Abstract
The occurrence and progression of breast cancer (BCa) are complex processes involving multiple factors and multiple steps. The tumor microenvironment (TME) plays an important role in this process, but the functions of immune components and stromal components in the TME require further elucidation. In this study, we obtained the RNA-seq data of 1086 patients from The Cancer Genome Atlas (TCGA) database. We calculated the proportions of tumor-infiltrating immune cells (TICs) and immune and stromal components using the CIBERSORT and ESTIMATE methods, and we screened differentially expressed genes (DEGs). Univariate Cox regression analysis of overall survival was performed on the DEGs, and a protein–protein interaction network of their protein products was generated. Finally, the hub gene CD5 was obtained. High CD5 expression was found to be associated with longer survival than low expression. Gene set enrichment analysis showed that DEGs upregulated in the high-CD5 expression group were mainly enriched in tumor- and immune-related pathways, while those upregulated in the low-expression group were enriched in protein export and lipid synthesis. TIC analysis showed that CD5 expression was positively correlated with the infiltration of CD8+ T cells, activated memory CD4+ T cells, gamma delta T cells, and M1 macrophages and negatively correlated with the infiltration of M2 macrophages. CD5 can increase anticancer immune cell infiltration and reduce M2 macrophage infiltration. These results suggest that CD5 is likely a potential prognostic biomarker and therapeutic target, providing novel insights into the treatment and prognostic assessment of BCa.
Keywords: Breast cancer, CD5, Tumor microenvironment, Tumor immune infiltrates
Introduction
Breast cancer (BCa) is one of the most common malignant tumors and the leading cause of death among women. Cancer statistics from the U.S. show 279,100 new BCa cases and 42,690 deaths documented in 2020 [1]. Surgery, chemotherapy, and radiotherapy are currently the main treatment methods for BCa. Although there have been continuous improvements in diagnostic and treatment techniques that have significantly prolonged survival, there is still a risk of disease progression, indicated by relapse, metastasis, or drug resistance. Early diagnosis and exploration of new treatment methods facilitate improvement in the prognosis and survival of BCa patients. Therefore, more effective biomarkers must be identified to guide the diagnosis, treatment, and prognosis of BCa [2].
The tumor microenvironment (TME) is the internal environment surrounding a tumor and is an important factor affecting tumor behavior. The TME not only contains tumor cells but also contains various infiltrating immune cells. Studies have shown that a variety of immune cells, including T cells, B cells, macrophages, and natural killer cells, are involved in the apoptosis, proliferation, invasion, and metastasis of tumor cells [3]. BCa is a type of tumor with infiltration of inflammatory cells, and its occurrence and progression are closely related to the immune cells present [4]. Therefore, the study of tumor-infiltrating immune cells (TICs) has led to breakthroughs in the current treatment of BCa. For example, infiltrating T cells activated in TME in BCa express high levels of programmed death-1 (PD-1); This led to sustained activation of the PD-1 pathway in TME and suppression of T cell function, preventing the immune system from killing tumor cells. Recent advances in immunotherapy include immune checkpoint inhibitors, and related drugs such as Pembrolizumab and Atezolizumab have been approved for certain types of triple-negative breast cancer (TNBC), often used in combination with chemotherapy to enhance anti-tumor response. Studies have shown that there is an immunosuppressive state in triple-negative breast cancer, and CD5 may affect the efficacy of PD-1 inhibitors by affecting T cell function. CD5 is expressed in T cells and is involved in regulating the immune response. Studies have found that CD5 may also be expressed in some breast cancer cells, which may be related to its immune escape mechanism [5–7]. Combining CD5 with other immune-related biomarkers (such as PD-L1, CTLA-4, etc.) allows for a more comprehensive assessment of the tumor immune microenvironment and identification of possible immune escape mechanisms. Therapeutic strategies targeting CD5 may play a role in future immunotherapies, especially in regulating T cell function and improving anti-tumor response.
In this study, we used The Cancer Genome Atlas (TCGA) database to explore the TME-associated immune gene CD5; this gene plays an important role in the regulation of TME effects and tumor immunity and can affect the survival of BCa patients. It is a member of the scavenger receptor cysteine-rich (SRCR) family. The main role of CD5 is to distinguish T cell and B cell (B1a) subpopulations as a marker of such subpopulations [8]. CD5 is a cell-surface glycoprotein with unknown functions. In addition to its expression in T cells and B cells, CD5 is expressed in natural killer cells, monocytes, and macrophages. Early studies on CD5 mainly focused on T cells and B cells because their expression of CD5 is associated with their antigen-specific receptor complexes [9]. Recently, a growing number of studies have shown that CD5 is associated with the occurrence and progression of a variety of tumors, including chronic lymphocytic leukemia, non-small-cell lung cancer (NSCLC), B-cell leukemia, melanoma, and thymic cancer [10–13]. Evidence indicates that CD5 overexpression is correlated with improved prognosis in patients with NSCLC and melanoma [14, 15]. However, the role of CD5 in the microenvironment of BCa remains unclear. Therefore, CD5 may be an unexplored BCa biomarker related to immune cell regulation.
Materials and methods
Data resources
The transcriptome RNA-seq data of BCa patients and corresponding clinical data were obtained from the TCGA database (https://portal.gdc.cancer.gov/), including 111 normal samples and 1053 tumor samples. The total number of cases after exclusion of cases with incomplete information was 1086 (retrieved on December 24, 2020). The breast cancer tissue specimens used in this study were taken from fresh tissue collected during surgery at the Breast Disease Treatment Center of the Affiliated Hospital of Qinghai University between May 2021 and December 2021, and were identified as breast cancer or normal para-cancerous tissue by pathologists after surgery. The study was submitted to and approved by the Ethics Committee of Qinghai University Hospital, and the included samples followed the guidelines of the Declaration of Helsinki. The data for the trial came from publicly available databases and informed consent was obtained for use. The human study cohort included 10 patients. All patients participating in this study provided written informed consent.
Calculation of the immune score and stromal score
The immune score and stromal score in the TME were analyzed using the “estimate” package in the R language (version 4.0.3, https://www.r-project.org/) [16]. We combined the immune score and stromal score with the follow-up information of the BCa patients to conduct survival analysis. P < 0.05 was considered to indicate statistical significance.
Screening of differentially expressed genes (DEGs)
Based on the median immune score and stromal score, tumor samples were divided into a high-expression group and a low-expression group. DEGs were screened using the “Limma” package in the R language. |log2FC|> 1 and a false discovery rate (FDR) < 0.05 were used as the cutoff criteria for DEGs [17].
Functional enrichment analysis of DEGs
Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analyses were performed on the DEGs using the “clusterProfiler”, “enrichplot”, and “ggplot2” packages in R language. Both P < 0.05 and q < 0.05 were needed to indicate a significantly enriched term [18].
Construction of a DEG heat map
A heat map of DEGs was constructed using the pheatmap package in the R language.
Construction of a protein–protein interaction (PPI) network
A PPI network of DEGs was constructed via the online STRING (https://www.string-db.org) analysis website. The top 20 genes in terms of the number of connection nodes were identified using the cytoHubba plugin in Cytoscape software.
Univariate Cox regression analysis
Univariate Cox regression analysis was used to explore the DEGs associated with the overall survival (OS) rate of BCa patients. P < 0.05 was considered to indicate statistical significance. The screened genes were intersected with the genes screened according to the PPI network, yielding four target genes, including CD5.
CD5 expression and survival analysis
We performed a pan-cancer analysis of CD5 expression using the DiffExp module in the Tumor Immune Estimation Resource (TIMER) database (https://cistrome.shinyapps.io/timer/). The Gene Expression Profiling Interactive Analysis (GEPIA) database (http://gepia.cancer-pku.cn/) was used to verify the expression status of CD5 in various cancers. The prognostic value of CD5 in various cancers was evaluated via the KM plotter database (http://kmplot.com/analysis/). The Kaplan–Meier method was used to analyze the relationship between CD5 expression and OS rate in BCa patients. P < 0.05 was used as the threshold for identifying a significant difference.
Analysis of CD5 expression and clinical case characteristics
The association between CD5 expression and clinical stage, molecular staging, and lymph node metastasis of breast cancer patients was analyzed using the Ualcan database (http://ualcan.path.uab.edu/).
CD5 promoter methylation analysis
Changes in CD5 promoter methylation levels in breast cancer patients were assessed using the Ualcan database (https://ualcan.path.uab.edu/). In addition, the association between CD5 promoter methylation levels and OS was assessed. P < 0.05 was considered statistically significant.
Immunohistochemistry
Immunohistochemical staining was performed as previously described, except that the primary antibody was used at a 1:1000 dilution and incubated for 24 h. The primary and secondary antibodies were purchased from Abcam (Cambridge, MA, USA).
Western blotting
Western blot analysis was performed as previously described. The primary and secondary antibodies were purchased from Abcam (Cambridge, MA, USA). The primary antibody was used at a 1:600 dilution and the secondary antibody was incubated for 1 h.
TIC profile
We used the CIBERSORT (https://cibersort.stanford.edu/) algorithm to calculate the correlations between CD5 expression and the expression abundance of 22 types of infiltrating immune cells in BCa samples. P < 0.05 was used as the screening threshold for the BCa samples. The screened samples were subjected to a final analysis to calculate the correlations between TICs and CD5 expression [19].
Results
Correlations between the immune and stromal scores and the survival of BCa patients
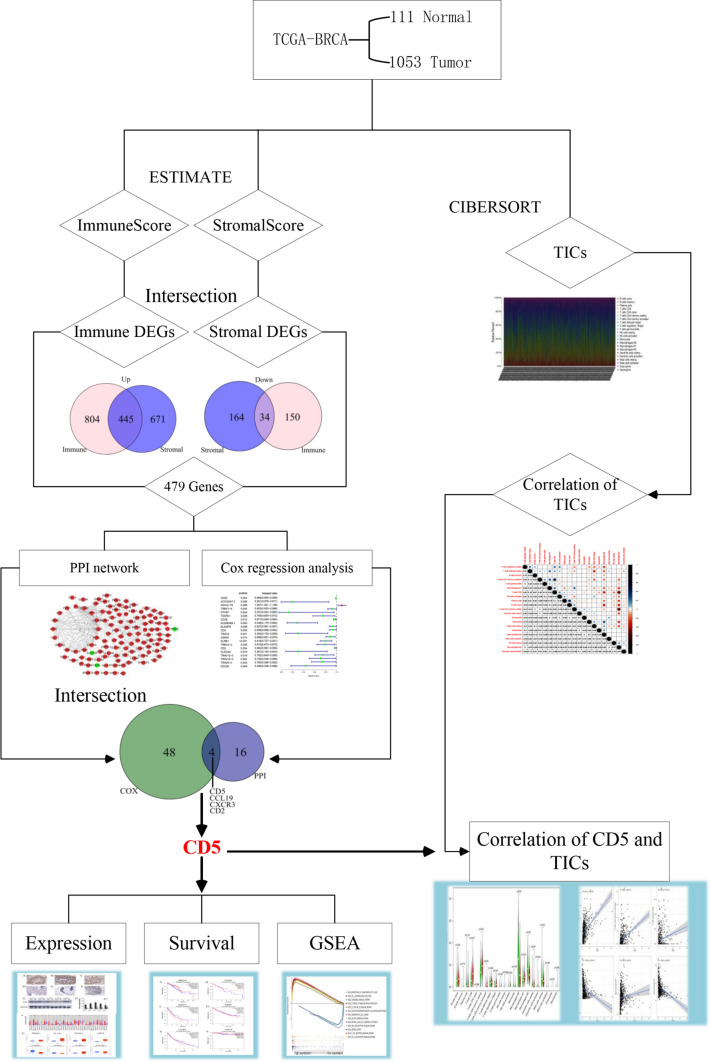
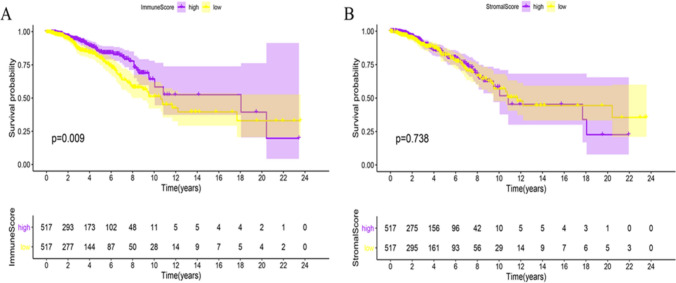
To assess the correlation of the immune score and the stromal score with the survival of BCa patients, Kaplan–Meier survival analysis was performed. A brief flowchart of our study is shown in (Fig. 1). As shown in (Fig. 2A, B), the immune score was significantly correlated with the OS of the patients (P = 0.009); the stromal score was not significantly correlated with the patients’ OS (P = 0.738). These results suggest that the immune score is more suitable for predicting the prognosis of BCa patients.
Fig. 1.
Analytical flowchart of this study
Fig. 2.
Relationships of the immune score and stromal score with the prognosis of BCa patients. A Immune score. A high immune score indicates a good prognosis. B Stromal score
Screening of DEGs
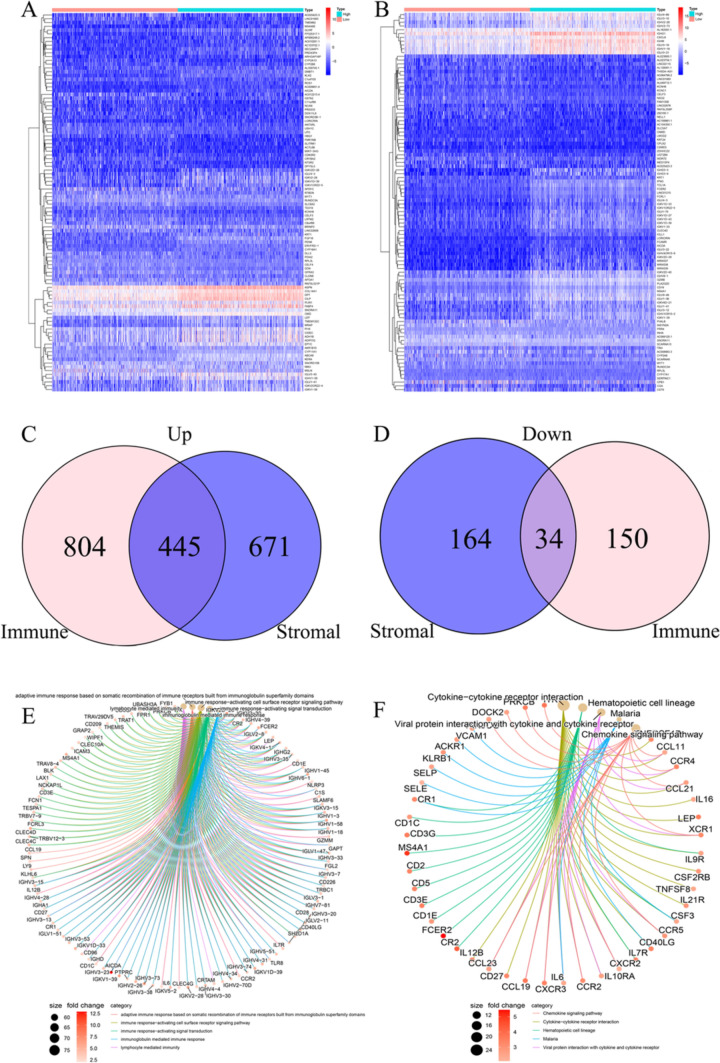
A total of 1433 DEGs were identified between immune score groups (high vs. low); 1249 were upregulated, while 184 were downregulated. A total of 1314 DEGs were identified between stromal score groups (high vs. low); 1116 were upregulated, while 198 were downregulated (Fig. 3A, B). Their intersection in a Venn diagram yielded 445 common upregulated genes and 34 common downregulated genes (Fig. 3C, D).
Fig. 3.
Heat map, Venn diagrams, and GO and KEGG for differentially expressed genes. A, B Heat maps of DEGs obtained for the immune score and stromal score, respectively. C Venn diagram of upregulated DEGs according to the immune score and the stromal score. D Venn diagram of downregulated DEGs according to the immune score and the stromal score. E, F GO and KEGG analyses of DEGs shared by the immune score and stromal score
Functional enrichment analysis of DEGs related to both immune score and stromal score
GO analysis showed that DEGs were most enriched in immune-related functions (Fig. 3E), including the immune response-activating cell surface receptor signaling pathway, the adaptive immune response based on somatic recombination of immune receptors built from immunoglobulin superfamily domains, lymphocyte-mediated immunity, the humoral immune response, B cell–mediated immunity and the humoral immune response were mediated by circulating immunoglobulin. Similarly, KEGG analysis showed that DEGs were mainly enriched in immune-related pathways, including cytokine-cytokine receptor interaction, chemokine signaling pathway, cell adhesion molecules, primary immunodeficiency, and hematopoietic cell lineage (Fig. 3F).
Identification of PPI network core genes and univariate Cox analysis
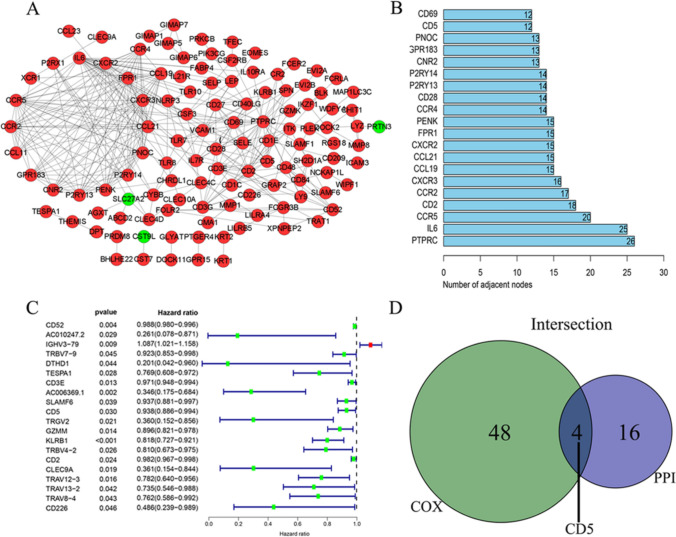
We constructed a PPI network based on the STRING data in the software Cytoscape and selected the 20 genes with the most connecting nodes as hub genes (Fig. 4A, B). Cox regression analysis showed that a total of 49 genes were closely related to the OS of BCa patients (Fig. 4C). The intersection of the core genes of the PPI network and the results of the Cox regression analysis yielded four critical hub genes, including CD5 (Fig. 4D).
Fig. 4.
Construction of the PPI network of DEGs and univariate Cox regression analysis. A Construction of the PPI network of DEGs. B Top 20 interacting genes in terms of the number of connection nodes in the PPI network. C Results of Cox regression analysis. D The intersection of the hub genes screened by the PPI network and the Cox analysis results. Finally, four hub genes, including CD5, were identified
Pan-cancer analysis of CD5 expression
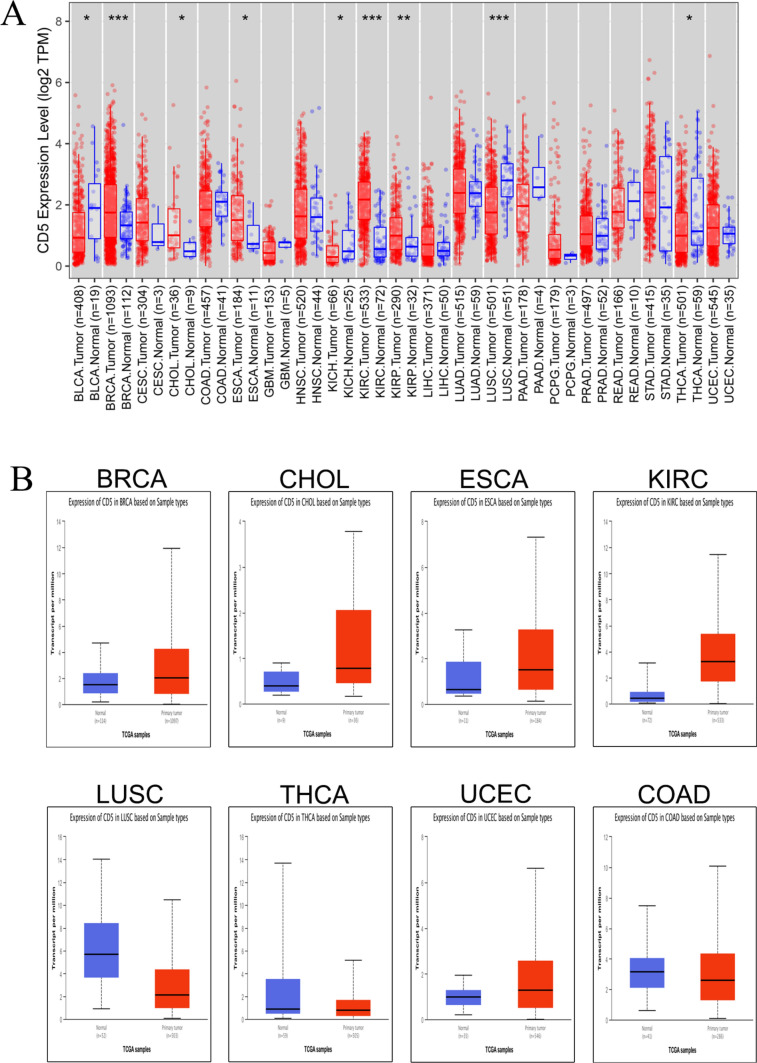
To explore the possible role of CD5 in cancer development, we looked at CD5 expression level in 21 cancers using the TIMER database. As shown in Fig. 5A, CD5 was differentially expressed in normal and cancerous tissues of nine cancers. CD5 expression was significantly increased in cancer tissues compared to normal tissues in five cancer types (Breast invasive carcinoma, Cholangiocarcinoma, Esophageal carcinoma, Kidney renal clear cell carcinoma and Kidney renal papillary cell carcinoma) and significantly decreased in four cancer types (Bladder Urothelial Carcinoma, Kidney Chromophobe, Lung squamous cell carcinoma and Thyroid carcinoma). In addition, we examined CD5 expression in 18 cancers using the GEPIA database and found that CD5 was differentially expressed in normal and cancerous tissues of nine cancers (Fig. 5B). Compared with normal tissues, CD5 expression was significantly higher in cancer tissues of Breast invasive carcinoma, Cholangiocarcinoma, Esophageal carcinoma, Kidney renal clear cell carcinoma, and Uterine Corpus Endometrial Carcinoma, but significantly lower in cancer tissues of Lung squamous cell carcinoma, Thyroid carcinoma, and Colon adenocarcinoma. Overall, our results indicate that CD5 is upregulated in Breast invasive carcinoma, Cholangiocarcinoma, Esophageal carcinoma, and Kidney renal clear cell carcinoma, but is downregulated in Lung squamous cell carcinoma and Thyroid carcinoma. Thus, CD5 likely plays an important role in these six cancers.
Fig. 5.
Pan-cancer analysis of CD5 expression. A Comparative analysis of CD5 gene expression levels in different cancer types. B Analysis of the prognostic value of CD5 gene in different cancer types
Prognostic value of CD5
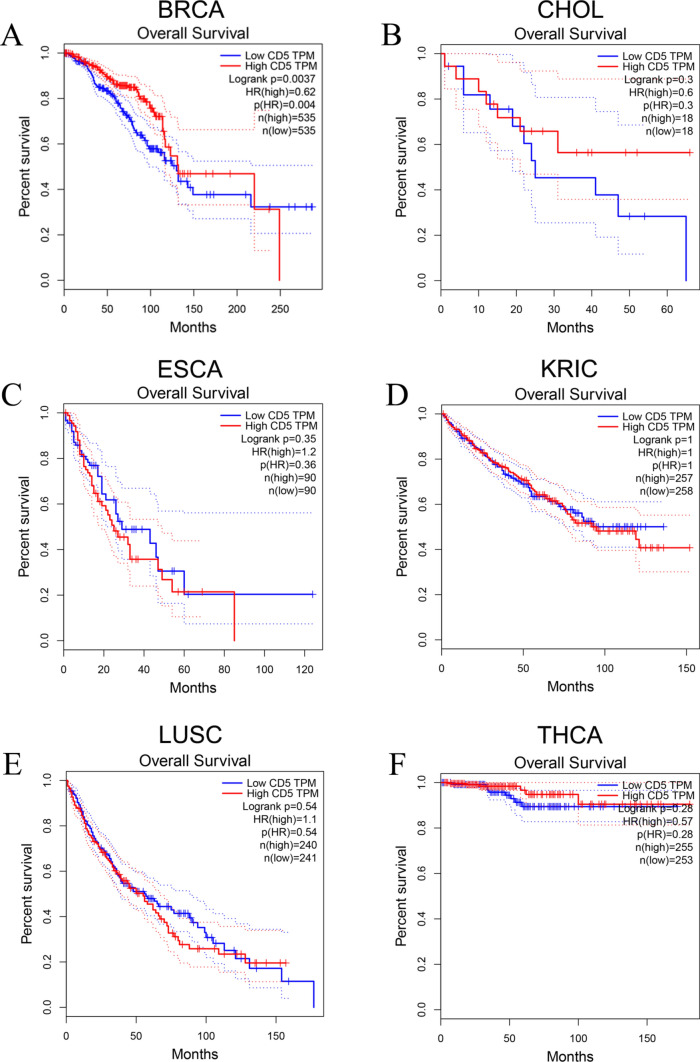
To examine the relationship between CD5 expression and disease prognosis, we performed survival analysis in context of CD5 expression in BRCA, CHOL, ESCA, KIRC, LUSC, and THCA. OS was used as the main prognostic indicator. High CD5 expression predicted a good prognosis for breast cancer patients (P < 0.05), but not for the other five cancers (Fig. 6A–F).
Fig. 6.
Correlation between overall survival and CD5 expression in cancer patients. A The survival differed significantly between the BRCA patients with high or low CD5 expression (P < 0.05). B–F Differential CD5 expression in patients with CHOL, ESCA, KRIC, LUSC and THCA was not significantly correlated with patient OS
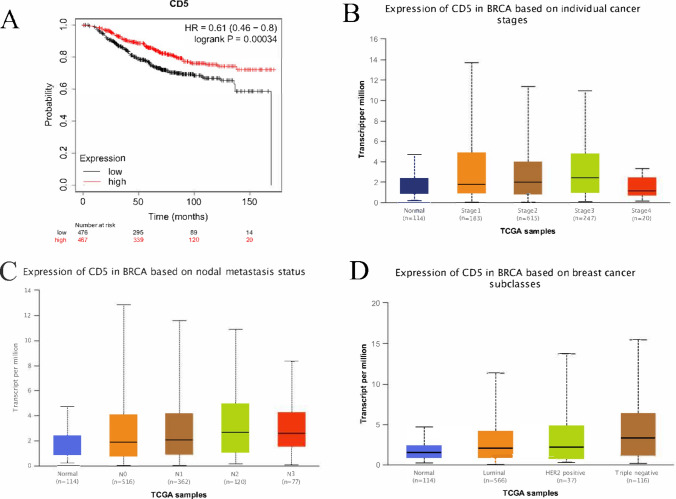
To validate these results, we repeated the survival analysis using an independent dataset from the GEO database (GSE1456 is a dataset of breast cancer). Our results showed that high CD5 expression was associated with improved patient OS (Fig. 7A). Furthermore, compared with normal breast tissue, CD5 expression was significantly correlated with clinicopathological features such as tumor stage and lymph node metastasis status. CD5 was significantly overexpressed in stage 3, N2 lymph node metastasis, and triple-negative breast cancer (Fig. 7B–D).
Fig. 7.
Analysis of an independent data set (GSE1456) supports the correlation between CD5 expression and survival and its association with clinicopathological characteristics. A An independent data set from the GEO database (GSE1456) was used for validating the results of survival analysis. We observed a correlation between high CD5 expression and improved OS in patients (P < 0.05). B–D Analysis of CD5 expression and clinicopathological features showed that CD5 expression is significantly correlated with cancer stage and lymph node metastasis status (P < 0.05)
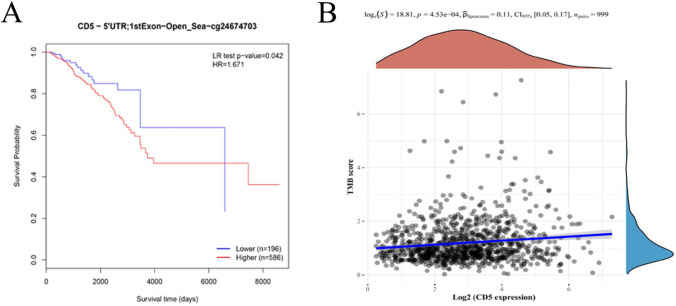
Notably, we observed an abnormally low level of CD5 promoter methylation in breast cancer. Survival analysis showed that patients with lower CD5 promoter methylation level had a longer OS (Fig. 8A). In addition, a significant positive correlation was observed between CD5 expression and TMB (Fig. 8B). Taken together, these results suggest that CD5 expression is strongly associated with breast cancer development and patient survival status.
Fig. 8.
MethSurv graphical output generated by CpG cg24674703 in BRCA samples, and the association of CD5 expression with TMB. A KM plot of cg24674703-CD5 using BRCA samples dichotomized using the MaxStat method. Blue represents CD5 low expression group, and red represents CD5 high expression group (P < 0.05). B CD5 expression was significantly and positively correlated with TMB (P < 0.05)
CD5 expression validation
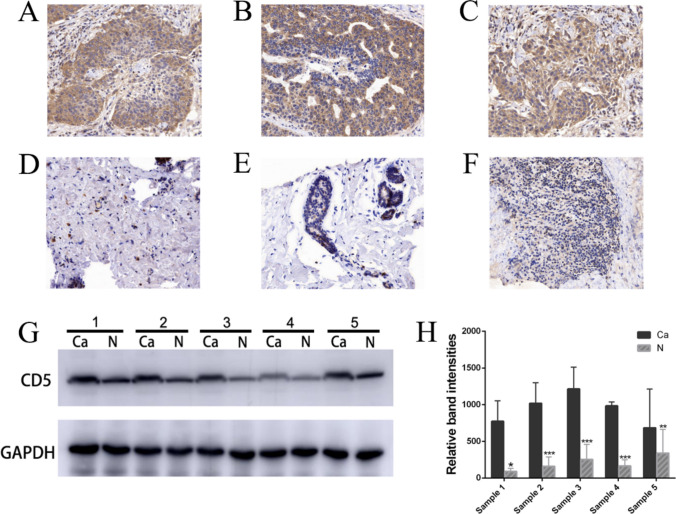
To validate the results of our bioinformatics analysis, we performed immunohistochemical staining and western blot analysis of fresh clinical tissue specimens. Consistent with our bioinformatics analysis results, the expression levels of CD5 in breast cancer tissue samples were significantly higher compared to normal tissue (P < 0.05), as confirmed by immunohistochemical staining and western blot analysis results (Fig. 9A–H).
Fig. 9.
Immunohistochemical staining and western blot analysis confirmed higher CD5 expression in cancer tissue compared to normal breast tissue. Fresh tissue specimens from either breast cancer tissue (A–C) or normal breast tissue (D–F) were subjected to immunohistochemical staining and western blot analysis (G, H). In A–C CD5 was highly expressed in breast cancer cells, and a few lymphocytes also had CD5 expression. Gray-scale value analysis of western blot analysis results
Gene set enrichment analysis (GSEA)
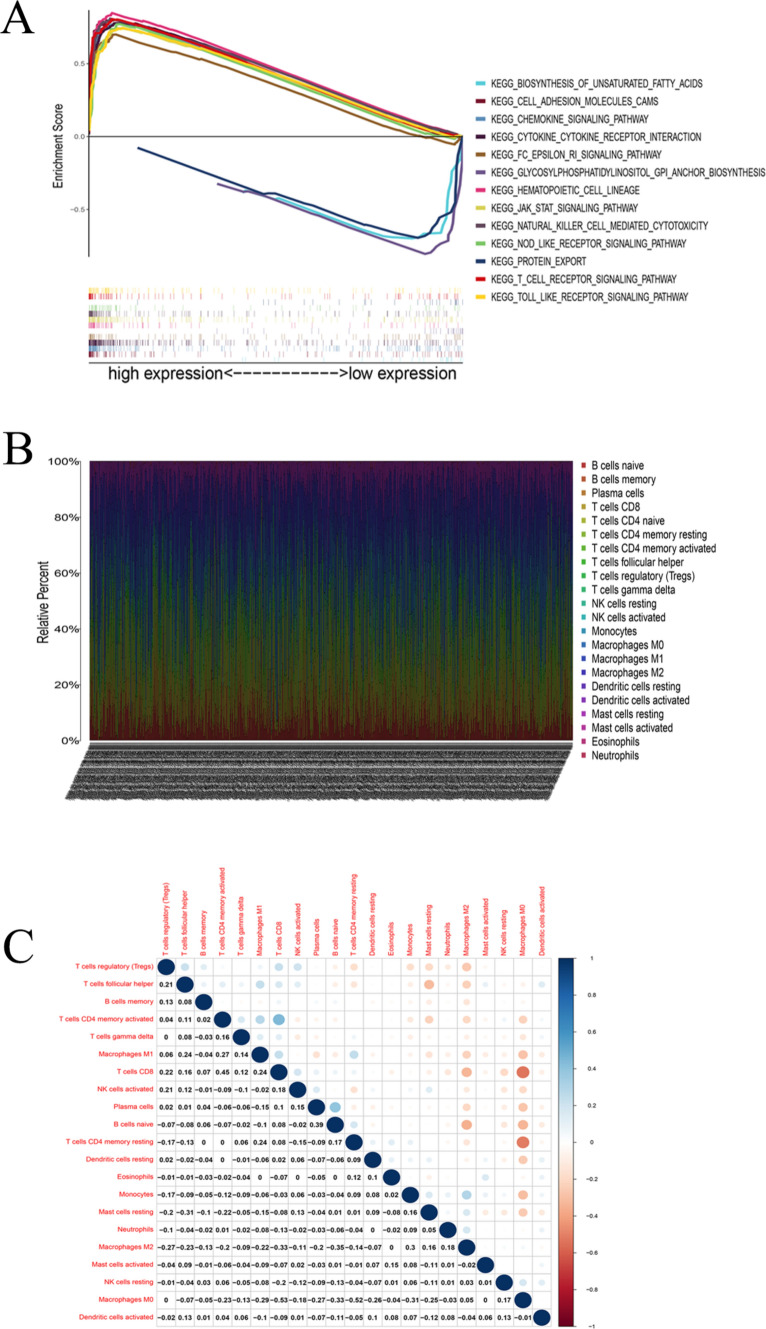
GSEA showed that the high-CD5 expression group was mainly enriched in natural killer cell-mediated cytotoxicity, the T cell receptor signaling pathway, the B cell receptor signaling pathway, leukocyte transendothelial migration, pathways in cancer, the Jak-STAT signaling pathway, the VEGF signaling pathway, the MAPK signaling pathway, and other pathways related to tumor and immune responses. In contrast, low-CD5-expressing tissues were more enriched in the protein export, glycosylphosphatidylinositol synthesis, and lipid synthesis pathways (Fig. 10A).
Fig. 10.
The expression distribution of 22 TICs in BCa samples and the correlation analysis between each of the 22 TICs, and GSEA correlation analysis of CD5 expression level. A The top 10 most significantly enriched pathways in the samples from the high-CD5 expression group and the only three significantly enriched pathways in the samples from the low-CD5 expression group. B The proportion of distribution of 22 TICs in the BCa sample. C Correlation analysis between TICs. Heat map showing the pairwise correlations of all 22 types of immune cells. Red represents a positive correlation, and blue represents a negative correlation
Correlation analysis between CD5 and TICs
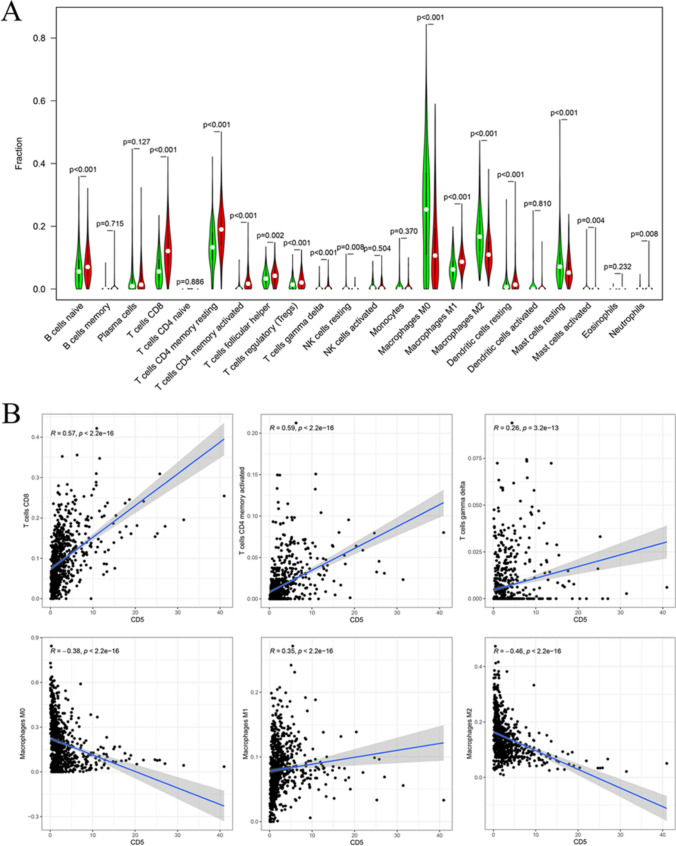
To further explore the association between CD5 and the immune microenvironment, we first analyzed the relative abundance of TIC subpopulations using the CIBERSORT algorithm. Based on this result, 22 immune cell profiles of BCa samples were constructed. Then, differential analysis and correlation analysis were performed. The results showed that the levels of a total of 15 types of immune cells were correlated with CD5 expression (Fig. 10B, C). Among them, the levels of CD8+ T cells, activated memory CD4+ T cells, gamma delta T cells, and M1 macrophages were positively correlated with CD5 expression, while M0 and M2 macrophages were negatively correlated with CD5 expression (Fig. 11A, B).
Fig. 11.
Correlation analysis between TICs and CD5 expression. A Violin plot showing the results of the correlation analysis between CD5 expression and the 22 types of TICs in BCa samples. B Scatter plot showing the immune cells whose proportions were most significantly correlated with CD5 expression. Among them, CD8+ T cells, activated memory CD4+ T cells, gamma delta T cells, and M1 macrophages were positively correlated with CD5 expression, while M0 and M2 macrophages were negatively correlated with CD5 expression. Red color represents high CD5 expression
Discussion
The biological behavior of tumors is not determined only by the nature of tumor cells themselves; the TME also plays an important role in the occurrence, progression, and metastasis of tumors [20]. The TME is a local internal environment composed of immune cells, interstitial cells, various cytokines, and tumor cells with local infiltration [21]. Extensive immune cell infiltration is evident in the local microenvironments of BCa, colorectal cancer, lung cancer, and other cancers. Local infiltration by these immune cells plays an important role in tumor immune escape and can directly affect the prognosis and survival of cancer patients [22–24]. Comprehensive assessment of TME features such as the distribution, density, and functional status of immune cells, can reveal new key markers that can regulate the TME; these factors can be targeted to prevent TME development or promote the transformation of the TME from a tumor-friendly to a tumor-suppressive type.
The innate ability of the immune system to detect and kill tumor cells is a very effective weapon against malignant tumors. Increasing evidence has confirmed that immune biomarkers are critical for cancer prevention and diagnosis and the selection of appropriate immunotherapy strategies. For example, immune checkpoint inhibitors targeting PD-1 and PDL-1 have achieved good therapeutic effects in a variety of tumors, including BCa [5].
CD5 is a member of the cysteine-rich scavenger receptor family. It is a 67-kDa transmembrane glycoprotein molecule that is mainly expressed in T cells, some B cell subpopulations, macrophages, and dendritic cell subpopulations [25–27]. As a lymphocyte surface receptor, CD5 is involved in the development, differentiation, and survival of lymphocytes. It is also a signal transduction receptor associated with the antigen-specific clonotype receptors of T cells and B1a cells in the immune synaptic center; CD5 can modulate the activation or death signals delivered by specific antigens, thus regulating the immune response [28].
In lymphocytes, CD5 is an important cell surface receptor that plays a key role in regulating the function of the immune system.The interaction of CD5 overexpression with tumor-infiltrating T cells has attracted considerable research attention. On the one hand, CD5 overexpression may promote the enhancement of anti-tumor immune responses. In the tumor microenvironment, tumor-infiltrating T cells usually show high levels of CD5 expression, which may reflect the activation status of T cells in tumor immune surveillance. This high expression of CD5 correlates with enhanced activity of T cells, which in turn may lead to a more effective anti-tumor response [29]. On the other hand, CD5 overexpression may also be triggered by an increase in the number of tumor-infiltrating T cells, and alterations in the tumor microenvironment may prompt T cells to up-regulate CD5. this phenomenon suggests a complex relationship between tumor progression and immune response. Specifically, certain cytokines secreted by tumor cells may stimulate T cells in the surrounding environment to express higher levels of CD5, thereby affecting their function and proliferative capacity [30]. Therefore, an in-depth exploration of CD5 expression and its relationship with tumor-infiltrating T cells will help to understand the immune escape mechanism of tumors, as well as provide possible targets for the development of new immunotherapeutic strategies.
This immunomodulatory feature of CD5 is closely associated with the occurrence and progression of tumors [31]. In small-cell lung cancer, high CD5 expression is associated with a good prognosis, possibly because high CD5 expression enhances the antitumor activity of T lymphocytes [14]. Similarly, genetic functional variations in CD5 can affect the survival rate of melanoma patients, and carrying the CD5 haplotype Pro224-Ala471 is associated with a better survival rate in melanoma patients [15]. The above findings demonstrate the important role of CD5 in tumor development and prognosis. However, the association between CD5 and the prognosis of BCa and its role in the TME of BCa remain unclear.
In this study, we found that CD5 was a TME-related immune gene that affected the survival of BCa patients, suggesting that CD5 may play a role in tumor immunity by regulating the TME around BCa. GSEA showed that high CD5 expression was closely correlated with tumor and immune response pathways, including natural killer cell-mediated cytotoxicity, the T cell receptor signaling pathway, the B cell receptor signaling pathway, leukocyte transendothelial migration, pathways in cancer, the Jak-STAT signaling pathway, the VEGF signaling pathway, and the MAPK signaling pathway. Low CD5 expression was more associated with protein export, glycosylphosphatidylinositol synthesis, and lipid synthesis pathways, indicating that high CD5 expression likely regulates the TME and thus influences tumor occurrence and progression through the multiple immune response-related pathways mentioned above.
In BCa patients, high expression of immune-related genes is associated with longer progression-free survival, even for BCa subtypes with a relatively high degree of malignancy, such as human epidermal growth factor receptor 2 (HER2)–positive BCa [32]. The significant correlation between immune score and overall survival (OS) suggests the important role of the immune microenvironment in tumor progression. A high immune score is generally associated with more effector T cell infiltration and predicts a better prognosis. This finding not only provides biomarkers for clinical prognosis, but may also influence treatment decisions, such as selecting immune checkpoint inhibitors or adjusting chemotherapy regimregimes. In early-stage, lymph node–negative, HER2-amplified BCa, increased expression levels of lymphocyte-related immune genes have been correlated with a reduction in the recurrence rate of BCa [33]. These reports are consistent with our results. Thus, changes in the expression levels of key immune genes might affect the prognosis of BCa patients.
By analyzing the data available on TCGA database, we found that CD5 expression in BCa tissue was markedly different from that in normal breast tissue. Patients with BCa and high CD5 expression levels had better prognoses and higher survival rates. These results were validated by analyzing an independent GEO dataset as well as by performing immunohistochemical and western blot analyses of clinical breast cancer specimens.
We also found that CD5 was positively correlated with the proportion of T cells, M1 macrophages, and other immune cells in the TME, while it was negatively correlated with M2 macrophage infiltration. Different immune cells often play different roles in tumors. Therefore, we reviewed previous reports on the relationship between immune cells and the prognosis of BCa patients. Increased levels of T cell activation can significantly improve the OS rate of BCa patients [34]. An increase in CD4+ and CD8+ T cell infiltration indicates a good prognosis in the subpopulation of triple-negative BCa [35]. Tumor-associated macrophages can infiltrate a variety of tumors [36]. BCa patients with high expression of M1 macrophages have high immune activity and a good tumor immune microenvironment, as well as high expression of immune checkpoint molecules [37]. Cordyceps sinensis extract promotes M1 polarization in macrophages by activating the NF-κB pathway, thereby inhibiting BCa growth [38]. On the other hand, M2 macrophages play a greater role in promoting BCa progression than M1 macrophages. The ERK/STAT3 signaling pathway can drive the development and progression of BCa by stimulating M2-like polarization of macrophages [39]. M2 macrophages recruited by tumors promote gastric cancer and BCa metastasis through the secretion of CHI3L1 protein [40]. All of these reports are consistent with our observations. The positive or negative correlations between CD5 and these immune cell populations suggest that CD5 has an important impact on the immune activity status of the TME.
Conclusions
In summary, by analyzing the transcriptome data of BCa in the TCGA database, CD5 was identified as a key immune gene associated with the TME. High CD5 expression suggests a better prognosis for BCa patients and is a potential prognostic biomarker for BCa. According to the results of the GSEA and TIC analysis, we can speculate that CD5 has an immunomodulatory effect in the tumor environment and can be used as a biomarker and regulator of the immune status of the TME. One limitation of this study is that the results require further verification through experiments. Another limitation is that the regulatory mechanisms of CD5 expression, as well as the specific regulatory pathways in which CD5 participates, require further exploration and elaboration.
Acknowledgements
Not applicable.
Author contributions
YZ, HHZ and WWW contribute equally to this work. JML and JDZ: Study concept and experimental design. WWW, MZW: collected the samples and medical history and performed IHC staining. YZ, GSS and ZL: Analysis and interpretation of data. YZ, HHZ and WWW: Writing of the manuscript, preparation of figures and statistical analysis. The authors read and approved this manuscript.
Funding
This work was supported by grants from the Thousand Talents of Program of High-end Innovation of Qinghai Province in China (for GuoShuang Shen).
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Declarations
Ethics approval and consent to participate
Not applicable.
Competing interests
The authors declare no competing interests.
Footnotes
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Yi Zhao, Hengheng Zhang and Wenwen Wang have contributed equally.
References
- 1.Siegel RL, Miller KD, Jemal A. Cancer statistics, 2020. CA Cancer J Clin. 2020;70(1):7–30. [DOI] [PubMed] [Google Scholar]
- 2.Jiao Y, Fu Z, Li Y, Zhang W, Liu Y. Aberrant FAM64A mRNA expression is an independent predictor of poor survival in pancreatic cancer. PLoS ONE. 2019;14(1): e0211291. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Li X, Wenes M, Romero P, Huang SC, Fendt SM, Ho PC. Navigating metabolic pathways to enhance antitumour immunity and immunotherapy. Nat Rev Clin Oncol. 2019;16(7):425–41. [DOI] [PubMed] [Google Scholar]
- 4.Wagner J, Rapsomaniki MA, Chevrier S, et al. A single-cell atlas of the tumor and immune ecosystem of human breast cancer. Cell. 2019;177(5):1330-1345.e18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Bertucci F, Gonçalves A. Immunotherapy in breast cancer: the emerging role of PD-1 and PD-L1. Curr Oncol Rep. 2017;19(10):64. [DOI] [PubMed] [Google Scholar]
- 6.Jiang C, Cao S, Li N, Jiang L, Sun T. PD-1 and PD-L1 correlated gene expression profiles and their association with clinical outcomes of breast cancer. Cancer Cell Int. 2019;19:233. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Ren X, Wu H, Lu J, et al. PD1 protein expression in tumor infiltrated lymphocytes rather than PDL1 in tumor cells predicts survival in triple-negative breast cancer. Cancer Biol Ther. 2018;19(5):373–80. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Alotaibi F, Vincent M, Min WP, Koropatnick J. Reduced CD5 on CD8(+) T cells in tumors but not lymphoid organs is associated with increased activation and effector function. Front Immunol. 2020;11: 584937. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Velasco-de Andrés M, Casadó-Llombart S, Català C, Leyton-Pereira A, Lozano F, Aranda F. Soluble CD5 and CD6: lymphocytic class I scavenger receptors as immunotherapeutic agents. Cells. 2020;9(12):2589. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Demir C, Kara E, Ekinci Ö, Ebinç S. Clinical and laboratory features of CD5-negative chronic lymphocytic leukemia. Med Sci Monit. 2017;23:2137–42. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Jaseb K, Purrahman D, Shahrabi S, Ghanavat M, Rezaeean H, Saki N. Prognostic significance of aberrant CD5 expression in B-cell leukemia. Oncol Rev. 2019;13(1):400. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Hosaka N, Ohe C, Miyasaka C, et al. The role of CD5 expression in thymic carcinoma: possible mechanism for interaction with CD5+ lymphoid stroma (microenvironment). Histopathology. 2016;68(3):450–5. [DOI] [PubMed] [Google Scholar]
- 13.Zhao P, Li L, Zhou S, et al. CD5 expression correlates with inferior survival and enhances the negative effect of p53 overexpression in diffuse large B-cell lymphoma. Hematol Oncol. 2019;37(4):360–7. [DOI] [PubMed] [Google Scholar]
- 14.Moreno-Manuel A, Jantus-Lewintre E, Simões I, et al. CD5 and CD6 as immunoregulatory biomarkers in non-small cell lung cancer. Transl Lung Cancer Res. 2020;9(4):1074–83. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Potrony M, Carreras E, Aranda F, et al. Inherited functional variants of the lymphocyte receptor CD5 influence melanoma survival. Int J Cancer. 2016;139(6):1297–302. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.A language and environment for statistical computing.
- 17.Ritchie ME, Phipson B, Wu D, et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015;43(7): e47. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Yu G, Wang LG, Han Y, He QY. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS. 2012;16(5):284–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Newman AM, Liu CL, Green MR, et al. Robust enumeration of cell subsets from tissue expression profiles. Nat Methods. 2015;12(5):453–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Noël A, Jost M, Maquoi E. Matrix metalloproteinases at cancer tumor-host interface. Semin Cell Dev Biol. 2008;19(1):52–60. [DOI] [PubMed] [Google Scholar]
- 21.Wang JJ, Lei KF, Han F. Tumor microenvironment: recent advances in various cancer treatments. Eur Rev Med Pharmacol Sci. 2018;22(12):3855–64. [DOI] [PubMed] [Google Scholar]
- 22.Soysal SD, Tzankov A, Muenst SE. Role of the tumor microenvironment in breast cancer. Pathobiology. 2015;82(3–4):142–52. [DOI] [PubMed] [Google Scholar]
- 23.Koi M, Carethers JM. The colorectal cancer immune microenvironment and approach to immunotherapies. Future Oncol. 2017;13(18):1633–47. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Mittal V, El Rayes T, Narula N, McGraw TE, Altorki NK, Barcellos-Hoff MH. The microenvironment of lung cancer and therapeutic implications. Adv Exp Med Biol. 2016;890:75–110. [DOI] [PubMed] [Google Scholar]
- 25.Yin X, Yu H, Jin X, et al. Human blood CD1c+ dendritic cells encompass CD5high and CD5low subsets that differ significantly in phenotype, gene expression, and functions. J Immunol. 2017;198(4):1553–64. [DOI] [PubMed] [Google Scholar]
- 26.Berland R, Wortis HH. Origins and functions of B-1 cells with notes on the role of CD5. Annu Rev Immunol. 2002;20:253–300. [DOI] [PubMed] [Google Scholar]
- 27.Korenfeld D, Gorvel L, Munk A, et al. A type of human skin dendritic cell marked by CD5 is associated with the development of inflammatory skin disease. JCI Insight. 2017; 2(18). [DOI] [PMC free article] [PubMed]
- 28.Cho JH, Sprent J. TCR tuning of T cell subsets. Immunol Rev. 2018;283(1):129–37. [DOI] [PubMed] [Google Scholar]
- 29.Drennan MB, et al. CD5 modulates T lymphocyte activation. Cell Immunol. 2019;345:104–12. [Google Scholar]
- 30.Liao Y, et al. The role of tumor microenvironment in the upregulation of CD5 in T cells. Front Immunol. 2020;11:68.32117240 [Google Scholar]
- 31.Brossard C, Semichon M, Trautmann A, Bismuth G. CD5 inhibits signaling at the immunological synapse without impairing its formation. J Immunol. 2003;170(9):4623–9. [DOI] [PubMed] [Google Scholar]
- 32.Matikas A, Lövrot J, Ramberg A, et al. Dynamic evaluation of the immune infiltrate and immune function genes as predictive markers for neoadjuvant chemotherapy in hormone receptor positive, HER2 negative breast cancer. Oncoimmunology. 2018;7(9): e1466017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Alexe G, Dalgin GS, Scanfeld D, et al. High expression of lymphocyte-associated genes in node-negative HER2+ breast cancers correlates with lower recurrence rates. Cancer Res. 2007;67(22):10669–76. [DOI] [PubMed] [Google Scholar]
- 34.Lu L, Bai Y, Wang Z. Elevated T cell activation score is associated with improved survival of breast cancer. Breast Cancer Res Treat. 2017;164(3):689–96. [DOI] [PubMed] [Google Scholar]
- 35.Matsumoto H, Thike AA, Li H, et al. Increased CD4 and CD8-positive T cell infiltrate signifies good prognosis in a subset of triple-negative breast cancer. Breast Cancer Res Treat. 2016;156(2):237–47. [DOI] [PubMed] [Google Scholar]
- 36.Komohara Y, Fujiwara Y, Ohnishi K, Takeya M. Tumor-associated macrophages: potential therapeutic targets for anti-cancer therapy. Adv Drug Deliv Rev. 2016;99(Pt B):180–5. [DOI] [PubMed] [Google Scholar]
- 37.Oshi M, Tokumaru Y, Asaoka M, et al. M1 Macrophage and M1/M2 ratio defined by transcriptomic signatures resemble only part of their conventional clinical characteristics in breast cancer. Sci Rep. 2020;10(1):16554. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Li J, Cai H, Sun H, et al. Extracts of Cordyceps sinensis inhibit breast cancer growth through promoting M1 macrophage polarization via NF-κB pathway activation. J Ethnopharmacol. 2020;260: 112969. [DOI] [PubMed] [Google Scholar]
- 39.Mu X, Shi W, Xu Y, et al. Tumor-derived lactate induces M2 macrophage polarization via the activation of the ERK/STAT3 signaling pathway in breast cancer. Cell Cycle. 2018;17(4):428–38. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Chen Y, Zhang S, Wang Q, Zhang X. Tumor-recruited M2 macrophages promote gastric and breast cancer metastasis via M2 macrophage-secreted CHI3L1 protein. J Hematol Oncol. 2017;10(1):36. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.