Abstract
Background
We previously identified in non-small cell lung cancer (NSCLC) patients an autoantibody to complement factor H (CFH) that is associated with non-metastatic disease and longer time to progression in patients with stage I disease. A recombinant human antibody, GT103, was cloned from single B cells isolated from patients with the autoantibody. GT103 inhibits tumor growth and establishes an antitumor microenvironment. The anti-CFH autoantibody and GT103 recognize the epitope PIDNGDIT within the SCR19 domain of CFH. Here, we asked if this autoantibody could have originally arisen as a humoral response to a similar epitope in a viral protein from a prior infection.
Methods
Homologous viral peptides with high sequence identity to the core PIDNGDIT epitope sequence were identified and synthesized. NSCLC patient plasma containing anti-CFH autoantibodies were assayed by ELISA against these peptides. GT103 was assayed on a 4345-peptide pathogen microarray.
Results
Epitopes similar to the GT103 epitope are present in several viruses, including human metapneumovirus-1 (HMPV-1) that contains a sequence within attachment glycoprotein G that differs by one amino acid. Anti-CFH autoantibodies in NSCLC patient plasma weakly bound to an HMPV-1 peptide containing the epitope. GT103 cross-reacted with multiple viral epitopes on a peptide microarray, with the top hits being peptides in the human endogenous retrovirus-K polymerase (HERV-K pol) protein and measles hemagglutinin glycoprotein. GT103 bound the viral HMPV-1, HERV-K pol, and measles epitope peptides but with lower affinity compared to the GT103 epitope peptide.
Conclusion
These findings suggest that memory B cells against a viral target could have affinity matured to produce an antibody that recognizes a similar epitope on tumor cells and exhibits antitumor properties.
Supplementary Information
The online version contains supplementary material available at 10.1007/s00262-025-03975-8.
Keywords: Autoantibodies, Viral epitopes, Antibody cross-reactivity, Molecular mimicry, Humoral immunity
Introduction
Autoantibodies are hallmarks of a variety of human diseases including, but not limited to, those categorized as autoimmune diseases. The human autoantigen atlas (http://biokb.ncpsb.org.cn/aagatlas_portal/index.php#) is comprised of 8045 autoantigens associated with 1073 diseases, including cancer and diseases of immune, musculoskeletal, connective tissue, neural, and cardiovascular systems [1]. Autoantibodies can be used to screen for or diagnose autoimmune diseases such as systemic lupus erythematosus [2], the seropositive form of rheumatoid arthritis [3], and type I diabetes [4], and have been explored as screening and prognostic biomarkers for cancer [5–7]. In addition, the identification of an antigen by the presence of autoantibodies against it holds the promise of development of therapies targeted to it. The Cancer Antigenic Peptide Database (De Duve Institute, Brussels, Belgium; https://caped.icp.ucl.ac.be/Peptide/list), currently contains 474 tumor-associated antigen (TAA) peptide entries, all of which have been shown to be presented on human leukocyte antigen (HLA) proteins and recognized by T cells. Unique antigens are those created by mutation that are particular to individuals and thus would likely not be targets for drug development. Shared neo-antigens include tumor-specific antigens, expressed in many tumors but not normal tissues, and tumor-associated antigens that are overexpressed in tumor tissue but also expressed in normal tissue. Viral peptides associated with human malignancy, such as Epstein–Barr virus and human papilloma virus are not included in this database but represent additional potential antigenic targets for therapy.
Interestingly, autoantibodies are also found in healthy individuals. LaBaer and colleagues compiled autoantibody profiles (the “autoantibodyome”) of 272 control subjects from case–control biomarker studies [8]. They studied a total of 8282 antigen reactivities and designated 77 autoantibodies, present in 10–47% of individuals, as “common.” The 77 autoantibodies were found at similar frequencies in the matched diseased cohorts; thus were not elicited by the disease under study. The number of different autoantibodies increased between the age ranges infancy—6 years and 6–12 years, and plateaued thereafter. Among the common autoantigens, 21 contained sequence matches to viral proteins. These sequence matches could reflect the phenomenon of molecular mimicry, in which antibodies reactive to viral antigens from previously encountered viruses cross-react with normal antigens [9].
Cross-reactivity of autoantibodies as well as T cell receptors (TCRs) against TAAs with viral antigens has also been reported. Using a bioinformatics approach, Ragone et al. [10] performed a global BLAST search with 99 TAA peptides from the Cancer Antigenic Peptide database against all human viral sequences and found 82 viral peptides sharing homology with TAA peptides; cross-reactivity of several of these pairs was confirmed in an ex vivo T cell stimulation assay. Chiou et al. [11] analyzed T cell receptor sequences enriched in non-small cell lung cancer (NSCLC) tumors and discovered that T cells with specificity to the epithelial protein TMEM161A were cross-reactive with epitopes from Epstein–Barr virus. Petrizzo et al. [12] found that neoantigen-specific T cells from hepatocellular carcinoma were cross-reactive with multiple pathogen epitopes.
GT103 is a human monoclonal antibody in clinical trials for NSCLC (ClinicalTrials.Gov IDs NCT04314089 and NCT05617313). GT103 is derived from an autoantibody that was discovered to be preferentially expressed in early stage as opposed to late stage NSCLC [13], and its presence in early stage NSCLC correlates with longer time to recurrence [14]. GT103 recognizes an epitope in the SCR19 domain of the complement regulatory protein complement factor H (CFH) [15]. This epitope is not accessible to antibody in native CFH but is recognized by antibody upon conformational change [15, 16]. GT103, by binding to CFH, causes tumor cell lysis by activating the classical complement pathway, and slows tumor growth in vivo with concomitant promotion of an antitumor microenvironment [16–18]. Since the epitope is masked in the native structure of CFH and is possibly only revealed in the tumor cell microenvironment, we wondered if the exposed epitope stimulates affinity maturation of memory B cells with specificity for a similar epitope originally perceived as foreign. In select patients, the autoantibody produces a beneficial phenotype. Autoimmune diseases have long been associated with cross-reactivity of viral epitopes with self epitopes [19], but this phenomenon in cancer is a more recent area of exploration. Here, we examine the possibility that anti-CFH autoantibodies could have arisen from prior exposure of individuals to a nonself epitope present in a virus.
Materials and methods
Blood sample collection
The Duke University Institutional Review Board (IRB) approved this study with the written consent of each subject. Thousands of blood samples from patients being seen at Duke clinics for treatment or screening for NSCLC were collected and the plasma stored in our IRB-approved −80°C repository. Samples from patients previously shown to be positive or negative for the presence of the anti-CFH autoantibody by the criteria of reference [14] were randomly selected for the current study.
Search criteria
A nonredundant Protein Blast search using PIDNGDIT as the query was run at the NCBI website https://blast.ncbi.nlm.nih.gov/Blast.cgi#. The results were limited to viruses (taxid:10,239).
ELISA
We performed peptide ELISA as previously described in detail [14] using a peptide containing the CFH epitope GPPPPIDNGDITSFPGGG-Lys(biotin) [15, 16] or related biotin-conjugated viral peptides as described in the figure legends. (The epitope in the CFH peptide is underlined.) Briefly, wells of a 96-well plate, precoated with NeutrAvidin (ThermoFisher; cat. # 15,129), were washed with PBS/0.1% Tween 20 (PBST) and 2 μg/ml peptide or biotin in PBST added. After a 30 min incubation followed by washing, the wells were loaded with patient plasma or GT103, diluted as described in the figure legends. After a 1.5 h incubation followed by washing, goat antihuman IgG gamma-chain-horseradish peroxidase secondary antibody (Chemicon; Sigma-Aldrich AP39P) diluted 1:8000 was added to the wells. After a 1 h incubation followed by washing, bound antibody was detected using ABTS/hydrogen peroxide. Absorbance was measured at 405 nm using a 96-well plate spectrophotometer. Mean absorbance from background binding of antibody to biotin-coated control wells was subtracted from mean absorbance of the corresponding test wells. Reactions were run in duplicate and P values calculated using Student’s T-test.
Epitope microarray analysis
We performed binding of GT103 to an infectious disease epitope microarray. The PEPperCHIP® Infectious Disease Epitope Microarray (PEPperPRINT GmbH) covers 3,760 linear B-cell epitopes from 196 different pathogens associated with infectious diseases, as referenced in the Immune Epitope Database (https://www.iedb.org). The 3,760 linear B-cell epitopes were converted into 4,345 different peptides printed in duplicate (8,690 peptide spots) and complemented by polio (KEVPALTAVETGAT, 42 spots), c-myc (EQKLISEEDL, 30 spots) and HA (YPYDVPDYAG, 40 spots) control peptides. The microarray was blocked with blocking buffer (Rockland Immunochemicals MB-070) for 30 min, then incubated with GT103 at a concentration of 1 µg/ml in 10% blocking buffer in phosphate-buffered saline, pH 7.4, 0.05% Tween 20 (PBST-05) for 16 h at 4 °C with orbital shaking. After washing in PBST-05 (3 × 10 s), the microarray was stained with DyLight680- and DyLight800-labeled secondary antibodies recognizing human IgG (H + L) and anti-HA control antibody, respectively, for 45 min at room temperature. After further washing, the microarray was read with an Innopsys InnoScan 710-IR microarray scanner with a scanning resolution of 20 µm and scanning gains of 20 at low laser power (680 nm, red) and 40 at high laser power (800 nm, green). Quantification of spot intensities and peptide annotation were done with PepSlide® Analyzer.
Results
The autoantibody to CFH that is associated with non-metastatic NSCLC, and the completely human monoclonal antibody GT103 from which it was derived, both recognize the epitope PIDNGDIT [15, 16]. To investigate the known prevalence of this epitope in other proteins, the GT103 peptide epitope (PIDNGDIT) was used as the query sequence versus known viral protein sequences using the NCBI BLAST algorithm. Supplementary Table 1 lists the hits resulting from this search, and Supplementary Fig. 1 shows representative alignments. The hits fell within over 80 HMPV-1 isolates that contained amino acid sequences that differed from the query by only one amino acid residue, as well as within human immunodeficiency virus-1. Hits also fell within bacteriophages, most frequently Caudoviricetes (Caudovirales), a class of phage species that is found in the human gut [20]. Hits appeared in a variety of proteins, including a viral exonuclease, a tail fiber-like protein, portal protein, attachment glycoprotein G, and envelope glycoprotein.
The similarity in amino acid sequence between the GT103 epitope and the related region in the attachment glycoprotein G of HMPV-1 led us to speculate that anti-CFH antibodies may have arisen from an immune reaction to HMPV-1. As a first step to explore this possibility, we compared binding of anti-CFH autoantibodies in patient plasma to a peptide containing the CFH epitope PIDNGDIT versus a peptide containing the similar HMPV-1 epitope PIDNPDIT. Both peptides were immobilized in an ELISA plate, which was then used to test human plasma samples that had been shown previously to be positive or negative for the presence of the CFH autoantibody. None of the plasma samples from NSCLC patients, whether CFH autoantibody positive or negative, when diluted 1:200, demonstrated detectable antibody binding to the HMPV-1 peptide (Fig. 1). In contrast, the autoantibody-positive samples bound the CFH peptide as expected while the autoantibody-negative samples did not. Binding of antibody from some of the autoantibody-positive samples to the HMPV-1 peptide was detected at 1:2 dilution of plasma (with mean A450 values of 0.8–1.0 for the CFH peptide versus 0.005–0.1 for the HMPV peptide in that assay).
Fig. 1.

Binding of plasma antibodies to CFH and HMPV-1 and peptides. Plasma samples from anti-CFH autoantibody-positive and autoantibody-negative patients were diluted 1:200 and tested by ELISA against CFH peptide (GPPPPIDNGDITSFPGGG-Lys(biotin)) and HMPV-1 peptide (TSTIPIDNPDITPNSGGG-Lys(biotin)) that contains a one amino acid difference (bold) in the consensus region (underlined). After background (biotin-coated well) subtraction, mean of duplicates are reported, with standard deviation given for the CFH peptide ELISA
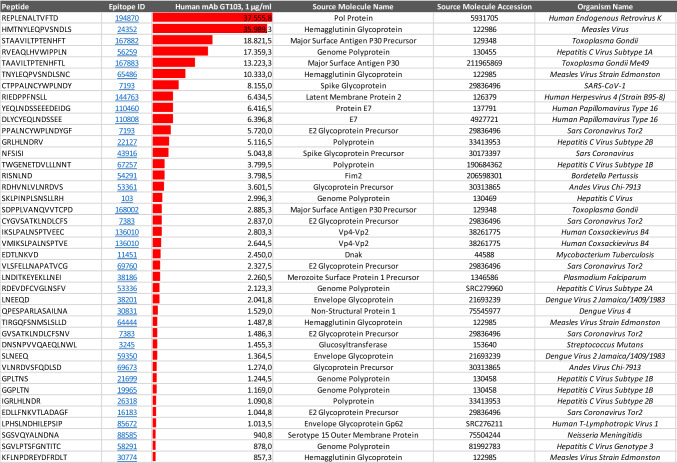
To further investigate the possibility that the anti-CFH autoantibody recognizes viral epitopes, and thus may have originated from an antiviral response, we had GT103 tested on the PEPperCHIP®, an infectious disease epitope peptide microarray. Incubation of GT103 with the array resulted in a moderate to strong antibody response against a number of pathogen peptides at high signal-to-noise ratios (Fig. 2). The top antibody responses were attributed to peptides REPLENALTVFTD (pol protein, human endogenous retrovirus-K), HMTNYLEQPVSNDLS (hemagglutinin glycoprotein, measles virus), TAAVILTPTENHFT (major surface antigen P30 precursor, Toxoplasma gondii), RVEAQLHVWIPPLN (hepatitis C virus subtype 1A), TNYLEQPVSNDLSNC (hemagglutinin glycoprotein, measles virus), and CTPPALNCYWPLNDY (spike glycoprotein, SARS-CoV-1). The array did not include HMPV-1 peptides.
Fig. 2.
Binding of GT103 to PEPperCHIP® Infectious Disease Epitope Microarray. The responses are sorted by decreasing spot intensity
The strongest responses were based on peptides REPLENALTVFTD and HMTNYLEQPVSNDLS that shared the common motifs LE(Q/N)x(L/V)(T/S). Similar motifs were also found with top peptides TAAVILTPTENHFT, RVEAQLHVWIPPLN, or TNYLEQPVSNDLSNC, albeit at slightly lower intensity levels. Therefore, we assume that the cross-reactivity of GT103 was driven by peptides with consensus motifs like 1LE(Q/N)x(L/V)(T/S)6, while consensus motifs like TENHFT or VEAQLH with an exchange of 1L, 3(Q/N) or 6(T/S) were also tolerated to some extent. (Superscripts denote amino acid residue position.) Even though the infectious disease epitopes recognized by GT103 in the microarray do not overtly resemble the CFH epitope (PIDNGDIT), the consensus motif LE(Q/N)x(L/V)(T/S) accurately reflects the amino acid class (i.e., hydrophobic, polar, or charged) within the CFH epitope.
To assess the binding of the anti-CFH antibody to the two top viral peptide hits, we tested epitope peptides from HERV-K and measles virus hemagglutinin protein, along with HMPV-1 and CFH peptides, by ELISA for recognition by GT103. The results are shown in Fig. 3. GT103 bound to all peptides, but with significantly lower affinity to the viral epitope peptides than to the CFH peptide. Binding of GT103 to the HMPV-1 peptide was at least two orders of magnitude weaker than binding to the CFH peptide.
Fig. 3.
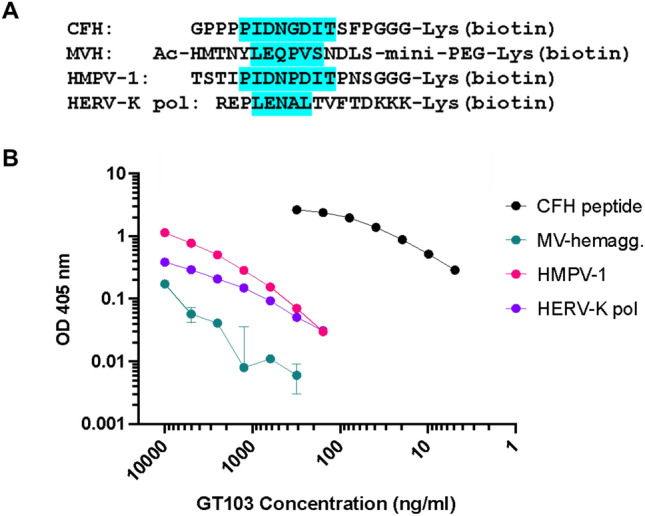
ELISA of GT103 binding to its epitope peptide (CFH peptide) compared to viral epitope peptides. A Amino acid sequences of the peptides used in the ELISA (with full epitope or consensus motif highlighted). B Binding of GT103 to viral peptides. Biotinylated peptides were immobilized in the wells of an ELISA plate and tested against twofold dilutions of GT103 from 10 to 0.005 µg/ml
Discussion
It is well established that solid tumors containing tertiary lymphoid structures (TLS), organized aggregates of dendritic cells, and T cells surrounded by B-cell follicles, are associated with a more favorable prognosis [21]. In these structures, B cells are clonally expanded and affinity matured in response to tumor antigens [22–26]. Hypothetically, the anti-CFH autoantibody in NSCLC patients could have originated in a number of ways: in one scenario, bone marrow-resident B cells with specificity toward the sterically obscured PIDNGDIT epitope in CFH escape deletion (having never been recognized as self), migrate into the tumor, organize into TLS, and undergo activation and expansion upon encountering the now-exposed epitope in the tumor environment. In another scenario, preexisting memory B cells with specificity toward PIDNGDIT-related epitopes in pathogens encountered during prior infection migrate into the tumor and are activated and expanded upon encountering the CFH epitope there. The anti-CFH autoantibody is also found in the blood of some of our lung cancer screening patients without overt cancer. In non-cancer patients, the anti-CFH autoantibody could have been produced in secondary lymphoid organs such as lymph nodes or spleen in response to antigens derived from nascent, undetectable tumors, or previously eradicated tumors.
Although the origin of the anti-CFH autoantibody may never be definitively known, the evidence presented in this paper supports the possibility that it could have evolved from a weak humoral reaction to a viral antigen, possibly one encountered in childhood, like HMPV-1 or measles, or in reaction to a viral vaccine. Another possibility is that the antibody could have originated in reaction to gut bacteriophage, as these have been shown to interact with innate immune cells [27], and many matches to the epitope sequence appear in Caudoviricetes proteins. The weak reactivity of the anti-CFH autoantibody or the recombinant derivative GT103 toward the viral epitopes tested contrasts with the strong binding (low nanomolar affinity) of GT103 to the CFH epitope [16]. B cells expressing the antibody in adults may have affinity matured and then produced an antibody that binds CFH stronger than the viral epitope that was originally encountered.
Regardless of the origin of the anti-CFH antibody, it is worth recognizing that autoantibodies can have beneficial, neutral, or detrimental effects on the health of the patient. The prevalence of autoantibodies in tumors, and even as part of the normal autoantibodyome, appears to be much higher than commonly thought, as seen in recent publications (8, 11, 25) and databases. Identification of autoantibodies with beneficial effects would allow for the development of antibody therapeutics, or preventative vaccines against infections and tumors based on the antigen. To further study this hypothesis, additional and more comprehensive studies with a broad array of pathogens and in vivo experiments are needed.
Supplementary Information
Below is the link to the electronic supplementary material.
Acknowledgements
We thank Mihai Azoitei and Daniel Marston of the Duke Human Vaccine Institute for enlightening discussions.
Author contributions
MJC presented conceptualization, methodology, and formal analysis; EBG provided conceptualization, writing—original draft, and writing—review and editing; KW performed writing—review and editing; EFP carried out conceptualization, project administration, and writing—review and editing
Funding
No funding was received for conducting this study.
Data availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
Declarations
Competing interests
E.F.P., E.B.G., and M.J.C. are founders of Grid Therapeutics. E.F.P. is the CEO of Grid Therapeutics. K.W. has no competing interests to declare that are relevant to the content of this article.
Footnotes
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
- 1.Wang D, Yang D, Yang L et al (2023) Human autoantigen atlas: searching for the hallmarks of autoantigens. J Proteome Res 22:1800–1815. 10.1021/acs.jproteome.2c00799 [DOI] [PubMed] [Google Scholar]
- 2.Yaniv G, Twig G, Shor DB et al (2015) A volcanic explosion of autoantibodies in systemic lupus erythematosus: a diversity of 180 different antibodies found in SLE patients. Autoimmun Rev 14:75–79. 10.1016/j.autrev.2014.10.003 [DOI] [PubMed] [Google Scholar]
- 3.Malmstrom V, Catrina AI, Klareskog L (2017) The immunopathogenesis of seropositive rheumatoid arthritis: from triggering to targeting. Nat Rev Immunol 17:60–75. 10.1038/nri.2016.124 [DOI] [PubMed] [Google Scholar]
- 4.Bian X, Wasserfall C, Wallstrom G et al (2017) Tracking the antibody immunome in type 1 diabetes using protein arrays. J Proteome Res 16:195–203. 10.1021/acs.jproteome.6b00354 [DOI] [PubMed] [Google Scholar]
- 5.Patel AJ, Tan TM, Richter AG, Naidu B, Blackburn JM, Middleton GW (2022) A highly predictive autoantibody-based biomarker panel for prognosis in early-stage NSCLC with potential therapeutic implications. Br J Cancer 126:238–246. 10.1038/s41416-021-01572-x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Qiu C, Wang X, Batson SA, Wang B, Casiano CA, Francia G, Zhang JY (2023) A luminex approach to develop an anti-tumor-associated antigen autoantibody panel for the detection of prostate cancer in racially/ethnically diverse populations. Cancers (Basel). 10.3390/cancers15164064 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Yang R, Han Y, Yi W, Long Q (2022) Autoantibodies as biomarkers for breast cancer diagnosis and prognosis. Front Immunol 13:1035402. 10.3389/fimmu.2022.1035402 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Shome M, Chung Y, Chavan R, Park JG, Qiu J, LaBaer J (2022) Serum autoantibodyome reveals that healthy individuals share common autoantibodies. Cell Rep 39:110873. 10.1016/j.celrep.2022.110873 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Tagliamonte M, Cavalluzzo B, Mauriello A, Ragone C, Buonaguro FM, Tornesello ML, Buonaguro L (2023) Molecular mimicry and cancer vaccine development. Mol Cancer 22:75. 10.1186/s12943-023-01776-0 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Ragone C, Manolio C, Cavalluzzo B et al (2021) Identification and validation of viral antigens sharing sequence and structural homology with tumor-associated antigens (TAAs). J Immunother Cancer. 10.1136/jitc-2021-002694 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Chiou SH, Tseng D, Reuben A et al (2021) Global analysis of shared T cell specificities in human non-small cell lung cancer enables HLA inference and antigen discovery. Immunity 54(586–602):e8. 10.1016/j.immuni.2021.02.014 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Petrizzo A, Tagliamonte M, Mauriello A et al (2018) Unique true predicted neoantigens (TPNAs) correlates with anti-tumor immune control in HCC patients. J Transl Med 16:286. 10.1186/s12967-018-1662-9 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Amornsiripanitch N, Hong S, Campa MJ, Frank MM, Gottlin EB, Patz EF Jr (2010) Complement factor H autoantibodies are associated with early stage NSCLC. Clin Cancer Res 16:3226–3231. 10.1158/1078-0432.CCR-10-0321 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Gottlin EB, Campa MJ, Gandhi R, Bushey RT, Herndon JE 2nd, Patz EF Jr (2022) Prognostic significance of a complement factor H autoantibody in early stage NSCLC. Cancer Biomark 34:385–392. 10.3233/CBM-210355 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Campa MJ, Gottlin EB, Bushey RT, Patz EF Jr (2015) Complement factor h antibodies from lung cancer patients induce complement-dependent lysis of tumor cells, suggesting a novel immunotherapeutic strategy. Cancer Immunol Res 3:1325–1332. 10.1158/2326-6066.CIR-15-0122 [DOI] [PubMed] [Google Scholar]
- 16.Bushey RT, Moody MA, Nicely NL et al (2016) A therapeutic antibody for cancer, derived from single human B cells. Cell Rep 15:1505–1513. 10.1016/j.celrep.2016.04.038 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Bushey RT, Saxena R, Campa MJ, Gottlin EB, He YW, Patz EF (2023) Antitumor immune mechanisms of the anti-complement factor H antibody GT103. Mol Cancer Ther 22:778–789. 10.1158/1535-7163.MCT-22-0723 [DOI] [PubMed] [Google Scholar]
- 18.Saxena R, Bushey RT, Campa MJ, Gottlin EB, Guo J, Patz EF Jr, He YW (2024) Promotion of an antitumor immune program by a tumor-specific. Complement-Act Antibody J Immunol 212:1589–1601. 10.4049/jimmunol.2300728 [DOI] [PubMed] [Google Scholar]
- 19.Johnson D, Jiang W (2023) Infectious diseases, autoantibodies, and autoimmunity. J Autoimmun 137:102962. 10.1016/j.jaut.2022.102962 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Liang G, Bushman FD (2021) The human virome: assembly, composition and host interactions. Nat Rev Microbiol 19:514–527. 10.1038/s41579-021-00536-5 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Domblides C, Rochefort J, Riffard C, Panouillot M, Lescaille G, Teillaud JL, Mateo V, Dieu-Nosjean MC (2021) Tumor-associated tertiary lymphoid structures: from basic and clinical knowledge to therapeutic manipulation. Front Immunol 12:698604. 10.3389/fimmu.2021.698604 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Centuori SM, Bhattacharya D (2022) Locally produced autoantibodies in cancer. Cell 185:1110–1111. 10.1016/j.cell.2022.03.006 [DOI] [PubMed] [Google Scholar]
- 23.Germain C, Gnjatic S, Tamzalit F et al (2014) Presence of B cells in tertiary lymphoid structures is associated with a protective immunity in patients with lung cancer. Am J Respir Crit Care Med 189:832–844. 10.1164/rccm.201309-1611OC [DOI] [PubMed] [Google Scholar]
- 24.Helmink BA, Reddy SM, Gao J et al (2020) B cells and tertiary lymphoid structures promote immunotherapy response. Nature 577:549–555. 10.1038/s41586-019-1922-8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Mazor RD, Nathan N, Gilboa A et al (2022) Tumor-reactive antibodies evolve from non-binding and autoreactive precursors. Cell 185(1208–22):e21. 10.1016/j.cell.2022.02.012 [DOI] [PubMed] [Google Scholar]
- 26.Wieland A, Patel MR, Cardenas MA et al (2021) Defining HPV-specific B cell responses in patients with head and neck cancer. Nature 597:274–278. 10.1038/s41586-020-2931-3 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Carroll-Portillo A, Lin HC (2019) Bacteriophage and the innate immune system: access and signaling. Microorganisms. 10.3390/microorganisms7120625 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.