Abstract
Aims—Loss of heterozygosity (LOH) at specific chromosomal regions strongly suggests the existence of tumour suppressor genes at the relevant segment. Frequent LOH on chromosome 5q has been reported in a wide variety of human tumours, including those of the lung. The aim of this study was to screen for LOH and to clarify the location of putative tumour suppressor genes on chromosome 5 implicated in the genesis and/or development of non-small cell lung carcinoma.
Methods—Thirty three patients with advanced non-small cell lung carcinoma were screened for LOH with a panel of 21 microsatellite DNA markers spanning the entire chromosome 5, using semi-automated fluorochrome based methodology.
Results—Twenty of the non-small cell lung carcinoma samples displayed LOH for one or more informative locus. LOH involving only 5q was found in 10 of 14 of the informative samples. Deletions involving 5p only were not present in the samples under study. There was no evidence of microsatellite instability in any of the analysed loci. These results indicate the presence of five distinct segments displaying high frequencies of deletion on chromosome 5, namely: 5q11.2–q12.2, 5q15 (D5S644 locus), 5q22.3–q23.1, 5q31.1, and 5q35.3. Eight of 14 samples had simultaneous interstitial deletions in at least two different regions. Moreover, concomitant deletion of three and four distinct regions was displayed in three of 14 and two of 14, respectively, of the informative samples.
Conclusion—Allelic deletion on chromosome 5 is a frequent event in patients with non-small cell lung carcinoma. These results suggest the involvement of these five regions, either independently or simultaneously, in both lung squamous cell carcinoma and lung adenocarcinoma.
Keywords: non-small cell lung carcinoma, chromosome 5, loss of heterozygosity
Inactivation of tumour suppressor genes appears to be one of the genetic mechanisms involved in the development of solid tumours. This process includes mutation of one allele, followed by a deletion of the remaining one (loss of heterozygosity; LOH) or homozygous deletion of both alleles. Allelic deletions detected as LOH have proved useful for mapping regions of DNA that contain tumour suppressor genes.
In non-small cell lung cancer the chromosomes most frequently affected by LOH are 3p, 5q, 9p, 13q, and 17p.1
Chromosome 5 is a frequent target of genetic alterations in several malignant diseases such as ovarian cancer,2 gastric cancer,3 oesophageal cancer,4 and malignant myeloid diseases.5 LOH on chromosome 5 has been reported in non-small cell lung carcinoma, affecting both the p and q arms. Deletions involving the long arm of chromosome 5 include the APC/MCC gene cluster,6–9 the 5q33–q35 region8 and, more recently, the 5q11–q13 region containing the mismatch repair gene hMSH3.10 A candidate tumour suppressor gene locus on the short arm of chromosome 5, del-27, was reported to be deleted in non-small cell lung carcinoma and maps to the chromosomal segment 5p12–p13.11 Moreover, chromosome 5 is described as being a molecular target of carcinogens in the development of lung cancer, and the proportion of bleomycin induced 5q aberrations that survive in patients with lung cancer is significantly higher than in healthy controls.12
In our study, 21 (CA)n microsatellite markers were analysed in 33 matched normal/tumour samples from patients with non-small cell lung carcinoma using fluorescent DNA technology. Our objective was to screen for LOH and to clarify the location of putative tumour suppressor genes on chromosome 5 implicated in the genesis and/or development of non-small cell lung carcinoma.
Methods
TISSUE SAMPLES
Thirty three non-small cell lung carcinomas (25 squamous cell carcinomas and eight adenocarcinomas) and matched control constitutional DNAs were obtained from the Portuguese Institute of Oncology Dr Francisco Gentil, in Lisbon, from patients with locally advanced or metastatic disease not submitted to previous chemotherapeutic/radiotherapeutic treatments. Tumour samples were collected by fibre-optic bronchoscopy and control DNA samples were obtained either from normal bronchial tissue not adjacent to the tumour or from blood. Both the neoplastic and normal cell content were analysed by histopathological methods. DNA from solid tissue was purified by proteinase K digestion, extracted with phenol/chloroform, and precipitated by ethanol. DNA from blood samples was obtained by the guanidine HCl method.
ALLELOTYPING OF SHORT TANDEM REPEATS
Twenty one short tandem repeat DNA markers for chromosome 5 (PE-ABD linkage set of markers) were obtained from Perkin Elmer-ABD (Foster City, California, USA) and amplified by the polymerase chain reaction (PCR) for LOH screening (table 1 ▶). One primer of each pair was fluorescently labelled with a fluorochrome (TET, FAM, HEX, or JOE) attached to the 5` end. PCR reactions were carried out in a total volume of 20 μl containing 1× reaction buffer (10× PCR Buffer II; Perkin Elmer-ABD), 0.06–0.5 μM of each primer, 200 μM of each deoxynucleotide triphosphate (Pharmacia, Buckinghamshire, UK), 0.5 U of DNA polymerase (AmpliTaq Gold; Perkin Elmer-ABD), 1.0–2.0 mM of MgCl2 (Perkin Elmer-ABD), and 100–200 ng of genomic DNA. PCR amplifications were performed in a Perkin Elmer 9600 GenAmp PCR system and the conditions were 12 minutes at 95°C, followed by 15 seconds at 94°C, 15 seconds at 55/60°C, and 30 seconds at 72°C for 10 cycles, and by 15 seconds at 89°C, 15 seconds at 55/60°C, and 30 seconds at 72°C for 30 cycles, with a final extension step of 10 minutes at 72°C. Negative controls were incorporated into every PCR set of amplification for the detection of contamination.
Table 1.
LOH at chromosome 5
| Locus | Map location | Distance to the next marker (cM) | Informative cases | LOH | % |
| D5S406 | 5p15.31 | 32.7 | 9 | 0 | 0.0 |
| D5S419 | 5p14.3 | 12.8 | 19 | 2 | 10.5 |
| D5S426 | 5p11 | 12.7 | 12 | 1 | 8.3 |
| D5S407 | 5q11.2 | 9.8 | 17 | 9 | 52.9 |
| D5S647 | 5q11.2 | 10.2 | 18 | 9 | 50.0 |
| D5S424 | 5q12.2 | 14.3 | 13 | 6 | 46.2 |
| D5S428 | 5q14.1 | 10.5 | 17 | 5 | 29.4 |
| D5S644 | 5q15 | 4.4 | 24 | 11 | 45.8 |
| D5S299 | 5q21.3 | 2.5 | 24 | 6 | 25.0 |
| D5S433 | 5q21.3 | 9.0 | 22 | 9 | 40.9 |
| D5S346 | 5q22.3 | 0.6 | 26 | 9 | 34.6 |
| D5S421 | 5q23.1 | 0.5 | 22 | 9 | 40.9 |
| MCC | 5q23.1 | 1.0 | 20 | 9 | 45.0 |
| D5S318 | 5q23.1 | 16.0 | 21 | 8 | 38.1 |
| D5S393 | 5q31.1 | 9.0 | 15 | 5 | 33.3 |
| D5S436 | 5q31.1 | 8.5 | 14 | 6 | 42.8 |
| D5S673 | 5q31.1 | 7.2 | 11 | 8 | 72.7 |
| D5S422 | 5q34 | 13.4 | 14 | 6 | 42.8 |
| D5S400 | 5q35.2 | 4.7 | 12 | 3 | 25.0 |
| D5S429 | 5q35.3 | 13.7 | 18 | 10 | 55.6 |
| D5S408 | 5q35.3 | 9.8 (qter) | 18 | 8 | 44.4 |
LOH, loss of heterozygosity.
PCR products were separated by 6% denaturing polyacrylamide gel electrophoresis in a 373A DNA sequencer (Perkin Elmer-ABD). All samples were tested at least twice in independent gel loadings. Microsatellite allele sizes were determined by reference to the internal lane standards 350-TAMRA or 350-ROX (Perkin Elmer-ABD) and the results were analysed using GeneScan 672 software (Perkin Elmer-ABD).13
Results and discussion
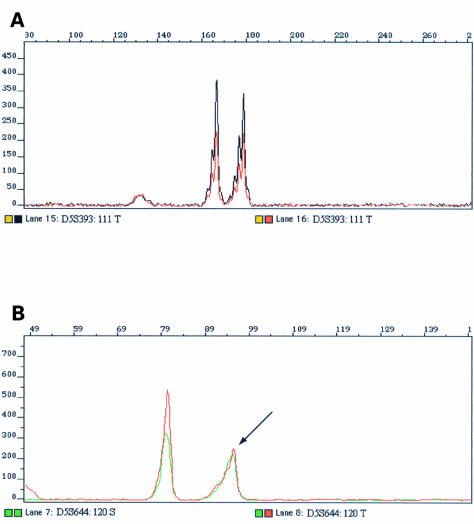
LOH was evaluated using the formula (T1 : T2/N1 : N2),14 with additional visual interpretation of both tumour and constitutional amplimer profiles (fig 1 ▶). LOH was defined as a reduction of at least 30% in one allele of the tumour DNA when compared with the constitutional DNA sample.15,16
Figure 1.
Retention and loss of heterozygosity (LOH). (A) Example of a sample showing retention of heterozygosity; the normal and tumour DNA profiles are identical. (B) Example of a sample displaying LOH; an arrow indicates a reduction in one of the tumour alleles when compared with the matched constitutional DNA.
LOH for one or more informative (heterozygous) locus was detected in 20 of the 33 non-small cell lung carcinoma samples. No significant differences in the proportions of samples displaying LOH were found between the squamous cell carcinoma samples and the adenocarcinoma samples (64% and 50%, respectively).
Six of 33 samples displayed LOH for all the informative loci, suggesting a reduction to monosomy of chromosome 5.
Among the samples with interstitial deletions (fig 2 ▶), 10 of 14 displayed LOH involving 5q loci only and four of 14 showed LOH involving both 5p and 5q loci. Samples displaying LOH involving only the short arm of chromosome 5 were not found. Although minimal regions of deletion have been identified previously on 5p, namely del-27 on 5p12–p1311 and the D5S667 locus on 5p15.2,17 the 5p loci deleted in our samples seem to be involved in larger deletions, encompassing both the p and q arms of chromosome 5. Similar results have been reported for ovarian cancer2 and tumours of the papilla of Vater,18 in which the few losses in the short arm of chromosome 5 were involved in the largest chromosomal deletions.
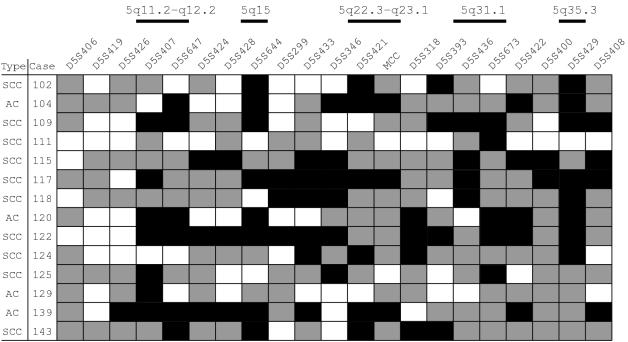
Figure 2.
Allelotyping results in non-small cell lung carcinoma. AC, adenocarcinoma; SCC, squamous cell carcinoma; black rectangles, loss of heterozygosity; white rectangles, retention of heterozygosity; grey rectangles, uninformative.
We also searched for microsatellite instability, in the form of deletion or expansion of dinucleotide repeats. None of the loci analysed displayed an alteration in the number of CA repeats and, therefore, we found no evidence of the RER+ phenotype in the cases under study. In fact, putting together our results and the ones reviewed elsewhere,10 the RER+ phenotype seems to be an infrequent event in non-small cell lung tumorigenesis.
Analysis of the deletion map (fig 2 ▶), together with the frequency of LOH obtained for each locus (table 1 ▶) allowed us to identify five regions on chromosome 5 displaying high rates of LOH.
The 5q11.2–q12.2 region, which encompasses the hMSH3 mismatch repair gene locus (5q11–q12), was deleted in approximately half of the informative cases. This gene is reported to be hemizygously deleted in 42% of non-small cell lung carcinoma samples,10 but no inactivating mutation in the residual hMSH3 allele was found. The hMSH3 protein is involved in heteromolecular complexes and, therefore, the authors suggested that it could play a role in lung tumorigenesis by a mechanism of dosage effect, affecting cellular functions other than the mismatch repair activity, because the RER+ phenotype was found very infrequently in the samples under study. Our data seem to agree with this hypothesis because we found no microsatellite instability in the samples with deletions encompassing the hMSH3 locus. Further studies to screen for mutations in the remaining allele are necessary to clarify the hMSH3 inactivation state.
LOH at the D5S644 locus (5q15) was detected in nearly half of the informative tumours. This region is proximal to the APC locus, so that this gene can be excluded as the candidate tumour suppressor gene in this region. To our knowledge, this is the first report implicating 5q15 in non-small cell lung carcinoma. A previous study in ovarian cancer2 defined a 22 cM interval on 5q13.1–q21, which also did not encompass the APC locus, supporting the hypothesis of a candidate tumour suppressor gene in this region of chromosome 5.
Our study also identified the 5q22.3–q23.1 region as being frequently deleted in non-small cell lung carcinoma. Analysis of the deletion map (fig 2 ▶; cases 102 and 139) indicates that the minimal region of deletion might be flanked by markers D5S346 and D5S318, defining a 2 cM segment that encompasses the APC and MCC loci. Ashton-Rickardt and colleagues6 and D'Amico and colleagues7 were the first authors to describe the APC/MCC region as being frequently deleted in lung cancer. Since then, several reports have indicated a high frequency of LOH at this region.8,9,11,19 Nevertheless, to date, no mutations have been reported in these genes.19 Moreover, several other genes have been cloned in this area,20 increasing the number of possible candidate tumour suppressor genes in this interval.
The 5q31.1 region has been reported as a target for frequent deletion in the myelodysplastic syndrome,21 oesophageal cancer,4 and gastric cancer.3 Within this region is the gene encoding interferon regulatory factor 1 (IRF-1), which functions as a tumour suppressor gene and is functionally inactivated in myelodysplastic syndrome22 and gastric cancer.23 The IRF-1 locus has not yet been implicated in lung tumorigenesis, but the high frequency of LOH on 5q31.1 obtained in our study makes IRF-1 a strong candidate for a tumour suppressor gene at this region.
The q telomere of chromosome 5 (5q33–q35) has been implicated previously in non-small cell lung carcinoma.8 Our study reveals a very high frequency of LOH in a more telomeric region, 5q35.3, not yet implicated in non-small cell lung carcinoma.
Loss of more than one region in the same tumour sample was seen in eight of 14 informative cases. We also found the concomitant deletion of three and four of the above mentioned regions in three of 14 and two of 14, respectively, of the informative samples. Deletions simultaneously involving two, three, and four of the above mentioned regions were found in both histological subtypes (fig 2 ▶), suggesting the involvement of more than one of these regions, either independently or simultaneously, in the genesis and/or development of non-small cell lung carcinoma.
Acknowledgments
This study was funded by PRODEP, CIENCIA/PRAXIS XXI, Liga Portuguesa Contra o Cancro-Núcleo Sul, CMDT/UNL. We are also grateful to Perkin Elmer-ABD for providing the linkage set of markers for chromosome 5.
References
- 1.Giaccone G. Oncogenes and antioncogenes in lung tumorigenesis. Chest 1996;109:130S–4S. [DOI] [PubMed] [Google Scholar]
- 2.Tavassoli M, Steingrimsdottir H, Pierce E, et al. Loss of heterozygosity on chromosome 5q in ovarian cancer is frequently accompanied by TP53 mutation and identifies a tumour suppressor gene locus at 5q13.1–21. Br J Cancer 1996;74:115–19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Tamura G, Ogasawara S, Nishizuka S, et al. Two distinct regions of deletion on the long arm of chromosome 5 in differentiated adenocarcinomas of the stomach. Cancer Res 1996;56:612–15. [PubMed] [Google Scholar]
- 4.Ogasawara S, Tamura G, Maesawa C, et al. Common deleted region on the long arm of chromosome 5 in esophageal carcinoma. Gastroenterology 1996;110:52–7. [DOI] [PubMed] [Google Scholar]
- 5.Horrigan SK, Westbrook CA, Kim AH, et al. Polymerase chain reaction-based diagnosis of del (5q) in acute myeloid leukemia and myelodysplastic syndrome identifies a minimal deletion interval. Blood 1996;88:2665–70. [PubMed] [Google Scholar]
- 6.Ashton-Rickardt PG, Wyllie AH, Bird CC, et al. MCC, a candidate familial polyposis gene in 5q.21, shows frequent allele loss in colorectal and lung cancer. Oncogene 1991;6:1881–6. [PubMed] [Google Scholar]
- 7.D'Amico D, Carbone DP, Johnson BE, et al. Polymorphic sites within the MCC and APC loci reveal very frequent loss of heterozygosity in human small cell lung cancer. Cancer Res 1992;52:1996–9. [PubMed] [Google Scholar]
- 8.Hosoe S, Uen K, Shigedo Y, et al. A frequent deletion of chromosome 5q21 in advanced small cell and non-small cell carcinoma of the lung. Cancer Res 1994;54:1787–90. [PubMed] [Google Scholar]
- 9.Fong KM, Zimmerman PV, Smith J. Microsatellite instability and other molecular abnormalities in non-small cell lung cancer. Cancer Res 1995;55:28–30. [PubMed] [Google Scholar]
- 10.Benachenhou N, Guiral S, Gorska-Flipot I, et al. High resolution deletion mapping reveals frequent allelic losses at the DNA mismatch repair loci hMLH1 and hMSH3 in non-small-cell lung cancer. Int J Cancer 1998;77:173–80. [DOI] [PubMed] [Google Scholar]
- 11.Wieland I, Böhm M, Arden KC, et al. Allelic deletion mapping on chromosome 5 in human lung carcinomas. Oncogene 1996;12:97–102. [PubMed] [Google Scholar]
- 12.Wu X, Zhao Y, Kemp BL, et al. Chromosome 5 aberrations and genetic predisposition to lung cancer. Int J Cancer 1998;79:490–3. [DOI] [PubMed] [Google Scholar]
- 13.Ziegle JS, Su Y, Corcoran KP, et al. Application of automated DNA sizing technology for genotyping microsatellite loci. Genomics 1992;14:1026–31. [DOI] [PubMed] [Google Scholar]
- 14.Cawkwell L, Bell SM, Lewis FA, et al. Rapid detection of allele loss in colorectal tumours using microsatellites and fluorescent DNA technology. Br J Cancer 1993;67:1262–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Marsh DJ, Zheng Z, Zedenius J, et al. Differential loss of heterozygosity in the region of the Cowden locus within 10q22–23 in follicular thyroid adenomas and carcinomas. Cancer Res 1997;57:500–3. [PubMed] [Google Scholar]
- 16.Conde AR, Martins G, Saraiva C, et al. Association of p53 genomic instability with the glutathione S-transferase null genotype in gastric cancer in the Portuguese population. J Clin Pathol: Mol Pathol 1999;52:131–4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Peralta RC, Casson AG, Wang R, et al. Distinct regions of frequent loss of heterozygosity of chromosome 5p and 5q in human esophageal cancer. Int J Cancer 1998;78:600–5. [DOI] [PubMed] [Google Scholar]
- 18.Achille A, Baron A, Zamboni G, et al. Chromosome 5 allelic losses are early events in tumours of the papilla of Vater and occur at sites similar to those of gastric cancer. Br J Cancer 1998;78:1653–60. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Cooper CA, Bubb VJ, Smithson N, et al. Loss of heterozygosity at 5q21 in non-small cell lung cancer: a frequent event but without evidence of apc mutation. J Pathol 1996;180:33–7. [DOI] [PubMed] [Google Scholar]
- 20.Ueno K, Kumagai T, Kijima T, et al. Cloning and tissue expression of cDNAs from chromosome 5q21–22 which is frequently deleted in advanced lung cancer. Hum Genet 1998;102:63–8. [DOI] [PubMed] [Google Scholar]
- 21.Le Beau MM, Espinosa III R, Neuman WL, et al. Cytogenetic and molecular delineation of the smallest commonly deleted region of chromosome 5 in malignant myeloid diseases. Proc Natl Acad Sci U S A 1993;90:5484–8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Harada H, Willison K, Sakakibara J, et al. Absence of the type-I IFN system in EC cells: transcriptional activator (IRF-1) and repressor (IRF-2) genes are developmentally regulated. Cell 1990;63:303–12. [DOI] [PubMed] [Google Scholar]
- 23.Nozawa H, Oda E, Ueda S, et al. Functionally inactivating point mutation in the tumour-suppressor IRF-1 gene identified in human gastric cancer. Int J Cancer 1998;77:522–7. [DOI] [PubMed] [Google Scholar]