Abstract
Background
Viral respiratory illnesses are the most common acute illnesses experienced and generally follow a predicted pattern over time. The severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) pandemic interrupted that pattern.
Methods
The HIVE (Household Influenza Vaccine Evaluation) study was established in 2010 to follow a cohort of Southeast Michigan households over time. Initially focused on influenza, surveillance was expanded to include other major respiratory pathogens, and, starting in 2015, the population was followed year round. Symptoms of acute illness were reported, and respiratory specimens were collected and tested to identify viral infections. Based on the known population being followed, virus-specific incidence was calculated.
Results
From 2015 to 2022, 1755 participants were followed in HIVE for 7785 person-years with 7833 illnesses documented. Before the pandemic, rhinovirus (RV) and common cold human coronaviruses (HCoVs) were the viruses most frequently identified, and incidence decreased with increasing age. Type A influenza was next but with comparable incidence by age. Parainfluenza and respiratory syncytial viruses were less frequent overall, followed by human metapneumoviruses. Incidence was highest in young children, but infections were frequently documented in all age groups. Seasonality followed patterns established decades ago. The SARS-CoV-2 pandemic disrupted these patterns, except for RV and, to a lesser extent, HCoVs. In the first 2 years of the pandemic, RV incidence far exceeded that of SARS-CoV-2.
Conclusions
Longitudinal cohort studies are important in comparing the incidence, seasonality, and characteristics of different respiratory viral infections. Studies documented the differential effect of the pandemic on the incidence of respiratory viruses in addition to SARS-CoV-2.
Keywords: respiratory infections, cohort studies, influenza, SARS-CoV-2, coronaviruses
The HIVE study began year-round surveillance of respiratory viruses in 2015. Prepandemic patterns of infection largely followed those recognized years ago. Rhinovirus incidence in the pandemic exceeded that of SARS-CoV-2. Except for common cold coronaviruses, infection with other viruses was infrequent.
Longitudinal cohort studies were used as an important tool for the study of acute respiratory infections even before the many infectious agents causing them were recognized [1]. The continuing need for cohort studies to answer many of the relevant questions was reemphasized in 2018 in the National Institutes of Health plan for improving the influenza vaccine [2]. That was a welcome reminder of the importance of those studies in defining not only the occurrence of influenza but also that of other respiratory viruses and their interaction. It has since resulted in the initiation or continuation of several longitudinal observational studies both in the United States and other parts of the world [3, 4].
The Household Influenza Virus Evaluation (HIVE) study at the University of Michigan began in the autumn of 2010 following the influenza pandemic of 2009. The original focus was mainly on influenza and the preventive effects of influenza vaccination [5]. The study gradually expanded to include other respiratory viruses, taking advantage of collected data and specimens. Thus, in addition to influenza transmission patterns and influenza vaccine effectiveness estimates, HIVE could provide insights into the other common respiratory viruses and their interaction [6]. Because of its original concentration on influenza, HIVE operated only during the colder months of the year in its first few years; a year-round collection of data and specimens did not begin until October 2015 [7]. The original approach was sufficient for studying influenza and most other significant respiratory viruses but limited the calculation of agent-specific year-round incidence, particularly of rhinoviruses (RVs) [8].
As we approached the completion of 10 years of study in 2020, the pandemic caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) began. Methods for specimen collection and pathogen identification were modified to allow the study to continue, making it possible to examine changes in infection occurrence during the pandemic against the background of previous virus activity [9]. Here we report on our observations, contrasting the 5-year prepandemic period with the first 2 years of the pandemic.
METHODS
Enrollment, Data Collection, and Response to the SARS-CoV-2 Restrictions
The methods used to study the HIVE cohort since follow-up began in 2010 have been published previously [7]. While they have remained largely consistent, some changes were made to improve surveillance and expand population eligibility before the start of the SARS-CoV-2 pandemic.
Throughout the study, households have been recruited from those in the Ann Arbor region receiving primary care from Michigan Medicine. From the 2010/2011 to 2015/2016 influenza seasons, eligible households included 4 or more participating members, at least 2 of whom were younger than 18 years. In subsequent years, newly recruited households contained 3 or more members including at least 1 child under 5 years (2016/2017) or 10 years starting in 2017/2018. In response to the SARS-CoV-2 pandemic, household eligibility was updated in 2020/2021 (to at least 3 household members—1 of whom was under 21 years) and 2021/2022 (to at least 3 household members—1 of whom was under 18 years). Additional changes in eligible household composition during the SARS-CoV-2 pandemic were made to support longitudinal retention, including permitting households to continue participating after their youngest member turned 18 years old and/or when only 2 household members—of any age—remained enrolled.
From the start of the study, each year in the spring or summer, households attended an enrollment or reengagement meeting. Adult household members provided informed consent for themselves and their children, and children 7 years or older also provided verbal assent before participating. Starting in 2016/2017, participating households consented for up to 5 years of participation (through 2021/2022) to increase retention between seasons. In summer 2021, all households were reconsented for participation for up to 7 years (through 2028/2029).
Since its inception, the study has been reviewed and approved by the institutional review board at the University of Michigan Medical School.
Acute Respiratory Illness Surveillance and Specimen Collection
Since 1 October 2014, the study has been operating year round. Starting with the 2015/2016 season, the surveillance year began on 1 July and ended on 30 June. Weekly email or telephone contacts were used to confirm the presence or absence of acute respiratory illness (ARI) in the household. Participants were instructed to report illnesses meeting a standard case definition: the onset of 2 or more ARI symptoms from anyone in the household, including cough, fever/feverishness, nasal congestion, chills, headache, body aches, and/or sore throat. Beginning in the 2014/2015 season, a separate set of age-specific symptom qualifiers were provided for participants under 3 years of age, including cough, fever/feverishness, nasal congestion, trouble breathing, fussiness, decreased appetite, and fatigue.
Until March 2020, participants with 2 or more of the specified symptoms were requested to visit the study clinic for their illness within 7 days of symptom onset. During this visit, the study team interviewed them and collected combined nasal and throat swabs (only nasal for children under age 3) for virus analysis. Study staff followed up with ill participants approximately 1 week after their illness visit to collect information about the course of the illness, including whether they contacted a health care provider regarding their illness.
After SARS-CoV-2 activity restrictions were put in place, participants with symptoms meeting the study case definition were instructed to self-collect their onset specimen for contactless home pickup by the study team. We had previously determined that self-collected specimens were comparable to those collected in the clinic [10]. Interviews to collect symptom data were carried out by telephone, and the subsequent follow-up was conducted as previously described.
Laboratory Testing
Respiratory specimens collected at illness visits were tested by reverse transcription polymerase chain reaction (RT-PCR) with a panel of respiratory viruses. For specimens collected before the 2016/2017 study year, testing was performed by singleplex RT-PCR using primers and probes developed by the Centers for Disease Control and Prevention (CDC) Division of Viral Diseases, Gastroenteritis, and Respiratory Viruses [7, 11]. Specimens collected from the 2016/2017 season onward were tested using the FTD Respiratory Pathogen 33 multiplex PCR kit (Fast Track Diagnostics) [12]. Specimens with positive results for RV and enterovirus on this panel were considered RV for the purposes of this analysis. RT-PCR for identification of SARS-CoV-2 was completed using primers and probes developed by the CDC (samples collected through October 2021) or the ThermoFisher TaqPath COVID-19/FluA/FluB Combo Kit (November 2021 through July 2022) [9].
Statistical Analysis
The frequencies and proportions of ARI and etiologic agents, along with study population and household sizes, were summarized by age category (<5, 6–11, 12–17, 18–49, and ≥50 years) for every month of the 2015/2016 through 2021/2022 influenza seasons, as well as by period. Periods were defined as prepandemic (2015/2016–2019/2020) and pandemic (2020/2021–2021/2022) to allow for assessment of virus transmission disruptions associated with 2020–2022 public health measures.
Illness reports meeting the specimen collection criteria with or without an associated specimen were eligible for inclusion in the analyses. Incidence rates (IRs) and their associated 95% confidence intervals (CIs) were calculated for ARIs and individual etiologic agents by period for each age category. IRs were calculated by totaling the number of ARIs and etiologic agents and dividing by overall and age-specific person-time contributed during the given period. Co-occurring agents were not treated as mutually exclusive. Statistical analyses were performed using R version 4.3.2 (R Foundation for Statistical Computing) and SAS software version 9.4 (SAS Institute).
Latent Class Methods for Analysis of Symptoms
Differentiating between largely overlapping symptom profiles is an ongoing challenge. For this reason, we performed a latent class analysis to generate and evaluate possible alternatives for syndromic profiles [13]. The latent class analysis included 7195 illness reports for which specimens were collected and tested. Eleven symptoms were included in class construction: fever, cough, fatigue, dyspnea, ear pain, wheezing, chills, sore throat, congestion, body aches, and headaches (Supplementary Table 1). The mean predicted probability of class membership was generally high and ranged from 84.7% (SD 13.7%) to 93.1% (SD 12.5%; Supplementary Table 2). The model that was selected contained 4 classes of illnesses based on general clusters and key combinations of symptoms (ie, symptoms were not exclusively assigned to a single class). All latent classes had a high frequency of cough. Specific symptoms of illnesses in each group are shown in Supplementary Table 1. Broadly, group 1 illnesses had frequent upper respiratory congestion (termed “upper respiratory congestion” for simplicity); group 2 illnesses had frequent fever and fatigue (“febrile malaise”); group 3 illnesses had frequent lower respiratory symptoms (eg, wheezing shortness of breath) with constitutional symptoms (“lower respiratory congestion”); and group 4 illnesses included aches and chills with nasal congestion (“congested myalgia”).
RESULTS
Population Under Study
The HIVE study began in 2010 following the influenza A(H1N1) pandemic and has continued since. Table 1 shows the number of households and individuals followed each year. Landmarks during this period include, in October 2014, the beginning of year-round surveillance, with the first full study year beginning in July 2015, and the start (in 2020) of the SARS-CoV-2 pandemic. Even during the decade preceding the pandemic, the size of the study population varied somewhat from year to year. The interquartile range (IQR) also changed somewhat, likely a result of changes in the eligibility of households. However, the age distribution remained relatively stable. Throughout, the percent female ranged from 49% to 52% and percent non-White from 5% to 13%. The advent of the pandemic resulted in changes not only in the way specimens were collected but also in school closures and the behavior of the population. These are described in detail elsewhere [9].
Table 1.
HIVE Cohort Structure Over 12 Influenza Seasons of Follow-up (2010–2022)
| Influenza Season | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Characteristic | 2010 2011 |
2011 2012 |
2012 2013 |
2013 2014 |
2014 2015 |
2015 2016 |
2016 2017 |
2017 2018 |
2018 2019 |
2019 2020 |
2020 2021 |
2021 2022 |
| Surveillance period | 1 Oct 30 Apr |
1 Dec 10 May |
1 Oct 13 May |
1 Oct 30 Apr |
1 Oct 30 Jun |
1 Jul 30 Jun |
1 Jul 30 Jun |
1 Jul 30 Jun |
1 Jul 30 Jun |
1 Jul 30 Jun |
1 Jul 31 Jul |
1 Aug 30 Jun |
| Households, n | 328 | 213 | 321 | 232 | 340 | 227 | 209 | 292 | 373 | 385 | 392 | 268 |
| Members, median (IQR) n | 4 (4–5) | 4 (4–5) | 4 (4–5) | 4 (4–5) | 4 (4–5) | 4 (4–5) | 4 (4–5) | 4 (3–5) | 4 (3–4) | 4 (3–4) | 4 (3–4) | 4 (3–4) |
| Participants, n | 1441 | 943 | 1426 | 1049 | 1431 | 996 | 894 | 1185 | 1488 | 1526 | 1565 | 994 |
| Age groups, y, n (%) | ||||||||||||
| <5 | 229 (15.9) | 108 (11.5) | 211 (14.8) | 119 (11.3) | 183 (12.8) | 97 (9.7) | 117 (13.1) | 218 (18.4) | 287 (19.3) | 263 (17.2) | 208 (13.3) | 130 (13.1) |
| 5–11 | 395 (27.4) | 290 (30.8) | 403 (28.3) | 316 (30.1) | 453 (31.7) | 327 (32.8) | 257 (28.7) | 298 (25.1) | 356 (23.9) | 386 (25.3) | 378 (24.2) | 238 (23.9) |
| 12–17 | 214 (14.9) | 151 (16.0) | 217 (15.2) | 180 (17.2) | 223 (15.6) | 173 (17.4) | 150 (16.8) | 166 (14.0) | 202 (13.6) | 201 (13.2) | 248 (15.8) | 144 (14.5) |
| 18–49 | 544 (37.8) | 351 (37.2) | 538 (37.7) | 383 (36.5) | 510 (35.6) | 351 (35.2) | 331 (37.0) | 452 (38.4) | 574 (38.6) | 594 (38.9) | 616 (39.4) | 391 (39.3) |
| 50+ | 59 (4.1) | 43 (4.6) | 57 (4.0) | 51 (4.9) | 62 (4.3) | 48 (4.8) | 39 (4.4) | 48 (4.1) | 69 (4.6) | 82 (5.4) | 115 (7.3) | 91 (9.2) |
Denominators are column totals at the start of each season's follow-up.
Abbreviation: IQR, interquartile range.
Findings from the initial 3 years of the study have been published along with research focused mainly on influenza [7, 14, 15]. The present analyses cover the seasons with year-round data collection, before and during the pandemic. Shown in Table 2 and Table 3 are the total number of specimens collected each period and those that tested positive for at least 1 virus. In addition to the distribution of detected viruses by age, results were calculated in terms of person-years of observation, accounting for variability in study population size in different age strata as well as variable times of observation.
Table 2.
Incidence Rates of Overall Illnesses and Respiratory Viral Agents Among HIVE Participants Over 5 Influenza Seasons of Follow-up (2015–2020)
| ARI Incidence in Hive Subjects by Age Group | ||||||
|---|---|---|---|---|---|---|
| Characteristic | Overall | <5 y | 5–11 y | 12–17 y | 18–49 y | 50+ y |
| Subjects, n | 2585 | 507 | 695 | 365 | 904 | 114 |
| Illnesses, n (%) | 6057 | 1465 (24.2) | 1614 (26.6) | 691 (11.4) | 2115 (34.9) | 172 (2.8) |
| Incidence (95% CI) per 100 py |
111.3 (108.5–114.1) | 178.2 (169.2–187.6) | 108.9 (103.7–114.4) | 82.5 (76.5–88.9) | 104.1 (99.7–108.7) | 63.6 (54.5–73.9) |
| Etiologic Agents | ||||||
| RV | ||||||
| Cases, n (%) | 2001 | 623 (31.1) | 508 (25.4) | 227 (11.3) | 588 (29.4) | 55 (2.7) |
| Incidence (95% CI) per 100 py | 36.8 (35.2–38.4) | 75.8 (70.0–82.0) | 34.3 (31.4–37.4) | 27.1 (23.7–30.9) | 28.9 (26.6–31.3) | 20.4 (15.3–26.5) |
| HCoVsa | ||||||
| Cases, n (%) | 650 | 169 (26.0) | 139 (21.4) | 65 (10.0) | 255 (39.2) | 22 (3.4) |
| Incidence (95% CI) per 100 py | 11.9 (11.0–12.9) | 20.6 (17.6–23.9) | 9.4 (7.9–11.1) | 7.8 (6.0–9.9) | 12.6 (11.1–14.2) | 8.1 (5.1–12.3) |
| IAV | ||||||
| Cases, n (%) | 397 | 74 (18.6) | 132 (33.2) | 51 (12.8) | 126 (31.7) | 14 (3.5) |
| Incidence (95% CI) per 100 py | 7.3 (6.6–8.0) | 9 (7.1–11.3) | 8.9 (7.5–10.6) | 6.1 (4.5–8.0) | 6.2 (5.2–7.4) | 5.2 (2.8–8.7) |
| IBV | ||||||
| Cases, n (%) | 152 | 24 (15.8) | 72 (47.4) | 23 (15.1) | 28 (18.4) | 5 (3.3) |
| Incidence (95% CI) per 100 py | 2.8 (2.4–3.3) | 2.9 (1.9–4.3) | 4.9 (3.8–6.1) | 2.7 (1.7–4.1) | 1.4 (.9–2.0) | 1.9 (.6–4.3) |
| RSV | ||||||
| Cases, n (%) | 239 | 104 (43.5) | 57 (23.8) | 20 (8.4) | 53 (22.2) | 5 (2.1) |
| Incidence (95% CI) per 100 py | 4.4 (3.9–5.0) | 12.7 (10.3–15.3) | 3.8 (2.9–5.0) | 2.4 (1.5–3.7) | 2.6 (2.0–3.4) | 1.9 (.6–4.3) |
| HMPV | ||||||
| Cases, n (%) | 153 | 53 (34.6) | 47 (30.7) | 14 (9.2) | 33 (21.6) | 6 (3.9) |
| Incidence (95% CI) per 100y | 2.8 (2.4–3.3) | 6.4 (4.8–8.4) | 3.2 (2.3–4.2) | 1.7 (.9–2.8) | 1.6 (1.1–2.3) | 2.2 (.8–4.8) |
| HPIVsa | ||||||
| Cases, n (%) | 284 | 116 (40.8) | 74 (26.1) | 18 (6.3) | 70 (24.6) | 6 (2.1) |
| Incidence (95% CI) per 100 py | 5.2 (4.6–5.9) | 14.1 (11.7–16.9) | 5.0 (3.9–6.3) | 2.1 (1.3–3.4) | 3.4 (2.7–4.4) | 2.2 (.8–4.8) |
| >1 Agentb | ||||||
| Cases, n (%) | 391 | 155 (39.6) | 109 (27.9) | 35 (9.0) | 85 (21.7) | 7 (1.8) |
| Incidence (95% CI) per 100 py | 7.2 (6.5–7.9) | 18.9 (16.0–22.1) | 7.4 (6.0–8.9) | 4.2 (2.9–5.8) | 4.2 (3.3–5.2) | 2.6 (1.0–5.3) |
Follow-up began upon enrollment or 1 July 2015 and ended at loss to follow-up, withdrawal, or 1 July 2020 (whichever was first).
Abbreviations: ARI, acute respiratory illness; CI, confidence interval; HCoVs, common cold human coronaviruses 229E, NL63, OC43, and HKU1; HMPV, human metapneumovirus; HPIVs, human parainfluenza viruses 1–4; IAV, influenza A virus; IBV, influenza B virus; py, person-years; RSV, respiratory syncytial virus; RV, rhinovirus.
aSamples were coded as HCoV positive or HPIV positive if any of the 4 subtypes of HCoV or HPIV serotypes were detected, respectively, regardless of the number of times they were detected.
bConstituent infecting agents were counted in agent-specific totals, as well.
Table 3.
Incidence Rates of Overall Illnesses and Respiratory Viral Agents Among HIVE Participants Over 2 Influenza Seasons of Follow-up (2020–2022)
| ARI Incidence in Hive Subjects by Age Group | ||||||
|---|---|---|---|---|---|---|
| Characteristic | Overall | <5 y | 5–11 y | 12–17 y | 18–49 y | 50+ y |
| Subjects, n | 1862 | 261 | 464 | 288 | 711 | 138 |
| Illnesses, n (%) | 1776 | 413 (23.3) | 458 (25.7) | 198 (11.2) | 605 (34.1) | 102 (5.7) |
| Incidence (95% CI) per 100 py |
75.8 (72.3–79.4) | 141.0 (127.7–155.3) | 79.7 (72.6–87.4) | 53.2 (46.1–61.2) | 66.3 (61.1–71.8) | 53.6 (43.7–65.1) |
| Etiologic Agent | ||||||
| RV | ||||||
| Cases, n (%) | 734 | 266 (36.2) | 204 (27.8) | 67 (9.1) | 175 (23.8) | 22 (3.0) |
| Incidence (95% CI) per 100 py | 31.3 (29.1–33.7) | 90.8 (80.2–102.4) | 35.5 (30.8–40.7) | 18.0 (14.0–22.9) | 19.3 (16.5–22.3) | 11.6 (7.2–17.5) |
| HCoVsa | ||||||
| Cases, n (%) | 195 | 38 (19.5) | 50 (25.6) | 18 (9.2) | 80 (41.0) | 9 (4.6) |
| Incidence (95% CI) per 100 py | 8.3 (7.2–9.6) | 13.0 (9.2–17.8) | 8.7 (6.5–11.5) | 4.8 (2.9–7.6) | 8.8 (6.9–10.9) | 4.7 (2.2–9.0) |
| IAV | ||||||
| Cases, n (%) | 30 | 8 (26.7) | 9 (30.0) | 3 (10.0) | 9 (30.0) | 1 (3.3) |
| Incidence (95% CI) per 100 py | 1.3 (.9–1.8) | 2.7 (1.2–5.4) | 1.6 (.7–3.0) | 0.8 (.2–2.4) | 1.0 (.5–1.9) | 0.5 (.0–2.9) |
| SARS-CoV-2 | ||||||
| Cases, n (%) | 232 | 22 (9.5) | 60 (25.9) | 31 (13.4) | 100 (43.1) | 19 (8.2) |
| Incidence (95% CI) per 100 py | 9.9 (8.7–11.3) | 7.5 (4.7–11.4) | 10.4 (8.0–13.4) | 8.3 (5.7–11.8) | 11.0 (8.9–13.3) | 10.0 (6.0–15.6) |
| RSV | ||||||
| Cases, n (%) | 28 | 13 (46.4) | 8 (28.6) | 4 (14.3) | 3 (10.7) | 0 (0.0) |
| Incidence (95% CI) per 100 py | 1.2 (.8–1.7) | 4.4 (2.4–7.6) | 1.4 (.6–2.7) | 1.1 (.3–2.8) | 0.3 (.1–1.0) | 0.0 (0.0–1.9) |
| HMPV | ||||||
| Cases, n (%) | 26 | 10 (38.5) | 7 (26.9) | 2 (7.7) | 7 (26.9) | 0 (0.0) |
| Incidence (95% CI) per 100 py | 1.1 (.7–1.6) | 3.4 (1.6–6.3) | 1.2 (.5–2.5) | 0.5 (.1–1.9) | 0.8 (.3–1.6) | 0.0 (0.0–1.9) |
| HPIVsa | ||||||
| Cases, n (%) | 53 | 17 (32.1) | 17 (32.1) | 6 (11.3) | 13 (24.5) | 0 (0.0) |
| Incidence (95% CI) per 100 py | 2.3 (1.7–3.0) | 5.8 (3.4–9.3) | 3.0 (1.7–4.7) | 1.6 (.6–3.5) | 1.4 (.8–2.4) | 0.0 (0.0–1.9) |
| >1 Agentb | ||||||
| Cases, n (%) | 97 | 30 (30.9) | 28 (28.9) | 9 (9.3) | 25 (25.8) | 5 (5.2) |
| Incidence (95% CI) per 100 py | 4.1 (3.4–5.1) | 10.2 (6.9–14.6) | 4.9 (3.2–7.0) | 2.4 (1.1–4.6) | 2.7 (1.8–4.0) | 2.6 (.9–6.1) |
Follow-up began upon enrollment or 1 July 2020 and ended at loss to follow-up, withdrawal, or 1 July 2022 (whichever was first).
Abbreviations: ARI, acute respiratory illness; CI, confidence interval; HCoVs, common cold coronaviruses 229E, NL63, OC43, and HKU1; HMPV, human metapneumovirus; HPIVs, human parainfluenza viruses 1–4; IAV, influenza A virus; py, person-years; RSV, respiratory syncytial virus; RV, rhinovirus; SARS-CoV-2, severe acute respiratory syndrome coronavirus 2.
aSamples were coded as HCoV positive or HPIV positive if any of the 4 subtypes of HCoV or HPIV serotypes were detected, respectively, regardless of the number of times they were detected.
bConstituent infecting agents were counted in agent-specific totals, as well.
More specimens were collected yearly in the prepandemic period than during the pandemic period even though the latter were collected at home instead of in the clinic. This mainly reflects the reduced occurrence of illnesses—particularly in the first year of the pandemic when nonpharmaceutical interventions were in extensive use and schools were closed, although other factors cannot be excluded. The more stable prepandemic period allows comparison of the frequencies of identification of all the respiratory viruses over a full 5 years of follow-up. RV was by far the most commonly identified agent. RV incidence decreased as age increased from 75.8 (95% CI, 70.0–82.0) infections per 100 person-years in the <5 year olds to 20.4 (95% CI, 15.3–26.5) in the ≥50 year olds except for a slight increase in younger adults—a previously noted pattern [16]. Interestingly, a somewhat similar pattern was seen with the second-most commonly identified group of viruses, the common cold human coronaviruses (HCoVs). These viruses are seasonal, infect children at a young age, and then reinfect the partially immune, similar to many other respiratory viruses [17]. In contrast, influenza virus A (IAV) incidence and, to a lesser extent, influenza virus B (IBV) incidence were flatter across the age groups. This was clear when calculated as incidence. Both respiratory syncytial virus (RSV) and human parainfluenza virus (HPIV) were by far the highest in incidence in the youngest age group. In addition, there were many symptomatic reinfections in adult age groups. Human metapneumovirus (HMPV) was similar but at a lower incidence. Coinfections were relatively frequent, especially at younger ages, at 18.9 (95% CI, 16.0–22.1) per 100 person-years in the youngest age group.
During the pandemic period (2020/2021–2021/2022), the most frequently identified virus overall and among each age group was RV at 31.3 (95% CI, 29.1–33.7) per 100 person-years. SARS-CoV-2 was identified somewhat more frequently than the HCoVs with relatively similar incidence across ages. However, they were much less frequently identified than RV. The other viruses were seen less frequently in these 2 years, and the numbers of identifications were too small for age-specific frequencies to be stable. No IBV was identified during the 2 years.
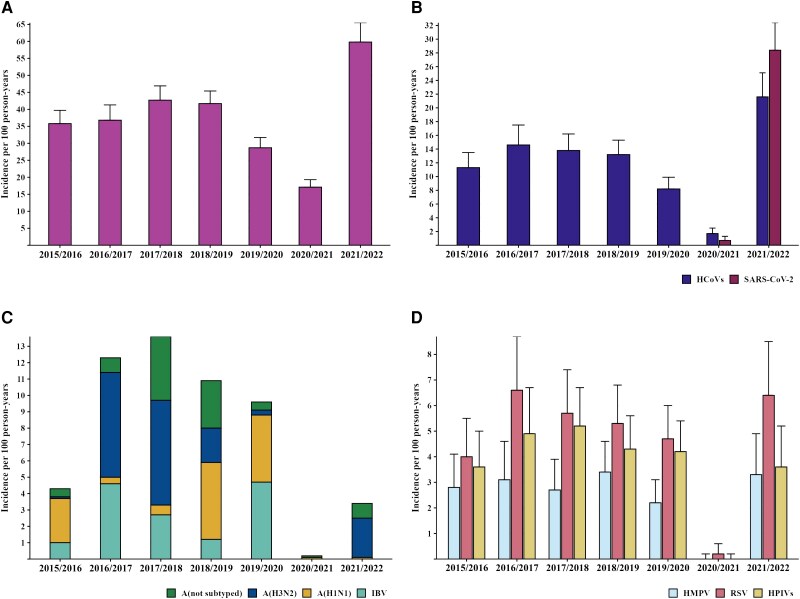
Year to Year and Seasonal Variation in Infection
The variation in the year-to-year incidence of virus identifications over the entire period can be seen in Figure 1A and 1B for more frequently detected viruses, and Figure 1C and 1D, using a different scale for less frequently detected viruses. The annual incidence of most respiratory illnesses showed relatively limited year-to-year variation before the pandemic. As an example, RV was demonstrably less frequent only in the 2019/2020 season, a year cut short by the start of the pandemic. Although at a lower frequency, a similar pattern was observed with the HCoVs. Influenza incidence was lowest in 2015/2016, a season in which A(H1N1) predominated; the vaccine used that year showed considerable effectiveness in this well-vaccinated population. RSV, HPIV, and HMPV (shown together in Figure 1D) displayed only minor variation in year-to-year incidence before the pandemic. During the first year of the pandemic, viruses in general were identified at low frequency with influenza viruses, RSV, HPIV, and HMPV almost absent. RV and HCoVs were at their highest frequency in the second year; there was a modest return of influenza and a greater return of the other viruses. During the first year of the pandemic, when nonpharmaceutical interventions were in place, SARS-CoV-2 (shown with the HCoVs in Figure 1B) was infrequently seen in the HIVE population. In the second year, it was still less frequent than RV and only slightly more frequent than HCoV.
Figure 1.
Incidence rates by season of selected viruses in the HIVE cohort between 2015 and 2022: (A) rhinovirus; (B) HCoVs and SARS-CoV-2; (C) IAV and IBV; and (D) RSV, HMPV, and HPIVs. Abbreviations: HCoVs, common cold human coronaviruses 229E, NL63, OC43, and HKU1; HMPV, human metapneumovirus; HPIVs, human parainfluenza viruses 1–4; IAV, influenza A virus; IBV, influenza B virus; RSV, respiratory syncytial virus; SARS-CoV-2, severe acute respiratory syndrome coronavirus 2. Error bars show the upper confidence limit of the incidence rate.
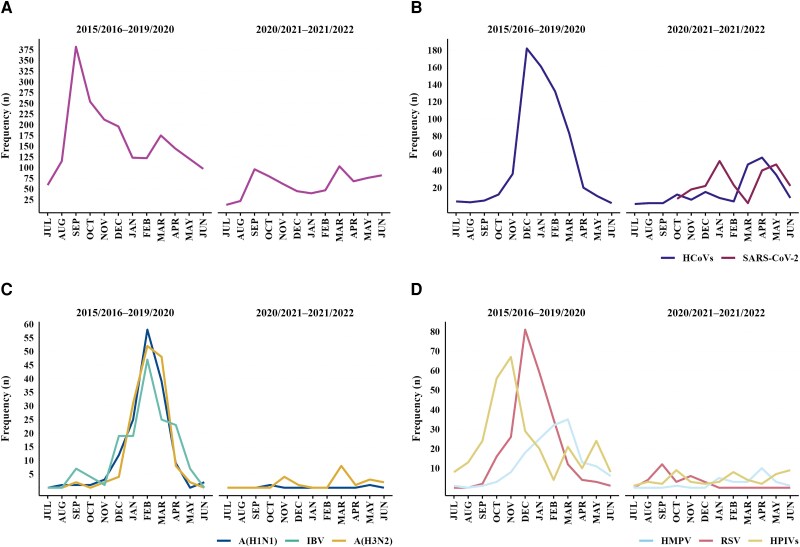
The actual months that the viruses were identified are shown in Figure 2. The figure shows the total number of detections of each virus per month over the 5 years preceding the pandemic and the 2 years of the pandemic. RV is known to be a year-round pathogen but particularly prevalent in the autumn when schools open, with sometimes a second smaller increase in the spring. That pattern was seen not only in the prepandemic period but also during the pandemic. In contrast, the HCoVs are sharply seasonal. In the prepandemic period, they were mostly seen between December and March but returned in later months during the pandemic. In the years preceding the pandemic, RSV peaked in December/January; HMPV peaked later in February/March; and all types of HPIV together did not show a clear seasonal pattern but peaked in October/November. These agents were not seen enough during the 2-year pandemic period to establish a pattern. The appearance of SARS-CoV-2 in the cohort was probably delayed by compliance with nonpharmaceutical interventions and showed little seasonality after that [9]. The periodicity of influenza outbreaks is well documented—sometimes peaking in the early winter and/or spring, with IBV usually delayed in appearance [18]. However, in the prepandemic period, both IAV and IBV peaked in the late winter. Epidemic curves for a more detailed examination in greater detail of the incidence of each virus from month to month throughout the prepandemic and pandemic periods are available in the Supplementary Material.
Figure 2.
Seasonality pre- and postpandemic of selected viruses in the HIVE cohort: (A) rhinovirus; (B) HCoVs and SARS-CoV-2; (C) IAV and IBV; and (D) RSV, HMPV, and HPIVs. Total detections per month have been summed in the prepandemic (2015–2020) and postpandemic (2020–2022) periods to compare cumulative seasonality. Abbreviations: HCoVs, common cold human coronaviruses 229E, NL63, OC43, and HKU1; HMPV, human metapneumovirus; HPIVs, human parainfluenza viruses 1–4; IAV, influenza A virus; IBV, influenza B virus; RSV, respiratory syncytial virus; SARS-CoV-2, severe acute respiratory syndrome coronavirus 2.
Illness Characteristics
There was no clear distinction in symptom profiles between various viruses (Supplementary Table 3), but patterns were more evident with latent class groupings (Table 4). Influenza illnesses were most often in the congested myalgia cluster (group 4) (42.4% of IAV illnesses and 45.0% of IBV illnesses). Congested myalgia was also common among SARS-CoV-2 illnesses (44.8%). Interestingly, while the remaining influenza illnesses were distributed among the other classes, SARS-CoV-2 had an equivalent share of illness in the upper respiratory congestion cluster (group 1), demonstrating the breadth of illness presentations as this virus continues to emerge. In contrast, RV, HCoVs, RSV, HMPV, and HPIVs were most frequently in the upper respiratory congestion cluster. Although often a driver of medical visits and cause for parental concern, lower respiratory congestion (group 3) was the smallest cluster overall among HIVE illnesses.
Table 4.
Participant Age and Viral Testing Results by Latent Class of Symptoms for Those With Any Viral Testing Result Without Coinfection (n = 6707)
| Characteristic | Upper Respiratory Congestion (n = 3058) | Febrile Malaise (n = 1059) | Lower Respiratory Congestion (n = 698) | Congested Myalgia (n = 1892) |
|---|---|---|---|---|
| Age, y | ||||
| <5 | 323 (21.3) | 983 (64.7) | 56 (3.7) | 158 (10.4) |
| 5–11 | 1000 (56.6) | 57 (3.2) | 141 (8.0) | 570 (32.2) |
| 12–17 | 445 (57.5) | 2 (0.3) | 95 (12.3) | 232 (30.0) |
| 18–49 | 1176 (49.0) | 16 (0.7) | 360 (15.0) | 848 (35.3) |
| ≥50 | 114 (46.5) | 1 (0.4) | 46 (18.8) | 84 (34.3) |
| IAV | 72 (24.4) | 34 (11.5) | 64 (21.7) | 125 (42.4) |
| H1N1a | 29 (24.4) | 13 (10.9) | 30 (25.2) | 47 (39.5) |
| H3N2a | 21 (16.0) | 16 (12.2) | 30 (22.9) | 64 (48.9) |
| IBV | 28 (23.3) | 9 (7.5) | 29 (24.2) | 54 (45.0) |
| SARS-CoV-2 | 90 (44.8) | 8 (4.0) | 13 (6.5) | 90 (44.8) |
| RSV | 79 (47.9) | 41 (24.9) | 22 (13.3) | 23 (13.9) |
| RV | 1216 (50.5) | 541 (22.5) | 208 (8.6) | 443 (18.4) |
| HMPV | 52 (38.8) | 26 (19.4) | 26 (19.4) | 30 (22.4) |
| HCoVs | 324 (52.9) | 86 (14.1) | 57 (9.3) | 145 (23.7) |
| HPIVs | 93 (38.6) | 60 (24.9) | 22 (9.1) | 66 (27.4) |
No. and row percentages are presented. Latent class analysis included the following symptoms: fever, cough, fatigue, dyspnea, ear pain, wheezing, chills, sore throat, congestion, body aches, and headaches. Groupings are based on general cluster trends and are not exclusive by symptom (see Supplement Table 1 for symptom profiles). Clusters are broadly defined as upper respiratory congestion (group 1); febrile malaise (group 2); lower respiratory congestion (group 3); and congested myalgia (group 4). All groupings were significantly different by virus per χ2 test.
Abbreviations: HCoVs, common cold human coronaviruses 229E, NL63, OC43, and HKU1; HMPV, human metapneumovirus; HPIVs, human parainfluenza viruses 1–4; IAV, influenza A virus; IBV, influenza B virus; RSV, respiratory syncytial virus; RV, rhinovirus; SARS-CoV-2, severe acute respiratory syndrome coronavirus 2.
aFor those for which IAV subtyping was available.
Most of the illnesses described in HIVE were mild, with only 13% of illnesses requiring a medical visit. IAV and IBV illnesses were most likely to lead to medical care (25.1% and 32.2%, respectively). This spectrum of illness is reflected in the latent class groupings: 19.8% of lower respiratory illnesses (group 3) required medical care, followed by 19.2% of febrile malaise illnesses (group 2), 17.6% of congested myalgia illnesses (group 4), and 8.2% of upper respiratory congestion illnesses (group 1). Gastrointestinal symptoms were reported during acute infection with all tested viruses. Influenza A(H3N2) illnesses had the most common reports of gastrointestinal symptoms (39.3%; 66/168), and RV illnesses had the fewest reports of gastrointestinal symptoms. Gastrointestinal symptoms were most commonly associated with group 4 illnesses (congested myalgia).
DISCUSSION
The SARS-CoV-2 pandemic is unique in many ways—not only in being caused by a coronavirus. It is the first pandemic that has been documented as being accompanied by disruption of the expected occurrence of other respiratory viruses. The influenza pandemic of 1968 occurred just after the time that many of these viruses were first identified and their behavior characterized. In that pandemic, and the pandemic of 2009, no perturbations of infection patterns and seasonality of other viruses were recognized. Indeed, in 1968/1969, extensive RSV transmission occurred at the same time as the first wave of A(H3N2) in early winter [19]. While there was some disruption in daily life associated with these pandemics, there were neither lockdowns nor deployment of nonpharmaceutical interventions as there were in the SARS-CoV-2 pandemic. It may be reasonable to attribute the extreme changes in infection patterns recently seen to these societal efforts to reduce SARS-CoV-2 transmission. However, the interruption of transmission of the other viruses was selective; RV returned very quickly and reestablished their usual seasonal pattern while SARS-CoV-2 was still transmitting extensively. This was somewhat surprising, based on the generally accepted concept that they are mainly transmitted by large droplets [20]. Less surprising was the return of the HCoVs, given, at least, their virologic relation to SARS-CoV-2. The other viruses have now returned, albeit much more slowly.
The overall seasonal patterns of the respiratory viruses seen prepandemic are remarkably similar to those described many decades ago, despite societal changes that have occurred, particularly those involving childcare. RV is prominent year round but peaks around September when schools open [20, 21]. The second-most identified viral group, the HCoVs, was almost completely limited to midwinter and early spring [11]. HPIV followed RV in the seasonal progression in the late autumn [22]. RSV mainly appeared afterward, with variation in exact time from year to year [23]. The less-frequently identified HMPV followed. Influenza, in this study, peaked in late winter, but it is recognized that it may move somewhat from year to year, with IBV usually following IAV subtypes [18]. What drives these patterns has been discussed extensively in terms of climate and human behavior [24].
Relative infection frequency of different viruses by age is often given as a percentage distribution of identifications. It is done largely because there is often no firm knowledge of the population involved. This is particularly true of hospital-based studies, which are the ones commonly performed. Only in a cohort study such as HIVE, with a known population, is it possible to derive the actual incidence of all symptomatic infections, whether causing medical consultation or not. In contrast to the percent distributions, incidence gives a much clearer picture of the effect of increasing age. It is also possible to show that the occurrence of viruses such as RSV—which was often thought of as a pathogen in young children and now in older individuals—is common to persons of all ages.
The latent class analyses were carried out to look for variation in symptoms as they occur in the population, which would not be appreciated by examination of frequency. It was possible to show that differences do exist even when controlling for age, such as influenza having different class distributions as compared to RV and HCoVs. There were also differences in the occurrence of gastrointestinal symptoms among viral infections. However, there is much overlap. This emphasizes the need for diagnostic testing for specific pathogens as we begin to have more antivirals available for specific therapy.
In addition to the results described in this report are HIVE-based studies of vaccine effectiveness, virus transmission, and correlates of protection based on secretory and cellular immunity [15, 25, 26]. The study now focuses on maintaining the same households on report for 5 or more years. This will increase the current ability to determine the duration of immunity and waning of protection. A great advantage of HIVE's approach is the ability to examine all the major respiratory viruses simultaneously using the same methods. These methods have allowed recognition of the continued major contribution of RV to illnesses even during the pandemic. A clear goal currently will be to document how and when the pattern of infection with all the major respiratory viruses fully returns and the place of SARS-CoV-2 infection in seasonal patterns. Only in a study with a broad approach and known population like HIVE's will that be truly possible.
Supplementary Data
Supplementary materials are available at The Journal of Infectious Diseases online (http://jid.oxfordjournals.org/). Supplementary materials consist of data provided by the author that are published to benefit the reader. The posted materials are not copyedited. The contents of all supplementary data are the sole responsibility of the authors. Questions or messages regarding errors should be addressed to the author.
Supplementary Material
Contributor Information
Arnold S Monto, Department of Epidemiology, University of Michigan School of Public Health, Ann Arbor, Michigan, USA.
Joshua E Foster-Tucker, Department of Epidemiology, University of Michigan School of Public Health, Ann Arbor, Michigan, USA.
Amy P Callear, Department of Epidemiology, University of Michigan School of Public Health, Ann Arbor, Michigan, USA.
Aleda M Leis, Department of Epidemiology, University of Michigan School of Public Health, Ann Arbor, Michigan, USA.
Elie-Tino Godonou, Department of Epidemiology, University of Michigan School of Public Health, Ann Arbor, Michigan, USA.
Matthew Smith, Department of Epidemiology, University of Michigan School of Public Health, Ann Arbor, Michigan, USA.
Rachel Truscon, Department of Epidemiology, University of Michigan School of Public Health, Ann Arbor, Michigan, USA.
Emileigh Johnson, Department of Epidemiology, University of Michigan School of Public Health, Ann Arbor, Michigan, USA.
Lara J Thomas, Department of Epidemiology, University of Michigan School of Public Health, Ann Arbor, Michigan, USA.
Mark S Thompson, Westat, Rockville, Maryland, USA.
Alicia M Fry, Influenza Division, National Center for Immunization and Respiratory Diseases, Centers for Disease Control and Prevention, Atlanta, Georgia, USA.
Brendan Flannery, Influenza Division, National Center for Immunization and Respiratory Diseases, Centers for Disease Control and Prevention, Atlanta, Georgia, USA.
Ryan E Malosh, Division of Immunizations, Michigan Department of Health and Human Services, Lansing, Michigan, USA.
Joshua G Petrie, Marshfield Clinic Research Institute, Marshfield, Wisconsin, USA.
Adam S Lauring, Department of Internal Medicine, University of Michigan Medical School, Ann Arbor, Michigan, USA.
Emily T Martin, Department of Epidemiology, University of Michigan School of Public Health, Ann Arbor, Michigan, USA.
Note
Financial support. This work was supported by the Centers for Disease Control and Prevention (grant numbers U01IP001034, R01AI097150, R56AI097150, and U01IP000474); and the National Institute of Health, National Institute of Allergy and Infectious Diseases (grant number 75N93021C00015).
References
- 1. Dingle JH, Badger GF, Feller AE, Hodges RG, Jordan WS, Rammelkamp CH. A study of illness in a group of Cleveland families. I. Plan of study and certain general observations. Am J Hyg 1953; 58:16–30. [DOI] [PubMed] [Google Scholar]
- 2. Erbelding EJ, Post DJ, Stemmy EJ, et al. A universal influenza vaccine: the strategic plan for the National Institute of Allergy and Infectious Diseases. J Infect Dis 2018; 218:347–54. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Huang QS, Bandaranayake D, Wood T, et al. Risk factors and attack rates of seasonal influenza infection: results of the Southern Hemisphere Influenza and Vaccine Effectiveness Research and Surveillance (SHIVERS) seroepidemiologic cohort study. J Infect Dis 2019; 219:347–57. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Heimonen J, Chow EJ, Wang Y, et al. Risk of subsequent respiratory virus detection after primary virus detection in a community household study—King County, Washington, 2019–2021. J Infect Dis 2024; 229:422–31. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Ohmit SE, Petrie JG, Malosh RE, et al. Influenza vaccine effectiveness in the community and the household. Clin Infect Dis 2013; 56:1363–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Monto AS, Malosh RE, Evans R, et al. Data resource profile: household influenza vaccine evaluation (HIVE) study. Int J Epidemiol 2019; 48:1040–1040g. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Monto AS, Malosh RE, Petrie JG, Thompson MG, Ohmit SE. Frequency of acute respiratory illnesses and circulation of respiratory viruses in households with children over 3 surveillance seasons. J Infect Dis 2014; 210:1792–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Monto AS, Bryan ER, Ohmit S. Rhinovirus infections in Tecumseh, Michigan: frequency of illness and number of serotypes. J Infect Dis 1987; 156:43–9. [DOI] [PubMed] [Google Scholar]
- 9. Fine SR, Bazzi LA, Callear AP, et al. Respiratory virus circulation during the first year of the COVID-19 pandemic in the household influenza vaccine evaluation (HIVE) cohort. Influenza Other Respir Viruses 2023; 17:e13106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Malosh RE, Petrie JG, Callear AP, Monto AS, Martin ET. Home collection of nasal swabs for detection of influenza in the household influenza vaccine evaluation study. Influenza Other Respir Viruses 2021; 15:227–34. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Monto AS, DeJonge PM, Callear AP, et al. Coronavirus occurrence and transmission over 8 years in the HIVE cohort of households in Michigan. J Infect Dis 2020; 222:9–16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Sakthivel SK, Whitaker B, Lu X, et al. Comparison of fast-track diagnostics respiratory pathogens multiplex real-time RT-PCR assay with in-house singleplex assays for comprehensive detection of human respiratory viruses. J Virol Methods 2012; 185:259–66. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Weller BE, Bowen NK, Faubert SJ. Latent class analysis: a guide to best practice. J Black Psychol 2020; 46:287–311. [Google Scholar]
- 14. Ohmit SE, Petrie JG, Malosh RE, Fry AM, Thompson MG, Monto AS. Influenza vaccine effectiveness in households with children during the 2012–2013 season: assessments of prior vaccination and serologic susceptibility. J Infect Dis 2015; 211:1519–28. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Malosh RE, Petrie JG, Callear A, et al. Effectiveness of influenza vaccines in the HIVE household cohort over 8 years: is there evidence of indirect protection? Clin Infect Dis 2021; 73:1248–56. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Monto AS, Ullman BM. Acute respiratory illness in an American community. The Tecumseh study. JAMA 1974; 227:164–9. [PubMed] [Google Scholar]
- 17. Petrie JG, Bazzi LA, McDermott AB, et al. Coronavirus occurrence in the household influenza vaccine evaluation (HIVE) cohort of Michigan households: reinfection frequency and serologic responses to seasonal and severe acute respiratory syndrome coronaviruses. J Infect Dis 2021; 224:49–59. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Malosh RE, McGovern I, Monto AS. Influenza during the 2010–2020 decade in the United States: seasonal outbreaks and vaccine interventions. Clin Infect Dis 2023; 76:540–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Monto AS, Davenport FM, Napier JA, Francis T. Modification of an outbreak of influenza in Tecumseh, Michigan by vaccination of schoolchildren. J Infect Dis 1970; 122:16–25. [DOI] [PubMed] [Google Scholar]
- 20. Jennings LC, Dick EC. Transmission and control of rhinovirus colds. Eur J Epidemiol 1987; 3:327–35. [DOI] [PubMed] [Google Scholar]
- 21. Fox JP, Cooney MK, Hall CE. The Seattle virus watch. V. Epidemiologic observations of rhinovirus infections, 1965–1969, in families with young children. Am J Epidemiol 1975; 101:122–43. [DOI] [PubMed] [Google Scholar]
- 22. Fry AM, Curns AT, Harbour K, Hutwagner L, Holman RC, Anderson LJ. Seasonal trends of human parainfluenza viral infections: United States, 1990–2004. Clin Infect Dis 2006; 43:1016–22. [DOI] [PubMed] [Google Scholar]
- 23. Kim HW, Arrobio JO, Brandt CD, et al. Epidemiology of respiratory syncytial virus infection in Washington, D.C. I. Importance of the virus in different respiratory tract disease syndromes and temporal distribution of infection. Am J Epidemiol 1973; 98:216–25. [DOI] [PubMed] [Google Scholar]
- 24. Lowen AC, Mubareka S, Steel J, Palese P. Influenza virus transmission is dependent on relative humidity and temperature. PLoS Pathog 2007; 3:1470–6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Petrie JG, Eisenberg MC, Ng S, et al. Application of an individual-based transmission hazard model for estimation of influenza vaccine effectiveness in a household cohort. Am J Epidemiol 2017; 186:1380–8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Petrie JG, Parkhouse K, Ohmit SE, Malosh RE, Monto AS, Hensley SE. Antibodies against the current influenza A(H1N1) vaccine strain do not protect some individuals from infection with contemporary circulating influenza A(H1N1) virus strains. J Infect Dis 2016; 214:1947–51. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.