | 17.49 | 29.06 | 17.43 | 0.769 |
| 14. | NON-HDL CHOLESTEROL (mg/dL) | 142.28 | 36.80 | 136.63 | 40.91 | 0.103 |
| 15. | ALKALINE PHOSPHATASE (U/L) | 82.67 | 25.94 | 83.67 | 28.56 | 0.681 |
| 16. | BILIRUBIN - TOTAL (mg/dL) | 0.73 | 0.32 | 0.68 | 0.34 | 0.090 |
| 17. | BILIRUBIN -DIRECT (mg/dL) | 0.15 | 0.09 | 0.17 | 0.1 | 0.018* |
| 18. | BILIRUBIN (INDIRECT) (mg/dL) | 0.57 | 0.26 | 0.51 | 0.27 | 0.011* |
| 19. | GAMMA GLUTAMYL TRANSFERASE (GGT) PHOTOMETRY (U/L) | 35.99 | 56.91 | 36.04 | 42.97 | 0.991 |
| 20. | SGOT / SGPT RATIO CALCULATED | 0.94 | 0.37 | 0.95 | 0.4 | 0.771 |
| 21. | ASPARTATE AMINOTRANSFERASE (SGOT ) PHOTOMETRY (U/L) | 28.86 | 12.23 | 29.33 | 13.72 | 0.685 |
| 22. | PROTEIN - TOTAL PHOTOMETRY (g/dL) | 7.23 | 0.49 | 7.08 | 0.41 | 0.0002* |
| 23. | ALBUMIN - SERUM PHOTOMETRY (gm/dL) | 4.35 | 0.36 | 4.3 | 0.31 | 0.096 |
| 24. | SERUM ALB/GLOBULIN RATIO CALCULATED | 1.53 | 0.23 | 1.58 | 0.29 | 0.032* |
| 25. | SERUM GLOBULIN (gm/dL) | 2.89 | 0.40 | 2.76 | 0.37 | 0.002* |
| 26. | HbA1c - (HPLC) (%) | 7.93 | 3.56 | 5.98 | 1.34 | 1.27E−14* |
SD: Standard deviation, N = Number of samples, * variables that have significant p-values < 0.05.
Sample collection
The blood samples for the biochemical test were initially drawn to diagnose the subjects as cases or controls in the present study. Additionally, the 2mL blood sample was collected in EDTA collection tubes from cases and controls after seeking informed consent from all the subjects for the isolation of DNA. The blood samples were stored at -20° C until further processing.
DNA isolation
The blood samples were processed further to isolate the genomic DNA. The DNA was isolated using the Qiagen DNA mini kit as per the manufacturer’s protocol. The qualitative and quantitative analysis of the DNA samples was performed using the agarose gel electrophoresis and Nanodrop (ThermoFisher) (Supplementary Fig. 1a,b,c).
Genotyping
The DNA samples of cases and controls were genotyped using an array designed exclusively for the CoGsI screening (Supplementary Figs. 2, 3, 4, 5). The samples were screened for a total of 2658 variants. The detailed information about the sample genotyping was adopted from our previous studies29,30.
Statistical analysis
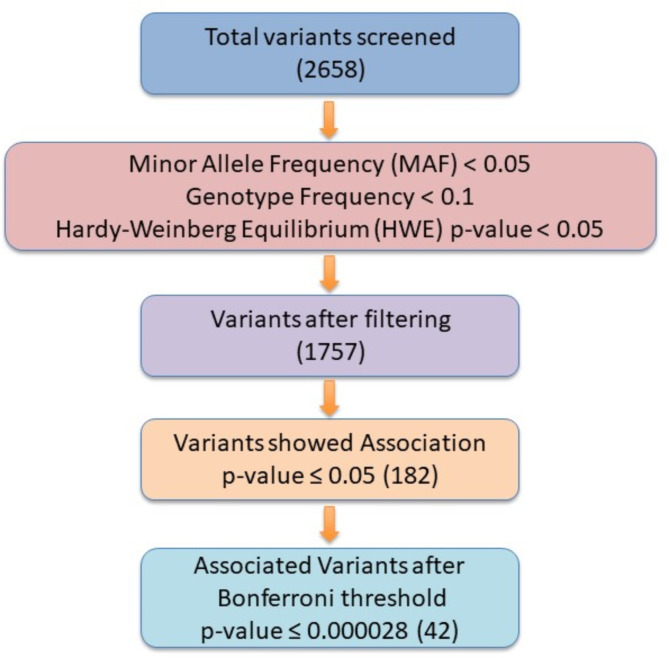
The statistical analysis of the genotyping data was conducted using the PLINK v1.0931. The association of the clinical parameters among cases and controls was performed using the SPSS v27 (IBM) and R programming. To ensure the robustness of our analysis, we applied stringent data filtration criteria, including a minor allele frequency threshold of less than 0.05, to exclude rare variants that may not provide sufficient statistical power for association tests based on the sample size of the present study, genotype frequency of less than 0.1 was considered to filter out variants with low homozygosity, and adherence to Hardy-Weinberg Equilibrium (HWE) with a significance level of less than 0.05 using PLINK v1.0931. Following these quality control measures, we observed that 1757 Single Nucleotide Polymorphisms (SNPs) remained for further analysis. We evaluated the association of these SNPs with T2D using comprehensive statistical assessments. Our investigation revealed a noteworthy outcome, with 182 genetic variants demonstrating significant associations with T2D (p-value < 0.05). To ensure the reliability of our findings, we rigorously corrected potential artifacts using the Bonferroni correction method (p-value ≤ 0.000028). The p-values of the association result have been adjusted with the confounding factors (age, gender, and BMI). To evaluate the correlation between the associated genetic variants with the physical activity level, occupational stress, and unhealthy diet, the Pearson correlation coefficient method was performed using Python script. Consequently, at the culmination of our analysis, we identified and confirmed 42 genetic variants after Bonferroni correction that maintain a robust and statistically significant association with the development of T2D. The variant filter criterion is represented in Fig. 1.
Fig. 1.
Flowchart depicting the variant filtering criteria used in the present study. The variant filtering criteria in the flowchart comprised of the minor allele frequency filter of < 0.05, genotype frequency of < 0.1, and Hardy Weinberg equilibrium of < 0.05. The association of the variants with T2D was observed at the Bonferroni threshold of ≤ 0.000028.
Functional annotation of associated variants/genes
The annotation of the variants for their functional role was evaluated using the Variant Effect Predictor tool (Ensembl)32. The associated genes were further annotated using the GENE2FUNC tool of the FUMA GWAS tool33. The gene expression was evaluated using the GTEx data for 54 tissue sets. In addition, the Gene ontology analysis for the role of genes in the biological processes was determined using the MSigDb data from the FUMA GWAS tool. The network analysis of the associated genes was performed using the GeneMANIA tool (https://genemania.org/)34.
Results
Clinical parameters
The cases and controls belong to the age group of 38 (± 7.62) and 40 (± 12.19) years, respectively. Eleven of 26 blood parameters were significantly associated with T2D in cases with p-values of < 0.05 (Table 1). The levels of vitamin D3 in cases and controls are 22.44 (± 15.34) and 29.72 (± 18.76) ng/ml, respectively, which falls in the insufficiency and normal range of vitamin D3 levels recommended for better health. All the individuals have Apolipoprotein B in the normal range < 110 mg/dL. The individuals have a moderate risk related to the Apo B/ ApoA1 ratio because they fall in the moderate range of 0.7–0.9. The HDL cholesterol levels of cases and controls were 42.24 (± 8.81) and 45.38 (± 12.16) mg/dL, which falls in the normal range. The levels of bilirubin in cases and controls were 0.15 (± 0.09) and 0.17 (± 0.1) for direct bilirubin and 0.57 (± 0.26) and 0.51 (± 0.27) for indirect bilirubin respectively, as compared to the normal range of 0.2–1.2 mg/dL. The total protein content for cases were 7.23 (± 0.49) and 7.08 (± 0.41) g/dL, respectively. Both groups fell within the 6–8.5 g/dL range for total protein. The normal range of serum albumin/ globulin ratio is 1–2, and the observed values for cases and controls were 1.53 ± 0.23 and 1.58 ± 0.29. Serum globulin levels in the cases and controls were 2.89 ± 0.40 and 2.76 ± 0.37 compared to the normal range of 2–3.5 g/dL. Regarding glycemic control, the average HbA1c level in cases was 7.93 (± 3.56), which falls in the diabetic range. On the other hand, for the controls, the HbA1c level was 5.98 ± 1.34, which falls in the normal range. This information summarises the key findings related to age, vitamin D3 levels, Apo-B, cardiovascular risk, HDL cholesterol, bilirubin, total protein, albumin/ globulin ratio, and diabetic levels in cases and controls.
The estimated power of the study was 96.3% (Supplementary Fig. 6), considering a p-value threshold of 0.000028, with 8% of the T2D prevalence35, 0.2 as minor allele frequency with an average odds ratio of 2. A power greater than 80%36; indicates very little chance of Type II errors, ensuring that the study is sensitive enough to detect even small effects and providing strong evidence of association with the selected sample size.
Genotyping
From the total of 2658 variants following the quality control measures, the present study has revealed 42 genetic variants demonstrating significant associations with T2D. The 42 variants were found to provide a strong risk predisposition to T2D. All these variants were associated with T2D risk with higher odds at a very high significance level (Table 2). Moreover, the variants were further evaluated for their epistatic effect (supplementary Table 1) and their association under additive (supplementary Table 2), dominant (supplementary Table 3), and recessive model (supplementary Table 4). The epistatic analysis revealed that the variant of the Calcium Voltage-Gated Channel Subunit Alpha1 C (CACNA1C) gene showed interaction with the Interleukin 1 Beta (IL1B), Fibrinogen Alpha Chain (FGA), T Cell Immunoglobulin And Mucin Domain Containing 4 (TIMD4), Major Histocompatibility Complex, Class I, G (HLA-G), 5-Hydroxytryptamine Receptor 1B (HTR1B), Poliovirus receptor-related 2 (PVRL2), and Transmembrane Serine Protease 6 (TMPRSS6). The interaction analysis further depicted the elevated risk at higher significance levels with T2D in the population. Most of the associated variants were intronic, while three of them, viz. rs1402467, rs6050, and rs713598, were missense variants that would probably alter the structure of the protein. In addition, variants rs1063320 and rs6296 were 3´ UTR and 5´ UTR variants, respectively, that might play an essential role in regulating gene expression. In addition, the interactive analysis of the 42 associated genetic variants with occupational stress, unhealthy diet, and physical activity was evaluated to determine the complex mechanism behind the predisposition of T2D in young Indian professionals. It was found that some of the genetic variants were found to be significantly interacting with these environmental factors in causing the T2D susceptibility in the studied population (Fig. 2, Supplementary Table 5).
Table 2.
Genetic variants found to be associated with T2D in studied cohort.
| SNP | Gene | Func cons | Codon | Effect allele | EAF in cases | EAF in controls | Alternate allele | OR (95%CI) | P adjusted |
|---|---|---|---|---|---|---|---|---|---|
| rs12027135 | TMEM57 | Intron variant | T | 0.6549 | 0.327 | A | 3.906 (3.11–4.906) | 5.85E-33 | |
| rs10918594 | RP11-227F8.2 | NA | C | 0.6092 | 0.4003 | G | 2.335 (1.873–2.911) | 2.94E-14 | |
| rs61759167 | PRDM16 | Intron variant | T | 0.3644 | 0.2538 | C | 1.686 (1.334–2.131) | 1.13E-05 | |
| rs1143623 | IL1B | 2KB upstream variant | C | 0.6761 | 0.3258 | G | 4.32 (3.432–5.437) | 2.68E-37 | |
| rs1402467 | SULT1C4 | Missense variant | D (Asp) > E (Glu) | G | 0.5458 | 0.3712 | C | 2.035 (1.635–2.534) | 1.66E-10 |
| rs6599234 | SCN5A | NA | A | 0.6831 | 0.3144 | T | 4.701 (3.728–5.927) | 3.60E-41 | |
| rs12493607 | TGFBR2 | Intron variant | C | 0.6496 | 0.3586 | G | 3.317 (2.647–4.155) | 3.12E-26 | |
| rs4533622 | CTNNB1 | Intron variant | A | 0.6127 | 0.4141 | C | 2.238 (1.795–2.789) | 5.13E-13 | |
| rs16998073 | PRDM8 | NA | T | 0.7042 | 0.3101 | A | 5.296 (4.187–6.699) | 1.14E-46 | |
| rs6050 | FGA | Missense variant | T [66] > A [67] | G | 0.5616 | 0.3434 | A | 2.449 (1.963–3.056) | 1.19E-15 |
| rs10520514 | TENM3 | Intron variant | T | 0.6004 | 0.404 | A | 2.216 (1.778–2.761) | 9.10E-13 | |
| rs1501908 | TIMD4 | None | C | 0.6849 | 0.3093 | G | 4.852 (3.846–6.122) | 1.16E-42 | |
| rs12654264 | HMGCR | Intron variant | A | 0.5827 | 0.423 | T | 1.905 (1.531–2.37) | 6.15E-09 | |
| rs1610696 | HLA-G | 500B Downstream variant | C | 0.6831 | 0.3081 | G | 4.841 (3.837–6.108) | 1.43E-42 | |
| rs1063320 | HLA-G | 3 Prime UTR variant | G | 0.6585 | 0.3081 | C | 4.33 (3.441–5.449) | 1.69E-37 | |
| rs6296 | HTR1B | 5 Prime UTR variant | C | 0.6426 | 0.3573 | G | 3.234 (2.583–4.049) | 2.80E-25 | |
| rs1801132 | ESR1 | Synonymous variant | G | 0.6144 | 0.3939 | C | 2.452 (1.965–3.058) | 1.03E-15 | |
| rs11755527 | BACH2 | Intron variant | G | 0.5792 | 0.4304 | C | 1.822 (1.465–2.266) | 6.24E-08 | |
| rs713598 | MGAM | Missense variant | A [Ala] > P [66] | C | 0.6725 | 0.3194 | G | 4.375 (3.476–5.508) | 6.49E-38 |
| rs6943555 | AUTS2 | Intron variant | A | 0.6743 | 0.3258 | T | 4.285 (3.405–5.393) | 6.00E-37 | |
| rs180242 | GNG11 | 2KB upstream variant | A | 0.6303 | 0.3813 | T | 2.766 (2.214–3.456) | 1.30E-19 | |
| rs6968865 | LOC101927609 | Intron variant | T | 0.5986 | 0.3939 | A | 2.294 (1.841–2.859) | 9.30E-14 | |
| rs4646244 | NAT2 | Intron variant | A | 0.6303 | 0.3396 | T | 3.314 (2.647–4.151) | 2.86E-26 | |
| rs1537415 | GLT6D1 | Intron variant | G | 0.7218 | 0.3157 | C | 5.626 (4.439–7.13) | 1.95E-49 | |
| rs7020673 | GLIS3 | Intron variant | C | 0.5475 | 0.428 | G | 1.617 (1.302–2.009) | 1.35E-05 | |
| rs11014166 | CACNB2 | Intron variant | T | 0.706 | 0.2879 | A | 5.94 (4.686–7.53) | 1.61E-52 | |
| rs12772424 | TCF7L2 | Intron variant | T | 0.6673 | 0.3093 | A | 4.477 (3.555–5.638) | 5.35E-39 | |
| rs10995271 | ZNF365 | Intron variant | C | 0.6127 | 0.3801 | G | 2.58 (2.068–3.22) | 2.44E-17 | |
| rs187238 | IL18 | 2KB upstream variant | C | 0.7377 | 0.2348 | G | 9.162 (7.143–11.75) | 1.26E-75 | |
| rs1800955 | DRD4 | 2KB upstream variant | C | 0.5792 | 0.4432 | T | 1.73 (1.391–2.15) | 7.48E-07 | |
| rs10848653 | CACNA1C | Intron variant | G | 0.5 | 0.274 | A | 2.65 (2.112–3.324) | 1.57E-17 | |
| rs7153648 | SIX1 | None | C | 0.7306 | 0.2753 | G | 7.142 (5.605–9.099) | 6.52E-62 | |
| rs684513 | CHRNA5 | Intron variant | G | 0.6074 | 0.3712 | C | 2.621 (2.1-3.271) | 7.52E-18 | |
| rs1558902 | FTO | Intron variant | A | 0.6655 | 0.3582 | T | 3.564 (2.84–4.472) | 5.43E-29 | |
| rs3865188 | RP11-2L4.1 | None | A | 0.6039 | 0.4078 | T | 2.214 (1.777–2.758) | 9.85E-13 | |
| rs11869286 | STARD3 | Intron variant | G | 0.6954 | 0.2992 | C | 5.347 (4.229–6.761) | 2.62E-47 | |
| rs4291 | ACE | 2KB upstream variant | T | 0.6281 | 0.3649 | A | 2.94 (2.35–3.678) | 1.21E-21 | |
| rs519113 | PVRL2 | Intron variant | C | 0.6831 | 0.2929 | G | 5.203 (4.118–6.573) | 4.63E-46 | |
| rs11672691 | AC011526.1 | Intron variant | A | 0.4542 | 0.3384 | G | 1.627 (1.304–2.03) | 1.52E-05 | |
| rs1884613 | RP5-881L22.5 | Intron variant | G | 0.6567 | 0.2929 | C | 4.617 (3.664–5.818) | 1.95E-40 | |
| rs5756506 | TMPRSS6 | Intron variant | C | 0.7332 | 0.2835 | G | 6.944 (5.451–8.847) | 3.80E-60 | |
| rs713875 | RP3-438O4.4 | Intron variant | G | 0.6232 | 0.3952 | C | 2.532 (2.028–3.159) | 1.07E-16 |
SNP: Single Nucleotide Polymorphism, Func Cons: Functional consequence, EAF: Effect Allele Frequency (represents the frequency of the allele tested for association in the studied population), OR: Odds ratio, CI: Confidence Interval, padjusted: Adjusted p- value for age, gender and BMI.
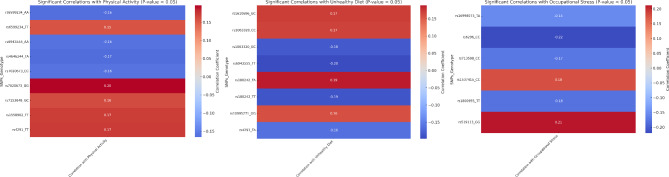
Fig. 2.
Correlations of significant variants and their genotypes with lifestyle/environmental factors including Physical Activity, Unhealthy Diet, and Occupational Stress in Young Indian Professionals.
Functional annotation
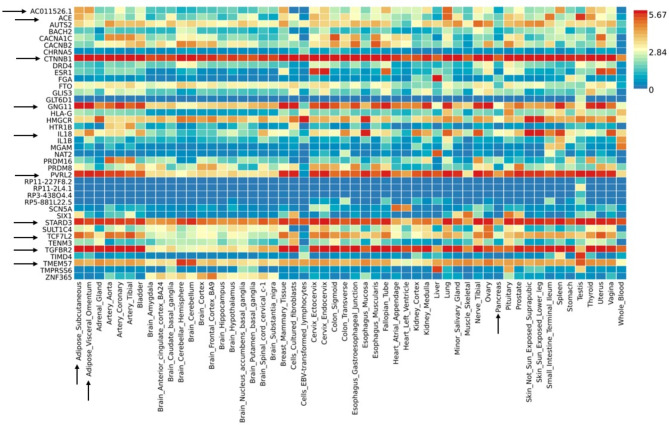
The genes of the associated variants were further annotated for their functional relevance. The heatmap was generated that depicts some interesting findings (Fig. 3). The genes have shown different expression profiles in the 54 tissues set. It was found that the genes including Angiotensin I Converting Enzyme (ACE), Catenin Beta 1 (CTNNB1), G Protein Subunit Gamma 11 (GNG11), Interleukin 18 (IL18), PVRL2, StAR Related Lipid Transfer Domain Containing 3 (STARD3), Transcription Factor 7 Like 2 (TCF7L2), Transforming Growth Factor Beta Receptor 2 (TGFBR2), and Transmembrane Protein 57 (TMEM57) were up-regulated in the subcutaneous adipose tissue, visceral omentum and pancreas. In contrast, the rest of the other genes were comparatively under-expressed in other tissue sets.
Fig. 3.
Heatmap representing the expression of the genes in 54 tissue sets data of GTEx database highlighting the possibly important genes and their expression in the specific tissues. The significantly associated genes of the present study were highlighted with the black arrow and their expression in the specific tissues including Adipose subcutaneous tissue, adipose visceral omentum tissue, and the pancreas tissue has been highlighted with a black arrow.
Furthermore, the gene ontology (GO) analysis for the role of genes in the biological processes was performed using the MSigDB data. It was observed that PR/SET Domain 16 (PRDM16), TGFBR2, CTNNB1, HTR1B, Estrogen Receptor 1 (ESR1), IL18, ACE, IL1B, SIX Homeobox 1 (SIX1), FTO Alpha-Ketoglutarate Dependent Dioxygenase (FTO), 3-Hydroxy-3-Methylglutaryl-CoA Reductase (HMGCR), Sodium Voltage-Gated Channel Alpha Subunit 5 (SCN5A), Calcium Voltage-Gated Channel Auxiliary Subunit Beta 2 (CACNB2), Dopamine Receptor D4 (DRD4), and CACNA1C were significantly associated with the brown fat cell differentiation, response to lipid calcium ion transmembrane transport, negative regulation of glucose transmembrane transport, lipid localisation, MAPK cascade, regulation of calcium ion import, calcium ion homeostasis, calcium ion transmembrane transport, and regulation of voltage-gated calcium channel activity (Supplementary Table 6). These results suggest their potential role in glucose homeostasis, insulin transport, and the risk of developing obesity, which ultimately leads to the development of T2D. Similarly, the GO analysis of the GWAS catalog reported genes using the Gene2Func function of FUMA GWAS tool has shown a significant association of PRDM16, TGFBR2, CTNNB1, PRDM8, ESR1, GLIS Family Zinc Finger 3 (GLIS3), CACNB2, Zinc Finger Protein 365 (ZNF365), TCF7L2, FTO, ACE, HMGCR, TIMD4, HLA-G, N-Acetyltransferase 2 (NAT2), and CACNB2 with various risk factors of T2D including blood pressure, LDL cholesterol levels, Apolipoprotein B levels, triglyceride levels, circulating leptin levels, fasting insulin levels, total cholesterol levels, and blood glucose levels (Supplementary Table 7).
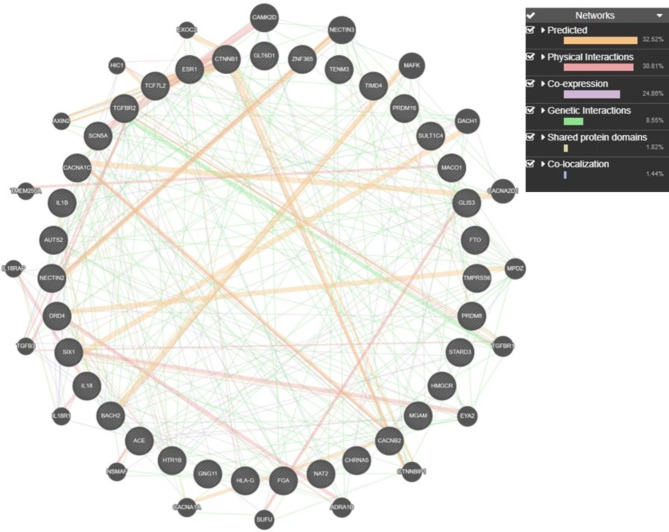
Moreover, the network analysis of the risk genes depicts the predicted interaction among the genes contributing about 32.52%, physical interaction of 30.81%, co-expression of 24.86%, genetic interaction of 8.55%, shared protein domains contributing about 1.82% and co-localisation of 1.44% (Fig. 4).
Fig. 4.
Network analysis depicting the gene-gene interaction of the associated genes depicting the genetic, physical, predicted interactions, shared protein domain, co-localisation, and co-expression. The different interactions have been shown in different colors as represented in the key provided in the figure. Majority of the interactions were found to predicted, physical and genetic in nature.
Discussion
The present study is the first corporate-based genetic screening of T2D in India. In this study, the variants of the genes that were previously reported to be associated with various lifestyle complexities, neurological, stress, and behavioural disorders were screened in the young professionals of the corporate sector. India is considered the diabetic capital of the world37. Many young people, specifically those residing in metropolitan areas, suffer from T2D, which is probably due to environmental factors, including their lifestyle habits and deskbound jobs. No efforts have been made to conduct genetic screening in the corporate sector to evaluate predisposition to T2D until now.
The present study has found a significant association of the eleven clinical parameters with T2D risk in the current population screened, viz. 25-OH Vitamin D, Apolipoprotein - B (APO-B), (APO B/A1) (Ratio), HDL Cholesterol – Direct, HDL / LDL Ratio, Bilirubin -Direct (mg/dL), Bilirubin (Indirect) (mg/dL), Protein - Total photometry, Serum ALB/Globulin ratio, Serum Globulin, and HbA1c - (HPLC). All these factors are critical clinical biomarkers for the indication of pre-diabetic and diabetic conditions.
Moreover, the present study identified the genetic variants in the TMEM57, RP11-227F8.2, PRDM16, IL1B, SULT1C4, SCN5A, TGFBR2, CTNNB1, PRDM8, FGA, TENM3, TIMD4, HMGCR, HLA-G, HTR1B, ESR1, BACH2, TAS2R38, Activator Of Transcription And Developmental Regulator AUTS2 (AUTS2), GNG11, LOC101927609, NAT2, GLT6D1, GLIS3, CACNB2, TCF7L2, ZNF365, IL18, DRD4, CACNA1C, SIX1, CHRNA5, FTO, RP11-2L4.1, STARD3, ACE, PVRL2, AC011526.1, RP5-881L22.5, TMPRSS6, and RP3-438O4.4 associated with risk of T2D. Interestingly, these candidate genes/loci were associated with various complex disorders/traits including diabetes, cardiovascular disease, cancer, dyslipidemia, taste, and many more38–43 where environmental factors contribute significantly to their pathogenesis.
Captivatingly, three missense variants (rs1402467, rs6050, and rs713598) in SULT1C4, FGA, and TAS2R38 were associated with the risk of T2D in the studied population. SULT1C4 is a sulfotransferase, and studies have found increased protein expression in individuals with fatty liver and obesity44, indicating its substantial role in T2D pathogenesis. Studies have also shown the association of the FGA gene with different types of diabetes, including T2D, type 1 diabetes, diabetic neuropathy, and gestational diabetes45–47. Similarly, the TAS2R38, which encodes a bitter taste receptor and is localised in the gustatory system, has been found to negatively regulate glucose homeostasis, which might lead to the risk of T2D41,48,49.
Moreover, two UTR variants (rs1063320 and rs6296) in HLA-G and HTR1B were associated with the risk of T2D in the studied population, and studies have also identified the association of HLA-G in the risk of gestational diabetes mellitus and type 1 diabetes50,51. The HTR1B, which is a negative regulator of serotonin, might act as a crucial factor for T2D pathogenesis52. It has been observed that serotonin levels have decreased in the brain in conditions like type 1 diabetes and T2D. The negative regulation of serotonin is linked to T2D through its impact on insulin secretion and glucose metabolism. The serotonin is vital for the control of metabolic processes including pancreatic beta cell insulin secretion. Studies indicate several ways in which serotonin might enhance glucose-stimulated insulin secretion (GSIS) via serotonylation, where serotonin links itself to proteins involved in insulin exocytosis. Serotonin pathways dysregulation, whether caused by environmental or hereditary elements, might compromise these metabolic processes, hence increasing the T2D risk53,54. Studies have also reported that increasing serotonin levels has an association with a positive effect on T2D condition52,55,56. The polymorphism in HTR1B was also associated with increased BMI in the African American ancestry35, a risk factor for T2D pathogenesis57. Similarly, other genes whose variants were found to be associated with the present study were also likely to have a role in glucose homeostasis, dyslipidemia, obesity, diabetic nephropathy, diabetic neuropathy, type 1 diabetes, and many more that are crucial risk factors for T2D susceptibility39,58–65.
The correlation analysis showed interesting findings where the GG genotype of rs7020673 exhibits a positive correlation (0.20) with less physical activity (Fig. 2), these findings corroborate with the association analyses where the recessive model was showing higher odds than other model i.e., 2.2 (1.5–3.1 at 95% CI, p-value 9.59E-06) (supplementary Table 4), implying that those carrying this genotype might be at greater risk to have T2D because of less exercise. Whereas, the TA genotype of rs4646244 showed a negative correlation (-0.17) (Fig. 2) with physical activity, potentially offering some protection against T2D by promoting higher activity levels. Furthermore, the rs1063320 showed a negative correlation (-0.18) (Fig. 2), indicating that GC genotype carriers with healthy eating habits are less likely to have T2D risk. On the other hand, the rs180242 TT genotype shows a positive correlation (0.19) with an unhealthy diet, implying a greater inclination for bad dietary practices that might increase the risk of T2D. Moreover, rs6296 CC genotype negative correlation (-0.22) with occupational stress (Fig. 2). On the other hand, the rs519113 GG genotype was positively correlated (0.21) with occupational stress (Fig. 2), thereby, possibly increasing T2D risk.
The GO analysis in the present study also indicates the role of associated genes with risk factors of T2D based on their role in regulating dyslipidemia, insulin transport, MAPK pathway, glucose homeostasis, and many more. The network analysis of the gene-gene interaction depicts the interplay of the genes. Most of the genes were found to interact with each other in terms of physical, co-expression, and genetic interaction. The key genes such as TMEM57, PRDM16, IL1B, SCN5A, TGFBR2, CTNNB1, HMGCR, HTR1B, ESR1, and FTO are observed to be interacting through various types of interactions. Predicted interactions show potential relationships that may not yet be experimentally confirmed but suggest implications for determining the similar functions of the interacting genes. Physical interactions indicate direct protein-protein interactions that play a crucial role in understanding the biochemical pathways. Similarly, co-expression of the genes expressed together suggests the involvement of the interacting genes in related biological processes. The genes that were found to show the genetic interactions highlight that their effect can be modified by another interacting gene, revealing their epistatic relationships. The shared protein domains hint at the functional similarities, while co-localisation highlights the proteins that share the same cellular compartment. These interactions provide insights into the genetic architecture of T2D, highlighting potential therapeutic targets and thereby guiding future research into the disease’s pathogenesis. Thus, the individual carrying variations in more than one of these genes would possibly be at a higher risk of developing T2D.
Moreover, the study provides valuable genetic and clinical information that is crucial for the development of personalised healthcare strategies and interventions that are specifically tailored to individuals’ distinct genetic profiles. This will ultimately aid in the prevention and management of T2D in the Indian population, particularly among young professionals who are dealing with the challenges of urban lifestyles and occupational stress.
Conclusion
The present study is the first to investigate genetic predispositions to T2D among young corporate professionals in India. In this study, 42 genetic loci were found to be significantly associated with the T2D risk, including missense variants in SULT1C4, FGA, and TAS2R38, and UTR variants in HLA-G and HTR1B. These genes are involved in pathways regulating glucose homeostasis, insulin signaling, obesity, inflammation, and stress response. The study highlights the role of genetic factors interacting with occupational stress, physical activity, and unhealthy diet. The findings of this study will pave the way for personalised risk assessments and tailored prevention strategies, emphasising the importance of further exploring gene-environment interactions to address the rising T2D burden in India. The replication studies are warranted to validate these findings and expand genomic research in understudied populations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Acknowledgements
All Authors acknowledge Mr. Masroor Lodi, Unlock Wellness Pvt. Ltd. for providing “Predictor” to manage clinical data and Mr. Adireddi Govind Rao, NMC Genetics India Pvt. Ltd. for providing the research facilities and Pensive.one for genomic data management. Authors acknowledge Mr. Hetender Singh (B-Ezy Diagnostics Pvt. Ltd.) and Dr. Megha Sharma, Assistant Editor (English), Directorate of Publications Division, M/o Information and Broadcasting, Government of India for assisting in the proofreading of the manuscript.
Author contributions
S.F.H., H.S., I.S., and V.S. conceptualize the idea, S.F.H. and T.K. assisted in sample and Data collection, G.R. and P.M. executed the necessary procedures for sample processing. LG generated the data. L.S., H.S. and V.S. did the data analyses and wrote the first draft. S.F.H., T.K., H.S., I.S. and V.S. collaboratively finalized and refined the manuscript.
Funding
No funding was received to conduct this study.
Data availability
All data described in this study are provided within the article and Supplementary Material. The raw genotyping and clinical data are available from the corresponding authors upon request.
Code availability
The code used in the manuscript is present in the public repository https://github.com/sharmavarun840.
Declarations
Competing interests
The authors declare no competing interests.
Ethical approval
All procedures performed were in accordance with the ethical standards of the institutional and/or national research committee and with the Helsinki Declaration 2013 and its later amendments or comparable ethical standards.
Disclosures
Shah Fahad Husami and Tavleen Kaur are employees of Unlock Wellness Pvt. Ltd. Love Gupta, Garima Rastogi, Pooja Meena, Varun Sharma, and Hemender Singh are employees of NMC Genetics India Pvt. Ltd, the research organisation that conducted the study.
Footnotes
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Contributor Information
Indu Sharma, Email: induadmo@gmail.com.
Hemender Singh, Email: hemender.singh4737@gmail.com.
Varun Sharma, Email: sharmavarun840@gmail.com.
References
- 1.de Candia, P. et al. Type 2 diabetes: How much of an autoimmune disease? Front. Endocrinol.10, 451 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Saad, B. et al. Introduction to diabetes and obesity. In Anti-diabetes and Anti-obesity Medicinal Plants and Phytochemicals: Safety, Efficacy, and Action Mechanisms 3–19 (2017).
- 3.Mancusi, C. et al. Insulin resistance the hinge between hypertension and type 2 diabetes. High. Blood Press. Cardiovasc. Prev.27, 515–526 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Bahari, N. I. et al. Issues and challenges in the primary prevention of type 2 diabetes mellitus: A systematic review. J. Prev.44(1), 105–125 (2023). [DOI] [PubMed] [Google Scholar]
- 5.Pradhan, P. et al. Genetic and epigenetic modifications in the pathogenesis of diabetic retinopathy: A molecular link to regulate gene expression. New. Front. Ophthalmol. (Lond.)2(5), 192 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Habtemariam, S. Medicinal Foods as Potential Therapies for Type-2 Diabetes and Associated Diseases: The Chemical and Pharmacological Basis of Their Action (Academic, 2019).
- 7.Uddin, M. J. et al. A comparison of machine learning techniques for the detection of type-2 diabetes mellitus: Experiences from Bangladesh. Information14(7), 376 (2023). [Google Scholar]
- 8.Hu, H. et al. A systematic review of the direct economic burden of type 2 diabetes in China. Diabetes Ther.6, 7–16 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Mambiya, M. et al. The play of genes and non-genetic factors on type 2 diabetes. Front. Public. Health7, 349 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.De Rosa, S. et al. Type 2 diabetes mellitus and cardiovascular disease: Genetic and epigenetic links. Front. Endocrinol.9, 2 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Prasad, R. B. & Groop, L. Genetics of type 2 diabetes—pitfalls and possibilities. Genes6(1), 87–123 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Kaul, N. & Ali, S. Genes, genetics, and environment in type 2 diabetes: Implication in personalized medicine. DNA Cell Biol.35(1), 1–12 (2016). [DOI] [PubMed] [Google Scholar]
- 13.Murea, M., Ma, L. & Freedman, B. I. Genetic and environmental factors associated with type 2 diabetes and diabetic vascular complications. Rev. Diabet. Stud. RDS9(1), 6 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Pandey, A., Chawla, S. & Guchhait, P. Type-2 diabetes: Current understanding and future perspectives. IUBMB Life67(7), 506–513 (2015). [DOI] [PubMed] [Google Scholar]
- 15.Saltiel, A. R. & Kahn, C. R. Insulin signalling and the regulation of glucose and lipid metabolism. Nature414(6865), 799–806 (2001). [DOI] [PubMed] [Google Scholar]
- 16.Romao, I. & Roth, J. Genetic and environmental interactions in obesity and type 2 diabetes. J. Am. Diet. Assoc.108(4), S24–S28 (2008). [DOI] [PubMed] [Google Scholar]
- 17.Pradeepa, R. & Mohan, V. Prevalence of type 2 diabetes and its complications in India and economic costs to the nation. Eur. J. Clin. Nutr.71(7), 816–824 (2017). [DOI] [PubMed] [Google Scholar]
- 18.Farag, Y. M. & Gaballa, M. R. Diabesity: An overview of a rising epidemic. Nephrol. Dial. Transpl.26(1), 28–35 (2011). [DOI] [PubMed] [Google Scholar]
- 19.Emmert-Fees, K. M. et al. Cost-effectiveness of a collaborative care model among patients with type 2 diabetes and depression in India. Diabetes Care46(1), 11–19 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Brar, S. et al. Cost of screening, out-of-pocket expenditure & quality of life for diabetes & hypertension in India. Indian J. Med. Res.157(6), 498 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Madhu, S. V. et al. Chronic stress, sense of coherence and risk of type 2 diabetes mellitus. Diabetes Metab. Syndr. Clin. Res. Rev.13(1), 8–23 (2019). [DOI] [PubMed]
- 22.Nagarathna, R. et al. Prevalence of diabetes and its determinants in the young adults Indian population-call for yoga intervention. Front. Endocrinol.11 (2020). [DOI] [PMC free article] [PubMed]
- 23.Perng, W. et al. Youth-onset type 2 diabetes: The epidemiology of an awakening epidemic. Diabetes Care46(3), 490–499 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Ali, O. Genetics of type 2 diabetes. World J. Diabetes. 4(4), 114–123 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Heraclides, A. et al. Psychosocial stress at work doubles the risk of type 2 diabetes in middle-aged women: Evidence from the Whitehall II study. Diabetes Care32(12), 2230–2235 (2009). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Cosgrove, M. P. et al. Work-related stress and type 2 diabetes: Systematic review and meta-analysis. Occup. Med. (Lond.)62(3), 167–173 (2012). [DOI] [PubMed] [Google Scholar]
- 27.Skol, A. D. et al. Joint analysis is more efficient than replication-based analysis for two-stage genome-wide association studies. Nat. Genet.38(2), 209–213 (2006). [DOI] [PubMed] [Google Scholar]
- 28.2. Diagnosis and classification of diabetes: Standards of Care in Diabetes-2024. Diabetes Care47(Suppl 1): S20–S42 (2024). [DOI] [PMC free article] [PubMed]
- 29.Bhat, K. G. et al. Preliminary genome wide screening identifies new variants associated with coronary artery disease in Indian population. Am. J. Transl. Res.14(7), 5124–5131 (2022). [PMC free article] [PubMed] [Google Scholar]
- 30.Bhat, K. G. et al. Pharmacogenomic evaluation of CYP2C19 alleles linking low clopidogrel response and the risk of acute coronary syndrome in indians. J. Gene Med. e3634 (2023). [DOI] [PubMed]
- 31.Purcell, S. et al. PLINK: A tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet.81(3), 559–575 (2007). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.McLaren, W. et al. The ensembl variant effect predictor. Genome Biol.17(1), 1–14 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Watanabe, K. et al. Functional mapping and annotation of genetic associations with FUMA. Nat. Commun.8(1), 1826 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Franz, M. et al. GeneMANIA update 2018. Nucleic Acids Res.46(W1), W60–W64 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Pradeepa, R. & Mohan, V. Epidemiology of type 2 diabetes in India. Indian J. Ophthalmol.69 (11), 2932–2938 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Hong, E. P. & Park, J. W. Sample size and statistical power calculation in genetic association studies. Genomics Inf.10 (2), 117–122 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Joshi, S. R. & Parikh, R. M. India; the diabetes capital of the world: Now heading towards hypertension. J. Assoc. Physicians India55(Y), 323 (2007). [PubMed] [Google Scholar]
- 38.Liu, T. et al. The relationship between gestational diabetes mellitus and interleukin 1beta gene polymorphisms in southwest of China. Med. (Baltim).99(43), e22679 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Chen, M. et al. CTNNB1/β-catenin dysfunction contributes to adiposity by regulating the cross-talk of mature adipocytes and preadipocytes. Sci. Adv.6(2), eaax9605 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Zaharieva, E. et al. Interleukin-18 serum level is elevated in type 2 diabetes and latent autoimmune diabetes. Endocr. Connect.7(1), 179–185 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Dotson, C. D. et al. Bitter taste receptors influence glucose homeostasis. PLoS One3(12), e3974 (2008). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Krijgsman, D. et al. HLA-G: A new immune checkpoint in cancer?. Int. J. Mol. Sci.21(12) (2020). [DOI] [PMC free article] [PubMed]
- 43.Xu, Z. Y., Jing, X. & Xiong, X. D. Emerging role and mechanism of the FTO Gene in Cardiovascular diseases. Biomolecules13(5) (2023). [DOI] [PMC free article] [PubMed]
- 44.Redan, B. W. et al. Altered transport and metabolism of phenolic compounds in obesity and diabetes: Implications for functional food development and assessment. Adv. Nutr.7(6), 1090–1104 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Hwang, J. Y. et al. Association between single nucleotide polymorphisms of the fibrinogen alpha chain (FGA) gene and type 2 diabetes Mellitus in the Korean population. Genomics Inf.7(2), 57–64 (2009). [Google Scholar]
- 46.Vojtková, J. et al. An association between fibrinogen gene polymorphisms and diabetic peripheral neuropathy in young patients with type 1 diabetes. Mol. Biol. Rep.48(5), 4397–4404 (2021). [DOI] [PubMed] [Google Scholar]
- 47.Pan, H. T. et al. Proteomics and bioinformatics analysis of cardiovascular related proteins in offspring exposed to gestational diabetes mellitus. Front. Cardiovasc. Med.9, 1021112 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Keller, M. et al. TAS2R38 and its influence on smoking behavior and glucose homeostasis in the German sorbs. PLoS One. 8(12), e80512 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Dotson, C. D. et al. Variation in the gene TAS2R38 is associated with the eating behavior disinhibition in old order amish women. Appetite54(1), 93–99 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Shobeiri, S. S. et al. Evaluation of soluble human leukocyte antigen-G in peripheral blood of pregnant women with gestational diabetes mellitus. Casp. J. Intern. Med.7(3), 178–182 (2016). [PMC free article] [PubMed] [Google Scholar]
- 51.Silva, H. P. et al. The association between the HLA-G 14-bp insertion/deletion polymorphism and type 1 diabetes. Genes Immun.17(1), 13–18 (2016). [DOI] [PubMed] [Google Scholar]
- 52.Sukhov, I. B. et al. The effect of prolonged intranasal administration of serotonin on the activity of hypothalamic signaling systems in male rats with neonatal diabetes. Cell. Tissue Biol.10(4), 314–323 (2016). [Google Scholar]
- 53.Ohara-Imaizumi, M. et al. Serotonin regulates glucose-stimulated insulin secretion from pancreatic β cells during pregnancy. Proc. Natl. Acad. Sci. U S A110(48), 19420–19425 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Yabut, J. M. et al. Emerging roles for serotonin in regulating metabolism: New implications for an ancient molecule. Endocr. Rev.40(4), 1092–1107 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Bennet, H. et al. Serotonin (5-HT) receptor 2b activation augments glucose-stimulated insulin secretion in human and mouse islets of Langerhans. Diabetologia59(4), 744–754 (2016). [DOI] [PubMed] [Google Scholar]
- 56.Kim, H. et al. Serotonin regulates pancreatic beta cell mass during pregnancy. Nat. Med.16(7), 804–808 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Corbin, L. J. et al. BMI as a modifiable risk factor for type 2 diabetes: Refining and understanding causal estimates using mendelian randomization. Diabetes65(10), 3002–3007 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Hromatka, B. S. et al. Genetic variants associated with motion sickness point to roles for inner ear development, neurological processes and glucose homeostasis. Hum. Mol. Genet.24(9), 2700–2708 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Wang, L. et al. TGF-Beta as a master regulator of diabetic nephropathy. Int. J. Mol. Sci.22(15) (2021). [DOI] [PMC free article] [PubMed]
- 60.Shojima, N. & Yamauchi, T. Progress in genetics of type 2 diabetes and diabetic complications. J. Diabetes Investig.14(4), 503–515 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Guo, H. et al. The expression of Tim-1 and Tim-4 molecules in regulatory T cells in type 1 diabetes. Endocrine68(1), 64–70 (2020). [DOI] [PubMed] [Google Scholar]
- 62.Wang, H. J. et al. Chronic HMGCR/HMG-CoA reductase inhibitor treatment contributes to dysglycemia by upregulating hepatic gluconeogenesis through autophagy induction. Autophagy11(11), 2089–2101 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Ereqat, S. et al. Estrogen receptor 1 gene polymorphisms (PvuII and XbaI) are associated with type 2 diabetes in Palestinian women. PeerJ7, e7164 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Marroquí, L. et al. BACH2, a candidate risk gene for type 1 diabetes, regulates apoptosis in pancreatic β-cells via JNK1 modulation and crosstalk with the candidate gene PTPN2. Diabetes63(7), 2516–2527 (2014). [DOI] [PubMed] [Google Scholar]
- 65.Chandak, G. R. et al. Common variants in the TCF7L2 gene are strongly associated with type 2 diabetes mellitus in the Indian population. Diabetologia50(1), 63–67 (2007). [DOI] [PubMed] [Google Scholar]
- 66.Auton, A. et al. A global reference for human genetic variation. Nature526(7571), 68–74 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Nassar, L. R. et al. The UCSC genome browser database: 2023 Update. Nucleic Acids Res.51(D1), D1188–D1195 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
All data described in this study are provided within the article and Supplementary Material. The raw genotyping and clinical data are available from the corresponding authors upon request.
The code used in the manuscript is present in the public repository https://github.com/sharmavarun840.