Abstract
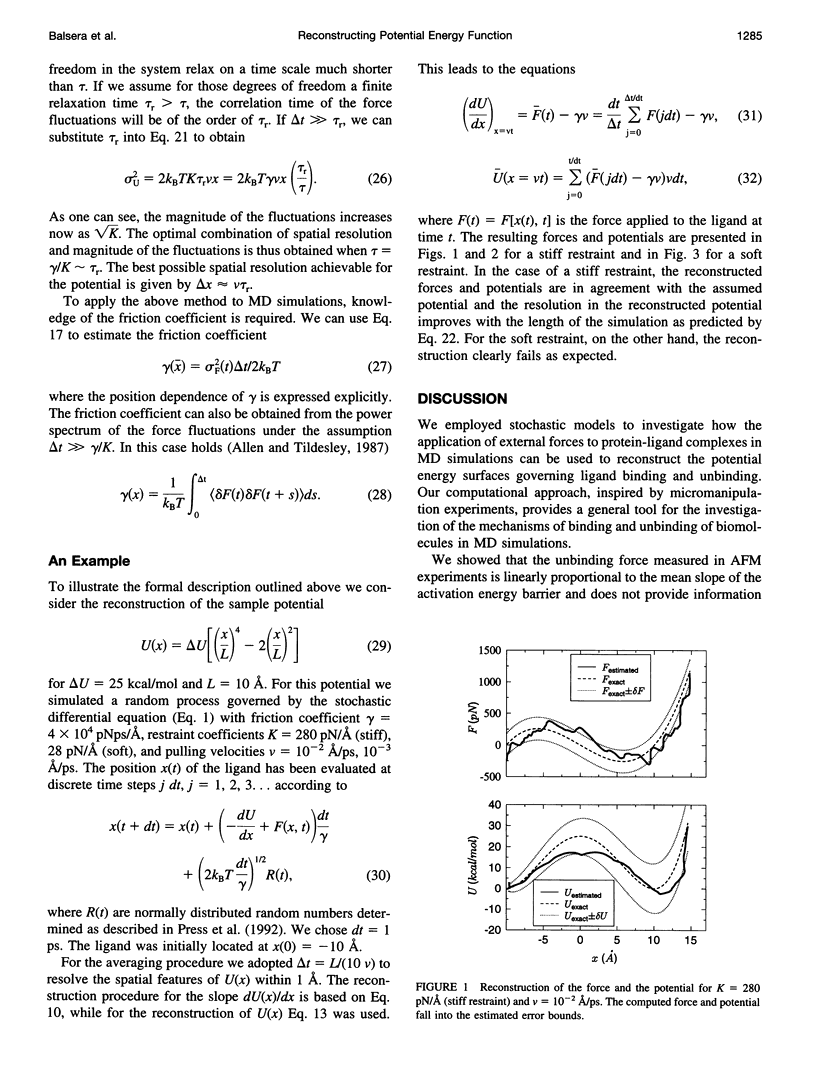
One-dimensional stochastic models demonstrate that molecular dynamics simulations of a few nanoseconds can be used to reconstruct the essential features of the binding potential of macromolecules. This can be accomplished by inducing the unbinding with the help of external forces applied to the molecules, and discounting the irreversible work performed on the system by these forces. The fluctuation-dissipation theorem sets a fundamental limit on the precision with which the binding potential can be reconstructed by this method. The uncertainty in the resulting potential is linearly proportional to the irreversible component of work performed on the system during the simulation. These results provide an a priori estimate of the energy barriers observable in molecular dynamics simulations.
Full text
PDF





Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Bell G. I. Models for the specific adhesion of cells to cells. Science. 1978 May 12;200(4342):618–627. doi: 10.1126/science.347575. [DOI] [PubMed] [Google Scholar]
- Chilkoti A., Boland T., Ratner B. D., Stayton P. S. The relationship between ligand-binding thermodynamics and protein-ligand interaction forces measured by atomic force microscopy. Biophys J. 1995 Nov;69(5):2125–2130. doi: 10.1016/S0006-3495(95)80083-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Evans E., Ritchie K. Dynamic strength of molecular adhesion bonds. Biophys J. 1997 Apr;72(4):1541–1555. doi: 10.1016/S0006-3495(97)78802-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Evans E., Ritchie K., Merkel R. Sensitive force technique to probe molecular adhesion and structural linkages at biological interfaces. Biophys J. 1995 Jun;68(6):2580–2587. doi: 10.1016/S0006-3495(95)80441-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Florin E. L., Moy V. T., Gaub H. E. Adhesion forces between individual ligand-receptor pairs. Science. 1994 Apr 15;264(5157):415–417. doi: 10.1126/science.8153628. [DOI] [PubMed] [Google Scholar]
- Grubmüller H., Heymann B., Tavan P. Ligand binding: molecular mechanics calculation of the streptavidin-biotin rupture force. Science. 1996 Feb 16;271(5251):997–999. doi: 10.1126/science.271.5251.997. [DOI] [PubMed] [Google Scholar]
- Izrailev S., Stepaniants S., Balsera M., Oono Y., Schulten K. Molecular dynamics study of unbinding of the avidin-biotin complex. Biophys J. 1997 Apr;72(4):1568–1581. doi: 10.1016/S0006-3495(97)78804-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moy V. T., Florin E. L., Gaub H. E. Intermolecular forces and energies between ligands and receptors. Science. 1994 Oct 14;266(5183):257–259. doi: 10.1126/science.7939660. [DOI] [PubMed] [Google Scholar]


