Abstract
Crop improvement by genetic modification remains controversial, one of the major issues being the potential for unintended effects. Comparative safety assessment includes targeted analysis of key nutrients and antinutritional factors, but broader scale-profiling or “omics” methods could increase the chances of detecting unintended effects. Comparative assessment should consider the extent of natural variation and not simply compare genetically modified (GM) lines and parental controls. In this study, potato (Solanum tuberosum) proteome diversity has been assessed using a range of diverse non-GM germplasm. In addition, a selection of GM potato lines was compared to assess the potential for unintended differences in protein profiles. Clear qualitative and quantitative differences were found in the protein patterns of the varieties and landraces examined, with 1,077 of 1,111 protein spots analyzed showing statistically significant differences. The diploid species Solanum phureja could be clearly differentiated from tetraploid (Solanum tuberosum) genotypes. Many of the proteins apparently contributing to genotype differentiation are involved in disease and defense responses, the glycolytic pathway, and sugar metabolism or protein targeting/storage. Only nine proteins out of 730 showed significant differences between GM lines and their controls. There was much less variation between GM lines and their non-GM controls compared with that found between different varieties and landraces. A number of proteins were identified by mass spectrometry and added to a potato tuber two-dimensional protein map.
Potato (Solanum tuberosum) is the fourth most important food crop consumed worldwide. Although the potato gene pool used by European breeders has for historical reasons been limited, a large number of very different cultivars have been developed (Bradshaw and Mackay, 1994). While most cultivated potato varieties are of the tetraploid species Solanum tuberosum, many varieties have had genes for desired characteristics, such as disease and pest resistance, introgressed from wild and cultivated Solanum species, which provide a large source of genetic diversity for breeding purposes.
For vegetatively propagated crops with complex genetics, such as the potato, genetic modification for crop improvement offers significant potential. Within Europe, at least, genetic modification still remains controversial. A major concern is the possibility of unintended effects caused, for example, by the site of transgene integration (e.g. interruption of important open reading frames or regulatory sequences), which could result in modified metabolism, novel fusion proteins, or other pleiotropic effects that could compromise safety (Kuiper et al., 2001; Cellini et al., 2004). This includes the production of new allergens or toxins. Arguably, unintended effects are less likely to be detected with conventional targeted analysis of a relatively limited number of molecules (Millstone et al., 1999) compared with nontargeted methods such as transcriptional, protein, and metabolite profiling. While transcriptomics provides, at least for specific plant species, the most complete coverage of potential unintended effects (Meyers et al., 2004), whole genome arrays are presently not available for all important food crops. As the levels of transcripts and proteins do not always correlate (Anderson and Seilhamer, 1997; Gygi et al., 1999), reliance on any one profiling technology for safety assessment is questionable.
Using two-dimensional electrophoresis (2-DE) and mass spectrometry (MS), it is often possible to visualize, quantify, and identify hundreds or even thousands of proteins in a given tissue or cell sample, and proteome analysis is increasingly used in functional plant studies (Canovas et al., 2004). Proteomics has the potential to quantify the levels of allergens, most of which are proteins, and detect possible posttranslational modifications. Several studies have demonstrated the capacity of 2-DE to characterize and distinguish varieties and genotypes and even to identify single mutations with multiple effects (for review, see de Vienne et al., 2001; Thiellement et al., 2002; Canovas et al., 2004).
There are few reports of total potato tuber proteins resolved by 2-DE and very little information on the extent of natural variation in the proteome caused by genetic background, environmental influences, and other factors. Consideration of the extent of natural variation in the proteome is obviously important in the comparative analysis of genetically modified (GM) crops because observed differences in GM lines might be random fluctuations or well within the normal variation observed in nonmodified material.
The aim of this work is to provide insight into the extent of variation in the potato tuber proteome by analyzing a number of potato genotypes. In addition, several previously characterized GM potato lines were studied for possible unintended effects.
RESULTS
2-DE Reference Map for Potato Tubers
The 2-DE gels of potato tuber proteins contained between 500 to 1,200 spots, depending on the genotype, with some variation in the number of spots between individual samples. A total of 77 proteins were tentatively identified by HPLC electrospray tandem mass spectrometry (MS/MS) using cultivar Desirée as the model genotype, and these were included in a 2-DE reference map (Table I; Fig. 1). In many cases, the identification was based on homologous proteins in other plant species, such as tomato (Lycopersicon esculentum) or Arabidopsis (Arabidopsis thaliana). Functions for the identified proteins were categorized as previously described by Bevan et al. (1998). Most of the identified proteins were present in relatively high amounts on 2-DE gels, and many of them were classified into functional groups, such as energy metabolism, protein destination, and storage or disease/defense responses.
Table I.
Potato tuber proteins identified from cultivar Desirée
The assigned spots are shown in Figure 1. The accession number/EST refers to database accession numbers of matching proteins or ESTs; the full peptide sequences are shown where the ESTs are not publicly available. The number of matched peptides and the functional group are also indicated. The first group includes proteins with statistically significant differences between GM lines or controls; the second group includes proteins that showed significant differences between some of the non-GM genotypes and had high loadings in the second, third, or fourth principal component; the third group includes other identified proteins. Where two or more proteins were identified in the same spot, all are listed.
| Spot | Identification | Accession No./EST | Peptides | Functional Group |
|---|---|---|---|---|
| Proteins with Significant Differences between the Non-GM Varieties and Landraces | ||||
| Second Component | ||||
| 4 | Thioredoxin peroxidase/peroxiredoxin | AF442385, AAP34571 | 6 | Disease/defense |
| 55 | RAN1, GTP binding nuclear protein | NP_197501, T15nr049 | 1 | Intracellular traffic |
| 56 | Cathepsin D inhibitor | AAG12337 | 1 | Disease/defense |
| 57 | Putative Gln amidotransferase/cyclase | BAC43102 | 1 | Metabolism |
| 58 | Expressed protein | NP_179517 | 2 | Unclassified |
| 59 | Patatin; Dehydrin homolog C17 | P15478, PSTFS38TH, PSEAF25TH; T07779 | 9; 3 | Protein destination and storage; disease/defense |
| 60 | EST (peptide sequences LGSHFVSENQDVSIK VAYSIVGPTHSPLR FSTSSSSTK YETGRPHSYK YETGRPHSYKLR IEKYETGRPHSYKLR) | T04nr049 | 6 | Unclassified |
| Third Component | ||||
| 6 | Triosephosphate isomerase, cytosolic | P12863, P48494 | 4 | Energy, metabolism |
| 14 | Enolase | P26300, AAL06912 | 9 | Energy |
| 15 | UTP-Glc-1-P uridylyltransferase | P19595, AAB71613 | 13 | Metabolism |
| 20 | Enolase | P26300, Q42971 | 13 | Energy |
| 43 | Enolase | JQ1186, P26300 | 8 | Energy |
| 44 | Enolase | P26300, JQ1186 | 10 | Energy |
| 45 | UTP-Glc-1-P uridylyltransferase | P19595 | 16 | Metabolism |
| 61 | 26S protease regulatory subunit 6A homolog (TBP-1) | P54776 | 12 | Protein destination and storage |
| 62 | Fru-bisP aldolase, cytoplasmic isozyme 2; Patatin; Phosphoenolpyruvate carboxylase kinase | P46257; P15478; AAF19403 | 1; 2; 1 | Energy; protein destination and storage; signal transduction |
| 63 | Ascorbate peroxidase | BAC22953, CAB58361, NP_195321 | 4 | Disease/defense |
| 64 | Enolase | JQ1186, P26300 | 8 | Energy |
| 65 | Asp-semialdehyde dehydrogenase precursor; Putative protein disulfide isomerase | AAG33078; BAB67990 | 1; 5 | Metabolism; protein destination and storage |
| 66 | Enolase | JQ1186, P26300 | 5 | Energy |
| 67 | Ascorbate peroxidase | CAB58361, BAC22953, NP_195321, PSHBN28TH | 6 | Disease/defense |
| Fourth Component | ||||
| 47 | Phytepsin precursor (aspartic proteinase) | P42210, PSTDE14TH | 4 | Protein destination and storage |
| 68 | Expressed protein/hypothetical protein/unknown protein | NP_568098, T02532, AAO18441, PSHBQ20TH | 3 | Unclassified |
| 69 | Patatin | P15478 | 3 | Protein destination and storage |
| 70 | Kunitz-type enzyme inhibitor S0C11 | AAL67830, T04nr016 | Disease/defense | |
| 71 | EST | PSTGU28TH | 2 | Unclassified |
| 72 | Aspartic proteinase | AAC49730, PSTES05TH, PSHEN78TH | 5 | Protein destination and storage |
| Proteins with Significant Differences between the GM Lines | ||||
| 54 | Aspartic protease | AAT77954 | 3 | Protein destination and storage |
| 70 | Kunitz-type enzyme inhibitor S0C11 | AAL67830, T04nr016 | 2 | Disease/defense |
| 73 | Patatin | T07592 | 2 | Protein destination and storage |
| 74 | Hsp20.1 protein | CAA12387, 4H3p01nr | 8 | Disease/defense |
| 76 | Chaperonin 21 precursor | AAF6029 | 7 | Protein destination and storage |
| 77 | EST (peptide sequences LAIGTGNTFNLVFIK YLDTNGEAGHVK NVASWFQIK KTGSYMYK) | T02nr073 | 4 | Unclassified |
| 78 | Unknown protein | NP_908355 | 2 | Unknown |
| 75, 79 | Unidentified | |||
| Other Identified Proteins | ||||
| 1 | Glc-3-P dehydrogenase, cytosolic | P26519, S72667 | 2 | Energy |
| 2 | Pathogenesis-related protein STH-2 | P17642 | 1 | Disease/defense |
| 3 | Kunitz-type enzyme inhibitor S0C11 | AAL67830, AF460237 | 4 | Disease/defense |
| 5 | Superoxide dismutase [Cu-Zn] | P27082, P14830 | 2 | Disease/defense |
| 7 | Malate dehydrogenase, cytosolic; Glc-3-P dehydrogenase | CAC10208; AAB54003 | 1; 1 | Energy |
| 8 | Dihydrolipoamide dehydrogenase precursor | AF295339 | 1 | Energy |
| 9 | Putative malate dehydrogenase | NP_171936 | 1 | Energy |
| 10 | Proteasome subunit α-type 6 | Q9XG77 | 2 | Protein destination and storage |
| 11 | Transcription factor homolog BTF3 | T16984 | 1 | Transcription |
| 12 | Nucleoside-diphosphate kinase | S47974 | 1 | Metabolism |
| 13 | Heat shock protein 17.6 | T07602 | 1 | Disease/defense |
| 16 | UTP-Glc-1-P uridylyltransferase | AAB71613 | 1 | Metabolism |
| 17 | Kunitz-type enzyme inhibitor S9C11 | AAL67830 | 1 | Disease/defense |
| 18 | Triosephosphate isomerase precursor, chloroplast | P48496, Q9M4S8 | 3 | Energy, metabolism |
| 19 | Putative Ala aminotransferase | AAM61453 | 1 | Metabolism |
| 21 | β-Cyano-Ala synthase-like protein | BAB20032 | 1 | Metabolism |
| 22 | 26S proteasome, non-ATPase regulatory subunit (RPN11), putative | NP_197745 | 1 | Protein destination and storage |
| 23 | Cystatin | AAF23126 | 2 | Disease/defense |
| 24 | Phytepsin precursor (aspartic proteinase) | P42210 | 1 | Protein destination and storage |
| 25 | Probable malate dehydrogenase | T06386 | 1 | Energy |
| 26 | Patatin | P15478 | 1 | Protein destination and storage |
| 27 | Patatin; Phosphoenolpyruvate carboxykinase | P15478; AAF19403 | 1; 1 | Protein destination and storage; signal transduction |
| 28 | Patatin; Phosphoenolpyruvate carboxykinase | P15478; AAF19403 | 3; 1 | Protein destination and storage; signal transduction |
| 29 | Patatin | AAK56395 | 3 | Protein destination and storage |
| 30 | Fructokinase | P37829 | 4 | Metabolism |
| 31 | Fructokinase | AAB51108 | 3 | Metabolism |
| 32 | Putative lactoylglutathione lyase | AAL07227 | 1 | Disease/defense |
| 33 | Phytepsin precursor (aspartic proteinase) | P42210 | 1 | Protein destination and storage |
| 34 | Cys proteinase precursor | S55923 | 1 | Protein destination and storage |
| 35 | Heat shock protein 17.6 | T07602 | 1 | Disease/defense |
| 36 | ATP synthase β-chain precursor, mitochondrial | Q01859, AAD03393 | 7 | Unclear classification |
| 37 | UTP-Glc-1-P uridylyltransferase | S31431 | 4 | Metabolism |
| 38 | Actin | P30172, Q96493 | 3 | Cell structure |
| 39 | Putative nascent polypeptide-associated complex α-chain/expressed protein | BAB89723, NP_196889 | 3 | Protein destination and storage |
| 40 | Putative nascent polypeptide-associated complex α-chain/expressed protein | BAB89723, NP_196889 | 2 | Protein destination and storage |
| 41 | Ascorbate peroxidase | CAA72247, CAB58361 | 2 | Disease/defense |
| 42 | Ascorbate peroxidase | CAA72247, CAB58361 | 2 | Disease/defense |
| 46 | Annexin p34 | AAC97494 | 2 | Disease/defense |
| 48 | dnaK-type molecular chaperone hsc-2/heat shock protein 70 kD | S14950, CAB72129, PSHDH15TH | 4 | Protein destination and storage |
| 49 | Aminopeptidase 2 precursor, chloroplast | Q42876, PSEAS06TH, PSHCI03TH | 2 | Protein destination and storage |
| 50 | Patatin | AAK56395 | 1 | Protein destination and storage |
| 51 | Putative NAD-dependent malate dehydrogenase, mitochondrial | CAD33240 | 2 | Energy |
| 52 | Gly-rich RNA-binding protein GRP1 | CAA73034 | 1 | Transcription |
| 53 | Dihydrolipoamide dehydrogenase precursor | AAN23154 | 2 | Metabolism |
Figure 1.
2-DE reference map of potato tuber proteins in cultivar Desirée. Proteins identified in this study are marked with arrows and numbers that correlate with protein identifications in Table I.
Tuber Protein Profiles of Potato Varieties and Landraces
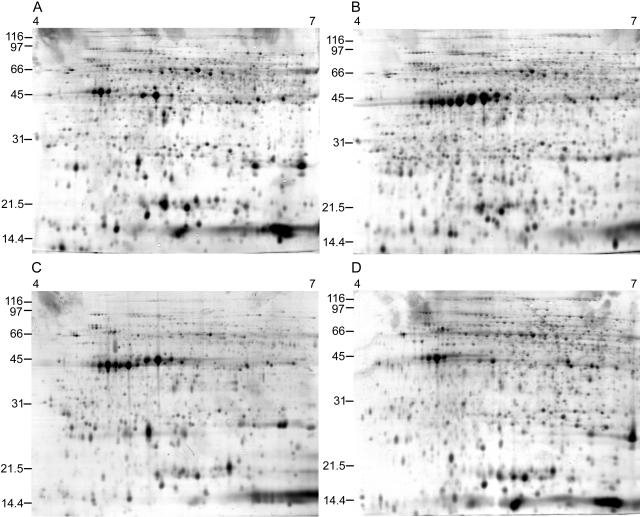
A total of 32 non-GM potato genotypes were selected to represent a range of genetic variation: 21 named cultivars of tetraploid potato, eight landraces, and three diploid genotypes, including accessions and named cultivars of Solanum phureja adapted for long-daylength conditions. Figure 2 shows 2-DEs for selected cultivars, landraces, and diploids. Between 800 to 1,200 polypeptides were detected, depending on genotype. A total of 1,932 polypeptides were detected when all spots in all genotypes were combined. There were clear qualitative and quantitative differences in polypeptide profiles between the genotypes (Fig. 2). One of the most obvious differences occurred in proteins with a Mr of approximately 40,000 to 45,000 and a pI of 4.5 to 5.5, which corresponds to various isoforms of patatin, the major storage protein of potato tuber (Pots et al., 1999; Schmidt et al., 2002); many genotypes showed distinctive patterns of putative patatin isoforms.
Figure 2.
2-DE images of tuber proteins of a range of potato genotypes. A, Cultivar Desirée; B, cultivar Maris Piper; C, landrace accession TBR3369 (1); D, S. phureja accession PHU4637 (1). Total potato tuber protein was run in 2-DE with isoelectric focusing (pH range 4–7) in the first dimension and 12% SDS-PAGE gels in the second dimension. Gels were stained with SYPRO Ruby.
To determine statistically significant differences in the quantities of individual protein spots in the different potato genotypes, ANOVA or the Kruskal-Wallis nonparametric test was employed as outlined in Table II. Analysis of individual proteins revealed that, for 1,077 out of 1,111 spots, the expression was significantly different among the genotypes. Interestingly, only 34 of the protein spots detected did not appear to differ significantly between genotypes. Among these proteins were those denoted as 6, 9, 18, 46, and 52 (Table I), i.e. proteins matched to two triosephosphate isomerases, a putative malate dehydrogenase, GRP1, and annexin p34. Proteins that were expressed in less than 27 of the 127 samples were not analyzed statistically because these proteins were rarely detected in all four replicates of the different potato genotypes. Genotype-specific proteins, i.e. spots expressed in only one or a few genotypes, were not examined any further; there were approximately 600 of these proteins.
Table II.
Statistical analysis of tuber protein profiles of potato genotypes
Protein spots were analyzed by ANOVA if they were detected in at least 88 of the 127 gels to satisfy the assumption of data normality. Spots were analyzed by the Kruskal-Wallis nonparametric test if they were detected in 27 to 87 of the gels. Spots that were analyzed by ANOVA were also subjected to PCA.
| No. Spots | No. Spots with Significant Differences between Genotypes (P < 0.01) | |
|---|---|---|
| Protein spots detected (in total including all genotypes) | 1,932 | |
| Spots analyzed in ANOVA and PCA | 393 | 363 |
| Spots analyzed by the Kruskal-Wallis test | 718 | 714 |
| Spots analyzed for differences between genotypes (in total) | 1,111 | 1,077 |
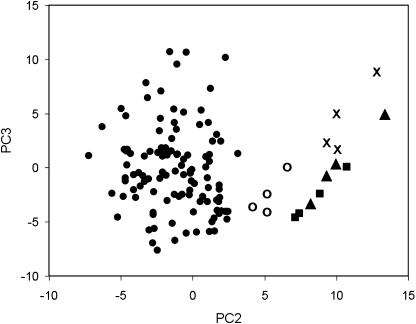
Principal component analysis (PCA) was assessed for its capacity to differentiate between genotypes based on protein quantities and to identify groups of proteins responsible for differentiating the genotypes. PCA was carried out on the 393 protein spots also analyzed by ANOVA (Table II) and was found to differentiate several genotypes (Fig. 3). For example, in the second and third components, the line TBR3302 (2) and the three S. phureja genotypes (accession PHU.4637 and named cultivars Inca Sun and Mayan Gold) separated from the tetraploid S. tuberosum-named varieties and landraces. In the fourth component, varieties Glenna, Morag, Maris Piper, and Pentland Javelin were slightly separated from the other varieties (data not shown), but the differences between named varieties were not as obvious. Nor were most landraces separated very clearly from other genotypes. Although total spot intensities were normalized across each gel to minimize possible subtle differences in protein loadings, staining, etc., the loadings for the first PCA component indicated that there were differences in the amount of protein or overall intensity, and thus it was not indicative of differences between the genotypes. Only proteins with a pI of 4 to 7 and a Mr of approximately 10,000 to 200,000 are visible on the gels in these conditions, provided they are soluble; therefore, the first component indicates that there are subtle differences in the amount of proteins detected within this range.
Figure 3.
Principal component scores for the tetraploid S. tuberosum-named varieties and landraces (•) and the landrace accession TBR3302 (2) (○), and for the diploid S. phureja accession PHU.4637 (X) and named cultivars Inca Sun (▴) and Mayan Gold (▪). Differences among the tetraploid S. tuberosum-named varieties and landraces were not equally clear.
Several of the proteins with very large positive or large negative loadings in PCA components two, three, and four were proteins that also differed among genotypes. These proteins were targeted for identification (Table I; Fig. 1). Among those contributing to the second component were several that were related to disease and defense responses. Most of the proteins with high loadings in the third component appeared to be involved in the glycolytic pathway, such as several enolases, triosephosphate isomerase, and Fru-bisP aldolase. This component also included two polypeptides with homologies to ascorbate peroxidases and UTP-Glc-1-P uridylyltransferases. Protein spots with low or high loadings in the fourth component included some proteinases, a Kunitz-type enzyme inhibitor, and some proteins or polypeptides matching expressed sequence tag (EST) sequences of currently unknown function.
Tuber Protein Profiles of GM Potato Lines
A range of GM potato lines was selected for proteomic analysis along with their appropriate controls. Qualitatively, the protein patterns on the 2-DE gels were similar across the lines (data not shown). ANOVA revealed significant differences (P < 0.01) in the quantities of seven protein spots (Fig. 1, spots 70 and 73–78) out of 240 analyzed (Table III). Analysis with the Kruskal-Wallis test also revealed that spot intensity was significantly different among the lines in two cases (Fig. 3, spots 54 and 79) out of 490. Seven of these proteins were identified by HPLC electrospray MS/MS (Table I; Fig. 3). These proteins included some defense-related proteins, as well as some proteins involved in protein destination and storage, and some proteins with unknown function. One of these spots was matched to a patatin protein; however, other spots that were previously identified as patatins did not appear to differ between lines. Also, other spots previously matched to Kunitz-type enzyme inhibitors and aspartic proteinases did not show significant differences among lines. Line-specific proteins, i.e. proteins expressed in all replicates of only one or few GM lines, were not found.
Table III.
Statistical analysis of tuber protein profiles of GM potato lines
Protein spots were analyzed by ANOVA if they were detected in at least 31 of the 40 gels to satisfy the assumption of data normality. Spots were analyzed by the Kruskal-Wallis nonparametric test if they were detected in 10 to 30 of the 40 gels. Spots that were analyzed by ANOVA were also subjected to PCA.
| No. Spots | No. Spots with Significant Differences between Lines (P < 0.01) | |
|---|---|---|
| Protein spots detected (in total including all lines) | 1,367 | |
| Spots analyzed in ANOVA and PCA | 240 | 7 |
| Spots analyzed by the Kruskal-Wallis test | 490 | 2 |
| Spots analyzed for differences between genotypes (in total) | 730 | 9 |
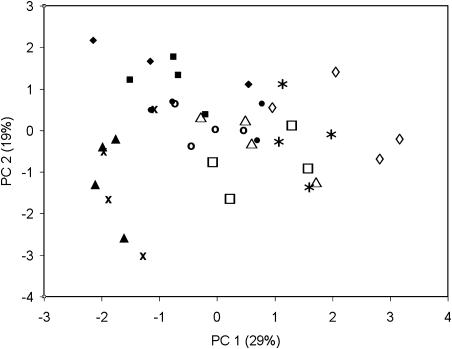
PCA was carried out on the 240 protein spots with at least 31 nonzero responses. No pronounced separation between the lines was observed in the plots of PCA scores (data not shown). PCA analysis was also carried out on protein sets where significant difference among potato lines was found by ANOVA. Again, there was no clear separation between the lines in the PCA plots (Fig. 4). The lines Mal1 2A and Mal1 5A were slightly separated from all the other lines in the first and second components, suggesting that these particular lines were primarily responsible for the seven protein spots identified as significantly different by ANOVA (Table III).
Figure 4.
Principal component scores for the GM lines and controls Desirée wild type (*), Desirée TC1 (□), SAM35S 3 (▵), SAM35S 1 (⋄), Mal1 2V1 (♦), Mal1 T41A (▪), Mal1 5A (▴), Mal1 2A (x), W2GBSS VO9 (○), and W2GBSS VO4 (•). Values in parentheses indicate the percentage of total variation accounted for each principal component.
Comparison of GM and Non-GM Potato Lines and Varieties
As the results indicated more differences between non-GM genotypes than between GM lines and their controls, some varieties and lines were selected for a second set of experiments. The GM line Sam35S 3, vector-only control W2 granule-bound starch synthase (GBSS) VO4, two wild-type control Desirée lines, the variety Maris Piper, and an accession of S. phureja (PHU.4637) were analyzed together, with three tubers of each genotype.
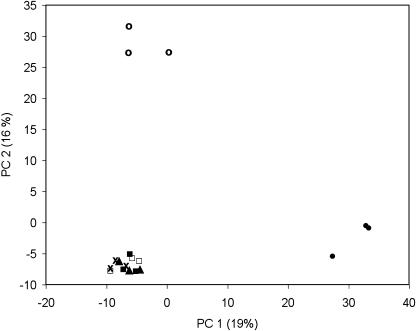
In the PCA plot (Fig. 5), the genotypes Maris Piper and S. phureja were clearly separated from each other and from all Desirée samples, whether GM or non-GM. No separation was observed between wild-type Desirée and transformed lines in the components (data not shown), again indicating that there was much less variation between Desirée and the GM lines than between the different nontransgenic varieties.
Figure 5.
Principal component scores for Desirée wild type (x and □), Sam35S 3 (▴), W2GBSS VO4 (▪), for the named variety Maris Piper (○), and S. phureja accession PHU.4637 (•). Values in parentheses indicate the percentage of total variation accounted for by each principal component.
DISCUSSION
The aim of this work is to gain insight into potato proteome diversity using a large selection of potato varieties and to assess the potential for 2-DE to detect significant changes in the proteome of transgenic potato.
It is clear that genotypic variation is extensive, with most of the proteins detected showing significant quantitative and qualitative differences between one or more varieties and landraces. The approach used also demonstrates the applicability of multivariate analysis for capturing most of the information present in the large set of data and for compressing a large number of possibly correlated variables into a smaller number of uncorrelated variables (Gottlieb et al., 2004). With PCA, a clear separation between several genotypes could be demonstrated. Protein spots that were found to contribute to this separation included proteins of several different functional categories. Furthermore, the levels of many proteins, such as defense-related proteins, may be affected by environmental conditions in the field experiment, contributing to variation in protein expression.
Compared with the natural variation observed in the non-GM samples, the effects of transformation on the proteome were considerably less pronounced. Indeed, statistical analysis showed no clear differences between the protein patterns of the GM lines and their controls. No new proteins unique to individual GM lines were observed. Therefore, on the basis of this analysis, there was no evidence for any major changes in protein pattern in the GM lines tested. This in itself is interesting, as some of the lines, e.g. Mal1 and Sam35S, produced extremely stunted plants with low tuber yield (Kumar et al., 1996; Taylor et al., 1998, 2000). However, nine proteins showing significant differences in expression were detected, and seven could be tentatively identified. It would require observations over several years and climatic conditions to confirm that these are truly unintended effects. Furthermore, such changes need not reflect a risk, particularly where the proteins are known to exist in other varieties or where the levels fall within the limits of natural variation in non-GM material. Corpillo et al. (2004) used a similar approach to assess GM and non-GM tomato lines and also did not find any evidence of differences in protein expression caused by genetic modification.
Proteomic profiling by 2-DE is a promising tool for screening purposes and, although the number of proteins that can be analyzed by 2-DE is still limited with respect to the predicted numbers of proteins present in the entire proteome of plants, it remains the most widely used tool for high-resolution protein separation and quantification. Like other profiling methods, proteomic screening is not yet in routine use when assessing the safety of GM products, but has the potential to reduce uncertainty by providing much more information on crop composition than targeted analysis alone. The combined development and application of validated metabolomic, proteomic, and transcriptomic approaches in plant biology will contribute to our knowledge of biological systems, but there may be clear benefits in the area of food safety.
MATERIALS AND METHODS
Plant Material
Non-GM potato (Solanum tuberosum) genotypes analyzed included the following (all general cultivars): Desirée (line 110), Record, Pentland Dell, Shelagh, Stirling, Torridon, Glenna, Morag, Eden, Maris Piper, Pentland Javelin, Cara, Pentland Crown, Brodick, Barbara, 91.MT.46 E 15, Pink Fir Apple, Golden Wonder, Lumpers, Fortyfold, and Anya. Lumpers and Fortyfold are old cultivars with no disease resistance introgressed from wild species. TBR.3369 1, 3, 4, and 5 are TBR lines with accession number 3369. Each clone is genetically unique but closely related to the rest within this subset. TBR.5646 1, 2, and 4 are the TBR lines with accession number 5646. TBR.3302 (2) is a TBR line with accession number 3302.PHU.4637 (1), Inca Sun, and Mayan Gold are diploid Solanum phureja lines. PHU4637 is a short-day clone, while the two others are long-day-adapted clones.
GM lines selected for the analyses have been developed at the Scottish Crop Research Institute (SCRI) over several years. All materials were planted as tubers, and the tubers were at least the second clonally propagated generation derived from the original transformation event. Where transgenic line numbers are provided, these indicate independent transgenic events with the constructs used. Agrobacterium-mediated transformation was used to generate the GM lines.
For cultivar Desirée, the control lines included: (1) wild-type tubers (line 50); (2) tubers generated from nonmodified plants produced via tissue culture (which included a callus phase; Des TC1); and (3) transgenic tubers transformed with an empty vector construct, either vector pBIN19 (Bevan, 1984) or its derivative pGPTV-Kan (Becker et al., 1992), containing the nptII gene but no target gene.
The following are GM lines of cultivar Desirée.
W2GBSS series. These were transformed to express an Aureobasidium gene (designated W2) in sense orientation, driven by the GBSS promoter (van der Leij et al., 1991), and a plastid-targeting sequence (Chambers et al., 1988) in the binary vector pGPTV-Kan. The W2 gene is derived from the filamentous fungus Aureobasidium pullulans and is believed to encode a glucan-branching enzyme. While the tubers show a waxy phenotype (high amylopectin content), this is not due to the W2 gene, which is not expressed (L.V.T. Shepherd, H.V.D. Davies, M.A. Taylor, and S. Tiller, unpublished data). In these experiments, only lines with empty vector constructs were used (W2GBSS VO4 and VO9).
Mal1 series. These contained the potato Mal1 gene introduced in sense or antisense orientation under control of two cauliflower mosaic virus (CaMV) 35S promoters in tandem (Sanders et al., 1987) in the binary vector pBIN19. Antisense lines Mal1 2A and 5A show extremely stunted growth in the field, but less so when grown under containment. The antisense lines have reduced activity of a glycoprotein-processing type II enzyme in tubers and show distinctive changes in leaf morphology caused by changes in cell wall structure (Taylor et al., 1998, 2000). Sense lines, such as the line Mal1 T41A, showed normal phenotype. The line Mal1 2V1 is a line with an empty vector construct.
SamDC series. These contained the potato S-adenosylmethionine decarboxylase (SamDC) gene in antisense orientation under control of two CaMV 35S promoters in the binary vector pBIN19 (Kumar et al., 1996). The CaMV 35S lines Sam35S 1 and 3 show stunted phenotype with reduced tuber numbers and dry matter content due to modified ethylene/polyamine metabolism following down-regulation of SamDC activity.
Sample Preparation
In 2000, five tubers of each independent line were grown in a randomized field plot according to standard agricultural practices at the SCRI. Each plot was replicated four times within the field. The five plants of each plot were harvested and the tubers were bulked together, resulting in four replicated samples of each line. For each replicate, a single average-sized tuber (usually between 80–100 g fresh weight per tuber, depending on the line and construct) was selected for analysis, resulting in four individual tuber samples for each line.
Total Protein Extraction
From each fresh tuber, a complete transverse slice of approximately 0.5 cm was taken across the middle of the tuber, frozen in liquid nitrogen, and ground in a mortar. Total protein was extracted from approximately 3 g of the powder, as previously described by Koistinen et al. (2002). The protein pellet was dissolved in 2-DE sample buffer containing 9.5 m urea, 2% (w/v) CHAPS, 1% (w/v) dithiothreitol (DTT), 0.64% (v/v) Bio-Lyte 5/7 ampholyte (Bio-Rad Laboratories, Hercules, CA), 0.16% (v/v) Bio-Lyte 3/10 ampholyte (Bio-Rad). Total protein was analyzed using the protein assay dye reagent (Bio-Rad).
Approximately 300 or 150 μg of total protein, depending on the experiment, were loaded on each 2-DE gel. Each gel corresponded to a single replicate sample from each potato line; thus, there were four replicate gels of each potato line or variety, except for the landrace TBR5646 (2), where there were three replicate gels.
2-DE
The first-dimension isoelectric focusing was performed using 24-cm immobilized pH gradient (IPG) strips (Amersham Biosciences, Uppsala) with a linear pH range of 4 to 7 in an Ettan IPGPhor isoelectric focusing system. The IPG strips were rehydrated overnight with total protein diluted in 8 m urea, 2% (w/v) CHAPS, 0.5% (v/v) IPG buffer 4 to 7, 0.28% (w/v) DTT, bromphenol blue up to a volume of 450 μL. After rehydration, the focusing was run using the following conditions: from 0 to 500 V in 1 min, from 4,000 to 8,000 V in 90 min, and, finally, 8,000 V until 52,000 Vh. After focusing, the strips were stored at −70°C and equilibrated at room temperature in 6 m urea, 50 mm Tris-HCl, pH 8.8, 30% (v/v) glycerol, 2% (w/v) SDS, 1% (w/v) DTT for 10 min, and another 10 min in the same buffer but with 2.5% (w/v) iodoacetamide replacing DTT. The second dimension was run in the Hoefer DALT system (Amersham Biosciences) in 19×23-cm homogeneous 12% SDS-PAGE gels, according to the manufacturer's instructions. The gels were run with constant 25- to 30-mA current overnight.
The gels were stained with SYPRO Ruby fluorescent stain (Bio-Rad), according to the manufacturer's instructions, except that 250 mL of the stain solution were used for each gel. Gel images were acquired with the FLA-3000 fluorescent image analyzer (Fuji Photo Film, Tokyo) using an excitation wavelength filter of 470 nm and an emission wavelength filter of 580 nm.
Image and Data Analysis
Gel image analysis was performed with PDQuest software (Bio-Rad). Protein spot intensities were normalized to the total intensity of valid spots to minimize possible errors due to differences in the amount of protein and staining intensity. For the gels run from the GM lines and their controls, the majority of spots with intensity value <121 had a quality score of 0 as given by PDQuest, while spots with intensity values >121 tended to have quality scores >0. On this basis, all spots with intensity value <121 and quality score <40 were considered as background noise, and their intensity value was set to 0. For the gels run from the non-GM varieties and landraces, only spots with intensity values >64 were considered. The spot intensities were transformed to square roots to normalize the data, and all subsequent statistical analyses were performed using these numbers.
The statistical methods used were ANOVA and Kruskal-Wallis to identify individual protein spots with significantly different expression levels and PCA to explore whether one or more lines or varieties would separate from others and to identify groups of proteins that, in combination, had different expression levels among the gels. For the GM lines, ANOVA was performed when spots were detected in at least 31 of the 40 gels. The Kruskal-Wallis nonparametric test was performed when spots were detected in 10 to 30 gels. The nonparametric test was used because the assumption of normality underlying ANOVA was violated due to the large number of zeros (no protein expression detected). For the potato varieties and landraces, ANOVA was performed when spots were detected in at least 88 of the 127 gels and the Kruskal-Wallis test when detected in 28 to 87 of the gels. In both cases, PCA was performed for the spots that were also analyzed by ANOVA. Statistical analysis was performed using the Genstat software package (NAG, Oxford).
Protein Identification by HPLC Electrospray MS/MS
For protein identification, 2-DE gels were stained with silver, according to Shevchenko et al. (1996), except that the gels were washed with a solution containing 40% (v/v) ethanol and 5% (v/v) acetic acid and then washed twice with water for 30 min. In-gel digestion was performed according to Koistinen et al. (2002) and the tryptic peptides were analyzed by two different MS systems.
Tryptic peptides were separated using the Ultimate/Famos capillary liquid chromatography (LC) system (LC Packings, Amsterdam). The sample was loaded onto a 300-μm i.d.×1-mm C18 PepMap (Dionex, Sunnyvale, CA) precolumn with a flow rate of 10 μL/min of 0.1% (v/v) acetic acid. After preconcentration and cleanup, the precolumn was automatically switched in-line with the PepMap C18 (3 μm, 75-μm i.d.×50 mm; Dionex) analytical column and the peptides were separated in a gradient of 2% to 40% (v/v) acetonitrile (ACN) in 40 min (0.1% [v/v] formic acid), the flow rate being 200 nL/min. The LC was connected to a mass spectrometer with a Protana platform (Protana, Odense, Denmark) using 30 μm PicoTip (New Objective, Woburn, MA). Mass spectra were recorded with a LCQ quadrupole ion trap mass spectrometer (ThermoQuest, San Jose, CA) using TriplePlay function: first full-scan mass spectrum was measured for range mass-to-charge ratio 615 to 2,000, the second scan was done to measure more exact Mr of the most abundant peptide signal in the first scan, and the third scan was done to measure the collision-induced MS/MS spectrum of the selected peptide. The spray needle was set to 2.4 to 3 kV in the positive ion mode. The inlet capillary temperature was 200°C. Other source parameters and spray position were optimized with the tryptic digest of myoglobin. The peptides were identified with Xcalibur software (ThermoQuest) and the Sequest algorithm using the National Center for Biotechnology (NCBI) nonredundant protein database, The Institute for Genomic Research (TIGR) potato EST database, and a potato EST library. Matches of MS/MS spectra against sequences in the databases were also verified manually.
Alternatively, tryptic peptides were separated as above, except that they were loaded with a flow rate of 30 μL/min of 0.1% (v/v) formic acid and 2% (v/v) ACN and eluted with a linear gradient starting from 100% eluent A (0.1% [v/v] formic acid and 5% [v/v] ACN) to 30% of eluent B (0.1% [v/v] formic acid and 95% [v/v] ACN) in 40 min. The LC was connected to a mass spectrometer with a nanoES ion source (Protana) using 15 μm PicoTip (New Objective). The positive time-of-flight (TOF) mass spectra were recorded on a QSTAR XL hybrid quadrupole TOF instrument (Applied Biosystems, Foster City, CA) using information-dependent acquisition. A TOF MS survey scan was recorded for mass range mass-to-charge ratio 400 to 2,000 followed by MS/MS scans of the two most intense peaks. Typical ion spray voltage was in the range of 2.0 to 2.4 kV and N2 was used as collision gas. The peptides were identified using ProID software (Applied Biosystems).
Acknowledgments
The potato EST library was kindly supplied by Dr. J.P. van Dijk, RIKILT Institute of Food Safety, Wageningen, The Netherlands.
This work was supported by GMOCARE (contract no. QLK1–1999–00765), by SAFE FOODS (contract no. Food–CT–2004–506446), by the European Commission (grant no. QLRT–1999–00765), by the Scottish Executive Environment and Rural Affairs Department (SEERAD), and by the Finnish Cultural Foundation (grant to S.J.L.).
Article, publication date, and citation information can be found at www.plantphysiol.org/cgi/doi/10.1104/pp.105.060152.
References
- Anderson L, Seilhamer J (1997) A comparison of selected mRNA and protein abundances in human liver. Electrophoresis 18: 533–537 [DOI] [PubMed] [Google Scholar]
- Becker D, Kemper E, Schell J, Masterson R (1992) New plant binary vectors with selectable markers located proximal to the left T-DNA border. Plant Mol Biol 20: 1195–1197 [DOI] [PubMed] [Google Scholar]
- Bevan M (1984) Binary Agrobacterium vectors for plant transformation. Nucleic Acids Res 12: 8711–8721 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bevan M, Bancroft I, Bent E, Love K, Goodman H, Dean C, Bergkamp R, Dirkse W, Van Staveren M, Stiekema W, et al (1998) Analysis of 1.9 Mb of contiguous sequence from chromosome 4 of Arabidopsis thaliana. Nature 391: 485–488 [DOI] [PubMed] [Google Scholar]
- Bradshaw JE, Mackay GR (1994) Breeding strategies for clonally propagated potatoes. In JE Bradshaw, GR Mackay, eds, Potato Genetics. CABI, Wallingford, UK, pp 467–497
- Canovas FM, Dumas-Gaudot E, Recorbet G, Jorrin J, Mock H-P, Rossignol M (2004) Plant proteome analysis. Proteomics 4: 285–298 [DOI] [PubMed] [Google Scholar]
- Cellini F, Chesson A, Colquhoun I, Constable A, Davies HV, Engel KH, Gatehouse AMR, Kärenlampi S, Kok EJ, Leguay J-J, et al (2004) Unintended effects and their detection in genetically modified crops. Food Chem Toxicol 42: 1089–1125 [DOI] [PubMed] [Google Scholar]
- Chambers SP, Prior SE, Barstow DA, Minton NP (1988) The pMTL nic-cloning vectors. I. Improved pUC polylinker regions to facilitate the use of sonicated DNA for nucleotide sequencing. Gene 68: 139–149 [DOI] [PubMed] [Google Scholar]
- Corpillo D, Gardini G, Vaira AM, Basso M, Aime S, Accotto GP, Fasano M (2004) Proteomics as a tool to improve investigation of substantial equivalence in genetically modified organisms: the case of a virus-resistant tomato. Proteomics 4: 193–200 [DOI] [PubMed] [Google Scholar]
- de Vienne D, Bost B, Fiévet J, Zivy M, Dillmann C (2001) Genetic variability of proteome expression and metabolic control. Plant Physiol Biochem 39: 271–283 [Google Scholar]
- Gottlieb DM, Schultz J, Bruun SW, Jacobsen S, Søndergaard I (2004) Multivariate approaches in plant science. Phytochemistry 65: 1531–1548 [DOI] [PubMed] [Google Scholar]
- Gygi SP, Rochon Y, Franza BR, Aebersold R (1999) Correlation between protein and mRNA abundance in yeast. Mol Cell Biol 19: 1720–1730 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Koistinen KM, Hassinen VH, Gynther PAM, Lehesranta SJ, Keinänen SI, Kokko HI, Oksanen EJ, Tervahauta AI, Auriola S, Kärenlampi SO (2002) Birch PR-10c is induced by factors causing oxidative stress but appears not to confer tolerance to these agents. New Phytol 155: 381–391 [DOI] [PubMed] [Google Scholar]
- Kuiper HA, Kleter GA, Noteborn HPJM, Kok EJ (2001) Assessment of the food safety issues related to genetically modified foods. Plant J 27: 503–528 [DOI] [PubMed] [Google Scholar]
- Kumar A, Taylor MA, Arif SAM, Davies HV (1996) Potato plants expressing antisense and sense S-adenosylmethionine decarboxylase (SAMDC) transgenes show altered levels of polyamines and ethylene: antisense plants display abnormal phenotypes. Plant J 9: 147–158 [Google Scholar]
- Meyers BC, Galbraith DW, Nelson T, Agrawal V (2004) Methods for transcriptional profiling in plants. Be fruitful and replicate. Plant Physiol 135: 637–652 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Millstone E, Brunner E, Mayer S (1999) Beyond ‘substantial equivalence’. Nature 401: 525–526 [DOI] [PubMed] [Google Scholar]
- Pots AM, Gruppen H, Hessing M, van Boekel MA, Voragen AG (1999) Isolation and characterization of patatin isoforms. J Agric Food Chem 47: 4587–4592 [DOI] [PubMed] [Google Scholar]
- Sanders PR, Winter JA, Barnason AR, Fraley R (1987) Comparison of cauliflower mosaic virus 35S and nopaline synthase promoters in transgenic plants. Nucleic Acids Res 15: 1543–1558 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schmidt MH, Raulf-Heimsoth M, Posch A (2002) Evaluation of patatin as a major cross-reactive allergen in latex-induced potato allergy. Ann Allergy Asthma Immunol 89: 613–618 [DOI] [PubMed] [Google Scholar]
- Shevchenko A, Wilm M, Vorm O, Mann M (1996) Mass spectrometric sequencing of proteins from silver-stained polyacrylamide gels. Anal Chem 68: 850–858 [DOI] [PubMed] [Google Scholar]
- Taylor MA, George LA, Ross HA, Davies HV (1998) cDNA cloning and characterisation of an α-glucosidase gene from potato (Solanum tuberosum L.). Plant J 13: 419–425 [DOI] [PubMed] [Google Scholar]
- Taylor MA, Ross HA, McRae D, Stewart D, Roberts I, Duncan G, Wright F, Millam S, Davies HV (2000) A potato α-glucosidase gene encodes a glycoprotein-processing α-glucosidase II-like activity. Demonstration of enzyme activity and effects of down-regulation in transgenic plants. Plant J 24: 305–316 [DOI] [PubMed] [Google Scholar]
- Thiellement H, Zivy M, Plomion C (2002) Combining proteomic and genetic studies in plants. J Chromatogr B 782: 137–149 [DOI] [PubMed] [Google Scholar]
- van der Leij FR, Visser RG, Ponstein AS, Jacobsen E, Feenstra WJ (1991) Sequence of the structural gene for granule-bound starch synthase of potato (Solanum tuberosum L.) and evidence for a single point deletion in the amf allele. Mol Gen Genet 228: 240–248 [DOI] [PubMed] [Google Scholar]