Abstract
Impaired glucose uptake in the brain is an early presymptomatic manifestation of Alzheimer’s disease (AD), with symptom-free periods of varying duration that likely reflect individual differences in metabolic resilience. We propose a systemic “bioenergetic capacity”, the individual ability to maintain energy homeostasis under pathological conditions. Using fasting serum acylcarnitine profiles from the AD Neuroimaging Initiative as a blood-based readout for this capacity, we identified subgroups with distinct clinical and biomarker presentations of AD. Our data suggests that improving beta-oxidation efficiency can decelerate bioenergetic aging and disease progression. The estimated treatment effects of targeting the bioenergetic capacity were comparable to those of recently approved anti-amyloid therapies, particularly in individuals with specific mitochondrial genotypes linked to succinylcarnitine metabolism. Taken together, our findings provide evidence that therapeutically enhancing bioenergetic health may reduce the risk of symptomatic AD. Furthermore, monitoring the bioenergetic capacity via blood acylcarnitine measurements can be achieved using existing clinical assays.
Subject terms: Alzheimer's disease, Metabolomics
Bioenergetic capacity, the ability to maintain energy balance under stress, can be monitored using blood acylcarnitine profiles. Improving this capacity may slow Alzheimer’s disease progression.
Introduction
Dysregulation of bioenergetic pathways is a central feature of Alzheimer’s disease (AD), with detectable abnormalities occurring in the brain years prior to the symptomatic onset of AD dementia1. For example, studies investigating brain glucose uptake have consistently identified glucose hypometabolism as a presymptomatic metabolic manifestation of AD2–4. Additional evidence for early energetic dysregulation in AD comes from epidemiological studies, which have linked metabolic diseases, such as type 2 diabetes (T2D), and cardiovascular disease to a significantly increased risk of developing AD later in life5–9. The delay between the onset of symptomatic disease and apparent aberrations in energy pathways suggests the existence of a “bioenergetic capacity”, which can provide a temporary reserve that provides resilience from pathological symptoms of the disease.
Mitochondria are the essential cellular units of energy metabolism and are thus central to our proposed framework of energy-related resilience. They have been actively studied as potential targets for therapeutic intervention in AD10,11. Cellular energy supply through mitochondrial metabolism is fueled by three main routes that ultimately feed into the tricarboxylic acid (TCA) cycle: (1) glucose catabolism to pyruvate, (2) beta-oxidation of fatty acids to acetyl-CoA, and (3) the degradation of proteins into glucogenic and ketogenic amino acids (Fig. 1a). These three routes are tightly controlled through a complex system of receptors and intracellular signaling that respond to the current metabolic state at both the organismal and tissue level12. In our study, we reduce the complexity of mitochondrial energetics by focusing on metabolic states under overnight fasting conditions. As a result, within the triad of energy pathways, dietary glucose metabolism assumes a marginal role (similar to the above-mentioned state of reduced glucose uptake), while the metabolic routes of fatty acids and proteins become predominant.
Fig. 1. Concept of individual bioenergetic capacity mirroring impaired energy metabolism in the brain.
a The three main sources of mitochondrial energy metabolism: glucose, fatty acids, and proteins/amino acids, all of which ultimately feed into the TCA cycle. Common genetic variants in mitochondrial transporters and enzymes are assumed to define the inherited bioenergetic potential of each individual. Our study focuses on fasting individuals, largely removing the effect of dietary glucose and focusing on the fatty acid and protein routes. b Chain length-specific role of acylcarnitines as readouts for the bioenergetic capacity through the functionality, activity, and efficiency of mitochondrial energy metabolism; and examples of previously reported acylcarnitine level changes for AD-related phenotypes. c Integrated concept of bioenergetic capacity as the age-specific result of inherited bioenergetic potential and acquired modifiable metabolic functionality. Hypothetical trajectories for high and low inherited bioenergetic potential are shown, where deviations from the average are determined by modifiable lifestyle factors, such as physical activity, diet, health status, and other factors. Deviations from the overall population average are assumed to confer vulnerability or resilience to AD-related pathology and cognitive decline. AD Alzheimer’s disease, FAs fatty acids, AAs amino acids, CSF cerebrospinal fluid, TCA tricarboxylic acid, β-Ox. beta-oxidation, PDH Pyruvate dehydrogenase complex.
Acylcarnitines are a group of molecules whose blood levels provide accurate readouts specifically of the fatty acid and protein aspects of mitochondrial metabolism (Fig. 1b)13. They have long been used for the diagnosis of inborn errors of energy metabolism (IEEMs) through newborn screenings13,14. As a milder version of these deficiencies, genome-wide association studies (GWAS) have identified less penetrant single nucleotide polymorphisms (SNPs) that map to the same genes as in the IEEMs and show similar but weaker effects on blood acylcarnitine levels15–18. Acylcarnitine levels therefore serve as sensitive indicators of genetic influences on mitochondrial pathways. Beyond genetic variation, various acquired conditions influence mitochondrial pathways and are reflected in blood acylcarnitine levels. For example, increased levels of intermediate acylcarnitines from fatty acid and amino acid metabolism have been reported both in patients with T2D19,20 and in obese individuals with ketogenic branched-chain amino acid overload21. This suggests that the bioenergetic state of acylcarnitine levels might contain a modifiable component with consequences on disease risk. Interestingly, even during healthy aging, blood acylcarnitine levels reflect the age-dependent decrease in mitochondrial energetic capacity in both β-oxidation and TCA cycle pathways22–24. It has furthermore been proposed that blood acylcarnitine levels may be informative about the fatty acid oxidation status within the dorsolateral prefrontal cortex25. Alterations to acylcarnitine levels have also been described in the context of AD, with various studies discussing the biomarker potential of this metabolite class26–32.
Based on this combined evidence, we hypothesize that acylcarnitine profile in fasting blood provides a proxy of an individual’s bioenergy capacity and thus their resilience buffer. We assume that deviations in this bioenergetic capacity from the normal population average significantly modulate the risk for disease outcomes, including neurodegenerative diseases like AD (Fig. 1c). In this study we show that acylcarnitine profiles can be used to: (1) categorize individuals by resilience status along their bioenergetic capacity. (2) Disentangle modifiable and genetic contributions to this resilience. (3) Integrate these profiles with genetic variation into a prognostic instrument that is predictive of future cognitive trajectories.
Results
Bioenergetic subgroups in AD and their determining factors
We performed subgroup Identification (SGI)33 on 1531 Alzheimer’s Disease Neuroimaging Initiative (ADNI) participants using fasting serum profiles of 23 acylcarnitines measured through targeted metabolomics as representative readouts of individual bioenergetic capacity. Study characteristics and acylcarnitine descriptions can be found in Supplementary Data 1 and 2, respectively. Acylcarnitine levels were corrected for significant medication effects and ADNI study phase prior to clustering, but otherwise deliberately left uncorrected for any other potential confounders. SGI revealed a series of associations for cluster splits in the hierarchical tree relating to demographic variables, clinical diagnosis, and A-T-(N)-(C) measures34 (Fig. 2a).
Fig. 2. Acylcarnitine profiles stratify participants from the ADNI study in groups of different AD pathology.
a Acylcarnitine-based hierarchical clustering, with informative branches highlighted with solid-colored lines. Solid gray lines indicate cluster pairs that showed no significant associations. Clinical and demographic parameters at split points indicate significant differences between the individuals in the left and right subclusters (identified by number labels in the dendrogram) below that respective point. b–d Individuals in cluster 2 have lower CSF p-tau levels, higher brain glucose uptake assessed by FDG-PET, and better cognitive function measured by the ADAS-Cog. 13 subscale compared to cluster 3. e, f Further down on the left-hand side of the tree within favorable cluster 2, cluster 6 contains younger individuals with lower (worse) CSF Aβ1-42 compared to cluster 7. g–i On the right-hand-side of the tree, within cluster 3, cluster 8 contains a higher number of CSF amyloid-positive (indicated in the legend by “+”) individuals with clinical AD and a higher proportion of females compared to cluster 9. j We investigated factors impacting subgroup division by examining both non-modifiable (acylcarnitine-related SNPs) and modifiable (adjusted acylcarnitine levels) factors. The results reveal a substantial amount (40–60%) of variance explained by genetics-corrected acylcarnitine levels, primarily medium- and long-chain, with overall rather minor contributions from genetic factors. The epistatic interaction between rs17806888 and rs924135, which explained ~32% of the variance between clusters 2 and 3, is a notable exception. Both variants have been reported to significantly influence succinylcarnitine, highlighting a genetic link to the TCA cycle and amino acid-based energy metabolism. Variables marked with * have been centered to zero mean and scaled to unit variance. Abbreviations: CSF cerebrospinal fluid, FDG-PET Fluorodeoxyglucose-Positron Emission Tomography, ADAS-Cog. 13 Alzheimer’s Disease Assessment Scale—Cognitive Subscale 13, BMI body mass index. Box plots display the median (central line), interquartile range (box bounds), whiskers extending to the smallest and largest values within 1.5 times the interquartile range from the quartiles, and notches indicating the 95% confidence interval for the median. Details for each statistical test and the corresponding sample sizes are provided in Supplementary Data 3. All tests were two-sided and raw p values were reported. Source data are provided as a Source Data file.
The analysis split ADNI participants into two clusters, 2 and 3, at the top of the tree, with significantly different levels of p-tau in cerebrospinal fluid (CSF), glucose uptake measured by [18F] fluorodeoxyglucose-positron emission tomography (FDG-PET), and the 13-item AD assessment scale-cognitive subscale (ADAS-Cog. 13; Fig. 2b–d, Supplementary Data 3). Cluster 2 overall contained less affected individuals with significantly better cognitive function, higher brain glucose uptake, and lower CSF p-tau levels. This group was further divided into two distinct clusters: cluster 7, which on average was healthier with higher CSF Aβ1-42 levels and higher age, and cluster 6, a younger group showing signs of early pathological aging (Fig. 2e, f). On the other side of the tree, within cluster 3—which exhibited more progressed levels of AD biomarkers—there were further distinctions in clusters 8 and 9, which showed significant differences in the distribution of body mass index (BMI), sex, and diagnostic groups. (Fig. 2g–i). To rule out that these associations were driven by confounding factors, we repeated the analysis while adjusting for sex, age, BMI, copies of APOE ε4, and years of education. All associations remained significant (Supplementary Data 4).
To investigate the factors potentially influencing subgroup division, we examined the variance explained by non-modifiable and modifiable factors for each branching point (BP) in the tree. Specifically, we examined acylcarnitine-related SNPs reported by Shin et al.16 as non-modifiable factors (SNP list is provided in Supplementary Data 5, replication in ADNI in Supplementary Data 6) and modifiable acylcarnitine levels (i.e., residuals adjusted for genetic and other non-modifiable factors) to better understand their impact on subgroup separation. Overall, sample clustering was mainly determined by the modifiable, covariate-adjusted acylcarnitine levels, which accounted for a significant portion of group differences with explained variance values ranging from 40% to 60% for different cluster BPs (Fig. 2j). Notably, the most substantial contributions were observed for medium- and long-chain acylcarnitines, highlighting a likely modifiable role for beta-oxidation function35,36 (Supplementary Fig. 1).
Genetic influences only accounted for a small proportion of variance (Supplementary Data 7–9), except for one epistatic interaction effect of two SNPs: The interplay of rs17806888 (mapped to SUCLG2) and rs924135 (mapped to ABCC1), both of which have been reported to affect succinylcarnitine levels16, explained 32% of the initial partitioning of the data into cluster 2 and cluster 3 at the top of the tree (Fig. 2j and Supplementary Fig. 2). When we examined relations to AD, we observed that both single acylcarnitine-related SNPs and epistatic models were not strongly linked to AD and its biomarkers, although some associations were independently confirmed in the MayoLOAD and ROS/MAP studies (Supplementary Fig. 3 and Supplementary Data 10). The few genetic associations with AD parameters that we found were limited to influences on short-chain and dicarboxylic acylcarnitines.
These findings indicate that the genetic link between AD, acylcarnitine pathways, and the identified bioenergetic subgroups is primarily linked to the TCA cycle and amino acid-based energy metabolism.
In summary, these results suggest that individual bioenergetic capacity, represented by fasting serum acylcarnitine levels, can identify different groups of individuals. These include groups of study participants with less pronounced AD biomarker profiles, early pathological aging with decreased CSF Aβ1-42 levels and relatively low CSF p-tau levels, neurodegenerative processes accompanied by cognitive decline, and more advanced AD biomarker profiles. The stratification of study participants was primarily driven by the modifiable fraction of acylcarnitine levels involved in beta-oxidation, whereas the genetic component pointed towards TCA cycle-related mechanisms.
Bioenergetic age correlates with Alzheimer’s disease pathology
Blood acylcarnitine levels have previously been described to significantly correlate with age22. Extending this concept, we hypothesized that incorporating acylcarnitine levels into a “bioenergetic age”, which might deviate from a person’s chronological age, can provide a single integrated readout of an individual’s bioenergetic capacity. After cross-normalizing the different cohort datasets for better comparability (“Methods” and Supplementary Fig. 4), we fitted a multivariable linear model that regresses age on the fasting serum levels of acylcarnitines in the KORA cohort (Supplementary Fig. 5 and Supplementary Data 11). KORA is a predominantly healthy, population-based cohort without prevalent AD cases, rendering it an appropriate reference for modeling the average aging process. The correlation between bioenergetic age and chronological age in KORA showed an r = 62%. When applied to the ADNI cohort, correlation dropped to 28% (cognitively normal participants only), which was very similar to what we observed when replicating the bioenergetic age computation process in AGES-RS (r = 29%, Supplementary Data 12).
We then examined the different subgroups in ADNI concerning their bioenergetic age. First, we observed a gradual increase in bioenergetic age across the subgroups in the tree, ranging from cluster 6 with the lowest age, through clusters 7 and 5, to cluster 4 with the highest age (Fig. 3a, b). Furthermore, this analysis revealed a strong correlation between bioenergetic age and the first principal component of acylcarnitine levels, indicating that the primary axis of variation in the data and the clustering is associated with bioenergetic age (Supplementary Fig. 6). As we demonstrated in the previous section, the clustering seems primarily driven by modifiable factors, likely associated with individual beta-oxidation capacity. This finding thus suggests that bioenergetic age is also a modifiable factor, indicating that individuals could undergo interventions to transition into a different subgroup.
Fig. 3. Bioenergetic age as a readout of bioenergetic capacity and determinant of bioenergetic subgroups.
a Principal component (PC) analysis of acylcarnitine profiles shows that the first PC expectedly follows the cluster structure from Fig. 2. b Interestingly, the bioenergetic age predicted for the individuals within the dataset similarly corresponds with this cluster organization. c Individuals in the pathologically healthier cluster 2, although only slightly younger than those in cluster 3 in terms of chronological age, display a significantly reduced bioenergetic age. d Cluster 7, which is chronologically older than cluster 6 but demonstrates favorable disease pathology, presents a bioenergetic age that is younger than their chronological age. This observation suggests that cluster 7 may constitute a resilient subgroup of individuals. e Similar to the previous two examples, individuals characterized by an advanced bioenergetic age exhibit more pronounced Alzheimer’s disease pathology compared to those with a younger bioenergetic age. Box plots display the median (central line), interquartile range (box bounds), whiskers extending to the smallest and largest values within 1.5 times the interquartile range from the quartiles, and notches indicating the 95% confidence interval for the median. Details for each statistical test and the corresponding sample sizes are provided in Supplementary Data 13. All tests were two-sided and raw p values were reported. Source data are provided as a Source Data file.
Bioenergetic age highlighted interesting relationships between subgroups and AD pathology throughout the tree (Supplementary Data 13). First, at the top split, we observed that while clusters 2 and 3 showed only a moderate difference in chronological age, there was a remarkable difference in bioenergetic age, with cluster 2 being substantially younger (Fig. 3c). This was in line with the observation that cluster 3 displayed progressed disease pathology compared to cluster 2. Further down the tree, bioenergetic age confirmed that the overall healthier cluster number 7 was substantially younger than the group’s chronological age, offering a potential explanation for their beneficial phenotypes (Fig. 3d). Importantly, this observed effect was not due to cluster 7 consisting of younger participants or a lower number of symptomatic individuals. Rather, we found that individuals within cluster 7 consistently exhibit the same chronological but significantly younger bioenergetic age than other participants irrespective of their diagnostic group (Supplementary Fig. 7). In clusters 8 and 9, we observe the same effect: Bioenergetically older individuals displayed increased AD pathology (Fig. 3e). This was further confirmed by significant associations of bioenergetic age with AD biomarkers across the A-T-(N)-(C) spectrum (Supplementary Data 14), and consistent findings for cognitive function and gray matter volume in AGES-RS (Supplementary Data 15).
In summary, we found that the acylcarnitine-based bioenergetic age metric, which we propose as a potential readout of a person’s bioenergetic capacity, showed strong associations with AD biomarkers beyond the natural aging process. Furthermore, our results suggest that bioenergetic age, which appears to be influenced by modifiable factors related to beta-oxidation function, can be a target for interventions to improve energy- and aging-related outcomes.
Bioenergetic age and succinylcarnitine-linked genotypes modulate future cognitive decline
We next assessed whether the bioenergetic age estimator can predict the trajectory of cognitive decline in AD, and whether this would allow us to identify subgroups of individuals that might particularly benefit from interventions.
We found that baseline bioenergetic age was significantly associated with the rate of cognitive decline in three different cognitive measures in the ADNI study (Fig. 4a–c). Specifically, bioenergetically younger individuals demonstrated a significantly slower decline over 5 years (Supplementary Data 16). This result was replicated using longitudinal diagnosis data from the AGES-RS study (Fig. 4d, Supplementary Data 15), providing further evidence that younger bioenergetic age is protective against cognitive decline. Interestingly, these findings were largely independent of APOE ε4 status, with the single exception of the ADNI composite score for memory, for which the interaction between APOE ε4 status and the bioenergetic age metric reached nominal significance (P = 0.0160).
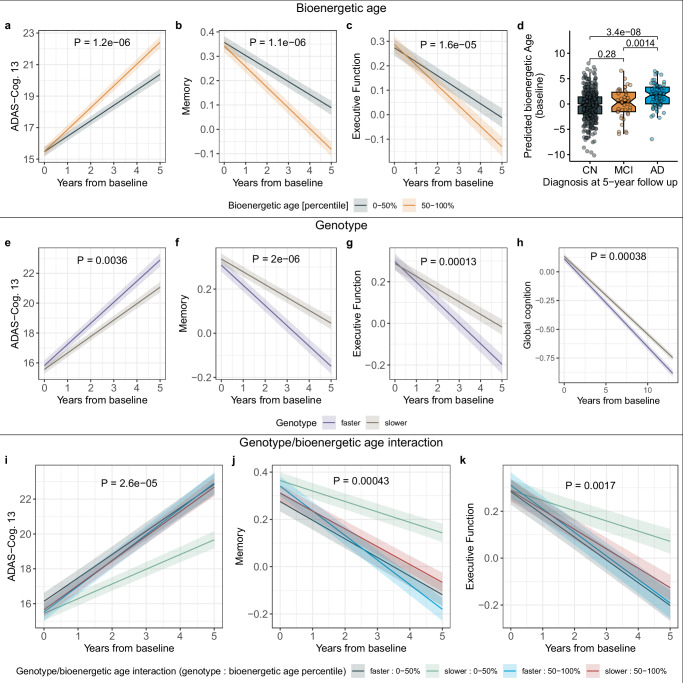
Fig. 4. Bioenergetic age and succinylcarnitine-linked genotypes modulate the rate of cognitive decline.
a–c Bioenergetically younger individuals displayed a slower rate of cognitive decline compared to bioenergetically older individuals in the ADNI cohort. Median-split was only applied for visualization, reported P values are for the continuous variable. d Replication in the AGES study, by comparing bioenergetic age at baseline with clinical AD diagnosis after 5 years. e–g Individuals with an unfavorable genotype configuration assessed by the combination of two SNPs, rs17806888 and rs924135, showed an accelerated rate of cognitive decline. h Replication of the genetic signal in the ROS/MAP study. i–k Interaction analysis: Only individuals with favorable bioenergetic age and favorable genotypes showed a slower cognitive decline. This insinuates that individuals with unfavorable bioenergetic age but favorable genotypes could substantially benefit from targeted intervention. The p values provided represent the contrast between bioenergetic age groups within the favorable (slower) genotype group. Ribbons around linear fits represent ±1 standard error around the estimate. Box plots display the median (central line), interquartile range (box bounds), whiskers extending to the smallest and largest values within 1.5 times the interquartile range from the quartiles, and notches indicating the 95% confidence interval for the median. Details for statistical tests and the corresponding sample sizes/degrees of freedom are provided in Supplementary Data 15 (d) and 16 (a–c, e–k). All tests were two-sided and raw p values were reported. Source data are provided as a Source Data file.
We subsequently examined longitudinal cognitive trajectories in connection with the combination of SNPs rs17806888 and rs924135. These SNPs accounted for a substantial proportion of variance in the clustering (Fig. 2j). Since two SNPs can generate multiple genotype combinations, we identified a representative genotype by evaluating all possible groupings of allele combinations against the rest. We selected the grouping that had the most significant associations across three cognitive measures: ADAS-Cog 13, memory, and executive function (Supplementary Data 17). The genotype identified in this manner appears to significantly modulate cognitive decline (Fig. 4e–g). We attempted to replicate this grouping, which had similar genotype frequencies as in ADNI (59% vs. 58% in ROS/MAP; Supplementary Fig. 8), using trajectories of global cognition over 13 years in the ROS/MAP cohort. Interestingly, significant results only emerged when the analysis was restricted to individuals exhibiting clinically evident cognitive changes, such as a switch in cognitive classification from normal to mild cognitive impairment (MCI), or MCI to AD (Fig. 4h). These results suggest that specific genotype combinations related to short-chain acylcarnitines can influence the rate of future cognitive decline. Nonetheless, this effect might only be pertinent to individuals who are susceptible to cognitive alterations.
Finally, we examined the relationship between cognitive decline and a combination of bioenergetic age and the genotype groupings within the ADNI cohort. Remarkably, it seems that the protective influence of advantageous genotype combinations is limited to individuals with a younger bioenergetic age (Fig. 4i–k). This observation points to a highly interesting subgroup of individuals, those with beneficial genotypes but older bioenergetic age, who could potentially see considerable benefits from early interventions designed to decrease their predicted bioenergetic age. This subgroup constitutes ~30% of the ADNI participants. Replication analysis of this finding was not possible, since none of the available datasets except ADNI had combined fasting acylcarnitine measurements, genotyping, and longitudinal cognitive data.
Interventions targeting the bioenergetic capacity might be as effective as amyloid antibodies
To estimate the potential clinical significance of modulating bioenergetic age, we conducted a simulated clinical trial in the ADNI cohort. We evaluated two metrics: (1) the potential clinical benefit of improving bioenergetic age in general, and (2) the potential benefit specifically in the group with high bioenergetic age but protective genotypes at rs17806888 and rs924135. We followed the protocol of the recent lecanemab trial as a reference study, with only minor deviations (see “Methods”)37. The 0–25% quartile of bioenergetic age at baseline was considered as “treatment” and the 75–100% quartile as “control” group (n = 191 each).
Treatment and control groups showed significant differences for all considered outcomes. This included the Clinical Dementia Rating—Sum of Boxes score (CDR-SB; primary outcome in the lecanemab trial), the ADAS-Cog. 13, the Mini-Mental State Examination (MMSE), and the Functional Activities Questionnaire (FAQ) (see Supplementary Data 18). In addition to symptom severity, the treatment and control group also significantly differed in distributions of chronological age, sex, and AD medication intake. There was no significant difference in APOE ε4 status (n = 102 vs. 108) or presence of the genotype groupings for rs17806888 and rs924135 (n = 101 vs. 96) between groups.
We then estimated mean changes in clinical outcomes over an 18-month period (the duration of the lecanemab trial) for each treatment group using a mixed model for repeated measurements and adjusted least-squares mean changes (Table 1). In addition to the covariates used in the lecanemab trial, the model was corrected for years of education, chronological age, and sex due to the significant differences observed for group characteristics. In addition, to account for the significant correlation between bioenergetic and chronological age, we included the interaction of chronological age and time as a covariate. We noted statistically significant effects across outcomes comparable to those reported for lecanemab, despite the significantly lower power in this analysis compared to the lecanemab trial (which had n > 800 per group). Evaluating the treatment vs. control groups in the group with protective genotypes supports the hypothesized particular benefit in the group with higher bioenergetic age, with average mean differences exceeding those for the groups without stratification by genotype (Table 1, right).
Table 1.
End points of the simulated 18-month clinical trial
| All genotypes | Genotypes [slower decline] | |||
|---|---|---|---|---|
| Outcome | 0–25% quartile | 75–100% quartile | 0–25% quartile | 75–100% quartile |
| CDR-SB | ||||
| Adjusted mean change | 0.906 | 1.441 | 0.751 | 1.647 |
| Adjusted mean change difference | −0.534 (−0.816 to −0.252) | −0.896 (−1.285 to −0.507) | ||
| P value | 2.14 × 10−4 | 7.09 × 10−6 | ||
| ADAS-Cog. 13 | ||||
| Adjusted mean change | 2.520 | 3.942 | 2.317 | 4.680 |
| Adjusted mean change difference | −1.422 (−2.458 to −0.387) | −2.362 (−3.805 to −0.920) | ||
| P value | 7.14 × 10−3 | 1.37 × 10−3 | ||
| MMSE | ||||
| Adjusted mean change | −1.061 | −1.803 | −0.738 | −2.086 |
| Adjusted mean change difference | 0.742 (0.246 to 1.237) | 1.349 (0.669 to 2.028) | ||
| P value | 3.39 × 10−3 | 1.07 × 10−4 | ||
| FAQ | ||||
| Adjusted mean change | 2.542 | 3.999 | 2.146 | 4.104 |
| Adjusted mean change difference | −1.458 (−2.202 to −0.713) | −1.958 (−2.996 to −0.919) | ||
| P value | 1.32 × 10−4 | 2.32 × 10−4 | ||
Discussion
In this study, we investigated the association between AD pathology and bioenergetic capacity, approximated using fasting serum acylcarnitine levels. This bioenergetic capacity is influenced by a combination of modifiable and autosomal mitochondrial genes and naturally declines with age. Historically, acylcarnitine profiles have been widely used in clinical practice for detecting inborn errors in mitochondrial energy metabolism38. Moreover, numerous previous studies have shown the relevance of blood acylcarnitine levels and related mitochondrial pathways in metabolic disease and AD28–32,39,40. The present study combines these earlier findings with a translational framework of bioenergetic capacity with the potential for targeted interventions, based on collective evidence from more than 9000 individuals.
Hierarchical clustering-based SGI classified participants along the progression of their AD biomarkers. This stratification was predominantly driven by the modifiable, non-genetic fraction of acylcarnitine levels, specifically medium- and long-chain acylcarnitines associated with beta-oxidation function, accounting for 40–60% of variance explained. Notably, the division into the initial two main clusters was additionally influenced by an interaction between two genetic variants associated with succinylcarnitine, an intermediate of the TCA cycle, and amino acid-based energy metabolism. These observations indicate that beta-oxidation might be a promising target for intervention, whereas targeting the TCA cycle may be complicated by complex genetic influences.
To integrate our findings into a unified score approximating an individual’s bioenergetic capacity, we derived a “bioenergetic age” metric. This metric showed strong correlations with AD biomarkers, including brain glucose uptake, cognitive function, and disease progression. Furthermore, the two-SNP genotype related to succinylcarnitine, which influenced the clustering, was also predictive of cognitive decline. This is in line with previous findings reporting an association of genetic variation in the SUCLG2 gene with cognitive decline41. Combined analysis of the bioenergetic age with the two-SNP model revealed that certain allele combinations appeared to result in resilience against cognitive decline, but only in individuals with a younger bioenergetic age. Furthermore, a simulated clinical trial indicates that effect sizes of potential treatments targeting the bioenergetic capacity might be comparable to the recently approved anti-amyloid drug lecanemab. Throughout all results, it is evident that acylcarnitines as a group correlate significantly with disease biomarkers and severity, underscoring their potential relevance in AD pathology. Thus, with further clinical validation, acylcarnitines could be leveraged to improve disease staging and potentially guide targeted therapeutic interventions.
Based on our results, we propose the following model of bioenergetic dysregulation in AD: as individuals age, their bioenergetic capacity decreases. This decrease accelerates with the onset of AD due to dysfunctional glucose uptake in the brain. The body then relies on alternative energy sources, primarily lipids via beta-oxidation and eventually protein-based energy production. Individuals with larger bioenergetic capacity, due to favorable genetics and maintained metabolic health, can temporarily compensate for these changes, resulting in resilience. However, when beta-oxidation can no longer provide sufficient energy, the body resorts to amino acid-based energy production. Only at that stage, amino acid metabolism will be negatively affected by genetic influences on related transporters and enzymes. This results in a ripple effect, increasing vulnerability to pathological processes and leading to accelerated disease progression. Overall, the bioenergetic capacity may thus be likened to a metabolic reserve providing a significant source of resilience against the disease. This is in line with previous research, showing the brain can exhibit a metabolic signature of cognitive resilience in individuals over 80 years old42.
Our present study has several limitations: (1) a major goal of this work was to demonstrate the relevance of acylcarnitine metabolism in AD. To this end, we focused on classical biomarkers, such as CSF Aβ1-42, p-tau, FDG-PET, and cognitive measures. Future studies are needed to consider additional, emerging evidence suggesting a complex landscape of mixed pathologies extending beyond the amyloid cascade hypothesis43,44. Of note, our primary clinical outcomes for assessing potential resilience were cognitive function and activities of daily living. Regardless of the precise molecular etiology of the disease, improvements in these areas are considered the most important outcomes for patients suffering from dementia. (2) Our simulated clinical trial was not a properly matched randomized clinical trial (RCT), which is a result of the bioenergetic age being significantly associated with demographics, biomarker status, and symptom severity. Consequently, the trial groups were significantly different in these regards, which would be avoided in a real RCT. To address this, we employed a complex statistical model incorporating various confounding and interaction effects to mitigate as many of these factors as possible. Future studies are needed to further corroborate our findings. (3) Our conclusions are based on fasting serum acylcarnitine levels, which do not specify affected brain regions or cell types. Although earlier findings show correlations between acylcarnitine levels in the blood and the dorsolateral prefrontal cortex25, the general association between peripheral and central metabolism is often still hypothetical. Further research is needed to determine the exact relationship and relevance to specific brain functions, including beta-oxidation and TCA cycle efficiency in different cell types.
Our key proposition—that the reduction of bioenergetic age will increase bioenergetic capacity and thus resilience in a genotype-specific manner—requires further validation in independent cohorts due to the rarity of studies that combine genetic, metabolomics, and longitudinal cognitive data. Once established as a robust marker of mitochondrial health, bioenergetic age combined with the two-SNP genotype could be utilized to select individuals for targeted interventions. Interventions to boost the bioenergetic capacity might include (1) low-carb or ketogenic diets, which directly influence mitochondrial beta-oxidation through nutritional lipids45, (2) physical activity, which is known to beneficially affect energy metabolism and mitochondrial fitness46,47, and (3) the use of drugs like Metformin, which was originally used to treat insulin resistance and T2D but recently has increasingly been shown to have additional beneficial effects, including the improvement of mitochondrial health48,49. Importantly, the bioenergetic age score could serve as a monitoring tool for such interventions, using established, cost-efficient, and fast acylcarnitine-measuring technologies, for example, based on dried blood spots50. While the ultimate long-term benefits of an intervention study can only be seen after decades, bioenergetic aging assessed through such minimally invasive acylcarnitine measurements is expected to predict the success or failure of long-term interventions within a substantially shorter time frame.
Methods
ADNI study
The ADNI was launched in 2003 as a public-private partnership, led by Principal Investigator Michael W. Weiner, MD. The primary goal of ADNI has been to test whether serial magnetic resonance imaging, positron emission tomography, other biological markers, and clinical and neuropsychological assessment can be combined to measure the progression of MCI and early AD. For up-to-date information, see www.adni-info.org. Written informed consent was obtained at enrollment, which included permission for analysis and data sharing. Consent forms were approved by each participating site’s institutional review board.
Data was obtained from the AMP-AD Knowledge Portal (https://adknowledgeportal.synapse.org), see “Data Availability Statement” and the ADNI database at https://adni.loni.usc.edu. The AMP-AD Knowledge Portal is the distribution site for data, analysis results, analytical methodology, and research tools generated by the AMP-AD Target Discovery and Preclinical Validation Consortium and multiple Consortia and research programs supported by the National Institute on Aging.
Metabolomics data was available for 1681 participants. Samples were profiled with the targeted Biocrates p180 kit (Biocrates, Innsbruck, Austria). Metabolomics data processing largely followed a previously published protocol32: Of the 186 metabolites covered by the platform, four were removed due to technical issues, leaving a total of 182 metabolites for further analysis. Samples were distributed across 23 plates. Each plate included NIST Standard Reference samples. Twenty-two metabolites with large numbers of missing values (>40%) were excluded and plate batch effects were removed by cross-plate mean normalization using NIST sample metabolite concentrations. The sample set moreover contained blinded duplicated measurements for 19 samples (ADNI-1) and blinded triplicated measurements for 17 samples (ADNI-GO and -2) distributed across plates. These duplicated and triplicated study samples were used to remove 20 metabolites with coefficients of variation >20% or intra-class correlation <65%. Biological replicates were then averaged, and non-fasting participants (n = 108) were excluded. We imputed missing metabolite data using half the value of the lower limit of detection per metabolite, log2-transformed metabolite concentrations, centered and scaled distributions to a mean of zero and unit variance and winsorized single outlying values to 3 standard deviations. Mahalanobis distance was used for the detection of multivariable subject outliers. Applying a critical Chi-square value of P < 0.01 resulted in the removal of 42 samples. Finally, metabolites were adjusted for significant medication effects using stepwise backwards selection (for details see Toledo et al.30). The final dataset contained 140 metabolite measurements, covering 23 acylcarnitine species (Supplementary Data 2), for 1531 individuals.
Representative phenotypes across A-T-(N)-(C) measures34 were used as clinical phenotypes, including baseline levels of CSF amyloid β1-42 (CSF Aβ1-42, A), CSF p-tau (T), ROI-based FDG-PET measures of average glucose uptake across the left and right angular, left and right temporal, and bilateral posterior cingulate regions (N), and ADAS-Cog. 13 scores (C). Diagnostic groups were coded as follows: 1 = cognitively normal (CN) individuals and individuals with subjective memory complaints (SMC); 2 = early MCI (EMCI); 3 = EMCI with CSF Aβ1-42 pathology; 4 = late MCI (LMCI); 5 = LMCI with CSF Aβ1-42 pathology; 6 = AD cases; 7 = AD cases with CSF Aβ1-42 pathology. Sex, age, BMI, years of education, and copies of APOE ε4 were included as covariates in cross-sectional association tests. Epistatic analyses were additionally adjusted for the ADNI study phase. Longitudinal analyses of cognitive trajectories were adjusted for age and diagnosis at baseline, sex, copies of APOE ε4, education, and ADNI study phase. Longitudinal analyses were restricted to 5 years of follow-up to retain statistical power (minimal n = 378).
Whole genome genotyping was available for 1548 ADNI participants, with 1378 participants having overlapping metabolomics data. Genotyping data were collected using the Illumina Human 610-Quad, HumanOmni Express, and HumanOmni 2.5M BeadChips. Pre-imputation quality control procedures included filtering for SNP call rate <95%, Hardy-Weinberg equilibrium test p value < 1 × 10−6, minor allele frequency <1%, participant call rate <95%, and discordance between reported and inferred sex. Non-Hispanic Caucasian participants were selected using HapMap 3 genotype data and multidimensional scaling analysis. Genotype imputation was performed for each BeadChip type separately using the Haplotype Reference Consortium reference Panel r1.1.
ROS/MAP study
The Religious Order Study (ROS) and the Rush Memory and Aging Project (MAP) studies51 are longitudinal cohort studies of aging and AD, conducted by the Rush Alzheimer’s Disease Center and designed to be used in joint analyses to maximize sample size. Both studies were approved by an Institutional Review Board at Rush University Medical Center. All participants signed an informed consent and a repository consent to allow their biospecimens and data to be used for ancillary studies. Further, all participants signed an Anatomic Gift Act for organ donation for research. More details can be found at www.radc.rush.edu.
Imputed genome-wide genotype data for 2059 study participants was obtained from the AMP-AD Knowledge Portal (https://adknowledgeportal.synapse.org), see “Data Availability Statement”. A description of this data, including quality control procedures and imputation, was previously published52. We further included phenotypic data on clinical diagnosis at death, global cognition during lifetime, amyloid-β and paired helical filament (PHF)-tau protein load in brain tissue, global burden of AD neuropathology (mean of neuritic plaques, diffuse plaques, and neurofibrillary tangles).
Epistatic analyses were adjusted for sex, age at death, education, post-mortem interval, and number of copies of APOE ε4 as covariates. Longitudinal analysis of cognitive trajectories was adjusted for baseline age (instead of age at death) and clinical diagnosis, while post-mortem interval was omitted here. Longitudinal analysis was restricted to 13 years of follow-up, as at this time point we had a similar sample size left (n = 368) as in the longitudinal analyses in ADNI.
MayoLOAD study
The Mayo Study of late-onset AD (MayoLOAD)53 is a case/control study from three different series: Mayo Clinic Jacksonville, Mayo Clinic Rochester, and Mayo Clinic Brain Bank series. The study was approved by the appropriate institutional review board and appropriate informed consent was obtained from all participants. Preprocessed genotype data for 2067 participants were obtained from the AMP-AD Knowledge Portal (https://adknowledgeportal.synapse.org), see “Data Availability Statement”. Briefly, samples with call rates <90% were removed. In addition, samples were discarded based on mismatch between inferred and reported sex. Further, samples were filtered based on inferred relatedness to ensure that the resultant sample set represents unrelated individuals. SNPs with call rates <90%, minor allele frequencies <0.01, and/or Hardy-Weinberg P values < 0.001 were eliminated. Total genotyping rate after filtering was 99.2%. Genotypes were then imputed using the 1000 genomes phase 3 reference panel54 by first prephasing genotypes using SHAPEIT2 (v 2.12)55 and then imputing using IMPUTE2 (v 2.3.2)56.
Data on clinical diagnosis was numerically coded into four categories: 1 = controls with no evidence for AD-related neuropathology; 2 = clinically normal controls without neuropathology assessment; 3 = clinical AD without neuropathology confirmation; 4 = clinical diagnosis of AD dementia with neuropathology-confirmed AD. Covariates for the study included sex, age at death, and number of copies of APOE ε4.
KORA study
The Cooperative Research in the Region of Augsburg (KORA) study is a population-based sample from the general population living in the region of Augsburg, Southern Germany57. Here we used data from the KORA F4 study, the first follow-up examination of KORA S4 in 2006–2008. The study, including the protocols for subject recruitment and assessment and the informed consent for participants, was reviewed and approved by the local ethical committee (Bayerische Landesärztekammer).
Metabolomics data was available for 3029 predominantly healthy participants with data on fasting serum acylcarnitine levels, sex, age, and BMI. Metabolic profiling was conducted using the targeted Biocrates p150 kit (Biocrates, Innsbruck, Austria), a precursor of the p180 kit where measurements of acylcarnitines are performed analogously as on the p180 kit. A detailed description of the processing of metabolomics data is provided in Mittelstrass et al.58. Briefly, similar to the ADNI procedure, filters for coefficient of variation (<25%), missingness (<10%), and correlation of repeated measurements (>50%) were applied to remove metabolites of limited measurement quality. Multivariable subject outliers were identified using the Mahalanobis distance, remaining missing values were imputed, and data was log-transformed for subsequent analyses. Data for 22 out of 23 acylcarnitines investigated in ADNI (Supplementary Data 2) were available here, with the exception being levels of C4:1, which was not reliably measured in KORA.
AGES-Reykjavik study
The Age, Gene/Environment Susceptibility-Reykjavik Study (AGES-RS) is an epidemiologic study focusing on four biologic systems: vascular, neurocognitive (including sensory), musculoskeletal, and body composition/metabolism59. AGES-RS was approved by the National Bioethics Committee in Iceland that acts as the Institutional Review Board for the Icelandic Heart Association (approval number: VSN-00-063) and by the National Institute on Aging Intramural Institutional Review Board. A multistage consent is obtained in AGES-RS to cover participation, use of specimens and DNA, and access to administrative records.
Fasting serum-based measurements for the 23 acylcarnitines generated with the targeted Biocrates p180 kit (Biocrates, Innsbruck, Austria) were available for 575 AGES-RS participants at baseline, of which 544 had a follow-up diagnostic assessment after 5 years. As in ADNI, data was batch-normalized, log2-transformed, centered, scaled, and adjusted for medication effects. All analyses in AGES-RS were adjusted for age, sex, education, and copies of APOE ε4. In the analysis of gray matter volume, intracranial volume was included as an additional covariate.
Comparability of AD case ascertainment in ADNI and AGES-RS
Both ADNI and AGES-RS use a threshold-based identification of individuals who are screen-positive for cognitive impairment. This includes the MMSE and a test for logical memory from the Wechsler Adult Intelligence Scale—Revised59–61 Case ascertainment further includes a larger battery of cognitive tests and further includes assessment of neuropsychiatric symptoms/depression and (instrumental) activities of daily functioning. Consensus diagnosis of possible and probable AD in both cohorts was based on the criteria defined by the National Institute of Neurological and Communicative Disorders and Stroke–Alzheimer’s Disease and Related Disorders Association (NINCDS-ADRDA). Both cohorts have been meta-analyzed before, including in the International Genomics of Alzheimer’s Project (IGAP)62.
Subgroup identification analysis (SGI)
We used the SGI software package33 for automatic SGI. For the present study, the analysis consisted of the following three steps. First, the data matrix was standardized (mean 0, standard deviation 1 for each metabolite) before analysis. The samples were then hierarchically clustered based on their acylcarnitine profiles using the Euclidean distance metric and Ward linkage63. This resulted in a dendrogram, i.e., a binary tree that provides the hierarchical structure of sample similarities. Second, we performed an association analysis with AD-related phenotypes and the 32 acylcarnitine-associated SNPs. Each BP in the dendrogram provides two subgroups of participants, from the top BP that separates the entire dataset into two parts over smaller clusters down to the bottom where subgroups consist of only a few samples each. For each BP, an association test of the respective left vs. right cluster was performed using linear regression for A-T-(N)-(C) measures, and ordinal regression for copies of APOE ε4 and the numeric coding of diagnosis. To avoid low-powered statistical tests, BPs were only tested if both of the two underlying subgroups were larger than N = 77 samples (valid cluster pairs), which corresponds to 5% of all samples. Third, we adjusted p values for multiple testing by correcting the number of valid cluster pairs s = 11. Since a dendrogram’s clusters are nested, non-overlapping groups, the s statistical tests performed are strictly independent. Thus, control of the family-wise error rate can be achieved using a Bonferroni-like multiple testing correction by adjusting each p value by a factor of s (corresponding to an adjusted threshold of p ≤ 4.55 × 10−3).
Collection of SNPs and SNP combinations associated with blood acylcarnitine levels
We obtained a list of 32 acylcarnitine-associated SNPs (Supplementary Data 5) from a large GWAS study performed on a total of 7824 individuals from the KORA and TwinsUK studies16. The generalizability of the effects of these SNPs on ADNI was tested using a targeted genetic association screening, where we tested for influences of all 32 SNPs against all 23 acylcarnitine species assuming an additive genetic model. Only age and sex were included as covariates, following the protocol of the reference study. Associations were considered to be significant if they had an FDR-adjusted p value ≤ 0.05.
The reference GWAS further lists multi-SNP combinations of one lead SNP per associated locus that in concert explained the largest fraction of the heritable population variance of single acylcarnitine concentrations in blood. We used these multi-SNP combinations in epistatic modeling.
Epistatic analysis
To determine multi-locus epistatic associations with the clustering and AD-related outcomes, we ran an epistasis model using SNP combinations for each acylcarnitine with significant associations in more than one locus. For these multi-locus genetic models, we combined SNP genotypes in all combinations into aggregated genotypes h to test pure genetic interactions; for example, if SNP1 has alleles TA for an individual, and SNP2 has alleles CC, then the aggregated genotype is TACC. Since each SNP can have up to three different genotypes (major allele homozygote, heterozygote, minor allele homozygote), the aggregation of N SNPs can yield up to 3N combinations. For each outcome (cluster pairs and AD-related phenotypes) y and aggregated genotype h, we then computed the following two statistical models: the base model M1 regressing the outcome in question (dependent variable) on the set of confounders only; and the full model M2 that includes h in addition to confounders. The models use ordinal regression with log-log link functions and logistic regression for binary outcomes. The genetic variable h is treated as a factor and is thus expanded into a binary indicator variable for each factor level during model fitting. If a factor level is observed in less than 10 participants, it was omitted due to lack of statistical power. To control for spurious effects of overly fractionated factor variables with many different SNP combinations, we additionally capped the maximum number of degrees of freedom (DFs). This was achieved by imposing a ridge-type penalty following the suggestions described by Harrell64 and setting the maximum number of DFs to 15. Statistical significance was finally assessed using a likelihood ratio test between M1 and M2.
Analysis of explained variance
The variance between cluster pairs explained by significant genetic effects (both for single SNPs and multi-SNP combinations) was estimated using McKelvey’s measure, which has been described as a robust approach for logistic regression models65. To estimate the variance explained by each of the 23 acylcarnitines, we calculated a linear mixed model including a random intercept for each BP, as implemented in the variancePartition R package66. To account for potential confounding through non-modifiable factors, we included all significant SNPs, the interaction between rs17806888 and rs924135, as well as age, sex, BMI, and Education as covariates in the model. Analysis of explained variance was restricted to cluster pairs for which we observed significant differences in AD-related phenotypes or demographic variables.
Normalization of acylcarnitine levels across cohorts
To enable comparability of semi-quantitative acylcarnitine levels across cohorts, we performed a reference group-based cross-normalization for all 22 acylcarnitines that were measured in KORA, ADNI, and AGES-RS. We first extracted all healthy female individuals from the three cohorts. Additional selection parameters in KORA besides female sex included age 60–72 years and BMI 23.31–29.72 kg/m2. In ADNI and AGES-RS we included all cognitively normal females with an age of 72 years or younger. With the assumption that the real acylcarnitine level distributions are similar for these reference groups, measurement values were then centered and scaled in each cohort separately. In the final step, we filtered out multivariate outliers (n = 3; all ADNI) based on the Mahalanobis distance (see more details in the description of ADNI data). The remaining reference subjects were used to rescale z-scored acylcarnitine concentrations in the respective complete cohorts.
Computation and analysis of bioenergetic age
After normalizing acylcarnitine levels to these reference groups, we calculated a linear regression model using the reference-transformed 22 acylcarnitines to predict chronological age in KORA. To investigate the robustness of the model, we performed 10-fold, 3 times repeated cross-validation (Supplementary Data 11). The coefficients of the model are provided in Supplementary Data 19. We then applied this model to ADNI and AGES-RS using reference-transformed acylcarnitine levels. In this analysis, we did not adjust acylcarnitine levels for the ADNI study phase, as participants in ADNI-GO/2 were on average 2.65 years younger than participants in ADNI-1 (P = 5.76 × 10−13), such that adjustment would have confounded the reference set-based rescaling. The z-scored difference between chronological age and bioenergetic age was derived by subtracting their z-scored transformations, centering to zero, and scaling to unit variance. Differences between ADNI cluster pairs associated with AD-related phenotypes and the three age measures (chronological age, predicted bioenergetic age, and their delta) were assessed using linear regression without adjustment for any additional variables. Cross-sectional associations of predicted bioenergetic age with A-T-(N)-(C) measures in ADNI, as well as with total gray matter volume, cognition, and clinical diagnosis in AGES-RS were tested using linear regression while adjusting for all relevant covariates (see study-specific sections), including chronological age.
Analysis of longitudinal cognitive trajectories
For analyses of cognitive trajectories in ADNI, we included the composite ADNI scores for memory (ADNI-MEM)67 and executive function (ADNI-EF)68 in addition to the ADAS-Cog. 13, which have been described to be more sensitive to subtle cognitive changes and have been used in studying resilience to AD before69. We tested longitudinal associations with predicted bioenergetic age using linear mixed-effects models with cognitive scores as dependent variables. The explanatory variable of interest was the interaction of bioenergetic age and time. Models were adjusted for relevant covariates (see description of ADNI data), including chronological age at baseline, and allowed for random intercepts for each participant. For binarization of the SNP combination of rs17806888 and rs924135 into two groups (slower vs. faster progression of cognitive decline), we used the same linear mixed effect model (replacing bioenergetic age with genotype aggregates) and iterated over all binary combinations of aggregated genotypes. We removed aggregates with less than 20 observations to avoid spurious associations, leaving us with six aggregated genotypes. We then selected the combination that showed the highest significance across the three cognitive scores using Fisher’s sum of logs method70. The three-way interaction analysis was performed in the same way (explanatory variable of interest: time × bioenergetic age × SNP grouping), while for the calculation of the association p value, we used an ANOVA, where the reduced model omitted the interaction with time. In AGES-RS, we used standard linear regression to test for an association between baseline predicted bioenergetic age and pheno-conversion after 5 years, while adjusting for relevant covariates. To replicate the genotype grouping obtained in ADNI in ROS/MAP, we applied the same (in terms of predictors/covariates) linear mixed-effects model as in ADNI using global cognition as outcome. As replication in the full subset of the ROS/MAP cohort available here failed, we selected a subset of n = 1081 participants (a total of 1936 with cognitive and genetic data available) where a clinically relevant change in cognitive status (from cognitively normal to MCI or AD, or from MCI to AD) was noticed in any follow-up visit (1–26 years, meantime until change = 5.71 years).
Simulated clinical trial to estimate the potential treatment effects targeting the bioenergetic capacity
For the simulated trial, we restricted ADNI participants to cognitively normal controls with subjective memory complaints, individuals with MCI, and mild AD cases, which had data available both at baseline and the 1-year and 2-year follow-up visits (final follow-up for mild AD cases in ADNI). Cognitively normal controls without any complaints were excluded to allow comparisons with the lecanemab trial37. Participant characteristics are provided in Supplementary Data 18. The treatment and control groups were defined as the 0–25% quartile and the 75–100% quartile in bioenergetic age at baseline, respectively. As outcomes, we used the CDR-SB and ADAS-Cog. 13 (for comparisons to the lecanemab trial, although here the ADAS-Cog. 14 was used), as well as the MMSE and the FAQ score to cover a broad spectrum of evaluations of both cognitive function and activities of daily living.
The analysis for outcome changes from baseline to 24 months followed the lecanemab trial protocol with minor adjustments. As in the reference trial, we used linear mixed effect models for repeated measures (including participant ID as random intercept) modeling the longitudinal outcome scores as dependent variables and the following explanatory variables: baseline outcome score, trial group, visit (year after baseline), baseline diagnosis (cognitively normal, MCI, AD), use of memantine and/or acetylcholinesterase inhibitors, copies of APOE ε4, racial/ethnic category, baseline outcome score-by-visit interaction to model the differences in progression, and trial group-by-visit interaction as the variable of interest. Deviating from the lecanemab protocol, we included years of education, sex, chronological age, and chronological age-by-visit interaction as additional fixed effects to account for non-matched group characteristics. To calculate contrasts within the genotype grouping of rs17806888 and rs924135 associated with slower cognitive decline, we additionally included genotype grouping, genotype grouping-by-trial group interaction, and genotype grouping-by-visit interaction as covariates and the three-way interaction genotype grouping-by-trial group-by-visit as variable of interest. Finally, we estimated adjusted least-squares mean changes and adjusted least-squares mean change differences across outcomes from the 24-month model restricting trial time to 18 months.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Supplementary information
Description of Additional Supplementary Files
Source data
Acknowledgements
This work was supported by the National Institutes of Health/the National Institute of Aging through grants RF1AG058942, RF1AG059093, U01AG061359, U19AG063744, and R01AG069901. In addition, the following funding sources are acknowledged: ADNI: data collection and sharing for this project was funded by the Alzheimer’s Disease Neuroimaging Initiative (ADNI) (National Institutes of Health Grant U01 AG024904) and DOD ADNI (Department of Defense award number W81XWH-12-2-0012). ADNI is funded by the National Institute on Aging, the National Institute of Biomedical Imaging and Bioengineering, and through generous contributions from the following: AbbVie, Alzheimer’s Association; Alzheimer’s Drug Discovery Foundation; Araclon Biotech; BioClinica, Inc.; Biogen; Bristol-Myers Squibb Company; CereSpir, Inc.; Cogstate; Eisai Inc.; Elan Pharmaceuticals, Inc.; Eli Lilly and Company; EuroImmun; F. Hoffmann-La Roche Ltd and its affiliated company Genentech, Inc.; Fujirebio; GE Healthcare; IXICO Ltd.; Janssen Alzheimer Immunotherapy Research & Development, LLC.; Johnson & Johnson Pharmaceutical Research & Development LLC.; Lumosity; Lundbeck; Merck & Co., Inc.; Meso Scale Diagnostics, LLC.; NeuroRx Research; Neurotrack Technologies; Novartis Pharmaceuticals Corporation; Pfizer Inc.; Piramal Imaging; Servier; Takeda Pharmaceutical Company; and Transition Therapeutics. The Canadian Institutes of Health Research is providing funds to support ADNI clinical sites in Canada. Private sector contributions are facilitated by the Foundation for the National Institutes of Health (www.fnih.org). The grantee organization is the Northern California Institute for Research and Education, and the study is coordinated by the Alzheimer’s Therapeutic Research Institute at the University of Southern California. ADNI data are disseminated by the Laboratory for Neuro Imaging at the University of Southern California. Data collection and sharing for this project was further funded by the Alzheimer’s Disease Metabolomics Consortium (National Institute on Aging R01AG046171, RF1AG051550, and 3U01AG024904-09S4). ROS/MAP: study data were provided by the Rush Alzheimer’s Disease Center, Rush University Medical Center, Chicago. Data collection was supported through funding by NIA grants P30AG10161 (ROS), R01AG15819 (ROSMAP; genomics and RNAseq), R01AG17917 (MAP), R01AG30146, R01AG36042 (5hC methylation, ATACseq), RC2AG036547 (H3K9Ac), R01AG36836 (RNAseq), R01AG48015 (monocyte RNAseq) RF1AG57473 (single nucleus RNAseq), U01AG32984 (genomic and whole exome sequencing), U01AG46152 (ROSMAP AMP-AD, targeted proteomics), U01AG46161(TMT proteomics), U01AG61356 (whole genome sequencing, targeted proteomics, ROSMAP AMP-AD), the Illinois Department of Public Health (ROSMAP), and the Translational Genomics Research Institute (genomic). Additional phenotypic data can be requested at www.radc.rush.edu. MayoLOAD: Study data were provided by the Mayo Clinic Alzheimer’s Disease Genetic Studies, led by Dr. Nilüfer Ertekin-Taner and Dr. Steven G. Younkin, Mayo Clinic, Jacksonville, FL using samples from the Mayo Clinic Study of Aging, the Mayo Clinic Alzheimer’s Disease Research Center, and the Mayo Clinic Brain Bank. Data collection was supported through funding by NIA grants P50 AG016574, R01 AG032990, U01 AG046139, R01 AG018023, U01 AG006576, U01 AG006786, R01 AG025711, R01 AG017216, R01 AG003949, NINDS grant R01 NS080820, CurePSP Foundation, and support from Mayo Foundation. KORA: the KORA study was initiated and financed by the Helmholtz Zentrum München – German Research Center for Environmental Health, which is funded by the German Federal Ministry of Education and Research (BMBF) and by the State of Bavaria. AGES-RS: the AGES-RS study has been funded by NIH contract N01-AG-12100, the NIA Intramural Research Program, Hjartavernd (the Icelandic Heart Association) contract HHSN271201200022C, and the Althingi (the Icelandic Parliament). The Alzheimer's Disease Neuroimaging Initiative (ADNI): Data used in the preparation of this article were obtained from the ADNI database (adni.loni.usc.edu). As such, the investigators within the ADNI contributed to the design and implementation of ADNI and/or provided data but did not participate in the analysis or writing of this report. A complete listing of ADNI investigators can be found at: http://adni.loni.usc.edu/wp-content/uploads/how_to_apply/ADNI_Acknowledgement_List.pdf. The Alzheimer's Disease Metabolomics Consortium (ADMC): Data and methods used in the preparation of this article were obtained from the ADMC. The full list of contributing scientists is available at https://sites.duke.edu/adnimetab/team/.
Author contributions
Conceptualization and methodology: M.A., M.B., G.K., and J.K.; data analysis: M.A., M.B., and T.W.; metabolomics data generation and QC: M.A., G.K., R.W.S., J.A., A.P., V.G., L.J.L., K.S., and R.K.D; genotype data generation and QC: M.A., P.L.D.J., N.E.T., D.A.B., K.N., and A.J.S.; visualization: M.A., M.B., G.K., and J.K.; supervision: R.K.D., A.J.S., M.A., G.K., J.K., P.L.D.J. N.E.T., D.A.B., V.G., L.J.L., K.S., and P.M.D.; funding acquisition: R.K.D., A.J.S., M.A., G.K., P.L.D.J., N.E.T., D.A.B., and P.M.D.; writing—original draft: M.A., M.B., G.K., J.K., and P.M.D.; results interpretation: M.A., M.B., K.S., G.K., and J.K.; writing—review and editing: all authors. All authors read and approved the final manuscript. ADNI and ADMC consortia provided data and analysis protocols.
Peer review
Peer review information
Nature Communications thanks Stefano Sensi, Russell Swerdlow and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Data availability
KORA data can be accessed upon request at https://helmholtz-muenchen.managed-otrs.com/external/. AGES-RS data can be accessed upon request according to informed consent at https://hjarta.is/en/research/ages-phase-1/. All other omics datasets are available via the AD Knowledge Portal (https://adknowledgeportal.org). The AD Knowledge Portal is a platform for accessing data, analyses, and tools generated by the Accelerating Medicines Partnership (AMP-AD) Target Discovery Program and other National Institute on Aging (NIA)-supported programs to enable open-science practices and accelerate translational learning. The data, analyses, and tools are shared early in the research cycle without a publication embargo on secondary use. Data are available for general research use according to the following requirements for data access and data attribution (https://adknowledgeportal.org/DataAccess/Instructions). For access to the content described in this manuscript see: ADNI metabolomics data from the AbsoluteIDQ-p180 kit is available at the AD Knowledge Portal under 10.7303/syn5592519 (ADNI-1) and 10.7303/syn9705278 (ADNI-GO/-2), the full complement of clinical and demographic data for the ADNI cohorts are hosted on the LONI data sharing platform and can be requested at https://adni.loni.usc.edu/data-samples/adni-data/#AccessData. ROS/MAP imputed genotype data are available at the AD Knowledge Portal under 10.7303/syn3157329, study meta-data, basic covariates, and clinical variables are available under 10.7303/syn3157322. The full complement of clinical and demographic data for the ROS/MAP cohorts is hosted on the Rush Alzheimer’s Disease Center data-sharing platform and can be requested at www.radc.rush.edu. MayoLOAD genotyping data are available at the AD Knowledge Portal under 10.7303/syn3157238, study meta-data and covariates are available under 10.7303/syn3205821.6. Source data are provided with this paper.
Code availability
All R code required to conduct the analyses reported here is available at https://github.com/compneurobio/bioenergetic_capacity.
Competing interests
A.J.S. is a member of the Scientific Advisory Board of Bayer Oncology and of the Dementia Advisory Board of Siemens Medical Solutions USA, Inc.; A.J.S received in-kind support from Avid Radiopharmaceuticals, a subsidiary of Eli Lilly (PET tracer precursor); A.J.S. received Editorial Office Support as Editor-in-Chief from Springer-Nature Publishing (Brain Imaging and Behavior). P.M.D. has received research grants (through Duke University) from Lilly, Avanir, Bausch, Alzheimer’s Drug Discovery Foundation; P.M.D. has received advisory fees from Verily, Otsuka, Genomind, Cogniciti, Clearview, VitaKey, Neuronix, Neuroglee, and Transposon Therapeutics; P.M.D. owns shares or options in UMethod, Evidation Health, Transposon, Marvel Biome, and Advera Health; P.M.D. serves on the board of Apollo and is a co-inventor (through Duke University) on patents relating to dementia biomarkers, metabolomics, and therapies. R.K.D holds equity in Metabolon Inc. M.A., R.K.D., and G.K. are co-inventors (through Duke University/Helmholtz Zentrum München) on patents on applications of metabolomics in diseases of the central nervous system; M.A., R.K.D., G.K., and J.K. hold equity in Chymia LLC and IP in PsyProtix/atai Life Sciences N.V. that are exploring the potential for therapeutic applications targeting mitochondrial metabolism in treatment-resistant depression. JK is a confounder of iollo, advisor to Everfur, and advisor/part-time employee at ExactRx. The remaining authors declare no competing interests.
Footnotes
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Lists of authors and their affiliations appear at the end of the paper.
Contributor Information
Matthias Arnold, Email: matthias.arnold@helmholtz-muenchen.de.
Jan Krumsiek, Email: jak2043@med.cornell.edu.
The Alzheimer’s Disease Neuroimaging Initiative:
P. Murali Doraiswamy, Kwangsik Nho, Andrew J. Saykin, and Rima Kaddurah-Daouk
Alzheimer’s Disease Metabolomics Consortium:
Matthias Arnold, P. Murali Doraiswamy, Kwangsik Nho, David A. Bennett, Andrew J. Saykin, Karsten Suhre, Rima Kaddurah-Daouk, Gabi Kastenmüller, and Jan Krumsiek
Supplementary information
The online version contains supplementary material available at 10.1038/s41467-025-57032-0.
References
- 1.Mosconi, L. et al. Pre-clinical detection of Alzheimer’s disease using FDG-PET, with or without amyloid imaging. J. Alzheimers Dis.20, 843–854 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Heiss, W. D. et al. Positron emission tomography findings in dementia disorders: contributions to differential diagnosis and objectivizing of therapeutic effects. Keio J. Med.38, 111–135 (1989). [DOI] [PubMed] [Google Scholar]
- 3.Heiss, W. D., Szelies, B., Kessler, J. & Herholz, K. Abnormalities of energy metabolism in Alzheimer’s disease studied with PET. Ann. N. Y. Acad. Sci.640, 65–71 (1991). [DOI] [PubMed] [Google Scholar]
- 4.Kessler, J., Herholz, K., Grond, M. & Heiss, W. D. Impaired metabolic activation in Alzheimer’s disease: a PET study during continuous visual recognition. Neuropsychologia29, 229–243 (1991). [DOI] [PubMed] [Google Scholar]
- 5.Chatterjee, S. & Mudher, A. Alzheimer’s disease and type 2 diabetes: a critical assessment of the shared pathological traits. Front. Neurosci.12, 383 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Lee, H. J. et al. Diabetes and Alzheimer’s disease: mechanisms and nutritional aspects. Clin. Nutr. Res.7, 229 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Leibson, C. L. et al. The risk of dementia among persons with diabetes mellitus: a population-based cohort study. Ann. N. Y. Acad. Sci.826, 422–427 (1997). [DOI] [PubMed] [Google Scholar]
- 8.Santos, C. Y. et al. Pathophysiologic relationship between Alzheimer’s disease, cerebrovascular disease, and cardiovascular risk: a review and synthesis. Alzheimer's Dement.7, 69–87 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Tublin, J. M., Adelstein, J. M., Del Monte, F., Combs, C. K. & Wold, L. E. Getting to the heart of Alzheimer's disease. Circ. Res.124, 142–149 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Cenini, G. & Voos, W. Mitochondria as potential targets in Alzheimer disease therapy: an update. Front. Pharmacol.10, 902 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Onyango, I. G., Dennis, J. & Khan, S. M. Mitochondrial dysfunction in Alzheimer’s disease and the rationale for bioenergetics based therapies. Aging Dis.7, 201–214 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Turner, N., Cooney, G. J., Kraegen, E. W. & Bruce, C. R. Fatty acid metabolism, energy expenditure and insulin resistance in muscle. J. Endocrinol.220, T61–T79 (2014). [DOI] [PubMed] [Google Scholar]
- 13.Rinaldo, P., Cowan, T. M. & Matern, D. Acylcarnitine profile analysis. Genet. Med.10, 151–156 (2008). [DOI] [PubMed] [Google Scholar]
- 14.CDC. Using tandem mass spectrometry for metabolic disease screening among newborns. MMWR Recomm. Rep.50, 1–34. https://www.cdc.gov/mmwr/preview/mmwrhtml/rr5003a1.htm (2001). [PubMed]
- 15.Long, T. et al. Whole-genome sequencing identifies common-to-rare variants associated with human blood metabolites. Nat. Genet.49, 568–578 (2017). [DOI] [PubMed] [Google Scholar]
- 16.Shin, S. Y. et al. An atlas of genetic influences on human blood metabolites. Nat. Genet.46, 543–550 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Suhre, K. et al. Human metabolic individuality in biomedical and pharmaceutical research. Nature477, 54–60 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Draisma, H. H. M. et al. Genome-wide association study identifies novel genetic variants contributing to variation in blood metabolite levels. Nat. Commun.6, 7208 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Batchuluun, B. et al. Elevated medium-chain acylcarnitines are associated with gestational diabetes mellitus and early progression to type 2 diabetes and induce pancreatic β-cell dysfunction. Diabetes67, 885–897 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Schooneman, M. G., Vaz, F. M., Houten, S. M. & Soeters, M. R. Acylcarnitines: reflecting or inflicting insulin resistance? Diabetes62, 1–8 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Newgard, C. B. et al. A branched-chain amino acid-related metabolic signature that differentiates obese and lean humans and contributes to insulin resistance. Cell Metab.9, 311–326 (2009). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Jarrell, Z. R. et al. Plasma acylcarnitine levels increase with healthy aging. Aging12, 13555–13570 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Sun, N., Youle, R. J. & Finkel, T. The mitochondrial basis of aging. Mol. Cell61, 654–666 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Yu, Z. et al. Human serum metabolic profiles are age dependent. Aging Cell11, 960–967 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Huguenard, C. J. C. et al. APOE ε4 and Alzheimer’s disease diagnosis associated differences in L-carnitine, GBB, TMAO, and acylcarnitines in blood and brain. Curr. Res. Transl. Med.71, 103362 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Mapstone, M. et al. Plasma phospholipids identify antecedent memory impairment in older adults. Nat. Med.20, 415–418 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Casanova, R. et al. Blood metabolite markers of preclinical Alzheimer’s disease in two longitudinally followed cohorts of older individuals. Alzheimer's Dement.12, 815–822 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Ciavardelli, D. et al. Medium-chain plasma acylcarnitines, ketone levels, cognition, and gray matter volumes in healthy elderly, mildly cognitively impaired, or Alzheimer’s disease subjects. Neurobiol. Aging43, 1–12 (2016). [DOI] [PubMed] [Google Scholar]
- 29.Cristofano, A. et al. Serum levels of acyl-carnitines along the continuum from normal to Alzheimer’s Dementia. PLoS ONE11, e0155694 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Toledo, J. B. et al. Metabolic network failures in Alzheimer’s disease: a biochemical road map. Alzheimer’s Dement.13, 965–984 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Van Der Velpen, V. et al. Systemic and central nervous system metabolic alterations in Alzheimer’s disease. Alzheimers Res Ther.11, 1–12 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Arnold, M. et al. Sex and APOE ε4 genotype modify the Alzheimer’s disease serum metabolome. Nat. Commun.11, 1148 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Buyukozkan, M., Suhre, K. & Krumsiek, J. SGI: automatic clinical subgroup identification in omics datasets. Bioinformatics38, 573–576 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Jack, C. R. et al. NIA-AA Research Framework: toward a biological definition of Alzheimer’s disease. Alzheimer’s Dement.14, 535–562. 10.1016/j.jalz.2018.02.018 (2018). [DOI] [PMC free article] [PubMed]
- 35.Wolf, M. et al. Production and release of acylcarnitines by primary myotubes reflect the differences in fasting fat oxidation of the donors. J. Clin. Endocrinol. Metab.98, E1137–E1142 (2013). [DOI] [PubMed] [Google Scholar]
- 36.Lehmann, R. et al. Medium chain acylcarnitines dominate the metabolite pattern in humans under moderate intensity exercise and support lipid oxidation. PLoS ONE5, e11519 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Van Dyck, C. H. et al. Lecanemab in early Alzheimer’s disease. N. Engl. J. Med.388, 142–143 (2023). [DOI] [PubMed] [Google Scholar]
- 38.Gucciardi, A. et al. Analysis and interpretation of acylcarnitine profiles in dried blood spot and plasma of preterm and full-term newborns. Pediatr. Res.77, 36–47 (2015). [DOI] [PubMed] [Google Scholar]
- 39.Nho, K. et al. Association of altered liver enzymes with Alzheimer disease diagnosis, cognition, neuroimaging measures, and cerebrospinal fluid biomarkers. JAMA Netw. Open2, e197978 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Adams, S. H. et al. Plasma acylcarnitine profiles suggest incomplete long-chain fatty acid beta-oxidation and altered tricarboxylic acid cycle activity in type 2 diabetic African-American women. J. Nutr.139, 1073–1081 (2009). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Ramirez, A. et al. SUCLG2 identified as both a determinator of CSF Aβ1–42 levels and an attenuator of cognitive decline in Alzheimer’s disease. Hum. Mol. Genet.23, 6644–6658 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Arenaza-Urquijo, E. M. et al. The metabolic brain signature of cognitive resilience in the 80+: beyond Alzheimer pathologies. Brain142, 1134–1147 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Kepp, K. P., Robakis, N. K., Høilund-Carlsen, P. F., Sensi, S. L. & Vissel, B. The amyloid cascade hypothesis: an updated critical review. Brain146, 3969–3990 (2023). [DOI] [PubMed] [Google Scholar]
- 44.Granzotto, A. & Sensi, S. L. Once upon a time, the Amyloid Cascade Hypothesis. Ageing Res. Rev.93, 102161 (2024). [DOI] [PubMed] [Google Scholar]
- 45.Zhu, H. et al. Ketogenic diet for human diseases: the underlying mechanisms and potential for clinical implementations. Signal Transduct. Target Ther.7, 11 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Hawley, J. A., Hargreaves, M., Joyner, M. J. & Zierath, J. R. Integrative biology of exercise. Cell159, 738–749 (2014). [DOI] [PubMed] [Google Scholar]
- 47.Hargreaves, M. & Spriet, L. L. Skeletal muscle energy metabolism during exercise. Nat. Metab.2, 817–828 (2020). [DOI] [PubMed] [Google Scholar]
- 48.Kulkarni, A. S., Gubbi, S. & Barzilai, N. Benefits of metformin in attenuating the hallmarks of aging. Cell Metab.32, 15–30 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Yang, M. et al. Inhibition of mitochondrial function by metformin increases glucose uptake, glycolysis and GDF-15 release from intestinal cells. Sci. Rep.11, 2529 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Zakaria, R., Allen, K. J., Koplin, J. J., Roche, P. & Greaves, R. F. Advantages and challenges of dried blood spot analysis by mass spectrometry across the total testing process. EJIFCC27, 288 (2016). [PMC free article] [PubMed] [Google Scholar]
- 51.Bennett, D. A. et al. Religious orders study and rush memory and aging project. J. Alzheimer’s Dis.64, S161–S189. 10.3233/JAD-179939 (2018). [DOI] [PMC free article] [PubMed]
- 52.De Jager, P. L. et al. A genome-wide scan for common variants affecting the rate of age-related cognitive decline. Neurobiol. Aging33, 1017.e1–1017.e15 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Carrasquillo, M. M. et al. Genetic variation in PCDH11X is associated with susceptibility to late-onset Alzheimer’s disease. Nat. Genet.41, 192–198 (2009). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Auton, A. et al. A global reference for human genetic variation. Nature526, 68–74 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Delaneau, O., Zagury, J. F. & Marchini, J. Improved whole-chromosome phasing for disease and population genetic studies. Nat. Methods10, 5–6 (2013). [DOI] [PubMed] [Google Scholar]
- 56.Howie, B., Marchini, J. & Stephens, M. Genotype imputation with thousands of genomes. G3 Genes, Genomes, Genet.1, 457–470 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Holle, R., Happich, M., Löwel, H. & Wichmann, H. E. KORA–a research platform for population based health research. Gesundheitswesen67, S19–S25 (2005). [DOI] [PubMed] [Google Scholar]
- 58.Mittelstrass, K. et al. Discovery of sexual dimorphisms in metabolic and genetic biomarkers. PLoS Genet.7, e1002215 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Harris, T. B. et al. Age, gene/environment Susceptibility–Reykjavik Study: multidisciplinary applied phenomics. Am. J. Epidemiol.165, 1076–1087 (2007). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Wechsler, D. WAIS-R Manual: Wechsler Adult Intelligence Scale-Revised (Psychological Corporation, 1981).
- 61.Petersen, R. C. et al. Alzheimer’s Disease Neuroimaging Initiative (ADNI): clinical characterization. Neurology74, 201–209 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Kunkle, B. W. et al. Genetic meta-analysis of diagnosed Alzheimer’s disease identifies new risk loci and implicates Aβ, tau, immunity and lipid processing. Nat. Genet.51, 414–430 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Ward, J. H. Hierarchical grouping to optimize an objective function. J. Am. Stat. Assoc.58, 236–244 (1963). [Google Scholar]
- 64.Harrell, F. E. Multivariable modeling strategies. 63–102. 10.1007/978-3-319-19425-7_4 (2015).
- 65.DeMaris, A. Explained variance in logistic regression. Sociol. Methods Res.31, 27–74 (2002). [Google Scholar]
- 66.Hoffman, G. E. & Schadt, E. E. variancePartition: interpreting drivers of variation in complex gene expression studies. BMC Bioinform.17, 483 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Crane, P. K. et al. Development and assessment of a composite score for memory in the Alzheimer’s Disease Neuroimaging Initiative (ADNI). Brain Imaging Behav.6, 502–516 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Gibbons, L. E. et al. A composite score for executive functioning, validated in Alzheimer’s Disease Neuroimaging Initiative (ADNI) participants with baseline mild cognitive impairment. Brain Imaging Behav.6, 517–527 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Van Loenhoud, A. C. et al. Cognitive reserve and clinical progression in Alzheimer disease: a paradoxical relationship. Neurology93, E334–E346 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Fisher, R. A. Statistical Methods for Research Workers 11th ed. rev. https://psycnet.apa.org/record/1925-15003-000 (Oliver and Boyd, 1925).
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Description of Additional Supplementary Files
Data Availability Statement
KORA data can be accessed upon request at https://helmholtz-muenchen.managed-otrs.com/external/. AGES-RS data can be accessed upon request according to informed consent at https://hjarta.is/en/research/ages-phase-1/. All other omics datasets are available via the AD Knowledge Portal (https://adknowledgeportal.org). The AD Knowledge Portal is a platform for accessing data, analyses, and tools generated by the Accelerating Medicines Partnership (AMP-AD) Target Discovery Program and other National Institute on Aging (NIA)-supported programs to enable open-science practices and accelerate translational learning. The data, analyses, and tools are shared early in the research cycle without a publication embargo on secondary use. Data are available for general research use according to the following requirements for data access and data attribution (https://adknowledgeportal.org/DataAccess/Instructions). For access to the content described in this manuscript see: ADNI metabolomics data from the AbsoluteIDQ-p180 kit is available at the AD Knowledge Portal under 10.7303/syn5592519 (ADNI-1) and 10.7303/syn9705278 (ADNI-GO/-2), the full complement of clinical and demographic data for the ADNI cohorts are hosted on the LONI data sharing platform and can be requested at https://adni.loni.usc.edu/data-samples/adni-data/#AccessData. ROS/MAP imputed genotype data are available at the AD Knowledge Portal under 10.7303/syn3157329, study meta-data, basic covariates, and clinical variables are available under 10.7303/syn3157322. The full complement of clinical and demographic data for the ROS/MAP cohorts is hosted on the Rush Alzheimer’s Disease Center data-sharing platform and can be requested at www.radc.rush.edu. MayoLOAD genotyping data are available at the AD Knowledge Portal under 10.7303/syn3157238, study meta-data and covariates are available under 10.7303/syn3205821.6. Source data are provided with this paper.
All R code required to conduct the analyses reported here is available at https://github.com/compneurobio/bioenergetic_capacity.