Abstract
Nucleic acid conjugation methodologies involve linking the nucleic acid sequence to other (bio)molecules covalently. This typically allows for nucleic acid property enhancement whether it be for therapeutic purposes, biosensing, etc. Here, we report a streamlined, aqueous compatible, on‐column conjugation methodology using nucleic acids containing a site‐specific amino‐modifier. Both monophosphates and carboxylates were amenable to the conjugation strategy, allowing for the introduction of a variety of useful handles including azide, aryl, and hydrophobic groups in DNA. We find that an on‐column approach is superior to post‐synthetic template‐directed synthesis, mainly with respect to product purification and recovery.
Keywords: Nucleic acid conjugation, Phosphoramidate linkage, Amide linkage, Solid-support conjugation
We report an aqueous compatible conjugation methodology for nucleic acids that feature amino‐modifiers, enabling the synthesis of phosphoramidate‐ or amide‐containing oligonucleotide conjugates. Various ligand handles, including those containing azides and metal chelators, are suitable for the conjugation strategy. Our approach offers a streamlined, efficient and robust strategy for ligand conjugation to nucleic acid strands in mild aqueous conditions.
Introduction
Nucleic acid conjugation is an attractive avenue for imparting additional chemical or biological properties to synthetic oligonucleotides.[ 1 , 2 ] Although canonical nucleic acid production is desirable for many applications, including synthetic PCR primers, there exist many applications that benefit from ligand conjugation. Oligonucleotide therapeutics, such as siRNA and antisense oligonucleotides, stand out as a significant category of nucleic acids modified with ligands, especially in light of the recent siRNA‐conjugate approvals by the FDA and other regulatory bodies. [3]
Solid‐phase synthesis is a versatile pathway for the introduction of an armamentarium of chemical modifications within the nucleic acid scaffold (both internal and terminal positions). Numerous examples of chemical modifications have been reported, many of which are now commercially available as phosphoramidite building blocks. Importantly, the chemical modifications need not necessarily resemble the chemical structure of canonical nucleic acid units. Conversely, conjugation techniques can be exploited when the ligand is incompatible with standard synthetic strategies involved in the chemical preparation of nucleic acids. Conjugation strategies may also be utilized for late‐stage modification of substrates (e. g. post‐synthesis of the nucleic acid).
There exists a variety of conjugation methodologies reported in the literature. For instance, the thiol‐maleimide strategy is widely utilized for producing (bio)conjugates between a compound containing a thiol moiety and another containing a maleimide. [4] This conjugation is particularly popular in preparing antibody‐drug candidates, and has found applications in generating nucleic acid conjugates. One drawback is the reversibility of the covalent linkage via a retro 1,4‐addition. [4] The copper‐catalyzed cycloaddition of an azide‐containing compound with an alkyne‐containing one to generate a 1,2,3‐triazole linkages demonstrates another avenue to generate conjugates.[ 5 , 6 ] This so‐called “click” chemistry has been explored in many different contexts, including oligonucleotides. [7] One difficulty with this strategy is installing azides in oligonucleotides, as these can undergo reduction to the corresponding amines in the presence of phosphines (P(III)), commonly known as the Staudinger reduction reaction. [8] As a result, most have resorted in installing the alkyne moiety in the oligonucleotide, instead of the azide. This is not to say that azides have not been incorporated directly in nucleic acids.[ 9 , 10 , 11 ] Recently reviewed by the Müller group, common strategies include functionalizing an azide‐containing unit to the solid support prior to solid‐phase synthesis of the oligonucleotide. [12] Alternatively, the azide may be installed post‐synthetically by converting the free 5′‐OH into a leaving group and performing a substitution reaction using NaN3. [13] Additionally, the use of a bromohexylene phosphoramidite can be employed, and further functionalized using NaN3.[ 14 , 15 ] Nucleobase modifications, such as 6‐chloro‐9‐(β‐D‐ribofuranosyl)‐purines, can also be exploited to introduce azide functionality. [16] The Dahma group explored the use of a reverse synthesis methodology, followed by H‐phosphonate chemistry to ultimately introduce a 3′‐azidothymidyl unit at the 3′‐end of DNA. [17] Aigner et al. used a 2′‐azido‐modified nucleoside containing a 2‐chlorophenyl protected 3′‐monophosphate as a reagent to incorporate within RNA strands. [18] All of these pathways generate the canonical phosphodiester linkage, while requiring fairly anhydrous conditions.
The Richert group, on the other hand, made use of a terminal amino group (either 5′ or 3′) to couple with an oxyazabenzotriazole‐activated amino‐protected mononucleotide. [19] This synthetic strategy enabled the preparation of P3′→N5′, or the N3′→P5′ phosphoramidate‐containing DNA. Reactions were performed in a template‐directed fashion using a primer strand immobilized to a solid‐support, and required a deblock step after each unit addition. Interestingly, it is unknown, to the extent of our knowledge, whether the azidomethyloxycarbonyl (Azoc) protecting group is capable of cycloaddition with an alkyne‐containing compound. Although the azide was part of a protecting group handle that gets removed, we asked whether a similar post‐synthetic approach could be a viable option for the incorporation of other azide‐containing nucleotide units. In addition, phosphoramidate linkages are particularly interesting as they are more sensitive to acid‐mediated hydrolysis relative to canonical phosphodiester bonds. As such, the introduction of site‐specific phosphoramidate inserts may be utilized to cleave oligomers at discrete locations.
Taken together, we sought to develop a simple and efficient methodology to prepare nucleic acid conjugates containing a variety of useful handles, using nucleic acid amino‐modifiers that are readily available. Our initial strategy involved using a DNA, or RNA, primer containing a 3′‐amino terminal to perform nonenzymatic RNA template‐directed synthesis (TDS). We opted to use the 3′‐azido‐3′‐deoxythymidine (AZT) as a cheap and readily accessible source of an azide‐containing compound. TDS reactions were performed using a 2‐methylimidazole activated AZT mononucleotide, or using the unactivated AZT 5′‐O‐monophosphate in the presence of 1‐ethyl‐3‐(3‐dimethylaminopropyl)carbodiimide (EDC) as a condensation reagent, to generate a N3′→P5′‐phosphoramidate product containing a 3′‐terminal azido functionality. The untemplated conjugation was also possible albeit with slower kinetics and lower yields. We then asked whether an on‐column approach could be exploited, given the ease in scaling and purification (Figure 1). Our post‐synthetic on‐column approach enables a versatile methodology to incorporate various handles such as azido, aryl, hydrophobic and chelating functionality compatible with aqueous conditions. We were able to employ monophosphate‐ and carboxylate‐containing ligands to form phosphoramidate and amide linkages, respectively. Substrates were deprotected using standard ammonium hydroxide treatment, followed by conventional purification/isolation techniques. The development of an efficient post‐synthetic on‐column conjugation approach using mild aqueous conditions is reported. Ultimately, we focused on using commercially available building blocks, or easily synthesized precursors, to increase the accessibility of this methodology.
Figure 1.
Aqueous compatible on‐column functionalization to prepare oligonucleotides containing a phosphoramidate or amide linkages.
Results and Discussion
Synthesis of 3′‐Azido Precursors
3′‐Azido‐3′‐deoxythymidine (AZT, Zidovudine), used as an antiretroviral medication for the treatment of HIV/AIDS, is a readily and commercially available azide‐containing nucleoside for use in the laboratory. [20] Many other azide‐containing nucleosides are commercial, but tend to typically be expensive. The incorporation of azido functionality in nucleic acids can be cumbersome, as preparing the conventional phosphoramidite building block in the presence of an azide group is difficult, albeit not impossible.[ 9 , 21 ] We attempted to prepare the 5′‐phosphoramidite of AZT with no success, presumably due to the production of the iminophosphorane intermediate via the P(III)‐mediated reduction of the azide group (data not shown). Next, we attempted to prepare the H‐phosphonate derivative, but recoveries were sluggish in our hands. Capitalizing on the enzymatic‐mediated incorporation of the AZT triphosphate was another possible avenue, as shown previously.[ 22 , 23 ] However, we feared that this approach could be challenging for larger scales. As a result, we opted to explore an alternative fully chemical methodology, which involved P(V) intermediates in the form of a simple monophosphate.
The synthesis of the AZT 5′‐phosphoroimidazolide was accomplished in two steps (Scheme 1). First, the 5′‐O‐monophosphate of AZT (compound 1) was accessible via a straightforward phosphorylation using excess POCl3 in pyridine at 0 °C. Although the classical method reported by Yoshikawa is suitable for this step, we opted for this alternative procedure given the lack of a 3′‐hydroxyl moiety and exocylic amino group in AZT. [24] Purification was readily achievable by automated reverse‐phase flash chromatography, using triethylammonium bicarbonate buffer (solvent A) and acetonitrile (solvent B), resulting in the triethylammonium salt version of monophosphate 1. [25] The desire to retain the azido group precluded us from using the conventional Ph3P‐mediated dehydrative redox reaction between monophosphate 1 and 2‐methylimidazole to prepare phosphoroimidazolide 2. [8] We noted that 1‐ethyl‐3‐(3‐dimethylaminopropyl)carbodiimide (EDC) had been previously utilized to activate 5′‐O‐monophosphates, and sought to use this procedure to produce our desired phosphoroimidazolide 2.[ 26 , 27 , 28 , 29 ] Fortunately, this pathway produced the desired mononucleotide activated with 2‐methylimidazole (compound 2, see Supplementary Information for synthetic details). In 2017, the Szostak group reported 2‐aminoimidazole as a superior leaving group to 2‐methylimidazole for template‐directed synthesis of RNA on RNA templates. Unfortunately, we were unable to efficiently produce the 2‐aminoimidazole‐activated monomer using an EDC‐mediated reaction. To increase the generalizability of our methodology, we also prepared the 5‐methylcytidine mononucleotide 4 using the well‐established convertible nucleotide strategy, followed by Yoshikawa‐type phosphorylation.[ 30 , 31 ] The 5′‐phosphoroimidazolide was not prepared for reasons that are described below.
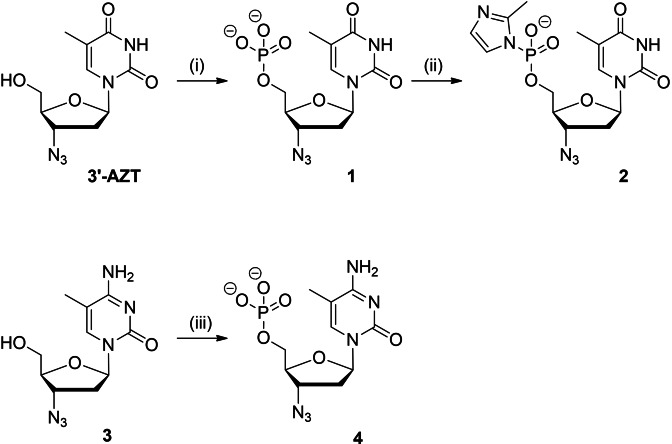
Scheme 1.
Synthesis of activated 5′‐phosphoroimidazolide‐3′‐azido‐3′‐deoxythymidine 2 and −3′‐azido‐2′, 3′‐dideoxy‐5‐methylcytidine 5′‐O‐monophosphate 4. Reagents and conditions: (i) POCl3 (4.0 eq.), Py, 0 °C, 10 min; (ii) 2‐methylimidazole (4 eq.), EDC⋅HCl (5 eq.), NEt3 (10 eq.) DMF, 22 °C, 24 h; (iii) POCl3 (4.0 eq.), NEt3 (2.0 eq.), trimethylphosphate, 0 °C, 2 h.
Template‐Directed Synthesis of DNA and RNA Containing a 3′‐Terminal Azido Group
With the 2‐methylimidazole activated monomer 2 in hand, we asked whether we could produce nucleic acids containing a 3′‐azido‐3′‐deoxythymidine unit using TDS. Chemical TDS is advantageous as it provides a fully post‐synthetic strategy without requiring enzymes. A FAM‐labeled DNA (or RNA) primer strand containing a 3′‐amino‐3′‐deoxy group was prepared (primerNH2), using previously reported procedures, and hybridized to an RNA template. [32] We opted for the 3′‐amino terminated primers as these are superior to those containing a 3′‐hydroxyl in TDS reactions, which is particularly useful to improve the reaction rates of thymidinyl activated mononucleotides.[ 33 , 34 , 35 , 36 ] The TDS reaction of a DNA primer on an RNA template was initiated by the addition of monomer 2 (Figure 2), and showed virtually full conversion to the +1 product by PAGE in 24 hours (see Supplementary Figure S19–21 for other conditions tested). As expected, switching the DNA primer to an RNA primer increased the reaction kinetics, with almost full conversion of the +1 product within 3 h at 5 mM Mg2+ and full conversion within 1 h with 100 mM Mg2+ (Supplementary Figures S22 and S23). The presence of a 3′‐azide group causes chain termination in the primer extension, as observed with enzymatic incorporation of AZT, preventing over‐extension often observed with amino systems.[ 33 , 35 , 36 ] The kinetics of the reaction were slower than observed previously for similar activate mononucleotide substrates containing a 3′‐amino, instead of a 3′‐azido, and may be due to a number of reasons.[ 35 , 36 ] First, primers were terminated with a 3′‐amino‐2′,3′‐dideoxythymidyl unit, which presumably reduces +1 extension efficiency relative to cytidyl and guanosyl terminals in TDS (see Supplementary Figure S24 for ddG terminated RNA). [35] To test this hypothesis, the RNA primer sequence version was synthesized with a 3′‐amino‐2′,3′‐dideoxyguanidyl terminal, and demonstrated improved product yield relative to the DNA or RNA sequence terminated with 3′‐amino‐2′,3′‐dideoxythymidyl residue. Second, previous work has shown that activated ribonucleotides predominantly switch to the C3′‐endo A‐form conformation upon binding to an RNA template, from a mixture C2′‐endo and C3′‐endo conformation, for productive primer extension. [37] AZT in solution has almost equal conformation populations, which could impact the kinetics. [38]
Figure 2.
Template‐directed synthesis (TDS) reaction of a DNA primer on an RNA template using (A) 2‐methylimidazole activated monomer 2. (i) Reaction conditions were DNA primer (1 μM), RNA template (2.5 μM), MgCl2 (5 mM), monomer (40 mM), HEPES buffer (200 mM, pH 8.0). PAGE analysis illustrating reaction time‐course of (ii) templated and (iii) untemplated reaction. (B) EDC‐mediated condensation reaction using monophosphate 1. (i) Reaction conditions were DNA primer (5 μM), RNA template (10 μM), MgCl2 (20 mM), 5′‐O‐monophosphate 1 (100 mM), EDC (400 mM) for 8 hours in HEPES buffer (200 mM, pH 8.0). (ii) PAGE analysis of a control experiment showcasing the effect of removing the template (−T), monomer (−M), magnesium (−Mg2+), and EDC (−EDC). “C” is for the primer‐strand only control. (iii) Same conditions as (ii) but at 3 °C instead of 22 °C.
Next, we investigated whether the incorporation of the desired azido modification was possible without the preparation of the phosphoroimidazolide, as these reactants have lower shelf stability [35] compared to identical compounds containing solely the monophosphate group (e. g. compound 1 vs. compound 2, Scheme 1). We opted to use EDC as the condensing reagent given our success in preparing phosphoroimidazolide 2, and widespread utility for aqueous‐based conjugation. In addition, others have activated mononucleotides with imidazoles using EDC in aqueous conditions, [28] and EDC‐mediated template‐directed ligation has been demonstrated previously.[ 39 , 40 , 41 ] We thus reacted a DNA primerNH2‐ RNA template duplex with monophosphate 1 in the presence of EDC (Figure 2) at two temperatures (3 °C and 22 °C). Surprisingly, the templated and untemplated reactions yielded significant product and by‐product formation, respectively, within 8 h. A control experiment demonstrated that both the monomer and EDC were required for conjugation. Removal of the template caused significant reduction in yield, albeit reaching 25 % product within 8 h, with 20 % primer remaining. Lowering the temperature seemingly generated less by‐products for the 8 h mark, particularly for the untemplated reaction.
Satisfied with these conversions, we investigated the possibility of increasing the scale of the reaction and isolating the target product strand. Unfortunately, the presence of high amounts of salt from the reaction medium (e. g. MgCl2, Na+‐HEPES buffer, EDC⋅HCl and mononucleotides) prevented the development of a robust and efficient pathway of recovering the desired product by SAX‐HPLC. Including a precipitation step in the process resulted in reduced recoveries. In addition, we noted the presence of side‐products for the EDC‐mediated reaction, particularly at longer reaction times, as documented by others previously. [41] As a result, we did not explore alternative purification techniques such as PAGE and size‐exclusion chromatography.
On‐Column Post‐Synthetic Conjugation of DNA
These limitations prompted us to explore an alternative strategy. Recognizing that solid‐phase synthesis offers a myriad of advantages, including ease of purification and retention of the nucleobases/phosphodiester backbone protecting groups, we sought to explore this avenue for the incorporation of azide modifications, and perhaps other useful chemical handles. We first synthesized the commercially available 5′‐(4‐methoxytritylamino)‐5′‐deoxythymidine‐3′‐O‐[(2‐cyanoethyl)‐(N,N‐diisopropyl)]‐phosphoramidite, according to reported procedures, [42] and incorporated this modification at the 5′‐end of a model DNA sequence (Figure 3). Unmasking the 5′‐amino group required prolonged treatment with 3 % TCA in DCM (twice the amount suggested by the standard manufacturer protocol for canonical 5′‐O‐(4,4′‐dimethyoxytrityl)‐protected amidites ensured full deprotection). The support‐bound DNA strand was then treated with monophosphate 1 (Scheme 1, see Supplementary Information for synthetic details) and EDC in water containing 50 % v/v DMF for 8 hours. The DMF was introduced to aid in solubilizing the support‐bound oligomer containing protecting groups. Standard deprotection conditions, followed by analysis SAX‐HPLC revealed the appearance of a new product peak in low‐to‐moderate yield (Supplementary Figure S43). As expected, characterization by LR‐MS was consistent with our expected product. Introduction of aqueous HEPES buffer pH 8.0 and N‐methylimidazole (N‐MeIm) in the reaction significantly improved the conversion to the product (see Supplementary Figure S48 and S49). The conversion was also affected by choice of organic solvent, with better conversions observed when using 25 % MeCN instead of 50 % DMF (Supplementary Figure S49 and S50). Using these optimized conditions, we were able to couple both monophosphates 1 and 6 efficiently (Figure 3, see Supplementary Scheme S1 for synthetic details of compound 6).
Figure 3.
On‐column EDC‐mediated conjugation using a 5′‐amino DNA construct. (A) (i) Reaction scheme of the on‐column amino‐modifier conjugation strategy using a 5′‐O‐monophosphate to generate a P5′→N5′ phosphoramidate linkage. (ii) SAX‐HPLC trace of (a) the crude starting material strand containing a 5′‐amino terminal and (b) the product strand containing the azido functional group. Values indicate retention times (min). (iii) Low‐resolution electrospray ionization mass spectrum (negative mode) of the purified product strand showcasing m/z signals in agreement with the expected m/z. (B) (i) The reaction scheme of the on‐column amino‐modifier conjugation strategy using a 3′‐O‐monophosphate generating a P3′→N5′ phosphoramidate linkage. (ii) SAX‐HPLC trace of (a) the crude starting material strand containing a 5′‐amino terminal and (b) the product strand containing the azido functional group. Values indicate retention times (min). (iii) Low‐resolution mass spectrum of product strand showcasing m/z signals in agreement with the expected m/z. The solid‐support was treated with 100 mM monophosphate, 100 mM N‐MeIm, 800 mM EDC in 250 mM HEPES pH 8.0 containing 25 % MeCN (v/v) and allowed to react for 8 h. The solid support was treated a second time with a freshly prepared coupling solution, after thorough washing of the solid‐support bound oligonucleotide.
Content with the product formation, we employed protected nucleoside 5′‐O‐[(2‐cyanoethyl)‐(N,N‐diisopropyl)]‐phosphoramidites (so‐called reverse‐amidites) to produce a support‐bound DNA containing a 3′‐amino terminal (Figure 4). The detritylated DNA was reacted with monophosphate 1, EDC and N‐MeIm in 250 mM HEPES pH 8.0 buffer containing 25 % (v/v) MeCN for 8 h. Unexpectedly, we observed lower conversions for the 3′‐amino terminal relative to a similar reaction with the 5′‐amino. Two 8 h conjugation treatments were performed to optimize the conversion, and we were pleased to observe significantly higher product formation (Figure 4). All subsequent conjugations were performed with two coupling steps (fresh coupling solution added after 8 h) to increase robustness of the technique and ensure optimal product formation. Shorter reaction times may result in similar conjugation conversions, but this is most‐likely substrate dependent. Changing the sequence to include a fluorescein tag and cytidyl units provided similar conjugation efficacy (Supplementary Figure S37). A conjugation procedure was repeated with the cytidyl analogue 4 (Figure 1), as opposed to monophosphate 1, and performed adequately (Supplementary Figure S58).
Figure 4.
On‐column EDC‐mediated conjugation using a 3′‐amino terminal oligonucleotide. (i) Reaction scheme of the on‐column amino‐modifier conjugation strategy using a 5′‐O‐monophosphate to generate a N3′→P5′ phosphoramidate linkage. (ii) SAX‐HPLC trace of (a) the crude starting material strand containing a 5′‐amino terminal and (b) the product strand containing the azido functional group. Values indicate retention times (min). (iii) Low‐resolution electrospray ionization mass spectrum (negative mode) of the purified product strand showcasing m/z signals in agreement with the expected m/z. On‐column EDC‐mediated conjugation using 3′‐amino DNA construct and 5′‐O‐monophosphate. The solid‐support was treated with a coupling solution of 100 mM monomer, 100 mM N‐MeIm, 800 mM EDC in 250 mM HEPES pH 8.0 containing 25 % MeCN (v/v) and allowed to react for 8 h. The solid support was treated a second time with a freshly prepared coupling solution, after thorough washing of the solid‐support bound oligonucleotide.
Fauster et al. developed an approach to incorporate 2′‐azido modifications in nucleic acids, by using an o‐chlorophenyl‐protected 3′‐O‐monophosphate derivative (see protected monophosphate 9 as an example, Supplementary Information) in presence of a powerful condensation reagent, 1‐(mesitylsulfonyl)‐3‐nitro‐1H‐1,2,4‐triazole (MSNT), under anhydrous conditions. [18] We asked whether we could generate a phosphoramidate linkage using similar conditions. We used compound 9 due to the ease of synthetic accessibility (see Supplementary Information for synthetic details). Somewhat surprisingly, low conversions to the desired product were observed when reacting a 3′‐amino terminated oligomer with protected monophosphate 9 using MSNT (Supplementary Figure S62). Perhaps the free amine was reacting directly with MSNT generating the corresponding sulfonamide derivative. Importantly, acceptable conversions were observed for conjugating protected monophosphate 9 with an oligomer containing a 3′‐OH terminal, instead of the 3′‐amino terminated sequence, as a positive control (Supplementary Figure S65). We then prepared the unprotected 3′‐O‐monophosphate (compound 8) version of what was used by Fauster et al. to have the ability of incorporating an azide functionality internally. Repeating our optimized procedure with the unprotected 3′‐O‐monophosphate 8 yielded good conversions to the azido‐containing oligomer using a 5′‐amino terminated DNA (Figure 5).
Figure 5.
On‐column EDC‐mediated conjugation using a 5′‐amino DNA construct. (i) Reaction scheme of the on‐column amino‐modifier conjugation strategy using a 3′‐O‐monophosphate nucleoside to generate a P3′→N5′ phosphoramidate linkage. (ii) SAX‐HPLC trace of (a) the crude starting material strand containing a 5′‐amino terminal and (b) the product strand containing the azido functional group. The synthesis retained the DMTr. Values indicate retention times (min). (iii) Low‐resolution electrospray ionization mass spectrum (negative mode) of the purified product strand showcasing m/z signals in agreement with the expected m/z. On‐column EDC‐mediated conjugation using 3′‐amino DNA construct and 3′‐O‐monophosphate. The solid‐support was treated with a coupling solution of 100 mM monophosphate, 100 mM N‐MeIm, 800 mM EDC in 250 mM HEPES pH 8.0 containing 25 % MeCN (v/v) and allowed to react for 8 h. The solid support was treated a second time with a freshly prepared coupling solution, after thorough washing of the solid‐support bound oligonucleotide.
Ribonucleoside 5′‐O‐monophosphates are readily and commercially available. We envisioned employing uridine 5′‐O‐monophosphate (UMP) as an economical agent for synthesizing branched oligonucleotides (Figure 6). As a model experiment, we conjugated the 3′‐amino terminated DNA with UMP and extended both the 2′‐ and 3′‐positions with regular 3′‐O‐phosphoramidites. Prolonged coupling times (10 min) with 100 mM amidite solutions were employed (as opposed to the standard 50 mM concentrations we typically use) to optimize the sterically encumbered branching. We were pleased to observe that the major product was in agreement with our desired product by MS. The single monophosphate coupling was also repeated using cytidine 5′‐O‐monophosphate (CMP) showcasing that the exocyclic protecting group need not be preinstalled for conjugation (Supplementary Figure S71).
Figure 6.
On‐column EDC‐mediated conjugation using 3′‐amino DNA construct and uridine 5′‐O‐monophosphate (UMP), follow by the simultaneous extension at both the 2′‐ and 3′‐OH. (i) Reaction scheme of the on‐column amino‐modifier conjugation strategy using a riboside 5′‐O‐monophosphate to generate a N3′→P5′ phosphoramidate linkage capable of subsequent branching. PG refers to the cyanoethyl group. (ii) SAX‐HPLC trace of (a) the crude starting material strand containing a 3′‐amino terminal, (b) the product strand containing the uridine unit, and (c) the product strand resulting for subsequent extension of two thymidinyl units at the 2′‐ and 3′‐positions. Values indicate retention times (min). (iii) Low‐resolution electrospray ionization mass spectrum (negative mode) of the purified product strands showcasing m/z signals in agreement with the expected m/z. (a) After the coupling of the UMP unit and (b) after the branch extension.
As this conjugation methodology was untemplated, we suspected that virtually any amino modifier could be used in lieu of the amino sugars employed above. To test this hypothesis, a commercially available MMTr‐protected hexanolamine phosphoramidite was constructed and incorporated at the 5′‐end of a model DNA strand. Conjugating monophosphate 1 produced a dangling azido‐containing nucleotide in acceptable yields (Figure 7).
Figure 7.
On‐column EDC‐mediated conjugation using a 5′‐hexylamino‐containing DNA construct with monophosphate 1. (i) Reaction scheme of the on‐column amino‐modifier conjugation strategy using a 5′‐O‐monophosphate nucleoside to generate a phosphoramidate linkage. (ii) SAX‐HPLC trace of (a) the crude starting material strand containing a 5′‐hexylamino terminal and (b) the product strand containing the azido functional group. Values indicate retention times (min). (iii) Low‐resolution electrospray ionization mass spectrum (negative mode) of the purified product strand showcasing m/z signals in agreement with the expected m/z. The solid‐support was treated with a coupling solution of 100 mM monophosphate, 100 mM N‐MeIm, 800 mM EDC in 250 mM HEPES pH 8.0 containing 25 % MeCN (v/v) and allowed to react for 8 h. The solid support was treated a second time with a freshly prepared coupling solution, after thorough washing of the solid‐support bound oligonucleotide.
With a robust aqueous‐based conjugation methodology in hand, we tested whether we could conjugate carboxylates in the same manner as monophosphates, resulting in an amide linkage instead of a phosphoramidate. Many similar strategies have been reported[ 1 , 42 , 43 , 44 , 45 , 46 , 47 , 48 ], but seldomly under aqueous conditions using an on‐column approach, to the extent of our knowledge. Using our previously optimized conditions from above, we treated a 5′‐amino modified DNA with potassium benzoate (Figure 8). Unfortunately, conjugation was unsatisfactory. Replacing the 25 % (v/v) MeCN with 50 % (v/v) DMF resulted in significantly improved coupling, furnishing a major product with slightly higher retention time via SAX‐HPLC. The product strand MS analysis aligned with our calculated values. We then successfully conjugated a para‐substituted bromobenzoic acid group with a 5′‐amino containing DNA (see Supplementary Figure S81). This example could serve as a useful cross‐coupling partner, as previously demonstrated for other aryl bromides.[ 49 , 50 , 51 ] As many previous reports have focused on the conjugation of peptides to nucleic acids, we tested an Fmoc‐protected glycine as an amino acid example, which showed excellent conjugation after standard deprotection conditions (Figure 8 and Supplementary Figure S83). Next, we tested oleic acid as a hydrophobic handle using a 3′‐amino containing DNA (Figure 8). Realizing that oleic acid is capable of forming higher order structures in HEPES pH 8.0, it was crucial to maintain a high ratio of organic solvent content as to inhibit the formation of micelles/vesicles. We were pleased to observe successful conjugation, albeit with slightly lower conversions (Figure 8). Nevertheless, product purification was straightforward, and we are currently working with such conjugates to localize nucleic acid substrates at the surface of membrane bilayers. Recently, the Desaulnier group reported an elegant preparation of folate‐labeled siRNAs, using a folate derivative phosphoramidite, to enhance targeted delivery to folate receptor‐expressing cancer cell lines. [52] We wondered whether we could apply our post‐synthetic conjugation strategy to pteroic acid, a folate derivative. Initial conjugation using the 5′‐amino‐modified oligonucleotide was suboptimal. Once more, we suspected that poor solubility of the precursor caused inefficient conjugation. Using 50 % (v/v) DMSO overcame this hurdle and provided the desired product in satisfactory yield (see Supplementary Figure S87).
Figure 8.
On‐column EDC‐mediated conjugation of amino‐containing DNA and carboxylates. (A) (i) Reaction scheme of the on‐column 5′‐amino‐modified DNA conjugation strategy using commercially available carboxylate handles such as potassium benzoate and Fmoc‐protected glycine (Fmoc‐Gly‐OH), generating an amide linkage. (ii) SAX‐HPLC trace of (a) the crude starting material strand containing a 5′‐amino terminal, (b) the crude reaction from the conjugation with potassium benzoate, and (c) the crude reaction from the conjugation with Fmoc‐Gly‐OH. Values indicate retention times (min). (iii) Low‐resolution mass spectrum of the purified product strands showcasing m/z signals in agreement with the expected m/z for (a) conjugation with potassium benzoate and (b) conjugation with Fmoc‐Gly‐OH. (B) Same as (A) but using a 3′‐amino modified DNA conjugated with oleic acid. Optimal conditions were as follows: 100 mM carboxylate, 100 mM N‐MeIm, 800 mM EDC in 100 mM HEPES pH 8.0 containing 50 % DMF (v/v). Treatment was repeated with fresh coupling solution after 8 h, after thorough washing of the solid‐support bound oligonucleotide.
As a final example, we aimed to conjugate 2,2′,2′′,2′′′‐(1,4,7,10‐tetraazacyclododecane‐1,4,7,10‐tetrayl)tetraacetic acid (DOTA) to amino‐modified oligomers. DOTA, and DOTA‐like derivatives, are of particular interest as they can chelate trivalent metallic isotopes with high affinity, typically used for imaging purposes.[ 53 , 54 ] Research groups have developed robust pathways to incorporate DOTA (and DOTA‐like molecules) in nucleic acid and nucleic acid‐like structures.[ 55 , 56 , 57 , 58 , 59 , 60 ] These tend to be typically laborious and/or costly to conjugate this desirable moiety. We first used a t‐butyl protected version of DOTA containing a single free carboxylate for controlled conjugation, which performed adequately under our reaction conditions (Figure 9). Importantly, the deprotection conditions were carried out in a 2 M NaOH: methanol mixture (1 : 4, v/v) at 65 °C for 10 hours prior to neutralization and purification via SAX‐HPLC. Additionally, we repeated the conjugation using a mixed sequence and observed similar conversions as illustrated in Figure 9 (Supplementary Figure S91). The unprotected version of DOTA (a more‐affordable alternative) was also subject to conjugation, and we were pleased to observe moderate‐to‐high product formation, with the appearance of an additional byproduct (relative to the t‐butyl protected DOTA precursor). Purification of the DOTA‐containing product was readily achievable by SAX‐HPLC (Supplementary Figure S93).
Figure 9.
On‐column EDC‐mediated conjugation of 5′‐amino‐modified DNA and DOTA. (A) (i) Reaction scheme of the on‐column conjugation strategy using commercially available protected DOTA, generating an amide linkage. (ii) SAX‐HPLC trace of (a) the crude starting material strand containing a 5′‐amino terminal, (b) the crude reaction from the conjugation reaction with DOTA. (iii) Low‐resolution electrospray ionization mass spectrum (negative mode) of the purified product strands showcasing m/z signals in agreement with the expected m/z for DOTA conjugate. Optimal conditions were as follows: 100 mM carboxylate, 100 mM N‐MeIm, 800 mM EDC in 100 mM HEPES pH 8.0 containing 50 % DMF (v/v). Treatment was repeated with fresh coupling solution after 8 h, after thorough washing of the solid‐support bound oligonucleotide. Note that the expected m/z includes a potassium adduct (as previously observed. [48]
An interesting feature of the phosphoramidate linkage is their enhanced susceptibility to acid‐mediated hydrolysis, relative to canonical phosphodiester bonds. [61] These could be theoretically utilized as a means to cleave the newly formed conjugate on‐demand. We are currently investigating the application of nucleic acid conjugates containing one, or multiple, acid‐labile cleavage sites for material science purposes and within modular nanostructures.
On‐Column Post‐Synthetic Conjugation of RNA
We asked whether our on‐column conjugation methodology was applicable to RNA. As a preliminary model example, we prepared an RNA sequence containing a 5′‐amino‐5′‐deoxythymidyl unit (5′‐ NH2dT r(UUU AAA AAA AA)). We initially tested the conjugation with either monophosphate 6 or potassium benzoate. After the conjugation reaction, the solid support was treated with a solution of 4 : 1 (v/v) ammonium hydroxide : ethanol at 55 °C for 17 h, followed by concentration to dryness in a DNA concentrator. The desilylation was carried out in standard conditions (3 : 1 (v/v) NEt3⋅3HF:DMSO at 65 °C for 2 h), followed by a precipitation step using cold n‐butanol. To our surprise, only the amide coupling supplied the desired conjugate as the major product (Supplementary Figure S96). The phosphoramidate conjugate reaction seemingly contained starting material oligomer and a major unknown (by)‐product with slightly longer retention time by SAX‐HPLC.
To test whether the monophosphate conjugation reaction was influenced by the presence of the 2′‐O‐TBDMS groups, and/or the workflow associated with the desilylation, we prepared a sequence using 2′‐O‐Me phosphoramidites, instead of 2′‐O‐TBDMS protected phosphoramidites. Conjugating CMP with the 2′‐O‐Me‐containing sequence (Supplementary Figure S100), followed by the standard DNA workflow (described above) revealed good conversions to the desired phosphoramidate‐containing conjugate. Amide coupling using potassium benzoate was also successful (Supplementary Figure S102). Current efforts are geared towards making the conjugation methodology effective on canonical RNA sequences. Perhaps, as observed previously, the desilylation conditions caused phosphoramidate linkage degradation. [62] Using desilylation conditions compatible with phosphoramidate linkages did not, however, mitigate the issues (data not shown). It may be that the support‐bound silyl‐protected RNA is simply insoluble in the tested solvent systems, and warrants further investigation.
Conclusions
We report a streamlined on‐column conjugation methodology using nucleic acids containing site‐specific amino‐modifiers. Both monophosphates and carboxylates were amenable to the conjugation strategy in DNA, allowing for the introduction of a variety of useful handles including azide, aryl, and hydrophobic groups. Importantly, easily synthesized compounds and/or commercially available starting materials were used to make this strategy more‐appealing to groups lacking organic chemistry expertise/infrastructure. In our limited search, it was easier to locate commercial sources of useful carboxylate‐containing compounds than those containing monophosphates. Our on‐column approach offers a mild aqueous‐compatible means of conjugation, using the readily available coupling reagent EDC. Most of the reaction reagents are conveniently removed by washing the solid‐support containing the covalently bound oligomer of interest. As anticipated, we observed that the yield of conjugation is influenced by solvent conditions, presumably because the chemical handle must be soluble for optimal reactivity. Standard deprotection protocols for nucleic acids are used on the conjugate. Of course, a notable limitation of this technique is that the conjugate handle must withstand the conditions of oligomer cleavage and deprotection.
Conflict of Interests
The authors declare no conflict of interest.
1.
Supporting information
As a service to our authors and readers, this journal provides supporting information supplied by the authors. Such materials are peer reviewed and may be re‐organized for online delivery, but are not copy‐edited or typeset. Technical support issues arising from supporting information (other than missing files) should be addressed to the authors.
Supporting Information
Saraya J. S., Sammons S. R., O'Flaherty D. K., ChemBioChem 2025, 26, e202400643. 10.1002/cbic.202400643
Data Availability Statement
The data that support the findings of this study are available from the corresponding author upon reasonable request.
References
- 1. Klabenkova K., Fokina A., Stetsenko D., Molecules 2021, 26, 5420. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Lönnberg H., Bioconjugate Chem. 2009, 20, 1065–1094. [DOI] [PubMed] [Google Scholar]
- 3. Egli M., Manoharan M., Nucleic Acids Res. 2023, 51, 2529–2573. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Renault K., Fredy J. W., Renard P. Y., Sabot C., Bioconjugate Chem. 2018, 29, 2497–2513. [DOI] [PubMed] [Google Scholar]
- 5. Rostovtsev V. V., Green L. G., Fokin V. V., Sharpless K. B., Angew. Chem. Int. Ed. 2002, 41, 2596–2599. [DOI] [PubMed] [Google Scholar]
- 6. Tornøe C. W., Christensen C., Meldal M., J. Org. Chem. 2002, 67, 3057–3064. [DOI] [PubMed] [Google Scholar]
- 7. Fantoni N. Z., El-Sagheer A. H., Brown T., Chem. Rev. 2021, 121, 7122–7154. [DOI] [PubMed] [Google Scholar]
- 8. Leffler J. E., Temple R. D., J. Am. Chem. Soc. 1967, 89, 5235–5246. [Google Scholar]
- 9. Fomich M. A., Kvach M. V., Navakouski M. J., Weise C., Baranovsky A. V., Korshun V. A., Shmanai V. V., Org. Lett. 2014, 16, 4590–4593. [DOI] [PubMed] [Google Scholar]
- 10. Pourceau G., Meyer A., Vasseur J. J., Morvan F., J. Org. Chem. 2009, 74, 6837–6842. [DOI] [PubMed] [Google Scholar]
- 11. Santner T., Hartl M., Bister K., Micura R., Bioconjugate Chem. 2014, 25, 188–195. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Müggenburg F., Müller S., Chem. Rec. 2022, 22, e202100322. [DOI] [PubMed] [Google Scholar]
- 13. Miller G. P., Kool E. T., J. Org. Chem. 2004, 69, 2404–2410. [DOI] [PubMed] [Google Scholar]
- 14. Lietard J., Meyer A., Vasseur J. J., Morvan F., Tetrahedron Lett. 2007, 48, 8795–8798. [Google Scholar]
- 15. Lietard J., Meyer A., Vasseur J. J., Morvan F., J. Org. Chem. 2008, 73, 191–200. [DOI] [PubMed] [Google Scholar]
- 16. Nainar S., Beasley S., Fazio M., Kubota M., Dai N., Corrêa I. R., Spitale R. C., ChemBioChem 2016, 17, 2149–2152. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Yamada K., Wahba A. S., Bernatchez J. A., Ilina T., Martínez-Montero S., Habibian M., Deleavey G. F., Götte M., Parniak M. A., Damha M. J., ACS Chem. Biol. 2015, 10, 2024–2033. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Aigner M., Hartl M., Fauster K., Steger J., Bister K., Micura R., ChemBioChem 2011, 12, 47–51. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Kaiser A., Spies S., Lommel T., Richert C., Angew. Chem. Int. Ed. 2012, 51, 8299–8303. [DOI] [PubMed] [Google Scholar]
- 20. Bozzi A., D'Andrea G., Brisdelli F., Curr. Clin. Pharmacol. 2008, 3, 20–37. [DOI] [PubMed] [Google Scholar]
- 21. Lingala S., Nordstrøm L. U., Mallikaratchy P. R., Tetrahedron Lett. 2019, 60, 211–213. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Sluis-Cremer N., Sheen C. W., Zelina S., Torres P. S. A., Parikh U. M., Mellors J. W., Antimicrob. Agents Chemother. 2007, 51, 48–53. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Michailidis E., Marchand B., Kodama E. N., Singh K., Matsuoka M., Kirby K. A., Ryan E. M., Sawani A. M., Nagy E., Ashida N., Mitsuya H., Parniak M. A., Sarafianos S. G., J. Biol. Chem. 2009, 284, 35681–35691. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Yoshikawa M., Kato T., Takenishi T., Bull. Chem. Soc. Jpn. 1969, 42, 3505–3508. [Google Scholar]
- 25. Li L., Prywes N., Tam C. P., O'Flaherty D. K., Lelyveld V. S., Izgu E. C., Pal A., Szostak J. W., J. Am. Chem. Soc. 2017, 139, 1810–1813. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Shabarova Z. A., Biochimie 1988, 70, 1323–1334. [DOI] [PubMed] [Google Scholar]
- 27. Chu B. C. F., Wahl G. M., Orgel L. E., Nucleic Acids Res. 1983, 11, 6513–6529. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Li L., Prywes N., Tam C. P., O'Flaherty D. K., Lelyveld V. S., Izgu E. C., Pal A., Szostak J. W., J. Am. Chem. Soc. 2017, 139, 1810–1813. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29. Taylor P., Ivanovskaya M. G., Gottikh M. B., Shabarova Z. A., Nucleosides Nucleotides 1987, 6, 913–934. [Google Scholar]
- 30. Gogoi K., Gunjal A. D., Phalgune U. D., Kumar V. A., Org. Lett. 2007, 9, 2697–2700. [DOI] [PubMed] [Google Scholar]
- 31. Xu Y.-Z., Zheng Q., Swann P., J. Org. Chem. 1992, 57, 3839–3845. [Google Scholar]
- 32. Eisenhuth R., Richert C., J. Org. Chem. 2009, 74, 26–37. [DOI] [PubMed] [Google Scholar]
- 33. O'Flaherty D. K., Zhou L., Szostak J. W., J. Am. Chem. Soc. 2019, 141, 10481–10488. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Zhou L., O'Flaherty D. K., Szostak J. W., Angew. Chem. Int. Ed. 2020, 59, 15682–15687. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Zhang S., Zhang N., Blain J. C., Szostak J. W., J. Am. Chem. Soc. 2013, 135, 924–932. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Zhang S., Blain J. C., Zielinska D., Gryaznov S. M., Szostak J. W., Proc. Natl. Acad. Sci. USA 2013, 110, 17732–17737. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. Zhang N., Zhang S., Szostak J. W., J. Am. Chem. Soc. 2012, 134, 3691–3694. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Jagannadh B., Reddy D. V., Kunwar A. C., Biochem. Biophys. Res. Commun. 1991, 179, 386–391. [DOI] [PubMed] [Google Scholar]
- 39. Zielinski W. S., Orgel L. E., Nature 1987, 327, 346–347. [DOI] [PubMed] [Google Scholar]
- 40. von Kiedrowsk G., Angew. Chem. Int. Ed. Engl. 1986, 98, 932–935. [Google Scholar]
- 41. Obianyor C., Newnam G., Clifton B. E., Grover M. A., Hud N. V., ChemBioChem 2020, 21, 3359–3370. [DOI] [PubMed] [Google Scholar]
- 42. Tetzlaff C. N., Schwope I., Bleczinski C. F., Steinberg J. A., Richert C., Tetrahedron Lett. 1998, 39, 4215–4218. [Google Scholar]
- 43. Gras M., Adler P., Smietana M., Chem. A Eur. J. 2024, 30, e202401069. [DOI] [PubMed] [Google Scholar]
- 44. Wiener J., Kokotek D., Rosowski S., Lickert H., Meier M., Sci. Rep. 2020, 10, 1457. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45. Basu S., Wickstrom E., Tetrahedron Lett. 1995, 36, 4943–4946. [Google Scholar]
- 46. Honcharenko D., Druceikaite K., Honcharenko M., Bollmark M., Tedebark U., Strömberg R., ACS Omega 2021, 6, 579–593. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47. McMinn D. L., Greenberg M. M., J. Am. Chem. Soc. 1998, 120, 3289–3294. [Google Scholar]
- 48. Aaronson J. G., Klein L. J., Momose A. A., O'Brien A. M., Shaw A. W., Tucker T. J., Yuan Y., Tellers D. M., Bioconjugate Chem. 2011, 22, 1723–1728. [DOI] [PubMed] [Google Scholar]
- 49. Omumi A., Beach D. G., Baker M., Gabryelski W., Manderville R. A., J. Am. Chem. Soc. 2011, 133, 42–50. [DOI] [PubMed] [Google Scholar]
- 50. Li J.-Y., Huang H., Bioconjugate Chem. 2018, 29, 3841–3846. [DOI] [PubMed] [Google Scholar]
- 51. Ding Y., Clark M. A., ACS Comb. Sci. 2015, 17, 1–4. [DOI] [PubMed] [Google Scholar]
- 52. Salim L., Desaulniers J. P., Org. Biomol. Chem. 2023, 21, 3365–3372. [DOI] [PubMed] [Google Scholar]
- 53. Sosabowski J. K., Mather S. J., Nat. Protoc. 2006, 1, 972–976. [DOI] [PubMed] [Google Scholar]
- 54. Baranyai Z., Tircsó G., Rösch F., Eur. J. Inorg. Chem. 2020, 2020, 36–56. [Google Scholar]
- 55. Wang X., Milne M., Martínez F., Scholl T. J., Hudson R. H. E., RSC Adv. 2017, 7, 45222–45226. [Google Scholar]
- 56. Hovinen J., Nucleosides Nucleotides Nucleic Acids 2007, 26, 1459–1462. [DOI] [PubMed] [Google Scholar]
- 57. Cai J., Shapiro E. M., Hamilton A. D., Bioconjugate Chem. 2009, 20, 205–208. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58. Jaakkola L., Ylikoski A., Hovinen J., Bioconjugate Chem. 2006, 17, 1105–1107. [DOI] [PubMed] [Google Scholar]
- 59. Peuralahti J., Jaakkola L., Mukkala V. M., Hovinen J., Bioconjugate Chem. 2006, 17, 855–859. [DOI] [PubMed] [Google Scholar]
- 60. Kiviniemi A., Mäkelä J., Mäkilä J., Saanijoki T., Liljenbäck H., Poijärvi-Virta P., Lönnberg H., Laitala-Leinonen T., Roivainen A., Virta P., Bioconjugate Chem. 2012, 23, 1981–1988. [DOI] [PubMed] [Google Scholar]
- 61. Pourshahian S., Gryaznov S. M., Anal. Chem. 2019, 91, 11154–11161. [DOI] [PubMed] [Google Scholar]
- 62. Tago N., Katolik A., Clark N. E., Montemayor E. J., Seio K., Sekine M., Hart P. J., Damha M. J., J. Org. Chem. 2015, 80, 10108–10118. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
As a service to our authors and readers, this journal provides supporting information supplied by the authors. Such materials are peer reviewed and may be re‐organized for online delivery, but are not copy‐edited or typeset. Technical support issues arising from supporting information (other than missing files) should be addressed to the authors.
Supporting Information
Data Availability Statement
The data that support the findings of this study are available from the corresponding author upon reasonable request.