Abstract
The immune response is an intricate system that involves the complex connection of cellular and molecular components, each with distinct functional specialisations. It has a distinct capacity to adjust and mould the immune response in accordance with specific stimuli, influenced by both genetic and environmental factors. The presence of genetic diversity, particularly across different ethnic and racial groups, significantly contributes to the impact of incidence of diseases, disease susceptibility, autoimmune disorders, and cancer risks in specific regions and certain populations. Environmental factors, including geography and socioeconomic status, further modulate the variety of the immune system responses. These, in turn, affect the susceptibility to infectious diseases and development of autoimmune disorders. Despite the complexity of the relationship, there remains a gap in understanding the specificity of immune indices across races, immune reference ranges among populations, highlighting the need for deeper understanding of immune diversity for personalized approaches in diagnostics and therapeutics. This review systematically organizes these findings, with the goal of emphasizing the potential of targeted interventions to address health disparities and advance translational research, enabling a more comprehensive strategy. This approach promises significant advancements in identifying specific immunological conditions, focusing on personalized interventions, through both genetic and environmental factors.
Keywords: Immune system, Diversity, Population, Autoimmunity, Cancer, Infection
1. Introduction
The human immune system is a complex network of cells, tissues, and organs that work in concert to defend the body against pathogens [1]. This complicated system exhibits substantial variability among individuals, influencing disease susceptibility, treatment response, and overall health outcomes [2].
Recent research has highlighted the critical importance of understanding immune system diversity. This diversity is shaped by a multitude of factors, including genetics, epigenetics, environmental exposures, and lifestyle [3]. Genetic variations, particularly among different populations, significantly impact disease prevalence in specific regions and the vulnerability of certain populations to various illnesses [4]. For instance, race can serve as a useful indicator for certain diseases when a higher prevalence is observed within specific racial groups [5].
Utilizing technological advancements such as genomics, bioinformatics, and other cutting-edge technologies, numerous studies have demonstrated immunological differences across races, including variations in immune responses, susceptibility to infections, epidemiology of autoimmune diseases, and the characteristics of certain cancers [6,7,8,9]. However, significant gaps persist in our understanding of how these racial differences impact disease diagnosis, treatment decisions, and clinical outcomes, especially regarding test indicators and normal ranges.
By uncovering the complexities of immune diversity, researchers can develop more personalized and effective strategies for disease prevention, diagnosis, and treatment. This review will explore the key factors contributing to immune system diversity, including genetic variations, epigenetic modifications, and environmental influences. Moreover, we will highlight the scarcity of data on immune reference ranges across different racial and geographic populations. Addressing these knowledge gaps will facilitate future research and contribute to a more accurate and personalized understanding of immunological health in diverse populations.
2. Method
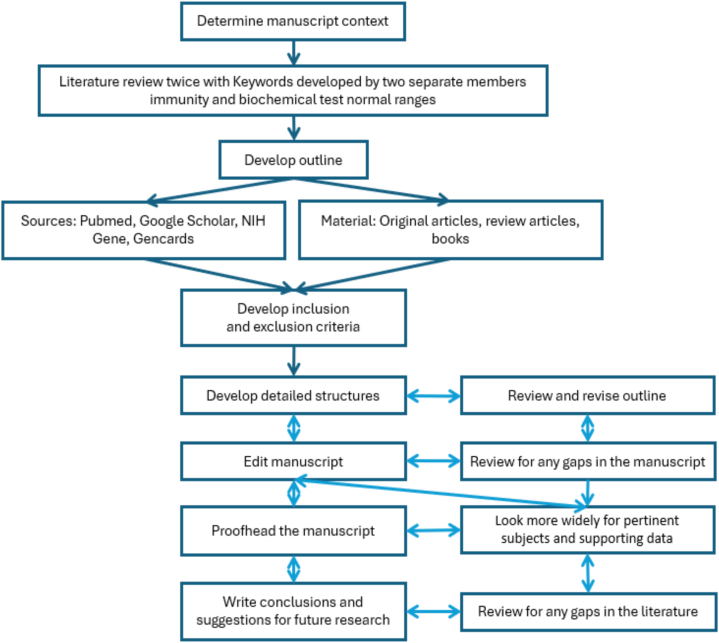
The ideation stage is characterized by an assessment of the existing literature on the topic of interest, as well as producing concepts for the article's fundamental format. The second stage, on the other hand, is distinguished by the ongoing iterative review that occurs in between the stages of review and revised outline, review for any gaps in the manuscript, look more widely for pertinent subjects and supporting data, review for any gaps in the literature (Fig. 1).
Fig. 1.
The flowchart describes the workflow with two distinct stages: the ideation stage and the writing stage.
3. Autoimmune disorders and dysregulated immune responses
3.1. The mechanism behind autoimmunity
Autoimmunity is the phenomenon in which the immune system launches an attack on its own cells. Our adaptive immune system employs B- and T-lymphocytes and their products to generate a targeted response, which, occasionally towards external pathogens. Simultaneously, it also possesses methods to establish tolerance towards "self" antigens, hence preventing an immunological response that attacks the host cells. Autoimmunity arises when this tolerance is disrupted [10]. The corruption of immune system may be initially rooted from, or contributed by, a myriad of both intrinsic and extrinsic factors including genetic predisposition, environmental triggers, and the overall diversity of the immune system (Table 1; Fig. 2).
Table 1.
Summary of important genetic polymorphisms and their related diseases or immune responses.
| Diseases with increased risk | Races | Gene | Gene characteristics | Related Features | References |
|---|---|---|---|---|---|
| Cancer | |||||
| Ovarian cancer | Caucasian | BRCA2 | BRCA2 6174delT | Higher risk familial breast-ovarian cancer | (238) |
| Breast cancer | Caucasian | BRCA1 | BRCA1 185delAG, BRCA1 5382insC | Higher risk familial breast-ovarian cancer | (239) |
| BRCA2 | BRCA2 6174delT | Higher risk familial breast-ovarian cancer | (238) | ||
| Mongoloid | FOXA1 (HNF3A) | Methylation and expression | Poor prognosis - ER (−) | (240) | |
| Prostate cancer | High FOXA1 expression | Poor prognosis | (241) | ||
| Negroid | Missense mutations (F254V and H247L) | Highest incidence, mortality | (242) | ||
| Caucasian | TMPRSS2 | TMPRSS2 mutations | Higher risk | (243) | |
| Caucasian | ERG | ERG mutations | Higher risk | (244) | |
| Skin cancer | Caucasian | SLC24A5 | SLC24A5 rs1426654AA, SLC45A2 rs16891982GG | Higher risk | (245) |
| Autoimmune disease | |||||
| Type I Diabetes | Mongoloid | PTPN22 | C1858T | Higher risk | (246) |
| Caucasian | IFIH1 | rs1990760 | Presents higher rate, during childhood or early adulthood | (247) | |
| Mongoloid | HLA | HLA-DQA1 0101/0102, HLA-DQB1 0301 | Higher risk | (246) | |
| RA | Caucasian | SLC22A4/5 | L503F SNP | Erosive joint damage compared to other ethnicities. higher rates of these autoantibodies | (248) |
| Mongoloid | HLA | HLA-DRB1∗0101 and HLA-DRB1∗0405 | Higher risk | (249) | |
| PTPN22 | C1858T | Higher risk | (246) | ||
| TNFAIP3 | rs6920220 | Higher risk | (250) | ||
| TAGAP | rs394581 | Higher risk | |||
| CCR6 | rs3093023 | Higher risk | |||
| IL-6 | IL-6-174G/C | Higher risk | (251) | ||
| SLC22A4 | slc2F1/slc2F2 SPN | Higher risk | (252) | ||
| SLE | Caucasian | SLC15A4 | rs10847697 and rs1385374 | Higher risk of renal involvement | (253) |
| Negroid | TLRs related genes expression | Elevated TLR3/4/7/8/9 related genes expression | Younger age, more severe symptoms, early mortality | (254) | |
| Mongoloid | GTF2IRD1-GTF2I | 7q11.23 (rs73366469) | Higher risk | (255) | |
| Cystic Fibrosis | Caucasian | CFTR | R553X stop mutation | Higher risk | (256) |
| 1677delTA mutation | Higher risk | (257) | |||
| W1282X | Higher risk | ||||
| F508del | Higher risk | ||||
| W1316X mutation | Higher risk | (258) | |||
| Negroid | DF508 mutation | Higher risk and chronic respiratory failure | |||
| G551D mutation | Higher risk | ||||
| Kawasaki disease | Mongoloid | ORAI | rs3741596 | Higher risk | (259) |
| Infectious disease | |||||
| COVID-19 | Mongoloid | LZTFL1 | rs17713054 | Doubles the risk of respiratory failure and death | (260) |
| Malaria | Negroid | FCGR2B | Minor allele frequency (MAF) of rs1050501 | Substantial protection against severe malaria | (261) |
| Caucasian | FCGR2B | Weak resistance to Malaria | |||
| Caucasian | ATP2B4 | ATP2B4 variations | Weak resistance to Malaria | ||
| ABO genes | ABO genes variations | Weak resistance to Malaria | |||
| Caucasian | ACKR1 | Third mutation, unstable protein (Arg89Cys: cytosine - > thymidine at position 265) | Plasmodium vivax susceptibility | (262) | |
| Negroid | Single T to C substitution at nucleotide −33 | Plasmodium vivax resistance | (263) | ||
| Caucasian | G6PD | G6PD Mediterranean | Reduce the parasite load faster | (264) | |
| Negroid | G6PD Mediterranean | Reduce the parasite load faster | (265) | ||
| HIV | Caucasian | CCR5 | CCR5-Delta32 homozygous | Protection against HIV-1 | (266) |
| CCR5-Delta32 heterozygous | Reduced risk infected with HIV-1 | ||||
| Mongoloid | HLA | HLA-B57 and HLA-B27 | Maintain viral load without treatment | (267) | |
| Negroid | Maintain viral load without treatment | ||||
| Negroid | TRIM5α | TRIM5α 136Q | Stronger resistance again HIV | (268) | |
| Negroid | APOBEC3G | H186R in exon 4 | Virus replication prevention | (269) | |
| Negroid | KIR - HLA interaction | KIR3DS1 - HLA-B Bw4-80Ile | Reducing the risk of infection or slowing the progression of disease | (270) | |
| Tuberculosis | Mongoloid | NRAMP1 | Asp543Asn | Higher risk or natural resistance | (271) |
| Negroid | Higher risk or natural resistance | ||||
| Mongoloid | NRAMP1 | 3′ UTR | Protection against infection | (272) | |
| Negroid | Protection against infection | ||||
| Mongoloid | NRAMP1 | INT4 | Protection against infection | (273) | |
| Negroid | Protection against infection | (274) | |||
Fig. 2.
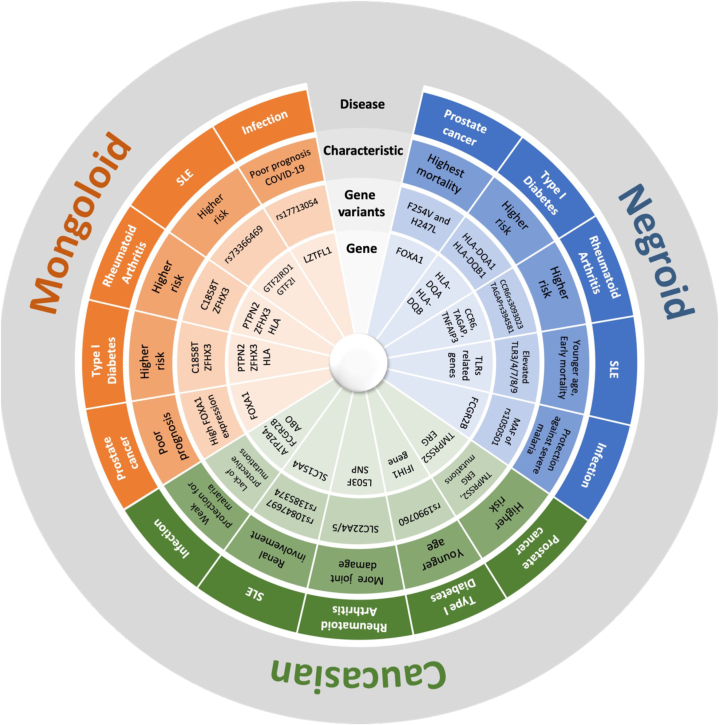
The Involvement of Genetic Factors in Racial and Ethnic Differences in Diseases.
A. Prostate Cancer: FOXA1 influences androgen receptor signaling critical for prostate cancer cell survival. Mongoloids have poor outcomes with FOXA1 overexpression, Negroids show high incidence due to FOXA1 mutations, and Caucasians have increased risk linked to TMPRSS2 and ERG mutations [241,275].
B. Type I Diabetes: T1DM involves B cell destruction and varies by race, with unique genetic risk markers: C1858T, ZFHX3, and HLA in Mongoloids; HLA-DQA103:01-HLA-DQB102:01 in Negroids; and rs1990760 and IFIH1 in Caucasians [246,276,277].
C. Rheumatoid Arthritis (RA) and Systemic Lupus Erythematosus (SLE): Caucasians face severe RA linked to SLC22A4/5 SNPs, while African individuals often experience earlier onset and more severe SLE due to elevated TLR expression [254,278].
D. Infections: Mongoloids have doubled risks of severe COVID-19 due to rs17713054 and LZTFL1. Negroids are more protected against severe malaria linked to rs1050501, while Caucasians show weak malaria resistance associated with the Fc-GammaR pathway [260,261,279,280].
3.1.1. The breakdown of immune central and peripheral tolerance
The mechanisms, by which autoimmune conditions are generated, are complicated. The breakdown of immune central and peripheral tolerance can contribute significantly to the rise of autoimmune disorders. In the thymus, developing T cells undergo selection processes to eliminate those that recognize self-antigens with high affinity (negative selection) [11]. Autoimmune regulator gene (AIRE) on the chromosome 21 plays the key role in this complex process. It is a transcription factor that plays a crucial role in establishing immune tolerance, specifically in the thymus. It is primarily expressed in medullary thymic epithelial cells (mTECs) [12]. AIRE enables mTECs to express a wide array of tissue-specific antigens (TSAs) that are usually restricted to peripheral tissues (e.g., insulin from the pancreas, thyroid-specific proteins). By promoting the presentation of these TSAs on MHC molecules, AIRE exposes developing T cells to a broader range of self-antigens, thereby allowing the immune system to identify and eliminate T cells that react strongly to these antigens [13]. When developing T cells strongly bind to these TSA-MHC complexes, they undergo apoptosis through clonal deletion. This process ensures that potentially self-reactive T cells, which could cause autoimmune diseases, are removed during thymic development [14]. When these “errorneous” cells escape this negative selection, peripheral tolerance emerges and acts as a secondary defense mechanism, which happens in the peripheral tissues and lymphoid organs, involving regulatory T cells (Tregs), apoptosis of self-reactive lymphocytes, and mechanisms that induce anergy (functional inactivation) [15]. Tregs play a crucial role in suppressing autoreactive T cells in peripheral tissues. Tregs are primarily derived from the thymus and express the transcription factor FOXP3, which is crucial for their development and function [16]. FOXP3+ Tregs can inhibit the ability of dendritic cells (DCs) and other antigen-presenting cells (APCs) to activate effector T cells, partly through interactions involving the immune checkpoint CTLA-4 (Cytotoxic T-Lymphocyte Associated Protein 4). CTLA-4, expressed by Tregs, competes with CD28 for binding to the costimulatory molecules CD80 and CD86 on APCs, leading to reduced T-cell activation [17]. Beside inhibiting T-cells via immune checkpoints, these T-regs also secrete cytokines such as IL-10, TGF-β, and IL-35, which inhibit the activity of effector T cells (e.g., Th1, Th2, Th17 cells) and reduce inflammation [18]. Escaped self-reactive lymphocytes undergo apoptosis via the activation of FAS receptor (CD95/APO-1) and FASL (FAS ligand), which are a surface a death receptor and its ligand expressed on T-cells and NK cells [19]. When binding, they lead to recruitment and activation of caspase-8, which in turn leads to apoptosis, characterized by DNA fragmentation, membrane blebbing, and eventual cell death of the overactive immune cells [20]. Anergy in peripheral T cells is induced when T cells recognize self-antigens without receiving the necessary co-stimulatory signals from APCs (e.g., CD28 binding to B7). This lack of a second signal leads to the expression of genes like EGR2 and EGR3 that suppress T-cell activation [21]. They also upregulate inhibitory receptors like PD-1 (Programmed cell death protein 1), which further suppresses their activation and function, preventing autoimmunity [22]. Failure of any of the components along the process of central or peripheral tolerance may lead to the rise of autoimmune conditions.
3.1.2. Loss of immune regulation
Loss of immune regulation is among the mechanisms inducing autoimmunity, which may manifest as inappropriate expression of cytokine genes or the epigenetic changes that come as an aftermath. Genetic diversity in cytokine genes significantly influences the regulation of immune responses by affecting the production, secretion, and receptor interactions of cytokines. For instance, single nucleotide polymorphisms (SNPs) - variations in a single nucleotide that occur at a specific position in the genome - in cytokine genes can alter cytokine expression levels or receptor binding affinity, impacting immune regulation [23]. Specifically, SNPs in the promoter region of the IL-10 gene, such as −1082A/G, −819T/C, and −592A/C, can affect the transcriptional activity of the gene [24]. IL-10 is an anti-inflammatory cytokine that helps regulate immune responses by inhibiting pro-inflammatory cytokines like TNF-α and IL-1β [25]. Variations in IL-10 levels can therefore impact the balance between pro- and anti-inflammatory responses [25]. The IL-23R (IL-23 receptor), is part of the receptor complex that binds IL-23, a cytokine crucial for the maintenance and expansion of Th17 cells. Th17 cells produce IL-17, a pro-inflammatory cytokine that plays a key role in the immune response against extracellular pathogens and in the pathogenesis of several autoimmune conditions [26].
3.1.3. Epigenetics
Aside from genetic, epigenetic factor also contribute to the manifestation of autoimmune conditions. Epigenetics contributes significantly to the emergence of autoimmune diseases by modifying gene expression without altering the underlying DNA sequence. Alterations in DNA methylation patterns can influence the expression of genes involved in immune regulation. It refers to the addition of a methyl group to cytosine bases in DNA, typically at CpG dinucleotides, which represses gene transcription by modifying the chromatin structure [27]. Histone modification is another process that promotes alteration in chromatin structure, therefore can lead to autoimmunity. They include the processes of acetylation, methylation, phosphorylation, and ubiquitination of translational products [28]. Finally, non-coding RNAs (ncRNAs) - functional RNA molecule that is not translated into a protein, particularly microRNAs (miRNAs) and long non-coding RNAs (lncRNAs), play crucial roles in the regulation of immune responses and have been implicated in the pathogenesis of autoimmune diseases [29]. The complex interactions between these epigenetic modifications and genetic predispositions highlight the importance of understanding epigenetic factors to develop therapeutic interventions for autoimmune conditions.
3.2. How immunological diversity manifests in autoimmune diseases
3.2.1. Autoimmune regulator gene (AIRE)
As mentioned, the dysfunction of AIRE may lead to peripheral and central tolerance breakdown. The mutation of AIRE, including a wide range of nonsense and missense mutations in the coding regions, induce an autoimmune condition named Autoimmune polyendocrinopathy candidiasis-ectodermal dystrophy (APS-1 or APECED) - a rare, monogenic autoimmune condition that typically presents in childhood [30]. APS-1 is characterized by a triad of chronic mucocutaneous candidiasis (fungal infections), hypoparathyroidism, and adrenal insufficiency (Addison's disease). Patients often develop additional autoimmune conditions, including type 1 diabetes and autoimmune thyroiditis – which are the aftermaths of autoreactive T cells [31].
The immunologic diversity is expressed through the multivariants of AIRE mutation. Different mutations were seen in different populations, which displays the multicolor picture of APS-1 presentation. For example, the R257X mutation, the most common mutation of AIRE is particularly common in Finland, Norway, and other Scandinavian countries. It is the most frequently observed mutation in these regions, representing over 80 % of AIRE mutations in Finnish APS-1 patients [32,33]. This mutation is associated with a classic APS-1 phenotype, characterized by a triad of chronic mucocutaneous candidiasis, hypoparathyroidism, and adrenal insufficiency. Finnish patients tend to exhibit these typical symptoms of APS-1 starting in childhood [34]. The AIRE R139X mutation is frequently seen in APS-1 patients from Italy and Sardinia, with a prevalence of around 50 % among APS-1 cases in these regions, with a clinical presentation similar to that of the R257X mutation [35]. Meanwhile, the Y85C mutation is highly prevalent among Iranian Jewish individuals, representing the most common AIRE mutation in this specific group. This mutation accounts for over 90 % of APS-1 cases in this population. This mutation is associated with a severe APS-1 phenotype. The high prevalence of this mutation in a relatively isolated population suggests a founder effect, where the mutation spread from a small ancestral population [36].
3.2.2. Interleukin-2 receptor gamma (IL2RG)
Severe Combined Immunodeficiency Syndrome (SCID), one of the inborn errors of immunity, may also arise from various autoimmune defects. The most common cause of SCID are mutations in the IL2RG gene located in the X chromosome; hence the disorder is termed X-linked SCID (X-SCID) [37]. IL2RG encodes the interleukin-2 receptor gamma chain (γc), also known as CD132, which is a critical component of multiple cytokine receptors, including those for interleukin-2 (IL-2), IL-4, IL-7, IL-9, IL-15, and IL-21. These cytokines are vital for the development, differentiation, and function of immune cells, such as T cells, B cells, and NK cells. The role of IL2RG is especially important in maintaining immune homeostasis and preventing autoimmune responses [38,39]. Mutations of the gene greatly break the balance and homeostasis of inner immunological environment, characterized by a near-complete absence of T cells and NK cells, and non-functional B cells. X-SCID patients are highly vulnerable to infections due to their immunodeficiency. While autoimmunity is not the primary presentation, the lack of Tregs and functional immune regulation can lead to dysregulated immune responses if partial immune function remains. The absence of proper immune regulation can manifest as autoimmunity or immune dysregulation in some cases [40].
The different variants of the gene can also lead to disparities in clinical phenotypes. For example, nonsense and frameshift mutations of the gene often cause a complete loss of IL2RG function, leading to classic X-linked SCID (X-SCID), which is characterized by a near-complete absence of T cells and NK cells and non-functional B cells (T– B + NK– phenotype). This classic presentation results in severe immunodeficiency, making patients highly vulnerable to life-threatening infections from infancy [41]. Meanwhile, partically dysfunction of the protein can be seen in milder forms of mutations, including missense variants. The impact of missense mutations depends on their location within the IL2RG protein. For instance, missense mutations in critical domains (e.g., the extracellular or transmembrane domains) can significantly impair receptor assembly or signaling, resulting in a phenotype similar to classic SCID; however, missense mutations in less critical regions (e.g. T121N) may result in hypomorphic or leaky mutations, where some residual IL2RG function is retained. This leads to a milder phenotype called atypical or leaky SCID, where patients may have low numbers of T cells or NK cells, but some functional immune responses remain, resulting in a later onset of symptoms and a more varied clinical presentation, including autoimmunity and milder infections [42].
Not only IL2RG, but SCID can also be caused by other genetic changes, mostly autosomal recessive ones. These include IL2RG, JAK3, DCLRE1C, RAG1, RAG2, IL7R, ADA, CD3D, CD3E, CD3Z, DOCK2, AK2, and TTC7A. Interestingly, studies have shown that the genotype manifestations not only lead to differences in SCID phenotypes, but they may also have different regional distributions [41,43,44,45]. Native American populations, such as the Navajo, was found to exhibit SCID due to an exclusive mutation of DCLRE1C gene, or Artemis mutation [46]. The phenotype of this mutation is called Athabascan SCID (SCIDA). It is an autosomal recessive pattern, commonly seen in Athabascan-speaking Native Americans. It is characterized by a deficiency of both T and B cells, while NK cells remain present, resulting in a T(−)B(−)NK(+) immunodeficiency [46]. In another setting, SCID prevalence remains high among Middle East and Northern Africa (MENA) regions due to the autosomal recessiveness in heredity, which mostly results from the common practice of consanguinity [47,48,49,50]. In MENA populations, autosomal recessive mutations of RAG1/RAG2 genes account for the majority of SCID cases, which manifests a founder effect on these populations [51].
3.2.3. Interleukin-23 (IL-23)
The elevation of IL-23 level is known to have a close association with Crohn's disease, an autoimmune disorder that mainly affects gastrointestinal systems [52]. The polymorphism in IL-23R can either upregulate or downregulate the risk of developing Crohn's disease, which was widely studied. The rs11209026 (Arg381Gln) variant, a missense SNP in the IL-23R gene has been known to be associated with a reduced risk of Crohn's disease; on the other hand, the rs10889677 (G/A in 3′ UTR) variant has been associated with an increased risk of Crohn's disease and ulcerative colitis [53,54]. The Arg381Gln polymorphism results in a structural change in the IL-23 receptor, which leads to altered signal transduction through the JAK/STAT pathway, particularly STAT3 activation. The reduced signaling decreases the differentiation and survival of Th17 cells, leading to a lower production of IL-17 and IL-22, thereby reducing inflammation and the risk of autoimmune responses [55]. On the contrary, variations in the 3′ UTR can influence the stability of mRNA and, subsequently, the expression levels of the IL-23 receptor [55].
3.2.4. Epigenetics
Epigenetic modification may also greatly alter the manifestation of systemic disorders. Systemic Lupus Erythematosus (SLE) is among those that possesses established evidence of the mechanisms. In SLE, T cells exhibit global DNA hypomethylation, especially in CpG islands of certain genes involved in immune activation. This hypomethylation leads to overexpression of immune activation genes, such as ITGAL (CD11a) and CD40L, which contribute to enhanced T cell activation, increased production of autoantibodies, and subsequent immune system attack on self-tissues. CD40L, in particular, plays a role in B cell activation and differentiation, leading to the production of pathogenic autoantibodies. Hypomethylation-induced overexpression of CD40L enhances B-T cell interactions, thereby contributing to autoimmunity [56,57]. Moreover, abnormal histone modifications, particularly histone acetylation, are closely linked to the dysregulation of immune genes. Histone acetylation is generally associated with transcriptional activation, and in SLE, it contributes to the overexpression of pro-inflammatory genes, leading to a breakdown of immune tolerance [58]. Hyperacetylation of histones H3 and H4 at the promoter regions of pro-inflammatory genes like IFN-α and TNF-α in immune cells, particularly dendritic cells. This epigenetic alteration results in increased expression of these cytokines, contributing to the chronic inflammatory environment that drives lupus pathogenesis [59].
HOTAIR (HOX transcript antisense intergenic RNA) is a lncRNA involved in chromatin remodeling. In SLE, HOTAIR is overexpressed in peripheral blood mononuclear cells (PBMCs). This overexpression is associated with the downregulation of TNF-α-induced protein 3 (TNFAIP3), an anti-inflammatory molecule that plays a role in inhibiting NF-κB signaling. Reduced TNFAIP3 expression leads to prolonged activation of NF-κB, resulting in increased inflammation and autoimmunity [60,61]. Another miRNA playing an crucial role in SLE is miR-155, a pro-inflammatory miRNA that plays a role in the activation of B cells, T cells, and macrophages [62]. It is involved in regulating the immune response by targeting SHIP1 and SOCS1, both negative regulators of immune signaling pathways [63]. miR-155 is upregulated in T cells, B cells, and dendritic cells in SLE, leading to hyperactivation of these cells. Increased miR-155 levels promote the production of autoantibodies by B cells, contributing to the autoimmune response. This miRNA has been shown to enhance T cell activation and survival, further driving autoimmunity [64,65]. This has led to the implication of the blockage of miR-15 in preclinical models, and results have shown reduction in autoimmune activity in SLE by dampening the activation of immune cells involved in the disease [66].
Another lncRNA that has been studied in SLE is growth arrest-specific 5 (GAS-5), a long lncRNA associated with multiple regulatory functions, particularly in immune responses, cellular apoptosis, and inflammatory processes [67]. GAS-5 is located on chromosome 1q25.1, hosting several small nucleolar RNAs (snoRNAs) within its introns. It is crucial in regulating the glucocorticoid receptor (GCR) pathway. By mimicking the glucocorticoid response elements, GAS-5 modulates the sensitivity of immune cells to glucocorticoids, which are commonly used in SLE treatment [68]. Genetic polymorphisms in GAS-5, such as the rs145204276 variant, have been identified as protective against SLE. Individuals with the ID + genotype of this polymorphism exhibit higher GAS-5 expression levels and reduced disease severity [69]. Interestingly, GAS-5 levels in CD4+ T cells are elevated in SLE patients with specific clinical manifestations, such as pleurisy and rashes, as well as in those with positive anti-dsDNA antibodies and low complement C3 levels. This paradoxical increase highlights its potential as a biomarker for SLE diagnosis and disease activity monitoring. Elevated GAS-5 in these contexts may reflect a compensatory mechanism attempting to restore immune balance [70].
The data elucidates the multifaceted mechanisms underlying autoimmunity, highlighting the complex interactions between genetic predispositions, epigenetic modifications, and environmental influences in the disruption of immune tolerance. Through examining diverse pathways - such as central and peripheral tolerance, immune regulation via cytokine signaling, and population-specific genetic variations - the heterogeneity of autoimmune disease etiology is highlighted. Notably, gaps in current literature persist, particularly regarding the precise role of epigenetic modifications in autoimmune pathogenesis, as well as the impact of genetic diversity on disease presentation across populations. These areas warrant further investigation to inform the development of more precise, individualized therapeutic interventions [71].
4. The influence of immune diversity on cancer
4.1. The mechanisms behind immune system alterations in cancer development
The development of cancer involves a complex contribution of genetic, epigenetic, and environmental factors that can alter immune system function, leading to a breakdown in the body's ability to detect and destroy malignant cells [72,73]. These factors impact various components of the immune system, from the genes regulating immune responses to the microenvironment in which immune cells operate [74,75].
4.1.1. Genetic mutations
Genetic mutations play a crucial role in shaping immune responses against cancer [76] (Table 1; Fig. 2). For example, mutations in oncogenes such as KRAS or MYC lead to excessive cell proliferation, which not only drives cancer development but also overwhelms the immune system, making it more difficult for immune cells to recognize and eliminate tumor cells [77]. At the same time, tumor suppressor genes like TP53, often called the “guardian of the genome”, are critical for maintaining normal immune surveillance [78]. When TP53 is mutated, cancer cells can avoid apoptosis (programmed cell death) and proliferate unchecked [79]. This loss of tumor suppressor function is a common feature in many cancers, from lung to breast cancer, and directly contributes to immune evasion [80]. Moreover, mutations in immune-regulatory genes can significantly weaken immune defenses [81]. For instance, mutations in the HLA (human leukocyte antigen) system, which is responsible for presenting antigens to T-cells, may impair antigen presentation, allowing cancer cells to "hide" from the immune system [82]. Additionally, alterations in immune checkpoint pathways, such as PD-1/PD-L1 and CTLA-4, can inhibit the immune system's ability to attack tumors [83]. Cancer cells often exploit these pathways by overexpressing PD-L1, which binds to PD-1 receptors on T-cells and suppresses their activity, thereby preventing an effective immune response [84].
4.1.2. Epigenetic changes
Epigenetic changes, which do not alter the DNA sequence but affect gene expression, play a significant role in modulating immune responses in cancer [85]. DNA methylation, one of the most studied epigenetic mechanisms, often leads to the silencing of critical genes involved in immune activation and tumor suppression [86]. For instance, hypermethylation of genes that regulate the immune system, such as MHC class I molecules, results in decreased antigen presentation on tumor cells, allowing them to escape immune recognition [87]. This mechanism is seen in various cancers, including colorectal and lung cancer, where the epigenetic silencing of MHC molecules prevents cytotoxic T-cells from identifying and destroying cancer cells [88]. Histone modifications, another epigenetic mechanism, also influence immune responses by changing the chromatin structure and gene accessibility [89]. For example, in some cancers, histone deacetylases (HDACs) reduce the expression of genes responsible for activating immune responses, such as cytokines and chemokines, which recruit and activate immune cells at the tumor site [90]. Drugs that inhibit HDACs are being investigated as potential therapies to restore immune function and boost anti-tumor immunity [91].
4.1.3. MicroRNAs
MicroRNAs (miRNAs) are small, non-coding RNAs that regulate gene expression post-transcriptionally. Abnormal miRNA expression in cancer can affect immune regulation [92]. For instance, overexpression of miR-155 has been associated with immune suppression by downregulating immune-checkpoint inhibitors, leading to increased immune evasion by tumor cells [93]. In some cancers, such as lymphoma, miR-155 overexpression contributes to a suppressive immune microenvironment that promotes tumor progression [94].
4.1.4. Environmental influences
Environmental influences such as chronic inflammation, carcinogens, and alterations in the microbiome profoundly impact the immune system's capacity to prevent or promote cancer [95].
4.1.4.1. Chronic inflammation
Chronic inflammation is one of the most well-documented risk factors for cancer [96]. Conditions like inflammatory bowel disease (IBD) and hepatitis lead to a continuous state of immune activation, which eventually exhausts immune cells, particularly T-cells and macrophages [97]. Inflammation produces cytokines such as IL-6 and TNF-α, which promote DNA damage in epithelial cells and support a pro-tumorigenic environment [98]. As immune cells become dysfunctional over time, they may fail to eliminate cells that have accumulated oncogenic mutations [99]. Chronic inflammation is particularly associated with cancers such as colorectal cancer (from IBD) and liver cancer (from chronic hepatitis infection) [100].
4.1.4.2. Carcinogens
Exposure to carcinogens including tobacco smoke, ultraviolet radiation, and industrial chemicals can directly damage DNA and immune cells [101]. For example, tobacco smoke contains compounds that induce mutations in both lung epithelial cells and immune cells, impairing the latter's ability to clear pre-cancerous and cancerous cells [102]. In skin cancer, UV radiation not only causes direct DNA mutations but also suppresses local immune responses by inducing immunosuppressive cytokines like TGF-β, which promotes immune tolerance to the growing tumor [103].
4.1.4.3. Microbiome
The microbiome, particularly the gut microbiome, has emerged as a key regulator of immune function [104]. Dysbiosis, or an imbalance in microbial species, can promote a pro-inflammatory or immunosuppressive environment [105]. In colorectal cancer, for instance, an increase in Fusobacterium nucleatum has been linked to immune evasion by cancer cells [106]. This bacterium can inhibit the activity of natural killer (NK) cells and cytotoxic T-cells, which are essential for immune-mediated cancer control [107]. Similarly, microbial metabolites such as short-chain fatty acids can modulate the immune system by affecting the function of T-cells and regulatory T-cells (Tregs), either promoting anti-tumor immunity or fostering a suppressive environment that favors tumor growth [108].
The relationship between genetic, epigenetic, and environmental factors is often intertwined, where one influences the other [109] (Fig. 3). For example, individuals with genetic predispositions, such as mutations in DNA repair genes like BRCA1 or BRCA2, are more vulnerable to environmental carcinogens, as their cells are less able to repair damage caused by toxins [110]. This interaction leads to an increased risk of both genetic mutations and immune dysregulation, further compounding cancer risk [111]. Additionally, environmental factors can induce epigenetic modifications that affect immune regulation [112]. For instance, cigarette smoke has been shown to cause DNA methylation changes in immune cells, leading to the suppression of key immune-regulatory genes [113]. These epigenetic alterations reduce the immune system's ability to detect and destroy cancer cells, contributing to the progression of cancers like lung cancer [114].
Fig. 3.
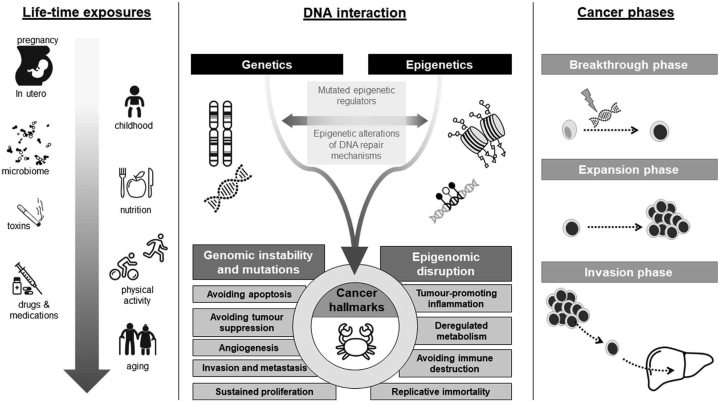
Epigenetic influence through a lifetime of exposure, DNA interactions and cancer development.
Epigenetic alterations occur throughout a human life, from in utero exposure (left panel) until ageing processes start. The DNA interaction (middle panel) that occurs are mutually interacting and influencing the cancer hallmarks for cancer risk and progression, for which current research seeks to explore the exact mechanisms a contribution throughout the phases (right panel) of cancer development. The illustration is generic and unique patterns of exposure, risk and influence may occur to each specific cancer type [281]. Source from: Drake TM and Søreide K. Cancer epigenetics in solid organ tumors: A primer for surgical oncologists. European Journal of Surgical Oncology. 2019; 45(5):736-46.
4.2. The role of immune response variability in cancer incidence across populations
The Global Database on Cancer Incidence offers valuable insights into disparities in cancer rates among populations worldwide, underscoring the influence of genetics, lifestyle, and environmental exposures on cancer risk. For instance, lung cancer incidence among men in high-income countries like the United States reaches approximately 57.3 per 100,000, with particularly high rates among African American men compared to other racial groups [115]. Similarly, breast cancer incidence in developed nations, including the United States and Western Europe, is reported at about 90–120 per 100,000 among women, whereas in several African and Asian regions, rates are lower at 20–40 per 100,000 [116]. Colorectal cancer also shows marked differences; Western nations report incidence rates of 40–50 per 100,000, which contrasts with significantly lower rates observed in many developing regions [117]. Among prostate cancer cases, African American men experience the highest incidence globally, with rates reaching up to 120 per 100,000, attributed in part to genetic factors and healthcare access disparities [118]. Lastly, hepatocellular carcinoma (HCC) rates are notably higher in East Asia and Sub-Saharan Africa, exceeding 30–40 per 100,000 due to risk factors like chronic hepatitis B infection and aflatoxin exposure [119]. This variation in incidence rates across different demographics underscores the critical need for targeted prevention strategies informed by population-specific data.
4.2.1. Lung cancer
Lung cancer, as one of the leading causes of cancer mortality worldwide, demonstrates significant differences in incidence patterns across populations, largely shaped by variations in smoking habits, environmental exposures, and genetic susceptibilities [115]. In high-income nations such as the United States, lung cancer remains prevalent among men, closely linked to high smoking rates, prolonged exposure to occupational carcinogens, and historical air quality concerns [116]. Comparatively, certain Asian populations, despite having lower overall smoking rates, exhibit elevated lung cancer risk due to a higher prevalence of epidermal growth factor receptor (EGFR) mutations, which are commonly found in non-smokers. These mutations drive tumor growth through cell proliferation and survival pathways, making EGFR-targeted therapies particularly relevant in these groups [120]. This contrast highlights how, in Western populations, lung cancer risk is predominantly environmentally driven, whereas in Asian populations, genetic mutations contribute more significantly, especially among non-smokers [121]. Outcomes and treatment options also vary among populations due to these genetic and environmental differences. For instance, EGFR-targeted therapies, including tyrosine kinase inhibitors (TKIs) like osimertinib, show higher efficacy in patients with EGFR mutations—predominantly seen in East Asian populations—leading to improved survival rates in these individuals compared to standard chemotherapy [122]. In contrast, Western populations benefit more broadly from immune checkpoint inhibitors, such as PD-1 and PD-L1 inhibitors, which have shown enhanced efficacy in patients with smoking-related lung cancers that tend to carry high mutational burdens [123]. Screening strategies also reflect population-specific risks. In the U.S. and other high-income countries, low-dose computed tomography (LDCT) screening is recommended for high-risk individuals, primarily heavy smokers aged 50 to 80, to detect lung cancer at earlier stages and improve survival outcomes [124]. However, in regions with a high prevalence of EGFR mutations among non-smokers, there is a growing interest in genetic testing and personalized screening approaches tailored to identify at-risk individuals who may not qualify for traditional smoking-related screenings [122]. These population-specific treatment and screening strategies highlight the necessity of a nuanced approach to lung cancer management, integrating environmental and genetic risk factors for optimal outcomes across diverse demographics.
4.2.2. Breast cancer
Similar to lung cancer, breast cancer incidence highlights the complex interplay of lifestyle, genetic, and environmental factors that shape cancer risk across diverse populations [116]. In high-income countries, including the United States and Western Europe, breast cancer incidence rates are substantially higher, influenced by lifestyle factors like dietary patterns, delayed childbearing, lower parity, and increased use of hormone replacement therapy, which are associated with prolonged estrogen exposure [125]. By contrast, many Asian and African nations report lower incidence rates, a difference partly attributed to differing reproductive behaviors, lower obesity rates, and generally reduced alcohol consumption, which are protective factors against breast cancer development [126]. One possible explanation for higher breast cancer rates in specific groups is a higher frequency of mutations in the breast cancer susceptibility genes BRCA1 and BRCA2, particularly among Ashkenazi Jewish individuals. These mutations occur in 2.0%–2.5 % of Ashkenazi Jewish men and women—approximately five times the frequency found in the general population [127]. Outcomes and treatment approaches for breast cancer also vary between populations due to differences in tumor biology, genetic factors, and healthcare access. For instance, in some African American and Hispanic populations in the United States, breast cancer tends to present at more advanced stages and with more aggressive subtypes, such as triple-negative breast cancer (TNBC), leading to poorer prognoses and limited treatment options [128]. Targeted therapies, such as HER2 inhibitors and hormone therapies, have markedly improved outcomes in populations where tumors express specific receptors, like HER2 and estrogen receptors; however, TNBC, which lacks these receptors, requires alternative approaches such as chemotherapy or emerging immunotherapy options [129]. Screening strategies are also adapted based on population-specific risks and healthcare resources. In developed countries, mammography is widely used, especially for women aged 50 and above, to detect breast cancer at earlier, more treatable stages, resulting in improved survival rates [130]. In contrast, countries with fewer resources may rely on clinical breast exams or opportunistic screening, which, although less sensitive, can still help in early detection. Furthermore, for high-risk groups, such as those with BRCA mutations, personalized screening regimens that include MRI alongside mammography are often recommended to enhance early detection, highlighting the importance of an individualized approach to breast cancer screening [131]. Tailoring breast cancer management based on genetic predisposition, environmental influences, and healthcare resources is essential to addressing disparities and improving outcomes across diverse populations.
4.2.3. Colorectal cancer
Building on the population-specific influences seen in lung and breast cancers, colorectal cancer (CRC) similarly demonstrates significant geographic and population-based variation due to a complex interplay of dietary, genetic, and lifestyle factors. In Western nations, the risk of CRC is higher, driven largely by dietary habits such as high consumption of red and processed meats, which are associated with increased carcinogenic compounds in the colon, and low intake of fiber, which reduces beneficial gut microbiota diversity and protective bowel transit times [132]. In contrast, lower CRC rates in many developing regions correlate with diets rich in plant-based foods, which may confer protective effects due to higher fiber and antioxidant intake [133]. Sporadic CRC, which accounts for about 75 % of cases, arises without an apparent inherited link, while familial CRC comprises roughly 25 %, indicating a genetic contribution, shared exposures, or both [134]. Countries like Western Europe, North America, and Australia have high CRC incidence rates, while regions such as sub-Saharan Africa, South America, and Asia show lower rates. However, CRC rates are rising in developing regions adopting Westernized diets and lifestyles [135]. Mechanistically, CRC progression is influenced by two major genomic instability pathways: chromosomal instability (CIN) and microsatellite instability (MIN), which contribute to mutations in key genes like APC, p53, and SMAD4, promoting tumorigenesis [136]. Inherited syndromes, such as Familial Adenomatous Polyposis (FAP), which involves APC gene mutations, further underscore the genetic contribution to CRC risk, especially in familial cases [137]. Outcomes and treatment options for CRC vary across populations due to these risk differences. In Western countries, earlier diagnosis through widespread screening programs, such as colonoscopy and fecal immunochemical tests (FIT), has improved CRC survival rates by enabling detection at earlier stages [138]. In regions with limited healthcare access, CRC tends to be diagnosed at more advanced stages due to a lack of routine screening and lower public awareness, which worsens outcomes and limits treatment options [117]. For high-risk groups, including those with a family history of CRC or known genetic predispositions, personalized screening protocols are often recommended, sometimes starting earlier in life or occurring at more frequent intervals to manage risk more effectively [139].
4.2.4. Prostate cancer
Extending the theme of geographic and population-specific differences in cancer, prostate cancer also exhibits notable disparities in incidence and outcomes across populations. Prostate cancer rates are highest in North America, with African American men experiencing particularly elevated incidence and mortality rates, making them one of the most affected demographic groups globally [115]. For example, men of West African descent, including African Americans, have a 1.7 times higher incidence of prostate cancer and a 2.4 times greater mortality rate than European Americans, with African American men specifically showing a high mortality rate of 37.4 per 100,000, significantly exceeding the rates seen in non-Hispanic whites (19.3 per 100,000) and other ethnic groups [140]. Molecular variations such as PTEN deletions and SPINK1 overexpression, which are more common in African American men, are associated with higher Gleason scores and aggressive disease, likely contributing to poorer outcomes in this population [140]. Additionally, environmental and lifestyle factors, such as high-fat diets and sedentary lifestyles typical in Western societies, exacerbate prostate cancer risk, possibly due to increased levels of circulating androgens and insulin-like growth factors, which can stimulate tumor growth [141]. Conversely, men in Asian countries generally have lower prostate cancer rates, potentially due to a diet rich in phytoestrogens from soy products, which may offer protective effects against prostate tumor growth [142]. Outcomes in prostate cancer vary significantly between populations, influenced by both biology and healthcare access. African American men, despite a higher predisposition to aggressive prostate cancer subtypes, often face poorer outcomes due to delayed diagnoses and limited access to advanced treatments [143]. Treatment options vary according to the cancer stage and patient characteristics, with common therapies including surgery, radiation, hormone therapy, and, in more advanced cases, chemotherapy and immunotherapy. Recent advances in precision medicine, such as targeting the androgen receptor signaling pathway, have shown promise in patients with castration-resistant prostate cancer, a form that disproportionately affects African American men [144]. Screening strategies for prostate cancer differ globally, shaped by resource availability and specific population risk profiles. In the United States, prostate-specific antigen (PSA) testing is common among high-risk individuals, especially African American men and those with a family history of prostate cancer, often beginning screening in their 40s or early 50s [145]. However, in many European countries, prostate cancer screening remains controversial due to concerns over overdiagnosis and overtreatment, with recommendations varying by country. Asian countries, where prostate cancer incidence is lower, often have limited formal screening programs, focusing instead on opportunistic screenings in clinical settings [146].
4.2.5. Hepatocellular carcinoma
Following similar population-specific patterns observed in prostate and colorectal cancers, hepatocellular carcinoma (HCC) displays a distinct geographic distribution due to varying risk factors and healthcare conditions across regions. HCC is notably more prevalent in areas with high rates of chronic hepatitis B (HBV) and hepatitis C (HCV) infections, particularly in East Asia and Sub-Saharan Africa. For instance, in China, HCC incidence reaches about 18 cases per 100,000, largely due to high HBV prevalence and vertical transmission from mother to child, which leads to early, chronic infections and significantly raises cancer risk. In Sub-Saharan Africa, where HBV rates are similarly high, compounded by aflatoxin exposure from contaminated grains and nuts, adding a unique carcinogenic factor to these regions [147]. In contrast, liver cancer rates in Western countries are generally lower, about 6–8 per 100,000, but have been rising due to the increasing prevalence of non-alcoholic fatty liver disease (NAFLD) associated with obesity, diabetes, and metabolic syndrome, conditions that promote liver inflammation and carcinogenesis [148]. Outcomes and treatment options for HCC vary significantly by region, influenced by healthcare access, availability of specialized treatments, and disease stage at diagnosis. In high-incidence areas like East Asia, many cases present at later stages due to limited access to screening, resulting in poorer outcomes [149]. In wealthier nations, however, early-stage HCC is more frequently detected through routine screening in high-risk patients, allowing for curative treatments like surgical resection, liver transplantation, or local ablative therapies [150]. Nevertheless, advanced HCC cases in Western countries are increasingly treated with systemic therapies, including tyrosine kinase inhibitors and immune checkpoint inhibitors, which have shown promise in extending survival but remain costly and less accessible in low-income regions [151]. Screening strategies for HCC also vary by population risk and healthcare infrastructure. In East Asia and Sub-Saharan Africa, screening efforts often focus on HBV-infected individuals to detect early signs of HCC, though public health limitations can restrict reach. In Western countries, guidelines recommend biannual ultrasonography for high-risk patients, particularly those with cirrhosis or long-standing viral hepatitis, to detect HCC at a stage amenable to curative interventions [152,153].
In conclusion, the intricate relationship between genetic mutations, epigenetic changes, and environmental exposures significantly impacts the immune system's ability to respond to cancer and is vital for addressing cancer disparities [154]. These factors weaken immune surveillance, promote immune tolerance, and create an environment conducive to tumor development and progression [155]. Gaining a deeper understanding of these processes is essential for advancing therapeutic approaches, such as immunotherapies and prevention strategies, that aim to restore or enhance immune function in cancer patients to effectively manage cancer across diverse populations [156].
5. Immune diversity and its role in shaping infectious disease susceptibility
5.1. The mechanisms behind immune system diversity significantly influence the impact of infectious diseases
5.1.1. Genetic diversity and epigenetic modifications
Genetic diversity in immune-regulatory genes, particularly within the HLA complex, plays a critical role in susceptibility to infections (Table 1). For example, the HLA-B∗57 allele is associated with slower progression of HIV infection due to enhanced CD8+ T-cell responses [157]. Conversely, certain HLA variants, such as HLA-DRB1∗1501, are linked to increased susceptibility to tuberculosis [158]. Additionally, mutations in innate immune receptors, such as TLR4 polymorphisms, can affect bacterial infection outcomes, including a heightened risk for septic shock in Gram-negative infections [159]. Furthermore, recent research has highlighted the role of RNA N6-methyladenosine (m6A) modification in immune regulation. Dysregulation of m6A regulators influences immune responses, as seen in periodontitis, where m6A-modified RNA patterns alter immunocyte abundance and affect the expression of critical immune genes, including HLA-B [160]. This example of m6A modification underlines the importance of epigenetic regulators in modulating immune responses across different infectious diseases.
The role of epigenetic modifications in modulating immune responses during infections is also crucial. Pathogens like Mycobacterium tuberculosis exploit host epigenetic mechanisms, such as DNA methylation, to suppress immune gene expression and evade detection [161]. Chronic viral infections, such as hepatitis B and C, alter histone acetylation patterns to downregulate interferon-stimulated genes, thereby reducing antiviral immunity [162]. miRNAs also contribute to immune evasion by viruses like Epstein-Barr virus, wherein viral miRNAs suppress immune surveillance pathways to promote viral persistence [163]. As highlighted in periodontitis, m6A modifications regulate immune microenvironments by influencing the activity of key immune-regulatory genes like ALKBH5 and ELAVL1, which are involved in monocyte abundance and cytokine signaling [160]. These findings expand our understanding of how epigenetic modifications, beyond DNA methylation and histone acetylation, play a critical role in immune responses to infections.
Genetic and epigenetic factors also play pivotal roles in determining individual responses to vaccination. Polymorphisms in HLA genes influence the strength of immune responses post-vaccination, with certain alleles correlating with lower or higher antibody titers. For instance, studies have shown that genetic variability in HLA alleles affects the body's reaction to the hepatitis B vaccine, where certain alleles result in weaker responses [164]. Similarly, variations in genes related to cytokine production, such as IL-10 and IL-6, can influence vaccine efficacy by altering inflammatory responses. Epigenetic factors also impact vaccine outcomes. DNA methylation can influence the development of immune memory, which is essential for the long-term protection conferred by vaccines. Research into epigenetic regulation of immune cells following vaccination has shown that DNA methylation changes can modulate the expression of genes critical for vaccine-induced immune responses, such as those involved in B-cell and T-cell activation [165]. Vaccination can also influence the epidemiology of other infectious diseases. For example, the Bacillus Calmette–Guérin (BCG) vaccine, primarily used against tuberculosis, has been observed to reduce mortality from other infections such as respiratory and gastrointestinal diseases by stimulating trained immunity—a form of enhanced innate immune response [166]. This cross-protection demonstrates how vaccination can reshape immune responses beyond its intended target, potentially reducing the burden of various infectious diseases in vaccinated populations.
5.1.2. Environmental influences
Environmental influences significantly shape immune responses to pathogens. In pathogen-rich environments, genetic adaptations have evolved to counteract endemic infectious diseases. A prominent example is the sickle cell trait (HbAS), which provides partial resistance to Plasmodium falciparum malaria [167]. Similarly, in populations with high burdens of helminth or other parasitic infections, immune tolerance mechanisms may develop, resulting in a higher threshold for inflammation but reduced efficacy against viral infections such as HIV [168]. Environmental exposures to pollutants and allergens also alter immune responses, as evidenced in urban populations with increased rates of respiratory infections and asthma exacerbations [169].
5.1.3. Socioeconomic conditions
Socioeconomic conditions are closely linked to infectious disease rates. Overcrowded living conditions, poor hygiene practices, malnutrition, and limited access to healthcare all contribute to the spread of diseases such as tuberculosis, which thrives in such environments [170]. Poor hygiene, especially in areas lacking proper sanitation infrastructure, exacerbates the spread of diarrheal diseases and respiratory infections. Lifestyle factors, including smoking, impair mucociliary clearance and lung immunity, predisposing individuals to respiratory infections such as pneumonia and chronic obstructive pulmonary disease (COPD)-related exacerbations [171]. Alcohol consumption also weakens immune function, increasing susceptibility to infections like bacterial pneumonia and hepatitis [172].
5.1.4. Nutrition
Nutrition plays an equally critical role in determining immune response. Deficiencies in key micronutrients, such as vitamin A and zinc, are linked to impaired immune function, leading to increased susceptibility to infections such as pneumonia and diarrhea [173]. Malnutrition has been particularly associated with diminished immune responses to vaccines. In malnourished populations, children often exhibit weaker responses to vaccines like measles and rotavirus, making these populations more vulnerable to vaccine-preventable diseases [173]. Additionally, poor sanitation and hygiene contribute significantly to infection rates, particularly in low-income settings. The “hygiene hypothesis” suggests that early exposure to a wide variety of microbes can promote a balanced immune response, reducing the risk of allergies and autoimmune diseases [174]. However, overcrowded living conditions and poor access to clean water exacerbate the spread of infectious diseases, such as tuberculosis and diarrheal diseases [170].
5.1.5. Microbiome
The microbiome's composition can either bolster or impair immune responses to pathogens. For instance, individuals with a disrupted gut microbiome are more susceptible to Clostridium difficile infections following antibiotic treatment [175]. Research indicates that gut microbiota diversity influences the efficacy of influenza vaccines, with a healthy microbiome linked to better antibody responses [176]. Additionally, early exposure to a diverse range of microorganisms, as observed in rural populations, may promote a more balanced immune system capable of managing both infections and inflammatory conditions [174].
5.2. Population-level variations in infectious disease response: The impact of immune system diversity
Immune diversity, shaped by genetic, environmental, and socioeconomic factors, has a profound impact on the prevalence and outcomes of infectious diseases, with notable disparities across populations. For example, the global burden of malaria is highest in Sub-Saharan Africa, accounting for approximately 95 % of malaria cases and deaths, with children under five and pregnant women being particularly vulnerable [177]. Similarly, tuberculosis (TB) incidence varies significantly by region, with the highest rates seen in South Asia and Sub-Saharan Africa, where TB incidence rates can exceed 200 per 100,000 people, compared to under 10 per 100,000 in most high-income countries [178]. Racial disparities are also prominent in diseases like COVID-19, where African American and Hispanic communities in the United States experienced higher hospitalization and mortality rates, largely attributed to socioeconomic factors and pre-existing health disparities, showing nearly 2–3 times higher COVID-19 mortality than White Americans [179]. Herpes viruses also demonstrate population-specific variations, with herpes simplex virus type 2 (HSV-2) seroprevalence being approximately 50 % among African populations compared to around 20 % in Europe, reflecting both genetic susceptibility and differing environmental exposures [180]. HIV incidence, which remains concentrated in specific regions and demographics, shows a similarly stark contrast, with African Americans in the United States having an incidence rate approximately eight times higher than White Americans, while sub-Saharan Africa accounts for roughly two-thirds of global cases [181]. This diversity in immune response and disease susceptibility across populations underscores the need for targeted approaches in disease prevention and management.
5.2.1. Malaria
Malaria's impact varies greatly due to genetic diversity, environmental conditions, and population-specific immune responses. Sub-Saharan Africa experiences the highest malaria burden, with 95 % of global cases, whereas incidence is much lower in Southeast Asia and Latin America due to both environmental and genetic factors [177]. Inter-individual immune response variation highlights the need for more personalized treatments. For example, the sickle cell trait (HbAS) provides 60–90 % protection against severe malaria among African populations by altering red blood cell structure to reduce Plasmodium survival [182]. Southeast Asian populations, where glucose-6-phosphate dehydrogenase (G6PD) deficiency is prevalent, also exhibit some malaria resistance but face treatment challenges due to drug-induced hemolysis risks with primaquine [183]. Global databases show that malaria susceptibility and response to treatment are shaped by genetic polymorphisms. West African populations carrying HLA-B∗53 demonstrate increased protection against severe malaria, while TNF and IL10 variants influence inflammatory responses, impacting disease outcomes across different groups [184]. Genome-wide studies have identified additional variants, such as those in CR1 and FcγRIIa, affecting immune function and parasite clearance, supporting the importance of tailored therapies and diagnostics [185]. These genetic differences complicate efforts to develop universal vaccines. The RTS,S/AS01 vaccine (Mosquirix), the first malaria vaccine approved for public use, demonstrates variable efficacy across populations. Host genetic factors, including specific HLA alleles, may affect immune response to the vaccine [186]. Additionally, G6PD deficiency alters drug efficacy and safety, highlighting pharmacogenomics' role in treatment customization [187]. Future strategies to combat malaria should integrate genetic, environmental, and regional data to improve outcomes in high-burden areas through personalized approaches, diagnostics, and more adaptable vaccines.
5.2.2. Tuberculosis
Following similar patterns observed in other infectious diseases, tuberculosis (TB) incidence varies significantly across populations due to inter-individual immune response diversity influenced by both genetic and socioeconomic factors. Globally, TB is most prevalent in sub-Saharan Africa and South Asia, where incidence rates reach up to 226 per 100,000 in Africa compared to approximately 58 per 100,000 in Europe [178]. This regional variation underscores the role of individual immune responses, which can determine infection severity and susceptibility to TB. Variability in genes associated with immune response, such as the HLA-DRB1∗1501 allele found in certain Asian populations, correlates with increased TB risk by influencing the body's ability to recognize and respond to Mycobacterium tuberculosis [188]. This diversity challenges universal treatments for TB and supports the need for more personalized approaches. Individuals in crowded or low-nutrition environments, common in low-income regions, face a significantly higher TB risk due to compromised immunity and increased exposure to the disease [189]. In these settings, weakened immune responses—especially among malnourished children—make individuals more prone to active TB, underscoring the need for population-specific preventive and therapeutic strategies. Genomic studies, including GWAS and functional genomics, have identified genetic polymorphisms that impact TB susceptibility, such as those in the TLR2 and NRAMP1 genes, which play key roles in innate immune responses to TB. Polymorphisms in these genes have been associated with differences in pathogen recognition, inflammation, and disease progression, suggesting potential for pharmacogenomic approaches in high-risk groups [190]. This genetic variability also impacts diagnostic reliability and vaccine efficacy, as immune diversity may alter individual responses to TB vaccines like BCG, reducing their effectiveness in certain populations [191]. Moreover, genetic variants influence TB drug efficacy and toxicity. For example, variations in NAT2 and CYP2E1 genes affect isoniazid metabolism, leading to differing rates of adverse effects and drug resistance. Addressing these genetic factors could optimize drug selection and reduce toxicity for affected populations [192]. Future TB management should consider genetic, environmental, and socioeconomic factors to develop diagnostic tools, vaccines, and therapies better suited to diverse populations globally.
5.2.3. Coronavirus disease 2019
COVID-19 also highlighted significant differences in infection severity and mortality across racial and ethnic groups. In the United States, mortality rates were highest among African Americans, Hispanics, and Native Americans, with rates reaching 90.7, 77.7, and 82.5 per 100,000, respectively, compared to lower rates in non-Hispanic Whites [193]. Such disparities underscore the role of individual immune response variability, which influences susceptibility and outcomes. Globally, COVID-19 infection rates are highest in densely populated areas and regions with limited healthcare access, further highlighting the need for region-specific approaches. Vaccination efforts introduced additional challenges, as immune diversity affects both vaccine efficacy and individual responses. For instance, variations in immune-related genes, particularly within the HLA system and cytokine-regulating genes, can impact how effectively individuals respond to mRNA vaccines like Pfizer-BioNTech and Moderna. These vaccines stimulate an immune response by delivering mRNA that instructs cells to produce a viral protein. However, genetic differences in HLA—which is crucial for presenting viral antigens to immune cells—can affect the strength and duration of this immune response. Variations in cytokine-related genes, which control immune signaling proteins, also influence the body's initial response and long-term immunity after vaccination [194]. The emergence of variants like Delta and Omicron has further complicated vaccine efficacy, as immune gene polymorphisms can influence how well an individual's immune system produces neutralizing antibodies, affecting immunity durability [195]. Additionally, genetic variants related to drug metabolism, such as those in CYP3A4, may alter vaccine side effects, influencing safety and effectiveness across different groups [196]. Moving forward, COVID-19 management may benefit from personalized vaccine protocols that consider individual genetic and environmental factors to optimize protection and minimize adverse effects across diverse populations.
5.2.4. Herpes simplex virus type 2
Building on observed patterns of infection disparities, herpes simplex virus type 2 (HSV-2) shows marked variations in prevalence and outcomes across global populations. Sub-Saharan Africa has the highest HSV-2 prevalence, with approximately 40 % of the population affected, whereas rates in Europe and the Americas are notably lower, between 10 and 20 % [197]. Inter-individual differences in immune response, particularly in genes linked to innate immunity, significantly influence susceptibility and disease progression. For example, genetic polymorphisms in the TLR2 gene, which encodes a receptor essential for detecting viral components, can modulate the body's initial immune response to HSV-2, impacting infection severity. Additionally, variations in the IFN-λ gene, which influences interferon production, affect antiviral defense across populations, contributing to differences in disease outcomes [198]. Global data underscores the disparity in HSV-2 prevalence between regions, largely influenced by socioeconomic factors. Lower socioeconomic status, limited access to sexual health services, and constrained resources for hygiene and healthcare in sub-Saharan Africa and parts of Latin America exacerbate HSV-2 transmission and disease burden. In these regions, crowded living conditions and reduced access to preventive care contribute to higher infection rates and more severe outcomes, as access to early diagnosis and antiviral treatments remains limited [180]. The challenge of developing universal treatments for HSV-2 is compounded by genetic diversity affecting immune response and drug efficacy. For instance, variations in TLR2 and IFN-λ genes can influence the effectiveness of antiviral therapies, as immune system modulation by these genes may alter how well the body responds to treatment. Functional genomics and pharmacogenomic studies indicate that genetic diversity can impact drug efficacy and toxicity, underscoring the potential for tailored therapies based on individual genetic profiles [199]. Immune diversity also complicates global screening and vaccination strategies for HSV-2. Diagnostic tools and future vaccine development must account for varying immune responses across populations to be effective. While no HSV-2 vaccine currently exists, research is ongoing, and immune response variation poses a hurdle for creating a one-size-fits-all solution. For instance, HLA gene variations, which play a role in antigen presentation, could impact vaccine effectiveness by altering the immune system's ability to recognize and respond to HSV-2 antigens [200].
5.2.5. Human immunodeficiency virus
Human immunodeficiency virus (HIV) also exemplifies how genetic diversity and immune response variability can shape disease susceptibility, progression, and treatment outcomes across populations. African populations carry the largest HIV burden globally, with approximately 70 % of cases, revealing vast inter-individual immune response differences. For instance, specific HLA alleles, like HLA-B∗57, are linked to slower HIV progression in these populations by enhancing CD8+ T-cell responses, which can more effectively control viral replication [201]. Such genetic variations underscore the need for a more personalized approach to HIV treatment. Globally, regions like North America and Europe see lower HIV prevalence and mortality rates, in part due to wider access to antiretroviral therapy (ART), illustrating socioeconomic and healthcare disparities between regions. Limited ART access, coupled with inadequate healthcare infrastructure, remains a significant challenge in sub-Saharan Africa, where higher HIV mortality persists [202]. Beyond socioeconomic factors, environmental contributors such as malnutrition and high rates of co-infections, particularly with tuberculosis, worsen HIV outcomes in low-income regions, leading to compounded morbidity and mortality [203]. Advances in genomics, including genome-wide association studies (GWAS), have identified multiple genetic polymorphisms associated with HIV susceptibility and ART response. Beyond HLA alleles, variations in genes like CCR5 – which codes for a receptor HIV uses to enter cells – affect susceptibility and disease outcomes. The CCR5-Δ32 variant, which is less common in African populations but more frequent in European groups, provides partial resistance to HIV, highlighting regional differences in genetic defense mechanisms [204]. Immune diversity poses additional challenges for HIV diagnostics and vaccine development, as varied immune responses across populations impact vaccine efficacy. Genetic variants also influence ART efficacy, toxicity, and resistance. For example, polymorphisms in genes like CYP2B6 and ABCB1, involved in drug metabolism and transport, can lead to differences in drug concentrations and side effects, affecting treatment outcomes and adherence in genetically diverse populations [205].
Immune diversity, influenced by genetic predispositions, environmental exposures, socioeconomic conditions, and healthcare access, profoundly impacts infectious disease susceptibility, progression, and outcomes across populations. These disparities underscore the importance of moving beyond universal approaches to disease prevention and treatment, as the biological and social determinants of health significantly differ worldwide. For instance, polymorphisms in genes like HLA, CCR5, and TLR2 affect immune responses to diseases such as HIV, tuberculosis, and malaria, shaping each population's unique vulnerability and response to treatment [206]. Socioeconomic factors further influence disease outcomes, as limited access to healthcare, poor nutrition, and high exposure to pathogens in lower-income areas exacerbate morbidity and mortality, despite available treatments [207]. Understanding and addressing these multifactorial disparities is essential for developing public health strategies that incorporate genetic, environmental, and social variables, fostering more equitable and effective disease management globally. Such a personalized approach holds promise for optimizing vaccine efficacy, improving diagnostic accuracy, and minimizing adverse drug reactions, thereby enhancing patient outcomes and reducing disease burden in high-risk populations [208].
6. The future of immunity: personalized interventions and mitigating disparities
6.1. Uncovering the power of immune profiling
The future of immunology hinges on harnessing insights gleaned from comprehensive immune profiling to develop targeted therapeutic strategies. This necessitates delving deeper through large-scale genomic studies, epigenome-wide association studies (EWAS), and multi-omics approaches, which integrate genomics, transcriptomics, proteomics, and metabolomics to create a more complete picture of immune function and its regulatory mechanisms [209,210,211,212]. Additionally, longitudinal studies tracking immune profiles over time can provide invaluable information regarding the dynamic nature of immune diversity.
For instance, these methods have been instrumental in identifying genetic variations that influence immune responses to infections such as malaria, tuberculosis, HIV, and COVID-19, in different ethnic groups, highlighting the importance of population-specific data in shaping therapeutic interventions [210]. Moreover, the role of epigenetics is increasingly recognized, the International Human Epigenome Consortium provided critical insights into how epigenetic modifications can affect immune regulation and disease progression. With the aim to create a comprehensive catalog of high-quality reference epigenomes for various primary human cell types, such as DNA methylation and histone modifications, the collaborative efforts have led to significant advances in the field, including cancer and autoimmune disorders, setting the stage for future research that integrates epigenomics with other data, such as proteomics and transcriptomics [211]. In addition to cross-sectional studies, longitudinal research tracking immune profiles over time offers invaluable information about the dynamic nature of immune diversity. These studies are essential for uncovering how immune responses evolve in different physiological states, such as aging, chronic inflammation, or during infections. Single-cell data science, with its ability to analyze individual immune cells in detail, further enhances our understanding by revealing heterogeneity within immune cell populations and uncovering previously hidden cellular subtypes and states [209]. Together, these advanced methods and approaches are poised to reshape the future of personalized immunotherapy and preventative medicine (Fig. 4).
Fig. 4.
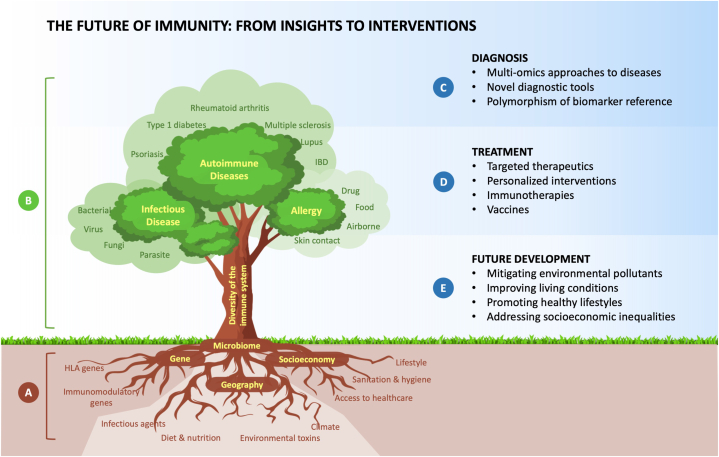
Exploring the Complexity of Immune System Diversity: Drivers, Responses, and Medical Applications.
A. The human immune system can be conceptualized as a complex network with three primary drivers: genetic variations; geographic factors, including infectious agents, diet, nutrition, climate, and environmental toxins; and socioeconomic factors such as access to healthcare, sanitation, hygiene, and lifestyle. These primary drivers interact with one another and significantly influence the body's microbiome, which acts as a protective interface for the immune system.
B. This strong foundation allows the immune system to develop robustly, resulting in individual diversity. Such diversity manifests in varied expressions of infectious diseases, autoimmune disorders, and allergies.
C. Understanding this conceptual framework suggests that medical researchers should adopt multi-omics approaches to diseases, develop novel diagnostic tools, and consider the polymorphism of biomarkers based on racial and regional factors.
D. Investigating the interplay between the drivers and responses of the immune system forms the basis for the development of targeted therapeutics, personalized interventions, immunotherapies, and vaccines.
E. Furthermore, examining the interplay between diseases and socioeconomic factors through the lens of immune interactions highlights the future of preventive healthcare. This approach focuses on reducing environmental pollutants, improving living conditions, promoting healthy lifestyles, and addressing socioeconomic disparities.
6.2. From diversity to personalized medicine
A deeper understanding of immune diversity, encompassing both genetic variations and intrinsic and extrinsic factors, holds immense promise for the development of personalized therapies. This knowledge can inform the creation of targeted immunotherapies for autoimmune diseases and more effective vaccines for infectious diseases. For example, recognizing how specific gene variants can influence immunity helps modify treatments to individuals or groups with certain genetic profiles [213], certain metabolic enzymes are now understood to influence the immune system's ability to detect and respond to viral infections [8], these variations must be accounted for in the development of both therapies and vaccines to ensure their efficacy across diverse populations [214]. Bridging the gap between fundamental research and clinical applications is paramount in translating these insights into effective healthcare solutions. Translational research, which turns findings from immune diversity studies into practical tools, holds extensive potential to revolutionize diagnostics, therapeutic targets, and interventions. This approach could lead to the development of novel diagnostic assays, which offer promising strategies for managing autoimmune diseases by restoring balance to the immune system, that predict patient responses to vaccines or therapies, or the identification of new therapeutic targets for modulating immune responses, particularly in chronic diseases such as autoimmune disorders or cancer [215,216,217], which significantly improve patient outcomes. Studies have shown that human genetic variants and age are the strongest predictors of immune responses to common pathogens and vaccines, and human immune system has been shaped by genetic adaptation to environmental pressures, resulting in significant variability in immune function across populations. By recognizing individual variations in immune responses, healthcare professionals and researchers can design personalized treatment plans that better address the underlying disease processes and are more efficacious across diverse populations [3,209,218].
6.3. Addressing health disparities in immunity
Understanding the role of environmental factors, epigenetics, and sociodemographic characteristics (race, geographic location, socioeconomic status) in shaping immune variation is crucial for addressing health disparities within public health strategies. The National Institute of Allergy and Infectious Diseases (NIAID) highlights the disproportionate burden of certain diseases among minority populations in the United States, who may face higher rates of diseases such as HIV/AIDS, non-communicable chronic diseases. Limitation to healthcare access, environmental exposures, socioeconomic inequalities, the report emphasized the need for targeted research, public health strategies, and interventions to improve health outcomes among minority groups [219]. A recent study investigated how socioeconomic status and ethnicity are linked to immunosenescence, the gradual deterioration of the immune system that associated with aging, and they found that individuals from lower socioeconomic status backgrounds and certain racial/ethnic groups (especially African Americans and Hispanics) experience accelerated immunosenescence. This contributes to increased susceptibility to infectious diseases and chronic conditions in older age [220]. Furthermore, a recent systematic review which examined the impact of geographical accessibility to healthcare on the spread and management of infectious diseases in low- and middle-income countries reveals that poor access to healthcare facilities, especially in rural areas, significantly worsens health outcomes, particularly for infectious diseases like tuberculosis, malaria, and HIV [221]. Initiatives aimed at mitigating environmental pollution, improving living conditions, promoting healthy lifestyles, enhancing access to healthcare, and reducing socioeconomic inequalities can potentially lessen observed disparities in immune responses [221,222,223,224,225,226,227]. These efforts can have a positive impact on population-level immune health. The studies call for policies and healthcare interventions to mitigate the social and economic factors, the need for improved healthcare infrastructure, transportation systems, and telemedicine solutions to bridge the gap in accessibility and exacerbating immune decline in these populations. Additionally, comparative studies across diverse populations can shed light on patterns of immune variation, revealing both commonalities and differences between demographic groups. Collaborative efforts involving international consortia are essential to capture the full spectrum of human immune diversity.
6.4. Overcoming diversity in infectious diseases with herd immunity
Herd immunity, a cornerstone concept in public health, arises when a substantial portion of a population becomes immune to a specific infectious disease, leading to a dramatic reduction in the risk of person-to-person transmission within a community [228]. This indirect protection benefits vulnerable individuals who are not yet vaccinated or immunocompromised.
However, herd immunity is a dynamic process. Over time, natural or vaccine-derived immunity can wane due to factors such as mortality or the emergence of new susceptible individuals entering the population through birth or migration [229]. It is important to note that herd immunity does not guarantee complete individual protection with their own specific immunity property. People who are not immune remain susceptible to infection and its potential consequences, even in populations with high herd immunity levels.
The successful eradication of smallpox [230] and polio [231] serves as a testament to the power of herd immunity achieved through extensive vaccination programs. Herd immunity also plays a critical role in mitigating the spread of other diseases like pertussis, influenza, pneumococcal disease, cholera [232], and rotavirus [233].
The level of vaccination coverage required to achieve herd immunity varies depending on the infectiousness of the disease, measured by the number of secondary infections caused by a single infected person. For instance, measles, where one person can infect up to 18 others, requires a herd immunity threshold of approximately 95 % [234,235]. Conversely, SARS-CoV-2, the virus responsible for COVID-19, has a lower transmission rate, with an average infected person transmitting the virus to two or three others [236], therefore, achieving herd immunity for COVID-19 might be possible with a vaccination coverage of around 60 % of the population [237].
7. Limitations of the study
Our review has several limitations. First, because of the limitations of a review length, we cannot express enough the broad immunity diversity and environment interaction context, but to state the general main points. For instances, specific roles of immune system components, or gene variations, epigenetic modifications and multivariable elements on each of the specific mechanisms or diseases between geographical and racial influences. Second, a clear association of complex interplay in immune response pathophysiology between internal and external determinants is not fully discussed on practical applications despite highlighted potential for precision medicine. Third, scope of data comparison analysis is limited due to the diversity of research in medical advances and many kinds of diseases, which can be subject to confounding variables, biases and study designs, as well as differences in disease prevalence burden between regions.
8. Conclusion
The immune system, characterized by its vast complexity, remains insufficiently comprehended. It becomes clear that the complexity of the immune system demands a customized approach in both research and clinical practice. Over extended periods of evolution, a multitude of genetic and non-genetic influences have significantly altered its original state. The expansion of reference samples, biobanks, and datasets is imperative to adequately capture the extensive genetic diversity present across populations. Racialization of immune indices is necessary for practical application in clinical practice. At present, immunoassays continue to rely on manufacturers, despite awareness of variations across races and geographic areas. Laboratory indicators assessing general status related to immunity are still commonly used across populations, despite differences between the underlying system and exposure factors. Although numerous studies have shown racial variations, there is still no specific agreement on normal constants common to ethnic groups, or the same ethnic group in diverse places. By integrating these insights into personalized and population-specific healthcare strategies, the future of immunology will not only enhance individual patient treatments and outcomes but also help reduce health disparities across diverse communities.
Funding
VinUniversity - Funding number: VUNI.2021.SG15 - Funding recipient: Chien Dinh Huynh
CRediT authorship contribution statement
Kim Han Hoang Nguyen: Writing – review & editing, Writing – original draft, Conceptualization. Nghi Vinh Le: Writing – review & editing, Writing – original draft, Conceptualization. Phuc Hoang Nguyen: Writing – review & editing, Writing – original draft, Conceptualization. Hien Hau Thi Nguyen: Writing – review & editing, Writing – original draft, Conceptualization. Duy Mai Hoang: Writing – review & editing, Conceptualization. Chien Dinh Huynh: Writing – review & editing, Writing – original draft, Supervision, Investigation, Conceptualization.
Declaration of Competing Interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
References
- 1.Justiz Vaillant A.A., Sabir S., Jan A. StatPearls Publishing; 2024. Physiology, Immune Response. StatPearls: Treasure Island (FL) [PubMed] [Google Scholar]
- 2.Marshall J.S., Warrington R., Watson W., Kim H.L. An introduction to immunology and immunopathology. Allergy Asthma Clin. Immunol. 2018;14(Suppl 2):49. doi: 10.1186/s13223-018-0278-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Liston A., Humblet-Baron S., Duffy D., Goris A. Human immune diversity: from evolution to modernity. Nat. Immunol. 2021;22(12):1479–1489. doi: 10.1038/s41590-021-01058-1. [DOI] [PubMed] [Google Scholar]
- 4.Orrù V., Steri M., Sidore C., Marongiu M., Serra V., Olla S., et al. Complex genetic signatures in immune cells underlie autoimmunity and inform therapy. Nat. Genet. 2020;52(10):1036–1045. doi: 10.1038/s41588-020-0684-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.King Thomas J., Mir H., Kapur N., Singh S. Racial differences in immunological landscape modifiers contributing to disparity in prostate cancer. Cancers. 2019;11(12) doi: 10.3390/cancers11121857. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Vuckovic D., Bao E.L., Akbari P., Lareau C.A., Mousas A., Jiang T., et al. The polygenic and monogenic basis of blood traits and diseases. Cell. 2020;182(5):1214. doi: 10.1016/j.cell.2020.08.008. 31.e11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Chen M.H., Raffield L.M., Mousas A., Sakaue S., Huffman J.E., Moscati A., et al. Trans-ethnic and ancestry-specific blood-cell genetics in 746,667 individuals from 5 global populations. Cell. 2020;182(5):1198. doi: 10.1016/j.cell.2020.06.045. 213.e14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Meyer D., Vr C.A., Bitarello B.D., Dy C.B., Nunes K. A genomic perspective on HLA evolution. Immunogenetics. 2018;70(1):5–27. doi: 10.1007/s00251-017-1017-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Dendrou C.A., Petersen J., Rossjohn J., Fugger L. HLA variation and disease. Nat. Rev. Immunol. 2018;18(5):325–339. doi: 10.1038/nri.2017.143. [DOI] [PubMed] [Google Scholar]
- 10.Baranzini S.E. In: Genomic and Personalized Medicine. second ed. Ginsburg G.S., Willard H.F., editors. Academic Press; 2013. Chapter 70 - autoimmune disorders; pp. 822–838. [Google Scholar]
- 11.Westerberg L.S., Klein C., Snapper S.B. Breakdown of T cell tolerance and autoimmunity in primary immunodeficiency--lessons learned from monogenic disorders in mice and men. Curr. Opin. Immunol. 2008;20(6):646–654. doi: 10.1016/j.coi.2008.10.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Zhao B., Chang L., Fu H., Sun G., Yang W. The role of autoimmune regulator (AIRE) in peripheral tolerance. J Immunol Res. 2018;2018 doi: 10.1155/2018/3930750. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Mathis D., Benoist C. A decade of AIRE. Nat. Rev. Immunol. 2007;7(8):645–650. doi: 10.1038/nri2136. [DOI] [PubMed] [Google Scholar]
- 14.Anderson M.S., Su M.A. AIRE expands: new roles in immune tolerance and beyond. Nat. Rev. Immunol. 2016;16(4):247–258. doi: 10.1038/nri.2016.9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Renz H., Herz U. Immune mechanisms of peripheral tolerance. Nutr. Res. 1998;18(8):1327–1333. [Google Scholar]
- 16.Fontenot J.D., Gavin M.A., Rudensky A.Y. Foxp3 programs the development and function of CD4+CD25+ regulatory T cells. Nat. Immunol. 2003;4(4):330–336. doi: 10.1038/ni904. [DOI] [PubMed] [Google Scholar]
- 17.Hori S., Nomura T., Sakaguchi S. Control of regulatory T cell development by the transcription factor Foxp3. Science. 2003;299(5609):1057–1061. doi: 10.1126/science.1079490. [DOI] [PubMed] [Google Scholar]
- 18.Sakaguchi S., Yamaguchi T., Nomura T., Ono M. Regulatory T cells and immune tolerance. Cell. 2008;133(5):775–787. doi: 10.1016/j.cell.2008.05.009. [DOI] [PubMed] [Google Scholar]
- 19.Nagata S., Golstein P. The Fas death factor. Science. 1995;267(5203):1449–1456. doi: 10.1126/science.7533326. [DOI] [PubMed] [Google Scholar]
- 20.Strasser A., Jost P.J., Nagata S. The many roles of FAS receptor signaling in the immune system. Immunity. 2009;30(2):180–192. doi: 10.1016/j.immuni.2009.01.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Li S., Miao T., Sebastian M., Bhullar P., Ghaffari E., Liu M., et al. The transcription factors Egr2 and Egr3 are essential for the control of inflammation and antigen-induced proliferation of B and T cells. Immunity. 2012;37(4):685–696. doi: 10.1016/j.immuni.2012.08.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Shi L., Chen S., Yang L., Li Y. The role of PD-1 and PD-L1 in T-cell immune suppression in patients with hematological malignancies. J. Hematol. Oncol. 2013;6(1):74. doi: 10.1186/1756-8722-6-74. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Chen D.-P., Wen Y.-H., Lin W.-T., Hsu F.-P., Yu K.-H. Exploration of the association between the single-nucleotide polymorphism of co-stimulatory system and rheumatoid arthritis. Front. Immunol. 2023;14 doi: 10.3389/fimmu.2023.1123832. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Jung J.H., Song G.G., Kim J.H., Choi S.J. The associations between interleukin 10 polymorphisms and susceptibility to autoimmune uveitis - a meta-analysis. Cent. Eur. J. Immunol. 2019;44(3):246–252. doi: 10.5114/ceji.2019.89596. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Moore K.W., de Waal Malefyt R., Coffman R.L., O'Garra A. Interleukin-10 and the interleukin-10 receptor. Annu. Rev. Immunol. 2001;19:683–765. doi: 10.1146/annurev.immunol.19.1.683. [DOI] [PubMed] [Google Scholar]
- 26.Bunte K., Beikler T. Th17 cells and the IL-23/IL-17 Axis in the pathogenesis of periodontitis and immune-mediated inflammatory diseases. Int. J. Mol. Sci. 2019;20(14) doi: 10.3390/ijms20143394. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Moore L.D., Le T., Fan G. DNA methylation and its basic function. Neuropsychopharmacology. 2013;38(1):23–38. doi: 10.1038/npp.2012.112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Alaskhar Alhamwe B., Khalaila R., Wolf J., von Bülow V., Harb H., Alhamdan F., et al. Histone modifications and their role in epigenetics of atopy and allergic diseases. Allergy Asthma Clin. Immunol. 2018;14(1):39. doi: 10.1186/s13223-018-0259-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Kumar D., Sahoo S.S., Chauss D., Kazemian M., Afzali B. Non-coding RNAs in immunoregulation and autoimmunity: technological advances and critical limitations. J. Autoimmun. 2023;134 doi: 10.1016/j.jaut.2022.102982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Heino M., Peterson P., Kudoh J., Shimizu N., Antonarakis S.E., Scott H.S., et al. APECED mutations in the autoimmune regulator (AIRE) gene. Hum. Mutat. 2001;18(3):205–211. doi: 10.1002/humu.1176. [DOI] [PubMed] [Google Scholar]
- 31.Peterson P., Peltonen L. Autoimmune polyendocrinopathy syndrome type 1 (APS1) and AIRE gene: new views on molecular basis of autoimmunity. J. Autoimmun. 2005;25:49–55. doi: 10.1016/j.jaut.2005.09.022. [DOI] [PubMed] [Google Scholar]
- 32.Pellegrino M., Bellacchio E., Dhamo R., Frasca F., Betterle C., Fierabracci A. A novel homozygous mutation of the AIRE gene in an APECED patient from Pakistan: case report and review of the literature. Front. Immunol. 2018;9 doi: 10.3389/fimmu.2018.01835. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Skrabic V., Skrabic I., Skrabic R., Roje B., Simunovic M. Clinical characteristics in the longitudinal follow-up of APECED syndrome in southern Croatia—case series. Genes. 2022;13(4):558. doi: 10.3390/genes13040558. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Ach T., Hadami B.Y., Ghariani N., Said ElMabrouk R., Ben Abdelkrim A., Kacem M., et al. Gastrointestinal manifestations in autoimmune polyendocrinopathy-candidiasis-ectodermal dystrophy (APECED) patient: major effect on treatment and prognosis. Endocrinol Diabetes Metab Case Rep. 2023;2023(2) doi: 10.1530/EDM-22-0352. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Betterle C., Greggio N.A., Volpato M. Clinical review 93: autoimmune polyglandular syndrome type 1. J. Clin. Endocrinol. Metab. 1998;83(4):1049–1055. doi: 10.1210/jcem.83.4.4682. [DOI] [PubMed] [Google Scholar]
- 36.Björses P., Halonen M., Palvimo J.J., Kolmer M., Aaltonen J., Ellonen P., et al. Mutations in the AIRE gene: effects on subcellular location and transactivation function of the autoimmune polyendocrinopathy-candidiasis-ectodermal dystrophy protein. Am. J. Hum. Genet. 2000;66(2):378–392. doi: 10.1086/302765. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Puck J.M., Pepper A.E., Henthorn P.S., Candotti F., Isakov J., Whitwam T., et al. Mutation analysis of IL2RG in human X-linked severe combined immunodeficiency. Blood. 1997;89(6):1968–1977. [PubMed] [Google Scholar]
- 38.Wan C.-K., Leonard W.J. In: Encyclopedia of Immunobiology. Ratcliffe M.J.H., editor. Academic Press; Oxford: 2016. Γc family cytokines: role in immune cell development and function; pp. 509–518. [Google Scholar]
- 39.Zhang Y., Su J. Interleukin-2 family cytokines: an overview of genes, expression, signaling and functional roles in teleost. Dev. Comp. Immunol. 2023;141 doi: 10.1016/j.dci.2023.104645. [DOI] [PubMed] [Google Scholar]
- 40.Justiz Vaillant A.A., Sabir S., Jan A. StatPearls Publishing; 2023. Severe Combined Immunodeficiency. StatPearls: Treasure Island (FL) [Google Scholar]
- 41.Kumrah R., Vignesh P., Patra P., Singh A., Anjani G., Saini P., et al. Genetics of severe combined immunodeficiency. Genes Dis. 2020;7(1):52–61. doi: 10.1016/j.gendis.2019.07.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Dvorak C.C., Haddad E., Heimall J., Dunn E., Buckley R.H., Kohn D.B., et al. The diagnosis of severe combined immunodeficiency (SCID): the primary immune deficiency treatment Consortium (PIDTC) 2022 definitions. J. Allergy Clin. Immunol. 2023;151(2):539–546. doi: 10.1016/j.jaci.2022.10.022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Bousfiha A., Jeddane L., Al-Herz W., Ailal F., Casanova J.L., Chatila T., et al. The 2015 IUIS phenotypic classification for primary immunodeficiencies. J. Clin. Immunol. 2015;35(8):727–738. doi: 10.1007/s10875-015-0198-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Jyonouchi S., Jongco A.M., Puck J., Sullivan K.E. Immunodeficiencies associated with abnormal newborn screening for T cell and B cell lymphopenia. J. Clin. Immunol. 2017;37(4):363–374. doi: 10.1007/s10875-017-0388-4. [DOI] [PubMed] [Google Scholar]
- 45.Picard C., Al-Herz W., Bousfiha A., Casanova J.L., Chatila T., Conley M.E., et al. Primary immunodeficiency diseases: an update on the classification from the international union of immunological societies expert committee for primary immunodeficiency 2015. J. Clin. Immunol. 2015;35(8):696–726. doi: 10.1007/s10875-015-0201-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Li L., Moshous D., Zhou Y., Wang J., Xie G., Salido E., et al. A founder mutation in Artemis, an SNM1-like protein, causes SCID in Athabascan-speaking Native Americans. J. Immunol. 2002;168(12):6323–6329. doi: 10.4049/jimmunol.168.12.6323. [DOI] [PubMed] [Google Scholar]
- 47.Akbayram S., Sari N., Akgün C., Doğan M., Tuncer O., Caksen H., et al. The frequency of consanguineous marriage in eastern Turkey. Genet Couns. 2009;20(3):207–214. [PubMed] [Google Scholar]
- 48.Al-Mousa H., Al-Saud B. Primary immunodeficiency diseases in highly consanguineous populations from Middle East and North Africa: epidemiology, diagnosis, and care. Front. Immunol. 2017;8:678. doi: 10.3389/fimmu.2017.00678. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Ben Halim N., Ben Alaya Bouafif N., Romdhane L., Kefi Ben Atig R., Chouchane I., Bouyacoub Y., et al. Consanguinity, endogamy, and genetic disorders in Tunisia. J Community Genet. 2013;4(2):273–284. doi: 10.1007/s12687-012-0128-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.El-Mouzan M.I., Al-Salloum A.A., Al-Herbish A.S., Qurachi M.M., Al-Omar A.A. Regional variations in the prevalence of consanguinity in Saudi Arabia. Saudi Med. J. 2007;28(12):1881–1884. [PubMed] [Google Scholar]
- 51.Greenberg-Kushnir N., Lee Y.N., Simon A.J., Lev A., Marcus N., Abuzaitoun O., et al. A large cohort of RAG1/2-deficient SCID patients-clinical, immunological, and prognostic analysis. J. Clin. Immunol. 2020;40(1):211–222. doi: 10.1007/s10875-019-00717-1. [DOI] [PubMed] [Google Scholar]
- 52.Cui X., Jiao C., Wang D., Ye Z., Ma J., Tang N., et al. Elevated levels of IL-27 are associated with disease activity in patients with crohn's disease. Mediators Inflamm. 2021;2021 doi: 10.1155/2021/5527627. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Duerr R.H., Taylor K.D., Brant S.R., Rioux J.D., Silverberg M.S., Daly M.J., et al. A genome-wide association study identifies IL23R as an inflammatory bowel disease gene. Science. 2006;314(5804):1461–1463. doi: 10.1126/science.1135245. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Zhu Y., Jiang H., Chen Z., Lu B., Li J., Shen X. Genetic association between IL23R rs11209026 and rs10889677 polymorphisms and risk of Crohn's disease and ulcerative colitis: evidence from 41 studies. Inflamm. Res. 2020;69(1):87–103. doi: 10.1007/s00011-019-01296-y. [DOI] [PubMed] [Google Scholar]
- 55.Abdollahi E., Tavasolian F., Momtazi-Borojeni A.A., Samadi M., Rafatpanah H. Protective role of R381Q (rs11209026) polymorphism in IL-23R gene in immune-mediated diseases: a comprehensive review. J. Immunot. 2016;13(3):286–300. doi: 10.3109/1547691X.2015.1115448. [DOI] [PubMed] [Google Scholar]
- 56.Hedrich C.M., Mäbert K., Rauen T., Tsokos G.C. DNA methylation in systemic lupus erythematosus. Epigenomics. 2017;9(4):505–525. doi: 10.2217/epi-2016-0096. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Jeffries M.A., Sawalha A.H. Epigenetics in systemic lupus erythematosus: leading the way for specific therapeutic agents. Int J Clin Rheumtol. 2011;6(4):423–439. doi: 10.2217/ijr.11.32. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Adams D.E., Shao W.H. Epigenetic alterations in immune cells of systemic lupus erythematosus and therapeutic implications. Cells. 2022;11(3) doi: 10.3390/cells11030506. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Gautam P., Sharma A., Bhatnagar A. Global histone modification analysis reveals hypoacetylated H3 and H4 histones in B Cells from systemic lupus erythematosus patients. Immunol. Lett. 2021;240:41–45. doi: 10.1016/j.imlet.2021.09.007. [DOI] [PubMed] [Google Scholar]
- 60.Mei X., Zhang B., Zhao M., Lu Q. An update on epigenetic regulation in autoimmune diseases. J Transl Autoimmun. 2022;5 doi: 10.1016/j.jtauto.2022.100176. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Rasuli E., Javidi-Aghdam K., Akbarzadeh-Khiavi M., Abdshah A., Gadakchi L., Jafarpour M., et al. Immunoregulatory role of AC007278.3 and HOTAIR long non-coding RNAs in lupus nephritis: potential biomarkers and therapeutic targets. Mol. Biol. Rep. 2024;51(1):1075. doi: 10.1007/s11033-024-10019-4. [DOI] [PubMed] [Google Scholar]
- 62.El-Shaer O.S., Sabry J.H., Mahgoub M.Y., Hamed N.A., Nour El Din D.M., Ameen S.G. MiR-155 expression is a potential biomarker of systemic lupus erythematosus diagnosis and disease activity prediction. Meta Gene. 2020;26 [Google Scholar]
- 63.Yu J., Mei J., Zuo D., Zhang M., Yu S., Li F., et al. Inflammatory factor-mediated miR-155/SOCS1 signaling axis leads to Treg impairment in systemic lupus erythematosus. Int. Immunopharm. 2024;141 doi: 10.1016/j.intimp.2024.113013. [DOI] [PubMed] [Google Scholar]
- 64.Pashangzadeh S., Motallebnezhad M., Vafashoar F., Khalvandi A., Mojtabavi N. Implications the role of miR-155 in the pathogenesis of autoimmune diseases. Front. Immunol. 2021;12 doi: 10.3389/fimmu.2021.669382. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Schell S.L., Rahman Z.S.M. miRNA-mediated control of B cell responses in immunity and SLE. Front. Immunol. 2021;12 doi: 10.3389/fimmu.2021.683710. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Qu B., Shen N. miRNAs in the pathogenesis of systemic lupus erythematosus. Int. J. Mol. Sci. 2015;16(5):9557–9572. doi: 10.3390/ijms16059557. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Ji J., Dai X., Yeung S.J., He X. The role of long non-coding RNA GAS5 in cancers. Cancer Manag. Res. 2019;11:2729–2737. doi: 10.2147/CMAR.S189052. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Afrashteh Nour M., Ghorbaninezhad F., Asadzadeh Z., Baghbanzadeh A., Hassanian H., Leone P., et al. The emerging role of noncoding RNAs in systemic lupus erythematosus: new insights into the master regulators of disease pathogenesis. Ther Adv Chronic Dis. 2023;14 doi: 10.1177/20406223231153572. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Liu C.H., Lu Y.L., Huang H.T., Wang C.F., Luo H.C., Wei G.J., et al. Association of LncRNA-GAS5 gene polymorphisms and PBMC LncRNA-GAS5 level with risk of systemic lupus erythematosus in Chinese population. J. Cell Mol. Med. 2021;25(7):3548–3559. doi: 10.1111/jcmm.16438. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Suo Q.F., Sheng J., Qiang F.Y., Tang Z.S., Yang Y.Y. Association of long non-coding RNA GAS5 and miR-21 levels in CD4(+) T cells with clinical features of systemic lupus erythematosus. Exp. Ther. Med. 2018;15(1):345–350. doi: 10.3892/etm.2017.5429. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Mazzone R., Zwergel C., Artico M., Taurone S., Ralli M., Greco A., et al. The emerging role of epigenetics in human autoimmune disorders. Clin. Epigenet. 2019;11(1):34. doi: 10.1186/s13148-019-0632-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Hanahan D., Weinberg R.A. Hallmarks of cancer: the next generation. Cell. 2011;144(5):646–674. doi: 10.1016/j.cell.2011.02.013. [DOI] [PubMed] [Google Scholar]
- 73.Schreiber R.D., Old L.J., Smyth M.J. Cancer immunoediting: integrating immunity's roles in cancer suppression and promotion. Science. 2011;331(6024):1565–1570. doi: 10.1126/science.1203486. [DOI] [PubMed] [Google Scholar]
- 74.Chen D.S., Mellman I. Elements of cancer immunity and the cancer–immune set point. Nature. 2017;541(7637):321–330. doi: 10.1038/nature21349. [DOI] [PubMed] [Google Scholar]
- 75.Gajewski T.F., Schreiber H., Fu Y.-X. Innate and adaptive immune cells in the tumor microenvironment. Nat. Immunol. 2013;14(10):1014–1022. doi: 10.1038/ni.2703. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Stratton M.R., Campbell P.J., Futreal P.A. The cancer genome. Nature. 2009;458(7239):719–724. doi: 10.1038/nature07943. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Vita M., Henriksson M. The Myc oncoprotein as a therapeutic target for human cancer. Semin. Cancer Biol. 2006;16(4):318–330. doi: 10.1016/j.semcancer.2006.07.015. [DOI] [PubMed] [Google Scholar]
- 78.Lane D.P. Cancer. p53, guardian of the genome. Nature. 1992;358(6381):15–16. doi: 10.1038/358015a0. [DOI] [PubMed] [Google Scholar]
- 79.Vousden K.H., Lane D.P. p53 in health and disease. Nat. Rev. Mol. Cell Biol. 2007;8(4):275–283. doi: 10.1038/nrm2147. [DOI] [PubMed] [Google Scholar]
- 80.Oren M. Decision making by p53: life, death and cancer. Cell Death Differ. 2003;10(4):431–442. doi: 10.1038/sj.cdd.4401183. [DOI] [PubMed] [Google Scholar]
- 81.Janeway C.A.J., Travers P., Walport M., Shlomchik M.J. fifth ed. Garland Science; New York: 2001. Immunobiology: the Immune System in Health and Disease. [Google Scholar]
- 82.Carretero R., Romero J.M., Ruiz-Cabello F., Maleno I., Rodriguez F., Camacho F.M., et al. Analysis of HLA class I expression in progressing and regressing metastatic melanoma lesions after immunotherapy. Immunogenetics. 2008;60(8):439–447. doi: 10.1007/s00251-008-0303-5. [DOI] [PubMed] [Google Scholar]
- 83.Pardoll D.M. The blockade of immune checkpoints in cancer immunotherapy. Nat. Rev. Cancer. 2012;12(4):252–264. doi: 10.1038/nrc3239. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Topalian S.L., Hodi F.S., Brahmer J.R., Gettinger S.N., Smith D.C., McDermott D.F., et al. Safety, activity, and immune correlates of anti-PD-1 antibody in cancer. N. Engl. J. Med. 2012;366(26):2443–2454. doi: 10.1056/NEJMoa1200690. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Jones P.A., Baylin S.B. The epigenomics of cancer. Cell. 2007;128(4):683–692. doi: 10.1016/j.cell.2007.01.029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Koch A., Joosten S.C., Feng Z., de Ruijter T.C., Draht M.X., Melotte V., et al. Analysis of DNA methylation in cancer: location revisited. Nat. Rev. Clin. Oncol. 2018;15(7):459–466. doi: 10.1038/s41571-018-0004-4. [DOI] [PubMed] [Google Scholar]
- 87.Meissner A., Gnirke A., Bell G.W., Ramsahoye B., Lander E.S., Jaenisch R. Reduced representation bisulfite sequencing for comparative high-resolution DNA methylation analysis. Nucleic Acids Res. 2005;33(18):5868–5877. doi: 10.1093/nar/gki901. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Garrido F., Aptsiauri N., Doorduijn E.M., Garcia Lora A.M., van Hall T. The urgent need to recover MHC class I in cancers for effective immunotherapy. Curr. Opin. Immunol. 2016;39:44–51. doi: 10.1016/j.coi.2015.12.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Li Y., Seto E. HDACs and HDAC inhibitors in cancer development and therapy. Cold Spring Harb Perspect Med. 2016;6(10) doi: 10.1101/cshperspect.a026831. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Margueron R., Reinberg D. The Polycomb complex PRC2 and its mark in life. Nature. 2011;469(7330):343–349. doi: 10.1038/nature09784. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.West A.C., Johnstone R.W. New and emerging HDAC inhibitors for cancer treatment. J. Clin. Invest. 2014;124(1):30–39. doi: 10.1172/JCI69738. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Oliveto S., Mancino M., Manfrini N., Biffo S. Role of microRNAs in translation regulation and cancer. World J. Biol. Chem. 2017;8(1):45–56. doi: 10.4331/wjbc.v8.i1.45. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Jiang S., Zhang H.W., Lu M.H., He X.H., Li Y., Gu H., et al. MicroRNA-155 functions as an OncomiR in breast cancer by targeting the suppressor of cytokine signaling 1 gene. Cancer Res. 2010;70(8):3119–3127. doi: 10.1158/0008-5472.CAN-09-4250. [DOI] [PubMed] [Google Scholar]
- 94.Rai D., Kim S.W., McKeller M.R., Dahia P.L., Aguiar R.C. Targeting of SMAD5 links microRNA-155 to the TGF-beta pathway and lymphomagenesis. Proc Natl Acad Sci U S A. 2010;107(7):3111–3116. doi: 10.1073/pnas.0910667107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Grivennikov S.I., Greten F.R., Karin M. Immunity, inflammation, and cancer. Cell. 2010;140(6):883–899. doi: 10.1016/j.cell.2010.01.025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Karin M., Lawrence T., Nizet V. Innate immunity gone awry: linking microbial infections to chronic inflammation and cancer. Cell. 2006;124(4):823–835. doi: 10.1016/j.cell.2006.02.016. [DOI] [PubMed] [Google Scholar]
- 97.Padoan A., Musso G., Contran N., Basso D. Inflammation, autoinflammation and autoimmunity in inflammatory bowel diseases. Curr. Issues Mol. Biol. 2023;45(7):5534–5557. doi: 10.3390/cimb45070350. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98.Balkwill F., Mantovani A. Inflammation and cancer: back to Virchow? Lancet. 2001;357(9255):539–545. doi: 10.1016/S0140-6736(00)04046-0. [DOI] [PubMed] [Google Scholar]
- 99.Coussens L.M., Werb Z. Inflammation and cancer. Nature. 2002;420(6917):860–867. doi: 10.1038/nature01322. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.El-Serag H.B. Epidemiology of viral hepatitis and hepatocellular carcinoma. Gastroenterology. 2012;142(6):1264–12673.e1. doi: 10.1053/j.gastro.2011.12.061. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.Hecht S.S. Lung carcinogenesis by tobacco smoke. Int. J. Cancer. 2012;131(12):2724–2732. doi: 10.1002/ijc.27816. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102.Cheng T.Y., Cramb S.M., Baade P.D., Youlden D.R., Nwogu C., Reid M.E. The international epidemiology of lung cancer: latest trends, disparities, and tumor characteristics. J. Thorac. Oncol. 2016;11(10):1653–1671. doi: 10.1016/j.jtho.2016.05.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Halliday G.M., Byrne S.N., Damian D.L. Ultraviolet A radiation: its role in immunosuppression and carcinogenesis. Semin. Cutan. Med. Surg. 2011;30(4):214–221. doi: 10.1016/j.sder.2011.08.002. [DOI] [PubMed] [Google Scholar]
- 104.Gopalakrishnan V., Spencer C.N., Nezi L., Reuben A., Andrews M.C., Karpinets T.V., et al. Gut microbiome modulates response to anti-PD-1 immunotherapy in melanoma patients. Science. 2018;359(6371):97–103. doi: 10.1126/science.aan4236. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105.Zitvogel L., Ma Y., Raoult D., Kroemer G., Gajewski T.F. The microbiome in cancer immunotherapy: diagnostic tools and therapeutic strategies. Science. 2018;359(6382):1366–1370. doi: 10.1126/science.aar6918. [DOI] [PubMed] [Google Scholar]
- 106.Kostic A.D., Chun E., Robertson L., Glickman J.N., Gallini C.A., Michaud M., et al. Fusobacterium nucleatum potentiates intestinal tumorigenesis and modulates the tumor-immune microenvironment. Cell Host Microbe. 2013;14(2):207–215. doi: 10.1016/j.chom.2013.07.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107.Rubinstein M.R., Wang X., Liu W., Hao Y., Cai G., Han Y.W. Fusobacterium nucleatum promotes colorectal carcinogenesis by modulating E-cadherin/β-catenin signaling via its FadA adhesin. Cell Host Microbe. 2013;14(2):195–206. doi: 10.1016/j.chom.2013.07.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 108.Luu M, Riester Z, Baldrich A, Reichardt N, Yuille S, Busetti A, Klein M, Wempe A, Leister H, Raifer H, Picard F, Muhammad K, Ohl K, Romero R, Fischer F, Bauer CA, Huber M, Gress TM, Lauth M, Danhof S, Bopp T, Nerreter T, Mulder IE, Steinhoff U, Hudecek M, Visekruna A. Microbial short-chain fatty acids modulate CD8+ T cell responses and improve adoptive immunotherapy for cancer. Nat Commun. 2021 Jul 1;12(1):4077. doi: 10.1038/s41467-021-24331-1. PMID: 34210970; PMCID: PMC8249424. [DOI] [PMC free article] [PubMed]
- 109.Feinberg A.P., Ohlsson R., Henikoff S. The epigenetic progenitor origin of human cancer. Nat. Rev. Genet. 2006;7(1):21–33. doi: 10.1038/nrg1748. [DOI] [PubMed] [Google Scholar]
- 110.Narod S.A. BRCA mutations in the management of breast cancer: the state of the art. Nat. Rev. Clin. Oncol. 2010;7(12):702–707. doi: 10.1038/nrclinonc.2010.166. [DOI] [PubMed] [Google Scholar]
- 111.Lord C.J., Ashworth A. BRCAness revisited. Nat. Rev. Cancer. 2016;16(2):110–120. doi: 10.1038/nrc.2015.21. [DOI] [PubMed] [Google Scholar]
- 112.Christensen B.C., Houseman E.A., Marsit C.J., Zheng S., Wrensch M.R., Wiemels J.L., et al. Aging and environmental exposures alter tissue-specific DNA methylation dependent upon CpG island context. PLoS Genet. 2009;5(8) doi: 10.1371/journal.pgen.1000602. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113.Tsaprouni L.G., Yang T.P., Bell J., Dick K.J., Kanoni S., Nisbet J., et al. Cigarette smoking reduces DNA methylation levels at multiple genomic loci but the effect is partially reversible upon cessation. Epigenetics. 2014;9(10):1382–1396. doi: 10.4161/15592294.2014.969637. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 114.Herbst R.S., Heymach J.V., Lippman S.M. Lung cancer. N. Engl. J. Med. 2008;359(13):1367–1380. doi: 10.1056/NEJMra0802714. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 115.Siegel R.L., Miller K.D., Jemal A. Cancer statistics, 2020. CA Cancer J Clin. 2020;70(1):7–30. doi: 10.3322/caac.21590. [DOI] [PubMed] [Google Scholar]
- 116.Bray F., Ferlay J., Soerjomataram I., Siegel R.L., Torre L.A., Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68(6):394–424. doi: 10.3322/caac.21492. [DOI] [PubMed] [Google Scholar]
- 117.Arnold M., Sierra M.S., Laversanne M., Soerjomataram I., Jemal A., Bray F. Global patterns and trends in colorectal cancer incidence and mortality. Gut. 2017;66(4):683–691. doi: 10.1136/gutjnl-2015-310912. [DOI] [PubMed] [Google Scholar]
- 118.Shenoy D., Packianathan S., Chen A.M., Vijayakumar S. Do African-American men need separate prostate cancer screening guidelines? BMC Urol. 2016;16(1):19. doi: 10.1186/s12894-016-0137-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 119.El–Serag H.B., Rudolph K.L. Hepatocellular carcinoma: epidemiology and molecular carcinogenesis. Gastroenterology. 2007;132(7):2557–2576. doi: 10.1053/j.gastro.2007.04.061. [DOI] [PubMed] [Google Scholar]
- 120.Wu Y.L., Tsuboi M., He J., John T., Grohe C., Majem M., et al. Osimertinib in resected EGFR-mutated non-small-cell lung cancer. N. Engl. J. Med. 2020;383(18):1711–1723. doi: 10.1056/NEJMoa2027071. [DOI] [PubMed] [Google Scholar]
- 121.Laguna J.C., García-Pardo M., Alessi J., Barrios C., Singh N., Al-Shamsi H.O., et al. Geographic differences in lung cancer: focus on carcinogens, genetic predisposition, and molecular epidemiology. Ther Adv Med Oncol. 2024;16 doi: 10.1177/17588359241231260. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 122.Midha A., Dearden S., McCormack R. EGFR mutation incidence in non-small-cell lung cancer of adenocarcinoma histology: a systematic review and global map by ethnicity (mutMapII) Am. J. Cancer Res. 2015;5(9):2892–2911. [PMC free article] [PubMed] [Google Scholar]
- 123.Hellmann M.D., Paz-Ares L., Bernabe Caro R., Zurawski B., Kim S.W., Carcereny Costa E., et al. Nivolumab plus ipilimumab in advanced non-small-cell lung cancer. N. Engl. J. Med. 2019;381(21):2020–2031. doi: 10.1056/NEJMoa1910231. [DOI] [PubMed] [Google Scholar]
- 124.Krist A.H., Davidson K.W., Mangione C.M., Barry M.J., Cabana M., Caughey A.B., et al. Screening for lung cancer: US preventive services task force recommendation statement. JAMA. 2021;325(10):962–970. doi: 10.1001/jama.2021.1117. [DOI] [PubMed] [Google Scholar]
- 125.Colditz G.A., Bohlke K. Priorities for the primary prevention of breast cancer. CA Cancer J Clin. 2014;64(3):186–194. doi: 10.3322/caac.21225. [DOI] [PubMed] [Google Scholar]
- 126.Porter P. "Westernizing" women's risks? Breast cancer in lower-income countries. N. Engl. J. Med. 2008;358(3):213–216. doi: 10.1056/NEJMp0708307. [DOI] [PubMed] [Google Scholar]
- 127.Warner E., Foulkes W., Goodwin P., Meschino W., Blondal J., Paterson C., et al. Prevalence and penetrance of BRCA1 and BRCA2 gene mutations in unselected Ashkenazi Jewish women with breast cancer. J Natl Cancer Inst. 1999;91(14):1241–1247. doi: 10.1093/jnci/91.14.1241. [DOI] [PubMed] [Google Scholar]
- 128.Dietze E.C., Sistrunk C., Miranda-Carboni G., O'Regan R., Seewaldt V.L. Triple-negative breast cancer in African-American women: disparities versus biology. Nat. Rev. Cancer. 2015;15(4):248–254. doi: 10.1038/nrc3896. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 129.Dent R., Trudeau M., Pritchard K.I., Hanna W.M., Kahn H.K., Sawka C.A., et al. Triple-negative breast cancer: clinical features and patterns of recurrence. Clin. Cancer Res. 2007;13(15 Pt 1):4429–4434. doi: 10.1158/1078-0432.CCR-06-3045. [DOI] [PubMed] [Google Scholar]
- 130.Nelson H.D., O'Meara E.S., Kerlikowske K., Balch S., Miglioretti D. Factors associated with rates of false-positive and false-negative results from digital mammography screening: an analysis of registry data. Ann. Intern. Med. 2016;164(4):226–235. doi: 10.7326/M15-0971. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 131.Harkness E.F., Astley S.M., Evans D.G. Risk-based breast cancer screening strategies in women. Best Pract. Res. Clin. Obstet. Gynaecol. 2020;65:3–17. doi: 10.1016/j.bpobgyn.2019.11.005. [DOI] [PubMed] [Google Scholar]
- 132.Chan A.T., Giovannucci E.L. Primary prevention of colorectal cancer. Gastroenterology. 2010;138(6):2029. doi: 10.1053/j.gastro.2010.01.057. 43.e10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 133.Burkitt D.P. Epidemiology of cancer of the colon and rectum. Cancer. 1971;28(1):3–13. doi: 10.1002/1097-0142(197107)28:1<3::aid-cncr2820280104>3.0.co;2-n. [DOI] [PubMed] [Google Scholar]
- 134.Sameer A.S. Colorectal cancer: molecular mutations and polymorphisms. Front. Oncol. 2013;3:114. doi: 10.3389/fonc.2013.00114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 135.Key T.J., Schatzkin A., Willett W.C., Allen N.E., Spencer E.A., Travis R.C. Diet, nutrition and the prevention of cancer. Public Health Nutr. 2004;7(1a):187–200. doi: 10.1079/phn2003588. [DOI] [PubMed] [Google Scholar]
- 136.Fearon E.R., Vogelstein B. A genetic model for colorectal tumorigenesis. Cell. 1990;61(5):759–767. doi: 10.1016/0092-8674(90)90186-i. [DOI] [PubMed] [Google Scholar]
- 137.Talseth-Palmer B.A. The genetic basis of colonic adenomatous polyposis syndromes. Hered. Cancer Clin. Pract. 2017;15(1):5. doi: 10.1186/s13053-017-0065-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 138.Rex D.K., Boland C.R., Dominitz J.A., Giardiello F.M., Johnson D.A., Kaltenbach T., et al. Colorectal cancer screening: recommendations for physicians and patients from the U.S. Multi-society task force on colorectal cancer. Gastroenterology. 2017;153(1):307–323. doi: 10.1053/j.gastro.2017.05.013. [DOI] [PubMed] [Google Scholar]
- 139.Ladabaum U., Dominitz J.A., Kahi C., Schoen R.E. Strategies for colorectal cancer screening. Gastroenterology. 2020;158(2):418–432. doi: 10.1053/j.gastro.2019.06.043. [DOI] [PubMed] [Google Scholar]
- 140.Johnson J.R., Woods-Burnham L., Hooker S.E., Jr., Batai K., Kittles R.A. Genetic contributions to prostate cancer disparities in men of West African descent. Front. Oncol. 2021;11 doi: 10.3389/fonc.2021.770500. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 141.Giovannucci E., Rimm E.B., Colditz G.A., Stampfer M.J., Ascherio A., Chute C.G., et al. A prospective study of dietary fat and risk of prostate cancer. J Natl Cancer Inst. 1993;85(19):1571–1579. doi: 10.1093/jnci/85.19.1571. [DOI] [PubMed] [Google Scholar]
- 142.Applegate C.C., Rowles J.L., Ranard K.M., Jeon S., Erdman J.W. Soy consumption and the risk of prostate cancer: an updated systematic review and meta-analysis. Nutrients. 2018;10(1) doi: 10.3390/nu10010040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 143.DeSantis C.E., Miller K.D., Goding Sauer A., Jemal A., Siegel R.L. Cancer statistics for african Americans, 2019. CA Cancer J Clin. 2019;69(3):211–233. doi: 10.3322/caac.21555. [DOI] [PubMed] [Google Scholar]
- 144.Ferraldeschi R., Welti J., Luo J., Attard G., de Bono J.S. Targeting the androgen receptor pathway in castration-resistant prostate cancer: progresses and prospects. Oncogene. 2015;34(14):1745–1757. doi: 10.1038/onc.2014.115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 145.Wolf A.M., Wender R.C., Etzioni R.B., Thompson I.M., D'Amico A.V., Volk R.J., et al. American Cancer Society guideline for the early detection of prostate cancer: update 2010. CA Cancer J Clin. 2010;60(2):70–98. doi: 10.3322/caac.20066. [DOI] [PubMed] [Google Scholar]
- 146.Kimura T., Egawa S. Epidemiology of prostate cancer in Asian countries. Int. J. Urol. 2018;25(6):524–531. doi: 10.1111/iju.13593. [DOI] [PubMed] [Google Scholar]
- 147.McGlynn K.A., Petrick J.L., El-Serag H.B. Epidemiology of hepatocellular carcinoma. Hepatology. 2021;73(Suppl 1):4–13. doi: 10.1002/hep.31288. Suppl 1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 148.White D.L., Kanwal F., El-Serag H.B. Association between nonalcoholic fatty liver disease and risk for hepatocellular cancer, based on systematic review. Clin. Gastroenterol. Hepatol. 2012;10(12):1342. doi: 10.1016/j.cgh.2012.10.001. 59.e2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 149.Yang J.D., Roberts L.R. Hepatocellular carcinoma: a global view. Nat. Rev. Gastroenterol. Hepatol. 2010;7(8):448–458. doi: 10.1038/nrgastro.2010.100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 150.Bruix J., Reig M., Sherman M. Evidence-based diagnosis, staging, and treatment of patients with hepatocellular carcinoma. Gastroenterology. 2016;150(4):835–853. doi: 10.1053/j.gastro.2015.12.041. [DOI] [PubMed] [Google Scholar]
- 151.Llovet J.M., Kelley R.K., Villanueva A., Singal A.G., Pikarsky E., Roayaie S., et al. Hepatocellular carcinoma. Nat. Rev. Dis. Prim. 2021;7(1):6. doi: 10.1038/s41572-020-00240-3. [DOI] [PubMed] [Google Scholar]
- 152.Marrero J.A., Kulik L.M., Sirlin C.B., Zhu A.X., Finn R.S., Abecassis M.M., et al. Diagnosis, staging, and management of hepatocellular carcinoma: 2018 practice guidance by the American association for the study of liver diseases. Hepatology. 2018;68(2):723–750. doi: 10.1002/hep.29913. [DOI] [PubMed] [Google Scholar]
- 153.Melendez-Torres J., Singal A.G. Early detection of hepatocellular carcinoma: roadmap for improvement. Expert Rev. Anticancer Ther. 2022;22(6):621–632. doi: 10.1080/14737140.2022.2074404. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 154.Hanahan D., Weinberg Robert A. Hallmarks of cancer: the next generation. Cell. 2011;144(5):646–674. doi: 10.1016/j.cell.2011.02.013. [DOI] [PubMed] [Google Scholar]
- 155.Dunn G.P., Old L.J., Schreiber R.D. The immunobiology of cancer immunosurveillance and immunoediting. Immunity. 2004;21(2):137–148. doi: 10.1016/j.immuni.2004.07.017. [DOI] [PubMed] [Google Scholar]
- 156.Ribas A., Wolchok J.D. Cancer immunotherapy using checkpoint blockade. Science. 2018;359(6382):1350–1355. doi: 10.1126/science.aar4060. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 157.Lewis D.E., Giorgi J.V. Immunology of HIV infection. Int. Rev. Immunol. 1990;7(1):1–13. doi: 10.3109/08830189009061761. [DOI] [PubMed] [Google Scholar]
- 158.Gaseitsiwe S., Valentini D., Mahdavifar S., Reilly M., Ehrnst A., Maeurer M. Peptide microarray-based identification of Mycobacterium tuberculosis epitope binding to HLA-DRB1∗0101, DRB1∗1501, and DRB1∗0401. Clin. Vaccine Immunol. 2010;17(1):168–175. doi: 10.1128/CVI.00208-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 159.Arbour N.C., Lorenz E., Schutte B.C., Zabner J., Kline J.N., Jones M., et al. TLR4 mutations are associated with endotoxin hyporesponsiveness in humans. Nat. Genet. 2000;25(2):187–191. doi: 10.1038/76048. [DOI] [PubMed] [Google Scholar]
- 160.Zhang X., Zhang S., Yan X., Shan Y., Liu L., Zhou J., et al. m6A regulator-mediated RNA methylation modification patterns are involved in immune microenvironment regulation of periodontitis. J. Cell Mol. Med. 2021;25(7):3634–3645. doi: 10.1111/jcmm.16469. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 161.Khadela A., Chavda V.P., Postwala H., Shah Y., Mistry P., Apostolopoulos V. Epigenetics in tuberculosis: immunomodulation of host immune response. Vaccines (Basel) 2022;10(10) doi: 10.3390/vaccines10101740. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 162.Ma H., Yan Q.Z., Ma J.R., Li D.F., Yang J.L. Overview of the immunological mechanisms in hepatitis B virus reactivation: implications for disease progression and management strategies. World J. Gastroenterol. 2024;30(10):1295–1312. doi: 10.3748/wjg.v30.i10.1295. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 163.Skalsky R.L., Cullen B.R. Viruses, microRNAs, and host interactions. Annu. Rev. Microbiol. 2010;64:123–141. doi: 10.1146/annurev.micro.112408.134243. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 164.Li Z.K., Nie J.J., Li J., Zhuang H. The effect of HLA on immunological response to hepatitis B vaccine in healthy people: a meta-analysis. Vaccine. 2013;31(40):4355–4361. doi: 10.1016/j.vaccine.2013.06.108. [DOI] [PubMed] [Google Scholar]
- 165.Fu H., Pickering H., Rubbi L., Ross T.M., Zhou W., Reed E.F., et al. The response to influenza vaccination is associated with DNA methylation-driven regulation of T cell innate antiviral pathways. Clin Epigenetics. 2024;16(1):114. doi: 10.1186/s13148-024-01730-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 166.Kleinnijenhuis J., Quintin J., Preijers F., Joosten L.A., Ifrim D.C., Saeed S., et al. Bacille Calmette-Guerin induces NOD2-dependent nonspecific protection from reinfection via epigenetic reprogramming of monocytes. Proc Natl Acad Sci U S A. 2012;109(43):17537–17542. doi: 10.1073/pnas.1202870109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 167.Allison A.C. Protection afforded by sickle-cell trait against subtertian malareal infection. Br. Med. J. 1954;1(4857):290–294. doi: 10.1136/bmj.1.4857.290. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 168.Bethony J., Brooker S., Albonico M., Geiger S.M., Loukas A., Diemert D., et al. Soil-transmitted helminth infections: ascariasis, trichuriasis, and hookworm. Lancet. 2006;367(9521):1521–1532. doi: 10.1016/S0140-6736(06)68653-4. [DOI] [PubMed] [Google Scholar]
- 169.Guarnieri M., Balmes J.R. Outdoor air pollution and asthma. Lancet. 2014;383(9928):1581–1592. doi: 10.1016/S0140-6736(14)60617-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 170.WHO . World Health Organization; Geneva: 2020. Global Tuberculosis Report 2020. [Google Scholar]
- 171.Rovina N., Koutsoukou A., Koulouris N.G. Inflammation and immune response in COPD: where do we stand? Mediators Inflamm. 2013;2013 doi: 10.1155/2013/413735. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 172.Szabo G., Saha B. Alcohol's effect on host defense. Alcohol Res. 2015;37(2):159–170. doi: 10.35946/arcr.v37.2.01. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 173.Moss W.J. Measles. Lancet. 2017;390(10111):2490–2502. doi: 10.1016/S0140-6736(17)31463-0. [DOI] [PubMed] [Google Scholar]
- 174.Strachan D.P. Hay fever, hygiene, and household size. Bmj. 1989;299(6710):1259–1260. doi: 10.1136/bmj.299.6710.1259. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 175.Kassam Z., Lee C.H., Yuan Y., Hunt R.H. Fecal microbiota transplantation for Clostridium difficile infection: systematic review and meta-analysis. Am. J. Gastroenterol. 2013;108(4):500–508. doi: 10.1038/ajg.2013.59. [DOI] [PubMed] [Google Scholar]
- 176.Harris V.C., Haak B.W., Handley S.A., Jiang B., Velasquez D.E., Hykes B.L., et al. Effect of antibiotic-mediated microbiome modulation on rotavirus vaccine immunogenicity: a human, randomized-control proof-of-concept trial. Cell Host Microbe. 2018;24(2):197–207.e4. doi: 10.1016/j.chom.2018.07.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 177.WHO . World Health Organization; Geneva: 2021. World Malaria Report 2021. [Google Scholar]
- 178.WHO . World Health Organization; Geneva: 2021. Global Tuberculosis Report 2021. [Google Scholar]
- 179.Price-Haywood E.G., Burton J., Fort D., Seoane L. Hospitalization and mortality among black patients and white patients with covid-19. N. Engl. J. Med. 2020;382(26):2534–2543. doi: 10.1056/NEJMsa2011686. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 180.Looker K.J., Magaret A.S., Turner K.M., Vickerman P., Gottlieb S.L., Newman L.M. Global estimates of prevalent and incident herpes simplex virus type 2 infections in 2012. PLoS One. 2015;10(1) doi: 10.1371/journal.pone.0114989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 181.CDC . Centers for Disease Control and Prevention; 2021. HIV Surveillance Report 2019. [Google Scholar]
- 182.Williams T.N., Weatherall D.J. World distribution, population genetics, and health burden of the hemoglobinopathies. Cold Spring Harb Perspect Med. 2012;2(9) doi: 10.1101/cshperspect.a011692. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 183.Nkhoma E.T., Poole C., Vannappagari V., Hall S.A., Beutler E. The global prevalence of glucose-6-phosphate dehydrogenase deficiency: a systematic review and meta-analysis. Blood Cells Mol. Dis. 2009;42(3):267–278. doi: 10.1016/j.bcmd.2008.12.005. [DOI] [PubMed] [Google Scholar]
- 184.Band G., Le Q.S., Clarke G.M., Kivinen K., Hubbart C., Jeffreys A.E., et al. Insights into malaria susceptibility using genome-wide data on 17,000 individuals from Africa, Asia and Oceania. Nat. Commun. 2019;10(1):5732. doi: 10.1038/s41467-019-13480-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 185.Stoute J.A. Complement receptor 1 and malaria. Cell Microbiol. 2011;13(10):1441–1450. doi: 10.1111/j.1462-5822.2011.01648.x. [DOI] [PubMed] [Google Scholar]
- 186.Laurens M.B. The promise of a malaria vaccine-are we closer? Annu. Rev. Microbiol. 2018;72:273–292. doi: 10.1146/annurev-micro-090817-062427. [DOI] [PubMed] [Google Scholar]
- 187.Howes R.E., Battle K.E., Satyagraha A.W., Baird J.K., Hay S.I. G6PD deficiency: global distribution, genetic variants and primaquine therapy. Adv. Parasitol. 2013;81:133–201. doi: 10.1016/B978-0-12-407826-0.00004-7. [DOI] [PubMed] [Google Scholar]
- 188.Chang S.T., Linderman J.J., Kirschner D.E. Effect of multiple genetic polymorphisms on antigen presentation and susceptibility to Mycobacterium tuberculosis infection. Infect. Immun. 2008;76(7):3221–3232. doi: 10.1128/IAI.01677-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 189.Hargreaves J.R., Boccia D., Evans C.A., Adato M., Petticrew M., Porter J.D. The social determinants of tuberculosis: from evidence to action. Am J Public Health. 2011;101(4):654–662. doi: 10.2105/AJPH.2010.199505. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 190.Azad A.K., Lloyd C., Sadee W., Schlesinger L.S. Challenges of immune response diversity in the human population concerning new tuberculosis diagnostics, therapies, and vaccines. Front. Cell. Infect. Microbiol. 2020;10:139. doi: 10.3389/fcimb.2020.00139. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 191.Andersen P., Doherty T.M. The success and failure of BCG - implications for a novel tuberculosis vaccine. Nat. Rev. Microbiol. 2005;3(8):656–662. doi: 10.1038/nrmicro1211. [DOI] [PubMed] [Google Scholar]
- 192.Sim E., Abuhammad A., Ryan A. Arylamine N-acetyltransferases: from drug metabolism and pharmacogenetics to drug discovery. Br. J. Pharmacol. 2014;171(11):2705–2725. doi: 10.1111/bph.12598. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 193.CDC . Centers for Disease Control and Prevention; 2021. COVID Data Tracker. [Google Scholar]
- 194.Lopez Bernal J., Andrews N., Gower C., Robertson C., Stowe J., Tessier E., et al. Effectiveness of the Pfizer-BioNTech and Oxford-AstraZeneca vaccines on covid-19 related symptoms, hospital admissions, and mortality in older adults in England: test negative case-control study. Bmj. 2021;373 doi: 10.1136/bmj.n1088. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 195.Liu Y., Rocklöv J. The reproductive number of the Delta variant of SARS-CoV-2 is far higher compared to the ancestral SARS-CoV-2 virus. J Travel Med. 2021;28(7) doi: 10.1093/jtm/taab124. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 196.McColl E.R., Croyle M.A., Zamboni W.C., Honer W.G., Heise M., Piquette-Miller M., et al. COVID-19 vaccines and the virus: impact on drug metabolism and pharmacokinetics. Drug Metab. Dispos. 2023;51(1):130–141. doi: 10.1124/dmd.122.000934. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 197.WHO . World Health Organization; Geneva: 2024. Herpes Simplex Virus. [Google Scholar]
- 198.Yin Y., Favoreel H.W. Herpesviruses and the type III interferon system. Virol. Sin. 2021;36(4):577–587. doi: 10.1007/s12250-020-00330-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 199.Poole C.L., James S.H. Antiviral therapies for herpesviruses: current agents and new directions. Clin Ther. 2018;40(8):1282–1298. doi: 10.1016/j.clinthera.2018.07.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 200.Johnston C., Koelle D.M., Wald A. Current status and prospects for development of an HSV vaccine. Vaccine. 2014;32(14):1553–1560. doi: 10.1016/j.vaccine.2013.08.066. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 201.Kiepiela P., Leslie A.J., Honeyborne I., Ramduth D., Thobakgale C., Chetty S., et al. Dominant influence of HLA-B in mediating the potential co-evolution of HIV and HLA. Nature. 2004;432(7018):769–775. doi: 10.1038/nature03113. [DOI] [PubMed] [Google Scholar]
- 202.UNAIDS . 2021. Global HIV & AIDS Statistics — Fact Sheet. UNAIDS 2024 Epidemiological Estimates. [Google Scholar]
- 203.Semba R.D., Tang A.M. Micronutrients and the pathogenesis of human immunodeficiency virus infection. Br. J. Nutr. 1999;81(3):181–189. doi: 10.1017/s0007114599000379. [DOI] [PubMed] [Google Scholar]
- 204.Samson M., Libert F., Doranz B.J., Rucker J., Liesnard C., Farber C.M., et al. Resistance to HIV-1 infection in caucasian individuals bearing mutant alleles of the CCR-5 chemokine receptor gene. Nature. 1996;382(6593):722–725. doi: 10.1038/382722a0. [DOI] [PubMed] [Google Scholar]
- 205.Wyen C., Hendra H., Vogel M., Hoffmann C., Knechten H., Brockmeyer N.H., et al. Impact of CYP2B6 983T>C polymorphism on non-nucleoside reverse transcriptase inhibitor plasma concentrations in HIV-infected patients. J. Antimicrob. Chemother. 2008;61(4):914–918. doi: 10.1093/jac/dkn029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 206.Tishkoff S.A., Verrelli B.C. Patterns of human genetic diversity: implications for human evolutionary history and disease. Annu Rev Genomics Hum Genet. 2003;4:293–340. doi: 10.1146/annurev.genom.4.070802.110226. [DOI] [PubMed] [Google Scholar]
- 207.Farmer P.E., Nizeye B., Stulac S., Keshavjee S. Structural violence and clinical medicine. PLoS Med. 2006;3(10) doi: 10.1371/journal.pmed.0030449. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 208.Ashley E.A. Towards precision medicine. Nat. Rev. Genet. 2016;17(9):507–522. doi: 10.1038/nrg.2016.86. [DOI] [PubMed] [Google Scholar]
- 209.Lähnemann D., Köster J., Szczurek E., McCarthy D.J., Hicks S.C., Robinson M.D., et al. Eleven grand challenges in single-cell data science. Genome Biol. 2020;21(1):31. doi: 10.1186/s13059-020-1926-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 210.Naidoo L., Arumugam T., Ramsuran V. Host genetic impact on infectious diseases among different ethnic groups. Adv. Genet. 2023;4(4) doi: 10.1002/ggn2.202300181. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 211.Stunnenberg H.G., Hirst M. The international human epigenome Consortium: a blueprint for scientific collaboration and discovery. Cell. 2016;167(5):1145–1149. doi: 10.1016/j.cell.2016.11.007. [DOI] [PubMed] [Google Scholar]
- 212.Zach M., Greslehner G.P. Understanding immunity: an alternative framework beyond defense and strength. Biol. Philos. 2023;38(1):7. doi: 10.1007/s10539-023-09893-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 213.Netea M.G., Domínguez-Andrés J., Barreiro L.B., Chavakis T., Divangahi M., Fuchs E., et al. Defining trained immunity and its role in health and disease. Nat. Rev. Immunol. 2020;20(6):375–388. doi: 10.1038/s41577-020-0285-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 214.Brodin P., Davis M.M. Human immune system variation. Nat. Rev. Immunol. 2017;17(1):21–29. doi: 10.1038/nri.2016.125. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 215.Barreiro L.B., Quintana-Murci L. From evolutionary genetics to human immunology: how selection shapes host defence genes. Nat. Rev. Genet. 2010;11(1):17–30. doi: 10.1038/nrg2698. [DOI] [PubMed] [Google Scholar]
- 216.Ghobadinezhad F., Ebrahimi N., Mozaffari F., Moradi N., Beiranvand S., Pournazari M., et al. The emerging role of regulatory cell-based therapy in autoimmune disease. Front. Immunol. 2022;13 doi: 10.3389/fimmu.2022.1075813. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 217.Quintana-Murci L. Understanding rare and common diseases in the context of human evolution. Genome Biol. 2016;17(1):225. doi: 10.1186/s13059-016-1093-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 218.Scepanovic P., Alanio C., Hammer C., Hodel F., Bergstedt J., Patin E., et al. Human genetic variants and age are the strongest predictors of humoral immune responses to common pathogens and vaccines. Genome Med. 2018;10(1):59. doi: 10.1186/s13073-018-0568-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 219.NIAID Diseases disproportionately affecting minorities. 2023. https://www.niaid.nih.gov/research/diseases-disproportionately-affecting-minorities [Available from:
- 220.Noppert G.A., Stebbins R.C., Dowd J.B., Aiello A.E. Socioeconomic and race/ethnic differences in immunosenescence: evidence from the health and retirement study. Brain Behav. Immun. 2023;107:361–368. doi: 10.1016/j.bbi.2022.10.019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 221.Hierink F., Okiro E.A., Flahault A., Ray N. The winding road to health: a systematic scoping review on the effect of geographical accessibility to health care on infectious diseases in low- and middle-income countries. PLoS One. 2021;16(1) doi: 10.1371/journal.pone.0244921. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 222.Flandroy L., Poutahidis T., Berg G., Clarke G., Dao M.-C., Decaestecker E., et al. The impact of human activities and lifestyles on the interlinked microbiota and health of humans and of ecosystems. Sci. Total Environ. 2018;627:1018–1038. doi: 10.1016/j.scitotenv.2018.01.288. [DOI] [PubMed] [Google Scholar]
- 223.Hinata N., Fujisawa M. Racial differences in prostate cancer characteristics and cancer-specific mortality: an overview. World J Mens Health. 2022;40(2):217–227. doi: 10.5534/wjmh.210070. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 224.Schakel L., Veldhuijzen D.S., Crompvoets P.I., Bosch J.A., Cohen S., van Middendorp H., et al. Effectiveness of stress-reducing interventions on the response to challenges to the immune system: a meta-analytic review. Psychother. Psychosom. 2019;88(5):274–286. doi: 10.1159/000501645. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 225.Scheffer D.D.L., Latini A. Exercise-induced immune system response: anti-inflammatory status on peripheral and central organs. Biochim. Biophys. Acta, Mol. Basis Dis. 2020;1866(10) doi: 10.1016/j.bbadis.2020.165823. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 226.Suzuki T., Hidaka T., Kumagai Y., Yamamoto M. Environmental pollutants and the immune response. Nat. Immunol. 2020;21(12):1486–1495. doi: 10.1038/s41590-020-0802-6. [DOI] [PubMed] [Google Scholar]
- 227.Xu Z., Jiang W., Huang W., Lin Y., Chan F.K.L., Ng S.C. Gut microbiota in patients with obesity and metabolic disorders - a systematic review. Genes Nutr. 2022;17(1):2. doi: 10.1186/s12263-021-00703-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 228.Desai A.N., Majumder M.S. What is herd immunity? JAMA. 2020;324(20):2113. doi: 10.1001/jama.2020.20895. [DOI] [PubMed] [Google Scholar]
- 229.Shelly A., Gupta P., Ahuja R., Srichandan S., Meena J., Majumdar T. Impact of microbiota: a paradigm for evolving herd immunity against viral diseases. Viruses. 2020;12(10) doi: 10.3390/v12101150. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 230.Lane J.M. Mass vaccination and surveillance/containment in the eradication of smallpox. Curr. Top. Microbiol. Immunol. 2006;304:17–29. doi: 10.1007/3-540-36583-4_2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 231.Ali M., Emch M., von Seidlein L., Yunus M., Sack D.A., Rao M., et al. Herd immunity conferred by killed oral cholera vaccines in Bangladesh: a reanalysis. Lancet. 2005;366(9479):44–49. doi: 10.1016/S0140-6736(05)66550-6. [DOI] [PubMed] [Google Scholar]
- 232.Mast T.C., Wang F.T., Su S., Seeger J.D. Evidence of herd immunity and sustained impact of rotavirus vaccination on the reduction of rotavirus-related medical encounters among infants from 2006 through 2011 in the United States. Pediatr. Infect. Dis. J. 2015;34(6):615–620. doi: 10.1097/INF.0000000000000702. [DOI] [PubMed] [Google Scholar]
- 233.Kim T.H., Johnstone J., Loeb M. Vaccine herd effect. Scand. J. Infect. Dis. 2011;43(9):683–689. doi: 10.3109/00365548.2011.582247. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 234.Fine P., Eames K., Heymann D.L. “Herd immunity”: a rough guide. Clin. Infect. Dis. 2011;52(7):911–916. doi: 10.1093/cid/cir007. [DOI] [PubMed] [Google Scholar]
- 235.Metcalf C.J.E., Ferrari M., Graham A.L., Grenfell B.T. Understanding herd immunity. Trends Immunol. 2015;36(12):753–755. doi: 10.1016/j.it.2015.10.004. [DOI] [PubMed] [Google Scholar]
- 236.Liu Y., Gayle A.A., Wilder-Smith A., Rocklöv J. The reproductive number of COVID-19 is higher compared to SARS coronavirus. J Travel Med. 2020;27(2) doi: 10.1093/jtm/taaa021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 237.Kwok K.O., Lai F., Wei W.I., Wong S.Y.S., Tang J.W.T. Herd immunity - estimating the level required to halt the COVID-19 epidemics in affected countries. J. Infect. 2020;80(6):e32–e33. doi: 10.1016/j.jinf.2020.03.027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 238.Neuhausen S., Gilewski T., Norton L., Tran T., McGuire P., Swensen J., et al. Recurrent BRCA2 6174delT mutations in Ashkenazi Jewish women affected by breast cancer. Nat. Genet. 1996;13(1):126–128. doi: 10.1038/ng0596-126. [DOI] [PubMed] [Google Scholar]
- 239.Levy-Lahad E., Catane R., Eisenberg S., Kaufman B., Hornreich G., Lishinsky E., et al. Founder BRCA1 and BRCA2 mutations in Ashkenazi Jews in Israel: frequency and differential penetrance in ovarian cancer and in breast-ovarian cancer families. Am. J. Hum. Genet. 1997;60(5):1059–1067. [PMC free article] [PubMed] [Google Scholar]
- 240.Espinal A.C., Buas M.F., Wang D., Cheng D.T., Sucheston-Campbell L., Hu Q., et al. FOXA1 hypermethylation: link between parity and ER-negative breast cancer in African American women? Breast Cancer Res. Treat. 2017;166(2):559–568. doi: 10.1007/s10549-017-4418-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 241.Dong H.-Y., Ding L., Zhou T.-R., Yan T., Li J., Liang C. FOXA1 in prostate cancer. Asian J. Androl. 2023;25(3):287–295. doi: 10.4103/aja202259. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 242.Huang F.W., Mosquera J.M., Garofalo A., Oh C., Baco M., Amin-Mansour A., et al. Exome sequencing of african-American prostate cancer reveals loss-of-function ERF mutations. Cancer Discov. 2017;7(9):973–983. doi: 10.1158/2159-8290.CD-16-0960. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 243.Magi-Galluzzi C., Tsusuki T., Elson P., Simmerman K., LaFargue C., Esgueva R., et al. TMPRSS2–ERG gene fusion prevalence and class are significantly different in prostate cancer of caucasian, african-american and Japanese patients. Prostate. 2011;71(5):489–497. doi: 10.1002/pros.21265. [DOI] [PubMed] [Google Scholar]
- 244.Rosen P., Pfister D., Young D., Petrovics G., Chen Y., Cullen J., et al. Differences in frequency of ERG oncoprotein expression between index tumors of Caucasian and African American patients with prostate cancer. Urology. 2012;80(4):749–753. doi: 10.1016/j.urology.2012.07.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 245.Reis L.B., Bakos R.M., Vianna F.S.L., Macedo G.S., Jacovas V.C., Ribeiro-Dos-Santos A.M., et al. Skin pigmentation polymorphisms associated with increased risk of melanoma in a case-control sample from southern Brazil. BMC Cancer. 2020;20(1):1069. doi: 10.1186/s12885-020-07485-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 246.Rochmah N., Arief F., Faizi M., Basuki S. The association between PTPN22 C1858T gene polymorphism and type 1 diabetes mellitus: an Indonesian study. Ann. Med. 2023;55(1):1211–1215. doi: 10.1080/07853890.2023.2190162. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 247.GeneCards. IFIH1 Gene - Interferon Induced with Helicase C Domain 1. https://www.genecards.org/cgi-bin/carddisp.pl?gene=IFIH1.
- 248.Barton A., Eyre S., Bowes J., Ho P., John S., Worthington J. Investigation of the SLC22A4 gene (associated with rheumatoid arthritis in a Japanese population) in a United Kingdom population of rheumatoid arthritis patients. Arthritis Rheum. 2005;52(3):752–758. doi: 10.1002/art.20877. [DOI] [PubMed] [Google Scholar]
- 249.Wakitani S., Murata N., Toda Y., Ogawa R., Kaneshige T., Nishimura Y., et al. The relationship between HLA-DRB1 alleles and disease subsets of rheumatoid arthritis in Japanese. Rheumatology. 1997;36(6):630–636. doi: 10.1093/rheumatology/36.6.630. [DOI] [PubMed] [Google Scholar]
- 250.Hughes L.B., Reynolds R.J., Brown E.E., Kelley J.M., Thomson B., Conn D.L., et al. Most common single-nucleotide polymorphisms associated with rheumatoid arthritis in persons of European ancestry confer risk of rheumatoid arthritis in African Americans. Arthritis Rheum. 2010;62(12):3547–3553. doi: 10.1002/art.27732. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 251.Shao M., Xie H., Yang H., Xu W., Chen Y., Gao X., et al. Association of interleukin-6 promoter polymorphism with rheumatoid arthritis: a meta-analysis with trial sequential analysis. Clin. Rheumatol. 2022;41(2):411–419. doi: 10.1007/s10067-021-05886-2. [DOI] [PubMed] [Google Scholar]
- 252.Ren T-l, Han Z-j, Yang C-j, Hang Y-x, Fang D-y, Wang K., et al. Association of SLC22A4 gene polymorphism with rheumatoid arthritis in the Chinese population. J. Biochem. Mol. Toxicol. 2014;28(5):206–210. doi: 10.1002/jbt.21554. [DOI] [PubMed] [Google Scholar]
- 253.Wang C., Ahlford A., Järvinen T.M., Nordmark G., Eloranta M.L., Gunnarsson I., et al. Genes identified in Asian SLE GWASs are also associated with SLE in Caucasian populations. Eur. J. Hum. Genet. 2013;21(9):994–999. doi: 10.1038/ejhg.2012.277. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 254.Harley J.B., Kelly J.A. Genetic basis of systemic lupus erythematosus: a review of the unique genetic contributions in African Americans. J. Natl. Med. Assoc. 2002;94(8):670–677. [PMC free article] [PubMed] [Google Scholar]
- 255.Sun C., Molineros J.E., Looger L.L., Zhou X.J., Kim K., Okada Y., et al. High-density genotyping of immune-related loci identifies new SLE risk variants in individuals with Asian ancestry. Nat. Genet. 2016;48(3):323–330. doi: 10.1038/ng.3496. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 256.Hull J., Shackleton S., Harris A. The stop mutation R553X in the CFTR gene results in exon skipping. Genomics. 1994;19(2):362–364. doi: 10.1006/geno.1994.1070. [DOI] [PubMed] [Google Scholar]
- 257.Petrova N., Balinova N., Marakhonov A., Vasilyeva T., Kashirskaya N., Galkina V., et al. Ethnic differences in the frequency of CFTR gene mutations in populations of the European and North caucasian part of the Russian federation. Front. Genet. 2021;12 doi: 10.3389/fgene.2021.678374. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 258.Araújo F.G., Novaes F.C., Santos N.P., Martins V.C., Souza S.M., Santos S.E., et al. Prevalence of deltaF508, G551D, G542X, and R553X mutations among cystic fibrosis patients in the North of Brazil. Braz. J. Med. Biol. Res. 2005;38(1):11–15. doi: 10.1590/s0100-879x2005000100003. [DOI] [PubMed] [Google Scholar]
- 259.Onouchi Y., Fukazawa R., Yamamura K., Suzuki H., Kakimoto N., Suenaga T., et al. Variations in ORAI1 gene associated with kawasaki disease. PLoS One. 2016;11(1) doi: 10.1371/journal.pone.0145486. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 260.Downes D.J., Cross A.R., Hua P., Roberts N., Schwessinger R., Cutler A.J., et al. Identification of LZTFL1 as a candidate effector gene at a COVID-19 risk locus. Nat. Genet. 2021;53(11):1606–1615. doi: 10.1038/s41588-021-00955-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 261.Gelabert P., Olalde I., de-Dios T., Civit S., Lalueza-Fox C. Malaria was a weak selective force in ancient Europeans. Sci. Rep. 2017;7(1):1377. doi: 10.1038/s41598-017-01534-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 262.Yazdanbakhsh K., Rios M., Storry J.R., Kosower N., Parasol N., Chaudhuri A., et al. Molecular mechanisms that lead to reduced expression of Duffy antigens. Transfusion. 2000;40(3):310–320. doi: 10.1046/j.1537-2995.2000.40030310.x. [DOI] [PubMed] [Google Scholar]
- 263.Hamblin M.T., Di Rienzo A. Detection of the signature of natural selection in humans: evidence from the duffy blood group locus. Am. J. Hum. Genet. 2000;66(5):1669–1679. doi: 10.1086/302879. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 264.Awab G.R., Aaram F., Jamornthanyawat N., Suwannasin K., Pagornrat W., Watson J.A., et al. Protective effect of Mediterranean-type glucose-6-phosphate dehydrogenase deficiency against Plasmodium vivax malaria. Elife. 2021;10 doi: 10.7554/eLife.62448. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 265.da Rocha J., Othman H., Tiemessen C.T., Botha G., Ramsay M., Masimirembwa C., et al. G6PD variant distribution in sub-Saharan Africa and potential risks of using chloroquine/hydroxychloroquine based treatments for COVID-19. Pharmacogenomics J. 2021;21:649–656. doi: 10.1038/s41397-021-00242-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 266.Ni J., Wang D., Wang S. The CCR5-delta32 genetic polymorphism and HIV-1 infection susceptibility: a meta-analysis. Open Med. 2018;13:467–474. doi: 10.1515/med-2018-0062. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 267.Kiepiela P., Leslie A.J., Honeyborne I., Ramduth D., Thobakgale C., Chetty S., et al. Dominant influence of HLA-B in mediating the potential co-evolution of HIV and HLA. Nature. 2004;432(7018):769–775. doi: 10.1038/nature03113. [DOI] [PubMed] [Google Scholar]
- 268.Javanbakht H., An P., Gold B., Petersen D.C., O'Huigin C., Nelson G.W., et al. Effects of human TRIM5α polymorphisms on antiretroviral function and susceptibility to human immunodeficiency virus infection. Virology. 2006;354(1):15–27. doi: 10.1016/j.virol.2006.06.031. [DOI] [PubMed] [Google Scholar]
- 269.Compaore T.R., Ouattara A., Baguiya A., Traore L., Zouré A.A., Ouedraogo H., et al. APOBEC3G role on HIV infection in Africa: a systematic review. Am. J. Mol. Biol. 2024;14:25–42. [Google Scholar]
- 270.Martin M.P., Gao X., Lee J.-H., Nelson G.W., Detels R., Goedert J.J., et al. Epistatic interaction between KIR3DS1 and HLA-B delays the progression to AIDS. Nat. Genet. 2002;31(4):429–434. doi: 10.1038/ng934. [DOI] [PubMed] [Google Scholar]
- 271.Li X., Yang Y., Zhou F., Zhang Y., Lu H., Jin Q., et al. SLC11A1 (NRAMP1) polymorphisms and tuberculosis susceptibility: updated systematic review and meta-analysis. PLoS One. 2011;6(1) doi: 10.1371/journal.pone.0015831. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 272.Liu Y., Zhao E., Zhu L., Zhang D., Wang Z. 3'UTR polymorphisms in NRAMP1 are associated with the susceptibility to pulmonary tuberculosis: a MOOSE-compliant meta-analysis. Medicine (Baltim.) 2019;98(23) doi: 10.1097/MD.0000000000015955. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 273.Hu B., Wang Y., Wang Z., He X., Wang L., Yuan D., et al. Association of SLC11A1 polymorphisms with tuberculosis susceptibility in the Chinese han population. Front. Genet. 2022;13 doi: 10.3389/fgene.2022.899124. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 274.Bellamy R., Beyers N., McAdam K.P.W.J., Ruwende C., Gie R., Samaai P., et al. Genetic susceptibility to tuberculosis in Africans: a genome-wide scan. Proc. Natl. Acad. Sci. USA. 2000;97(14):8005–8009. doi: 10.1073/pnas.140201897. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 275.Teng M., Zhou S., Cai C., Lupien M., He H.H. Pioneer of prostate cancer: past, present and the future of FOXA1. Protein Cell. 2021;12(1):29–38. doi: 10.1007/s13238-020-00786-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 276.Soetjipto Rochmah N., Faizi M., Hisbiyah Y., Endaryanto A. HLA-DQA1 and HLA-DQB1 gene polymorphism in Indonesian children with type I diabetes mellitus. Appl. Clin. Genet. 2022;15:11–17. doi: 10.2147/TACG.S348115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 277.Onengut-Gumuscu S., Chen W.M., Robertson C.C., Bonnie J.K., Farber E., Zhu Z., et al. Type 1 diabetes risk in african-ancestry participants and utility of an ancestry-specific genetic risk score. Diabetes Care. 2019;42(3):406–415. doi: 10.2337/dc18-1727. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 278.El Shahawy A., Abdel Wahab Mohammad L., Nahla M., Gaballah N.N.M.G., M Elalawi S. Association of solute carrier 22 member 4 gene polymorphism with rheumatoid arthritis. Zagazig University Medical Journal. 2024;30(2):533–540. [Google Scholar]
- 279.Jiang H., Promchan K., Lin B.R., Lockett S., Chen D., Marshall H., et al. LZTFL1 upregulated by all-trans retinoic acid during CD4+ T cell activation enhances IL-5 production. J. Immunol. 2016;196(3):1081–1090. doi: 10.4049/jimmunol.1500719. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 280.Willcocks L.C., Carr E.J., Niederer H.A., Rayner T.F., Williams T.N., Yang W., et al. A defunctioning polymorphism in FCGR2B is associated with protection against malaria but susceptibility to systemic lupus erythematosus. Proc Natl Acad Sci U S A. 2010;107(17):7881–7885. doi: 10.1073/pnas.0915133107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 281.Drake T.M., Søreide K. Cancer epigenetics in solid organ tumours: a primer for surgical oncologists. Eur. J. Surg. Oncol. 2019;45(5):736–746. doi: 10.1016/j.ejso.2019.02.005. [DOI] [PubMed] [Google Scholar]