Abstract
The cervical cell classification technique can determine the degree of cellular abnormality and pathological condition, which can help doctors to detect the risk of cervical cancer at an early stage and improve the cure and survival rates of cervical cancer patients. Addressing the issue of low accuracy in cervical cell classification, a deep convolutional neural network A2SDNet121 is proposed. A2SDNet121 takes DenseNet121 as the backbone network. Firstly, the SE module is embedded in DenseNet121 to increase the model’s focus on the nucleus region, which contains important diagnostic information, and reduce the focus on redundant information. Secondly, the sizes of the convolutional kernel and pooling window of the Stem layer are adjusted to adapt to the characteristics of the cervical cell images, so that the model can extract the local detailed information more effectively. Finally, the Atrous Dense Block (ADB) is constructed, and four ADB modules are integrated into DenseNet121 to enable the model to acquire global and local salient feature information. The accuracy of A2SDNet121 for two and seven-classification tasks on the Herlev dataset is 99.75% and 99.14%, respectively. The accuracy for two, three, and five-classification tasks on the SIPaKMeD dataset reaches 99.55%, 99.75% and 99.22%, respectively. Compared with other state-of-the-art algorithms, the A2SDNet121 model performs better in the multi-classification task of cervical cells, which can significantly improve the accuracy and efficiency of cervical cancer screening.
Keywords: Cervical cells, Convolutional neural network, Image classification, Multi-scale features, Attention mechanism
Subject terms: Cancer, Cancer screening
Introduction
Cervical cancer is a serious health threat faced by many women in developing countries. According to the “Global Cancer Statistics 2022”1 released by the International Agency for Research on Cancer of the World Health Organization, there were 660,000 new cervical cancer cases and 350,000 deaths in 2022. In China, there were approximately 150,000 new cases of cervical cancer and nearly 60,000 deaths, accounting for 22.7% and 17.1% of the global total, respectively. As can be seen, China has about one fifth of the world’s cervical cancer patients, and the incidence and mortality rates remain high. Cervical cancer usually refers to the abnormal changes in cervical epithelial cells after infection by human papillomavirus, resulting in the formation of precancerous lesions and ultimately developing into cancer. If the disease is diagnosed in the precancerous stage or earlier, cervical cancer can be prevented and controlled more effectively.
Currently, the discrimination criteria for cervical cell classification are reported by The Bethesda System (TBS)2014 version2. The specific classifications are as follows: negative for intraepithelial lesion or malignancy (NILM), atypical squamous cells of undetermined significance (ASC-US), low-grade squamous intraepithelial lesion (LSIL), atypical squamous cells, cannot exclude HSIL (ASC-H), and high-grade squamous intraepithelial lesion (HSIL). Proper classification of cervical cells can determine the degree of cellular abnormality and pathology, which can play a key role in pre-cancer screening for cervical cancer. Therefore, cervical cell screening is an important tool in the prevention of cervical cancer and its precancerous lesions, and is essential for saving lives.
In the past, cervical cell screening relied on the experience of technicians, which had the disadvantages of cumbersome, time-consuming and inefficient. Therefore, automatic screening techniques have attracted much attention3–5. At present, cervical cell automatic screening models are divided into traditional machine learning methods and deep learning methods, in terms of their technical implementation. Traditional machine learning methods require manual feature extraction. In contrast, deep learning methods do not require a complex process to achieve end-to-end lesion recognition, resulting in significant improvements in classification performance.
The ability to extract local detailed features and multi-scale features is a key factor that affects the performance of deep learning-based image classification models. Especially when dealing with complex medical images, the combination of these two types of features can improve the model’s sensitivity to details and understanding of global information. For cervical cell images, the performance of deep learning classification models has been enhanced to some extent, providing strong support for early screening and accurate diagnosis of cervical cancer. However, the models are still deficient in extracting local detailed features and capturing multi-scale information, which influences the classification performance and still exists certain gap with the needs of clinical applications.
To address the problems, a deep learning model A2SDNet121 for implementing multi-classification task of cervical cells is proposed to further improve the performance of the classification model. The main contributions of this paper are as follows: (1) A2SDNet121 utilizes the dense connection network DenseNet1216 as the backbone network to extract fine-grained features of cervical cell images, and achieves feature reuse through the dense connection mechanism. (2) The A2SDNet121 model enhances the attention to the nuclear region of cells by introducing the SE module7, which can reduce the influence of redundant information on the classification results. (3) The strategy of adjusting the Stem layer enables A2SDNet121 to better capture the local detailed features in the image. (4) A2SDNet121 utilizes atrous dense blocks based on atrous convolution8 to aggregate multi-scale contextual information and acquire multi-scale features.
Related works
In 2017, Zhang et al.9 introduced the Convolutional Neural Network (CNN) into the task of cervical cell classification for the first time. Since then, various CNN-based models have been widely applied in medical image analysis. In recent years, the main cervical cell classification models include U-Net, MobileNet, AlexNet, VGG, ResNet, DenseNet, and so on.
In 2019, Promworn et al.10 conducted a comparative study based on AlexNet, ResNet101, Vgg19_bn, DenseNet161, screzeneetl_1 models to evaluate their performance in the classification task. The experimental results on the Herlev dataset showed that DenseNet161 model is the best deep learning model, and the accuracy of binary classification and seven-classification are 94.38% and 68.54%, respectively. In the same year, Muhammed Talo11 classified the cell images in the cervical smear using the DenseNet model, evaluating the model on the SIPaKMeD dataset. The average classification accuracy of the model without data augmentation is 98.96%±0.12. In 2020, Elima et al.12 proposed a Fully Convolutional Neural Network (FCN) model based on U-Net architecture, which incorporated residual blocks, dense connection blocks, and fully convolutional layers in the encoding-decoding blocks. The accuracy of binary classification on the Herlev dataset reaches 98.8%. In 2021, Shi et al.13 proposed a novel classification method based on Graph Convolutional Network (GCN). The method clusters and analyzes the CNN features of all cell images, then constructs a graph structure, finally generates the relation-aware feature representation using GCN. The accuracy on the SIPaKMeD dataset achieves 98.73%. In 2022, Chen et al.14 introduced a lightweight CNN architecture that utilizes knowledge distillation to improve its representation capability. The seven-classification experiments on the Herlev dataset showed that the accuracy of Xception is improved by 0.86%. In the same year, Waly et al.15 developed a new intelligent deep convolutional neural network for cervical cancer detection and classification (IDCNN-CDC) model. The model used the Tsallis entropy technique with the dragonfly optimization (TE-DFO) algorithm to segment the image, and then extracted the features by SqueezeNet model. The accuracy of seven-classifications on the Herlev dataset reaches 97.96%. In addition, Kundu et al.16 proposed a method to extract deep features using CNN, which combined metaheuristics to eliminate redundant features to realize the selection and classification of the optimal feature subset. The accuracy of five-classification on the SIPaKMeD dataset reached 98.94%. In 2023, Das et al.17 proposed an Oppositional-based Harmony Search Algorithm (O-bHSA). The algorithm was composed of CNN and a metaheuristic optimization algorithm. The accuracy of five-classification on the SIPaKMeD dataset reaches 97.93%. Xu et al.18 combined Faster R-CNN, shallow feature enhancement network, and generative adversarial network to improve the classification performance. The maximum accuracy on the mixed datasets of SIPaKMeD and Herlev reaches 99.81%.
In order to get the desired classification results, deep learning techniques need a large amount of labeled sample data. However, cervical cell images, as a kind of medical images, the establishment of dataset containing a large number of labeled images is difficult. In recent years, researchers have solved the small sample classification problem by means of transfer learning and generative adversarial networks. In 2020, Chen et al.19 proposed a Transfer Learning based Snapshot Ensemble (TLSE) method. The advantage was that the integration benefits were obtained in a single CNN model through ensemble learning, meanwhile the introducing of transfer learning solved the problem of less data and improved the generalization ability. In 2022, Lin et al.20 designed an algorithm using super-resolution and ordinal regression, which was implemented by using generative adversarial network. The accuracy of binary classification and seven-classification on the Herlev dataset is 98.1% and 97.6% respectively. The algorithm was competitive even when the data is small and the resolution is low. In the same year, Gao et al.21 proposed a method combining transfer learning and knowledge distillation to improve the network’s generalization ability under limited data by transferring feature knowledge between ResNet18 network and data. The accuracy of five-classification on the SIPaKMeD dataset reaches 98.52%. In 2024, Song et al.22 proposed an effective worst-case enhancement algorithm to help classifiers learn more information from underrepresented datasets, thus improving the generalization ability of classifiers in small sample cases. The accuracy of binary classification and five-classification on the SIPaKMeD dataset are 90.7% and 88.9%, respectively.
The purpose of feature fusion is to combine feature information from different levels to improve the model performance. Some researchers have performed cervical cell classification by fusing features generated from different models or features generated from different scales. In 2021, Rahaman et al.23 proposed a feature fusion technique based on four models, VGG16, VGG19, XceptionNet and ResNet50, to accurately classify cervical cells. The accuracy of binary classification, three-class classification and five-classification on the SIPaKMeD dataset is 99.85%, 99.38% and 99.14%, respectively. The accuracy of binary classification and seven-classification on the Herlev dataset is 98.32% and 90.32%, respectively. In 2022, Qin et al.24 proposed DSRNet50 network based on ResNet50. The network first incorporated the packet convolution to reduce the computational burden, then utilized the attention channel to focus on useful features and suppress useless features, and finally fused the feature information extracted from the residual structure and the dense connection structure. This method achieves 98.90% accuracy for seven-classification on the Herlev dataset. In the same year, Qin et al.25 designed a fusion model consisting of one secondary task and two primary tasks. The secondary task enhanced primary tasks through merging the underlying features. Combining the two primary tasks, two-classification and five-classification, realized the mutual reinforcement to mitigate the influence of unreliable labels. The accuracy of binary and five-classification on the SIPaKMeD dataset are 98.96% and 98.67%, respectively. In 2023, Chowdary et al.26 adopted a feature extraction model based on the fusion of VGG19, VGG-F, and CaffeNet models and a multi-layer perceptrons classification model to classify cervical cells. The accuracy of seven-classification on the Herlev dataset reaches 98.39%, and the accuracy of five-classification on the SIPaKMeD dataset reaches 99.16%. Hao et al.27 proposed a classification method based on cervical cell feature fusion, which generated new features by integrating shallow features extracted manually and deep features extracted from VGG16 network. The accuracy of binary classification and seven-classification on the Herlev dataset is 98.1% and 92.3%, respectively. In addition, Wu et al.28 proposed a cervical cell classification framework based on improved Convolutional Neural Networks-Lagrangian Support Vector Machine (CNN-LSVM). First, the strong features extracted from the combination method were fused with the features from the LeNet5 model, then the fused features were fed into LSVM classifier based on Adaboost optimization for classification. The accuracy of binary classification and seven-classification on the Herlev dataset is 99.5% and 94.2%, respectively.
In the above literature, various deep learning-based approaches have been proposed for realizing the cervical cell classification task. Although these methods have solved the classification challenge of small sample data to some extent and improved the classification performance, the extraction of local details and multi-scale features is still insufficient. Therefore, the accuracy of the classification model still needs to be further improved to meet the practical requirements of clinical applications.
Dataset and preprocessing
Dataset
We use the Herlev dataset29 and the SIPaKMeD dataset30 to conduct cervical cell classification experiments.
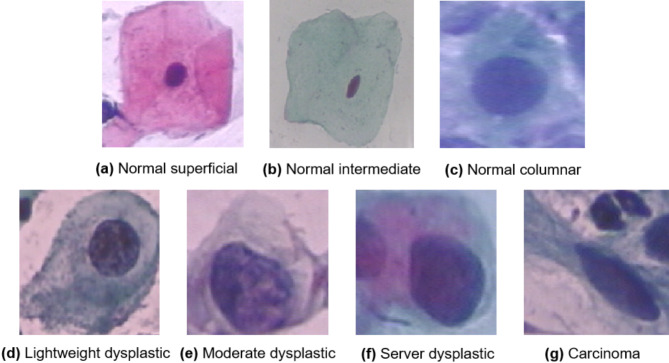
The Herlev dataset consists of 917 single-cell cervical cytopathology images. The image resolution is 0.201 μm/pixel and the average image size is 156 × 140 pixels. Each image is category labeled by two cytotechnicians and one physician. There are seven types of cervical cells including Normal superficial, Normal intermediate, Normal columnar, Lightweight dysplastic, Moderate dysplastic, Server dysplastic, and Carcinoma. The first three can be further classified as normal cells and the last four as abnormal cells. Example images of the seven types of cervical cells in this dataset are shown in Fig. 1.
Fig. 1.
Seven types of cervical cell images in Herlev dataset.
The SIPaKMeD dataset consists of 4049 single-cell cervical cytopathology images. Each image is category labeled by cytopathologists. The dataset shows five types of cervical cells. The superficial intermediate cells and parabasal cells are two types of normal cells. The koilocytotic cells and dyskeratotic cells are two types of abnormal cells. The metaplastic cells are benign cells and also belong to abnormal ones, in which virtually all cervical precancerous and cancerous lesions occur. Example images of the five types of cervical cells in this dataset are shown in Fig. 2.
Fig. 2.
Five types of cervical cell images in SIPaKMeD dataset.
Preprocessing
The cervical cell datasets used in this paper have a small number and suffer from data class imbalance. Problems such as overfitting, poor robustness, and weak generalization ability may occur when training the network model. Data augmentation techniques can solve this problem by increasing the number of training samples. The common data augmentation operations include rotation, flipping, color transformation, scale transformation and noise transformation. Cervical cells usually have symmetry and small variations in size and morphology structure. Due to the particularity of cervical cells, the transformation operations (color, scale and noise) can adversely affect the morphological and staining features of cervical cells, thereby reducing the classification performance of the model. We adopt the operations of flipping and rotating31 to expand the dataset. The two augmentation techniques can both maintain the original features of the cells and be capable of enhancing the robustness of the model. The expanded images are uniformly processed to the size of 224 × 224. The Herlev and SIPaKMeD datasets after data augmentation are shown in Tables 1 and 2, respectively.
Table 1.
Number of cervical cell images of each category in Herlev dataset before and after data augmentation.
| Category | Class | Cell type | Original number | Number of expansions |
|---|---|---|---|---|
| Normal | 1 | Normal superficial | 74 | 592 |
| 2 | Normal intermediate | 70 | 560 | |
| 3 | Normal columnar | 98 | 588 | |
| Abnormal | 4 | Lightweight dysplastic | 182 | 546 |
| 5 | Moderate dysplastic | 146 | 584 | |
| 6 | Server dysplastic | 197 | 591 | |
| 7 | Carcinoma | 150 | 600 | |
| Total | 917 | 4061 | ||
Table 2.
Number of cervical cell images of each category in SIPaKMeD dataset before and after data augmentation.
| Category | Class | Cell type | Original number | Number of expansions |
|---|---|---|---|---|
| Normal | 1 | Superficial | 831 | 4986 |
| 2 | Parabasal | 787 | 4722 | |
| Abnormal | 3 | Koilocytotic | 825 | 4950 |
| 4 | Dyskeratotic | 813 | 4878 | |
| Benign | 5 | Metaplastic | 793 | 4758 |
| Total | 4049 | 24,294 | ||
Methods
A2SDNet121 classification model
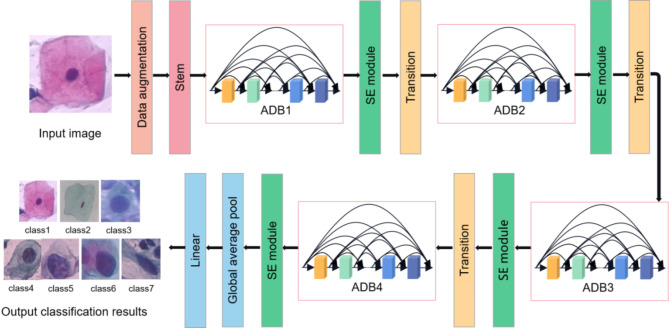
We propose a deep convolutional neural network model, A2SDNet121, to realize cervical cell classification automatically. The framework of the model is shown in Fig. 3. It takes DenseNet121 as the backbone network, including Stem layer, Atrous Dense Block (ADB), Squeeze-and-Excitation (SE) module, Transition layer, Average pooling layer and Fully connected layer. The SE module is introduced to enhance the correlation between channels. Feature utilization is improved by selectively highlighting information-rich features. The main function of the Stem layer is to extract the image features. By reducing the size of the Stem layer, the model is able to capture finer local features of the cervical cell images. ADB is used for multi-scale feature extraction to further aggregate multi-scale contextual information. These adjustments and improvements will help to enhance network performance and provide more accurate results for image classification tasks.
Fig. 3.
Framework of the A2SDNet121 classification model.
SE module
The SE module is an attention mechanism designed to automatically adjust the channel feature response, which can highlight important features and increase their weight, thereby improving the network’s attention to important information in the input image.
The nuclei of cervical cells contain a great deal of diagnostic information such as shape and colour. The nuclei of normal cells are usually small and dense, uniform in colour and normal in structure. While abnormal cells often have swollen, irregular nuclei with blurred borders. Their chromatin may appear abnormal hyperplasia or deletion, resulting in uneven colour and easy to misdetect. The SE module incorporated in the proposed model pays more attention to the nucleus region of cervical cells when predicting cervical cells, which reduces the focus on redundant information and improves the model classification accuracy.
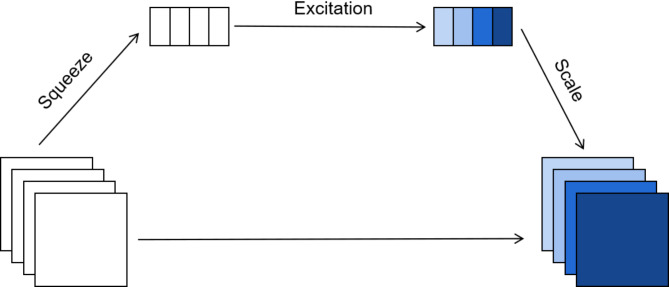
The structure of the SE module is shown in Fig. 4, including three key operational steps: Squeeze, Excitation and Scale.
Fig. 4.
Structure of SE module.
Squeeze. The Squeeze operation uses global average pooling for feature compression, synthesizing the feature map from H×W×C to 1 × 1×C. The operation extracts global information from each channel and provides a global perspective for subsequent channel weight learning, enhancing attention to the nuclear region. The squeeze operation is calculated as follows:
| 1 |
where,  denotes the average value of the f th feature map,
denotes the average value of the f th feature map, 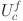 denotes the cth two-dimensional matrix in the feature map, and W and H denote the width and height of the feature map, respectively.
denotes the cth two-dimensional matrix in the feature map, and W and H denote the width and height of the feature map, respectively.
-
(2)
Excitation. The Excitation operation processes global information through fully connected layers and activation functions, learning the weights of each channel that reflect the dependencies between channels and their importance in the nucleus region, which enables the network to automatically identify and reinforce key features. The calculation formula is as follows:
| 2 |
where, represents the sigmoid function,
represents the sigmoid function, 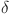 represents the ReLU function, and W1 and W2 represent the two fully connected layers.
represents the ReLU function, and W1 and W2 represent the two fully connected layers.
-
(3)
Scale. The Scale operation applies the learned channel weights to the original feature map and adjusts the feature response channel by channel, which can enhance the key features of the cell nucleus region and suppress the influence of irrelevant regions, thereby ensuring precise attention to the cell nucleus region. The formula for the Scale operation is as follows:
| 3 |
where, 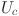 represents the feature map of each channel,
represents the feature map of each channel, 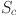 represents the weight coefficient of each feature channel obtained from the excitation operation, and
represents the weight coefficient of each feature channel obtained from the excitation operation, and represents the feature map with feature weight.
represents the feature map with feature weight.
Stem layer
The Stem layer includes a convolution layer and a pooling layer. In this paper, both layers are optimized to capture finer local features of images and improve the effect of feature extraction.
Convolution layer
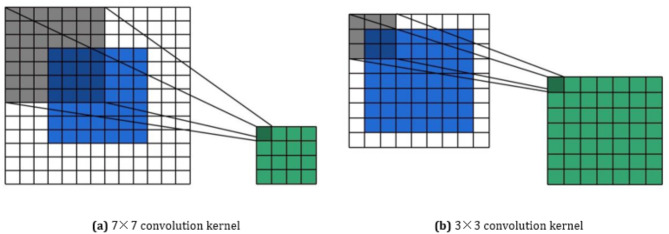
The convolutional layer of the Stem layer in DenseNet121 network consists of only 1 convolutional kernel, with a size of 7 × 7, a stride of 2 and a padding of 3. The size of the convolutional kernel determines the size of the receptive field when extracting the feature, thus affects the model’s ability to perceive the local image. Although the large convolutional kernel has a large receptive field, its ability to capture subtle local features is limited, especially when dealing with the small size structure of cervical cell nuclei, which can easily dilute feature information. Therefore, the proposed model adjusts the size of convolution kernel of Stem layer to 3 × 3, with the stride of 1 and the padding of 1. Figure 5 shows the convolution process of 7 × 7 and 3 × 3 convolution kernel. The improved convolutional layer significantly enhances the ability to perceive details in local areas, which can capture key features of cervical cell nuclei more effectively and reduce redundant interference from background information at the same time.
Fig. 5.
Convolution process of 7 × 7 and 3 × 3 convolution kernel.
Pooling layer
The Stem layer of the DenseNet121 network uses the max pooling method for its pooling layer. The pooling window size is 3 × 3, the stride is 2, and the padding is 1. The size of the pooling window determines the number of features to be pooled within each window, which affects the ability of the model to capture local features of the image. Although the larger pooling window can effectively reduce the size of feature maps, it may lead to the loss of key features such as local textures and edges of cervical cell nuclei. In this paper, according to the size features of cervical cell nuclei, the window size of the max pooling operation is adjusted to 2 × 2, the stride remains unchanged at 2, and the padding is adjusted to 0. Figure 6 shows the original and the modified max pooling processes. The improved pooling layer reduces the resolution of feature maps while minimizing information loss, which can better preserve important local feature information and enhance the model’s perception ability of subtle regions.
Fig. 6.
Pooling process.
Atrous dense block
Atrous convolution
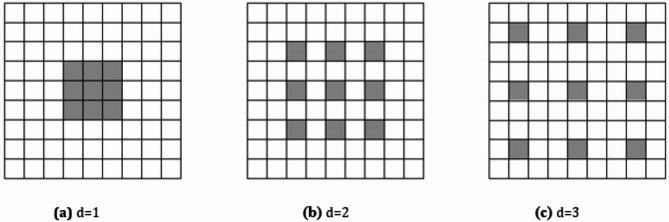
Compared to ordinary convolution, Atrous Convolution introduces a dilation rate d, which indicates the number of rows and columns with weight 0 inserted into the convolution kernel. Figure 7 (a) shows a general convolution, that is, a special form of atrous convolution with a dilation rate of 1. The size of its receptive field is 3 × 3. Figure 7 (b) shows an atrous convolution with a dilation rate of 2, and the size of the receptive field is 5 × 5. Figure 7 (c) is an atrous convolution with a dilation rate of 3, and the size of the receptive field is 7 × 7. In atrous convolution, the larger the dilation rate, the bigger the receptive field, and the more feature information can be obtained. Atrous convolution can increase the receptive field while maintaining the computational efficiency of the network, and is an effective method to balance the depth of the network and the size of the receptive field. This mechanism allows the model to maintain sensitivity to local details while obtaining global contextual information.
Fig. 7.
Schematic diagram of atrous convolution with different dilation rate.
The size of the equivalent convolution kernel is related to the size of original convolution kernel:
| 4 |
The formula for calculating the size of receptive field is as follows:
| 5 |
where, 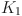 and
and 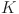 denotes the size of the equivalent and the original convolution kernel, respectively.
denotes the size of the equivalent and the original convolution kernel, respectively. 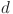 denotes the dilation rate, and
denotes the dilation rate, and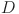 denotes the size of the receptive field.
denotes the size of the receptive field.
Dense block
The Dense Block of DenseNet121 is composed of multiple convolution layers, each of which contains one 1 × 1 convolution kernel and one 3 × 3 convolution kernel. The feature maps obtained from each convolutional layer are of the same size. A dense connection between layers is used to achieve feature reuse and solve the problem of information loss. Figure 8 shows the structure of the Dense Block containing four convolutional layers.
Fig. 8.
Structure of Dense Block.
The input of the DenseNet121 network at layer l is:
| 6 |
The output of DenseNet121 at layer l is:
| 7 |
where, 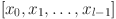 indicates the feature maps generated in layer 0, layer 1,…, layer l-1.
indicates the feature maps generated in layer 0, layer 1,…, layer l-1.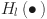 denotes the composite function of successive operations including batch normalization, ReLU activation function, and convolution operation.
denotes the composite function of successive operations including batch normalization, ReLU activation function, and convolution operation.  denotes the number of channels in the input layer, and
denotes the number of channels in the input layer, and 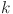 denotes the growth rate of the DenseNet121 network.
denotes the growth rate of the DenseNet121 network.
Atrous dense block
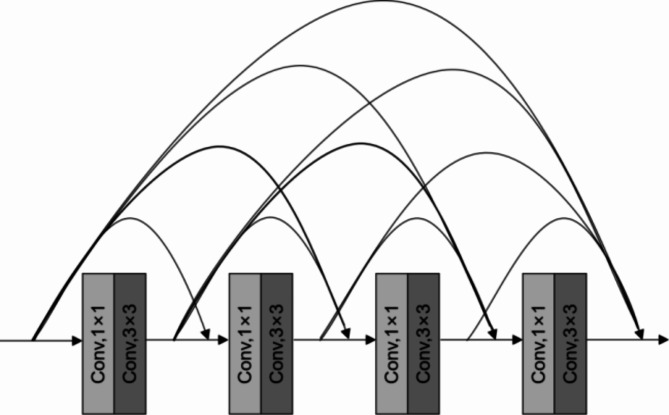
The A2SDNet121 model proposed in this paper consists of four Atrous Dense Block (ADB), denoted as ADB1-ADB4. The four ADB modules, in order, contain 6, 12, 16, and 24 convolutional layers, respectively. ADB is an improvement on the Dense Block. The last three layers of the Dense Block are adjusted to be atrous convolutional layers dilation rate d of 1, 2, and 3, respectively. The remaining layers remain unchanged and are still standard convolutional layers. Figure 9 shows the improved ADB structure with 4 convolutional layers as an example. ADB can obtain global and local salient feature information by using the dense connection of standard convolutional layers and different atrous convolutional layers, achieving efficient fusion of multi-scale features. Standard convolutional layers focus on extracting local detailed features such as edges and textures. Atrous convolutional layers effectively capture global contextual information by expanding the receptive field, significantly enhancing the expressive power of global features. The combination can extract more diverse multi-scale feature content.
Fig. 9.
Structure of ADB.
Experimental design
Evaluation metrics
Five evaluation metrics are used to evaluate the performance of classification models, including Accuracy (Acc), Precision (Pre), Specificity (Spe), Recall (Rec) and F1-score. Accuracy reflects the overall performance of the classification model. Precision and Recall reflect the model’s ability to recognize positive samples, and Specificity response the ability to recognize negative samples. F1-score denotes the harmonic mean of Precision and Recall. The formula for each metric is as follows:
| 8 |
| 9 |
| 10 |
| 11 |
| 12 |
where, TP (True Positive) refers to the number of positive samples correctly classified. FP (False Positive) refers to the number of negative samples that are incorrectly labelled as positive ones. TN (True Negative) refers to the number of negative samples correctly classified. FN (False Negative) refers to the number of positive samples that are incorrectly labelled as negative.
Experimental environment
We choose Python as the programming language and conduct all the experiments on NVIDIA RTX5000 GPU. The deep learning framework of PyTorch is employed to implement and train the classification models. For fair comparison, the same training schedule is designed for all classification models. Models adopt cross-entropy loss function, SGD optimizer, 300 epochs of training rounds. The initial learning rate is set to 0.0001, the decay rate is set to 0.1, and the batch size is set to 8. The other parameters are set to the default settings of the PyTorch framework.
The learning rate scheduler adopts the StepLR strategy to achieve the function of adjusting the learning rate according to the step size. The calculation formula is as follows:
| 13 |
where, lr is the initial learning rate, gamma is the attenuation factor, and epoch is the number of epochs of the current training. // is the division operator, which means rounding down the division result to the nearest integer. step_size indicates the adjustment interval, that is, how many epochs to adjust the learning rate.
Experimental results and analysis
Selection of the backbone network
Classic classification networks based on CNN architecture include AlexNet32, ResNet5033, ShuffleNet V234, MobileNetV235 and DenseNet121. With the development of deep learning, the performance of AlexNet is gradually surpassed by more complex network, which makes it less frequently adopted in tasks with high precision requirements. ShuffleNet V2 and MobileNetV2 are suitable for resource constrained environments due to their lower requirements for computing and memory. ResNet50 effectively alleviates the problems of gradient vanishing and exploding in deep networks by introducing residual connections, which can support deeper structures and is appropriate for complex feature learning tasks. The advantage of DenseNet121 lies in its dense connectivity strategy, which ensures that the output of each layer is connected to the outputs of all previous layers. The strategy significantly improves the efficiency of information flow and gradient propagation, making DenseNet121 particularly suitable for feature extraction in image classification tasks.
In order to select the most suitable network for cervical cell classification, we conduct the backbone network selection experiment of seven classifications on the Herlev dataset. The performance indexes of each network are shown in Table 3. The data in the table show that DenseNet121 has the highest accuracy, and the amount of the floating-point operations (FLOPs) and the number of parameters (Params) are both moderate. In contrast, ShuffleNet V2 and MobileNetV2 have inferior accuracy, although they are lower in FLOPs and Params. Therefore, considering the classification accuracy and computational efficiency, DenseNet121 is chosen as the backbone network.
Table 3.
Evaluation metrics of seven-classification task on the Herlev dataset for five classic networks.
| Network | Acc/% | Pre/% | Spe/% | Rec/% | F1-score/% | FLOPs/G | Params/M |
|---|---|---|---|---|---|---|---|
| AlexNet | 59.46 | 64.81 | 93.10 | 65.66 | 62.31 | 0.71 | 57.03 |
| ResNet50 | 77.84 | 82.54 | 96.09 | 79.70 | 80.39 | 4.13 | 23.52 |
| ShuffleNet V2 | 72.79 | 77.87 | 95.26 | 76.02 | 76.74 | 0.44 | 0.35 |
| MobileNetV2 | 78.02 | 81.19 | 96.18 | 80.04 | 80.35 | 0.33 | 2.23 |
| DenseNet121 | 85.33 | 85.23 | 97.55 | 85.26 | 85.11 | 2.90 | 6.96 |
Ablation experiment
To verify the effectiveness of each component in the network model A2SDNet121 proposed in this paper, we implement ablation experiments. DenseNet121 is used as the baseline. Then adds the SE module to DenseNet121,which is named “SDNet121”. Reduces the sizes of the convolutional layer and the pooling layer in the Stem layer of the SDNet121 model, and obtains the model “2SDNet121”. Finally, introduces the atrous convolution to 2SDNet121 model by constructing ADB module, and obtains the proposed model “A2SDNet121”. The above four models are tested on the Herlev dataset for seven-classification task, and the evaluation metrics are shown in Fig. 10.
Fig. 10.
Evaluation metrics of seven-classification task on the Herlev dataset for four classification models.
Observing the values of the indicators in Fig. 10, it can be seen that the accuracy of the baseline network DenseNet121 is only 85.33%. After incorporating the SE module, the accuracy of SDNet121 is increased by 12.47%, which is a significant improvement. On this basis, after reducing the convolutional kernel size and pooling window size of the Stem layer, the accuracy of the model is improved by 0.73%. After adding the ADB module, the accuracy is increased by another 0.74%, and the final model A2SDNet121 reaches an accuracy of 99.27%. By analyzing the data in the figure, the following conclusions can be drawn that the embedding of the SE module can significantly enhance the feature utilization rate, the adjustment of the Stem layer can obtain more local features, and the ADB module based on atrous convolution can obtain more multi-scale features. These improvements effectively enhance the classification performance of the model.
Visual analysis
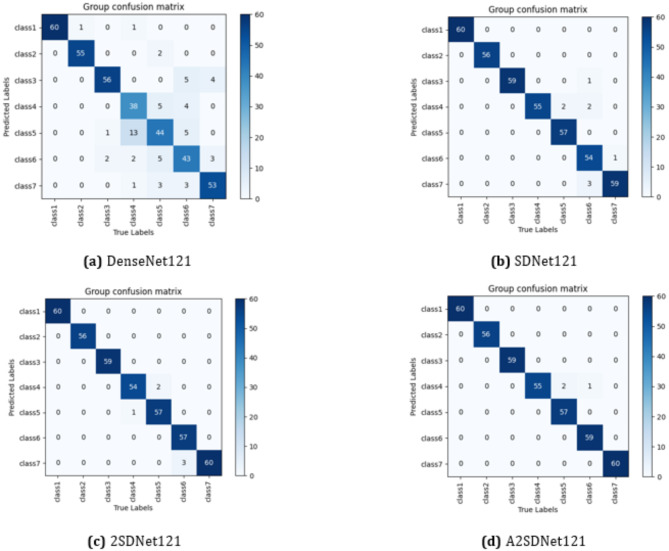
The confusion matrices of the above four models for the seven-classification task on the Herlev dataset are shown in Fig. 11. In Fig. 11(a), the DenseNet121 correctly identifies 349 images and incorrectly identifies 60 images. In Fig. 11(b), the SDNet121 model correctly identifies 400 images and incorrectly identifies 9 images. In Fig. 11(c), the 2SDNet121 model correctly identifies 403 images and incorrectly identifies 6 images. In Fig. 11(d), the A2SDNet121 model has 406 correctly recognized images and 3 incorrectly recognized images.
Fig. 11.
Confusion matrices of the four models for seven-classification task on the Herlev dataset.
We also conduct binary classification experiments on the Herlev dataset using the A2SDNet121 model. The evaluation metrics are shown in Fig. 12, and the confusion matrix in Fig. 13. As can be seen from Figs. 12 and 13, the accuracy of binary classification reaches 99.75%, and only one normal cell is incorrectly identified. The data fully prove the excellent performance of the proposed model.
Fig. 12.
Evaluation metrics of A2SDNet121 for two-classification task on the Herlev dataset.
Fig. 13.
Confusion matrix of A2SDNet121 for two-classification task on the Herlev dataset.
Effectiveness of the SE module in SDNet121
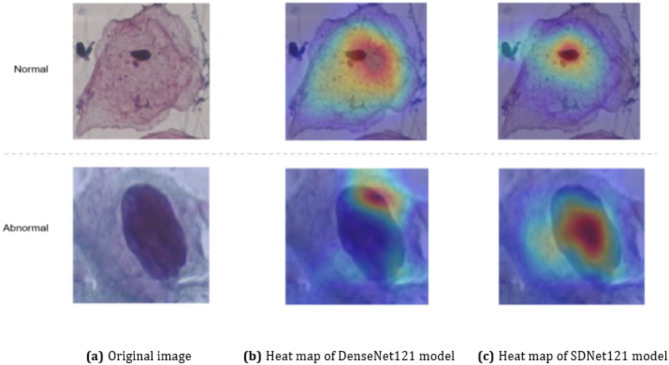
The ablation experiments have shown that the SE module enables the SDNet121 network to significantly improve all the evaluation metrics compared with the baseline network. In order to further validate the effectiveness and enhance the interpretability of the SDNet121 network, we adopt the technique of Gradient-weighted Class Activation Mapping (Grad-CAM)36 technique to compare the feature extraction ability of the baseline model DenseNet121 with that of SDNet121. The results of the visualized heat map are shown in Fig. 14. The three images in the first row of Fig. 14 are the normal cervical cell image and its visualized heat maps of the two models, while the second row are the abnormal cell image and its visualized heat maps. When making predictions for cervical cells, the SDNet121 model pays more attention to key regions (nuclei) with diagnostic significance compared to DenseNet121, and considerably less attention to redundant information. This suggests that the SE module enables SDNet121 to significantly improve its ability to identify and locate key regions, which makes it more suitable for the task of cervical cell classification.
Fig. 14.
Visualized heat maps based on Grad-CAM.
Validation of model stability
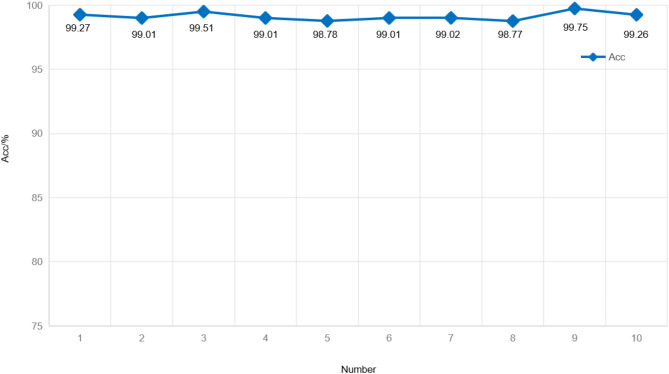
In order to verify the stability of the proposed A2SDNet121 model, we employ the ten-fold cross-validation to implement seven-classification experiment on the Herlev dataset. The dataset is divided into ten equal parts, and one of which is selected as the test set and the rest as the training set for each training. The test set is chosen without repetition to ensure that all images in the dataset have appeared in the test set after ten rounds of training. Figure 15 shows the curve of the classification accuracy of A2SDNet121 in ten-fold cross-validation. The data in the graph shows that the accuracy of the A2SDNet121 model fluctuates from 98.77 to 99.75%, with an average accuracy of 99.14%. The model demonstrates strong stability.
Fig. 15.
Accuracy curve of ten-fold cross-validation on the Herlev dataset for A2SDNet121.
Validation of model generalization
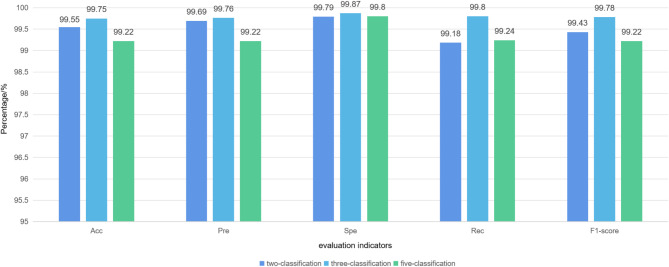
To verify the generalization performance of the A2SDNet121 model, two-, three- and five-classification experiments are carried out on the SIPaKMeD dataset. The evaluation metrics of A2SDNet121 for each classification task are shown in Fig. 16. The accuracy of A2SDNet121 reaches 99.55%, 99.75%, and 99.22% for two-, three- and five-classification on the SIPaKMeD dataset, respectively. This indicates that the A2SDNet121 model has a strong ability to predict unknown data.
Fig. 16.
Evaluation metrics of A2SDNet121for two-, three- and five-classification tasks on the SIPaKMeD dataset.
Comparison with other methods
To analyze the advantages of the proposed A2SDNet121 model for classification tasks, we compare it with some current excellent methods.
Table 4 shows the performance comparison data of A2SDNet121 and comparison methods in seven-classification task on the Herlev dataset. By comparing the evaluation indicators in Table, it can be found that only the precision value of the A2SDNet121 model is 0.03% lower than that of the DSRNet50 model. While the accuracy, specificity, recall, and F1-score of A2SDNet121 are all higher than other methods. Especially in terms of accuracy, it has improved by 0.24% compared to the second best method DSRNet50. This proves the superiority of the A2SDNet121 model in classification performance.
Table 4.
Evaluation metrics of A2SDNet121 with other state-of-the-art methods for seven-classification task on the Herlev dataset.
| Method | Acc/% | Pre/% | Spe/% | Rec/% | F1-score/% |
|---|---|---|---|---|---|
| HDFF23 | 90.32 | 91.50 | \ | 91.10 | 91.60 |
| DSRNet5024 | 98.90 | 99.20 | \ | 98.80 | 99.00 |
| Lin20 | 97.60 | \ | 97.00 | 98.00 | \ |
| IDCNN-CDC15 | 97.96 | 97.84 | \ | 98.20 | 97.81 |
| Chowdary26 | 98.39 | \ | 97.65 | 98.97 | \ |
| Hao27 | 92.30 | 92.30 | \ | 92.30 | 92.30 |
| Wu28 | 94.20 | \ | \ | 94.00 | 94.10 |
| VTCNet37 | 95.95 | 96.18 | \ | 95.95 | 95.97 |
| Fang38 | 94.55 | 94.13 | \ | 91.66 | 92.78 |
| A2SDNet121 | 99.14 | 99.17 | 99.86 | 99.14 | 99.15 |
Table 5 shows the performance comparison data of A2SDNet121 and comparison methods in five-classification task on the SIPaKMeD dataset. The data show that only the recall metric of the A2SDNet121 model is 0.56% lower than Shi’s, and the other metrics are higher than those of the comparison methods. The experimental data indicate that the comprehensive performance of A2SDNet121 is superior in the cervical cell multi-classification task.
Table 5.
Evaluation metrics of A2SDNet121 with other state-of-the-art methods for five-classification task on the SIPaKMeD dataset.
| Method | Acc/% | Pre/% | Spe/% | Rec/% | F1-score/% |
|---|---|---|---|---|---|
| HDFF23 | 99.14 | 99.20 | \ | \ | 99.00 |
| Shi13 | 98.37 | \ | 99.60 | 99.80 | \ |
| Qin25 | 98.67 | \ | 99.67 | 98.65 | 98.67 |
| Kundu16 | 98.94 | 98.79 | \ | 98.80 | 98.79 |
| Gao21 | 98.52 | \ | 98.68 | 98.53 | 98.59 |
| Chowdary26 | 99.16 | \ | 99.75 | 99.15 | \ |
| Das17 | 97.93 | 98.00 | \ | 97.00 | 98.00 |
| VTCNet37 | 95.95 | 96.18 | \ | 95.95 | 95.97 |
| Fang38 | 94.55 | 94.13 | \ | 91.66 | 92.78 |
| Hemalatha39 | 98.13 | 97.40 | \ | 97.60 | 97.40 |
| RES_DCGAN40 | 92.00 | 92.00 | \ | 92.00 | 92.00 |
| Shandilya41 | 99.11 | 98.69 | \ | 98.63 | 98.47 |
| A2SDNet121 | 99.22 | 99.22 | 99.80 | 99.24 | 99.22 |
Discussion
The current deep learning models for cervical cell classification tasks are not strong enough to capture the local detailed features and multi-scale information contained in cell images. To solve this problem, The A2SDNet121 model is proposed which is improved on the structure of DenseNet121 and more suitable for the features of cervical cell images.
The accuracy, precision, specificity, recall, and F1-score of A2SDNet121 for the seven-classification task on the Herlev dataset achieves 99.14%, 99.17%, 99.86%, 99.14% and 99.15%, respectively. The experimental data of ten-fold cross-validation indicate the model’s stability and reliability.
The five metrics of A2SDNet121 for the five-classification task on the SIPaKMeD dataset achieves 99.22%, 99.22%, 99.80%, 99.24% and 99.22%, respectively. The experimental data of cross-dataset validation demonstrate that the model can still maintain superior classification performance for unknown data.
In addition, we employ the visualization heat map based on the Grad-CAM technique to explain the decision-making process of A2SDNet121. The results show that the SE module can significantly improve the model’s ability to locate the nucleus region in cervical cell images, laying the foundation for accurate extraction of local and multi-scale features.
Although the quantitative and qualitative evaluation results indicate that A2SDNet121 has superior comprehensive performance, there are still some limitations requiring further research. First, the model needs to be generalized to the field of classification involving overlapping cells39. Second, in the case of high cost of labeled samples, the idea of semi-supervised learning42 can be borrowed to enhance the classification performance of the model by combining medical imaging technology43,44 and generative adversarial networks45,46 to generate high-quality pseudo-annotated samples. Finally, in order to comprehensively evaluate the performance of the model in real-world scenarios and enhance its credibility, we will introduce clinical data from partner hospitals in the future to further validate the practicality of the model.
Conclusion
In order to improve the accuracy and efficiency of the cervical cell multi-classification tasks, a deep convolutional neural network called A2SDNet121 is proposed in this paper. A2SDNet121 utilizes the SE module to rearrange the feature channels of the feature maps, improving the model’s focus on the nucleus region of cervical cells. The finer-grained local features of the cell images are extracted through the strategy of adjusting the size of convolutional kernels and the pooling window of the Stem layer. The global and local features are fused by the constructed ADB module. Experiments on multiple classification tasks on the Herlev dataset and the SIPaKMeD dataset have shown that the A2SDNet121 model performs well in the cervical cell multi-classification tasks, and can be used for large-scale pre-cancer screening of cervical cancer, which is promising for clinical applications.
Acknowledgements
The authors would like to thank Herlev University Hospital and the Technical University of Denmark for providing the source images from the Herlev dataset. We also express our gratitude to the SIPaKMeD dataset(https://www.cs.uoi.gr/~marina/sipakmed.html) for providing the source images. This work was supported in part by the Science and Technology Development Project of Jilin Province, China, under Grant 20220101123JC.
Author contributions
Y.Z.: Data curation, software, methodology, validation, visualization, writing-original draft, writing review, and editing. C.Y.N.: Conceptualization, Formal analysis, methodology, writing-review and editing.W.J.Y.: Investigation, visualization, validation.
Data availability
The data that support the findings of this study are available from the corresponding author, upon reasonable request.
Declarations
Competing interests
The authors declare no competing interests.
Footnotes
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
- 1.Bray, F. et al. Global Cancer statistics 2022: GLOBOCAN estimates of incidence and Mortality Worldwide for 36 cancers in 185 countries. Cancer J. Clin.74, 229–263. 10.3322/CAAC.21834 (2024). [DOI] [PubMed] [Google Scholar]
- 2.Nayar, R. & Wilbur, D. C. The pap test and Bethesda 2014. J. Am. Soc. Cytopathol.4, 170–180. 10.1016/j.jasc.2015.03.003 (2015). [DOI] [PubMed] [Google Scholar]
- 3.Kitchener, H. C. et al. Automation-assisted versus manual reading of cervical cytology (MAVARIC): a randomised controlled trial. Lancet Oncol.12, 56–64. 10.1016/s1470-2045(10)70264-3 (2011). [DOI] [PubMed] [Google Scholar]
- 4.Zhang, L. et al. Automation-assisted cervical cancer screening in manual liquid‐based cytology with hematoxylin and eosin staining. Cytometry Part. A. 85, 214–230. 10.1002/cyto.a.22407 (2013). [DOI] [PubMed] [Google Scholar]
- 5.William, W., Ware, A., Basaza-Ejiri, A. H. & Obungoloch, J. A review of image analysis and machine learning techniques for automated cervical cancer screening from pap-smear images. Comput. Methods Programs Biomed.164, 15–22. 10.1016/j.cmpb.2018.05.034 (2018). [DOI] [PubMed] [Google Scholar]
- 6.Huang, G., Liu, Z., van der Maaten, L. & Weinberger, K. Q. Densely connected convolutional networks. Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition. 4700–4708 (2017).
- 7.Hu, J., Shen, L. & Sun, G. Squeeze-and-Excitation Networks. roceedings of the IEEE Conference on Computer Vision and Pattern Recognition. 7132–7141, (2018).
- 8.Yu, F. & Koltum, V. Multi-Scale Context Aggregation by Dilated Convolutions. CoRR. (2015).
- 9.Zhang, L. et al. DeepPap: deep convolutional networks for cervical cell classification. IEEE J. Biomedical Health Inf.21, 1633–1643. 10.1109/jbhi.2017.2705583 (2017). [DOI] [PubMed] [Google Scholar]
- 10.Promworn, Y., Pattanasak, S., Pintavirooj, C. & Piyawattanametha, W. Comparisons of Pap smear classification with deep learning models. in: IEEE 14th International Conference on Nano/Micro Engineered and Molecular Systems (NEMS). 282–285 (2019). (2019).
- 11.Talo, M. Diagnostic classification of cervical cell images from pap smear slides. Acad. Perspective Procedia. 2, 1043–1050. 10.33793/acperpro.02.03.116 (2019). [Google Scholar]
- 12.Hussain, E., Mahanta, L. B., Das, C. R., Choudhury, M. & Chowdhury, M. A shape context fully convolutional neural network for segmentation and classification of cervical nuclei in pap smear images. Artif. Intell. Med.10710.1016/j.artmed.2020.101897 (2020). [DOI] [PubMed]
- 13.Shi, J. et al. Cervical cell classification with graph convolutional network. Comput. Methods Programs Biomed.19810.1016/j.cmpb.2020.105807 (2021). [DOI] [PubMed]
- 14.Chen, W., Gao, L., Li, X. & Shen, W. Lightweight convolutional neural network with knowledge distillation for cervical cells classification. Biomed. Signal Process. Control. 7110.1016/j.bspc.2021.103177 (2022).
- 15.Ibrahim Waly, M., Yacin Sikkandar, M., Abdelkader Aboamer, M., Kadry, S. & Thinnukool, O. Optimal deep convolution neural network for cervical Cancer diagnosis model. Computers Mater. Continua. 70, 3295–3309. 10.32604/cmc.2022.020713 (2022). [Google Scholar]
- 16.Kundu, R. & Chattopadhyay, S. Deep features selection through genetic algorithm for cervical pre-cancerous cell classification. Multimedia Tools Appl.82, 13431–13452. 10.1007/s11042-022-13736-9 (2022). [Google Scholar]
- 17.Das, N., Mandal, B., Santosh, K. C., Shen, L. & Chakraborty, S. Cervical cancerous cell classification: opposition-based harmony search for deep feature selection. Int. J. Mach. Learn. Cybernet.14, 3911–3922. 10.1007/s13042-023-01872-z (2023). [Google Scholar]
- 18.Xu, L., Cai, F., Fu, Y. & Liu, Q. Cervical cell classification with deep-learning algorithms. Med. Biol. Eng. Comput.61, 821–833. 10.1007/s11517-022-02745-3 (2023). [DOI] [PubMed] [Google Scholar]
- 19.Chen, W., Li, X., Gao, L. & Shen, W. Improving computer-aided cervical cells classification using transfer learning based Snapshot Ensemble. Appl. Sci.1010.3390/app10207292 (2020).
- 20.Lin, Z. et al. Classification of cervical cells leveraging simultaneous super-resolution and ordinal regression. Appl. Soft Comput.11510.1016/j.asoc.2021.108208 (2022).
- 21.Gao, W. et al. Cervical cell image classification-based knowledge distillation. Biomimetics710.3390/biomimetics7040195 (2022). [DOI] [PMC free article] [PubMed]
- 22.Song, Y., Zou, J., Choi, K. S., Lei, B. & Qin, J. Cell classification with worse-case boosting for intelligent cervical cancer screening. Med. Image. Anal. 91. 10.1016/j.media.2023.103014 (2024). [DOI] [PubMed]
- 23.Rahaman, M. M. et al. A deep learning-based framework for the classification of cervical cells using hybrid deep feature fusion techniques. Comput. Biol. Med.13610.1016/j.compbiomed.2021.104649 (2021). DeepCervix. [DOI] [PubMed]
- 24.Qin, M. et al. Efficient Cervical Cell Lesion Recognition Method Based on Dual Path Network. Wireless Communications and Mobile Computing. 1–11, (2022). 10.1155/2022/8496751 (2022).
- 25.Qin, J., He, Y., Ge, J., Liang, Y. A. & Multi-Task Feature Fusion Model for cervical cell classification. IEEE J. Biomedical Health Inf.26, 4668–4678. 10.1109/jbhi.2022.3180989 (2022). [DOI] [PubMed] [Google Scholar]
- 26.Chowdary, G. J. & Yogarajah, P. G, S., M, P. Nucleus segmentation and classification using residual SE-UNet and feature concatenation approach incervical cytopathology cell images. Technology in Cancer Research & Treatment. 22, (2023). 10.1177/15330338221134833 [DOI] [PMC free article] [PubMed]
- 27.Hao, X. et al. Cervical cell deep-learning automatic classification method based on fusion features. Multimedia Tools Appl.82, 33183–33202. 10.1007/s11042-023-14973-2 (2023). [Google Scholar]
- 28.Wu, N., Jia, D., Zhang, C. & Li, Z. Cervical cell classification based on strong feature CNN-LSVM network using Adaboost optimization. J. Intell. Fuzzy Syst.44, 4335–4355. 10.3233/jifs-221604 (2023). [Google Scholar]
- 29.Jantzen, J., Norup, J., Dounias, G. & Bjerregaard, B. Pap-smear benchmark data for pattern classification. Proceedings of the Conference on Nature inspired Smart Information Systems (NiSIS 1–9 (2005). (2005).
- 30.Plissiti, M. E. et al. in. 25th IEEE International Conference on Image Processing (ICIP). 3144–3148 (2018). (2018).
- 31.Wang, Y. et al. Automatic lung nodule detection using multi-scale dot nodule-enhancement filter and weighted support vector machines in chest computed tomography. Plos One. 1410.1371/journal.pone.0210551 (2019). [DOI] [PMC free article] [PubMed]
- 32.Krizhevsky, A., Sutskever, I. & Hinton, E. G. ImageNet classification with deep convolutional neural networks. Commun. ACM. 60, 84–90 (2017). [Google Scholar]
- 33.He, K., Zhang, X., Ren, S. & Sun, J. Deep Residual Learning for Image Recognition IEEE Conference on Computer Vision and Pattern Recognition (CVPR). (2016). (2016).
- 34.Ma, N., Zhang, X., Zheng, H. T. & Sun, J. ShuffleNet V2: practical guidelines for efficient CNN architecture design. Proceedings of the European Conference on Computer Vision (ECCV). 116–131, (2018).
- 35.Sandler, M., Howard, A., Zhu, M., Zhmoginov, A. & Chen, L. C. in IEEE/CVF Conference on Computer Vision and Pattern Recognition 4510–4520 (2018). (2018).
- 36.Selvaraju, R. R. et al. Grad-CAM:visual explanations from deep networks via gradient-based localization. Int. J. Comput. Vision. 128, 336–359 (2020). [Google Scholar]
- 37.Li, M. et al. A feature Fusion DL Model based on CNN and ViT for the classification of cervical cells. Int. J. Imaging Syst. Technol.3410.1002/ima.23161 (2024).
- 38.Fang, M., Fu, M., Liao, B., Lei, X. & Wu, F. X. Deep integrated fusion of local and global features for cervical cell classification. Comput. Biol. Med.17110.1016/j.compbiomed.2024.108153 (2024). [DOI] [PubMed]
- 39.K, H. & Dhandapani, V. V. A, A. G. CervixFuzzyFusion for cervical cancer cell image classification. Biomed. Signal Process. Control. 8510.1016/j.bspc.2023.104920 (2023).
- 40.Wubineh, B. Z., Rusiecki, A. & Halawa, K. Classification of cervical cells from the pap smear image using the RES_DCGAN data augmentation and ResNet50V2 with self-attention architecture. Neural Comput. Appl.36, 21801–21815. 10.1007/s00521-024-10404-x (2024). [Google Scholar]
- 41.Shandilya, G. et al. Enhancing advanced cervical cell categorization with cluster-based intelligent systems by a novel integrated CNN approach with skip mechanisms and GAN-based augmentation. Sci. Rep.1410.1038/s41598-024-80260-1 (2024). [DOI] [PMC free article] [PubMed]
- 42.Zhao, S., He, Y., Qin, J., Wang, Z. & Urzua, U. A. Semi-supervised Deep Learning Method for Cervical Cell Classification. Analytical Cellular Pathology. 1–12, (2022). 10.1155/2022/4376178 (2022). [DOI] [PMC free article] [PubMed]
- 43.Han, X. et al. The applications of magnetic particle imaging: from cell to body. Diagnostics1010.3390/diagnostics10100800 (2020). [DOI] [PMC free article] [PubMed]
- 44.Zou, Y. et al. Precision matters: the value of PET/CT and PET/MRI in the clinical management of cervical cancer. Strahlenther. Onkol.10.1007/s00066-024-02294-8 (2024). [DOI] [PubMed] [Google Scholar]
- 45.Xu, H., Li, Q. & Chen, J. Highlight removal from a single Grayscale Image using attentive GAN. Appl. Artif. Intell.3610.1080/08839514.2021.1988441 (2022).
- 46.Jia, Y., Chen, G. & Chi, H. Retinal fundus image super-resolution based on generative adversarial network guided with vascular structure prior. Sci. Rep.1410.1038/s41598-024-74186-x (2024). [DOI] [PMC free article] [PubMed]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
The data that support the findings of this study are available from the corresponding author, upon reasonable request.