ABSTRACT
Bacteriophages are viruses infecting bacteria. The vast majority of them bear a tail, allowing host recognition, cell wall perforation, and DNA injection into the host cytoplasm. Using electron cryo-microscopy (cryo-EM) and single particle analysis, we determined the organization of the tail proximal extremity of siphophage T5 that possesses a long flexible tail and solved the structure of its tail terminator protein p142 (TrP142). It allowed us to confirm the common evolutionary origin between T5 TrPp142 and other known or putative TrPs from siphophages, myophages, and bacterial tail-like machines, despite very poor sequence conservation. By also determining the structure of the T5 tail proximal extremity after interaction with T5 bacterial receptor FhuA, we showed that no conformational changes occur in TrPp142 and confirmed that the infection signal transduction is not carried by the tube itself. We also investigated the location of T5 Neck1 or tail completion protein p143 (TCPp143) and showed, thanks to a combination of cryo-EM and structure prediction using Alphafold2, that it is not located at the capsid-to-tail interface as suggested by its position in the genome, but instead, very unexpectedly, on the side of T5 tail tip, and that it appears to be monomeric. Based on structure comparison with other putative TCPs predicted structures, this feature could not be shared by other TCPs and questions the affiliation of p143 to this family of protein.
IMPORTANCE
Bacteriophages, viruses infecting bacteria, are the most abundant living entities on Earth. They are present in all ecosystems where bacteria develop and are instrumental in the regulation, diversity, evolution, and pathogeny of microbial populations. Moreover, with the increasing number of pathogenic strains resistant to antibiotics, virulent phages are considered a serious alternative or complement to classical treatments. 96% of all phages present a tail that allows host recognition and safe channeling of the DNA to the host cytoplasm. We present the atomic model of the proximal extremity of the siphophage T5 tail, confirming structural similarities with other phages. This structure, combined with results previously published and further explored, also allowed a review and a discussion on the role and localization of a mysterious tail protein, the tail completion protein, which is known to be present in the phage tails, but that was never identified in a phage structure.
KEYWORDS: bacteriophage T5, cryo-electron microscopy, tail terminator protein, tail completion protein
INTRODUCTION
Bacteriophages—phages for short—are viruses infecting bacteria. They represent the most abundant biological entity on our planet. A large majority of them are composed of a capsid filled with double-stranded DNA and a tail, which can be long and contractile (myophage), long and flexible (siphophage), or short (podophage). The distal extremity of the tail bares the host-recognition apparatus, which also allows the perforation of the bacterium cell wall and the safe channeling of the DNA into the host cytoplasm. In long-tailed phages, the assembly pathway of the capsid and the tail are independent, and DNA-filled capsids attach onto functional tails to form fully assembled virions at the end of the lytic cycle. In siphophages, functional tails are produced after capping of the tail tube by the tail terminator proteins (TrP), of which siphophage λ gpU is the prototype (1). In myophages, tails can be capped both by a tube terminator and by a sheath terminator, as in T4 (2), or only by a tail terminator, which caps both the tube and the sheath, as in P1 (3) or Pam3 (4). In some phages, the tail needs the addition of the tail completion protein (TCP) to form fully functional tails, of which λ gpZ is the prototype (5). In some publications, both names are used interchangeably to designate the tail terminator. Empirically, it has been observed that phage proteins within the tail are arranged in a similar order to their coding genes in the genome (5, 6). Thus, it is expected that the TCP, also called Ne1 (Neck protein type 1) in (6, see also https://phrogs.lmge.uca.fr/), encoded by a gene usually located downstream of the capsid module and upstream of the TrP of the tail module, itself upstream of the tail tube protein, would also be located at the capsid-to-tail interface, the neck (5–7).
Phage T5 that infects E. coli belongs to the morphotype siphophage, which accounts for the majority of tailed phages. In the structure module of its genome, under the control of the same promoter and located between the head completion protein gene and the tail tube protein gene, are found two genes, p142 and p143 that were attributed to the TrP and the TCP, respectively (Fig. 1A and H) (8): TrPp142 was indeed localized within the virion in the neck using cross-linking with anti-TrPp142 IgG and immunogold labeling combined with EM imaging (8). A similar approach for TCPp143 proved unsuccessful: TCPp143 could be overexpressed in some conditions, but the protein precipitated on the Ni-NTA column, preventing the obtention of purified protein to raise antibodies (9). We recently determined the structure of the T5 tail tip—the distal extremity of the tail—before and after interaction with its receptor (10), where we proposed that a monomer of TCPp143 be attached to the Baseplate Hub Protein pb3 (BHPpb3). Here, we present the cryo-EM structure of phage T5 tail proximal extremity at a pseudo-atomic resolution, also before and after interaction with the phage receptor. TrPp142 is resolved as a hexameric ring, interacting with the last tail tube protein TTPpb6 ring. No available density could account for TCPp143 in this region of the tail.
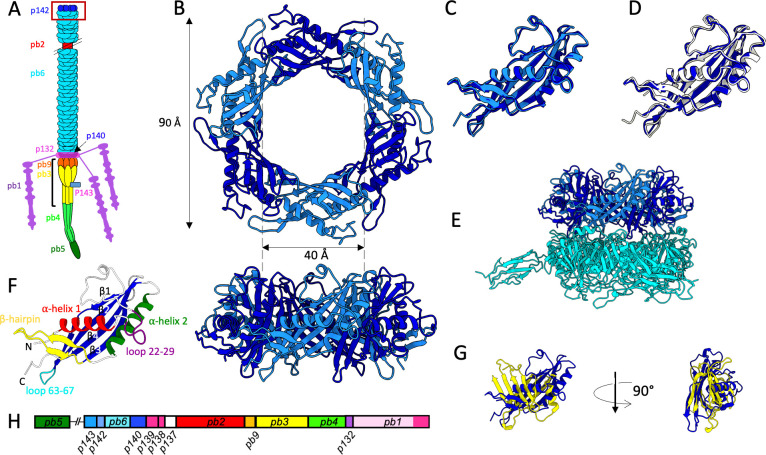
Fig 1.
(A) Scheme of T5 tail organization. The proximal end of the tail (formed of Trpp142 and TTPpb6 rings) is framed in red and the upper part of the distal complex (comprising TCPp143, DTPpb9, and BHPpb3) is indicated by a black accolade. (B) Top (upper panel) and side (lower panel) views of the Trpp142 hexamer as found at the distal extremity of the tail. Two consecutive monomers are not related by a C6 symmetry and are displayed in two different shades of blue. The outer and inner dimensions of the ring are indicated. (C) Superimposition of two Trpp142 consecutive, non-symmetry related, monomers. (D) Superimposition of two Trpp142 monomers from native tails (blue) and tails after interaction with receptor FhuA (white). (E) Side view of the proximal extremity of the tail, featuring the Trpp142 hexamer (in shades of blue) sitting on top of the TTPpb6 trimer (in cyan). (F) Annotated model of Trpp142, indicating its main secondary structures. C and N-termini are also indicated. (G) Superimposition of Trpp142 (blue) and a tail tube domain from TTPpb6 (yellow). (H) Map of T5 tail structural proteins genes.
RESULTS AND DISCUSSION
Structure of the proximal extremity of phage T5 tail
T5 tails, purified from a capsid defective mutant, were produced, vitrified for cryo-EM characterization, and imaged for the data collection on a high-end electron microscope (Fig. S1A) (10). Image analysis yielded a map of T5 tail proximal extremity, containing a trimeric ring of the tail TTPpb6, on top of which sits a hexameric ring formed by the TrPp142 (Fig. 1E). The estimated resolution of this map is 3.9 Å (Fig. S1C and D), which allowed us to build pseudo-atomic models for both TrPp142 hexamer and a TTPpb6 trimer. TrPp142 is a 161-residues protein, mainly formed by a four-strand antiparallel β-sheet that lines the lumen of the tube, flanked by two α-helices, and a short loop between β-strands 1 and 2. An 18-residue β-hairpin links the second α-helix to the β-sheet (Fig. 1F). As already discussed in the case of phage λ TrP gpU (11), TrPp142 fold appears to be related to the canonical tail tube domain (TTD) fold, a common building block present in numerous phage proteins (12, 13), although clear differences are present (Fig. S1G). It indeed exhibits a similar β-sheet/helix/loop fold, but a second α-helix is present in TrPp142 while a second β-sheet is present in the TTD. Also, the orientation of the helices is different (Fig. 1G). The interaction between two neighboring TrPp142 monomers is mostly mediated through strand β1, N-terminus, and C-terminus for monomer 1, and strand β3, α-helix 2, and the 22–29 loops for monomer 2. The interaction between the hexameric TrPp142 ring and the trimeric TTPpb6 ring under it is ensured by the tip of loop 63–67, which inserts in two similar pockets resulting in the domain duplication of TTPpb6 (14) and is mostly mediated by electrostatic interactions (Fig. S2). As the TrPp142 hexameric ring sits on top of a TTPpb6 trimeric ring, it exhibits a C3/pseudo-C6 symmetry and two consecutive TrPp142 monomers are not strictly identical (Fig. 1B and C). However, TTPpb6 also exhibits a C3/pseudo-C6 symmetry as it results in domain duplication (14) (excluding its Ig-like domain); thus, the two consecutive TrPp142 monomers are almost superimposable (rmsd = 1.02 Å over 156 residues). This is reminiscent of what was observed for T5 distal tail protein DTPpb9, which also assembles as a hexamer sitting above a trimer (10).
Interestingly, purified TrPs crystallize as biologically relevant hexamers (PDB 3FZB (1) 2GJV and 4ACV). Also, a previous study showed that over-expression of TrPp142 with a His6-tag in the N-terminus leads to soluble, monomeric protein while over-expression of TrPp142 with a His6-tag in the C-terminus (TrPp142-Cter) leads to the formation of fibers, of a diameter similar to that of T5 tails (9). This illustrates the fact that phage structural proteins have not only evolved to adopt a specific structure, allowing them to be building blocks for the phage but also to bear inherent dynamics when monomeric. This allows them to remain monomeric, even at high concentrations, and to polymerize and assemble only in presence of their partners, to form a fully functional virion. Polymerization however also occurs when conditions are extreme, as in a crystallization drop. In the case of TrPp142-Cter, the tag addition somehow modifies the dynamics of the protein, which induces aberrant polymerization upon overexpression. TrPp142-Cter fibers do not look like empty tubes, as would be expected by the piling of TrPp142 rings. It could be that the C-terminal tag folds back in the lumen of the tube, filling it and explaining why affinity purification failed (9). Several structures of TTPpb6 were previously solved, in different contexts: either as a crystallized monomer, within the tail tube, or interacting with the DTPpb9 (10, 14). Our study adds the structure of TTPpb6 in contact with the TrPp142 to these known structures. Overall, there are very little variations between the different TTPpb6 structures and TTPpb6 in interaction with TrPp142 superimposes almost perfectly with TTPpb6 interacting with another TTPpb6 or with DTPpb9 (rmsd of 0.655 Å and 0.576 Å, respectively, over the 464 residues).
DALI searches link Trpp142 to several phage TrPs from siphophages as well as from myophages, bacterial tail-like systems related to myo- or siphophages or prophage proteins (Fig. 2A through D; Table S1), with DALI Z-scores ranging from 23.9 to 5.4 and alignment rmsd from 1.2 to 4.2 Å, even though sequence identity is non-existent (7 to 18%). This confirms that TrPs form a family of largely conserved proteins among sipho-, myophages, and related machinery, despite no sequence homologies. Several structures of siphophage TrPs are available, either from purified protein or within the phage context: gp17 from SPP1 (15–17), gpU from λ (1, 18–20)(Fig. 3C and D), gp18 from R4C (21), and gp119 from phage DT57C (22), determined by NMR, X-ray crystallography, and/or cryo-EM. In phago structures confirm that TrPs indeed assemble as a hexamer, at the proximal extremity of the tail tube, thereby terminating tube polymerization and providing the surface for interaction with the capsid (17–19, 22). Deletion of this protein has various phenotypes: in phage λ, capsids are produced along with very long, TMP-less, polytails, resulting from the endless polymerization of the TTP (23); in myophages T4 and P2, deletion mutants of the tube terminators gp3 and gpR, respectively, result in tails with a longer tube, whereas deletion mutants of the sheath terminators gp15 and gpS, respectively, bear longer sheaths (24–26); in myophages Mu and SPO1, deletions mutants of the TrP K and 8, respectively, result in longer tails. In contracted tails, the tubes protrude from the capsid-proximal extremity of the tails rather than from the base plate (27, 28). In siphophage SPP1, normal-length tails are assembled in deletion mutant of the TrP gp17, which however cannot attach to full capsids (29). Thus, TrPs are suggested to both allow to stop the tail polymerization at the right length, together with the TMP (30), but also to allow interaction with the capsid (29). Some TrPs are larger proteins and include additional domains, for example interacting with the sheath of the myophage tails when they also fill the sheath termination role. Interestingly, the PVC16 TrP C-terminus additional domain appears to share structural similarity with the TrPp142 fold, indicating domain duplication, a common “LEGO” strategy among phages and phage-related machines (13).
Fig 2.
Comparison of TrPp142 with TrPs homologous proteins using DALI pairwise. TrPp142 is displayed in all panels, colored in dark blue, and superimposed to a selection of homologous proteins of different categories. (A) Siphophages and siphophage-like machines, phage λ (dark green), GTA (light green). (B) Myophages, phage T4 (yellow), phage XM1 (white). (C) Phage-like machines related to myophages: PyocinR2 (red), PVC (orange). (D) Miscellaneous: Type4 fimbrial biogenesis protein PilO from P. aeruginosa (gray).
Fig 3.
Isosurface view of the non-sharpened cryo-EM maps of T5 tail proximal extremity, in its native state (A, gray) and after interaction with the bacterial receptor FhuA (B, cyan). Left: side interior view and right: top view. The position and thickness of the top view slice are indicated by black accolades in the side view slice. TMPpb2 is indicated with red arrows in A and is absent in B. The framed panel in A shows the upper part of the tail tube extremity at a higher threshold, with TMPpb2 running up to the very end of the tube. (C) Slices of a C3 map of the proximal extremity of the T5 tail tube at different positions, showing TMPpb2 inside the lumen.
The tape measure protein TMPpb2
The unsharpened cryo-EM map of the T5 tail proximal extremity shows a 25 Å diameter large, cylindrical, and ill-defined density filling the tail tube lumen to the top of TrPp142, which disappears from the map after interaction with FhuA (Fig. 3A and B). This density most probably belongs to the N-terminus of the Tape Measure Protein pb2 (TMPpb2), which fills the entire tail tube in native phages and purified tails (14). Unlike for the distal part of the tail, the resolution of the central density in this proximal extremity of the tail is too low to propose a stoichiometry, yet alone a model of the TMP. However, as a trimer was solved for the C-terminus of the protein (10), is it reasonable to propose that a trimer is also present in the N-terminus. Also, and despite the low resolution, different slices taken along the cryo-EM map of the T5 tail proximal extremity do not seem to hint at a higher oligomeric state than a trimer of TMPpb2 (Fig. 3C).
Indeed, TMP stoichiometry has remained uncertain for many decades. In 1973, Zweig and Cummings measured the relative abundance of the different proteins of phage T5, based on a band densitometry analysis of radiolabeled phage proteins after SDS-PAGE (31). TMPpb2, BHPpb3, and the straight fiber protein pb4 were observed to represent 1.5%, 1.5%, and 1%, respectively, of the structural proteins of phage T5. In 1988, McCorquodale and Warner interpreted these data and concluded that TMPpb2, BHPpb3, and pb4 were present as pentamers in the virion (32). In 1974, Casjens and Hendrix proposed a stoichiometry of 6–7 per virion for TMPgpH of phage λ, based on densitometry measurements of phage protein bands after SDS-PAGE and Coomassie blue or Fast green staining (33). Both in T5 (10, 22) and λ (18, 19), however, a trimer of the TMP was solved in the C-terminus of the protein, in tight interaction with the BHP, as for siphophages 80α (34). For T5, this is consistent with Zweig and Cummings’ results, as BHPpb3 and pb4 were also found to be trimers (10). A stoichiometry of six has been however proposed for the TMP of myophages Pam3 (4) and A-1(L)(35). More data, at higher resolution, is thus needed to clarify this point.
TMPpb2 reaches the proximal extremity of the tube (Fig. 3A), indicating a possible direct interaction with the DNA in the full phage, as seen in other siphophages (e.g., SPP1 (15), λ (18)). As previously proposed (10, 14), the expulsion of the TMP would be part of the signal transmission pathway that starts with host recognition by the receptor binding protein, continues with cell wall perforation, and ends with DNA ejection. All along the tube and except for the 35 last residues in the tail tip, TMPpb2 does not interact with the tube wall (10, 14), which can explain the low resolution observed for that protein; this is the case in all phage tail structures determined to date.
Structure of the proximal extremity of the tail after interaction with receptor FhuA
We also imaged T5 tails after interaction with E. coli receptor FhuA reconstituted into nanodiscs (Tail-FhuA) with cryo-EM (10). Image analysis yielded a second 3D map for the proximal extremity of Tail-FhuA, at an average resolution of 4 Å (Fig. S1C and D) allowing us to build pseudo-atomic models for TrPp142 and TTPpb6. At the resolution of our maps, no conformational change was observed in TrPp142 monomers or hexamer (rmsd 0.43 Å over 156 residues between Tail and Tail-FhuA individual monomers) (Fig. 1D), and neither in TTPpb6. For the latter, it was already shown that the T5 tail tube, formed of a stack of TTPpb6 trimeric rings, does not undergo any conformational changes upon interaction with FhuA and that the signal of the infection is not transmitted from the receptor to the capsid through the tube itself (14). TrPs homologous proteins from myophages or tail-like machines also show no conformational change upon infection (2, 36, 37).
Because TMPpb2 is expelled during the infection process (8, 10), the lumen of the tube is empty in Tail-FhuA (Fig. 3B), which constitutes the only difference between the two EM maps of the tail proximal extremity. To increase the number of particles and to achieve better resolution, we combined both EM data sets but it did not lead to any sensible improvement of the map.
The tail completion protein TCPp143
As mentioned in the introduction, TCPs are a highly conserved family of tail proteins, expected to be located in the neck of phages given the position of their genes in the tail module of phage genomes (5–7, see also https://phrogs.lmge.uca.fr/). Phage T5 TCP p143, TCPp143, is a 27.8 kDa protein, which gene is sandwiched between that of T5 Head Completion Protein and of TrPp142 (Fig. 1H). In the cryo-EM map of the T5 tail proximal extremity presented here, we could clearly trace TTPpb6 trimers and a hexamer of TrPp142 (Fig. 1E, 3A, and B). Inside the tube, low-resolution density, similar to that present along the whole tube, can reasonably be attributed to TMPpb2 (see above). Thus, there is no unassigned density in this proximal extremity map of the tail that could account for TCPp143.
TCPp143 is however present in the T5 virion and purified tails, as it was unambiguously detected by mass spectrometry (8, 10). We recently determined the high-resolution cryo-EM structure of the T5 tail tip. In the C3 map, at the base of BHPpb3, a faint density was visible. Intrigued, we calculated an unsymmetrized map, focusing on this density, and we could indeed solve a bona fide density, otherwise blurred by the imposed threefold symmetry, located at the base of the BHPpb3, in the groove between the closed tube and the first fibronectin domain (Fig. 4A; Fig. S2A) (10). This density appears only on one of the BHPpb3 monomers, which is completely unexpected considering the overall C3 symmetry of the rest of the tail. Indeed, the only other monomeric protein of the assembly is the Receptor-Binding Protein pb5, located at the extremity of the straight fiber (Fig. 1A)(38). The map resolution in this area (6–10 Å) did not allow us to build a protein model de novo in this density (Fig. S2A); however, the only unlocalized tail protein remained TCPp143 (10).
Fig 4.
(A) Isosurface view of an unsymmetrized and unsharpened cryo-EM map of the tail tip (side view). The BHPpb3 trimer is in yellow and the DTPpb9 hexamer is in orange. The additional density located on the side of the BHPpb3 trimer and corresponding to the monomeric protein TCPp143 is framed in black. (B–D) Fit of an AF2 predicted model in the additional density (B: unmodified AF2 model, C: flexible-fitted AF2 model, D: flexible-fitted AF2 model in the segmented map). Different views (from left to right: front view, side view, top slice, bottom slice) are shown. The model in panel B is colored according to its per-residue pLLDT/confidence measure. Color key interpretation is the following: 100 to 90 = high accuracy expected, 90 to 70 = backbone expected to be correctly modeled, 70 to 50 = low confidence/caution, 50 to 0 = should not be interpreted, a strong predictor of disordered regions.
We used Alphafold2 (AF2) (38) to predict TCPp143 structure and see whether it could be fitted into the unassigned density. The five AF2 predicted TCPp143 structures are rather heterogeneous and exhibit relatively low confidence levels (pLLDT between 30% and 90%, Fig. 5; Fig. S3A), which could indicate a highly flexible protein. AF2-TCPp143 seems however to be composed of two domains (residues 1–80 and 81–255) (Fig. S3B and C). The global size and shape of some of the AF2 predicted models correspond to the size and shape of the unattributed density (Fig. 4B) and flexible fitting led to a convincing fit of the predicted secondary structures (Fig. 4C; PDB as Suppl. file). The fitted structure is mostly composed of α-helices and loops, with a small β-sheet formed by three short strands (Fig. 5; Fig. S3A) (10). Thus, we proposed that this unallocated density indeed belonged to a monomer of TCPp143. Even though our results indicate that only one TCPp143 copy per phage is present, cryo-EM single particle analysis consisting of the averaging of a large number of individual particles, we cannot exclude the possibilities of zero, two, or three TCPp143 copies being present on T5 tails. If it were the case though, these occurrences would only account for a minority of particles, as there is no hint of more than one full occupancy in our unsymmetrized map. We cannot explain yet why only one TCPp143 molecule would bind to the T5 tail tip, as the BHPpb3 trimer follows a C3 symmetry and, at the resolution of our map, no difference in the three potential binding sites could be identified (Fig. S2B through D)(10).
Fig 5.
Alphafold2 predictions for different phage TCPs, colored according to their per-residue pLLDT (confidence measure), except for the flexible fitted TCPp143 (purple). The pLLDT color key interpretation is as in Fig. 2. The bottom panel models are rotated horizontally by 90° compared to the ones of the top panel. Uniprot entries: T5 (Q6QGE0), λ (P03731), SPP1 (O48447), HK97 (E9RJ97), TP901-1 (Q77K21), P2 (P36934), and Mu (Q01261).
In the EM map of the T5 tail tip after interaction with its receptor FhuA, TCPp143 is absent (Fig. S2E). This is not surprising given the position of TCPp143 at the interface between the tip of the tube and the straight fiber, as the large conformational changes induced by receptor binding, inducing the bending of the straight fiber on the side of the tube, completely re-arrange/destroy TCPp143-binding site (Fig. S2E). Given the asymmetry induced by TCPp143 in the tail tip, its presence could determine the direction in which the straight fiber bends. In any case, it would definitely detach upon infection and could play an active role, remaining to be determined, in the infection process.
To see whether the putative TCPs of different phages share more than a common position in the structural gene module in phage genomes, we predicted the structures of some TCPs (Fig. 5). A first striking difference with TCPp143 is that all structures are predicted with a high level of confidence (pLLDT >90 for most of the protein). A second interesting feature is that, even though varying in length, from 112 residues for TP901-1 TCP to 192 for λ TCP, these predicted structures all share common features, including two side-by-side long α-helices and a small β-sheet, and superimpose very well (Fig. S3D). AF2-TCPp143 seems to stand out from the other predicted TCPs in terms of structure and length (255 residues), and thus, it could maybe also stand out in terms of location and be located at a different position in the tail.
As far as we know, no structure of a phage TCP has been solved to date, and their function and localization within the tail remain unknown. Literature about TCPs is scarce, contradictory, and does not help to set the case about their localization or function. Indeed, TCP deletion sometimes leads to phages failing to assemble correctly (tail and capsid are produced but do not assemble) as in myophage P2 (26), siphophages TP901-1 (39), and SPP1 (40). In other phages, like siphophage λ (gpZ) (33) or myophage Mu (27), TCP deletion leads to normal-looking virions that however exhibit much lower infectivity. Such a dramatic functional difference is difficult to explain, especially considering that their predicted structures appear very similar. The different phenotype of TCP deleted mutants however does not stem from a different neck organization between siphophages and myophages, as in TCP less mutants, myophages Mu and P2 behave differently, as well as siphophages SPP1, TP-901-1, and λ. In the case of phages P2, TP901-1, and SPP1, the fact that TCP deleted mutants do not form complete virions suggests that the presence of this protein is a prerequisite to capsid attachment to the tail, and the most straightforward conclusion would be that it is located in the neck region (40). However, the structure of the SPP1 neck has been determined and, as for T5, unambiguously shows that its TCP-gp16.1 is not present (17). This is also the case for λ, in which TCP-gpZ is detected in the tail particle but not localized in the tail structure (18, 19), and for all myophage neck structures solved to date, where there is no trace of a protein that would be a TCP (e.g., 4, 41, 42). In λ, in which full virions are produced in a TCP-deleted mutant, it was suggested that TCP-gpZ allows for the correct DNA insertion into the tail after capsid and tail attachment (43), explaining the lack of infectivity of gpZ-deleted virions. TCPp143 is a small, monomeric protein, part of a highly symmetrical megadalton-sized complex, with a very unexpected location. Thus, special attention and sophisticated image processing were required to reveal it in the T5 tail. We would like to kindly encourage our colleagues to have a fresh look at their phage tail cryo-EM data, as this protein should also be present in a majority of phage tails. Increasing data would allow us to shed light on this still mysterious protein.
Conclusion
We present the structure of the proximal extremity of the phage T5 tail. In the densities available, we could trace, as expected, a hexamer of TrPp142 and a first trimer of TTPpb6. In the lumen of the tail tube before interaction with the receptor, densities of much lower resolution are present, which we attribute to TMPpb2. As for the many structures of proximal tail extremity or necks of either sipho- or myophages, no remaining densities, even faint, could correspond to the TCP. Instead, we could localize a medium-resolution unattributed density in the unsymmetrized map of the distal extremity of the T5 tail, at the base of a BHPpb3 monomer, in which a predicted structure of TCPp143 could be fitted (10). TCPs have been proposed to form a superfamily of neck proteins, based on iterative profile-profile comparison and genome organization analysis (5, 6, see also https://phrogs.lmge.uca.fr/). However, predicted structures of other sipho- and myophages TCPs share the same fold, whereas TCPp143 seems to stand out in several aspects (predicted structure and proposed location). Given the dispersity of phenotypes of TCP deleted phages, it is thus possible that TCPs adopt different roles and locations within different phages and would, in fact, not correspond to a homogeneous family of proteins.
MATERIALS AND METHODS
T5 tail purification
Tails were purified as in references (8, 10). Briefly, an E. coli strain F culture was infected with the mutant phage T5D20am30d, which bears an Amber mutation in the major capsid protein gene. After complete cell lysis and DNA digestion by the addition of DNase, cell debris and unlysed cells were removed by low-speed centrifugation, and T5 tails were precipitated by incubation with 0.5 M NaCl and 10% (wt/wt) PEG 6000 overnight at 4°C. After low-speed centrifugation, the pellet was resolubilized in buffer and purified on a glycerol step gradient. The gradient fractions containing the tails were diluted 10 times, and loaded onto an ion exchange column, and the tails were eluted by a 0–0.5 M NaCl linear gradient. Purified tails were incubated for 30 min with FhuA-loaded nanodiscs in a tail:FhuA-nanodisc ratio of 1:10 (vol/vol) at room temperature (10).
Cryo-EM sample preparation
Typically, 3.5 µL of T5 tails sample (with or without FhuA-nanodisc) was deposited on a freshly glow discharged (25 mA, 30 s) Cu/Rh 300 mesh Quantifoil R 2/1 EM grids and plunge-frozen in nitrogen-cooled liquid ethane using a ThermoFisher Mark IV Vitrobot device (100% humidity, 20°C, 5 s blotting time, blot force 0).
EM data acquisition
Respectively, 3,208 and 5,752 micrographs were collected for the tails alone and the FhuA-nanodisc incubated tails, over three different data collections. 40-frame movies were acquired on a ThermoFisher Scientific Titan Krios G3 TEM (European Synchrotron Radiation Facility, Grenoble, France) (44) operated at 300 kV and equipped with a Gatan Quantum energy filter coupled to a Gatan K2 summit direct electron detector. Automated data collection was performed using Thermo Fisher Scientific EPU software, with a typical defocus range of −1.0 to −3.0 µm and a total dose of 40 e-/Å2 per movie. A nominal magnification of 105.000× was used, resulting in a calibrated pixel size at the specimen level of 1.351 Å. The images used were the same as those used in reference (10) (Table S2).
EM image processing
Preprocessing and particle picking
Frame alignment was performed using Motioncor2 (45) keeping, respectively, frames 3 to 30 and 1 to 40 for Tail and Tail, and applying dose weighting. Contrast transfer function parameters were then determined using Gctf (46); manual particle picking was performed with EMAN2/e2helixboxer (47). The picking coordinates were consistently centered on the last visible ring at the proximal extremity of the tail tube, corresponding to the tail terminator p142 hexamer. All subsequent image processing was performed using Relion (versions 3.0 and 3.1) (48) (Table S2).
Proximal extremity of the tail (Tail and Tail-FhuA)
Tail and Tail-FhuA data were processed identically. After particle extraction (box size of 140 × 140 pixels2), a run of 2D classification was performed to get rid of bad particles (Fig. S1B). A run of 3D classification was then performed, using a cylinder (diameter 90 Å, height 188 Å) as an initial model, at the end of which a small additional fraction of the particles was discarded, ending up with a homogeneous data set of, respectively, 9,953 and 10,701 particles. 3D refinement with masking and C3 symmetry imposed followed, using the best 3D class as an initial model (low-pass filtered at 15 Å) yielded a C3 map of the proximal extremity of the tail, comprising the tail terminator p142 hexamer and the first TTPpb6 ring. After masking and sharpening, the overall estimated resolution of the maps, respectively, reached 3.88 Å and 4.05 Å (FSC0.143) for Tail and Tail-FhuA (Fig. S1D). For both maps, a local resolution map was calculated using Relion built-in local resolution tool (Fig. S1C).
Tail completion protein TCPp143
The full protocol used to obtain the cryo-EM maps for the T5 tip complex is detailed in Linares et al. (10). Here, we detail the steps that are specific to the map shown in this study. After particle extraction under T5 collar and 2D classification, a homogeneous data set of 9,290 particles was obtained. No 3D classification was performed. Using a 15 Å resolution map determined from a previous cryo-EM data collection (unpublished) as an initial model, a C3 reconstruction of the tip was calculated, from the second TTPpb6 ring to the beginning of the central fiber. Refined particles from the previous reconstruction were reextracted (box size of 200 pixels by 200 pixels) and recentered on the lower part of BHPpb3, on the side of which TCPp143 is located. After reclassification/selection, symmetry relaxation, and a new run of 3D refinement using suitable masking and no symmetry imposed, a new map of the central part of the tip complex was obtained, where densities for the monomeric TCPp143 were visible.
Protein model building
Atomic protein models were built into the two cryo-EM maps using the Coot software (49) by tracing the protein sequence into the densities and were then iteratively refined alternating Coot manual refinement and PHENIX (50) real space refine tool until convergence. For Trpp142, the model of the homologous Trp-gpU from siphophage λ (PDB: F3Z2) (1) was rigid-body fitted in the cryo-EM map of the proximal extremity of the tail and was used as a starting point for model building, while for TTPpb6 models, existing cryo-EM models (PDB: 7QG9) (10) were used as a starting point and were refined into the EM maps. Molprobity (51) was used for model quality assessment (Table S2).
All the predicted structures in this study were generated using AlphaFold2 (52) on ColabFold (https://colab.research.google.com/github/sokrypton/ColabFold/blob/main/AlphaFold2.ipynb) (53). In addition, for TCPp143, we fitted the most promising AF2 predicted model into the unsymmetrized tip cryo-EM map, then used a combination of Flex-EM (54)/Namdinator (55) (flexible fitting) and PHENIX (50) (real space refine), iteratively, to obtain a better model for this protein, with a convincing fit of most of its secondary structures.
ACKNOWLEDGMENTS
We thank Guy Schoehn for the general scientific and instrumentation support; Grégory Effantin for cryo-EM data collection; Emmanuelle Neumann, Daphna Fenel and Séraphine Degroux for technical help; Leandro Estrozi, Ambroise Desfosses, and Benoît Arragain for help and discussion on image analysis; and Aymeric Peuch for help with the use of the EM computing cluster. We acknowledge the European Synchrotron Radiation Facility for the provision of beam time on CM01. This work used the platforms of the Grenoble Instruct-ERIC center (ISBG; UMS 3518 CNRS-CEA-UGA-EMBL) within the Grenoble Partnership for Structural Biology (PSB), supported by FRISBI (ANR-10-INBS-05-02) and GRAL, financed within the University Grenoble Alpes graduate school (Ecoles Universitaires de Recherche) CBH-EUR-GS (ANR-17-EURE-0003). The IBS acknowledges integration into the Interdisciplinary Research Institute of Grenoble (IRIG, CEA).
This research was funded by the Agence Nationale de la Recherche, grant number Project ANR-20-CE11-0023. The electron microscope facility is supported by the Auvergne-Rhône-Alpes Region, the Fondation pour la Recherche Médicale (FRM), the Fonds FEDER, and the GIS-Infrastructures en Biologie Santé et Agronomie (IBiSA).
C.B. conceived the project. R.L. and C.B. prepared the T5 tails and FhuA-nanodiscs used for high-resolution cryo-EM. R.L. optimized cryo-grid preparation, processed the cryo-EM data, built the atomic models, and analyzed and interpreted the models with the help of C.B. R.L. and C.B. wrote the paper.
Contributor Information
Cécile Breyton, Email: cecile.breyton@ibs.fr.
Kristin N. Parent, Michigan State University, East Lansing, Michigan, USA
DATA AVAILABILITY
The cryo-EM density maps of the T5 tail proximal extremity presented in this study and the associated atomic coordinates have been, respectively, deposited in the Electron Microscopy Data Bank (EMDB) and Protein Data Bank (PDB) under the following accession codes: EMD-15967/PDB 8BCP (native state) and EMD-15968/PDB 8BCU (after interaction with FhuA). The unsymmetrized cryo-EM density map of the T5 tail tip has been deposited in the EMDB under accession code EMD-51232. TCPp143 AF2 flexible-fitted model is provided as supplemental material.
SUPPLEMENTAL MATERIAL
The following material is available online at https://doi.org/10.1128/jvi.01376-24.
PDB file of TCPp143 AlphaFold2 predictedmodel fitted into the free density of T5 tail tip.
Figures S1 to S3; Tables S1 and S2.
ASM does not own the copyrights to Supplemental Material that may be linked to, or accessed through, an article. The authors have granted ASM a non-exclusive, world-wide license to publish the Supplemental Material files. Please contact the corresponding author directly for reuse.
REFERENCES
- 1. Pell LG, Liu A, Edmonds L, Donaldson LW, Howell PL, Davidson AR. 2009. The X-ray crystal structure of the phage lambda tail terminator protein reveals the biologically relevant hexameric ring structure and demonstrates a conserved mechanism of tail termination among diverse long-tailed phages. J Mol Biol 389:938–951. doi: 10.1016/j.jmb.2009.04.072 [DOI] [PubMed] [Google Scholar]
- 2. Fokine A, Zhang Z, Kanamaru S, Bowman VD, Aksyuk AA, Arisaka F, Rao VB, Rossmann MG. 2013. The molecular architecture of the bacteriophage T4 neck. J Mol Biol 425:1731–1744. doi: 10.1016/j.jmb.2013.02.012 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Yang F, Wang L, Zhou J, Xiao H, Liu H. 2023. In situ structures of the ultra-long extended and contracted tail of Myoviridae phage P1. Viruses 15:1267. doi: 10.3390/v15061267 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Yang F, Jiang Y-L, Zhang J-T, Zhu J, Du K, Yu R-C, Wei Z-L, Kong W-W, Cui N, Li W-F, Chen Y, Li Q, Zhou C-Z. 2023. Fine structure and assembly pattern of a minimal myophage Pam3. Proc Natl Acad Sci U S A 120:e2213727120. doi: 10.1073/pnas.2213727120 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Davidson AR, Cardarelli L, Pell LG, Radford DR, Maxwell KL. 2012. Long noncontractile tail machines of bacteriophages. Adv Exp Med Biol 726:115–142. doi: 10.1007/978-1-4614-0980-9_6 [DOI] [PubMed] [Google Scholar]
- 6. Lopes A, Tavares P, Petit M-A, Guérois R, Zinn-Justin S. 2014. Automated classification of tailed bacteriophages according to their neck organization. BMC Genomics 15:1027. doi: 10.1186/1471-2164-15-1027 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Linares R, Arnaud C-A, Degroux S, Schoehn G, Breyton C. 2020. Structure, function and assembly of the long, flexible tail of siphophages. Curr Opin Virol 45:34–42. doi: 10.1016/j.coviro.2020.06.010 [DOI] [PubMed] [Google Scholar]
- 8. Zivanovic Y, Confalonieri F, Ponchon L, Lurz R, Chami M, Flayhan A, Renouard M, Huet A, Decottignies P, Davidson AR, Breyton C, Boulanger P. 2014. Insights into bacteriophage T5 structure from analysis of its morphogenesis genes and protein components. J Virol 88:1162–1174. doi: 10.1128/JVI.02262-13 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Noirclerc-Savoye M, Flayhan A, Pereira C, Gallet B, Gans P, Ebel C, Breyton C. 2015. Tail proteins of phage T5: investigation of the effect of the His6-tag position, from expression to crystallisation. Protein Expr Purif 109:70–78. doi: 10.1016/j.pep.2015.02.003 [DOI] [PubMed] [Google Scholar]
- 10. Linares R, Arnaud C-A, Effantin G, Darnault C, Epalle NH, Boeri Erba E, Schoehn G, Breyton C. 2023. Structural basis of bacteriophage T5 infection trigger and E. coli cell wall perforation. Sci Adv 9:eade9674. doi: 10.1126/sciadv.ade9674 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Cardarelli L, Pell LG, Neudecker P, Pirani N, Liu A, Baker LA, Rubinstein JL, Maxwell KL, Davidson AR. 2010. Phages have adapted the same protein fold to fulfill multiple functions in virion assembly. Proc Natl Acad Sci U S A 107:14384–14389. doi: 10.1073/pnas.1005822107 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Flayhan A, Vellieux FMD, Lurz R, Maury O, Contreras-Martel C, Girard E, Boulanger P, Breyton C. 2014. Crystal structure of pb9, the distal tail protein of bacteriophage T5: a conserved structural motif among all siphophages. J Virol 88:820–828. doi: 10.1128/JVI.02135-13 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Veesler D, Cambillau C. 2011. A common evolutionary origin for tailed-bacteriophage functional modules and bacterial machineries. Microbiol Mol Biol Rev 75:423–433, doi: 10.1128/MMBR.00014-11 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Arnaud C-A. 2017. Structure de la queue du phage T5 et mécanisme de perforation de l’enveloppe bactérienne par les Siphoviridae. University Grenoble-Alpes. [Google Scholar]
- 15. Lhuillier S, Gallopin M, Gilquin B, Brasilès S, Lancelot N, Letellier G, Gilles M, Dethan G, Orlova EV, Couprie J, Tavares P, Zinn-Justin S. 2009. Structure of bacteriophage SPP1 head-to-tail connection reveals mechanism for viral DNA gating. Proc Natl Acad Sci U S A 106:8507–8512. doi: 10.1073/pnas.0812407106 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Chagot B, Auzat I, Gallopin M, Petitpas I, Gilquin B, Tavares P, Zinn-Justin S. 2012. Solution structure of gp17 from the Siphoviridae bacteriophage SPP1: insights into its role in virion assembly. Proteins 80:319–326. doi: 10.1002/prot.23191 [DOI] [PubMed] [Google Scholar]
- 17. Chaban Y, Lurz R, Brasilès S, Cornilleau C, Karreman M, Zinn-Justin S, Tavares P, Orlova EV. 2015. Structural rearrangements in the phage head-to-tail interface during assembly and infection. Proc Natl Acad Sci U S A 112:7009–7014. doi: 10.1073/pnas.1504039112 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Xiao H, Tan L, Tan Z, Zhang Y, Chen W, Li X, Song J, Cheng L, Liu H. 2023. Structure of the siphophage neck-tail complex suggests that conserved tail tip proteins facilitate receptor binding and tail assembly. PLoS Biol 21:e3002441. doi: 10.1371/journal.pbio.3002441 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Wang C, Duan J, Gu Z, Ge X, Zeng J, Wang J. 2024. Architecture of the bacteriophage lambda tail. Structure 32:35–46. doi: 10.1016/j.str.2023.10.006 [DOI] [PubMed] [Google Scholar]
- 20. Edmonds L, Liu A, Kwan JJ, Avanessy A, Caracoglia M, Yang I, Maxwell KL, Rubenstein J, Davidson AR, Donaldson LW. 2007. The NMR structure of the gpU tail-terminator protein from bacteriophage lambda: identification of sites contributing to Mg(II)-mediated oligomerization and biological function. J Mol Biol 365:175–186. doi: 10.1016/j.jmb.2006.09.068 [DOI] [PubMed] [Google Scholar]
- 21. Huang Y, Sun H, Wei S, Cai L, Liu L, Jiang Y, Xin J, Chen Z, Que Y, Kong Z, Li T, Yu H, Zhang J, Gu Y, Zheng Q, Li S, Zhang R, Xia N. 2023. Structure and proposed DNA delivery mechanism of a marine roseophage. Nat Commun 14:3609. doi: 10.1038/s41467-023-39220-y [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Ayala R, Moiseenko AV, Chen T-H, Kulikov EE, Golomidova AK, Orekhov PS, Street MA, Sokolova OS, Letarov AV, Wolf M. 2023. Nearly complete structure of bacteriophage DT57C reveals architecture of head-to-tail interface and lateral tail fibers. Nat Commun 14:8205. doi: 10.1038/s41467-023-43824-9 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Katsura I, Tsugita A. 1977. Purification and characterization of the major protein and the terminator protein of the bacteriophage lambda tail. Virol (Auckl) 76:129–145. doi: 10.1016/0042-6822(77)90290-2 [DOI] [PubMed] [Google Scholar]
- 24. King J. 1968. Assembly of the tail of bacteriophage T4. J Mol Biol 32:231–262. doi: 10.1016/0022-2836(68)90007-7 [DOI] [PubMed] [Google Scholar]
- 25. Vianelli A, Wang GR, Gingery M, Duda RL, Eiserling FA, Goldberg EB. 2000. Bacteriophage T4 self-assembly: localization of gp3 and its role in determining tail length. J Bacteriol 182:680–688. doi: 10.1128/JB.182.3.680-688.2000 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Lengyel JA, Goldstein RN, Marsh M, Calendar R. 1974. Structure of the bacteriophage P2 tail. Virology (Auckl) 62:161–174. doi: 10.1016/0042-6822(74)90312-2 [DOI] [PubMed] [Google Scholar]
- 27. Grundy FJ, Howe MM. 1985. Morphogenetic structures present in lysates of amber mutants of bacteriophage Mu. Virology (Auckl) 143:485–504. doi: 10.1016/0042-6822(85)90388-5 [DOI] [PubMed] [Google Scholar]
- 28. Parker ML, Eiserling FA. 1983. Bacteriophage SPO1 structure and morphogenesis. I. Tail structure and length regulation. J Virol 46:239–249. doi: 10.1128/jvi.46.1.239-249.1983 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29. Auzat I, Petitpas I, Lurz R, Weise F, Tavares P. 2014. A touch of glue to complete bacteriophage assembly: the tail-to-head joining protein (THJP) family. Mol Microbiol 91:1164–1178. doi: 10.1111/mmi.12526 [DOI] [PubMed] [Google Scholar]
- 30. Katsura I. 1990. Mechanism of length determination in bacteriophage lambda tails. Adv Biophys 26:1–18. doi: 10.1016/0065-227x(90)90004-d [DOI] [PubMed] [Google Scholar]
- 31. Zweig M, Cummings DJ. 1973. Structural proteins of bacteriophage T5. Virol (Auckl) 51:443–453. doi: 10.1016/0042-6822(73)90443-1 [DOI] [PubMed] [Google Scholar]
- 32. McCorquodale J, Warner H. 1988. Bacteriophages T5 and related phagesp 439–476. In The viruses. Plenum Press, New York. [Google Scholar]
- 33. Casjens SR, Hendrix RW. 1974. Locations and amounts of major structural proteins in bacteriophage lambda. J Mol Biol 88:535–545. doi: 10.1016/0022-2836(74)90500-2 [DOI] [PubMed] [Google Scholar]
- 34. Kizziah JL, Manning KA, Dearborn AD, Dokland T. 2020. Structure of the host cell recognition and penetration machinery of a Staphylococcus aureus bacteriophage. PLoS Pathog 16:e1008314. doi: 10.1371/journal.ppat.1008314 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Yu R-C, Yang F, Zhang H-Y, Hou P, Du K, Zhu J, Cui N, Xu X, Chen Y, Li Q, Zhou C-Z. 2024. Structure of the intact tail machine of Anabaena myophage A-1(L). Nat Commun 15:2654. doi: 10.1038/s41467-024-47006-z [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Bárdy P, Füzik T, Hrebík D, Pantůček R, Thomas Beatty J, Plevka P. 2020. Structure and mechanism of DNA delivery of a gene transfer agent. Nat Commun 11:3034. doi: 10.1038/s41467-020-16669-9 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. Desfosses A, Venugopal H, Joshi T, Felix J, Jessop M, Jeong H, Hyun J, Heymann JB, Hurst MRH, Gutsche I, Mitra AK. 2019. Atomic structures of an entire contractile injection system in both the extended and contracted states. Nat Microbiol 4:1885–1894. doi: 10.1038/s41564-019-0530-6 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Degroux S, Effantin G, Linares R, Schoehn G, Breyton C. 2023. Deciphering bacteriophage T5 host recognition mechanism and infection trigger. J Virol 97:e0158422. doi: 10.1128/jvi.01584-22 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39. Stockdale SR, Collins B, Spinelli S, Douillard FP, Mahony J, Cambillau C, van Sinderen D. 2015. Structure and assembly of TP901-1 virion unveiled by mutagenesis. PLoS ONE 10:e0131676. doi: 10.1371/journal.pone.0131676 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40. Seul A, Brasilès S, Petitpas I, Lurz R, Campanacci V, Cambillau C, Weise F, Zairi M, Tavares P, Auzat I. 2021. Biogenesis of a bacteriophage long non-contractile tail. J Mol Biol 433:167112. doi: 10.1016/j.jmb.2021.167112 [DOI] [PubMed] [Google Scholar]
- 41. Li F, Hou C-F, Lokareddy RK, Yang R, Forti F, Briani F, Cingolani G. 2023. High-resolution cryo-EM structure of the Pseudomonas bacteriophage E217. Nat Commun 14:4052. doi: 10.1038/s41467-023-39756-z [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42. Wang Z, Fokine A, Guo X, Jiang W, Rossmann MG, Kuhn RJ, Luo Z-H, Klose T. 2023. Structure of Vibrio phage XM1, a simple contractile DNA injection machine. Viruses 15:1673. doi: 10.3390/v15081673 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43. Thomas JO, Sternberg N, Weisberg R. 1978. Altered arrangement of the DNA in injection-defective lambda bacteriophage. J Mol Biol 123:149–161. doi: 10.1016/0022-2836(78)90318-2 [DOI] [PubMed] [Google Scholar]
- 44. Kandiah E, Giraud T, de Maria Antolinos A, Dobias F, Effantin G, Flot D, Hons M, Schoehn G, Susini J, Svensson O, Leonard GA, Mueller-Dieckmann C. 2019. CM01: a facility for cryo-electron microscopy at the European Synchrotron. Acta Cryst D Struct Biol 75:528–535. doi: 10.1107/S2059798319006880 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45. Li X, Mooney P, Zheng S, Booth CR, Braunfeld MB, Gubbens S, Agard DA, Cheng Y. 2013. Electron counting and beam-induced motion correction enable near-atomic-resolution single-particle cryo-EM. Nat Methods 10:584–590. doi: 10.1038/nmeth.2472 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46. Zhang K. 2016. Gctf: Real-time CTF determination and correction. J Struct Biol 193:1–12. doi: 10.1016/j.jsb.2015.11.003 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47. Tang G, Peng L, Baldwin PR, Mann DS, Jiang W, Rees I, Ludtke SJ. 2007. EMAN2: an extensible image processing suite for electron microscopy. J Struct Biol 157:38–46. doi: 10.1016/j.jsb.2006.05.009 [DOI] [PubMed] [Google Scholar]
- 48. Zivanov J, Nakane T, Forsberg BO, Kimanius D, Hagen WJ, Lindahl E, Scheres SH. 2018. New tools for automated high-resolution cryo-EM structure determination in RELION-3. Elife 7:e42166. doi: 10.7554/eLife.42166 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49. Emsley P, Lohkamp B, Scott WG, Cowtan K. 2010. Features and development of Coot. Acta Cryst D Biol Cryst 66:486–501. doi: 10.1107/S0907444910007493 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50. Adams PD, Afonine PV, Bunkóczi G, Chen VB, Davis IW, Echols N, Headd JJ, Hung L-W, Kapral GJ, Grosse-Kunstleve RW, McCoy AJ, Moriarty NW, Oeffner R, Read RJ, Richardson DC, Richardson JS, Terwilliger TC, Zwart PH. 2010. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Cryst D Biol Cryst 66:213–221. doi: 10.1107/S0907444909052925 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51. Williams CJ, Headd JJ, Moriarty NW, Prisant MG, Videau LL, Deis LN, Verma V, Keedy DA, Hintze BJ, Chen VB, Jain S, Lewis SM, Arendall WB, Snoeyink J, Adams PD, Lovell SC, Richardson JS, Richardson DC. 2018. MolProbity: more and better reference data for improved all-atom structure validation. Protein Sci 27:293–315. doi: 10.1002/pro.3330 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52. Jumper J, Evans R, Pritzel A, Green T, Figurnov M, Ronneberger O, Tunyasuvunakool K, Bates R, Žídek A, Potapenko A, et al. 2021. Highly accurate protein structure prediction with AlphaFold. Nature New Biol 596:583–589. doi: 10.1038/s41586-021-03819-2 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53. Mirdita M, Schütze K, Moriwaki Y, Heo L, Ovchinnikov S, Steinegger M. 2022. ColabFold: making protein folding accessible to all. Nat Methods 19:679–682. doi: 10.1038/s41592-022-01488-1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54. Pandurangan AP, Topf M. 2012. RIBFIND: a web server for identifying rigid bodies in protein structures and to aid flexible fitting into cryo EM maps. Bioinformatics 28:2391–2393. doi: 10.1093/bioinformatics/bts446 [DOI] [PubMed] [Google Scholar]
- 55. Kidmose RT, Juhl J, Nissen P, Boesen T, Karlsen JL, Pedersen BP. 2019. Namdinator - automatic molecular dynamics flexible fitting of structural models into cryo-EM and crystallography experimental maps. IUCrJ 6:526–531. doi: 10.1107/S2052252519007619 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
PDB file of TCPp143 AlphaFold2 predictedmodel fitted into the free density of T5 tail tip.
Figures S1 to S3; Tables S1 and S2.
Data Availability Statement
The cryo-EM density maps of the T5 tail proximal extremity presented in this study and the associated atomic coordinates have been, respectively, deposited in the Electron Microscopy Data Bank (EMDB) and Protein Data Bank (PDB) under the following accession codes: EMD-15967/PDB 8BCP (native state) and EMD-15968/PDB 8BCU (after interaction with FhuA). The unsymmetrized cryo-EM density map of the T5 tail tip has been deposited in the EMDB under accession code EMD-51232. TCPp143 AF2 flexible-fitted model is provided as supplemental material.