Abstract
Anaphase promoting complex/cyclosome (APC/C)-mediated proteolysis is essential for chromosome segregation, mitotic exit, and G1 entry. Here, we show the importance of APC/C in the control of dTTP pool size in mammalian cells. Two enzymes, thymidine kinase 1 (TK1) and thymidylate kinase (TMPK), involved in dTTP formation are the targets of the APC/C pathway. We demonstrate that TMPK is recognized and degraded by APC/C–Cdc20/Cdh1-mediated pathways from mitosis to the early G1 phase, whereas TK1 is targeted for degradation by APC/C–Cdh1 after mitotic exit. Overexpression of wild-type TK1 and TMPK induces a four- to fivefold increase in the cellular dTTP pool without promoting spontaneous mutations in the hprt (hypoxanthine-guanine phosphoribosyl transferase) gene. In contrast, coexpression of nondegradable TK1 and TMPK expands the dTTP pool size 10-fold accompanied by a drastic dNTP pool imbalance. Most interestingly, disruption of APC/C proteolysis of TK1 and TMPK leads to growth retardation and a striking increase in gene mutation rate. We conclude that down-regulation of dTTP pool size by the APC/C pathway during mitosis and the G1 phase is an essential means to maintain a balanced dNTP pool and to avoid genetic instability.
Keywords: APC/C, cell cycle, dNTP pool imbalance, dTTP, genome stability
Cellular production of dTTP is a highly regulated process that is coordinated with DNA replication in the cell cycle (for review, see Reichard 1988). Using synchronized cultures, it has been shown that dTTP pool size in S-phase cells is 20-fold larger than that in G0-phase cells (Spyrou and Reichard 1988). This fluctuation can be attributed to cell cycle-dependent regulation of enzymes involved in dTTP formation (Navalgund et al. 1980; Bradshaw 1983; Sherley and Kelly 1988). There are two major pathways controlling dTTP synthesis in mammalian cells. In the de novo pathway, thymidylate synthetase (TS) catalyzes the rate-limiting step of converting dUMP to dTMP. Alternatively, thymidine kinase (TK), the key enzyme in the salvage pathway, catalyzes the transferal of the terminal phosphate of ATP to the 5′-hydroxyl group of thymidine to form dTMP. Subsequent phosphorylation of dTMP by thymidylate kinase (TMPK) gives rise to dTDP, which is then converted to dTTP by dNDP kinase (NDK) for DNA synthesis (for review, see Reichard 1988). Since TMPK is required for dTTP production in both salvage and de novo pathways, deletion of the TMPK gene is lethal in Saccharomyces cerevisiae (Sclafani and Fangman 1984; Su and Sclafani 1991). Presently, it is generally agreed that S-phase-specific transcriptional activation of TS, cytosolic TK1, and TMPK genes are the major mechanisms responsible for their up-regulation in S phase, thus stimulating dTTP pool expansion for DNA synthesis (Coppock and Pardee 1987; Sherley and Kelly 1988; Huang et al. 1994; DeGregori et al. 1995; Liang et al. 1995).
After completion of DNA replication, dTTP synthesis is no longer in great demand during the G2/M phase, and cellular dTTP drops to a low level until the mid-G1 phase of the next cell cycle. However, since TK1 and TS proteins still remain intact during the G2/M phase, it is of interest to know how the dTTP pool is down-regulated during this period. In regard to this, our laboratory has previously shown that human TK1 (hTK1) is phosphorylated on Ser13 in the mitotic phase (Chang et al. 1998), which by perturbing its active tertramerization status reduces its catalytic efficiency (Li et al. 2004). Upon mitotic exit, hTK1 is recognized by anaphase promoting complex/cyclosome (APC/C)–Cdh1 via its KEN box, which leads to APC/C-dependent degradation; hTK1 function is thus abolished at the entry to the G1 phase of the cell cycle (Ke and Chang 2004). Apparently, post-translational control by phosphorylation and APC/C-mediated proteolysis is involved in the down-regulation of the dTTP supply from the salvage pathway during mitotic progression. However, whether down-regulation of dTTP synthesis in mitosis is important in controlling cell growth remains an open question.
The APC/C ubiquitin ligase mediates degradation of cell cycle proteins during mitosis and G1 phase. It is well-known that Cdh1 and Cdc20 are two different activators that recognize substrates for APC/C-mediated proteolysis (for review, see Peters 2002). Cdc20 is transiently activated in mitosis to target cyclin B1 and securin for APC/C-mediated proteolysis through direct binding to the destruction box (D-box, RXXL) of these substrate proteins (Pfleger et al. 2001), whereas APC/C–Cdh1 is activated in mitotic exit and recognizes either a D- or KEN-box (KEN) (Pfleger and Kirschner 2000). To date, human Cdc6, Aurora-A kinase, Cdc20, Cdc25A, and Skp2 have been identified as the substrates of APC/C–Cdh1 in mammalian cells (Petersen et al. 2000; Donzelli et al. 2002; Littlepage and Ruderman 2002; Bashir et al. 2004; Wei et al. 2004). In addition to these cell cycle regulators, human TK1 and mouse RNR R2 (mrR2) are recognized by Cdh1 for APC/C-mediated proteolysis (Chabes et al. 2003a; Ke and Chang 2004). Accordingly, we hypothesized that the APC/C–Cdh1 pathway may be important in abrogating the dNTP supply at entry into G1 phase by simultaneously degrading rR2 and TK1. To test this hypothesis, here we defined the contribution of the APC/C-mediated pathway in controlling intracellular dTTP pool size.
In this study, we found that in addition to TK1, APC/C also targeted TMPK for degradation and demonstrated that disruption of APC/C-dependent proteolysis of both TK1 and TMPK led to a significant elevation of the dTTP pool in cells. Because dTTP can act as an allosteric activator and inhibitor of the reduction of rGDP and rCDP by ribonucleotide reductase (RNR), respectively (Eriksson et al. 1979; Reichard et al. 2000), elevation of the dTTP pool should also affect cellular levels of dGTP and dCTP. We therefore asked whether APC/C-dependent proteolysis of both hTK1 and hTMPK plays a role in regulation of a balanced supply of four dNTPs for DNA replication. Our results showed that expression of nondegradable TK1 and TMPK resulted in a decreased cell proliferation rate and a significant increase in the gene mutation rate, revealing a connection between a mitotic regulatory system and the important events that occur in the S phase.
Results
APC/C-mediated pathways regulate intracellular dTTP pool size
To determine the specific contribution of APC/C-mediated proteolysis to dTTP pool size, we depleted expression of Cdh1 and Cdc20 by siRNA experiments and analyzed the intracellular levels of dTTP. In comparison to the nontransfected cells as well as the cells transfected with two nonrelated siRNA, HeLa cells depleted of Cdh1 contained about four times the level of dTTP, accompanied by a significant increase in hTK1 protein, indicating that APC/C–Cdh1-mediated degradation of hTK1 does control the intracellular dTTP pool (Fig. 1A). Interestingly, silencing of Cdc20 expression by transfecting cells with siRNA of Cdc20 also increased cellular dTTP concentration without changing hTK1 protein level. Simultaneous knockdown of both Cdh1 and Cdc20 expression gave rise to a sevenfold increase in the dTTP pool in comparison to cells transfected with control siRNA (Fig. 1A). To know whether the observed increase of the dTTP pool was due to changes in the distribution of the population of cells in the cell cycle, we measured the cell cycle state by fluorescence-activated cell sorting (FACS) (Fig. 1B). The percentage of cells in G1, S, and G2/M phase was similar for cells transfected with siRNA of control and Cdc20, indicating that the increase of dTTP in cells depleted of Cdc20 is not due to the change of the cell cycle state. Although cells transfected with siRNAs of Cdh1 and Cdh1/Cdc20 had a higher percentage of cells in the S-phase fraction than the control cells, it is very unlikely that these modest differences in cell cycle distribution could cause the dramatic increases in the levels of TK1 expression and the dTTP pool. These experimental results not only suggest that APC/C–Cdh1 can control dTTP pool size through increasing accumulation of hTK1, but also imply that the APC/C–Cdc20-mediated pathway controls other regulator(s) involved in dTTP synthesis.
Figure 1.
Depletion of either Cdc20 or Cdh1 increases the intracellular dTTP pool size. (A) HeLa cells were transfected with siRNAs corresponding to nonrelevant firefly mRNA (C1 and C2) and Cdc20, Cdh1, or Cdh1 mRNA as indicated. (NT) No siRNA transfection; (Cdc20/Cdh1 siRNAs) combination of Cdc20 and Cdh1 siRNAs transfection. Forty-eight hours after transfection, cells were extracted for dTTP pool size determination and Western blotting with specific antibodies. Each data point represents a mean ± S.D. of three separate experiments. (B) The cell cycle profiles of siRNA-treated cells as indicated were analyzed by flow cytometry and the results were expressed as percentage of cells in G1, S, and G2/M phases.
Human TMPK is a direct target of both APC/C–Cdc20 and APC/C–Cdh1 complex
By analyzing the amino acid sequences of molecules involved in dTTP synthesis, including TMPK, NDK, TS, and dCMP deaminase, TMPK was found to contain several putative APC/C recognition sites. The two D-, one KEN-, and two D-like-boxes in the TMPK are all highly conserved throughout human, mouse, and rat (Supplementary Fig. 2), suggesting TMPK as a potential target for APC/C. To verify whether TMPK is a target of either Cdc20 or Cdh1, we tested the effect of overexpression of Cdc20 and Cdh1 on the expression of a Flag-tagged human TMPK (hTMPK) by cotransfecting an expression vector of FLAG-hTMPK with either pHA-Cdc20 or pHA-Cdh1 in HeLa cells. Overexpression of either Cdc20 or Cdh1 significantly suppressed the expression of FLAG-hTMPK in HeLa cells in a proteasome inhibitor-, LLnL, sensitive manner (Fig. 2A). This suggests that either Cdc20 or Cdh1 can act as a rate-limiting factor for proteasome-dependent degradation of hTMPK. Since enforced expression of Cdc20 and Cdh1 also enhanced in vivo ubiquitinylation of hTMPK (Supplementary Fig. 3), we then assessed whether Cdc20 and Cdh1 directly mediates ubiquitinylation of hTMPK by performing in vitro polyubiquitinylation assays with a reconstitution system containing purified APC/C, E1, and E2 (human UbcH10). As shown in Figure 2B, GST-hTMPK was polyubiquitinylated in a reaction mixture of purified E1, E2, and APC/C in the presence of either Cdc20 or Cdh1 (lanes 3,5). Inclusion of dominant-negative C114S mutant of UbcH10 prevented polyubiquitinylation of hTMPK in this system (Fig. 2B, lanes 4,6), indicating the specificity of the ubiquitinylation reaction. Similar to human UbcH10, the Xenopus UbcX also gave a similar pattern of ubiquitinylated ladders derived from hTMPK (Fig. 2B, lanes 7,8). Unlike hTMPK, polyubiquitinylation of GST-hTK1 was dependent on APC/C–Cdh1 but not APC/C–Cdc20 (Fig. 2B, lane 10 vs. lane 11). Based on these data, we concluded that both APC/C–Cdc20 and APC/C–Cdh1 are able to directly ubiquitinylate hTMPK and that hTK1 is only recognized by APC/C–Cdh1 and not APC/C–Cdc20.
Figure 2.
hTMPK is a novel substrate of APC/C–Cdc20 and APC/C–Cdh1. (A) HeLa cells were transfected with pCMV2FLAG-hTMPK and pEGFP with different amounts of either pHA-Cdc20 or pHA-Cdh1 as indicated. Empty vector was added to the transfection mixture to adjust the total amount of DNA for each transfection to be equal. Cells were treated without (–) or with (+) LLnL for 6 h before harvesting. Followed by transfection for 24 h, cells were extracted for Western blotting analysis. Expression of GFP indicated the transfection efficiency among different samples. (B) In vitro polyubiquitinylation was performed using GST-hTMPK or GST-hTK1 as the substrate in a reaction mixture containing E1, E2 (UbcX or UbCH10), APC/C, Cdc20, or Cdh1 as indicated. The CS represents the C114S mutant of Ubc10. The samples were separated by SDS-PAGE (5%–15% gradient) and analyzed by Western blotting using anti-GST antibody as indicated. The asterisk indicates the dimmer forms of hTK1 and hTMPK.
hTMPK precedes hTK1 for APC/C-mediated degradation during mitotic progression
To analyze endogenous hTMPK expression, we generated a specific hTMPK antibody. Using this antibody, we examined the cell cycle oscillation of hTMPK levels in HeLa S3 cells, which were synchronized at mitosis by nocodazole and released into G1. The level of hTMPK stayed very low in mitotic arrest and early G1 cells and did not rise until the mid-G1 phase (Fig. 3A). In contrast, the expression of hTK1 reached its maximal peak in the mitotic arrested stage, with expression diminishing abruptly in the early G1 phase, demonstrating that degradation of hTMPK occurs earlier than hTK1 during cell cycle progression (Fig. 3A). The cell cycle-dependent expression patterns of hCdh1 and other known APC/C substrates, including Cdc20, cyclin A 1, cyclin B1, hrR2, and securin, were also examined in the same set of samples. As a direct test of APC/C-mediated degradation of hTMPK and hTK1, we prepared mitotic and G1 extracts to perform in vitro degradation assays. We found that 35S-methionine-labeled hTMPK was destroyed in both mitotic and G1 extracts, while 35S-methionine-labeled hTK1 was degraded only in the G1 extract (Fig. 3B). The in vitro degradation of hTMPK in the mitotic extract was APC/C-specific because it was inhibited by addition of GST-(1–90) cyclin B1, which is a specific competitive APC inhibitor, but not by GST-cyclin B1 (ΔN90) devoid of D-box (Fig. 3C). In G1 extract, the wild-type GST-hTK1 but not hTK1 with a KEN-box mutation competed for hTMPK degradation, indicating the requirement of APC/C–Cdh1 for hTMPK degradation in G1 phase (Fig. 3D). Since APC/C–Cdc20 is only active in the mitotic phase and APC/C–Cdh1 is not activated until the early G1 phase, the results from these in vitro degradation experiments suggest that APC/C–Cdc20 and APC/C–Cdh1 target hTMPK for degradation in the mitotic and G1 phase, respectively. To confirm the involvement of Cdc20 and Cdh1 in destabilization of hTMPK in vivo, we then examined the effect of Cdc20, Cdh1, and Cdc20/Cdh1 knockdown on stability of hTMPK in HeLa cells. To that end, we used 35S-methionine to pulse-label cells that were transfected with control, Cdc20, Cdh1, or Cdc20/Cdh1 siRNAs, followed by chasing for 12 h. The lysates of 35S-methionine-labeled cells were immunoprecipitated by hTMPK antibody. The results showed that after the 12-h chase period, <10% of prelabeled hTMPK remained in control siRNA transfected cells, while >80% of prelabeled hTMPK was still present in cells transfected with siRNAs of Cdc20 or Cdh1 (Fig. 3E). Codepletion of Cdc20 and Cdh1 caused no decay of prelabeled TMPK during the 12-h chase. These results indicate that the half-life of hTMPK was significantly prolonged by depletion of either Cdc20 or Cdh1. In a parallel pulse-chase experiment, we found that depletion of Cdh1 but not of Cdc20 stabilized the pulse-labeled hTK1, consistently indicating the specific contribution of Cdh1 to hTK1 destabilization (Fig. 3E). Thus, silencing of Cdc20, Cdh1, or Cdc20/Cdh1 expression renders hTMPK more stabilized, thereby increasing its expression level. Taking these results together, we conclude that Cdc20 targets hTMPK for degradation in the G2/M phase, and Cdh1 maintains its destruction and targets TK1 for proteolysis in the early G1 phase.
Figure 3.
hTMPK is degraded in M and early G1 phase. (A) HeLa S3 cells were arrested by nocodazole treatment for 20 h, followed by replenishment with fresh medium to release cells from the G2/M arrest. Extracts prepared from HeLa cells released from G2/M arrest at various time points as indicated were analyzed by Western blotting with the specific antibodies. (Asn) Asynchronized state. (B) S35-methionine-labeled Myc-hTMPK was incubated with M- and G1-phase extracts as described in Materials and Methods for different times as indicated. Purified nondegradable cyclin B1 (ΔN90 cyclin B1) and hTK1 proteins were added in each extract to indicate the specificity of degradation in the assay. (C) Purified GST-tagged (1–90) cyclin B1 and (ΔN90) cyclin B1 were each added to the M extract for in vitro degradation assay of S35-methionine-labeled Myc-hTMPK. (D) Purified GST-tagged wild-type and KEN mt hTK1 were each added to the G1 extract for in vitro degradation assay of S35-methionine-labeled Myc-hTMPK. (E) HeLa cells were transfected with control, Cdc20, and Cdh1 siRNAs as indicated. (Cdc20/Cdh1) Combination of Cdc20 and Cdh1 siRNA transfection. Thirty-six hours after transfection, cells were metabolically labeled with 35S-methionine (250 μCi/mL) for 30 min, followed by chasing for 0 and 12 h. Cell lysates were immunoprecipitated by anti-hTK1 and anti-hTMPK antibodies separately. The immunocomplexes were resolved on 12% SDS-PAGE, followed by Western blotting and autoradiography. Normal rabbit IgG (IgG) was used a negative control for immunoprecipitation.
APC/C regulates the dTTP pool size through degradation of hTK1 and hTMPK
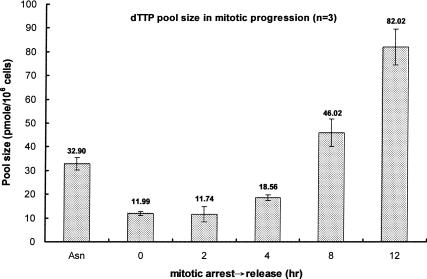
Because TMPK is required for dTTP formation from both salvage and de novo pathways, earlier degradation of TMPK than of TK1 may provide a means to minimize the dTTP supply in the mitotic phase. We then determined the dTTP levels in HeLa cells that were blocked in the mitotic phase by nocodazole treatment and released from the mitotic block. The level of dTTP was lowest in mitotic blocked cells, in which the expressed level of TMPK was also very low. Following release from nocodazole treatment, the dTTP pool rose as cells approached mid-G1 phase (Fig. 4), at which time hTMPK expression increased and hTK1 expression levels were still very low. Thus, the oscillation of TMPK expression levels during mitotic progression correlates with the fluctuation of the dTTP pool.
Figure 4.
Fluctuation of dTTP pool size during mitotic progression. HeLa S3 cells were arrested by nocodazole as described in the legend for Figure 3A. At different time points after release from mitotic block, cells were extracted for dTTP pool size determination as described in Materials and Methods. Each data point represents a mean ± S.D. of three separate experiments. (Asn) Asynchronized state.
We also silenced TMPK expression by siRNAs to determine whether the drastic expansion of dTTP pool size by depletion of Cdc20/Cdh1 in HeLa cells is dependent on TMPK. Clearly, removal of endogenous hTMPK expression abolished the Cdc20/Cdh1 siRNAs-mediated increase of the dTTP pool (Fig. 5). Depletion of hTK1 in Cdc20/Cdh1 knockdown cells also reduced the dTTP pool, albeit to a lesser degree as compared with hTMPK depletion. This result also suggests that APC/C–Cdh1-mediated degradation of TK1 contributes to a great proportion of the stimulatory effect on the dTTP pool by depletion of Cdc20/Cdh1 in HeLa cells. Western blot analysis showed that the expression levels of hTMPK and hTK1 were significantly increased by depletion of Cdc20/Cdh1, and cotransfection with siRNAs of hTMPK or hTK1 efficiently abolished each protein expression correspondingly (Fig. 5). These results provide direct evidence that hTK1 and hTMPK are responsible for up-regulation of the dTTP pool induced by Cdc20 and Cdh1 depletion in HeLa cells.
Figure 5.
TMPK is required for increment of the dTTP pool induced by inactivation of APC/C. HeLa cells were transfected with siRNAs as indicated. Forty-eight hours after transfection, cells were extracted for determination of dTTP pool size and Western blotting analysis.
The signal motifs for APC/C-mediated degradation of hTMPK
We next determined the signal motif responsible for targeting hTMPK for proteolysis. Each motif, including two D-box (RXXL), one KEN-box, and two D-box-like (RXXF), in hTMPK was mutated as schematically illustrated in Figure 6A. Since Cdh1 and Cdc20 is the rate-limiting factor for hTMPK degradation, we then examined the expression levels of each mutant protein in response to coexpression of Cdc20 and Cdh1. It appeared that mutation of the second D-box (D2 mt) rendered hTMPK resistant to Cdh1- or Cdc20-mediated destabilization (Fig. 6B). In addition, the mutant carrying mutations in the KEN-box (KEN mt) lost Cdh1 responsiveness, and also became less sensitive to Cdc20, while mutations to either of the D-box like motifs did not alter the sensitivity of hTMPK to Cdc20- or Cdh1-mediated destabilization. Mutation to the first D-box (D1) had no effect on Cdc20- or Cdh1-mediated degradation of TMPK (Fig. 6B). These results suggest that Cdc20 probably targets hTMPK for degradation by recognizing D-box at the D2 site, whereas Cdh1-mediated degradation of hTMPK requires the intactness of both the D2 and KEN motifs. In support of this notion, we found evidence that an intact D-box in the D2 site was required for GST-hTMPK bound to His-Cdh1 or His-Cdc20, and the KEN-box was additionally needed for binding to His-Cdh1 (Supplementary Fig. 4). On this basis, we then tested whether mutations in D2 or KEN sites affect APC/C–Cdh1- or APC/C–Cdc20-mediated polyubiquitinylation of GST-hTMPK protein by using a purified in vitro ubiquitinylation system. Consistently, polyubiquitinylated conjugates at higher molecular weight were seen for wild-type GST-hTMPK protein when APC/C–Cdc20 or APC/C–Cdh1 was included in the assay (Fig. 6D). For the D2 and D2/KEN mutants, no polyubiquitinylated conjugates was observed. For KEN mutant, polyubiquitinylation ladders could be seen in the reaction containing APC/C–Cdc20 but not APC/C–Cdh1. It seems that mutations of KEN-box impaired the hTMPK protein to be polyubiquitinylated by APC/C–Cdh1-mediated E3 reaction without affecting the Cdc20-mediated reaction. Considering the fact that KEN mutation impairs the functional activity of hTMPK (data not shown), it is possible that mutation of the KEN site may have resulted in a conformational change to hTMPK, which prevents the D-box from being recognized by Cdh1. Nonetheless, these results suggest that the D-box (D2 site) of hTMPK protein is a necessary motif for recognition by APC/C–Cdc20 and APC/C–Cdh1.
Figure 6.
D-box-dependent recognition of hTMPK by Cdc20 and Cdh1. (A) Schematic representation of mutants carrying mutation in various potential destruction signal motifs of hTMPK as indicated. (B) Cell extracts were prepared from LM-TK– cells that were transfected with wild-type (wt) or various mutants of pCMV2FLAG-hTMPK as indicated with (+) or without (–) pHA-Cdh1 or pHA-Cdc20, followed by SDS-PAGE and Western blotting analysis as described in the legend for Figure 2A. (C) In vitro polyubiquitinylation reactions of wild-type, D2 mt, KEN mt, and D2/KEN mt of hTMPK were performed as described in the legend for Figure 2B. The asterisk indicates the dimer forms of hTK1 and hTMPK.
Disruption of APC/C-dependent degradation of TK1 and TMPK induces a drastic dNTP imbalance
To further understand the effect of specific disruption of hTK1 and hTMPK degradation on dTTP pool size in a normal cell line, NIH 3T3 cells stably expressing similar levels of wild-type, loss-of-function, or APC/C resistant mutants of hTK1 and hTMPK were individually selected and pooled. Cells expressing wild-type hTK1 or hTMPK each gave a threefold rise in the dTTP pool relative to the mock cells, whereas expression of an ATP-binding-defective mutant of hTMPK (D15R) or thymidine-binding-defective mutant of hTK1 (RFN) did not alter the level of the dTTP pool (Fig. 7A). These results indicate that functional activities of both enzymes are the limiting factors for dTTP supply in NIH 3T3 cells. Expression of a nondegradable KEN mutant of hTK1 or D2 mutant of hTMPK led to a 30%–40% increase in the dTTP pool compared with their wild-type proteins (Fig. 7A). These moderate increases in dTTP could be due to low levels of endogenous hTK1 limiting dTMP supply in cells carrying overexpressed D2 mutant of hTMPK. Reciprocally, the level of endogenous TMPK may also have limited the elevation of the dTTP pool in NIH 3T3 cells overexpressing stabilized hTK1. If so, the effect of disrupting hTK1 and hTMPK degradation on the dTTP pool should be greater in cells overexpressing both hTK1 and hTMPK. Indeed, while cells overexpressing wild-type hTK1 and hTMPK (WT/WT) exhibited a fivefold increase in the dTTP pool, coexpression of nondegradable hTK1and hTMPK (KEN/D2) led to a >10-fold elevation in the dTTP pool (Fig. 7A). Given that both WT/WT and KEN/D2 cell lines expressed similar steady-state levels of hTK1 and hTMPK and that the enzymatic catalytic efficiencies of hTK1 (KEN) and hTMPK (D2) are similar to their wild-type proteins (data not shown), we attribute the significant difference in dTTP pool size between these two cell lines to the APC/C-mediated degradation capacity of hTK1 and hTMPK.
Figure 7.
Simultaneous disruption of hTK1 and hTMPK degradation elevates the dTTP pool size. (A) NIH 3T3 stable line cells were established as described in Materials and Methods. Cells were extracted for dTTP pool size measurement and Western blotting analysis as described in the legend for Figure 1. (B) NIH 3T3 cells stably expressing wild-type (WT/WT) and nondegradable mutant (KEN/D2) hTK1 and hTMPK were serum deprived and stimulated as indicated. (Asn) Asynchronized state; (SD) serum deprivation 24 h; (SD → SS) serum stimulation for 24 h after 24-h serum deprivation. Cells were harvested for Western blotting analysis and dTTP pool size determination as described in the legend for Figure 5. (C) Four dNTP pool sizes were determined in extracts from various 3T3 stable line cells (Mock, WT/WT, and KEN/D2) as described in Materials and Methods. Each data point represents a mean ± S.D. of three separate experiments. (D) The ratio of dGTP to dCTP in each cell line was calculated by dividing pool size of dGTP by that of dCTP. The dNTP pool imbalance induction folds were expressed by normalization of the dGTP/dCTP ratio of WT/WT and KEN/D2 cells with that of Mock cells.
Since KEN/D2 and WT/WT cells expressed similar steady-state levels of hTK1 and hTMPK, we then tested whether the expressed levels of these two proteins in these two cell lines were different in a growth-dependent manner, thus resulting in differences of dTTP accumulated in cells. For this purpose, we examined the expressed levels of hTK1, and hTMPK and the dTTP pool size in WT/WT and KEN/D2 cells under serum-deprivation and serum-stimulation conditions. For WT/WT cells, we observed down-regulation of hTK1, hTMPK, and the dTTP pool after serum deprivation; following serum stimulation, expression levels of both hTK1 and hTMPK were restored, accompanied by a four- to fivefold increase of the dTTP pool. As a contrast in KEN/D2 cells, levels of hTK1, hTMPK, and the dTTP pool remained elevated regardless of serum deprivation. Thus, the increase in dTTP accumulation in KEN/D2 cells correlated with the stabilized nature of hTK1 and hTMPK present in the cells.
We also measured the levels of three other dNTPs in WT/WT and KEN/D2 cells. In WT/WT cells, the dGTP pool was fourfold higher than that in the mock cells, while there were no significant differences in dATP and dCTP pools (Fig. 7C). In KEN/D2 cells, the dGTP pool size was expanded 12-fold as compared with that in the mock cells, while dATP was slightly increased and dCTP was decreased by 50% in comparison to the mock cells (Fig. 7C). Thus, a severe imbalance in dNTP pools was induced in KEN/D2 cells. If one considers the dGTP/dCTP ratio as a dNTP pool imbalance index, it is 3, 12, and 0.6 for cells expressing wild-type, nondegradable form, and the mock cells, respectively (Fig. 7D). Accordingly, the degree of dNTP pool imbalance induction is fivefold and 20-fold in cells expressing wild-type and nondegradable hTK1 and hTMPK, respectively. Thus, a fourfold augmentation of dNTP pool imbalance has been induced by disruption of mitotic degradation of TK1 and TMPK.
Analogous to this result, in HeLa cells depleted of Cdc20/Cdh1, both dGTP and dTTP pools were dramatically increased, accompanied by a significant decrease in the dCTP pool as compared with the cells transfected with the control siRNA (Fig. 8). Clearly, interference of mitotic degradation of endogenous hTK1 and hTMPK by depletion of Cdc20/Cdh1 caused a more pronounced effect on dNTP pool imbalance than what we observed in KEN/D2 cells; the dGTP pool was increased 28-fold while the dCTP pool decreased sevenfold, corresponding to an ∼200-fold induction of dNTP pool imbalance (Fig. 8). Thus, one can expect that deregulation of the APC/C pathway in a human cancer cell line containing elevated levels of hTK1 and hTMPK would cause a very severe induction of dNTP pool imbalance.
Figure 8.
Knockdown of Cdc20/Cdh1 in HeLa cells induces dNTP pool imbalance. HeLa cells were transfected with siRNAs against Cdc20 and Cdh1 simultaneously as described in the legend for Figure 1. Cells were extracted for the four dNTP pool size determinations as described in Materials and Methods. Each data point represents a mean ± S.D. of three separate experiments. (Control) Control siRNA transfection.
Stabilization of hTK1 and hTMPK suppresses cell growth and confers increased genetic instability
We then further compared the growth rates of WT/WT, KEN/D2, and the mock cells (Fig. 9A). KEN/D2 cells exhibited a much slower growth rate than the mock and WT/WT cells, whereas WT/WT cells revealed a faster growth rate. This indicates that the extent of dNTP pool changes observed in WT/WT cells indeed favors cell growth, and the degree of dNTP pool imbalance in KEN/D2 cells causes growth retardation. Given that dNTP pool imbalance is considered mutagenic (for review, see Kunz 1988; Meuth 1989), we further investigated whether dNTP pool imbalance induced by deregulation of hTK1 and hTMPK would promote genetic instability by measuring the spontaneous mutation rate of the hprt gene. Cultured cells (3 × 105) were incubated in medium containing 6-thioguanine (6-TG), by which the occurrence of resistant clones represents spontaneous mutations at the hprt locus. In parallel, cells (1 × 103) were incubated in medium in the absence of 6-TG to measure their colony formation efficiencies. Consistent with the observation in the growth rate measurement, cells expressing KEN/D2 exhibited the lowest colony formation efficiency (Fig. 9B). Upon 6-TG selection, the mock and WT/WT cells did not develop any resistant colonies, indicating that the extent of dNTP pool changes in WT/WT cells is insufficient to induce genetic instability, as revealed by the maintenance of hprt gene function. In contrast, two different clones of the KEN/D2 cells, despite poor cloning efficiencies, yielded >500 resistant colonies per dish under the same experimental conditions (Fig. 9B). RNA was prepared from 50 individual 6-TG resistant clones, and subjected to RT–PCR with the primers for the hprt gene followed by sequence analysis. Various types of mutations, including transition, insertion, deletion, and transversion, were found in the hprt coding sequences in these clones (Supplementary Table 1). Clearly, high frequencies of genetic mutations were observed in KEN/D2 cells. Similar results were also observed in experiments when Chinese hamster lung fibroblast (CHL) V79 cells were used to establish stable cell lines (Supplementary Fig. 5). Because the difference in mutation rate was so significant between WT/WT and KEN/D2 cells, this suggests that not only does APC/C-dependent degradation of TK1 and TMPK play an important role in reducing the dTTP pool, it also prevents dNTP pool imbalance, leading to genetic instability.
Figure 9.
Disruption of APC/C-mediated degradation of hTK1/hTMPK affects growth rate and induces genetic instability. (A) Growth analysis of various 3T3 stable line cells (Mock, WT/WT, and KEN/D2) was performed as described in Materials and Methods. Results shown here represent the average ± S.D. from three separate experiments. (B) Various 3T3 stable line (Mock, WT/WT, and KEN/D2) cells (3 × 105) were plated onto the culture dish in 6-TG (5 μg/mL) containing medium (+) for analyzing spontaneous mutation in the hprt locus. For comparing the colony formation efficiency for each line, 103 cells were in parallel grown in the medium without 6-TG (–).
Discussion
Research into the essential role of APC/C-dependent proteolysis in cell cycle progression has been mainly focused on its function in sister chromatid separation and mitotic exit through degradation of substrates, such as securin and mitotic cyclins (for review, see Harper et al. 2002). In this study, our findings demonstrated another important role of APC/C: the regulation of dTTP pool size through degradation of TK1 and TMPK during mitotic progression. We showed that this control mechanism is necessary for maintaining a balanced dNTP pool while its deregulation may lead to growth retardation and a marked increase in genetic mutation rate, a major cause of genetic instability.
The precise timing for degradation of hTK1 and hTMPK warrants down-regulation of dTTP during mitotic progression
Our previous study had shown that hTK1 is a substrate for APC/C–Cdh1, but not APC/C–Cdc20 E3 ligase (Ke and Chang 2004). Since TK1 is a rate-limiting enzyme for dTTP formation in the salvage pathway, knockdown of Cdh1 in this study resulted in a significant increase of dTTP in cells. However, it was surprising to observe that knockdown of Cdc20 also increased the dTTP pool in cells (Fig. 1A). Our results showed that APC/C–Cdc20 and APC/C–Cdh1 can target hTMPK for degradation by recognizing its D-box, the D2 site (Fig. 6). Given that TMPK is also necessary for dTTP formation from TS-mediated de novo pathway and that TS is unlikely to be an APC/C target because of lack of a destruction motif in its amino acid sequences, APC/C-dependent degradation of TMPK would, therefore, be important in controlling dTTP formation from both salvage and de novo pathways. In support of this view, our data showed that silencing TMPK expression completely abolished the increase in the dTTP pool induced by Cdc20/Cdh1 knockdown (Fig. 5).
Data from in vitro and in vivo degradation experiments demonstrated that hTMPK is degraded in M to early G1 phase, preceding the destruction of hTK1 (Fig. 3). It was already known that binding of Cdc20 to APC/C, triggering its activation, only takes place during mitosis from metaphase until telophase and is followed by Cdh1-mediated activation of APC/C (for review, see Peters 2002). Our observation that Cdc20 targets TMPK but not TK1 for APC/C-mediated degradation would explain the differences in expression levels of TMPK and TK1 in mitotic blocked cells, in which TMPK expression diminished while high levels of TK1 were still present. Theoretically, earlier degradation of TMPK ensures a low supply of dTTP supply in mitosis, at which time TK1 remains intact though less active. Also note that the presence of Cdh1-mediated proteolysis maintains destruction of TMPK in the early G1 phase. Therefore, the combinatorial activities of APC/C–Cdc20 and APC/C–Cdh1 ensure TMPK destruction throughout the entire period of mitotic progression. In agreement with this molecular control of TMPK degradation, we found that in HeLa cells, dTTP levels were kept very low in mitotic arrest until the mid-G1 phase of the next cycle, which coincided with the oscillation pattern of TMPK expression (Fig. 4). Of note, although APC/C–Cdh1 targets both TMPK and TK1 for degradation in the G1 phase, the levels of TMPK protein expression rise faster than those of TK1 after release from mitotic block. We speculate that this is because the levels of TMPK RNA increased in the mid-G1 phase while TK1 transcript level did not increase until the late G1 phase (data not shown).
Disrupting APC/C-mediated proteolysis of TK1 and TMPK leads to amplification of dNTP pool imbalance
Knowing that the D2 site of TMPK and the KEN-box of TK1 are the necessary motifs for APC/C targeting (Fig. 6), we established cell lines coexpressing wild-type (WT/WT) and nondegradable (KEN/D2) hTK1 and hTMPK at similar steady-state levels. As expected, KEN/D2 cells contained a much higher level of dTTP than WT/WT cells. Since serum deprivation significantly reduced expression levels of hTK1 and hTMPK and dTTP pool size in WT/WT but not in KEN/D2 cells, loss of mitotic degradation of hTK1 and hTMPK should be the main reason for greater amounts of the dTTP pool accumulated in KEN/D2 cells (Fig. 7B). In agreement with the fact that dTTP can allosterically regulate rGDP and rCDP substrate specificity of ribonucleotide reductase (RNR) (Eriksson et al. 1979; Reichard et al. 2000), the dNTP pool imbalance was largely amplified in KEN/D2 cells because of the greater extent of increase in dGTP pool size and the decrease in the dCTP pool. Since the expression levels of hTK1 and hTMPK in WT/WT and KEN/D2 cells were very similar to those observed in HeLaS3, K562, and HEK293T cells (data not shown), it is unlikely that the dNTP pool imbalance observed in KEN/D2 cells was a result of too much hTK1 and hTMPK being expressed in a nonphysiological range. Of note, depletion of Cdc20/Cdh1 in HeLa cells gave rise to an even greater dNTP pool imbalance due to much larger extents of dCTP depletion and dGTP increment (Fig. 8), further reinforcing the idea that down-regulation of dTTP by APC/C function is necessary for maintaining a balanced dNTP pool.
The importance of APC/C-mediated proteolysis of TK1 and TMPK in cell proliferation and genome integrity
The augmented dNTP pool imbalance seen in cells coexpressing nondegradable hTK1 and hTMPK clearly led to growth retardation. From analysis of the cell cycle profile, KEN/D2 cells displayed a pronounced delay in S → G2/M transition (data not shown), an indication of S-phase arrest. It has been well established that treatment of cells with high concentrations of thymidine stalls the DNA replication fork due to dCTP depletion (Studzinski and Lambert 1969; Bjursell and Reichard 1973; Bianchi et al. 1987). Therefore, it is likely that in the absence of APC/C control, the dTTP pool is elevated to a level mimicking that resulting from thymidine treatment, leading to slowing of DNA synthesis due to a significant reduction of the dCTP pool in these cells. In contrast, a faster growth rate was observed in cells coexpressing both wild-type hTK1 and hTMPK in which dTTP and dGTP pools were elevated and the dCTP pool remained similar to that in the mock cells. This result supports the notion that the slowed-down progression of the DNA replication fork resulting from elevation of dTTP requires depletion of dCTP (Bjursell and Reichard 1973; Bianchi et al. 1987). In addition, it also suggests that overexpression of degradable TK1 and TMPK produces dTTP to a range that would not deplete dCTP but enhances the rate of DNA synthesis, thereby promoting the cell growth.
In this study we also determined the effect of disrupting APC/C-mediated dTTP control on genetic stability by analyzing the spontaneous mutations in the hprt locus. Cells coexpressing nondegradable hTK1 and hTMPK gave rise to a striking increase of 6-TG resistant colonies as compared with no 6-TG resistant colony formation from cells coexpressing wild-type hTK1 and hTMPK under the same experimental condition (Fig. 9B). The tremendous difference in the mutation rate between KEN/D2 and WT/WT cells was surprising. We repeated the same experiment with CHL V79 cells and obtained virtually the same results (Supplementary Fig. 5). These data indicate that the dNTP pool imbalance resulting from disruption of APC/C degradation of TK1 and TMPK impairs the fidelity of DNA synthesis, possibly leading to DNA replication error continuing to accumulate during a slow growth cycle. In this regard, several recent reports have shown that mismatch repair and homologous recombination repair are the DNA repair mechanisms involved in reactivation of stalled replication forks due to dCTP depletion induced by thymidine treatment (Lundin et al. 2002; Mohindra et al. 2002, 2004; Bolderson et al. 2004). Since there was a dramatic increase of mutation frequencies in cells expressing nondegradable hTK1 and hTMPK and various types of mutation in hrpt were found in these 6-TG resistant clones, it is likely that the (dGTP + dTTP)/dCTP change in these cells upset the fidelity of DNA synthesis to a degree that was beyond the capacity of these two DNA repair systems.
Using yeast, it has been demonstrated that disruption of feedback inhibition of RNR leads to an increase in the level of dNTPs and significantly promotes cell survival after DNA damage, while the gene mutation rate in those surviving cells is clearly increased (Chabes et al. 2003b). Thus, cells containing elevated levels of the dNTP pool survive from DNA damage, but pay the price with loss of fidelity of DNA synthesis, resulting in genetic abnormalities. This study suggests that nucleotide pool regulation by RNR in yeast may be important in maintaining genomic integrity. Our data, however, also suggest that the balance of the dNTP pool in mammalian cells can be guarded by APC/C control through degradation of TK1 and TMPK. Without this control to down-regulate the dTTP pool during mitotic progression, a severe dNTP pool imbalance is generated in the next cell cycle that drives genetic changes leading to genetic instability, a characteristic of cancers. Herein, we propose that APC/C-mediated TK1 and TMPK degradation plays a vital role in securing dNTP pool balance during mitotic progression, which is essential for the maintenance of genomic integrity.
Materials and methods
Materials
Antiserum against hTMPK was produced by immunizing rabbits with purified GST-hTMPK protein and collected, after which polyclonal antibody was affinity-purified. Specificity of purified hTMPK antibody was verified by antigen competition assay (Supplementary Fig. 1). Anti-hTK1 polyclonal antibody was prepared as described previously (Chang et al. 1994). Anti-β-tubulin, anti-Flag, and anti-Cdc27 antibodies were obtained from Sigma. Anti-Cdc20, anti-securin, anti-hrR2, anti-cyclin B1, and anti-myc antibodies were purchased from Santa Cruz Biotechnology. Anti-HA was obtained from Roche. Anti-GFP and anti-cyclin A1 antibodies were purchased from Clontech and Transduction Laboratories, respectively. Anti-Cdh1 and anti-GST antibodies were obtained from Oncogene and Amersham Pharmacia, respectively. G418 and zeocin were purchased from Invitrogene. Nocodazole and 6-thioguanine were purchased from Sigma. LLnL was obtained from Calbiochem.
Construction of expression plasmids and site-directed mutagenesis
pCDNA3.1Zeo-hTMPK was generated by inserting the BamHI fragment of pGEX-2T-hTMPK (provided by Jin-Yuan Su, National Yang Ming University, Taiwan) into plasmid pCDNA3.1Zeo. The DNA fragment containing hTMPK cDNA was obtained by PCR amplification using pGEX-2T-hTMPK as a template and was subsequently subcloned into the HindIII site of plasmid pCMV2FLAG, producing pCMV2FLAG-hTMPK. Mutations of hTMPK were generated using Quick-Change Site-directed Mutagenesis kit (Stratagene) by specific mutated primers. Construction of hTK1 expression plasmid was described previously (Ke and Chang 2004). pHA-Cdc20 and pHA-Cdh1 were provided by Kristian Helin (European Institute of Oncology, Milan, Italy). Mouse E1 and UbcX expression plasmids were provided by Tim Hunt (ICRF Clare Hall Laboratory, South Mimms, United Kingdom). The human Ubc10 plasmids, including wild-type and C114S mutant, were provided by Joan V. Ruderman (Harvard Medical School, Boston, MA). pCS2-Myc-Cdc20 and pCS2-MycCdh1 were provided by Marc W. Kirschner (Harvard Medical School, Boston, MA). The baculovirus-expressed pFASTBacHis-Cdc20 and pFASTBacHis-Cdh1 were provided by Jan-Michael Peters (Research Institute of Molecular Biology, Vienna, Austria).
Cell culture, synchronization, transfection, growth analysis, and FACS analysis
HeLa, HeLa S3, LM-TK–, and NIH 3T3 cells were maintained in Dulbecco's modified Eagle's medium (DMEM, Life Technologies) supplemented with 10% fetal bovine serum plus 100 μg/mL streptomycin and 100 U/mL penicillin (Life Technologies) at 37°C under 5% CO2. For G2/M arrest, nocodazole was added to subconfluent HeLa S3 cells at a final concentration of 0.5 μg/mL for 20 h. To obtain early G1-phase cells, mitotic arrested cells were shaken off, washed with PBS, and incubated in fresh medium for an additional 3–5 h. For ectopic expression experiments, cells plated on a 60-mm dish were transfected with a mixture containing 3 μg of DNA and 18 μg of Lipofectamine (Invitrogene) according to the manufacturer's instructions. For measuring the growth rate, cells were plated at the density of 1 × 105 cells in a 10-cm dish and counted each day using the trypan blue dye exclusion method for 6 d. The FACS analysis was performed as described previously (Ke and Chang 2004). The cells were fixed in 70% (v/v) ethanol and the cell cycle profile was examined using a Becton Dickson FACScan flow cytometer and CellQuest software.
Pulse-chase experiment
For metabolic labeling, cells were washed with PBS twice and incubated in 5 mL of methionine-free DMEM for 1 h of deprivation. Cells were then incubated in 2 mL of fresh methionine-free DMEM containing dialyzed 10% FBS and 35S-methionine (500 μCi, Amersham Pharmacia) for 30 min. To terminate labeling, complete DMEM medium was added for a 0- and 12-h chase prior to harvesting. The 35S-methionine-labeled cell lysates were subjected to hTK1 and hTMPK immunoprecipitation, followed by SDS-PAGE, Western blot analysis, and autoradiography.
RNA interference experiments
RNA interference experiments were performed as described previously (Ke and Chang 2004). siRNAs against Cdc20 and Cdh1 were synthesized according to sequences described in the previous studies (Donzelli et al. 2002; Wei et al. 2004). siRNAs against hTK1 and hTMPK were purchased from Dharmacon siGenome SMART pools. Control siRNAs against firefly luciferase were obtained from Dharmacon. Transfection of siRNA was carried out using Oligofectamine (Invitrogene) according to the manufacturer's instructions.
Expression and purification of recombinant proteins
GST-hTMPK and GST-cyclin B1 proteins were expressed in Escherichia coli JM109, purified by glutathione 4B beads, and cleaved by thrombin according to the manufacturer's instruction. GST-hTK1, His-mouse E1, His-UbcX, and His-UbcH10 were purified as described previously (Ke and Chang 2004). His-Cdc20 and His-Cdh1 were expressed and purified in a baculovirus system according to the study described by Kramer et al. (1998).
In vitro degradation assay, in vitro polyubiquitinylation, and in vitro binding assays
Preparation of synchronized cell extracts for in vitro experiments and in vitro degradation assay was performed as described previously (Brandeis and Hunt 1996). One microliter of 35S-methionine-labeled in vitro translated Myc-tagged hTMPK was added to 10 μL of cell extracts supplemented with an energy-regeneration system (100 μg/mL cycloheximide, 2.5 mM ATP, 80 μg/mL creatine kinase, 40 mM creatine phosphate) and incubated at 30°C, followed by taking 1 μL of samples for SDS-PAGE and autoradiography. For competition, mitotic and G1-phase extracts were incubated with 5 μg of purified GST-cyclin B1 and GST-hTK1, respectively, for 0.5 h at 4°C before the in vitro degradation assay. Purification of APC/C by immunoprecipitation was performed according to the previous study (Kramer et al. 1998). In vitro polyubiquitinylation assay was performed as described previously (Ke and Chang 2004). Briefly, APC/C beads were activated by incubating with purified His-Cdc20 or His-Cdh1 for 1 h at 25°C and were then added into a reaction mixture containing 200 μg/mL mouse E1, 100 μg/mL UbcX or UbcH10, 1.25 mg/mL ubiquitin, 100 ng of substrate, and an energy-regeneration system in a total volume of 5 μL. All reaction mixtures were incubated for 1 h at 37°C, after which reaction mixtures were analyzed by 5%–15% gradient SDS-PAGE and Western blotting.
Determination of dNTP pool in whole cell extract
Cells (5 × 106) were washed with 10 mL of cold PBS twice and extracted by 1 mL of ice-cold 60% methanol for 1 h at –20°C, followed by centrifugation for 15 min at 14,000 × g. The supernatant was transferred to a fresh tube and dried under vacuum. The residue was dissolved in 200 μL of sterile water and stored at –20°C for later analysis. Determination of the dNTP pool size in each extract was based on DNA polymerase-catalyzed incorporation of radioactive dNTP into the synthetic oligonucleotide template method described by Sherman and Fyfe (1989). The reaction mixture (20 μL) contained 100 mM HEPES buffer (pH 7.5), 10 mM MgCl2, 0.5 units of E. coli DNA polymerase I Klenow fragment (Promega), 0.25 μM oligonucleotide template, 1 μL of dNTP extract, and 8.32 nM 3H-dATP (for dTTP, dCTP, and dGTP) or 3H-dTTP (for dATP). Incubation was carried out for 60 min at 25°C and spotted onto DE81 paper. The papers were dried, washed (3 × 10 min) with 5% Na2HPO4, and rinsed once with distilled water and once with 95% ethanol. After drying, radioactivity on the papers was measured in a liquid scintillation counter (Beckman).
Establishment of stable cell lines
NIH 3T3 cells were initially cultured in medium supplemented with HAT (100 μM sodium hypoxanthine, 0.4 μM aminopterin, 10 μM thymidine) for two generations before being used for the stable transfection experiments. This procedure ensures that cells contain the wild-type hprt gene. For stable transfection, NIH 3T3 cells were transfected with pCDNA3.1Neo-hTK1 or pCDNA3.1Zeo-hTMPK, respectively. Simultaneous expression of hTK1 and hTMPK was achieved by cotransfection of pCDNA3.1Neo-hTK1 and pCDNA3.1Zeo-hTMPK into NIH 3T3 cells. After transfection, cells were selected in the medium containing G418 (600 μg/mL, for hTK1) and zeocin (800 μg/mL, for hTMPK1) for 14 d. The antibiotic-resistant clones were selected, expanded, and further cultured in medium supplemented with adequate amounts of antibiotics.
Analysis for spontaneous mutagenesis of the hprt gene analysis
Analysis for spontaneous mutagenesis of the hprt gene was performed similarly to the method described in a previous study (Dare et al. 1995). Cells were plated at low density (103 cells/dish) and grown to a density 106 cells per dish before selection to minimize aggregation between different cell clones in our experimental condition. Cells (3 × 105) were then plated onto a 10-mm dish in the complete medium supplemented with 6-thioguanine (6-TG, 5 μg/mL) for 10 d. To measure the colony formation efficiency, 103 cells in parallel were plated onto another 10-mm dish in the medium without 6-TG. Colonies were fixed with ice-cold methanol and stained with 0.5% crystal violet. 6-TG resistant colonies were isolated and expanded, followed by RNA preparation for the subsequent RT–PCR of hprt mRNA. The PCR products were then purified from agarose gel and were subjected to the direct DNA sequencing.
Acknowledgments
We thank Kristian Helin for kindly providing HA-tagged Cdc20 and Cdh1 and Marc W. Kirschner for providing Myc-tagged Cdc20 and Cdh1 constructs. We are grateful to Wu-Nan Wen, Jin-Yuan Su, and Maria Zajac-Kaye for providing CHL V79 cell line, GST-hTMPK, and pCDNA3.1Zeo, respectively. We acknowledge Tim Hunt, Joan V. Ruderman, and Jan-Michael Peters for providing mouse E1, Xenopus UbcX, wild-type and C114S mutant human E2C/UbcH10, and His-tagged Cdc20 and Cdh1 expression plasmids. We appreciate Michael Brandeis and Ambrose Jong for helpful discussions. We also thank Drs. Ruey-Hwa Chen and Jeou-Yuan Chen for critical reading of the manuscript. This study was supported by grants NSC93-2752-B002-006-PAE and NSC94-3112-B-002-025 from National Science Council and NHRI-EX94-9421BI from National Health Research Institute, Taiwan (R.O.C).
Supplemental material is available at http://www.genesdev.org.
Article and publication are at http://www.genesdev.org/cgi/doi/10.1101/gad.1322905.
References
- Bashir T., Dorrello, N.V., Amador, V., Guardavaccaro, D., and Pagano, M. 2004. Control of the SCFskp2-cks1 ubiquitin ligase by the APC/CCdh1 ubiquitin ligase. Nature 428: 190–193. [DOI] [PubMed] [Google Scholar]
- Bianchi V., Pontis, E., and Reichard, P. 1987. Regulation of pyrimidine deoxyribonucleotide metabolism by substrate cycles in dCMP deaminase-deficient V79 hamster cells. Mol. Cell. Biol. 7: 4218–4224. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bjursell G. and Reichard, P. 1973. Effects of thymidine on deoxyribonucleoside triphosphate pools and deoxyribonucleic acid synthesis in Chinese hamster ovary cells. J. Biol. Chem. 248: 3904–3909. [PubMed] [Google Scholar]
- Bolderson E., Scorah, J., Helleday, T., Smythe, C., and Meuth, M. 2004. ATM is required for the cellular response to thymidine induced replication fork stress. Hum. Mol. Genet. 13: 2937–2945. [DOI] [PubMed] [Google Scholar]
- Bradshaw Jr H.D. 1983. Molecular cloning and cell cycle-specific regulation of a functional human thymidine kinase gene. Proc. Natl. Acad. Sci. 80: 5588–5591. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brandeis M. and Hunt, T. 1996. The proteolysis of mitotic cyclins in mammalian cells persists from the end of mitosis until the onset of S phase. EMBO J. 15: 5280–5289. [PMC free article] [PubMed] [Google Scholar]
- Chabes A.L., Pfleger, C.M., Kirschner, M.W., and Thelander, L. 2003a. Mouse ribonuclotide reductase R2 protein: A new target for anaphase-promoting complex–Cdh1-mediated proteolysis. Proc. Natl. Acad. Sci. 100: 3925–3929. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chabes A., Georgieva, B., Domkin, V., Zhao, X., Rothstein, R., and Thelander, L. 2003b. Survival of DNA damage in yeast directly depends on increased dNTP levels allowed by related feedback inhibition of ribonucleotide reductase. Cell 112: 391–401. [DOI] [PubMed] [Google Scholar]
- Chang Z.F., Huang, D.Y., and Hsue, N.C. 1994. Differential phosphorylation of human thymidine kinase in proliferating and M phase-arrested human cells. J. Biol. Chem. 269: 21249–21254. [PubMed] [Google Scholar]
- Chang Z.F., Huang, D.Y., and Chi, L.M. 1998. Serine 13 is the site of mitotic phosphorylation of human thymidine kinase. J. Biol. Chem. 273: 12095–12100. [DOI] [PubMed] [Google Scholar]
- Coppock D.L. and Pardee, A.B. 1987. Control of thymidine kinase mRNA during the cell cycle. Mol. Cell. Biol. 7: 2925–2932. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dare E., Zhang, L.H., Jenssen, D., and Bianchi, V. 1995. Molecular analysis of mutations in hprt gene of V79 hamster fibroblasts: Effects of imbalances in the dCTP, dGTP, and dTTP pools. J. Mol. Biol. 252: 514–521. [DOI] [PubMed] [Google Scholar]
- DeGregori J., Kowalik T., and Nevins, J.R. 1995. Cellular targets for activation by the E2F1 transcription factor include DNA synthesis- and G1/S-regulatory genes. Mol. Cell. Biol. 8: 4215–4224. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Donzelli M., Squatrito, M., Ganoth, D., Hershko, A., Pagano, M., and Draetta, G.F. 2002. Dual mode of degradation of Cdc25A phosphatase. EMBO J. 21: 4875–4884. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Eriksson S., Thelander, L., and Akerman, M. 1979. Allosteric regulation of calf thymus ribonucleoside diphosphate reductase. Biochemistry 18: 2948–2952. [DOI] [PubMed] [Google Scholar]
- Harper J.W., Burton, J.L., and Solomon, M.J. 2002. The anaphase-promoting complex: It's not just for mitosis any more. Genes & Dev. 16: 2179–2206. [DOI] [PubMed] [Google Scholar]
- Huang S.H., Tang, A., Drisco, B., Zhang, S.Q., Seeger, R., Li, C., and Jong, A. 1994. Human dTMP kinase: Gene expression and enzymatic activity coinciding with cell cycle progression and cell growth. DNA Cell Biol. 13: 461–471. [DOI] [PubMed] [Google Scholar]
- Ke P.Y. and Chang, Z.F. 2004. Mitotic degradation of human thymidine kinase 1 is dependent on the anaphase-promoting complex/cyclosome–Cdh1-mediated pathway. Mol. Cell. Biol. 24: 514–526. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kramer E.R., Geiffers, C., Holzl, G., Hengstschlager, M., and Peters, J.M. 1998. Activation of the human anaphase-promoting complex by proteins of the CDC20/Fizzy family. Curr. Biol. 8: 1207–1210. [DOI] [PubMed] [Google Scholar]
- Kunz B.A. 1988. Mutagenesis and deoxyribonucleotide pool imbalance. Mutat. Res. 200: 133–147. [DOI] [PubMed] [Google Scholar]
- Li J.L., Lu, C.Y., Ke, P.Y., and Chang, Z.F. 2004. Perturbation of ATP-induced tetramerization of human cytosolic thymidine kinase by substitution of serine-13 with aspartic acid at the mitotic phosphorylation site. Biochem. Biophys. Res. Commun. 313: 587–593. [DOI] [PubMed] [Google Scholar]
- Liang P., Averboukh, L., Zhu, W., Haley, T., and Pardee, A.B. 1995. Molecular characterization of the murine thymidylate kinase gene. Cell Growth Differ. 6: 1333–1338. [PubMed] [Google Scholar]
- Littlepage L.E. and Ruderman, J.V. 2002. Identification of a new APC/C recognition domain, the A box, which is required for the Cdh1-dependent destruction of the kinase Aurora-A during mitotic exit. Genes & Dev. 16: 2274–2285. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lundin C., Erixon, K., Arnaudeau, C., Schultz, N., Jenssen, D., Meuth, M., and Helleday, T. 2002. Different roles for nonhomologous end joining and homologous recombination following replication arrest in mammalian cells. Mol. Cell. Biol. 22: 5869–5878. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meuth M. 1989. The molecular basis of mutations induced by deoxyribonucleoside triphosphate pool imbalances in mammalian cells. Exp. Cell Res. 181: 305–316. [DOI] [PubMed] [Google Scholar]
- Mohindra A., Hays, L.E., Phillips, E.N., Preston, B.D., Helleday, T., and Meuth, M. 2002. Defects in homologous recombination repair in mismatch-repair-deficient tumour cell lines. Hum. Mol. Genet. 11: 2189–2200. [DOI] [PubMed] [Google Scholar]
- Mohindra A., Bolderson, E., Stone, J., Wells, M., Helleday, T., and Meuth, M. 2004. A tumour-derived mutant allele of XRCC2 preferentially suppresses homologous recombination at DNA replication forks. Hum. Mol. Genet. 13: 203–212. [DOI] [PubMed] [Google Scholar]
- Navalgund L.G., Rossana, C., Muench, A.J., and Johnson, L.F. 1980. Cell cycle regulation of thymidylate synthetase gene expression in cultured mouse fibroblasts. J. Biol. Chem. 255: 7386–7390. [PubMed] [Google Scholar]
- Peters J.M. 2002. The anaphase-promoting complex: Proteolysis in mitosis and beyond. Mol. Cell 9: 931–943. [DOI] [PubMed] [Google Scholar]
- Petersen B.O., Wagener, C., Marinoni, F., Kramer, E.R., Melixetian, M., Denchi, E.L., Gieffers, C., Matteucci, C., Peters, J.M., and Helin, K. 2000. Cell cycle-and cell growth-regulated proteolysis of mammalian CDC6 is dependent on APC–CDH1. Genes & Dev. 14: 2330–2343. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pfleger C.M. and Kirschner, M.W. 2000. The KEN box: An APC recognition signal distinct from the D box targeted by Cdh1. Genes & Dev. 14: 655–665. [PMC free article] [PubMed] [Google Scholar]
- Pfleger C.M., Lee, E., and Kirschner, M.W. 2001. Substrate recognition by the Cdc20 and Cdh1 components of the anaphase-promoting complex. Genes & Dev. 15: 2396–2407. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reichard P. 1988. Interactions between deoxyribonulceotide and DNA synthesis. Annu. Rev. Biochem. 57: 349–374. [DOI] [PubMed] [Google Scholar]
- Reichard P., Eliasson, R., Ingemarson, R., and Thelander, L. 2000. Cross-talk between the allosteric effector-binding sites in mouse ribonucleotide reductase. J. Biol. Chem. 275: 33021–33026. [DOI] [PubMed] [Google Scholar]
- Sclafani R.A. and Fangman, W.L. 1984. Yeast gene CDC8 encodes thymidylate kinase and is complemented by herpes thymidine kinase gene TK. Proc. Natl. Acad. Sci. 81: 5821–5825. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sherley J.L. and Kelly, T.J. 1988. Regulation of human thymidine kinase during the cell cycle. J. Biol. Chem. 263: 8350–8358. [PubMed] [Google Scholar]
- Sherman P.A. and Fyfe, J.A. 1989. Enzymatic assay for deoxyribonucleoside triphosphates using synthetic oligonucleotides as template primers. Anal. Biochem. 180: 222–226. [DOI] [PubMed] [Google Scholar]
- Spyrou G. and Reichard, P. 1988. Dynamic of the thymidine triphosphate pool during the cell cycle of synchronized 3T3 mouse fibroblast. Mutat. Res. 200: 37–43. [DOI] [PubMed] [Google Scholar]
- Studzinski G.P. and Lambert, W.C. 1969. Thymidine as a synchronizing agent I. Nucleic acid and protein formation in synchronous HeLa cultures treated with excess thymidine. J. Cell Physiol. 73: 109–117. [DOI] [PubMed] [Google Scholar]
- Su J.Y. and Sclafani, R.A. 1991. Molecular cloning and expression of the human deoxythymidylate kinase gene in yeast. Nucleic Acids Res. 10: 459–472. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wei W., Ayad, N.G., Wan, Y., Zhang, G.J., Kirschner, M.W., and Kaellin Jr., W.G. 2004. Degradation of the SCF component Skp2 in cell cycle phase G1 by anaphase-promoting complex. Nature 428: 194–198. [DOI] [PubMed] [Google Scholar]