Abstract
The interaction of NKG2D, a stimulatory receptor expressed on natural killer (NK) cells and activated CD8+ T cells, and its ligands mediates stimulatory and costimulatory signals to these cells. Here, we demonstrate that DNA-based vaccines, encoding syngeneic or allogeneic NKG2D ligands together with tumor antigens such as survivin or carcinoembryonic antigen, markedly activate both innate and adaptive antitumor immunity. Such vaccines result in highly effective, NK- and CD8+ T cell-mediated protection against either breast or colon carcinoma cells in prophylactic and therapeutic settings. Notably, this protection was irrespective of the NKG2D ligand expression level of the tumor cells. Hence, this strategy has the potential to lead to widely applicable and possibly clinically useful DNA-based cancer vaccines.
Keywords: NK cells, NKG2D ligands, T cells, survivin, Peyer's patches
One way to induce potent immune responses against tumors is to activate the key immune effector mechanisms. The NKG2D receptor–ligand interaction is a good candidate for such a task because it occurs at the crossroad between innate and adaptive immunity (1). NKG2D, a stimulatory lectin-like receptor, is expressed on natural killer (NK) cells, activated CD8+ T cells, γδ T cells, and activated macrophages (2). It mediates costimulatory signals for CD8+ T cells and stimulatory signals for NK cells and macrophages (3, 4). Ligands for NKG2D are related to class I major histocompatibility complex (MHC) molecules. In mice, NKG2D ligands include products of the retinoic acid early inducible-1 (RAE1) gene, H60 (2, 5), and UL16-binding protein-like transcript 1 (MULT1) (6). Importantly, in syngeneic mice, ectopic expression of NKG2D ligands causes NK cell-mediated rejection of transfected tumor cells (7, 8). It also primes cytotoxic T cells (CTLs), which are responsible for the rejection of subsequent challenges with tumor cells that lacked NKG2D ligands (7).
In this study, we tested our hypothesis that by engaging the NKG2D receptor, we would enhance the antitumor efficacy of DNA-based cancer vaccines by activating both innate and adaptive immunity. This task was accomplished in two different mouse model systems, a survivin-based DNA vaccine in syngeneic BALB/c mice (9) and a carcinoembryonic antigen (CEA)-based DNA vaccine in CEA-A2Kb double transgenic mice (10).
Survivin, a 16.5-kDa inhibitor of apoptosis protein, represents an almost ideal target for cancer vaccines because it is overexpressed by essentially all solid tumors and proliferating endothelial cells in the tumor vasculature. In contrast, it is poorly or only transiently expressed by normal adult tissues (11). Furthermore, overexpression of survivin in tumors is linked to decreased patient survival, increased tumor recurrence, and resistance to therapy (11), and spontaneous immune responses against survivin were recently demonstrated in various cancer patients (12). Survivin-based DNA vaccines were able to induce T cell-mediated antitumor responses without severe toxicity in mouse (9) and in clinical trials (13).
The 180-kDa oncofetal protein CEA is another appropriate vaccine target because it is overexpressed on a variety of tumors (14) but only expressed weakly on normal epithelial cells (15). This expression pattern, together with its immunogenicity, led to the development and evaluation of a variety of CEA whole gene-based or peptide-based vaccines (16, 17). In addition, techniques to measure CEA-specific CTL responses are well established (10, 18) and facilitate the verification of the specificity of immune responses induced by such vaccines.
Here, we demonstrate that engagement of the murine NKG2D receptor enhances both innate and adaptive immune responses induced by DNA-based vaccines and thereby augments their antitumor efficacies in both prophylactic and therapeutic settings.
Materials and Methods
Vector Construction and Transformation of Salmonella typhimurium. Expression vectors were constructed based on a pBudCE4.1 backbone (Invitrogen) as indicated. The double attenuated S. typhimurium (AroA-; dam-) strain RE88 was kindly provided by Remedyne Corporation (Santa Barbara, CA) and was transformed with DNA vaccine plasmids by electroporation as described in ref. 19.
Animals, Cell Lines, and Peptide. Female BALB/c mice, 6–8 weeks of age, were purchased from The Jackson Laboratory. C57BL/6J-CEA-A2Kb double transgenic mice were generated as described in refs. 10, 20, and 21. All animal experiments were performed according to the National Institutes of Health Guide for the Care and Use of Laboratory Animals.
The murine colon carcinoma cell line MC-38-CEA-A2Kb was generated as described in refs. 10 and 20. Murine colon carcinoma cell line CT-26 was provided by I. J. Fidler (MD Anderson Cancer Center, Houston). Yac-1 cells were purchased from American Type Culture Collection (Rockville, MD), and the murine breast carcinoma cell line D2F2 was kindly supplied by W.-Z. Wei (Wayne State University, Detroit). T2, a human HLA-A2+ cell line, was originally obtained from P. Cresswell (Yale University, New Haven, CT) and kindly provided by L. A. Sherman (The Scripps Research Institute). A plasmid containing the full-length murine NKG2D ligand-H60 and PE-conjugated NKG2D tetramer were generously gifted by A. Diefenbach and D. H. Raulet (University of California, Berkeley). CEA691 peptide (18) was kindly provided by E. Celis (Mayo Clinic, Rochester, MN) with >95% purity by HPLC.
Oral Immunization and Tumor Cell Challenge. Mice were immunized twice within a 2-week interval by gavage with 100 μl of PBS containing ≈5 × 108 doubly mutated S. typhimurium harboring the expression vectors. In prophylactic models, mice were challenged with tumor cells as indicated 2 weeks after the last vaccination and in therapeutic settings 5 days before the first vaccination.
Cytotoxicity and ELISPOT Assays. Cytotoxicity was measured by a standard 51Cr-release assay as described in ref. 22. ELISPOT assays were performed with an ELISPOT kit (BD Pharmingen) according to the instructions provided by the manufacturer.
Immunofluorescence. This procedure was performed on cryostat sections of Peyer's Patches fixed with acetone. Primary antibodies used were anti-mouse dendritic cell Ab, anti-mouse CD11c Ab (BD Pharmingen), recombinant mouse NKG2D/Fc Chimera (R & D Systems), and rabbit anti-human CEA Ab (DAKO). All secondary antibodies were purchased from Molecular Probes. Slides were mounted with antifade reagent (Molecular Probes) and examined with a confocal microscope (Axiovert 100TV, Zeiss).
In Vivo Depletions. These depletions were performed by i.p. injection of anti-CD8 Ab 2.43 (National Cell Culture Center, Brooklyn Center, MN) at 0.6 mg per mouse or anti-CD4 Ab GK1.5 (National Cell Culture Center) at 0.4 mg per mouse 1 day before tumor cell challenge and repeated weekly, or with antiasialo GM1 Ab (Wako Chemicals, Richmond, VA) at 20 μl per mouse 1 day before tumor challenge and repeated every 5 days.
Statistical Analysis. The statistical significance of differential findings between experimental groups and controls was determined by using Student's t test. Findings were regarded as significant if two-tailed P values were <0.05.
Results
NKG2D Ligand-H60 Improves Antitumor Efficacy of DNA Vaccines Encoding Murine Survivin. Expression vectors encoding H60 (pH60), survivin (pSurv), or both (pH60/Surv) were constructed on a pBudCE4.1 (pBud) backbone (Fig. 1A). The expression of survivin in transfected 293T cells was confirmed by Western blot analysis (Fig. 1B), and the expression of H60 was confirmed by positive staining with NKG2D tetramer (Fig. 1C). To evaluate the vaccine-induced H60 expression in vivo, mice were killed 24 h after pH60 vaccination, and cryostat sections of Peyer's patches were analyzed for NKG2D ligand expression. As shown in Fig. 1D, NKG2D ligands (red) were detected but only at low levels in mice vaccinated with the pBud control vector. This result is not surprising, because NKG2D ligand expression can be induced by infection or cellular stress (23, 24). However, in pH60-vaccinated mice, NKG2D ligands, presumably mostly H60, are expressed at much higher levels. Notably, NKG2D expression was found inside dendritic cells (DC), proving the successful delivery of H60 to DCs. However, H60 was also expressed by other cell types, presumably macrophages (25, 26).
Fig. 1.
Construction and expression of vectors encoding murine NKG2D ligand-H60 and survivin. (A) Expression vectors were constructed based on the pBudCE4.1 plasmid backbone. (B) Expression of murine survivin was detected by Western blotting analyses of pSurv-transfected 293T cells, by using rabbit anti-mouse survivin antibody. (C) Expression of H60 was demonstrated by flow cytometry using NKG2D tetramer. 293T cells were transfected with either pH60 (thick black line) or pBud vectors (thin gray line). (D) Expression of NKG2D ligands in Peyer's patches in vivo. Mice were vaccinated with either pBud (Left) or pH60 (Right). Green, DC Ab; red, NKG2D/Fc chimera; arrow-head, colocalization; *, both markers expressed on the same cells without colocalization.
To test our hypothesis that NKG2D ligand-H60 can enhance the antitumor efficacy of survivin-based DNA vaccines, vaccinated syngeneic mice were challenged with murine colon carcinoma cell line CT-26. CT-26 expressed survivin (Fig. 2A Left) and at best expressed NKG2D ligands weakly (Fig. 2 A Right). Tumor cells with low or negative NKG2D ligand expression had been reported to be more tumorigenic (7). Likewise, as shown in our experiment, tumors colonized in the lungs of all mice treated with PBS or pBud 25 days after challenge (Fig. 2B). Importantly, only a DNA vaccine encoding both a NKG2D ligand (H60) and survivin effectively protected the mice against tumor challenge (Fig. 2B).
Fig. 2.
NKG2D ligand H60 enhances the antitumor efficacy of survivin-based DNA vaccines in a murine CT-26 colon carcinoma model. (A) Endogenous expression (thick blue lines) in CT-26 cells of survivin (Left) or NKG2D ligand (Right). Thin red lines, staining controls. (B) Prophylactic setting. Vaccinated mice (n = 8) were challenged i.v. with 1 × 105 CT-26 colon carcinoma cells. (Upper) Representative lungs. (Lower) Average lung weight. Normal lung weight is ≈0.2 g. *, P < 0.00005, 0.001, 0.02, or 0.005 compared with PBS, pBud, pH60, or pSurv, respectively. Experiments were repeated 3 times with similar results. (C) In vivo depletion assays. Experiments were terminated 2 days earlier in CD8- and NK-depleted mice than in control and CD4-depleted animals. (D) Therapeutic setting. Mice were challenged i.v. with 1 × 105 CT-26 cells and later vaccinated. The results presented are the average of two separate experiments (n = 12 and 4).
In vivo depletions were performed to determine which cell population was responsible for this protection. Tumors developed so rapidly in CD8+ or NK cell depletion mice that the mice had to be killed 2 days earlier (Fig. 2C), suggesting that these effector cells are involved in the normal surveillance against CT-26 tumor cells. Moreover, these depletions abrogated most of the tumor protective effect induced by the pH60/Surv vaccine (Fig. 2C), indicating that the protection was mainly mediated by NK and CD8 T cells. In contrast, the depletion of CD4 cells from immune-competent mice protected against tumor cell challenge in both pBud and pH60/Surv groups (Fig. 2C), suggesting that CD4+ T cells contribute mainly to the negative regulation of antitumor immunity in our experimental system.
The DNA vaccines were also tested in therapeutic settings. Experimental pulmonary metastases were induced by i.v. injection of 1 × 105 CT-26 cells, which previously resulted in tumor colonization to lungs within 3 days (27). Mice were then vaccinated. Significantly, in each of two similar experiments, 75% of mice treated with the pH60/Surv vaccine survived (Fig. 2D), with most of them showing no or minimal tumor metastases (data not shown). In comparison, only 10% of mice survived in the empty pBud vector control group, all of them with >50% of their lung surface covered by fused tumor metastases (data not shown). The pSurv or pH60 vaccines failed to protect mice significantly.
Because survivin is overexpressed by essentially all solid tumor cells (11), the pH60/Surv vaccine should be effective against tumors of different origin. To test this contention, a breast cancer cell line, D2F2, was used for tumor cell challenge. D2F2 cells expressed survivin; they also expressed NKG2D ligands at an intermediate level (Fig. 3A). In the prophylactic setting (Fig. 3B), 12.5% of the mice were free of tumors in the pBud control group 30 days after tumor challenge, in agreement with a previous report on partial rejection of tumors expressing NKG2D ligands at intermediate levels (7). The number of tumor-free animals improved to 25% and 62.5% in pH60 or pSurv vaccinated groups of mice, respectively. Most importantly, only in the pH60/Surv group were 100% of mice completely free of tumors (Fig. 3B). In a therapeutic setting, a higher dose of D2F2 cells (3 × 105) was used to challenge mice and resulted in tumor development in all mice within 21 days in pBud group. In this case, vaccination with pH60/Surv induced the best therapeutic effect (Fig. 3C). Taken together, these data suggest that the pH60/Surv vaccine is optimally effective in both prophylactic and therapeutic settings against tumors of different origin, irrespective of their NKG2D ligand expression levels.
Fig. 3.
The pH60/Surv vaccine protects mice in a murine D2F2 breast carcinoma model. (A) Endogenous expression of survivin (Left) and NKG2D ligands (Center) by D2F2 cells (thick black lines), compared with NKG2D ligand positive control-Yac-1 cells (Right). Thin gray lines, staining controls. (B) Prophylactic setting. (C) Therapeutic setting. *, P < 0.01; *, P < 0.05 (compared with pBud); ***, P < 0.0002 and 0.05 (compared with pBud and pSurv, respectively).
The pH60/Surv Vaccine Activates both Innate and Adaptive Immune Responses. The known distribution and functions of NKG2D receptors (3, 4) and the results from NK and CD8 depletion experiments prompted us to directly assess whether coexpression of NKG2D ligand-H60 in the survivin-based DNA vaccine leads to enhanced NK- and CTL-mediated tumor cell killings. In fact, NK target cell killing was significantly enhanced in mice immunized with the pH60 vaccine and was further improved in pH60/Surv-vaccinated mice (Fig. 4A). Splenocytes isolated from pH60/Surv-vaccinated mice also exhibited the highest cytotoxicity against CT-26 target cells (Fig. 4B). This cytotoxicity was largely inhibited by Abs against CD8 or MHC class I molecules but not affected at all by anti-CD4 Ab (Fig. 4C), suggesting that the cytotoxicity detected was mediated mainly by MHC class I-restricted CD8+ T cells.
Fig. 4.
The pH60/Surv enhanced NK and CD8+ T cell activity. Freshly isolated or in vitro-stimulated splenocytes were tested in a standard 51Cr-release assay against Yac-1 (A) or CT-26 (B) target cells. ⋄, pBud group; ▪, pH60 group; ▵, pSurv group; •, pH60/Surv group. Experiments were repeated twice with similar results. (C) Cytotoxicity assays against CT-26 target cells in the absence or presence of blocking Abs. In vitro-stimulated splenocytes were originally isolated from pH60/Surv-vaccinated mice. •, no blocking Ab; shaded symbols, specific lysis in the presence of anti-CD4 (○), anti-CD8 (▵), or anti MHC class I (□) Abs. Experiments were repeated once with similar results.
To verify that the improved vaccine efficacy was indeed attributable to ligation of NKG2D receptor, another NKG2D ligand, RAE1, was also included in the vaccine, replacing H60. The expression of RAE was confirmed by flow cytometry (Fig. 5A). This pRAE1/Surv vaccine also proved superior in protecting mice from CT-26 challenge (Fig. 5B) and inducing NK (Fig. 5C) and CTL (Fig. 5D) responses. These data confirm that the combination vaccines enhance antitumor immunity by engaging the NKG2D receptor, suggesting a strong adjuvant effect of NKG2D ligands for the survivin-based DNA vaccine.
Fig. 5.
Another NKG2D ligand, RAE1, enhances the antitumor efficacy of survivin-based DNA vaccines. (A) Expression of RAE1 was demonstrated by flow cytometry using NKG2D tetramer. COS-7 cells were transfected with the pRAE (thick black line) or pBud control (thin gray line) vector. (B) In vivo CT-26 colon carcinoma model. Vaccinated mice (n = 8) were challenged i.v. with 1 × 105 CT-26 cells. *, P < 0.02 compared with pBud control group. Freshly isolated (C)or in vitro-stimulated (D) splenocytes were tested in a standard 51Cr-release assay against Yac-1 (C) or CT-26 (D), respectively. ⋄, pBud group; □, pRAE group; •, pRAE/Surv group.
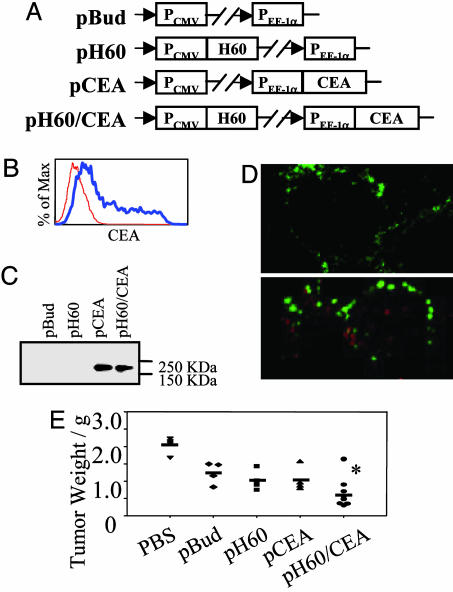
NKG2D Ligand H60 Enhances the Efficacy of CEA-Based DNA Vaccines. The difficulty in finding survivin-negative tumor cells prevented us from further demonstrating the specificity of the immune response. Consequently, we took advantage of our CEA-based DNA vaccine (16). We had already identified a specific HLA-A2-restricted CTL epitope in CEA-A2Kb double transgenic mice (10). Because H60 is not expressed in C57BL/6 mice (28), this approach was also selected to assess whether an allogeneic NKG2D ligand could enhance the efficacy of DNA-based vaccines and the feasibility for clinical applications. The corresponding DNA vaccine constructs are depicted in Fig. 6A, and the CEA expression of transfected 293T cells was confirmed by positive staining in both flow cytometric (Fig. 6B) and Western blot (Fig. 6C) analyses. Vaccine-induced CEA expression in vivo was further evaluated with cryostat sections of Peyer's patches from vaccinated mice. CEA expression was shown to localize to CD11c+ cells (Fig. 6D Lower), whereas control mice did not reveal any CEA expression in the Peyer's patches (Fig. 6D Upper).
Fig. 6.
NKG2D ligand H60 enhances the antitumor efficacy of CEA-based DNA vaccines. (A) Expression vectors were constructed based on the pBudCE4.1 backbone. (B) Flow cytometric analyses of 293T cells transfected with pCEA (thick blue line) or the empty pBud (thin red line) vector by using anti-CEA Ab. (C) Western blotting analyses of 293T cells transfected with different expression vectors by using anti-CEA Ab. (D) In vivo expression of CEA (red) in the Peyer's patches from pBud-vaccinated (Upper) or pCEA-vaccinated (Lower) mice. Each panel indicates two representative CD11c+ (green) dendritic cells. (E) Vaccinated CEA-A2Kb double transgenic mice were challenged s.c. with 6 × 105 MC-38-CEA-A2Kb cells and killed 28 days later. Tumor weights from each mouse are represented by individual symbols. Solid line, average tumor weights of each group. *, P < 0.0002 or 0.05 (compared with PBS or pBud control groups, respectively). Experiments were repeated three times with similar results.
Vaccine efficacy was further evaluated by a series of in vivo studies. CEA-A2Kb double transgenic mice were used because they are peripherally tolerant to CEA and express HLA-A2, enabling us to check vaccine efficacy and immune response against HLA-A2-restricted epitopes. Immunization with the pH60/CEA DNA vaccine effectively protected these transgenic mice from challenges of colon carcinoma cells MC-38 transfected with CEA and A2Kb (MC-38-CEA-A2Kb, Fig. 6E) but not from the parental MC-38 cells (data not shown). These findings demonstrate the specificity of the protection and suggest that the NKG2D ligand H60 also enhances the antitumor efficacy of CEA-based DNA vaccines.
Vaccination with pH60/CEA Enhances HLA-A2-Restricted, CEA691-Specific CTL Responses. As expected, splenocytes from pH60/CEA-vaccinated mice not only exhibited the highest killing against NK target cells (Fig. 7A) but also showed the most effective cytotoxicity against MC-38-CEA-A2Kb target cells (Fig. 7B) and not the parental MC-38 cells (Fig. 7C).
Fig. 7.
The pH60/CEA vaccine enhanced NK- and CEA-specific CTL responses. (A) Standard 51Cr release assays were performed with freshly isolated splenocytes against Yac-1 NK target cells. In vitro-stimulated splenocytes were tested in cytotoxicity assays against MC-38-CEA-A2Kb (B) or MC-38 (C) target cells. (A–C) ⋄, pBud group; ▪, pH60 group; ▵, pCEA group; •, pH60/CEA group. (D) Cytotoxicity of in vitro-stimulated splenocytes from pBud (open symbols) or pH60/CEA (closed symbols) groups against unloaded (▵) or CEA691-loaded (○) T2 target cells. (E) ELISPOT assays were performed with freshly isolated splenocytes either without (open bars) or in the presence of control peptide (kindly provided by J. Schlom of the National Institutes of Health; shaded bars) or CEA691 peptides (striped bars) at 10 μg/ml.
Next, we focused on the specific responses against one of the HLA-A2-restricted CTL epitopes, CEA691 (18). The pH60/CEA-vaccinated group displayed significant killing of CEA691-loaded HLA-A2+ T2 cells but not unloaded T2 cells (Fig. 7D). To further substantiate this finding, ELISPOT assays were performed with splenocytes freshly isolated from vaccinated CEA-A2Kb double transgenic mice. CEA691-specific, IFN-γ-secreting T cells were already present in pCEA-vaccinated mice; however, this reactivity was greatly enhanced in mice immunized with the pH60/CEA vaccine (Fig. 7E). Consequently, we demonstrated that the immune response induced by the pH60/CEA vaccine was still CEA-specific but had attained a higher level of efficacy.
Discussion
T cells targeting tumor-associated antigens are readily detectable in cancer patients and in those patients that received cancer vaccines, yet in most cases, such T cells fail to eradicate the tumor in vivo. Thus, it is clear that established tumors can induce immune tolerance through not well defined mechanisms (29). Consequently, we assumed that immunization strategies that employ different arms of the immune system should overcome such tolerance.
In our studies, DNA vaccines encoding tumor-associated antigens and the NKG2D ligands were transformed into double attenuated S. typhimurium (Dam-; AroA-), which target these genes to secondary lymphoid organs, i.e., Peyer's patches (19, 25). There, the bacteria can be phagocytosed by antigen-presenting cells such as DCs. In addition, Salmonella provide ligands for toll-like receptors or “danger signals,” which can induce DC activation and maturation. The genes encoded by vectors can be then transcribed, translated, expressed, and processed by DCs (30). Here, in fact, we demonstrate the in vivo expression of H60 or CEA encoded by our DNA vaccine in DCs located inside Peyer's patches, proving the efficiency of this delivery system.
DCs are known to support the tumoricidal activity of NK cells (31). In our vaccine setting, the interactions of NKG2D ligands expressed on DCs with the NKG2D receptor expressed on NK cells can directly activate these effector cells as reported in ref. 3. In addition cytokines secreted by activated DCs are likely to contribute to the robust NK activity detected in our pH60/Surv- or pH60/CEA-vaccinated mice.
DCs are the most proficient antigen-presenting cells (32) and have extraordinary capacity to activate naïve T cells (33). Once specific peptide/MHC complexes presented by DCs are recognized by the T cell receptor complex on CD8+ T cells, costimulatory signals provided by NKG2D–ligand interactions between these two populations will contribute to more effective T cell priming. This approach induced effective T cell responses, both in pH60/Surv-vaccinated syngeneic BALB/c mice and in pH60/CEA-vaccinated CEA-A2Kb double transgenic mice.
DCs, NK cells, and T cells are not independent in their functions but are subject to intense crosstalks. By secreting cytokines such as IFN-γ and TNF-α, activated NK cells can induce the maturation of DCs, which become stable and resistant to tumor-related suppressive factors, and show a strongly enhanced ability to induce Th1 and CTL responses (34). This interaction may contribute to the enhanced CTL killing observed in the pH60 (Figs. 4B and 7B) or pRAE (Fig. 5D) groups compared with control mice. CD8+ T cells are shown to induce maturation of DCs in the absence of CD4+ cells and CD40 ligation (35). By secreting IFN-γ, activated but not resting CD8+ T cells can induce differentiation of monocytes into DCs and restore the stimulatory capacity of IL-10-treated antigen-presenting cells (36). The secretion of IL-10 by tumor cells has been shown to be one of the mechanisms tumors use to escape immune surveillance (37). This CTL feedback can explain the enhanced NK killing of pSurv (Fig. 4A) or pCEA group (Fig. 7A) compared with the control group.
Besides the aforementioned, activated NK cells and CTLs can also enforce each other's activation directly. Activated NK cells boost the ongoing adaptive responses by producing IFN-γ, which promotes the Th1 polarization of antigen-specific T cells (38). Antigen-specific T cells can directly activate NK cells by secreting IL-2 (39), which can also provide signals required by NK cells in “helping” DCs (40). Thus, by delivering our vaccines with Salmonella, engaging NKG2D receptor, and providing tumor-associated antigens, our DNA vaccine can efficiently activate DCs, NK cells, and CTLs, presumably in Peyer's patches. The crosstalk between these different populations can reinforce their activation, giving the immune system the edge to overcome tumor-induced immune-suppression. These effects may be translated into the tumor protection induced by our vaccine in both prophylactic and therapeutic models.
Recently, regulatory T (Treg) cells, particularly CD4+CD25+ Treg cells, have reached the spotlight in tumor immunology. Depletion of CD4+CD25+ T cells in mice improves tumor clearance (41) and enhances the response to immune-based therapy (42). Tumor infiltrating Treg cells have been clinically associated with reduced survival (43). In our system, CD4+ T cells, presumably mediated by Treg cells, mainly contributed to the negative regulation of immune responses because their depletion led to better protection against tumor challenges. Treg cells, as all CD4+ T cells, do not express NKG2D receptor even after their activation (3). By engaging this receptor, our vaccine can achieve preferential activation of NK and CD8+ T cells and, thus, may tilt the balance toward immune surveillance and breakage of Treg-mediated peripheral tolerances to tumor-associated antigens. However, it is also possible that some CD4+ T cell subpopulations are required for optimal immune responses and generation of effective immune response and memory as reported in refs. 44 and 45.
We demonstrated here that our pH60/Surv vaccine effectively protected mice against two different tumors: CT-26 colon carcinoma cells, which express very low or negative levels of NKG2D ligands, and D2F2 breast carcinoma cells, which express NKG2D ligands at an intermediate level. However, the pH60/Surv vaccine worked in both models almost equally well, suggesting the potential utility of this vaccine for a variety of tumors, despite their varying NKG2D ligand expression levels. In addition, utilization of another NKG2D ligand, RAE1, in similar vaccines induced protection against tumor challenge, proving that the engagement of the NKG2D receptor is indeed responsible for the improved antitumor efficacy of such vaccines.
However, it is well known that NKG2D ligands display a great degree of polymorphism both in mice (46) and humans (47). Thus, to develop a widely applicable cancer vaccine, it should be determined whether an allogeneic NKG2D ligand can also serve as an adjuvant for DNA vaccines. Such an evaluation is facilitated by the fact that H60 is not expressed in C57BL/6 mice (28) and that a H-2Kb-restricted CTL response was reported to mediate graft-vs.-host disease (48). In this regard, in our pH60/CEA vaccine, H60 may provide an allogeneic stimulus as well as the stimulation and costimulation of NK and T cells. It is possible that the anti-H60 immune response is so dominant (49) that it overshadows the tumor-specific responses. However, our data indicate that the pH60/CEA vaccine successfully induced both potent NK- and CEA-specific CTL responses. These data suggest that allogeneic NKG2D ligands are also effective adjuvants and, thus, applicable to a variety of DNA-based cancer vaccines. Furthermore, the fact that such ligands enhance immune responses against a human tumor antigen like CEA suggests their potential for clinical applications. It is of course possible, given the NKG2D expression pattern and other differences between human and mouse (24), that the exact same strategy may not work with the same efficacy in humans. However, sufficient employment of both innate and adaptive arms of the immune system may still be a sensible way to overcome the hurdle created by the tumors.
In summary, we critically evaluated NKG2D ligands as adjuvants for DNA-based cancer vaccines. We demonstrated here that coexpression of NKG2D ligands in DNA-based cancer vaccines effectively enhances their antitumor efficacy by activating both innate and adaptive immune responses. Such DNA vaccines are well suited to eradicate residual tumor cells and to possibly boost an immune response against cancer relapse, thus making them relevant for future clinical applications. This contention is supported by the impressive antitumor effect achieved by our pH60/Surv DNA vaccine in a therapeutic setting featuring established metastases of murine breast and colon carcinoma.
Acknowledgments
We thank C. Dolman and D. Markowitz for excellent technical assistance and Kathy Cairns for editorial assistance with manuscript preparation. H.Z. is a fellow of the Susan G. Komen Breast Cancer Foundation. This study was supported by National Institutes of Health Grant CA83856 (to R.A.R.); Department of Defense Grants BC031079 (to R.A.R.), DAMD17-02-10562 (to R.X.), and DAMD17-02-1-0137 (to R.X.); and E. Merck, Darmstadt-Lexigen Research Center (Billerica, MA) Grant SFP1330 (to R.A.R.). This is The Scripps Research Institute's manuscript no. 16427-IMM.
Author contributions: H.Z., Y.L., J.-D.L., J.C.B., R.X., and R.A.R. designed research; H.Z., Y.L., J.-f.L., M.M., and N.M. performed research; F.J.P. contributed new reagents/analytic tools; H.Z., C.D.K., J.C.B., R.X., and R.A.R. analyzed data; and H.Z., C.D.K., J.C.B., R.X., and R.A.R. wrote the paper.
This paper was submitted directly (Track II) to the PNAS office.
Abbreviations: NK, natural killer; CEA, carcinoembryonic antigen; CTL, cytotoxic T cells; DC, dendritic cells; Treg, regulatory T.
References
- 1.Diefenbach, A. & Raulet, D. H. (2002) Immunol. Rev. 188, 9-21. [DOI] [PubMed] [Google Scholar]
- 2.Diefenbach, A., Jamieson, A. M., Liu, S. D., Shastri, N. & Raulet, D. H. (2000) Nat. Immunol. 1, 119-126. [DOI] [PubMed] [Google Scholar]
- 3.Jamieson, A. M., Diefenbach, A., McMahon, C. W., Xiong, N., Carlyle, J. R. & Raulet, D. H. (2002) Immunity 17, 19-29. [DOI] [PubMed] [Google Scholar]
- 4.Snyder, M. R., Weyand, C. M. & Goronzy, J. J. (2004) Trends Immunol. 25, 25-32. [DOI] [PubMed] [Google Scholar]
- 5.Cerwenka, A., Bakker, A. B., McClanahan, T., Wagner, J., Wu, J., Phillips, J. H. & Lanier, L. L. (2000) Immunity 12, 721-727. [DOI] [PubMed] [Google Scholar]
- 6.Carayannopoulos, L. N., Naidenko, O. V., Fremont, D. H. & Yokoyama, W. M. (2002) J. Immunol. 169, 4079-4083. [DOI] [PubMed] [Google Scholar]
- 7.Diefenbach, A., Jensen, E. R., Jamieson, A. M. & Raulet, D. H. (2001) Nature 413, 165-171. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Cerwenka, A., Baron, J. L. & Lanier, L. L. (2001) Proc. Natl. Acad. Sci. USA 98, 11521-11526. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Xiang, R., Mizutani, N., Luo, Y., Chiodoni, C., Zhou, H., Mizutani, M., Ba, Y., Becker, J. C. & Reisfeld, R. A. (2005) Cancer Res. 65, 553-561. [PubMed] [Google Scholar]
- 10.Zhou, H., Luo, Y., Mizutani, M., Mizutani, N., Becker, J. C., Primus, F. J., Xiang, R. & Reisfeld, R. A. (2004) J. Clin. Invest. 113, 1792-1798. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Altieri, D. C. (2003) Nat. Rev. Cancer 3, 46-54. [DOI] [PubMed] [Google Scholar]
- 12.Andersen, M. H., Pedersen, L. O., Becker, J. C. & Straten, P. T. (2001) Cancer Res. 61, 869-872. [PubMed] [Google Scholar]
- 13.Otto, K., Andersen, M. H., Eggert, A., Keikavoussi, P., Pedersen, L. O., Rath, J. C., Bock, M., Brocker, E. B., Straten, P. T., Kampgen, E., et al. (2005) Vaccine 23, 884-889. [DOI] [PubMed] [Google Scholar]
- 14.Shively, J. E. & Beatty, J. D. (1985) Crit. Rev. Oncol. Hematol. 2, 355-399. [DOI] [PubMed] [Google Scholar]
- 15.Thompson, J. A., Grunert, F. & Zimmermann, W. (1991) J. Clin. Lab. Anal. 5, 344-366. [DOI] [PubMed] [Google Scholar]
- 16.Xiang, R., Silletti, S., Lode, H. N., Dolman, C. S., Ruehlmann, J. M., Niethammer, A. G., Pertl, U., Gillies, S. D., Primus, F. J. & Reisfeld, R. A. (2001) Clin. Cancer Res. 7, Suppl., 856s-864s. [PubMed] [Google Scholar]
- 17.Huarte, E., Sarobe, P., Lu, J., Casares, N., Lasarte, J. J., Dotor, J., Ruiz, M., Prieto, J., Celis, E. & Borras-Cuesta, F. (2002) Clin. Cancer Res. 8, 2336-2344. [PubMed] [Google Scholar]
- 18.Kawashima, I., Hudson, S. J., Tsai, V., Southwood, S., Takesako, K., Appella, E., Sette, A. & Celis, E. (1998) Hum. Immunol. 59, 1-14. [DOI] [PubMed] [Google Scholar]
- 19.Luo, Y., Zhou, H., Mizutani, M., Mizutani, N., Reisfeld, R. A. & Xiang, R. (2003) Proc. Natl. Acad. Sci. USA 100, 8850-8855. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Clarke, P., Mann, J., Simpson, J. F., Rickard-Dickson, K. & Primus, F. J. (1998) Cancer Res. 58, 1469-1477. [PubMed] [Google Scholar]
- 21.BenMohamed, L., Krishnan, R., Longmate, J., Auge, C., Low, L., Primus, J. & Diamond, D. J. (2000) Hum. Immunol. 61, 764-779. [DOI] [PubMed] [Google Scholar]
- 22.Xiang, R., Lode, H. N., Dreier, T., Gillies, S. D. & Reisfeld, R. A. (1998) Cancer Res. 58, 3918-3925. [PubMed] [Google Scholar]
- 23.Groh, V., Bahram, S., Bauer, S., Herman, A., Beauchamp, M. & Spies, T. (1996) Proc. Natl. Acad. Sci. USA 93, 12445-12450. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Raulet, D. H. (2003) Nat. Rev. Immunol. 3, 781-790. [DOI] [PubMed] [Google Scholar]
- 25.Darji, A., Guzman, C. A., Gerstel, B., Wachholz, P., Timmis, K. N., Wehland, J., Chakraborty, T. & Weiss, S. (1997) Cell 91, 765-775. [DOI] [PubMed] [Google Scholar]
- 26.Xiang, R., Lode, H. N., Chao, T. H., Ruehlmann, J. M., Dolman, C. S., Rodriguez, F., Whitton, J. L., Overwijk, W. W., Restifo, N. P. & Reisfeld, R. A. (2000) Proc. Natl. Acad. Sci. USA 97, 5492-5497. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Xiang, R., Lode, H. N., Dolman, C. S., Dreier, T., Varki, N. M., Qian, X., Lo, K. M., Lan, Y., Super, M., Gillies, S. D., et al. (1997) Cancer Res. 57, 4948-4955. [PubMed] [Google Scholar]
- 28.Malarkannan, S., Shih, P. P., Eden, P. A., Horng, T., Zuberi, A. R., Christianson, G., Roopenian, D. & Shastri, N. (1998) J. Immunol. 161, 3501-3509. [PubMed] [Google Scholar]
- 29.Pardoll, D. (2003) Annu. Rev. Immunol. 21, 807-839. [DOI] [PubMed] [Google Scholar]
- 30.Schoen, C., Stritzker, J., Goebel, W. & Pilgrim, S. (2004) Int. J. Med. Microbiol. 294, 319-335. [DOI] [PubMed] [Google Scholar]
- 31.Fernandez, N. C., Lozier, A., Flament, C., Ricciardi-Castagnoli, P., Bellet, D., Suter, M., Perricaudet, M., Tursz, T., Maraskovsky, E. & Zitvogel, L. (1999) Nat. Med. 5, 405-411. [DOI] [PubMed] [Google Scholar]
- 32.Steinman, R. M. (1991) Annu. Rev. Immunol. 9, 271-296. [DOI] [PubMed] [Google Scholar]
- 33.Banchereau, J. & Steinman, R. M. (1998) Nature 392, 245-252. [DOI] [PubMed] [Google Scholar]
- 34.Kalinski, P., Giermasz, A., Nakamura, Y., Basse, P., Storkus, W. J., Kirkwood, J. M. & Mailliard, R. B. (2005) Mol. Immunol. 42, 535-539. [DOI] [PubMed] [Google Scholar]
- 35.Ruedl, C., Kopf, M. & Bachmann, M. F. (1999) J. Exp. Med. 189, 1875-1884. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Wirths, S., Reichert, J., Grunebach, F. & Brossart, P. (2002) Cancer Res. 62, 5065-5068. [PubMed] [Google Scholar]
- 37.Suzuki, T., Tahara, H., Narula, S., Moore, K. W., Robbins, P. D. & Lotze, M. T. (1995) J. Exp. Med. 182, 477-486. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Cooper, M. A., Fehniger, T. A., Fuchs, A., Colonna, M. & Caligiuri, M. A. (2004) Trends Immunol. 25, 47-52. [DOI] [PubMed] [Google Scholar]
- 39.Fehniger, T. A., Cooper, M. A., Nuovo, G. J., Cella, M., Facchetti, F., Colonna, M. & Caligiuri, M. A. (2003) Blood 101, 3052-3057. [DOI] [PubMed] [Google Scholar]
- 40.Mailliard, R. B., Son, Y. I., Redlinger, R., Coates, P. T., Giermasz, A., Morel, P. A., Storkus, W. J. & Kalinski, P. (2003) J. Immunol. 171, 2366-2373. [DOI] [PubMed] [Google Scholar]
- 41.Shimizu, J., Yamazaki, S. & Sakaguchi, S. (1999) J. Immunol. 163, 5211-5218. [PubMed] [Google Scholar]
- 42.Steitz, J., Bruck, J., Lenz, J., Knop, J. & Tuting, T. (2001) Cancer Res. 61, 8643-8646. [PubMed] [Google Scholar]
- 43.Curiel, T. J., Coukos, G., Zou, L., Alvarez, X., Cheng, P., Mottram, P., Evdemon-Hogan, M., Conejo-Garcia, J. R., Zhang, L., Burow, M., et al. (2004) Nat. Med. 10, 942-949. [DOI] [PubMed] [Google Scholar]
- 44.Janssen, E. M., Lemmens, E. E., Wolfe, T., Christen, U., von Herrath, M. G. & Schoenberger, S. P. (2003) Nature 421, 852-856. [DOI] [PubMed] [Google Scholar]
- 45.Sun, J. C. & Bevan, M. J. (2003) Science 300, 339-342. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Cerwenka, A., O'Callaghan, C. A., Hamerman, J. A., Yadav, R., Ajayi, W., Roopenian, D. C., Joyce, S. & Lanier, L. L. (2002) J. Immunol. 168, 3131-3134. [DOI] [PubMed] [Google Scholar]
- 47.Zwirner, N. W., Marcos, C. Y., Mirbaha, F., Zou, Y. & Stastny, P. (2000) Hum. Immunol. 61, 917-924. [DOI] [PubMed] [Google Scholar]
- 48.Choi, E. Y., Yoshimura, Y., Christianson, G. J., Sproule, T. J., Malarkannan, S., Shastri, N., Joyce, S. & Roopenian, D. C. (2001) J. Immunol. 166, 4370-4379. [DOI] [PubMed] [Google Scholar]
- 49.Choi, E. Y., Christianson, G. J., Yoshimura, Y., Sproule, T. J., Jung, N., Joyce, S. & Roopenian, D. C. (2002) Immunity 17, 593-603. [DOI] [PubMed] [Google Scholar]