Abstract
In previous work, we showed that telomeres of normal cells are organized within the 3D space of the interphase nucleus in a nonoverlapping and cell cycle-dependent manner. This order is distorted in tumor cell nuclei where telomeres are found in close association forming aggregates of various numbers and sizes. Here we show that c-Myc overexpression induces telomeric aggregations in the interphase nucleus. Directly proportional to the duration of c-Myc deregulation, we observe three or five cycles of telomeric aggregate formation in interphase nuclei. These cycles reflect the onset and propagation of breakage-bridge-fusion cycles that are initiated by end-to-end telomeric fusions of chromosomes. Subsequent to initial chromosomal breakages, new fusions follow and the breakage-bridge-fusion cycles continue. During this time, nonreciprocal translocations are generated. c-Myc-dependent remodeling of the organization of telomeres thus precedes the onset of genomic instability and subsequently leads to chromosomal rearrangements. Our findings reveal that c-Myc possesses the ability to structurally modify chromosomes through telomeric fusions, thereby reorganizing the genetic information.
Keywords: genomic instability, 3D nucleus, breakage-bridge-fusion
Multiple alterations accompany tumor initiation and progression resulting in the modulation of gene expression and in genomic instability. These interconnected changes occur within nuclei that harbor an altered 3D organization (1–3). In agreement with this concept, recent reports suggest tumor-associated changes of chromosomal organization in an altered 3D nucleus (3–8). However, mechanisms leading to structural changes of telomeres and chromosomes remain elusive.
We recently reported that the normal interphase nucleus has a unique 3D telomeric organization that is cell cycle dependent (9, 10). Telomeres are organized in a nonoverlapping manner and align into a central telomeric disk during the late G2 phase of the cell cycle (9). In contrast, tumor cells display an aberrant organization of telomeres that can be objectively measured in nuclei showing telomeric aggregates of various complexity and sizes (9).
Constitutive expression of c-Myc due to chromosomal translocations, mutation, or amplification contributes to the development and progression of many cancers (11, 12). c-Myc deregulation directly promotes genomic instability (13), causing locus-specific and karyotypic instability (14–18). Additionally, c-Myc induces illegitimate replication initiation (19, 20), DNA breakage (21), alterations of DNA repair (22, 23), and a low level of point mutations (24, 25). Effects of c-Myc on genomic instability are reversible after a transient experimental activation of c-Myc (15). However, c-Myc continues to generate instability after constitutive deregulation (16). In vivo, c-Myc deregulation directly initiates and promotes tumorigenesis (26–30). When c-Myc deregulation is abolished, in vivo tumorigenesis is reversible, provided that no additional mutations had occurred (29–34).
Prompted by the complexity of downstream genetic alterations that result from c-Myc deregulation, we investigated whether c-Myc affected the 3D organization of the mammalian interphase nucleus and whether this remodeling had an impact on genomic stability. We show that c-Myc deregulation causes remodeling of the 3D nuclear organization of telomeres and chromosomes, thus creating the topological conditions that initiate genomic instability.
Materials and Methods
Cells and Conditional Myc Activation. Culture conditions have been described for Ba/F3 (35) and PreB (36) cells. The plasmacytoma cell line MOPC460D was a gift of J. Mushinski (National Institutes of Health, Bethesda). Cell viability was determined by hemocytometer counts by using trypan blue. The primary mouse plasmacytoma DCPC21 was isolated from a BALB/c mouse (37). v-abl/myc-induced plasmacytomas (38) and primary lymphocytes were collected from BALB/c mice (Central Animal Care protocol 02-039).
To activate MycER (39) in Ba/F3 or PreB cells, 105 cells per ml were treated with 100 nM 4-hydroxytamoxifen (4HT). Cells were split 24 h before 4HT treatment. Non-4HT treated control cells were cultivated in ethanol, which is used to dissolve 4HT (25, 26, 39). Two different MycER activation schemes were performed. First, analyses of c-Myc-induced changes in 3D telomere organization were carried out after a single addition of 4HT that was left in the culture medium until its biological effects subsided (40–42). Nuclei were examined every 24 h over a 10-day period. A second time course was performed every 6 h for 120 h (Fig. 1). To enable a time-dependent analysis of Myc activation, 4HT was given for 2 or 12 h and was removed. Alternatively, 4HT was added every 12 h or was given once but left in the culture. MycER activation was determined by fluorescent immunohistochemistry.
Fig. 1.
MycER activation scheme. The effects of 4HT last 15–24 h in cell lines (40–42), as indicated by dashed lines. Cells were harvested every 6 h over a time period of 120 h. Mock-treated control cells were processed in parallel.
Immunohistochemistry (IHC). Fluorescent IHC of Myc protein was performed as described in ref. 43 by using a polyclonal anti-c-Myc antibody (N262; Santa Cruz Biotechnology) and a goat anti-rabbit IgG FITC antibody, each at a dilution of 1:100. Analysis was performed by using a Zeiss Axiophot 2 microscope. Images were acquired with a Cooke CCD SensiCam Camera.
Cell Death. Apoptotic bodies for control and MycER-activated cells were assessed by two independent observers who scored 300 DAPI-stained nuclei per time point in the presence or absence of MycER activation.
Telomere FISH. Ba/F3, PreB, and plasmacytoma cells were collected (200 × g for 10 min) and resuspended in PBS containing 3.7% formaldehyde (Fluka) and incubated for 20 min. Thereafter, the telomere FISH protocol was performed (9, 44) by using Cy3- or FITC-labeled PNA probes (DAKO). Three independent experiments were performed. At least 30 nuclei and 20 metaphases were examined per time point. Imaging of metaphases after telomere FISH was performed by using Zeiss Axioplan 2 with a cooled AxioCam HR B&W, DAPI, Cy3, or FITC filters in combination with Planapo 63x/1.4 oil objective lens. Images were acquired by using axiovision 3.1 (Zeiss) in multichannel mode. Because of the presence of multiple variables, the general linear modeling procedure was used. To test average aggregates among different groups, a two-way ANOVA test was performed for normality and robustness of the data. For details of all tests preformed, see Supporting Materials and Methods, which is published as supporting information on the PNAS web site.
3D Image Acquisition. At least 30 nuclei were analyzed for each time point. axiovision 3.1 with deconvolution module and rendering module were used. For every fluorochrome, the 3D image consists of a stack of 100 images with a sampling distance of 200 nm along the z and 107 nm in the xy direction. The constrained iterative algorithm option was used (45).
3D Image Analysis for Telomeres. Telomere measurements were done with teloview (9, 46). By choosing a simple threshold for the telomeres, a binary image is found. Based on that, the center of gravity of intensities is calculated for every object resulting in a set of coordinates (x, y, z) denoted by crosses on the screen. The integrated intensity of each telomere is calculated because it is proportional to the telomere length (47). The integration region is determined by growing a sphere on top of the found coordinate. After every step of growth (iteration), the sum under this volume (the telomere) is subtracted by the sum just surrounding it (background level). When the process of the growth of the sphere does not contribute to an integrated intensity increase, the algorithm stops and the integrated intensity of the telomere with an automatic background correction is obtained.
Chromosome Painting and Measurements of Chromosomal Overlap(s) in Interphase Nuclei. Chromosome painting was carried out as described in ref. 48 by using paints for mouse chromosomes 5 (Cy3), 13 (FITC), 7 (Cy3), 10 (FITC), and 17 (FITC) from Applied Spectral Imaging (Vista, CA). 3D image acquisition of painted nuclei was performed as described above. Measurements of chromosomal overlaps were performed after 3D image acquisition and constrained iterative deconvolution as follows: (i) based on the DAPI counterstain image, we determined the 3D boundary of the nuclear volume. Data outside that volume were ignored. (ii) For each one of the chromosomes, we determined an intensity threshold and referred only to voxels that were above the threshold that belonged to the specific chromosomes. The total volume occupied by each one of the chromosome pairs is measured (V1 and V2). (iii) The volume occupied by both chromosome pairs is measured, Vo. By dividing this value by V1 and by V2, the level of overlap relative to the total volume of each chromosome pair was measured, V0/V1, V0/V2 (for details, see Fig. 8 which is published as supporting information on the PNAS web site).
Spectral Karyotyping (SKY). Mouse SKY was performed by using a SKY system (Applied Spectral Imaging) (37). Twenty metaphases were examined per time point. Significant values for chromosomal rearrangements were determined after MycER activation. Mean total chromosomes and numbers of each chromosome observed for control and Myc-activated cells were compared over time by two-way ANOVA. In addition, statistical analyses were performed for the occurrence of translocations, breakages, and fusions over the experimental period of 120 h. P values of <0.05 were considered significant. Only the frequency procedure was used, followed by Fisher's exact test. The P value of the overall study was <0.0001.
Supporting Information. For additional information, see Figs. 9–12, Movies 1–3, and Tables 2–4, which are published as supporting information on the PNAS web site.
Results
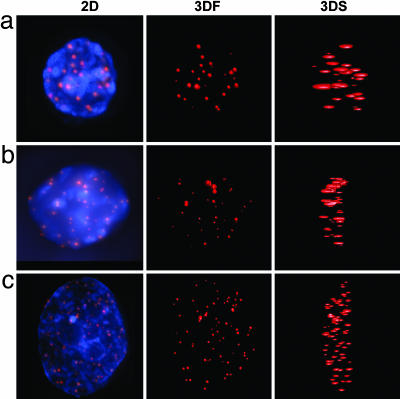
The 3D Organization of Telomeres Before c-Myc Activation. We examined whether c-Myc deregulation affected the 3D organization of telomeres in the interphase nucleus. To this end, we analyzed the effect of conditional c-Myc expression in two independent immortalized mouse B lymphocyte lines, Ba/F3 (35) and PreB (36), stably transfected with MycER (39). For both cell lines, we first evaluated the 3D organization of telomeres in nuclei of non-MycER-activated cells by using primary BALB/c B lymphocytes as a control. Consistent with our previous studies (9), telomeres of normal primary BALB/c B nuclei showed nonoverlapping telomere positions as determined by 3D imaging (Fig. 2a). Without MycER activation, both PreB and Ba/F3 interphase nuclei also displayed nonoverlapping telomere positions (Fig. 2 b and c, respectively). Therefore, the above cell lines were appropriate to study the effects of conditional c-Myc activation on the 3D telomeric organization.
Fig. 2.
Telomeric organization in interphase nuclei of primary and immortalized B lymphocytes without overlap in telomere positions. (a) Primary B cell nucleus. (b) Nucleus of near diploid PreB cell. (c) Nucleus of tetraploid Ba/F3 cell. Telomeres are shown in red; nuclei in blue. 3DF, 3D front view; 3DS, 3D side view.
c-Myc-Dependent Disruption of the 3D Telomeric Organization: Formation of Telomeric Aggregates (TAs) in Interphase Nuclei. We next analyzed the effect of conditional c-Myc expression on the 3D organization of telomeres. After a transient MycER activation with 4HT, nuclear c-Myc signals were observed in both PreB and Ba/F3 cells (Fig. 9 b and d). In non-4HT treated control cells, MycER was found in the cytoplasm (Fig. 9 a and c; see also ref. 39).
To determine whether c-Myc deregulation affected the 3D organization of telomeres, we performed time course experiments. In the first set of experiments, c-Myc deregulation and 3D telomeric organization were investigated in both PreB and Ba/F3 cells after a single 4HT treatment. Nuclei were analyzed after c-Myc deregulation at 0, 24, 48, 72, and 96 h and at 10 days and compared with nuclei from mock-treated control cells. In both cell lines, analyses of the 3D nuclear organization of telomeres revealed that c-Myc deregulation induced the formation of TAs. TAs are group(s) of telomeres that are found in clusters and, thus, in close association in the interphase nucleus. This 3D telomeric organization is distinct from the normal 3D organization of non-MycER-activated PreB, Ba/F3 cells and primary mouse lymphocytes (Fig. 2). Fig. 3 illustrates the presence of TAs in interphase nuclei of MycER-activated PreB and Ba/F3 cells (Fig. 3 b and d, respectively). Although such TAs had been observed in tumor cell nuclei previously (9), their presence in conditional c-Myc expressing cells is a previously uncharacterized finding.
Fig. 3.
c-Myc deregulation induces TAs in interphase nuclei of PreB and Ba/F3 cells shown at 72 h after 4HT-treatment. (a) Mock-treated PreB cells show nonoverlapping telomeres (red). (b) MycER-activated PreB cells with TAs (green arrow). (c) Mock-treated Ba/F3 cells show nonoverlapping telomeres. (d) MycER-activated Ba/F3 cells show the formation of TAs (green arrow).
c-Myc Induces Cycles of TAs in Interphase Nuclei. In subsequent experiments, we investigated the time relationship between c-Myc deregulation and the formation of TAs more closely. To this end, cells were harvested every 6 h over a time period of 120 h. We also varied the duration of conditional c-Myc expression (Fig. 1), confirming nuclear c-Myc staining as above (Fig. 9 and 11). Next, the 3D organization of telomeres was determined (Fig. 4). At this point, we focused on near diploid PreB cells only (49). Our positive controls were cells constitutively overexpressing c-Myc [mouse plasmacytomas (27) and a plasmacytoma line (Fig. 4Ae)]. Negative controls were mock-treated PreB cells (Fig. 4Aa).
Fig. 4.
c-Myc-induced telomeric aggregates appear in cycles. (A) Conditional c-Myc deregulation causes TA formation. (Aa) Negative control: non-Myc-deregulated PreB nucleus with nonoverlapping 3D telomeric nuclear positions. (bb–bd) TAs of various sizes and numbers are present after conditional c-Myc expression at any given time point of TA formation. Telomeres are shown in green; TAs by red arrows. (Ae) Positive control: plasmacytoma cell line, MOPC460D, with constitutive c-Myc deregulation due to T12;15, shows TAs. Similar results were obtained with primary plasmacytoma cells (data not shown). (B) c-Myc induces cycles of TAs. Fold increase in TAs over control levels during a period of 120 h. During this period, c-Myc had been up-regulated for different lengths of time (see Fig. 1). Black, 4HT given for 2 h and removed; red, 4HT administered for 12 h and removed; green, 4HT added once and not removed; blue, 4HT added at 0, 12, and 24 h; gray, control cells. The highest levels of TA formation and a single TA peak observed after consecutive activations of MycER are shown by arrows and an arrowhead, respectively. Error bars represent a 95% confidence interval of binomial distributions.
This time course confirmed that c-Myc deregulation induced TAs. Representative images show that TAs varied in size and numbers per MycER-activated PreB cell nucleus (Fig. 4A b–d, red arrows). High induction levels of TAs were observed at 30, 48, 72, and 96 h declining after 96 h (Fig. 4B, arrows). The highest levels of TA formation will hereafter be referred to as peaks of TAs. The 6-h time course performed over 120 h indicated that TAs formed in a c-Myc-dependent manner and showed a cyclic appearance (Fig. 4B). The number of TA cycles was directly linked to the duration of c-Myc deregulation. For example, 2 h of Myc activation induced three such cycles, whereas 12 h led to five cycles (Fig. 4B, black and red lines, respectively). 4HT, left in the culture medium until its biological effects on our cells subsided (Fig. 1), also induced five TA cycles (Fig. 4B, green line). In this context, repeated consecutive activations of MycER given every 12 h caused TAs in 96% of all nuclei. These cells died after 30 h (Fig. 4B, blue line) because of repeated cycles of c-Myc deregulation and not due to toxicity exerted by 4HT (50). Thus, only a single TA cycle is observed in this experimental setting (Fig. 4B, arrowhead). The increase in TAs and 3D volumes was significant (Table 3).
The c-Myc-Induced TA Cycles Represent Breakage-Bridge-Fusion (BBF) Cycles and Chromosomal Rearrangements. The cycles of c-Myc induced TAs in PreB nuclei showed similar periodicity for all c-Myc activation periods (Fig. 4B). We reasoned that these cycles might reflect both ongoing associations and dissociations of telomeres or BBF cycles. The BBF cycle could be induced by the breakage of dicentric chromosomes during anaphase-inducing apoptosis of cells having multiple or large TAs per nucleus. To address these possibilities, we first examined metaphase chromosomes at different times: prior, during, and after the peaks of TA formation for a 120-h period. We used both whole-genome analysis by mouse SKY and telomeric FISH of metaphase chromosomes. A significant level of dicentric chromosomes was noted (Fig. 5). Control cells had normal karyotypes (Fig. 12). In MycER-activated PreB cells, however, fusions had occurred. We show as example fusions at the telomeric ends of chromosomes 18 and 4 (Fig. 5a), red and green arrows) and between two chromosomes 1 (Fig. 5a, green arrow). Chromosome 1 was probably broken in the previous anaphase (Fig. 5a, green circle). An additional terminally deleted chromosome 1 is in the center of the same metaphase plate, and chromosomes 2, 3, and 7 reveal terminal deletions (Fig. 5a). Telomeric fusions involving both ends of chromosomes as well as sister chromatids were confirmed by telomeric FISH (Fig. 5b). Anaphase bridges and ring chromosomes were present (Fig. 5c) and data not shown).
Fig. 5.
Molecular cytogenetic evidence of BBF cycles in MycER-activated PreB cells. (a) SKY analysis reveals telomeric fusions and chromosome breakage. (a Upper) Metaphase, raw image (Left); metaphase, classified image (Center); and metaphase, inverted DAPI image (Right). (a Lower) Spectral karyotype. End-to-end fusion of chromosomes 18 and 4 (red arrow) and fusion of chromosome 1 with a broken piece of chromosome 1 (green arrow) are shown. One broken chromosome 1 is circled. Note additional broken chromosomes 1, 2, 3, and 7. (b) End-to-end fusions of chromosomes revealed by telomere FISH. (Upper) Centromeric fusion (see arrow and insert). (Lower) Telomeric fusion (see arrow and insert). (c) Anaphase bridges. (Upper) Short exposure of DAPI stained nucleus (100 msec). (Lower) Longer exposure (500 msec) of same image makes anaphase bridge visible (white arrow) but overexposes nuclei. (d) SKY illustrating chromosomal fusions (red arrow) and nonreciprocal translocations (white arrows). Broken chromosomes are also present (chromosomes 4, 6, 12, and 17).
The nature of c-Myc-induced 3D structural changes in interphase nuclei of conditionally Myc expressing cells was as follows: at peaks of TA formation and thereafter, a significant increase in end-to-end chromosomal fusions over control levels was observed. This result was followed by a significant increase in broken chromosomes and nonreciprocal translocations (Figs. 5d and 6 and Table 2). In conclusion, TA cycles unveil BBF cycles, namely the fusions of two chromosomes, consequently, the formation of dicentrics and their subsequent breakage in anaphase (Fig. 5). The cycles are induced by conditional Myc deregulation and lead to the onset of genomic instability, demonstrated by the chromosomal rearrangements resulting from these BBF cycles (Figs. 5 and 6 and Table 2).
Fig. 6.
Chromosomal aberrations in MycER-activated PreB cells over a period of 120 h after a single administration of 4HT. End-to-end fusions (blue) increase to 40% in the first 12 h. Over time, the percentage of fusions decreases. Translocations (orange) appear at 12 h and reach a maximum of 35% at 42 h. Telomere-free chromosomal end(s) (green) increase over time peaking at 30 h with 75% of metaphases having at least one telomere-free chromosomal end. Subsequently, the percentage of telomere-free chromosomal end(s) decreases. Q-FISH experiments confirmed healing of telomeric ends at later time points. The error bars show the 95% confidence interval for binomial distributions (51). Because of a confidence interval, the error bars are larger than expected when a standard error would have been used, which was not applicable in this situation. For details on each time point and aberration, see Table 2.
Next, we investigated whether cells with TAs died during the course of the experiments. If this possibility was the case, we would expect a correlation of cell death in Myc-activated cells at the peak of TA formation or shortly thereafter. The level of apoptosis was ≈2-fold higher in Myc-activated cells than in control cells (Table 1). There was no preference in apoptotic cell death for any specific time point during the 120 h. We concluded that BBF cycles, not apoptosis, contributed to the cycles of TA formation.
Table 1. Apoptosis levels in non-MycER and MycER-activated PreB cells.
| % apoptosis
|
|||
|---|---|---|---|
| Time, h | Controls | MycER-activated PreB cells | Fold increase |
| 0 | 3.0 | 3.0 | 1.0 |
| 12 | 2.0 | 4.0 | 2.0 |
| 24 | 6.0 | 12.0 | 2.0 |
| 30 | 4.0 | 10.0 | 2.5 |
| 42 | 2.0 | 8.0 | 4.0 |
| 48 | 5.0 | 10.0 | 2.0 |
| 66 | 4.0 | 11.0 | 2.75 |
| 72 | 3.0 | 8.0 | 2.7 |
| 84 | 3.0 | 5.0 | 1.7 |
| 96 | 3.0 | 3.0 | 1.0 |
| 102 | 2.0 | 3.0 | 1.5 |
3D Organization of Chromosomes in c-Myc Activated Interphase Nuclei. TAs and the initiation of BBF cycles with subsequent chromosomal rearrangements prompted us to investigate whether chromosomes were affected in their 3D nuclear positions during MycER activation. To this end, we examined the overlap of specific chromosomes over the 120-h period. SKY of MycER-activated PreB cells suggested chromosomal rearrangements involving chromosomes 7, 13, and 17. Additional rearrangements were found but did not reach significant levels (data not shown). We examined three combinations of chromosomes over a 96-h period. This period covered all peaks of TA formation (Fig. 4B). As shown in Fig. 7, we observed a change in overlaps between chromosomes 5 (red) and 13 (green) over the time course (Figs. 7 A and B). Both chromosomes were found in closer vicinity as the cells entered into the first TA cycle. Chromosomes 10 (green) and 7 (red) also showed increases in the percentage of overlap (Fig. 7 A and B), as did chromosomes 7 (red) and 17 (green) (Fig. 7 A and B). Representative 3D movies are shown in Movies 1–3.
Fig. 7.
Chromosome positions in Myc-activated nuclei. (A) Representative nuclei painted with chromosome paints over a period of 96 h after 4HT (Fig. 1). (Top) Chromosomes 5 and 13. (Middle) Chromosomes 7 and 10. (Bottom) Chromosomes 7 and 17. (B) Measurements of chromosomal overlaps in nuclei of c-Myc deregulated cells for chromosomes 5 and 13 (Left), 7 and 10 (Center), and 7 and 17 (Right) over a 96-h period.
Discussion
c-Myc Induces Telomeric Aggregates, Fusions, and BBF Cycles. Previous studies have shown that c-Myc triggers a complex network of genomic instability at the level of single genes (14, 15, 19) and whole chromosomes (16–18) (for review, see ref. 13). In addition, c-Myc induces illegitimate replication initiation (19, 20), chromosomal rearrangements (18), DNA breakage, alterations of DNA repair (21–23), and a low level of point mutations (24, 25). A previously uncharacterized mechanism underlying c-Myc-dependent genomic instability at the chromosomal level directly affects the integrity of the telomeres and was revealed in this study.
The clear periodicity of the TA cycles that was found with four different Myc-activating treatments suggested a biological relevant Myc-dependent process. Theoretically, cycles of Myc-induced TAs could reflect (i) nuclear remodeling with the transient association and subsequent dissociation of telomeres; (ii) end-to-end chromosomal fusions that initiate BBF cycles (52, 53); (iii) c-Myc induced cell death; and (iv) a combination of all of the above. Our data are consistent with BBF cycles and exclude apoptosis as a direct contributor to the TA cycles. Apoptosis occurred at equal levels throughout the study and consistently reached about twice the levels seen in the control cells. The loss of cells was compensated by a 2-fold increase in proliferation in MycER-activated PreB (19). These data also indicate that there is genetic separation of genomic instability and apoptosis as reported in ref. 54. Whether telomere associations and dissociations (55) contributed to the TA cycles is presently unknown.
Direct evidence of BBF cycles in the periodicity of TAs came from a detailed analysis of chromosomal fusions, breakage, and rearrangements observed over the time course of five TA cycles. We demonstrated the occurrence of end-to-end fusions that generated dicentric chromosomes and breaks during anaphase, leaving one chromosome or chromatid with a piece from another chromosome or chromatid. The resulting telomere-free ends continue to undergo fusions with other chromosomes, a cycle of events termed BBF cycle (52, 53). Experimental data support these events from fusions to breakages and nonreciprocal translocations. The periodicity of the TA cycles is consistent with a ≈12 h population doubling time of the PreB cells (19). Each peak of TAs is consistent with the repeated formation of TAs. Time points after the peak are in agreement with the breakage of dicentric chromosomes. Telomere-free ends initiate new BBF cycle(s) until no more telomere-free chromosomal end(s) persist.
From Telomeres to Chromosomal Rearrangements: A New Pathway of c-Myc-Dependent Genomic Instability. Muller (52) and McClintock (53) first described BBF cycles, a mechanism of chromosomal end-to-end fusion that contributes to the onset of genomic instability. BBF cycles contribute to deletions, gene amplification, nonreciprocal translocation, and overall genetic changes that are associated with tumorigenesis (56–63).
Our study showed that c-Myc is one key factor that initiates genomic instability through BBF cycles. Such BBF cycles in telomerase-positive immortalized mouse PreB cells (unpublished data) with long telomeres are distinct from BBF cycles reported for critically short telomeres (61, 64). Some TAs (but not necessarily all) represent fusions, as evident by the analysis of metaphase chromosomes. TAs and end-to-end fusions depended on time and levels of c-Myc activation. Analysis of frequencies of both events showed that they are closely linked. As the fusions initiate BBF cycles, the frequencies of breakage and nonreciprocal translocations increase over time.
A previously uncharacterized pathway of c-Myc-dependent genomic instability thus starts at the telomeric ends of the chromosomes. Both TAs and BBF cycles are the manifestation of deregulated Myc expression, leading to chromosomal rearrangements and subsequently to genomic instability.
Local chromosome movement increases chromosomal overlap in the nucleus. This temporal change in local positioning may permit the direct contact of chromosomal ends and facilitate recombinations and/or fusions. Such movements were observed after c-Myc deregulation and suggested an impact of the oncoprotein on local nuclear positioning of chromosomes. Chromosome movements were previously studied and found by others as well (65–69).
Several regulatory pathways involving oncogene deregulation may affect the 3D nuclear organization. Oncoproteins, including c-Myc, can alter the 3D nuclear organization and the organization of chromatin (70–72). They also affect the nuclear matrix. High mobility group protein I(Y) (HMGI(Y)) is a c-Myc-dependent nuclear matrix protein (73) with increased expression during neoplasia (2). The analysis of myc-binding sites in the human genome suggests that c-Myc binds to genes encoding nucleoskeletal components (74). Furthermore, constitutive c-Myc expression was shown to be associated with down-regulation of the telomere repeat binding protein TRF2 (10), a protein required for telomere capping and genome stability (75). Myc is also involved in the regulation of DNA repair (22, 23) and has been shown to induce DNA breakage (21). Thus taken together, many different c-Myc-dependent mechanisms could potentially affect the nuclear organization and, as shown here, converge at the telomeres.
Supplementary Material
Acknowledgments
We thank Dr. Michael Mowat for critical reading of this manuscript, Mary Cheang for statistical analyses, and Cheryl Taylor-Kashton and Landon Wark for deconvolution. The work was supported by the Canada Foundation for Innovation, the Canadian Institutes of Health Research, CancerCare Manitoba (S.M.), Fondation de France (Paris) and French Minister of Foreign Affairs (T.F.), Sander-Stiftung, and Deutsche Krebshilfe (P.B.), the Physics for Technology program of the Foundation for Fundamental Research in Matter, the Delft Inter-Faculty Research Center Life Tech, Cyttron, and the Delft Research program Life Science and Technology (Delft, The Netherlands).
Author contributions: S. Mai designed research; S.F.L., A.G., Z.L., F.K., T.C.Y.C., S. Moshir, V.M., and S. Mai performed research; B.J.V., Y.G., and I.T.Y. contributed new reagents/analytic tools; S.F.L., B.J.V., Y.G., A.G., Z.L., F.K., T.C.Y.C., S. Moshir, V.M., A.Y.C.C., P.D.K., T.F., P.B., and S. Mai analyzed data; and Y.G. and S. Mai wrote the paper.
This paper was submitted directly (Track II) to the PNAS office.
Abbreviations: 4HT, 4-hydroxytamoxifen; SKY, spectral karyotyping; TA, telomeric aggregate.
References
- 1.Pienta, K. J., Partin, A. W. & Coffey, D. S. (1989) Cancer Res. 49, 2525-2532. [PubMed] [Google Scholar]
- 2.Leman, E. S., Madigan, M. C., Brunagel, G., Takaha, N., Coffey, D. S. & Getzenberg, R. H. (2003) J. Cell Biochem. 88, 599-608. [DOI] [PubMed] [Google Scholar]
- 3.Zink, D., Fischer, A. H. & Nickerson, J. A. (2004) Nat. Rev. Cancer 4, 677-687. [DOI] [PubMed] [Google Scholar]
- 4.Parada, L. A., McQueen, P. G., Munson, P. J. & Misteli, T. (2002) Curr. Biol. 12, 1692-1697. [DOI] [PubMed] [Google Scholar]
- 5.Parada, L. & Misteli, T. (2002) Trends Cell Biol. 12, 425-432. [DOI] [PubMed] [Google Scholar]
- 6.Roix, J. J., McQueen, P. G., Munson, P. J., Parada, L. A. & Misteli, T. (2003) Nat. Genet. 34, 287-291. [DOI] [PubMed] [Google Scholar]
- 7.Parada, L. A., McQueen, P. G. & Misteli, T. (2004) Genome Biol. 5, R44. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Neves, H., Ramos, C., da Silva, M. G., Parreira, A. & Parreira, L. (1999) Blood 93, 1197-1207. [PubMed] [Google Scholar]
- 9.Chuang, T. C., Moshir, S., Garini, Y., Chuang, A. Y., Young, I. T., Vermolen, B., van den Doel, R., Mougey, V., Perrin, M., Braun, M., et al. (2004) BMC Biol. 2, 12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Ermler, S., Krunic, D., Knoch, T. A., Moshir, S., Mai, S., Greulich-Bode, K. M. & Boukamp, P. (2004) Eur. J. Cell Biol. 83, 681-690. [DOI] [PubMed] [Google Scholar]
- 11.Nesbit, C. E., Tersak, J. M. & Prochownik, E. V. (1999) Oncogene 18, 3004-3016. [DOI] [PubMed] [Google Scholar]
- 12.Potter, M. & Marcu, K. B. (1997) Curr. Top. Microbiol. Immunol. 224, 1-17. [DOI] [PubMed] [Google Scholar]
- 13.Mai, S. & Mushinski, J. F. (2003) J. Environ. Pathol. Toxicol. Oncol. 22, 179-199. [DOI] [PubMed] [Google Scholar]
- 14.Mai, S. (1994) Gene 148, 253-260. [DOI] [PubMed] [Google Scholar]
- 15.Mai, S., Hanley-Hyde, J. & Fluri, M. (1996) Oncogene 12, 277-288. [PubMed] [Google Scholar]
- 16.Mai, S., Fluri, M., Siwarski, D. & Huppi, K. (1996) Chromosome Res. 4, 365-371. [DOI] [PubMed] [Google Scholar]
- 17.Felsher, D. W. & Bishop, J. M. (1999) Proc. Natl. Acad. Sci. USA 96, 3940-3944. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Rockwood, L. D., Torrey, T. A., Kim, J. S., Coleman, A. E., Kovalchuk, A. L., Xiang, S., Ried, T., Morse, H. C., 3rd, & Janz, S. (2002) Oncogene 21, 7235-7240. [DOI] [PubMed] [Google Scholar]
- 19.Kuschak, T. I., Kuschak, B. C., Taylor, C. L., Wright, J. A., Wiener, F. & Mai, S. (2002) Oncogene 21, 909-920. [DOI] [PubMed] [Google Scholar]
- 20.Louis, S. F., Gruhne, B. & Mai, S. (2005) Prog. Oncog., in press.
- 21.Vafa, O., Wade, M., Kern, S., Beeche, M., Pandita, T. K., Hampton, G. M. & Wahl, G. M. (2002) Mol. Cell 9, 1031-1044. [DOI] [PubMed] [Google Scholar]
- 22.Hironaka, K., Factor, V. M., Calvisi, D. F., Conner, E. A. & Thorgeirsson, S. S. (2003) Lab. Invest. 83, 643-654. [DOI] [PubMed] [Google Scholar]
- 23.Karlsson, A., Deb-Basu, D., Cherry, A., Turner, S., Ford, J. & Felsher, D. W. (2003) Proc. Natl. Acad. Sci. USA 100, 9974-9979. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Partlin, M. M., Homer, E., Robinson, H., McCormick, C. J., Crouch, D. H., Durant, S. T., Matheson, E. C., Hall, A. G., Gillespie, D. A. & Brown, R. (2003) Oncogene 22, 819-825. [DOI] [PubMed] [Google Scholar]
- 25.Chiang, Y. C., Teng, S. C., Su, Y. N., Hsieh, F. J. & Wu, K. J. (2003) J. Biol. Chem. 278, 19286-19291. [DOI] [PubMed] [Google Scholar]
- 26.Adams, J. M., Harris, A. W., Pinkert, C. A., Corcoran, L. M., Alexander, W. S., Cory, S., Palmiter, R. D. & Brinster, R. L. (1985) Nature 318, 533-538. [DOI] [PubMed] [Google Scholar]
- 27.Potter, M. & Wiener, F. (1992) Carcinogenesis 13, 1681-1697. [DOI] [PubMed] [Google Scholar]
- 28.Pelengaris, S., Khan, M. & Evan, G. I. (2002) Cell 109, 321-334. [DOI] [PubMed] [Google Scholar]
- 29.Felsher, D. W. & Bishop, J. M. (1999) Mol. Cell 4, 199-207. [DOI] [PubMed] [Google Scholar]
- 30.Marinkovic, D., Marinkovic, T., Mahr, B., Hess, J. & Wirth, T. (2004) Int. J. Cancer 110, 336-342. [DOI] [PubMed] [Google Scholar]
- 31.D'Cruz, C. M., Gunther, E. J., Boxer, R. B., Hartman, J. L., Sintasath, L., Moody, S. E., Cox, J. D., Ha, S. I., Belka, G. K., Golant, A., Cardiff, R. D. & Chodosh, L. A. (2001) Nat. Med. 7, 235-239. [DOI] [PubMed] [Google Scholar]
- 32.Jain, M., Arvanitis, C., Chu, K., Dewey, W., Leonhardt, E., Trinh, M., Sundberg, C. D., Bishop, J. M. & Felsher, D. W. (2002) Science 297, 102-104. [DOI] [PubMed] [Google Scholar]
- 33.Karlsson, A., Giuriato, S., Tang, F., Fung-Weier, J., Levan, G. & Felsher, D. W. (2003) Blood 101, 2797-2803. [DOI] [PubMed] [Google Scholar]
- 34.Shachaf, C. M., Kopelman, A. M., Arvanitis, C., Karlsson, A., Beer, S., Mandl, S., Bachmann, M. H., Borowsky, A. D., Ruebner, B., Cardiff, R. D., et al. (2004) Nature 431, 1112-1117. [DOI] [PubMed] [Google Scholar]
- 35.Fest, T., Mougey, V., Dalstein, V., Hagerty, M., Milette, D., Silva, S. & Mai, S. (2002) Oncogene 21, 2981-2990. [DOI] [PubMed] [Google Scholar]
- 36.Mai, S., Hanley-Hyde, J., Rainey, G. J., Kuschak, T. I., Paul, J. T., Littlewood, T. D., Mischak, H., Stevens, L. M., Henderson, D. W. & Mushinski, J. F. (1999) Neoplasia 1, 241-252. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Wiener, F., Kuschak, T. I., Ohno, S. & Mai, S. (1999) Proc. Natl. Acad. Sci. USA 96, 13967-13972. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Wiener, F., Coleman, A., Mock, B. A. & Potter, M. (1995) Cancer Res. 55, 1181-1188. [PubMed] [Google Scholar]
- 39.Littlewood, T. D., Hancock, D. C., Danielian, P. S., Parker, M. G. & Evan, G. I. (1995) Nucleic Acids Res. 23, 1686-1690. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Grenman, S., Shapira, A. & Carey, T. E. (1988) Gynecol. Oncol. 30, 228-238. [DOI] [PubMed] [Google Scholar]
- 41.Grenman, S. E., Roberts, J. A., England, B. G., Gronroos, M. & Carey, T. E. (1988) Gynecol. Oncol. 30, 239-250. [DOI] [PubMed] [Google Scholar]
- 42.Mandlekar, S., Hebbar, V., Christov, K. & Kong, A. N. (2000) Cancer Res. 60, 6601-6606. [PubMed] [Google Scholar]
- 43.Fukasawa, K., Wiener, F., Vande Woude, G. F. & Mai, S. (1997) Oncogene 15, 1295-1302. [DOI] [PubMed] [Google Scholar]
- 44.Figueroa, R., Lindenmaier, H., Hergenhahn, M., Nielsen, K. V. & Boukamp, P. (2000) Cancer Res. 60, 2770-2774. [PubMed] [Google Scholar]
- 45.Schaefer, L. H., Schuster, D. & Herz, H. (2001) J. Microsc. 204, 99-107. [DOI] [PubMed] [Google Scholar]
- 46.Vermolen, B. J., Garini, Y., Mai, S., Mougey, V., Fest, T., Chunag, T. C. Y., Chuang, A. Y. C., Wark, L. & Young, I. T. (2005) Cytometry, in press. [DOI] [PubMed]
- 47.Poon, S. S., Martens, U. M., Ward, R. K. & Lansdorp, P. M. (1999) Cytometry 36, 267-278. [DOI] [PubMed] [Google Scholar]
- 48.Beatty, B., Mai, S. & Squire, J., eds. (2002) FISH: A Practical Approach (Oxford Univ. Press, Oxford).
- 49.Benedek, K., Chudoba, I., Klein, G., Wiener, F. & Mai, S. (2004) Chromosome Res. 12, 777-785. [DOI] [PubMed] [Google Scholar]
- 50.Smith, G., Taylor-Kashton, C., Dushnicky, L., Symons, S., Wright, J. & Mai, S. (2003) Neoplasia (Bratisl.) 5, 110-120. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Wilson, E. B. (1927) J. Am. Stat. Assoc. 22, 209-212. [Google Scholar]
- 52.Mueller, H. J. (1938) Collecting Net. 13, 181-198. [Google Scholar]
- 53.McClintock, B. (1941) Genetics 26, 234-282. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Fest, T., Guffei, A., Williams, G., Silva, S. & Mai, S. (2005) Oncogene 24, 2944-2953. [DOI] [PubMed] [Google Scholar]
- 55.Molenaar, C., Wiesmeijer, K., Verwoerd, N. P., Khazen, S., Eils, R., Tanke, H. J. & Dirks, R. W. (2003) EMBO J. 22, 6631-6641. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.DePinho, R. A. & Polyak, K. (2004) Nat. Genet. 36, 932-934. [DOI] [PubMed] [Google Scholar]
- 57.Artandi, S. E., Chang, S., Lee, S. L., Alson, S., Gottlieb, G. J., Chin, L. & DePinho, R. A. (2000) Nature 406, 641-645. [DOI] [PubMed] [Google Scholar]
- 58.Artandi, S. E. (2002) Trends Mol. Med. 8, 44-47. [DOI] [PubMed] [Google Scholar]
- 59.Smith, K. A., Stark, M. B., Gorman, P. A. & Stark, G. R. (1992) Proc. Natl. Acad. Sci. USA 89, 5427-5431. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Ciullo, M., Debily, M. A., Rozier, L., Autiero, M., Billault, A., Mayau, V., El Marhomy, S., Guardiola, J., Bernheim, A., Coullin, P., et al. (2002) Hum. Mol. Genet. 11, 2887-2894. [DOI] [PubMed] [Google Scholar]
- 61.Hande, M. P., Samper, E., Lansdorp, P. & Blasco, M. A. (1999) J. Cell Biol. 144, 589-601. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Murnane, J. P. & Sabatier, L. (2004) Bioessays 26, 1164-1174. [DOI] [PubMed] [Google Scholar]
- 63.Gisselsson, D., Jonson, T., Petersen, A., Strombeck, B., Dal Cin, P., Hoglund, M., Mitelman, F., Mertens, F. & Mandahl, N. (2001) Proc. Natl. Acad. Sci. USA 98, 12683-12688. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Londono-Vallejo, J. A. (2004) Cancer Lett. 212, 135-144. [DOI] [PubMed] [Google Scholar]
- 65.Zink, D. & Cremer, T. (1998) Curr. Biol. 8, R321-R324. [DOI] [PubMed] [Google Scholar]
- 66.Walter, J., Schermelleh, L., Cremer, M., Tashiro, S. & Cremer, T. (2003) J. Cell Biol. 160, 685-697. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Vourc'h, C., Taruscio, D., Boyle, A. L. & Ward, D. C. (1993) Exp. Cell Res. 205, 142-151. [DOI] [PubMed] [Google Scholar]
- 68.Ferguson, M. & Ward, D. C. (1992) Chromosoma 101, 557-565. [DOI] [PubMed] [Google Scholar]
- 69.Bridger, J. M., Boyle, S., Kill, I. R. & Bickmore, W. A. (2000) Curr. Biol. 10, 149-152. [DOI] [PubMed] [Google Scholar]
- 70.Fischer, A. H., Bond, J. A., Taysavang, P., Battles, O. E. & Wynford-Thomas, D. (1998) Am. J. Pathol. 153, 1443-1450. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Fischer, A. H., Chadee, D. N., Wright, J. A., Gansler, T. S. & Davie, J. R. (1998) J. Cell Biochem. 70, 130-140. [PubMed] [Google Scholar]
- 72.Chadee, D. N., Hendzel, M. J., Tylipski, C. P., Allis, C. D., Bazett-Jones, D. P., Wright, J. A. & Davie, J. R. (1999) J. Biol. Chem. 274, 24914-24920. [DOI] [PubMed] [Google Scholar]
- 73.Takaha, N., Hawkins, A. L., Griffin, C. A., Isaacs, W. B. & Coffey, D. S. (2002) Cancer Res. 62, 647-651. [PubMed] [Google Scholar]
- 74.Fernandez, P. C., Frank, S. R., Wang, L., Schroeder, M., Liu, S., Greene, J., Cocito, A. & Amati, B. (2003) Genes Dev. 17, 1115-1129. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.van Steensel, B., Smogorzewska, A. & de Lange, T. (1998) Cell 92, 401-413. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.