Abstract
Circulating tumor DNA (ctDNA) is nowadays considered a robust source to search for druggable tumoral genetic alterations, and in some specific settings liquid biopsy (LB) is already part of the diagnostics scenario and it has successfully implemented in the everyday practice. Three strengths make LB an extraordinary tool: i) to represent the complex molecular mosaicism that characterizes spatially heterogeneous malignancies; ii) to monitor in real-time the tumoral molecular landscape (i.e. to depict the longitudinal/temporal tumor evolution); iii) to ensure molecular profiling even in those cases in which tissue sampling is not feasible or not adequate. This review provides a snapshot of the current state of the art concerning ctDNA assay utility in gastric cancer (GC), testing its robustness as marker and seeking to understand the reasons for the delay in its application in clinical practice.
Keywords: Liquid biopsy, Gastric cancer, Circulating markers, ctDNA, CNAs
1. Introduction
The comprehensive molecular characterization and the development of novel molecular targeted drugs have revolutionized the history of many solid tumors, including gastric cancer (GC), leading to the transition to more specific and personalized treatment protocols. Despite the overall survival (OS), still remains poor, especially in advanced stages, therapeutic options for GC-targeted treatment, have increasingly improved in the last few years, and besides the well-established HER2-overexpression, several other druggable targets are emerging opening novel therapeutic options. Molecular pathologists are therefore faced with new challenges. They are required not only to assess these markers at diagnosis but also, experienced the need to monitor how these markers change throughout the disease evolution. In this setting LB was recognized as playing an important role. Indeed, LB can detect a broad range of molecular alterations including the presence of DNA mutations or copy number variations (CNV) in cancer driver genes, epigenetic alterations, transcriptomic and/or proteomic profile deregulation, and the identification of metabolite profile changes [1]. Considering that blood is not only in contact with the majority of tumor types but can also, be easily collected, it represents the gold standard for LB-based analysis, but several other body fluids can be interrogated [2].
1.1. Precision diagnostics in gastric cancer
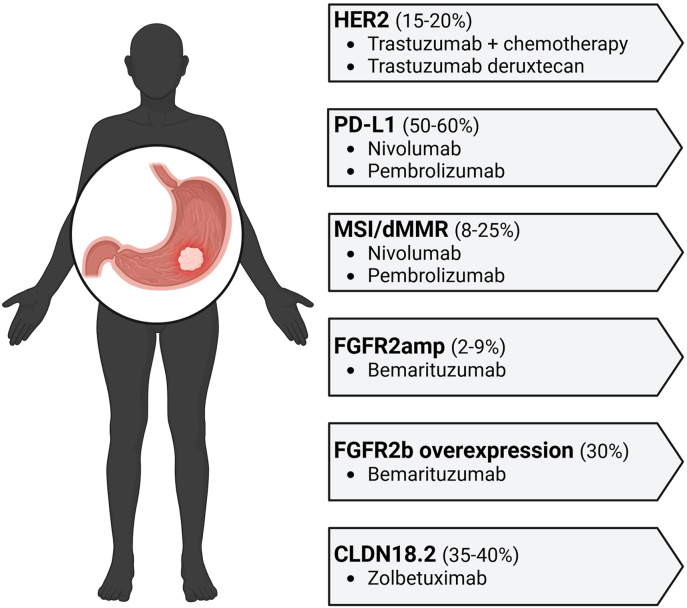
The Cancer Genome Atlas (TCGA) research network, ten years ago, paved the way for precision oncology in GC. Since this first seminal work that identified four distinct GC molecular subgroups, the number of actionable alterations has progressively improved [3]. To date, the human epidermal growth factor receptor 2 (HER2), microsatellite instability (MSI)/mismatch repair protein (MMR) status, and the expression of programmed death ligand 1 (PD-L1) should be routinely evaluated on GC specimens, according to the Combined Positive Score (CPS). HER2 overexpression is the first biomarker assessed in newly diagnosed GCs, with both a prognostic and predictive value [4]. According to the ASCO guidelines, HER2 and PD-L1 status should be assessed before starting first-line therapy in metastatic GC patients [5], while, the recently published Pan-Asian ESMO adapted guidelines stated that HER2, PD-L1, and MMR status should be available at diagnosis [6]. The seminal ToGA study demonstrated that the association of chemotherapy with a monoclonal antibody (MoAb) against HER2 (trastuzumab) improved the median OS (mOS) in patients with Erb-B2 Receptor Tyrosine Kinase 2 (ERBB2) amplification [7]. The percentage of patients who could benefit from this treatment (e.g. patients presenting ERBB2 amplification) stands for 10–20 % of all GCs [8]. Both PD-L1 assessment, and MMR protein analysis or microsatellite status definition can be considered in HER2-negative patients, and their evaluation allows the identification of patients eligible for the treatment with immune checkpoint inhibitors (ICIs). The prevalence of MSI/dMMR in GC is strongly related to the patients' provenience, and ranges from 4 to 8% in the Asiatic series to 23–28 % reported for Western countries [[9], [10], [11]], despite the benchmark remaining the molecular classification of TCGA that set the prevalence of MSI-high (MSI-H) GCs as 22 % of all GC cases [3]. Other opportunities are emerging, especially for HER2-negative patients, as Claudin 18.2 (CLDN18.2) over-expression. Although the two markers are not mutually exclusive, CLDN18.2 positivity was found to be numerically higher in HER2-negative patients compared with HER2-positive ones and prevalence in HER2-negative GCs rates from 35 to 40 % [12,13]. In light of promising results obtained by GLOW and SPOTLIGHT trials, the moAb zolbetuximab targeting CLDN18.2 was just approved by the European Medicine Agency. Moreover, other promising results are coming from the targeting of the FGFR2-driven signaling. Fibroblast growth factor 2 (FGFR2) is considered an important oncogenic driver, and the targeting of its aberrant signaling pathway represents a novel opportunity in the targeted therapy. FGFR2 gene amplification and the overexpression of the FGFR2b isoform protein (any moderate to strong membranous staining) were reported with a prevalence of 4 % and 29 % respectively, in the phase II FIGHT trial [14]. FIGHT trial final analysis, after 24-month follow-up, revealed that the combination of bemarituzumab with the standard mFOLFOX6 regimen, in HER2-negative advanced GC with FGFR2b overexpression and/or FGFR2 gene amplification improved both progression-free survival (9.5 vs 7.4 months), and mOS (19.2 vs 13.5 months) [15] (Fig. 1). Other alterations as Neurotrophic Receptor Tyrosine Kinase (NTRK) fusions, v-Raf murine sarcoma viral oncogene homolog B (BRAF) mutations, are under evaluation in ongoing clinical trials [16].
Fig. 1.
Principal molecular targets currently assessed in gastric cancer. Patients positive for HER2 expression are eligible for treatment with trastuzumab, a humanized recombinant monoclonal antibody able to bind to HER2 receptors, reducing its activity. According to the ToGA study, trastuzumab is used in combination with chemotherapy (fluorouracil plus cisplatin) in HER2-positive patients. Patients eligible for immunotherapy are selected on the basis of PD-L1 expression, calculated as CPS score and/or microsatellite status: MSI-high patients respond well to immunotherapy. The FIGHT trial investigated the activity of bemarituzumab as first-line treatment in metastatic patients harboring FGFR2 amplification or FGFR2b over-expression. The effect of the monoclonal antibody zolbetuximab against CLDN18.2 associated with chemotherapy (mFOLFOX6) was investigated in the SPOTLIGHT trial. Most of the cited molecular targets are almost mutually exclusive.
Although a good number of molecular targets are now available for GC, the main challenge in the targeted therapy is represented by its high molecular heterogeneity. For almost all GC, spatial and temporal heterogeneity in druggable targets was reported [17]. As demonstrated by Chao et al. nearly half (46 %) of the GC specimens presented a high intra-tumoral heterogeneity with clonal tumor cell populations coexisting at submillimeter distances [18]. Regarding CLDN18.2 expression, intra-tumoral heterogeneity was reported at around 31 % in primary, resectable GCs [19], while comparing CLDN18.2 expression in paired primary-metastatic specimens, the reported temporal heterogeneity was 25.2 % [20]. Even higher heterogeneity was reported for FGFR2b protein expression, with both an extensive intra-tumoral (55.5 %) and temporal (28 %) heterogeneity [21]. Even in PD-L1 assessment, studies suggested the need for at least five bioptic sampling [22]. As regards tumor heterogeneity, the better-characterized marker is HER2, and in routine clinical practice, six to eight samples, representative of neoplasia, are ideally required to confidentially evaluate HER2 status [23,24]. In such a context, is clear that novel technological approaches able to better represent the above-described heterogeneity are required. Digital spatial profiling (DSP) and LB may be important weapons, in the hands of molecular pathologists to underline spatial and temporal makers’ changes.
1.2. Biological and clinical strengths of liquid biopsy
Tissue biopsy (TB) is essential for the initial diagnosis and for the definition of tumor morphological features and biomarkers assessment. Thus, LB will be unable to replace TB in this stage. However, it holds several advantages. First, being minimally invasive, LB is a valid alternative to TB in metastatic setting for patients with inaccessible tumors or with a high risk of potentially life-threatening complications (e.g. hemorrhage, pneumothorax) or more in general, in elderly/debilitated patients presenting advanced disease. Moreover, it can be used as a tool to capture intra-tumor and inter-tumor heterogeneity. Another strength of LB is the possibility to perform serial samplings (e.g. longitudinal tumor monitoring) over time and detect changes in the tumor cell population in response to the therapeutic selective pressure. A wide range of clinical trials focusing on LB is underway. Over the past two years (2021–2022), approximately 200 new trials related to LB have been initiated in the field of oncology alone [25]. Most of the studies are focused on therapy response monitoring. Over the years LB assay increased progressively its robustness, becoming, nowadays, a valid tool when TB fails or is not feasible, in certain malignancies, such as non-small cell lung cancer (NSCLC). According to the guidelines of the College of American Pathologists (CAP), in the setting of NSCLC, LB can be employed, as an alternative to TB, to identify the presence of Epidermal Growth Factor Receptor (EGFR) T790M mutation, the most common resistance mechanism to EGFR-tyrosine kinase inhibitor (TKI) [26]. The same indication was also proposed by the recommendations of the AIOM–SIAPEC-IAP–SIBIOC–SIF Italian Scientific Societies, which not only recommended the employment of LB in NSCLC but also indicated possible novel application fields including i) breast cancer (BC) where LB is proposed to identify predictive biomarkers of response/resistance to treatment. In this setting, the PADA-1 trial demonstrated the benefit derived from the ESR1 mutations monitoring in plasma ctDNA from patients with advanced BC treated with aromatase inhibitors [27]. The guidelines also suggested ii) melanoma and iii) colorectal cancer (CRC), as novel fields of LB application to monitor treatment response, with BRAF and RAS genes as the most informative markers [28]. Although several clinical trials confirmed the potential role of LB in the GC setting, LB has not been codified in clinical practice, at this time.
1.3. Circulating tumor DNA (ctDNA) in gastric cancer
Circulating free DNA (cfDNA) is composed of small DNA fragments (mean length 166 bp) that are physiologically released by all cells, despite the main source being the hematopoietic system [29]. Circulating free DNA can be released by different mechanisms, which affect the length of the released DNA [30]. The component of cfDNA that is released by tumor cells is defined as circulating tumor DNA (ctDNA) and stands for less than 0.1 % to over 10 % of total cfDNA, depending on tumor type, grade of the malignancy, burden, clinical status (e.g. inflammatory status) and tumor dimensions [31]. In fact, for GC and many other tumors as well, the amount of ctDNA is strictly dependent on the stage. Fang and colleagues reported a number of cfDNA copies for mL three times higher in stage IV compared to earlier stages. While among less advanced stages no significative differences in the number of cfDNA copies were found [32]. Several studies assessed the different release rates between different cancer types. All the studies confirmed that cfDNA shedding is strictly dependent on primary: a high release rate was found in lung cancer, prostate, uterine and hepatocellular carcinoma (91%–77.1 %), while, it was lower in renal and thyroid cancers (56-41 %) and extremely low in gliomas (10 %) [33].
As for GC, cfDNA shedding rate is very similar to that of lung or colorectal cancer, for which LB is already recommended in clinical practice [34]. Circulating tumor DNA analysis allows the identification of a wide range of genomic alterations, including single nucleotide variants (SNVs), chromosomal rearrangements, and CNV (Fig. 2) [35].
Fig. 2.
Principal biomarkers currently investigated in blood from gastric cancer patients. Two different types of markers can be detected in blood: circulating tumor cells (CTC) and tumor-educated platelets (TEPs), in the plasma or serum isolated from blood three markers can be investigated: circulating tumor DNA (ctDNA), extracellular vesicles (EVs) and noncoding RNAs (ncRNAs).
1.3.1. Concordance between ctDNA and tissue biopsy
The concordance rate between LB and TB is set at around 76 % for NSCLC, while in GC is reported to be more heterogeneous [36]. As for many other malignancies, in GC most of the LB-TB correlations have been performed in advanced/metastatic cases. Considering only stage IV GC, the percentage of agreement ranges from 42 %, as reported by Kung et al., who analyzed 56 cases using a 29-genes NGS panel, to 89.5 % found in the VIKTORY clinical trial (NCT02299648) [37,38]. In the latter study, a NGS approach was used, but only three altered genes were considered, reporting percentages of concordance of 89.5 % for MET receptor tyrosine kinase (MET) amplifications, and 76.9 % and 75 % for respectively Kirsten Rat Sarcoma Viral Oncogene Homolog (KRAS) and Tumor Protein P53 (TP53) mutations. Focusing on GC-druggable targets, the concordance of ERBB2 amplification between plasma and tissue is still under debate. Two studies, both performed in stage IV patients, are available with different outcomes. Wang and colleagues reported a high concordance (91.1 %, k = 0.820) of ERBB2 amplification between ctDNA and tumor tissues, while in another work, with a very similar methodological strategy, Maron and co-workers reported a concordance markedly lower (61 %) [39,40]. A similar concordance rate (64 %) was also reported, by Shitara and co-workers in an exploratory analysis of the DESTINY-Gastric01 trial, conducted on advanced GC [41,42]. In the same setting: stage IV-GC, the Japanese plasma genomic profiling study GOZILA, revealed that ctDNA sequencing identified heterogeneous FGFR2 amplification that conventional TB failed to identify (prevalence 3.4 % in TB versus 14.8 in LB) [43]. As expected, the percentage of agreement dramatically decreases when less advanced stages are considered. He and co-workers considered LBs from both stage IV and III GCs, using a 605-genes NGS panel, and reported an agreement of 70.6 % and 30 %, respectively, indicating how the agreement drops dramatically, at less advanced stages [44]. Complete information about TB and LB comparisons discussed above are reported in Table 1.
Table 1.
Principal TB versus LB comparisons for ctDNA analysis in GC.
| Stage | % Agreement with TB | Investigated genes | Techniques | Numerosity of the cohort | Plasma volume | Reference |
|---|---|---|---|---|---|---|
| IV | 42 % | 29 genes | Targeted-NGS | 56 patients | 1 mL | 37 |
| IV | 89.5 % | MET gene | Targeted-NGS | 715 patients | 2 mL | 38 |
| 76.9 % | KRAS gene | 16 patients | ||||
| 75 % | TP53 gene | 13 patients | ||||
| ND | 91 % | ERBB2 gene | Targeted-NGS | 56 patients | 0.5 mL | 39 |
| Mixed | 61 % | ERBB2 gene | Targeted-NGS | 1630 patients | ND | 40 |
| IV | 64 % | ERBB2 gene | Targeted-NGS | 151 patients | ND | 41 |
| IV | 90.9 % | FGFR2 gene | Targeted-NGS | 365 patients | 3 mL | 43 |
| IV | 70.6 % | 605 genes | Targeted-NGS | 14 patients | 2 mL | 44 |
| III | 30.2 % | 10 patients |
1.3.2. ctDNA in gastric cancer early diagnosis
One of the major limitations in GC management is represented by the advanced stage at diagnosis, due to the lack of early specific symptoms and mass screening strategies for the general population. Despite some serological tests, including pepsinogen, gastrin-17, Helicobacter pylori antibody, carbohydrate antigens: 72-4 (CA72-4), 19-9 (CA19-9), 125 (CA125), 24-2 (CA24-2) and alpha-fetoprotein (AFP) are available as tools for low invasive screening, their specificity and sensitivity is limited [45]. Thus, ctDNA may be useful in improving early diagnosis of GC. Nevertheless, it is important to underline that for diagnostic purposes, in GC setting endoscopy associated with tissue biopsy is still mandatory. In comparative works, the analysis of ctDNA was reported as more sensitive than traditional serum markers (e.g. CEA, CA19-9, CA72-4) in early GC screening [46]. As demonstrated by Park and colleagues the mean quantity of ctDNA in plasma samples of GC patients was generally 2.4 fold higher than the respective age-matched healthy controls, with an estimated sensitivity and specificity of respectively 75 and 63 % [47]. However, the simple quantification of ctDNA is not enough to reach the diagnostic sensitivity, considering that increasing in ctDNA may be associated with different pathological conditions such as, for example, infections or inflammatory diseases [48]. CancerSEEK, a blood-based test for pan-cancer early diagnosis based on the analysis of both protein expression and gene alterations, showed high sensitivity in detecting GC at stages I-III. However, the sensitivity, which was around 70 %, was lower than that reported for ovary and liver cancers, despite being consistent with those reported for other malignancies such as pancreatic, colorectal, and lung cancer [49]. In another study the ratio between V-Myc Avian Myelocytomatosis Viral Oncogene Homolog (MYC) and the housekeeping gene Glyceraldehyde-3-Phosphate Dehydrogenase (GAPDH) by real-time PCR was assessed, achieving a sensitivity of 75 % in detecting GC at various stages. A high concordance with MYC amplification was evaluated in the matched tissue biopsies [50]. According to other malignancies, in the GC setting, the reliability of the analysis is strictly dependent on disease stage, which affects not only the tumor burden but also the ctDNA shedding. Yang et al. reported a ctDNA positivity of 68 % in stage III cases which dropped to 21 % in stage I-II malignancies [51], while Kim and colleagues established 90 ng/mL of cfDNA as a cut-off value to obtain high sensitivity (≥96 %) [52].
1.3.3. ctDNA in gastric cancer in treatment monitoring
Nowadays, treatment monitoring in GC is based on imaging and serological dosing of markers such as CA19-9 and CA72-4 [53]. The evaluation of molecular evolution of the disease during the treatment is mandatory considering the high temporal heterogeneity of GCs. Analysis of ctDNA was employed to monitor and evaluate the trastuzumab-resistant mechanism in GC treated with a combination of chemotherapy and anti-HER2 MoAb. The longitudinal analysis of ctDNA revealed the onset of secondary mutations, with variant allele frequence (VAF) values ≥ 2 %, involving mainly MAPK and RAS pathways [54]. In the DESTINY-Gastric01 trial, testing trastuzumab conjugated with deruxtecan (T-DXd) the employment of LB was useful to better identify patients benefitting from the treatment. The analysis of pre-treatment ctDNA revealed that patients presenting plasma amplification in MET, EGFR, or FGFR2 genes had a lower overall response rate (ORR). In the same context, the analysis of ctDNA at the end of the treatment proposed acquired DNA Topoisomerase I (TOP1) gene variants, as a possible mechanism of resistance to T-DXd [42]. The quantification of ctDNA by LB was shown to be useful also in verifying the response to immunotherapy. In a cohort of metastatic GC treated with pembrolizumab, the ctDNA content changes were found to be powerful predictors of both response and/or disease progression. Patients who experienced a decrease in ctDNA were associated with longer progression-free survival (PFS), and better ORR and disease control rate (DCR) [55]. Jin et al. not only demonstrated that patients who experienced a decline higher than 25 % in the maximal somatic variant allelic frequency had a longer PFS and higher response rate, but also correlated the presence of specific mutations (Transforming Growth Factor Beta Receptor 2: TGFBR2, Ras Homolog Family Member A: RHOA, and Phosphatidylinositol-3,4,5-Trisphosphate Dependent Rac Exchange Factor 2: PREX2) at baseline ctDNA with a shorter PFS. In the PANGEA clinical trial, the analysis of ctDNA was used as an alternative source, when TB was not available to monitor changes in tumor molecular profile during the different lines of therapy. The study demonstrated that following the temporal heterogeneity of the disease improved both the OS (15.7 vs 9 months) and the median time to treatment failure (13.6 vs 7.9 months). The search for genetic alterations in ctDNA can be also used as criteria for patients' selection. FIGHT, a randomized trial designed to investigate the efficacy and safety of an anti-FGFR antibody, was one of the first trials to enroll patients using a combination of ctDNA testing and tumor tissue immunohistochemistry [14].
1.3.4. Circulating tumor DNA for minimal residual disease in GC
Another important application of LB is the possibility to evaluate the presence of minimal residual disease (MRD) undetectable by imaging, after a curative approach. In this case, the employment of LB allows to avoid unnecessary administration of adjuvant therapies in low-risk patients and at the same time, identifies patients who might benefit from the adjuvant treatment [56]. LB analysis in patients with stage I-III GC after surgical resection demonstrated that the detection of ctDNA post-surgery (from 9 to 48 days after surgery, and before any adjuvant treatment) is strongly related to disease relapse [51]. Overall, 100 % of patients in which ctDNA was post-surgery detected, experienced recurrence, against 32 % in patients negative for ctDNA. Moreover, ctDNA positivity at any time point during the longitudinal post-surgery follow-up was associated with worse post-operative DFS and OS. The study demonstrated that the analysis of ctDNA could advance of six months the imaging-based detection of recurrence. The correlation between the detection of ctDNA after surgery and tumor relapse was further confirmed by Openshaw et al. demonstrating that patients positive for ctDNA post-surgery presented a median PFS of 298 days versus 1000 of the negative ones [57]. Possible applications of ctDNA analysis in GC are summarized in Fig. 3.
Fig. 3.
Analysis of ctDNA in gastric cancer. Principal advantages and issues hindering the introduction of ctDNA analysis in GC. Possible biomarkers to be tested and applications of LB applied to GC are summarized. Applications and biomarkers are enlisted starting from the more promising.
1.3.5. Methylation profile in gastric cancer released ctDNA
As described in several works, ctDNA may be investigated to study the methylation profile in the shedding tissue with a double analytical interest [58]. From a diagnostic point of view, the value of epigenetic alterations as biomarkers has been reported in many studies, which demonstrated that methylation was among the earliest and most comprehensive genomic aberrations and could correlate with clinical parameters in several cancers [59]. On the other hand, the unique tissue-specific methylation pattern can help to identify the primary shedding tumor. This is extremely useful in the identification of cancers of unknown primary (CUP), as well as the primary site of distant metastases [60]. Thus, ctDNA methylation signature is considered a promising screening marker for several cancers, including GC (Table 2). A cfDNA-based sequencing technology coupled with software able to recognize methylation patterns per region was able to detect cancer signals across multiple cancer types and predict the cancer signal origin with high accuracy [61]. While the overall sensitivity for all cancer types regardless of stage was 51.5 %, the sensitivity for GC was 66.7 %, similar to CRC and lung cancer (reporting a sensitivity of 82 % and 74.8 %, respectively). The sensitivity of the method was strictly dependent on the stage, ranging from 16 % for stage I to 100 % for stage IV. Obtained sensitivity, which is comparable with sensitivity reported for ctDNA tests based on evaluation of DNA mutation, may be a promising tool for early diagnosis, with the added value of high accuracy in the prediction of cancer signal origin.
Table 2.
Principal methylated genes in GC cancer.
| Gene | Source | Technique | Features | Reference |
|---|---|---|---|---|
| SOX17 | Serum | Methylation-specific PCR | Distinguish GC patients from healthy controls. Prognostic value: patients with non-methylated SOX17 present a better survival rate | [62] |
| XAF1 | Serum | Real-time methylation-specific PCR | Distinguish GC patients (methylated) from healthy controls (non-methylated) | [63] |
| TFPI2 | Serum | Methylation-specific PCR | TFPI2 methylation is associated with lymph node metastasis and distant metastasis | [64] |
| DAPK1 | Serum | Methylation-specific PCR. | Markers for screening and surveillance of GC | [65] |
| CDH1, GSTP1, | ||||
| CDKN2B, | ||||
| CDKN2B | ||||
| CDKN2B | Serum | Methylation-specific PCR | P16 methylation is a biomarker for early detection of gastric cancer | [66] |
| THBS1 | Serum | Methylation-specific PCR | Prognostic marker: aberrant THBS1 methylation is related to peritoneal dissemination, tumor progression, and poor prognosis | [67] |
| RUNX3 | Serum | Multiplex digital PCR | Prognostic value: methylated RUNX3 is associated with tumor size, massive submucosal invasion, and lympho-vascular invasion | [68] |
| ZIC1, HOXD10, RUNX3 | Plasma | Methylation-specific PCR | Distinguish GC and precancerous lesions in early screening protocols. | [69] |
| DOCK10, CABIN1, KCNQ5 | Plasma | Methylated CpG tandems amplification and sequencing (MCTA-Seq) | Discriminate early-stage GC | [70] |
| PCDH10, RASSF1A | Plasma | Methylation-specific PCR | Diagnostic marker: high frequency of methylation is reported in the GC group compared with the control. | [71] |
| Prognostic marker: cumulative survival is significantly lower in cases positive for RASSF1A methylation. | ||||
| Methylation of RASSF1A and PCDH10 is higher in late-stage patients and correlates with metastasis | ||||
| SEPT9, RNF180 | Plasma | Methylation-specific PCR | Prognostic marker: SEPT9 methylation is correlated with poorer GC survival | [72] |
| Diagnostic marker: identifies GC patients from controls | ||||
| RPRML | Plasma | Methylation-specific PCR | Methylated RPRML distinguishes patients with GC from controls | [73] |
| SFRP2 | Plasma | Digital PCR | SFRP2 methylation is associated with GC prognosis | [74] |
| ELMO1, ZNF569, C13orf18 | Plasma | Methylation-specific PCR | Diagnostic marker: discriminate GC from healthy controls | [75] |
| KCNQ5, C9orf50, CLIP4 | Plasma | Methylation-specific PCR | Diagnostic markers for GC early detection, with a sensitivity of 67.9 % and a specificity of 86.6 % | [76] |
| IRX1 | Plasma | Methylation-specific PCR | Distinguishes GC group from healthy controls. | [77] |
| Higher methylation levels are closely associated with TNM staging |
1.3.6. Pitfalls in ctDNA analysis
As for TB, the pre-analytical phase is crucial in LB to ensure the reliability of the obtained information. The key point to consider in approaching LB is the lysis of blood cells (e.g. leucocytes) that can release germinal DNA potentially diluting ctDNA and thus impacting the identification of potential mutations. To avoid leukocyte lysis, processing blood in a short time (within 1 h) is recommended. Furthermore, despite some authors utilizing serum as a source for cfDNA analysis due to its seemingly higher cfDNA content, it's important to note that serum may harbor genomic DNA contamination resulting from leukocyte lysis during the clotting process [78]. The second aspect to consider is that ctDNA has a relatively short half-life. Fast processing of the sample can avoid ctDNA degradation. However, if rapid processing is not possible, the use of collection tubes with preservatives can prevent analyte degradation [79]. Another aspect to consider is the sensitivity of the assay. Despite the recent technological improvements, LB often yields VAFs <1 %. Orthogonal studies have demonstrated that the major pitfalls (i.e., false positives and negatives) occur in the 0.1%–1% VAF range [80]. Even when high-sensitivity techniques are employed, ctDNA might not be detected, due to the low starting cfDNA input. Considering that one ng of human DNA contains about 300 haploid copies of genome, at least 3.3 ng of cfDNA is needed to have at least one copy of a mutation presenting a VAF of 0.1 %. In this case, the employment of extraction techniques that do not need a fixed plasma input could improve the cfDNA yield. Furthermore, LB can lead to an incorrect mutation classification due to clonal hematopoiesis (CH), a process that involves the accumulation of somatic mutations in hematopoietic stem cells due to aging, leading to clonal expansion of mutations in blood cells. These mutations (mainly found in DNA Methyltransferase 3 Alpha: DNMT3A, Tet Methylcytosine Dioxygenase 2: TET2, ASXL Transcriptional Regulator 1: ASXL1, Janus Kinase 2: JAK2, Protein Phosphatase, Mg2+/Mn2+ Dependent 1D: PPM1D, and TP53) in some cases overlap with common mutations in cancer driver genes and could be erroneously considered to arise from the tumor [81]. Retaining an aliquot of blood cells (buffy coat), to sequence cells in the presence of suspicious mutations, may be useful for the discrimination between CH-derived and tumor-derived mutations.
1.3.7. Techniques for ctDNA analysis in liquid biopsy
CtDNA represents only a minute fraction of cfDNA, making the detection of rare variants challenging (see Fig. 4). To optimally study ctDNA, technologies with a limit of detection (LOD) of around 0.1 % are required. Technologies employed for ctDNA analysis can be divided into PCR-based and NGS-based approaches. The two main advantages of PCR-based approaches are the lower cost and the simpler workflow, but PCR-based approaches can be used to analyze only one mutation per assay. When PCR-based technologies are applied to LB there are four major points to consider. The first one is cfDNA fragmentation; the majority of DNA fragments are about 160 bp in length, thus amplicon size must be adjusted to the length of cfDNA, designing assays with amplicons around 80 bp [82]. The second point is the need for high specificity in the detection of rare single mutations (VAF <0.1 %). To improve specificity several modifications to the traditional qPCR were introduced as ARMS-PCR (amplification-refractory mutation system PCR), PNA-LNA (peptide-nucleic-acid-locked nucleic acid) Clamp PCR, and COLD-PCR (co-amplification at lower denaturation temperature PCR) [[83], [84], [85]]. ARMS-PCR is based on the employment of allele-specific primers. The introduction of a mismatch at the 3’ end of the primer that alters the annealing temperature, associated with the employment of a Taq DNA polymerase without exonuclease activity, which inhibits the mismatch repair, allows a selective amplification. This technique is already used to improve selectivity in Therascreen EGFR Plasma RGQ PCR (Qiagen). PNA-LNA Clamp PCR employs peptide nucleic acids to hide the wild-type DNA sequence, preventing its amplification. Similarly, the selection of low-abundance mutated alleles in the presence of an excess of wild-type alleles is obtained in COLD-PCR, exploiting the different denaturation temperatures among the mutant and the wild-type allele. These qPCR variants can detect VAF as low as 0.1 %, achieving a significantly higher sensitivity than qPCR (VAF 5 %). Another point to consider is iii) the need for an absolute quantification of the genetic alterations. The absolute quantification by qPCR is obtained using standard curves, and assuming that samples and standards have the same amplification efficiency. Minimal changes in PCR efficiency, for example, due to the presence of PCR inhibitors, can markedly alter the accuracy of the quantification. The last point is represented by the need for high sensitivity. The introduction of digital PCR (dPCR), has effectively answered the last two points, allowing a highly sensitive quantitative detection of circulating markers. Three different dPCR technologies are currently available i) the droplet digital PCR (ddPCR), ii) the chamber-based digital PCR (cdPCR), and iii) the BEAMing (Beam, Emulsion, Amplification, Magnetics) dPCR. In both ddPCR and cdPCR, DNA molecules are partitioned, according to the Poisson distribution, within reaction units, and within them are amplified using TaqMan Probes or intercalating dyes. After the fluorescence detection, the ratio between the number of positive and negative partitions is performed and the absolute quantification is obtained. The main difference between these two systems is represented by the partition method: while ddPCR partitions are composed of thousands (∼20,000) of droplets generating a water-in-oil emulsion, in cdPCR partitions are physical wells on a solid support [86]. Thus, in cdPCR, the traditional droplet breaking, that can occur during pipetting or amplification is avoided improving the analytical performances. Despite some fluctuations depending on the gene, the LOD of ddPCR ranges from 0.07 to 0.2 % VAF. BEAMing, currently commercially available as OncoBEAM, is based on the employment of primers covalently bound to magnetic beads. Beads are compartmentalized into aqueous microdroplets where the amplification occurs. After beads purification, fluorescent probes specific for the wild-type or mutated sequences are attached. Each fluorescently labeled bead is then analyzed using a flow cytometer [87]. Compared to ddPCR and cdPCR, the turnaround time for BEAMing is longer (2 days), but offers higher sensitivity (LOD 0.02–0.04 %). In those cases that need be analyzed to analyzed for multiple genes simultaneously, NGS-based platformed are more suited. Currently, NGS platforms can analyze from a few genes to an entire exome or genome [88]. Two are the major issues to solve to achieve an optimal application of NGS to liquid biopsy: i) traditional NGS strategies are not able to detect variants below 1 %, and ii) sequencing errors or DNA polymerase errors can occur, limiting the confidence in calling mutations with VAFs <5 %. To reduce the error rate and ensure mutation calling with VAFs <0.1 %, some improvements as molecular barcoding have been adopted. In this case, unique molecular identifiers (UMI) are added before any PCR step, allowing the recognition of variant alleles present in the original sample errors introduced during library preparation. This approach, used for example in assays like Guardant360, is able to correct amplification and sequencing errors, achieving a sensitivity of 0.05 % allele fraction. In other cases, computational tools and bioinformatic analysis can be used to differentiate true positives from background noise in sequencing. Cancer personalized profiling by deep sequencing (CAPP-Seq) which utilizes DNA probes, designed for regions with high driver mutation frequencies, to hybridize and capture ctDNA is used by AVENIO ctDNA analysis kits (Roche Diagnostics). CAPP-Seq technology reduces the background noise and reaches a LOD of 0.02 % [89].
Fig. 4.
Principal technologies for ctDNA analysis. Technologies for ctDNA analysis can be divided into two main groups: PCR and NGS-based technologies, that differ for LOD, number of analyzed genes ad turnaround time (TAT).
1.4. Circulating tumor cells (CTC) in gastric cancer
CTCs are cancer cells released from the primary tumor into the bloodstream. A small fraction of them can survive in the circulation extravasating and initiating tumor formation in distant sites [90]. Despite not all CTCs having an invasive potential, all of them are suitable biomarkers to better understand the biological features of the shedding tumor. A large number of information can be obtained by CTCs that can be dissected from the transcriptomic, genomic, or proteomic point of view, both by bulk and single-cell analysis [91]. Currently, CTCs are mainly being investigated as prognostic biomarkers. In fact, CTC abundance has been associated with a worse prognosis in several cancer types. A meta-analysis of 14 retrospective studies on GC showed that CTC-positive patients had shorter OS and PFS than CTC-negative ones [92]. Other studies have investigated CTCs as a tool to monitor therapy effects. Despite the limited literature on CTC, all available studies confirmed that the increase of CTCs during longitudinal therapy outcome monitoring is associated with disease progression [93,94]. Recent advancements in analytical platforms have enabled the evaluation of the genomic, transcriptomic, or epigenetic profiles of circulating tumor cells (CTCs) at the single-cell level. Analysis of the transcriptomic profile of single-CTC revealed that most of them showed the overexpression of EMT-related transcription factors (Zinc Finger E-Box Binding Homeobox 2: ZEB2, Myocyte-Specific Enhancer Factor 2D: MEF2D, GATA Binding Protein 1: GATA1, and GATA Binding Protein 2: GATA2) and/or genes involved in chemoresistance (Cyclin Dependent Kinase Inhibitor 1A: CDKN1A and Cyclin Dependent Kinase Inhibitor 1B: CDKN1B). Noteworthy, CTC monitoring was shown to be useful also in evaluating immunotherapy response. Tan and colleagues detected PD‐L1 expression in CTCs from patients affected by different advanced cancers, including gastric or esophagogastric junction cancer, demonstrating that PD‐L1 levels in CTCs may be a clinically actionable marker for immunotherapy, and could predict the therapeutic response [95]. Similarly, CTCs expressing PD‐L1 were longitudinally monitored in unresectable or metastatic GC patients who received anti-HER2 plus anti-PD1 and chemotherapy (according to KEYNOTE-811) [96]. CTCs abundance is extremely limited and few (1–10) cells for milliliter of whole blood are expected in the metastatic setting [97]. In patients with advanced gastric or esophago-gastric junction cancer, according to the results of the PRODIGE 17-ACCORD 20 trial, the median number of detected CTCs, at baseline, was 1 (ranging, from 0 to 415) [98]. Due to their rarity, the isolation of CTCs represents the major challenge. At present, only one method, Cell Search System has been approved by FDA [99]. Cell Search System is based on the use of magnetic beads coated with an antibody against the epithelial marker: Epithelial Cell Adhesion Molecule (EpCAM). This technology presents important limitations, not least, the low or absent expression of EpCAM in several epithelial tumors, so to overcome this important issue and to avoid the bias of a prior knowledge of CTC surface markers, other isolation techniques, based on CTC biophysical proprieties are under evaluation [100]. Nowadays only ctDNA and CTCs have been approved by the FDA as integrative biomarkers in oncological patient management, while several other analytes are currently being investigated at the pre-clinical level, with very promising results [101].
1.5. Circulating cell-free RNA
Besides cfDNA, circulating cell-free RNA (cfRNA) has recently been proposed as an additional biomarker in LB. Despite for many years RNA was considered a complex marker to assess, especially in LB, thanks to technological implementation, messenger RNA present in the circulation (cf-mRNA) is gradually emerging also in LB [102]. Circulating RNA is a highly dynamic marker able to reflect temporal changes in organ-specific transcripts and with high informative potential. Moreover, as reported in the tissue context, the employment of cfRNA in LB is more suitable for gene fusion detection than cfDNA [103]. Although cfRNA appears to be a highly promising marker, particularly in the context of gene fusions or rearrangements (FGFR2, NTRK1/2/3, Anaplastic Lymphoma Receptor Tyrosine Kinase: ALK, RET Receptor Tyrosine Kinase: RET) that are considered druggable alterations, it poses a significant challenge. In fact, as for TB, RNA is less stable than DNA and more amenable to degradation. Moreover, cfRNA is a mixture of full-length RNA, fragmented RNA, non-coding RNAs, and ribosomal RNA [104]. In the circulating context, the latter, represents the most abundant RNA component that can overpower the signal of the more informative mRNA component. Studies investigating circulating transcriptome in LB are still limited, and most of them are focused on non-coding RNAs, which were historically considered less prone to degradation.
1.5.1. Circulating non-coding RNAs as markers in gastric cancer
Non-coding RNAs (ncRNAs) are a heterogeneous class of RNA-based molecules, that based on their length, are divided into two major categories: small ncRNAs (smaller than 200 nucleotides) and long ncRNAs (lncRNAs) (longer than 200 nucleotides). Non-coding RNAs are generally very stable in different body fluids thanks to i) the packing into biological vesicles (e.g. exosomes, microvesicles, and apoptotic bodies) or ii) the binding with lipoproteins and RNA-binding proteins [105]. So far, microRNAs (miRNAs), together with lncRNA, are the most investigated RNA-based biomarkers in LB, mainly for diagnostic and screening purposes.
1.5.2. Circulating miRNAs in gastric cancer
MiRNAs are single-strand sequences of 18–25 nucleotides in lengths that can regulate RNA translation through the binding with the 3’ untranslated region of the messenger RNAs. The volume of research dedicated to investigating miRNAs in GC is substantial, with the predominant focus being the identification of prospective diagnostic markers (Table 3). In a meta-analysis, all studies examining miRNAs as early diagnostic markers in GC were analyzed to identify the most frequently modulated miRNAs. MiRNAs presenting an expression pattern more reproducible among different studies were miR-20a, miR-223, and miR-421, and the oncomiRNA miR-21 which was tested in 13 works investigating deregulated miRNAs in GC-LBs, and in all of them reported as upregulated [106]. The major limitation of circulating miRNAs as biomarkers arises from the low concordance among studies, primarily stemming from the lack of standardization in analysis methods. As reported in Table 2, plasma and serum are indifferently employed, without considering that coagulation processes may affect the miRNA profile. That, associated with other technical issues, including blood centrifugation protocol, the extraction protocol, and the assay employed for miRNA detection, may affect final miRNA quantitation. Furthermore, the most extensively studied miRNAs exhibit physiological expression even in the absence of neoplastic conditions. In many instances, the pathological state is distinguished by an increased rate of miRNA release, which is however also, expressed in a non-pathological condition. Thus, studies aimed to identify the range of physiological levels of GC-related miRNAs are needed to create a baseline for each miRNA taking into account intra-patient variability (gender, age, ethnicity), and possible confounding factors (e.g. temporary inflammatory states, comorbidities). Given that miRNA profiles can be influenced by various physiological and pathological factors, utilizing a miRNA signature, consisting of sets of two or more miRNAs, may offer greater reliability in identifying GC. However, considering that i) several miRNA-based therapeutics are currently being tested in early-stage clinical trials and ii) miRNAs could be used as companion diagnostic biomarkers for these upcoming therapeutic molecules, a standardization of analytical protocols for miRNA quantification in LB is needed.
Table 3.
List of miRNAs modulated in the blood of GC patients.
| MicroRNA signature | Source | Features | Reference |
|---|---|---|---|
| hsa-miR-320a, hsa-miR-1260b, and hsa-miR-6515-5p | Serum | The signature distinguishes GC patients from non-cancer controls | [107] |
| hsa-miR-221-3p, hsa-miR-376c-3p, and hsa-miR-744-5p | Serum | The signature is upregulated in early GC patients | [108] |
| hsa-miR-106b and hsa- miR-21 | Serum | The signature differentiates high-risk from low-risk patients after H. pylori eradication | [109] |
| hsa-miR-21, hsa-miR-27a, hsa-miR-106b, hsa-miR-146a, hsa-miR-148a, hsa-miR-223, and hsa-miR-433 | Serum | The signature predicts the presence of lymph node metastasis in GC patients | [110] |
| hsa-miR-130a, hsa-miR-331, hsa-miR-19a, hsa- hsa-miR-223, hsa-miR-106a, miR-21, and hsa-miR-374 | Serum | The signature distinguishes gastric cancer patients from non-cancer controls | [111] |
| hsa-miR-21 | Serum | The miRNA is over-expressed in gastric cancer patients compared with non-cancer controls | [112] |
| hsa-miR-1, hsa-miR-20a, hsa-miR-27a, hsa-miR-34 and hsa-miR-423-5p | Serum | The signature is modulated in GC patients compared with non-cancer controls | [113] |
| hsa-miR-4257, hsa-miR-6785-5p, hsa-miR-187-5p, and hsa-miR-5739 | Serum | The signature can be considered a diagnostic marker to detect GC | [114] |
| hsa-miR-18a, hsa-miR-181b, and hsa-miR-335 | Serum | The signature is considered a diagnostic marker for early GC detection | [115] |
| hsa-miR-26a, hsa-miR-142-3p, hsa-miR-148a, and hsa-miR-195 | Plasma | The signature is down-regulated in GC patients compared with non-cancer controls | [116] |
| hsa-miR-17-5p, hsa-miR-21, hsa-miR-106a, hsa-miR-106b | Plasma | The signature is over-expressed in gastric cancer patients compared with non-cancer controls | [117] |
| hsa-miR-21, hsa-miR-93, hsa-miR-106a and hsa-miR-106b | Plasma | The signature distinguishes GC patients from non-cancer controls | [118] |
| hsa-miR-425-5p, hsa-miR-1180-3p, hsa-miR-122-5p, hsa-miR-24-3p, hsa-miR-4632-5p | Plasma | The signature is considered a marker for GC early diagnosis | [119] |
1.5.3. Circulating lncRNAs in gastric cancer
LncRNAs are transcripts longer than 200 nucleotides and are mostly transcribed by RNA polymerase II (Pol II). From a structural point of view, lncRNAs present similar features to that of mRNAs (5′ cap, exons, and poly-A tail); however, they cannot be translated into functional proteins, but act as gene expression regulators at transcriptional, posttranscriptional, and translational level. As with other non-coding RNAs, lncRNAs can be detected in the blood and could be potentially used as possible non-invasive cancer biomarkers. A meta-analysis conducted on 42 studies spanning from 2013 to 2020, focusing on circulating lncRNAs in GC, revealed a pooled sensitivity of 78 % and specificity of 75 %. These findings suggest that lncRNAs hold promise as potential non-invasive biomarkers for GC [120]. The list of dysregulated lncRNAs included 49 different lncRNAs, examined across all included studies; most of them upregulated in GC patients compared with healthy donors. The up-regulated lncRNA H19 in GC stood out for its reported sensitivity of 90.9 % and specificity of 100 %, highlighting its potential as a highly accurate biomarker. Other lncRNAs such as CTC-501O10.1, GC1, INHBASS1, and AK001058 exhibited high sensitivity ranging from 97 % to 90 %, but their specificity was comparatively lower, ranging from 83 % to 51 %. However, the metanalysis presented two biases, the first one concerns the study population, in fact, of 42 studies 37 were conducted on the Asiatic population, while only 5 included Caucasic patients. Another potential bias arises from the variety of sample sources investigated, including plasma, serum, and plasma/serum exosomes. Few studies have explored the prognostic role of circulating lncRNAs, such as the aforementioned H19. Despite H19's strong diagnostic potential, its levels do not appear to correlate with tumor progression [121]. High levels of lncRNA MIAT in the serum of GC patients correlate with a worse prognosis and shorter survival, proposing it as an independent prognostic marker for GC [122].
1.6. Circulating extracellular vesicles as markers in gastric cancer
Extracellular vesicles (EVs) are biological particles limited by a lipid bilayer, released by cells into biological fluids, which play a crucial role in intercellular communication. Based on their dimensions and biological origin, they can be divided into three main classes: microvesicles, exosomes, and apoptotic bodies. EVs are stable entities that can be obtained with non-invasive methods and most importantly they contain several biomarkers including DNA, RNA (both coding and non-coding RNA), and proteins. Thus, the analysis of circulating extracellular vesicles (cEVs) cargos may be a new frontier in liquid biopsy applied to oncological practice. Several studies demonstrated that GC-derived EVs had a key role not only in the first stages of tumor formation and proliferation but, also, in invasion, metastasis, and drug resistance [123]. A great body of literature now exists on investigating circulating miRNAs, lncRNAs, and more recently circRNAs derived from exosomes as diagnostic and prognostic markers in GC [124,125]. Despite the limited number of studies, also proteins, contained in the cEVs, emerge as an exceptionally intriguing marker. They constitute one of the most abundant components of exosomes and hold considerable promise for diagnostic and prognostic purposes. Several tumor-related proteins have been described in exosomes isolated from the blood of GC patients. A key area of interest in the GC setting is the ability to analyze critical protein markers, which are typically evaluated in FFPE tissue. For instance, Li et al. quantified exosomal-HER2 levels by ELISA (Enzyme-Linked ImmunoSorbent Assay) to assess HER2 status and predict the response to trastuzumab-based therapy [126]. Despite sensitivity (66.7 %) and specificity (74.2 %) needs to be improved, the quantification of exosomal HER2 levels may be useful to monitor trastuzumab-based treatment efficacy. Although data are still highly inconclusive, also exosomal PD-L1 may be considered a possible prognostic marker. Fan et al. demonstrated that circulating exosomal PD-L1 was significantly associated with the tumor stage and correlated with OS [127]. The primary limitation in utilizing exosomes as markers in diagnostics remains the lack of efficient and standardized exosome isolation methods in LB [128].
1.7. GC biomarkers in other biological fluids
Various body fluids offer opportunities for analyzing diagnostic, prognostic, and predictive biomarkers, including urine, saliva, bile, cerebrospinal fluid, pleural ascites, peritoneal fluid, amniotic fluid, breast milk, tears, synovial fluid, breath, and seminal fluid. While some fluids are attractive for their minimal invasiveness, others, though less accessible, may be in direct contact with the tumor, potentially harboring higher levels of biomarkers [129]. Indeed, while blood and its derivatives are typically considered the gold standard, the selection of alternative fluids is often guided by tumor localization. In the context of GC, alternative sources include gastric juice, gastric washes, or ascitic fluids in cases of advanced neoplasms. Despite the more invasive procedures needed to obtain gastric juice, it is considered a promising source because biomarkers are directly secreted from cells, bypassing the elimination by the liver [130]. Although the analysis of gastric juice may be challenging, due to low pH levels, some literature is available on the topic. Gareev et al. identified three miRNAs: miR-129-1-3p, miR-129-2-3p, and miR-421 as candidate GC biomarkers [131]. MicroRNA-21 represents a controversial case: it was reported up-regulated in blood by several studies, while in gastric juice, it was found to be down-regulated [132]. The lncRNA LINC00152 was successfully measured in gastric juice, reporting a differential expression between GC patients and healthy controls [133].
1.8. Current applications of liquid biopsy in clinical practice
LB is well established in the routine clinical practice for the management of NSCLC, but it also finds application in CRC, BC, and other cancers (46). Currently, the FDA approved five LB-based tests, mostly addressed to NSCLC, CRC, prostate cancer (PCa), and BC (Table 4) [134]. The first LB-based approved test is Cobas EGFR mutation Test v2 (Roche Diagnostics, 2016). The test allows the detection of exon 19 deletions or L858R mutation in exon 21 of the EGFR gene by a quantitative PCR (qPCR) analysis using ctDNA isolated from NSCLC patients [135]. Despite this test has revolutionized the history of ctDNA analysis, the number of detectable alterations is limited. Thanks to the introduction of novel sequencing techniques and the improvements in their LOD, in 2020, next-generation sequencing (NGS)-based LB tests were approved, including the Guardant360 CDx and FoundationOne Liquid CDx, both based on hybridization-based capture NGS [136]. The Guardant360 CDx can identify genetic alterations in 55 clinically relevant genes and is FDA-approved as a companion diagnostic for the identification of EGFR (exon 19 and 20), ERBB2/HER2 (SNVs and exon 20 insertions), and KRAS (G12C mutation) genes alterations in NSCLC and Estrogen Receptor 1 (ESR1) missense mutations in BC. FoundationOne Liquid CDx analyzed alterations in more than 300 pan-cancer genes and is currently approved as a companion diagnostic to identify patients with EGFR, ALK, MET, and Proto-Oncogene Tyrosine-Protein Kinase ROS (ROS1) alterations in NSCLC. More recently, other applications were added including advanced metastatic PCa cancer (Breast Cancer gene 1: BRCA1 and Breast Cancer gene 2: BRCA2 mutations), BC (phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit alpha: PIK3CA mutations), and metastatic CRC (Ataxia Telangiectasia Mutated: ATM and BRCA1/BRCA2) [[137], [138], [139]]. For BC and CRC, other tests are currently approved by the FDA: Therascreen PIK3CA RGQ PCR, a real-time qualitative PCR test for the detection of 11 mutations in the PIK3CA gene (BC patients), and Therascreen KRAS RGQ PCR Kit for the search of KRAS G12C mutation (CRC and NSCLC patients). Moreover, in CRC-affected subjects, Epi proColon can employed to analyze the ctDNA methylation profile of Septin 9 (SEPT9) promoter using real-time PCR [140].
Table 4.
List of FDA-approved tests, based on liquid biopsy.
| Name of test | Technique | Gene of interest | Type of cancer | Application |
|---|---|---|---|---|
| Cobas EGFR mutation Test v2 | qPCR | EGFR exons 19 and 21 | NSCLC | Erlotinib/osimertinib (EGFR) |
| Guardant360 CDx | NGS | 55 clinically relevant genes | NSCLC | Osimertinib/amivantamab (EGFR) |
| Breast cancer | Trastuzumab deruxtecan (ERBB2) | |||
| Sotorasib (KRAS) | ||||
| Elacestrant (ESR1) | ||||
| FoundationOne Liquid CDx | NGS | 300 pan-cancer genes | NSCLC, PCa BC, CRC | Niraparib/olaparib (BRCA1-2) |
| Encorafenib/ceritinib (BRAF) | ||||
| Osimertinib (EGFR) | ||||
| Alectinib/brigatinib/crizotinib/ceritinib (ALK) | ||||
| Capmatinib (MET) | ||||
| Entrectinib (ROS1) | ||||
| Therascreen PIK3CA RGQ PCR | Real-Time qPCR | PIK3CA | Breast cancer | Alpelisib (PIK3CA) |
| Epi proColon | Real-Time PCR | Methylation of SEPT9 promoter | CRC | Early detection of CRC |
1.9. Novel potential application for liquid biopsy
Considering the success of immunotherapy in several solid neoplasms, even more studies are focused on the evaluation of microsatellite status and/or the evaluation of the molecular tumor burden (TMB) in ctDNA. Most of the available ctDNA-based NGS tests are nowadays, able to detect MSI, with a good concordance with the respective FFPE tissue. For example, within the GOZILA study, performed on advanced GI cancers, underwent both plasma and tissue testing for MSI, the overall percent agreement was 98.2 %. Recently the TMB has become one of the primary markers to assess, to evaluate patients’ eligibility for ICI treatment. There is so, a need for the evaluation of the TMB with no invasive techniques: blood tumor mutational burden (bTMB) [141]. Jensen and colleagues proposed a low-coverage, genome-wide sequencing of cfDNA, to detect copy-number alterations (CNAs) in ctDNA for the calculation of a score, defined genomic instability number (GIN) [142]. GIN can capture genome-wide alterations in the circulating genome and its pattern during treatment was shown able to predict the responsiveness to immunotherapy.
2. Conclusions and future perspectives
Liquid biopsy nowadays has become a robust technique, that at least as far as ctDNA, is already part of clinical practice in some solid tumors. None of the current guidelines suggest the application of LB in GC, despite literature suggesting good i) shedding rate and ii) concordance with the respective TB, for advanced GCs. The employment of ctDNA for the treatment monitoring (PANGEA) or as criteria to enroll patients (FIGHT) in clinical trials, indicates that the time has come for the introduction of LB in the management of advanced GC. Stated that TB remains essential for the initial diagnosis, LB may successfully adjuvate TB in the therapy definition, overcoming the well-established spatial heterogeneity that characterizes GC and allowing real-time monitoring of tumor progression. However, not all centers are yet ready to approach LB, both because of the required technological platforms and the correct personnel training. The methylation profile in ctDNA and other circulating markers (cfRNA and CTC) which are providing intriguing results at the pre-clinical level, must be fine-tuned, to obtain a multiple-level and a more compressive non-invasive neoplasm characterization.
Author contributions
All authors equally contributed in writing and conceptualization of the manuscript. All authors have read and agreed to the published version of the manuscript.
Ethical approval
The proposed article is a review if the available literature, no ethical approval is required.
Funding
This review did not receive any funding.
Declaration of interest statement
I report research funding (to institution) from Roche, Astellas, and Diaceutics; personal honoraria as an invited speaker from Roche, Astellas, AstraZeneca, Lilly, Incyte, Bristol Myers Squibb, Agilent, Merck Serono, Pierre Fabre, GlaxoSmithKline, Novartis, and Amgen; and for participation in advisory boards of Amgen, Astellas, Roche, Merck Serono, GlaxoSmithKline, Novartis, and Janssen.
Acknowledgements
None.
References
- 1.Lone S.N., Nisar S., Masoodi T., Singh M., Rizwan A., Hashem S., et al. Liquid biopsy: a step closer to transform diagnosis, prognosis and future of cancer treatments. Mol Cancer. 2022;21(1):79. doi: 10.1186/s12943-022-01543-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Marrugo-Ramírez J., Mir M., Samitier J. Blood-based cancer biomarkers in liquid biopsy: a promising non-invasive alternative to tissue biopsy. Int J Mol Sci. 2018;19(10):2877. doi: 10.3390/ijms19102877. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.The Cancer Genome Atlas Research Network Comprehensive molecular characterization of gastric adenocarcinoma. Nature. 2014;513(7517):202–209. doi: 10.1038/nature13480. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Fassan M., Scarpa A., Remo A., De Maglio G., Troncone G., Marchetti A., et al. Current prognostic and predictive biomarkers for gastrointestinal tumors in clinical practice. Pathologica. 2020;112(3):248–259. doi: 10.32074/1591-951X-158. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Rajdev L., Kennedy E.B., Shah M.A. Immunotherapy and targeted therapy for advanced gastroesophageal cancer: ASCO guideline Q and A. JCO Oncol Pract. 2023;19(4):197–200. doi: 10.1200/OP.22.00818. [DOI] [PubMed] [Google Scholar]
- 6.Shitara K., Fleitas T., Kawakami H., Curigliano G., Narita Y., Wang F., et al. Pan-Asian adapted ESMO Clinical Practice Guidelines for the diagnosis, treatment and follow-up of patients with gastric cancer. ESMO Open. 2024;9(2) doi: 10.1016/j.esmoop.2023.102226. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Bang Y.J., Van Cutsem E., Feyereislova A., Chung H.C., Shen L., Sawaki A., et al. Trastuzumab in combination with chemotherapy versus chemotherapy alone for treatment of HER2-positive advanced gastric or gastro-oesophageal junction cancer (ToGA): a phase 3, open-label, randomised controlled trial. Lancet. 2010;376(9742):687–697. doi: 10.1016/S0140-6736(10)61121-X. [DOI] [PubMed] [Google Scholar]
- 8.Grillo F., Fassan M., Sarocchi F., Fiocca R., Mastracci L. HER2 heterogeneity in gastric/gastroesophageal cancers: from benchside to practice. World J Gastroenterol. 2016;22(26):5879. doi: 10.3748/wjg.v22.i26.5879. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Malapelle U., Parente P., Pepe F., De Luca C., Pisapia P., Sgariglia R., et al. Evaluation of micro satellite instability and mismatch repair status in different solid tumors: a multicenter analysis in a real world setting. Cells. 2021;10(8):1878. doi: 10.3390/cells10081878. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Wu H., Ma W., Jiang C., Li N., Xu X., Ding Y., et al. Heterogeneity and adjuvant therapeutic approaches in MSI-H/dMMR resectable gastric cancer: emerging trends in immunotherapy. Ann Surg Oncol. 2023;30(13):8572–8587. doi: 10.1245/s10434-023-14103-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Tan D.S., Kim Y.M., Lim M.C., Sho M., Lu C.H., Nagao S., et al. 101P Real-world prevalence of MSI-H/dMMR across 6 different tumor types in Asia. Ann Oncol. 2022;33 [Google Scholar]
- 12.Shah M.A., Shitara K., Ajani J.A., Bang Y.J., Enzinger P., Ilson D., et al. Zolbetuximab plus CAPOX in CLDN18.2-positive gastric or gastroesophageal junction adenocarcinoma: the randomized, phase 3 GLOW trial. Nat Med. 2023;29(8):2133–2141. doi: 10.1038/s41591-023-02465-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Shitara K., Lordick F., Bang Y.J., Enzinger P., Ilson D., Shah M.A., et al. Zolbetuximab plus mFOLFOX6 in patients with CLDN18.2-positive, HER2-negative, untreated, locally advanced unresectable or metastatic gastric or gastro-oesophageal junction adenocarcinoma (SPOTLIGHT): a multicentre, randomised, double-blind, phase 3 trial. Lancet Lond Engl. 2023;401(10389):1655–1668. doi: 10.1016/S0140-6736(23)00620-7. [DOI] [PubMed] [Google Scholar]
- 14.Wainberg Z.A., Enzinger P.C., Kang Y.K., Qin S., Yamaguchi K., Kim I.H., et al. Bemarituzumab in patients with FGFR2b-selected gastric or gastro-oesophageal junction adenocarcinoma (FIGHT): a randomised, double-blind, placebo-controlled, phase 2 study. Lancet Oncol. 2022;23(11):1430–1440. doi: 10.1016/S1470-2045(22)00603-9. [DOI] [PubMed] [Google Scholar]
- 15.Wainberg Z.A., Kang Y.K., Lee K.W., Qin S., Yamaguchi K., Kim I.H., et al. Bemarituzumab as first-line treatment for locally advanced or metastatic gastric/gastroesophageal junction adenocarcinoma: final analysis of the randomized phase 2 FIGHT trial. Gastric Cancer. 2024;27(3):558–570. doi: 10.1007/s10120-024-01466-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Pellino A., Brignola S., Riello E., Niero M., Murgioni S., Guido M., et al. Association of CLDN18 protein expression with clinicopathological features and prognosis in advanced gastric and gastroesophageal junction adenocarcinomas. J Personalized Med. 2021;11(11):1095. doi: 10.3390/jpm11111095. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Stahl P., Seeschaaf C., Lebok P., Kutup A., Bockhorn M., Izbicki J.R., et al. Heterogeneity of amplification of HER2, EGFR, CCND1 and MYC in gastric cancer. BMC Gastroenterol. 2015;15(1):7. doi: 10.1186/s12876-015-0231-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Chao J., Bedell V., Lee J., Li M.S., Chu P., Yuan Y.C., et al. Association between spatial heterogeneity within nonmetastatic gastroesophageal adenocarcinomas and survival. JAMA Netw Open. 2020;3(4) doi: 10.1001/jamanetworkopen.2020.3652. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Kim H.D., Choi E., Shin J., Lee I.S., Ko C.S., Ryu M.H., et al. Clinicopathologic features and prognostic value of claudin 18.2 overexpression in patients with resectable gastric cancer. Sci Rep. 2023;13(1) doi: 10.1038/s41598-023-47178-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Choi E., Shin J., Ryu M.H., Kim H.D., Park Y.S. Heterogeneity of claudin 18.2 expression in metastatic gastric cancer. Sci Rep. 2024;14(1) doi: 10.1038/s41598-024-68411-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Han N., Kim M.A., Lee H.S., Kim W.H. Evaluation of fibroblast growth factor receptor 2 expression, heterogeneity and clinical significance in gastric cancer. Pathobiology. 2015;82(6):269–279. doi: 10.1159/000441149. [DOI] [PubMed] [Google Scholar]
- 22.Yang J.H., Kim H., Roh S.Y., Lee M.A., Park J.M., Lee H.H., et al. Discordancy and changes in the pattern of programmed death ligand 1 expression before and after platinum-based chemotherapy in metastatic gastric cancer. Gastric Cancer. 2019;22(1):147–154. doi: 10.1007/s10120-018-0842-x. [DOI] [PubMed] [Google Scholar]
- 23.Ahn S., Ahn S., Van Vrancken M., Lee M., Ha S.Y., Lee H., et al. Ideal number of biopsy tumor fragments for predicting HER2 status in gastric carcinoma resection specimens. Oncotarget. 2015;6(35):38372–38380. doi: 10.18632/oncotarget.5368. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Pectasides E., Stachler M.D., Derks S., Liu Y., Maron S., Islam M., et al. Genomic heterogeneity as a barrier to precision medicine in gastroesophageal adenocarcinoma. Cancer Discov. 2018;8(1):37–48. doi: 10.1158/2159-8290.CD-17-0395. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Xie W., Suryaprakash S., Wu C., Rodriguez A., Fraterman S. Trends in the use of liquid biopsy in oncology. Nat Rev Drug Discov. 2023;22(8):612–613. doi: 10.1038/d41573-023-00111-y. [DOI] [PubMed] [Google Scholar]
- 26.Pasini L., Ulivi P. Liquid biopsy for the detection of resistance mechanisms in NSCLC: comparison of different blood biomarkers. J Clin Med. 2019;8(7):998. doi: 10.3390/jcm8070998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Bidard F.C., Hardy-Bessard A.C., Dalenc F., Bachelot T., Pierga J.Y., De La Motte Rouge T., et al. Switch to fulvestrant and palbociclib versus no switch in advanced breast cancer with rising ESR1 mutation during aromatase inhibitor and palbociclib therapy (PADA-1): a randomised, open-label, multicentre, phase 3 trial. Lancet Oncol. 2022;23(11):1367–1377. doi: 10.1016/S1470-2045(22)00555-1. [DOI] [PubMed] [Google Scholar]
- 28.Russo A., Incorvaia L., Del Re M., Malapelle U., Capoluongo E., Gristina V., et al. The molecular profiling of solid tumors by liquid biopsy: a position paper of the AIOM-SIAPEC-IAP-SIBioC-SIC-SIF Italian Scientific Societies. ESMO Open. 2021;6(3) doi: 10.1016/j.esmoop.2021.100164. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Sánchez-Herrero E., Serna-Blasco R., Robado De Lope L., González-Rumayor V., Romero A., Provencio M. Circulating tumor DNA as a cancer biomarker: an overview of biological features and factors that may impact on ctDNA analysis. Front Oncol. 2022;12 doi: 10.3389/fonc.2022.943253. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Stroun M., Lyautey J., Lederrey C., Olson-Sand A., Anker P. About the possible origin and mechanism of circulating DNA. Clin Chim Acta. 2001;313(1–2):139–142. doi: 10.1016/s0009-8981(01)00665-9. [DOI] [PubMed] [Google Scholar]
- 31.Parisi C., Tagliamento M., Belcaid L., Aldea M., Bayle A., Remon-Masip J., et al. Circulating tumor DNA in clinical trials for solid tumors: challenges and current applications. J Liq Biopsy. 2023;1 [Google Scholar]
- 32.Fang W., Lan Y., Huang K., Liu C., Hung Y., Lin C., et al. Clinical significance of circulating plasma DNA in gastric cancer. Int J Cancer. 2016;138(12):2974–2983. doi: 10.1002/ijc.30018. [DOI] [PubMed] [Google Scholar]
- 33.Vasseur D., Sassi H., Bayle A., Tagliamento M., Besse B., Marzac C., et al. Next-generation sequencing on circulating tumor DNA in advanced solid cancer: Swiss army knife for the molecular tumor board? A review of the literature focused on FDA approved test. Cells. 2022;11(12):1901. doi: 10.3390/cells11121901. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Zhang Y., Yao Y., Xu Y., Li L., Gong Y., Zhang K., et al. Pan-cancer circulating tumor DNA detection in over 10,000 Chinese patients. Nat Commun. 2021;12(1):11. doi: 10.1038/s41467-020-20162-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Danesi R., Lo Y.M.D., Oellerich M., Beck J., Galbiati S., Re M.D., et al. What do we need to obtain high quality circulating tumor DNA (ctDNA) for routine diagnostic test in oncology? – Considerations on pre-analytical aspects by the IFCC workgroup cfDNA. Clin Chim Acta. 2021;520:168–171. doi: 10.1016/j.cca.2021.05.033. [DOI] [PubMed] [Google Scholar]
- 36.Rodon Font N., No Garbarino Y., Díaz Castello O., Moya Amoros J., Barrios Sánchez P., Coroleu Lletget D., et al. Concordance analysis between liquid biopsy (ctDNA) and tumor DNA molecular profiles from panel-based next-generation sequencing. Rev Española Patol. 2022;55(3):156–162. doi: 10.1016/j.patol.2022.01.001. [DOI] [PubMed] [Google Scholar]
- 37.Kung C.Y., Fang W.L., Hung Y.P., Huang K.H., Chen M.H., Chao Y., et al. Comparison of the mutation patterns between tumor tissue and cell-free DNA in stage IV gastric cancer. Aging. 2023;15(3):777–790. doi: 10.18632/aging.204512. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Lee J., Kim S.T., Kim K., Lee H., Kozarewa I., Mortimer P.G.S., et al. Tumor genomic profiling guides patients with metastatic gastric cancer to targeted treatment: the VIKTORY umbrella trial. Cancer Discov. 2019;9(10):1388–1405. doi: 10.1158/2159-8290.CD-19-0442. [DOI] [PubMed] [Google Scholar]
- 39.Wang H., Li B., Liu Z., Gong J., Shao L., Ren J., et al. HER2 copy number of circulating tumour DNA functions as a biomarker to predict and monitor trastuzumab efficacy in advanced gastric cancer. Eur J Cancer. 2018;88:92–100. doi: 10.1016/j.ejca.2017.10.032. [DOI] [PubMed] [Google Scholar]
- 40.Maron S.B., Chase L.M., Lomnicki S., Kochanny S., Moore K.L., Joshi S.S., et al. Circulating tumor DNA sequencing analysis of gastroesophageal adenocarcinoma. Clin Cancer Res. 2019;25(23):7098–7112. doi: 10.1158/1078-0432.CCR-19-1704. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Shitara K., Bang Y.J., Iwasa S., Sugimoto N., Ryu M.H., Sakai D., et al. Trastuzumab deruxtecan in previously treated HER2-positive gastric cancer. N Engl J Med. 2020;382(25):2419–2430. doi: 10.1056/NEJMoa2004413. [DOI] [PubMed] [Google Scholar]
- 42.Shitara K., Bang Y.J., Iwasa S., Sugimoto N., Ryu M.H., Sakai D., et al. Trastuzumab deruxtecan in HER2-positive advanced gastric cancer: exploratory biomarker analysis of the randomized, phase 2 DESTINY-Gastric01 trial. Nat Med. 2024;30(7):1933–1942. doi: 10.1038/s41591-024-02992-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Jogo T., Nakamura Y., Shitara K., Bando H., Yasui H., Esaki T., et al. Circulating tumor DNA analysis detects FGFR2 amplification and concurrent genomic alterations associated with FGFR inhibitor efficacy in advanced gastric cancer. Clin Cancer Res. 2021;27(20):5619–5627. doi: 10.1158/1078-0432.CCR-21-1414. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.He W., Yang J., Sun X., Jiang S., Jiang J., Liu M., et al. Advantages and limitations of monitoring circulating tumor DNA levels to predict the prognosis of patients diagnosed with gastric cancer. Biomark Insights. 2022;17 doi: 10.1177/11772719221141525. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Chen C., Chen Q., Zhao Q., Liu M., Guo J. Value of combined detection of serum CEA, CA72-4, CA19-9, CA15-3 and CA12-5 in the diagnosis of gastric cancer. Ann Clin Lab Sci. 2017;47(3):260–263. [PubMed] [Google Scholar]
- 46.Qian C., Ju S., Qi J., Zhao J., Shen X., Jing R., et al. Alu-based cell-free DNA: a novel biomarker for screening of gastric cancer. Oncotarget. 2017;8(33):54037–54045. doi: 10.18632/oncotarget.11079. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Kim YS. Quantitative analysis of cell-free DNA in the plasma of gastric cancer patients. Oncol Lett 3(4):921-926. [DOI] [PMC free article] [PubMed]
- 48.Moati E., Taly V., Garinet S., Didelot A., Taieb J., Laurent-Puig P., et al. Role of circulating tumor DNA in gastrointestinal cancers: current knowledge and perspectives. Cancers. 2021;13(19):4743. doi: 10.3390/cancers13194743. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Cohen J.D., Li L., Wang Y., Thoburn C., Afsari B., Danilova L., et al. Detection and localization of surgically resectable cancers with a multi-analyte blood test. Science. 2018;359(6378):926–930. doi: 10.1126/science.aar3247. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Park KU, Lee HE, Park DJ, Jung EJ, Song J, Kim HH, et al. MYC quantitation in cell-free plasma DNA by real-time PCR for gastric cancer diagnosis. Clin Chem Lab Med 47(5):530-536. [DOI] [PubMed]
- 51.Yang J., Gong Y., Lam V.K., Shi Y., Guan Y., Zhang Y., et al. Deep sequencing of circulating tumor DNA detects molecular residual disease and predicts recurrence in gastric cancer. Cell Death Dis. 2020;11(5):346. doi: 10.1038/s41419-020-2531-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Kim K., Shin D.G., Park M.K., Baik S.H., Kim T.H., Kim S., et al. Circulating cell-free DNA as a promising biomarker in patients with gastric cancer: diagnostic validity and significant reduction of cfDNA after surgical resection. Ann Surg Treat Res. 2014;86(3):136. doi: 10.4174/astr.2014.86.3.136. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Roșu M.C., Ardelean A., Moldovan S.D., Faur F.I., Nesiu A., Totoloci B.D. The importance of CA 72-4 and CA 19-9 dosing in gastric cancer. J Med Life. 2023;16(2):186–188. doi: 10.25122/jml-2022-0173. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Wang Y., Zhao C., Chang L., Jia R., Liu R., Zhang Y., et al. Circulating tumor DNA analyses predict progressive disease and indicate trastuzumab-resistant mechanism in advanced gastric cancer. EBioMedicine. 2019;43:261–269. doi: 10.1016/j.ebiom.2019.04.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Kim S.T., Cristescu R., Bass A.J., Kim K.M., Odegaard J.I., Kim K., et al. Comprehensive molecular characterization of clinical responses to PD-1 inhibition in metastatic gastric cancer. Nat Med. 2018;24(9):1449–1458. doi: 10.1038/s41591-018-0101-z. [DOI] [PubMed] [Google Scholar]
- 56.Assi T., Khoury R., Ibrahim R., Baz M., Ibrahim T., Le Cesne A. Overview of the role of liquid biopsy in cancer management. Transl Oncol. 2023;34 doi: 10.1016/j.tranon.2023.101702. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Openshaw M.R., Ottolini B., Richards C.J., Guttery D., Shaw J.A., Thomas A. Developing the liquid biopsy in gastroesophageal adenocarcinoma: disease monitoring and detection of minimal residual disease. Ann Oncol. 2018;29 [Google Scholar]
- 58.Luo H., Wei W., Ye Z., Zheng J., Xu R hua. Liquid biopsy of methylation biomarkers in cell-free DNA. Trends Mol Med. 2021;27(5):482–500. doi: 10.1016/j.molmed.2020.12.011. [DOI] [PubMed] [Google Scholar]
- 59.Locke W.J., Guanzon D., Ma C., Liew Y.J., Duesing K.R., Fung K.Y.C., et al. DNA methylation cancer biomarkers: translation to the clinic. Front Genet. 2019;10:1150. doi: 10.3389/fgene.2019.01150. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Barefoot M.E., Loyfer N., Kiliti A.J., McDeed A.P., Kaplan T., Wellstein A. Detection of cell types contributing to cancer from circulating, cell-free methylated DNA. Front Genet. 2021;12 doi: 10.3389/fgene.2021.671057. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Klein E.A., Richards D., Cohn A., Tummala M., Lapham R., Cosgrove D., et al. Clinical validation of a targeted methylation-based multi-cancer early detection test using an independent validation set. Ann Oncol. 2021;32(9):1167–1177. doi: 10.1016/j.annonc.2021.05.806. [DOI] [PubMed] [Google Scholar]
- 62.Balgkouranidou I., Karayiannakis A., Matthaios D., Bolanaki H., Tripsianis G., Tentes A.A., Lianidou E., Chatzaki E., Fiska A., Lambropoulou M., Kolios G., Kakolyris S. Assessment of SOX17 DNA methylation in cell free DNA from patients with operable gastric cancer. Association with prognostic variables and survival. Clin Chem Lab Med. 2013;51(7):1505–1510. doi: 10.1515/cclm-2012-0320. [DOI] [PubMed] [Google Scholar]
- 63.Ling Z.Q., Lv P., Lu X.X., Yu J.L., Han J., Ying L.S., et al. Circulating methylated XAF1 DNA indicates poor prognosis for gastric cancer. Peng D, curatore. PLoS One. 2013;8(6) doi: 10.1371/journal.pone.0067195. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Hibi K., Goto T., Shirahata A., Saito M., Kigawa G., Nemoto H., et al. Detection of TFPI2 methylation in the serum of gastric cancer patients. Anticancer Res. 2011;31(11):3835–3838. [PubMed] [Google Scholar]
- 65.Lee T.L., Leung W.K., Chan M.W.Y., Ng E.K.W., Tong J.H.M., Lo K.W., et al. Detection of gene promoter hypermethylation in the tumor and serum of patients with gastric carcinoma. Clin Cancer Res Off J Am Assoc Cancer Res. 2002;8(6):1761–1766. [PubMed] [Google Scholar]
- 66.Abbaszadegan M.R., Moaven O., Sima H.R., Ghafarzadegan K., A’rabi A., Forghani M.N., et al. p16 promoter hypermethylation: a useful serum marker for early detection of gastric cancer. World J Gastroenterol. 2008;14(13):2055. doi: 10.3748/wjg.14.2055. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Hu X., Ling Z., Hong L., Yu Q., Li P., Ling Z. Circulating methylated THBS1 DNAs as a novel marker for predicting peritoneal dissemination in gastric cancer. J Clin Lab Anal. 2021;35(9) doi: 10.1002/jcla.23936. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Hideura E., Suehiro Y., Nishikawa J., Shuto T., Fujimura H., Ito S., et al. Blood free-circulating DNA testing of methylated RUNX3 is useful for diagnosing early gastric cancer. Cancers. 2020;26(4):789. doi: 10.3390/cancers12040789. 12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Lin Z., Luo M., Chen X., He X., Qian Y., Lai S., et al. Combined detection of plasma ZIC1, HOXD10 and RUNX3 methylation is a promising strategy for early detection of gastric cancer and precancerous lesions. J Cancer. 2017;8(6):1038–1044. doi: 10.7150/jca.18169. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Ren J., Lu P., Zhou X., Liao Y., Liu X., Li J., et al. Genome-scale methylation analysis of circulating cell-free DNA in gastric cancer patients. Clin Chem. 2022;68(2):354–364. doi: 10.1093/clinchem/hvab204. [DOI] [PubMed] [Google Scholar]
- 71.Pimson C., Ekalaksananan T., Pientong C., Promthet S., Putthanachote N., Suwanrungruang K., et al. Aberrant methylation of PCDH10 and RASSF1A genes in blood samples for non-invasive diagnosis and prognostic assessment of gastric cancer. PeerJ. 2016;4 doi: 10.7717/peerj.2112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Xu J., Song J., Wang T., Zhu W., Zuo L., Wu J., et al. A combination of methylation and protein markers is capable of detecting gastric cancer detection by combined markers. Epigenomics. 2021;13(19):1557–1570. doi: 10.2217/epi-2021-0080. [DOI] [PubMed] [Google Scholar]
- 73.Alarcón M., Olivares W., Córdova-Delgado M., Muñoz-Medel M., De Mayo T., Carrasco-Aviño G., et al. The reprimo-like gene is an epigenetic-mediated tumor suppressor and a candidate biomarker for the non-invasive detection of gastric cancer. Int J Mol Sci. 2020;21(24):9472. doi: 10.3390/ijms21249472. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Yan H., Chen W., Ge K., Mao X., Li X., Liu W., et al. Value of plasma methylated SFRP2 in prognosis of gastric cancer. Dig Dis Sci. 2021;66(11):3854–3861. doi: 10.1007/s10620-020-06710-8. [DOI] [PubMed] [Google Scholar]
- 75.Anderson B.W., Suh Y.S., Choi B., Lee H.J., Yab T.C., Taylor W.R., et al. Detection of gastric cancer with novel methylated DNA markers: discovery, tissue validation, and pilot testing in plasma. Clin Cancer Res. 2018;24(22):5724–5734. doi: 10.1158/1078-0432.CCR-17-3364. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Li H., Zhao G., Guo Y., Fang Y., Wang K., Ma Y., et al. Feasibility and reproducibility of a plasma-based multiplex DNA methylation assay for early detection of gastric cancer. Pathol Res Pract. 2022;238 doi: 10.1016/j.prp.2022.154086. [DOI] [PubMed] [Google Scholar]
- 77.Guo X., Liu W., Pan Y., Ni P., Ji J., Guo L., et al. Homeobox gene IRX1 is a tumor suppressor gene in gastric carcinoma. Oncogene. 2010;29(27):3908–3920. doi: 10.1038/onc.2010.143. [DOI] [PubMed] [Google Scholar]
- 78.Salvianti F., Gelmini S., Costanza F., Mancini I., Sonnati G., Simi L., et al. The pre-analytical phase of the liquid biopsy. N Biotech. 2020;55:19–29. doi: 10.1016/j.nbt.2019.09.006. [DOI] [PubMed] [Google Scholar]
- 79.Tivey A., Church M., Rothwell D., Dive C., Cook N. Circulating tumour DNA — looking beyond the blood. Nat Rev Clin Oncol. 2022;19(9):600–612. doi: 10.1038/s41571-022-00660-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Garcia J., Kamps-Hughes N., Geiguer F., Couraud S., Sarver B., Payen L., et al. Sensitivity, specificity, and accuracy of a liquid biopsy approach utilizing molecular amplification pools. Sci Rep. 2021;11(1) doi: 10.1038/s41598-021-89592-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Chan H.T., Chin Y.M., Nakamura Y., Low S.K. Clonal hematopoiesis in liquid biopsy: from biological noise to valuable clinical implications. Cancers. 2020;12(8):2277. doi: 10.3390/cancers12082277. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Helzer K.T., Sharifi M.N., Sperger J.M., Shi Y., Annala M., Bootsma M.L., et al. Fragmentomic analysis of circulating tumor DNA-targeted cancer panels. Ann Oncol. 2023;34(9):813–825. doi: 10.1016/j.annonc.2023.06.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Fouz M.F., Appella D.H. PNA clamping in nucleic acid amplification protocols to detect single nucleotide mutations related to cancer. Molecules. 2020;25(4):786. doi: 10.3390/molecules25040786. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Markou A., Tzanikou E., Ladas I., Makrigiorgos G.M., Lianidou E. Nuclease-assisted minor allele enrichment using overlapping probes-assisted amplification-refractory mutation system: an approach for the improvement of amplification-refractory mutation system-polymerase chain reaction specificity in liquid biopsies. Anal Chem. 2019;91(20):13105–13111. doi: 10.1021/acs.analchem.9b03325. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Galbiati S., Damin F., Burgio V., Brisci A., Soriani N., Belcastro B., et al. Evaluation of three advanced methodologies, COLD-PCR, microarray and ddPCR, for identifying the mutational status by liquid biopsies in metastatic colorectal cancer patients. Clin Chim Acta. 2019;489:136–143. doi: 10.1016/j.cca.2018.12.004. [DOI] [PubMed] [Google Scholar]
- 86.Visser E., De Kock R., Genet S., Borne B.V.D., Soud M.Y.E., Belderbos H., et al. Up-front mutation detection in circulating tumor DNA by droplet digital PCR has added diagnostic value in lung cancer. Transl Oncol. 2023;27 doi: 10.1016/j.tranon.2022.101589. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.O'Leary B., Hrebien S., Beaney M., Fribbens C., Garcia-Murillas I., Jiang J., et al. Comparison of BEAMing and droplet digital PCR for circulating tumor DNA analysis. Clin Chem. 2019;65(11):1405–1413. doi: 10.1373/clinchem.2019.305805. [DOI] [PubMed] [Google Scholar]
- 88.Tébar-Martínez R., Martín-Arana J., Gimeno-Valiente F., Tarazona N., Rentero-Garrido P., Cervantes A. Strategies for improving detection of circulating tumor DNA using next generation sequencing. Cancer Treat Rev. 2023;119 doi: 10.1016/j.ctrv.2023.102595. [DOI] [PubMed] [Google Scholar]
- 89.Kurtz D.M., Soo J., Co Ting Keh L., Alig S., Chabon J.J., Sworder B.J., et al. Enhanced detection of minimal residual disease by targeted sequencing of phased variants in circulating tumor DNA. Nat Biotechnol. 2021;39(12):1537–1547. doi: 10.1038/s41587-021-00981-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Visal T.H., Den Hollander P., Cristofanilli M., Mani S.A. Circulating tumour cells in the -omics era: how far are we from achieving the ‘singularity’? Br J Cancer. 2022;127(2):173–184. doi: 10.1038/s41416-022-01768-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Lin D., Shen L., Luo M., Zhang K., Li J., Yang Q., et al. Circulating tumor cells: biology and clinical significance. Signal Transduct Targeted Ther. 2021;6(1):404. doi: 10.1038/s41392-021-00817-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Li Z., Song M., Han S., Jin C., Yang J. The prognostic role of circulating tumor cells in gastric cancer: a meta-analysis. Front Oncol. 2022;12 doi: 10.3389/fonc.2022.963091. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Zhu Y., Chen N., Chen M., Cui X., Yang H., Zhu X., et al. Circulating tumor cells: a surrogate to predict the effect of treatment and overall survival in gastric adenocarcinoma. Int J Biol Markers. 2021;36(1):28–35. doi: 10.1177/1724600820981972. [DOI] [PubMed] [Google Scholar]
- 94.Liu Y., Ling Y., Qi Q., Lan F., Zhu M., Zhang Y., et al. Prognostic value of circulating tumor cells in advanced gastric cancer patients receiving chemotherapy. Mol Clin Oncol. 2017;6(2):235–242. doi: 10.3892/mco.2017.1125. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Tan Z., Yue C., Ji S., Zhao C., Jia R., Zhang Y., et al. Assessment of PD-L1 expression on circulating tumor cells for predicting clinical outcomes in patients with cancer receiving PD-1/PD-L1 blockade therapies. Oncol. 2021;26(12):e2227–e2238. doi: 10.1002/onco.13981. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Chong X., Li Y., Lu J., Feng X., Li Y., Zhang X. Tracking circulating PD-L1-positive cells to monitor the outcome of patients with gastric cancer receiving anti-HER2 plus anti-PD-1 therapy. Hum Cell. 2023;37(1):258–270. doi: 10.1007/s13577-023-00990-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97.Andree K.C., Van Dalum G., Terstappen L.W.M.M. Challenges in circulating tumor cell detection by the CellSearch system. Mol Oncol. 2016;10(3):395–407. doi: 10.1016/j.molonc.2015.12.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98.Pernot S., Badoual C., Terme M., Castan F., Cazes A., Bouche O., et al. Dynamic evaluation of circulating tumour cells in patients with advanced gastric and oesogastric junction adenocarcinoma: prognostic value and early assessment of therapeutic effects. Eur J Cancer. 2017;79:15–22. doi: 10.1016/j.ejca.2017.03.036. [DOI] [PubMed] [Google Scholar]
- 99.Miller M.C., Doyle G.V., Terstappen L.W.M.M. Significance of circulating tumor cells detected by the CellSearch system in patients with metastatic breast colorectal and prostate cancer. JAMA Oncol. 2010;2010:1–8. doi: 10.1155/2010/617421. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.Deng Z., Wu S., Wang Y., Shi D. Circulating tumor cell isolation for cancer diagnosis and prognosis. EBioMedicine. 2022;83 doi: 10.1016/j.ebiom.2022.104237. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.Kilgour E., Rothwell D.G., Brady G., Dive C. Liquid biopsy-based biomarkers of treatment response and resistance. Cancer Cell. 2020;37(4):485–495. doi: 10.1016/j.ccell.2020.03.012. [DOI] [PubMed] [Google Scholar]
- 102.Roskams-Hieter B., Kim H.J., Anur P., Wagner J.T., Callahan R., Spiliotopoulos E., et al. Plasma cell-free RNA profiling distinguishes cancers from pre-malignant conditions in solid and hematologic malignancies. npj Precis Oncol. 2022;6(1):28. doi: 10.1038/s41698-022-00270-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Albitar M., Zhang H., Charifa A., Ip A., Ma W., McCloskey J., et al. Combining cell-free RNA with cell-free DNA in liquid biopsy for hematologic and solid tumors. Heliyon. 2023;9(5) doi: 10.1016/j.heliyon.2023.e16261. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104.Chaddha M., Rai H., Gupta R., Thakral D. Integrated analysis of circulating cell free nucleic acids for cancer genotyping and immune phenotyping of tumor microenvironment. Front Genet. 2023;14 doi: 10.3389/fgene.2023.1138625. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105.Chen T. Circulating non-coding RNAs as potential diagnostic biomarkers in hepatocellular carcinoma. J Hepatocell Carcinoma. 2022;9:1029–1040. doi: 10.2147/JHC.S380237. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 106.Link A., Kupcinskas J. MicroRNAs as non-invasive diagnostic biomarkers for gastric cancer: current insights and future perspectives. World J Gastroenterol. 2018;24(30):3313–3329. doi: 10.3748/wjg.v24.i30.3313. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107.Yao Y., Ding Y., Bai Y., Zhou Q., Lee H., Li X., et al. Identification of serum circulating MicroRNAs as novel diagnostic biomarkers of gastric cancer. Front Genet. 2021;11 doi: 10.3389/fgene.2020.591515. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 108.yang Song M., feng Pan K., Su H juan, Zhang L., Ma J ling, you Li J., et al. Identification of serum MicroRNAs as novel non-invasive biomarkers for early detection of gastric cancer. Califano J, curatore. PLoS One. 2012;7(3) doi: 10.1371/journal.pone.0033608. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.Shiotani A., Murao T., Kimura Y., Matsumoto H., Kamada T., Kusunoki H., et al. Identification of serum miRNAs as novel non-invasive biomarkers for detection of high risk for early gastric cancer. Br J Cancer. 2013;109(9):2323–2330. doi: 10.1038/bjc.2013.596. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110.Kim S.Y., Jeon T.Y., Choi C.I., Kim D.H., Kim D.H., Kim G.H., et al. Validation of circulating miRNA biomarkers for predicting lymph node metastasis in gastric cancer. J Mol Diagn. 2013;15(5):661–669. doi: 10.1016/j.jmoldx.2013.04.004. [DOI] [PubMed] [Google Scholar]
- 111.Sierzega M., Kaczor M., Kolodziejczyk P., Kulig J., Sanak M., Richter P. Evaluation of serum microRNA biomarkers for gastric cancer based on blood and tissue pools profiling: the importance of miR-21 and miR-331. Br J Cancer. 2017;117(2):266–273. doi: 10.1038/bjc.2017.190. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 112.Wu J., Li G., Wang Z., Yao Y., Chen R., Pu X., et al. Circulating MicroRNA-21 is a potential diagnostic biomarker in gastric cancer. Dis Markers. 2015;2015:1–8. doi: 10.1155/2015/435656. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113.Liu R., Zhang C., Hu Z., Li G., Wang C., Yang C., et al. A five-microRNA signature identified from genome-wide serum microRNA expression profiling serves as a fingerprint for gastric cancer diagnosis. Eur J Cancer. 2011;47(5):784–791. doi: 10.1016/j.ejca.2010.10.025. [DOI] [PubMed] [Google Scholar]
- 114.Abe S., Matsuzaki J., Sudo K., Oda I., Katai H., Kato K., et al. A novel combination of serum microRNAs for the detection of early gastric cancer. Gastric Cancer. 2021;24(4):835–843. doi: 10.1007/s10120-021-01161-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 115.Izumi D., Zhu Z., Chen Y., Toden S., Huo X., Kanda M., et al. Assessment of the diagnostic efficiency of a liquid biopsy assay for early detection of gastric cancer. JAMA Netw Open. 2021;4(8) doi: 10.1001/jamanetworkopen.2021.21129. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 116.Qiu X., Zhang J., Shi W., Liu S., Kang M., Chu H., et al. Circulating MicroRNA-26a in plasma and its potential diagnostic value in gastric cancer. Li Z, curatore. PLoS One. 2016;11(3) doi: 10.1371/journal.pone.0151345. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 117.Tsujiura M., Ichikawa D., Komatsu S., Shiozaki A., Takeshita H., Kosuga T., et al. Circulating microRNAs in plasma of patients with gastric cancers. Br J Cancer. 2010;102(7):1174–1179. doi: 10.1038/sj.bjc.6605608. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 118.Zhao G., Jiang T., Liu Y., Huai G., Lan C., Li G., et al. Droplet digital PCR-based circulating microRNA detection serve as a promising diagnostic method for gastric cancer. BMC Cancer. 2018;18(1):676. doi: 10.1186/s12885-018-4601-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 119.Zhu X.L., Ren L.F., Wang H.P., Bai Z.T., Zhang L., Meng W.B., et al. Plasma microRNAs as potential new biomarkers for early detection of early gastric cancer. World J Gastroenterol. 2019;25(13):1580–1591. doi: 10.3748/wjg.v25.i13.1580. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 120.Cao F., Hu Y., Chen Z., Han W., Lu W., Xu J., et al. Circulating long noncoding RNAs as potential biomarkers for stomach cancer: a systematic review and meta-analysis. World J Surg Oncol. 2021;19(1):89. doi: 10.1186/s12957-021-02194-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121.Yörüker E.E., Keskin M., Kulle C.B., Holdenrieder S., Gezer U. Diagnostic and prognostic value of circulating lncRNA H19 in gastric cancer. Biomed Rep. 2018 Aug;9(2):181–186. doi: 10.3892/br.2018.1116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 122.Xu H., Zhou J., Tang J., Min X., Yi T., Zhao J., et al. Identification of serum exosomal lncRNA MIAT as a novel diagnostic and prognostic biomarker for gastric cancer. J Clin Lab Anal. 2020;34(8) doi: 10.1002/jcla.23323. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 123.Tang D., Liu S., Shen H., Deng G., Zeng S. Extracellular vesicles promote the formation of pre-metastasis niche in gastric cancer. Front Immunol. 2022;13 doi: 10.3389/fimmu.2022.813015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 124.Guo T., Tang X.H., Gao X.Y., Zhou Y., Jin B., Deng Z.Q., et al. A liquid biopsy signature of circulating exosome-derived mRNAs, miRNAs and lncRNAs predict therapeutic efficacy to neoadjuvant chemotherapy in patients with advanced gastric cancer. Mol Cancer. 2022;21(1):216. doi: 10.1186/s12943-022-01684-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 125.Jiang C., Zhang J., Wang W., Shan Z., Sun F., Tan Y., et al. Extracellular vesicles in gastric cancer: role of exosomal lncRNA and microRNA as diagnostic and therapeutic targets. Front Physiol. 2023;14 doi: 10.3389/fphys.2023.1158839. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 126.Li Q., Lv M., Lv L., Cao N., Zhao A., Chen J., et al. Identifying HER2 from serum‐derived exosomes in advanced gastric cancer as a promising biomarker for assessing tissue HER2 status and predicting the efficacy of trastuzumab‐based therapy. Cancer Med. 2023;12(4):4110–4124. doi: 10.1002/cam4.5269. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 127.Fan Y., Che X., Li Z., Qu X., Liu Y. The association of an exosomal form of PD-L1 with immunosuppressive activity and gastric cancer prognosis. J Clin Oncol. 2018;36(4_suppl) 47–47. [Google Scholar]
- 128.Yakubovich E.I., Polischouk A.G., Evtushenko V.I. Principles and problems of exosome isolation from biological fluids. Biochem Mosc Suppl Ser Membr Cell Biol. 2022;16(2):115–126. doi: 10.1134/S1990747822030096. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 129.Garzarelli V., Ferrara F., Primiceri E., Chiriacò M.S. Biofluids manipulation methods for liquid biopsy in minimally-invasive assays. MethodsX. 2022;9 doi: 10.1016/j.mex.2022.101759. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 130.Yamamoto H., Watanabe Y., Sato Y., Maehata T., Itoh F. Non-invasive early molecular detection of gastric cancers. Cancers. 2020;12(10):2880. doi: 10.3390/cancers12102880. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 131.Gareev I., Ahmad A., Wang J., Beilerli A., Ilyasova T., Sufianov A., et al. Gastric juice non-coding RNAs as potential biomarkers for gastric cancer. Front Physiol. 2023;14 doi: 10.3389/fphys.2023.1179582. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 132.Cui L., Zhang X., Ye G., Zheng T., Song H., Deng H., et al. Gastric juice MicroRNAs as potential biomarkers for the screening of gastric cancer. Cancer. 2013;119(9):1618–1626. doi: 10.1002/cncr.27903. [DOI] [PubMed] [Google Scholar]
- 133.Pang Q., Ge J., Shao Y., Sun W., Song H., Xia T., et al. Increased expression of long intergenic non-coding RNA LINC00152 in gastric cancer and its clinical significance. Tumour Biol J Int Soc Oncodevelopmental Biol Med. 2014;35(6):5441–5447. doi: 10.1007/s13277-014-1709-3. [DOI] [PubMed] [Google Scholar]
- 134.Cisneros-Villanueva M., Hidalgo-Pérez L., Rios-Romero M., Cedro-Tanda A., Ruiz-Villavicencio C.A., Page K., et al. Cell-free DNA analysis in current cancer clinical trials: a review. Br J Cancer. 2022;126(3):391–400. doi: 10.1038/s41416-021-01696-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 135.Kwapisz D. The first liquid biopsy test approved. Is it a new era of mutation testing for non-small cell lung cancer? Ann Transl Med. 2017;5(3) doi: 10.21037/atm.2017.01.32. 46–46. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 136.Lin C., Liu X., Zheng B., Ke R., Tzeng C.M. Liquid biopsy, ctDNA diagnosis through NGS. Life. 2021;11(9):890. doi: 10.3390/life11090890. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 137.Porta C., Pradelli L., Sicari E., Castellani S., Sivakumar S., Sokol E., et al. Liquid biopsy comprehensive genomic profiling of lung cancer in the Italian population: a real-world experience. Lung Cancer. 2023;185 doi: 10.1016/j.lungcan.2023.107359. [DOI] [PubMed] [Google Scholar]
- 138.Shahnoor S., Mansha M., Farhat S., Maryyum A., Khan A.N., Khan A.M. Advancing precision oncology in metastatic colorectal cancer: the food and drug administration approval of foundation one liquid CDx as a companion diagnostic a correspondence. Rare Tumors. gennaio. 2023;15 doi: 10.1177/20363613231204798. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 139.Bonfil R.D., Al-Eyd G. Evolving insights in blood-based liquid biopsies for prostate cancer interrogation. Oncoscience. 2023;10:69–80. doi: 10.18632/oncoscience.592. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 140.Shirley M. Epi proColon® for colorectal cancer screening: a profile of its use in the USA. Mol Diagn Ther. 2020;24(4):497–503. doi: 10.1007/s40291-020-00473-8. [DOI] [PubMed] [Google Scholar]
- 141.Raiber‐Moreau E., Portella G., Butler M.G., Clement O., Konigshofer Y., Hadfield J. Development and validation of blood tumor mutational burden reference standards. Genes Chromosomes Cancer. 2023;62(3):121–130. doi: 10.1002/gcc.23100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 142.Jensen T.J., Goodman A.M., Kato S., Ellison C.K., Daniels G.A., Kim L., et al. Genome-wide sequencing of cell-free DNA identifies copy-number alterations that can Be used for monitoring response to immunotherapy in cancer patients. Mol Cancer Therapeut. 2019;18(2):448–458. doi: 10.1158/1535-7163.MCT-18-0535. [DOI] [PubMed] [Google Scholar]