Abstract
Residual feed intake (RFI) is a key indicator of feed efficiency, critical for enhancing the economic sustainability of poultry production. However, the genetic and metabolic regulatory mechanisms of RFI remain unclear. This study analyzed the genome, liver transcriptome, metabolome, and lipidome of hens with low and high feed efficiency (N = 60) from the previously established RFI divergent broiler lines (F15). Our results revealed pronounced genetic differentiation between low RFI (LRFI) and high RFI (HRFI) lines and identified genomic signatures of selection associated with feed efficiency. Transcriptomic analysis showed differential expression of genes involved in neural regulation and lipid metabolism. Notably, LRFI chickens exhibited reduced hepatic lipid accumulation, which was associated with decreased fatty acid metabolism and increased cholesterol metabolism (P < 0.05). The lipidomic analysis uncovered distinct profiles of glycerophospholipids (e.g., PE-P and PC-O) and sphingolipids (e.g., ceramides), which were more abundant in LRFI chickens (P < 0.05) and strongly correlated with key lipid metabolism processes (P < 0.05). Despite improved feed efficiency, LRFI chickens demonstrated signs of increased oxidative stress. Moreover, integrative analyses revealed that genes such as MGAT5, GABRA4, and LRRC4C, exhibiting strong selection signatures and higher expression in the LRFI line (P < 0.05), were identified as key regulators of lipid metabolism, potentially contributing to the observed differences in feed efficiency. This comprehensive study highlights the synergistic effect of genetics and lipid metabolism in driving feed utilization efficiency in chickens, establishing a scientific foundation for breeding strategies aimed at improving feed efficiency in poultry production.
Keywords: Chicken, RFI, Genetics, Lipid metabolism, Multiomics
Introduction
Feed efficiency is a critical economic trait in the poultry industry, directly impacting the profitability of farms. Improving feed efficiency during production not only reduces feeding costs and increases farming revenue but also decreases greenhouse gas emissions, promoting sustainable development in animal husbandry (Nkrumah et al., 2006). Residual feed intake (RFI), defined as the difference between an animal's actual feed intake and the expected intake required for maintenance and growth, is a key indicator of measuring feed efficiency, with lower RFI (LRFI) values indicating higher production efficiency (Koch et al., 1963). RFI is a moderately heritable trait controlled by genetics, and the presence of sufficient genetic variation within populations facilitates the selection for improved feed efficiency (Yi et al., 2018). Selective breeding for LRFI in chickens can enhance production efficiency, reduce abdominal fat deposition, and maintain body weight and meat quality (Willems et al., 2019). Therefore, RFI is considered a suitable indicator for assessing feed efficiency and is widely used in the breeding of high-quality chickens.
Identifying candidate genes associated with important traits helps establish precise molecular breeding strategies, improving breeding outcomes and efficiency, and promoting sustainable animal husbandry. GWAS and transcriptome sequencing in broiler populations have identified significant SNP loci and candidate genes associated with RFI, including ELOVL2, TRPA1, NSUN3, EPHA6, and AGK, as well as numerous other genes in tissues like the breast muscle, liver, and duodenum (Li et al., 2021; Wen et al., 2021). These genes are mainly involved in pathways related to appetite, lipid metabolism, digestion, energy homeostasis, and biosynthesis (Yang et al., 2020; Karimi et al., 2022). Despite the identification of many candidate genes, there is no consensus due to significant differences between breeds and lines. This discrepancy is primarily attributed to the fact that RFI is a complex quantitative trait that integrates various biological pathways regulated across multiple tissues. Focusing on a more specific biological process can aid in elucidating the feed efficiency trait.
It has been reported that characteristics of hepatic lipid metabolism, such as lipid composition and content, enzyme activity, hormone levels and the expression of lipid metabolism-related genes, vary in animals and poultry with different feed efficiencies (Alexandre et al., 2015; Jin et al., 2020; Sun et al., 2023). On the other hand, altering the lipid content or composition in feed can also impact animal growth performance and feed efficiency, as well as influence the expression of lipid metabolism-related genes (Crespo and Esteve-Garcia, 2001; Fan et al., 2021).Therefore, hepatic lipid metabolism influences feed efficiency in chickens through multifaceted regulatory mechanisms. The liver is the primary organ in chickens for lipid synthesis, processing, degradation, and redistribution (Nafikov and Beitz, 2007; Badmus et al., 2022; Guo et al., 2023). Hepatic lipid metabolism includes fatty acid synthesis, lipid transport, and lipid digestion (Reyer et al., 2017). Specifically, endogenous and exogenous free fatty acids (FFA) and glycerol are esterified to form triglycerides (TG) that are stored in the liver, to be later released into the bloodstream (Schulze et al., 2017; Friedman et al., 2018). TG combine with apolipoproteins and cholesterol to form very low-density lipoproteins (VLDL), which participate in lipid transport (Feingold, 2022). The liver is also the main site for bile acid synthesis, which plays a positive role in lipid digestion by emulsifying fats, promoting their transport and absorption in the intestines (Perino et al., 2021; Rezen et al., 2022). Identifying candidate genes and lipids that affect hepatic lipid metabolism can provide a theoretical basis for improving feed efficiency in chickens by reducing lipid deposition.
In previous research, our team has focused on Huiyang Bearded (HB) chickens to develop two divergent broiler selection lines for RFI since 2011 (Zou et al., 2023). Through multiple generations of selective breeding, significant phenotypic and genotypic segregation has emerged between the high and low RFI lines. This study aims to systematically investigate the molecular mechanisms by which genetic and metabolic factors synergistically influence RFI using multi-omics approaches. Additionally, it seeks to identify candidate genes and key lipids associated with RFI, providing a theoretical foundation for breeding feed-efficient poultry.
Materials and methods
Ethics statement
All birds in this study were handled in accordance with the ethical standards and approved protocols of the Institutional Animal Care and Use Committee of the Institute of Animal Science, Guangdong Academy of Agricultural Sciences, Guangzhou, China (Approval No. 2024013).
Animals and sample collection
This study utilized two divergent RFI selection lines (HRFI and LRFI) derived from a pure population of Huiyang Bearded (HB) chickens. All chickens were reared under uniform conditions from hatching, and the environmental conditions were standardized, with the temperature gradually reduced from 32°C at hatching to 22°C by six weeks of age, decreasing weekly. Throughout the experimental period, each bird was individually housed in a cage, with the temperature kept above 20°C and the relative humidity maintained at 50 % to 70 %. The lighting schedule was set to 16 hours of light and 8 hours of darkness (16L:8D). Within the first 155 days after hatching, all chickens were vaccinated against Marek's disease (MD), infectious bursal disease (IBD), Newcastle disease (ND), avian influenza (H5 and H9), fowl pox, infectious laryngotracheitis (ILT), infectious coryza (IC), and avian encephalomyelitis according to standard immunization protocols. Individual body weight (BW) and feed intake were measured on 1 d, and at 6 w, 8 w, and 10 w of age. RFI for each individual was calculated as the difference between observed and expected feed intake from 6 w to 10 w, following the breeding scheme and RFI calculation method previously documented (Zou et al., 2023), and the ingredients and nutrient composition of the diet are provided in Supplementary Table S1 (Xie et al., 2024). All hens were fed the same standard diet with unrestricted access to feed and water. In the F15 generation, 240 hens were maintained in the high line and 370 in the low line. At 70 weeks of age (490 days), 30 hens were randomly selected from each line. After weighing their body weight, the chickens were euthanized by cervical dislocation. Pentobarbital sodium (P3761, Sigma-Aldrich, St. Louis, MO, USA) was used to anesthetize the birds prior to cervical dislocation. The anesthetic was prepared at a concentration of 40 mg/mL, and a dose of 40 mg/kg body weight was administered via intraperitoneal injection. Liver tissue samples were immediately collected, weighed, flash-frozen in liquid nitrogen, and stored at −80°C for subsequent sequencing and biochemical analyses.
DNA extraction and sequencing
Twenty biological replicates were performed for DNA sequencing. Genomic DNA was extracted from the liver using the DNeasy Blood & Tissue Kit (69506, Qiagen, Hilden, Germany). The concentration and quality of the genomic DNA were assessed using a NanoDrop 2000 Spectrophotometer (Thermofisher, Waltham, MA, USA). Subsequently, the NEBNext DNA Library Prep Reagent Set (#E6000, New England BioLabs, Ipswich, MA, USA) was employed for constructing DNA libraries. Following library preparation, sequencing was carried out on an Illumina NovaSeq 6000 platform (Illumina, San Diego, CA, USA), producing 150 bp paired-end reads (LC-Bio Technology Co., Ltd, Hangzhou, China).
Reads alignment and variant detection
To reduce read bias, raw data underwent filtering based on the following criteria: (1) removing reads with > 10 % unidentified nucleotides; (2) excluding reads with > 10 adapter-aligned nucleotides, allowing ≤ 10 % mismatches; and (3) ensuring an average base quality above 20 (Phred-like score). High-quality sequences were then aligned to the Gallus gallus reference genome (GRCg7b) using the BWA MEM algorithm (v0.7.8) (Li and Durbin, 2009). SAMtools (v1.3) (Li et al., 2009) facilitated the conversion, indexing, and sorting of mapping results into BAM files. Duplicate read pairs were removed using Picard (v1.94) to refine the alignment. Single nucleotide polymorphisms (SNP) calling for each accession was performed using GATK HaplotypeCaller (v4.0.4) (McKenna et al., 2010), followed by further filtering based on specified criteria. PLINK (v1.9) was employed to filter SNP data (Purcell et al., 2007), setting parameters of sample call rate > 0.9, SNP call rate > 0.9, and MAF > 0.01. Remaining SNPs and individuals were imputed via BEAGLE (v5.2) (Browning et al., 2021), followed by a reanalysis with PLINK (v1.9). After processing, each sample retained 5,593,366 SNPs on average, which were then phased, filtered, and annotated using SNPEff (v5.2) (Cingolani et al., 2012) and the chicken reference genome. Each individual achieved a sequencing depth of over 10 ×, with an average genome coverage exceeding 98.0 %. After filtering, an average of 5,593,366 SNPs were retained for analysis (Supplementary Table S2).
Population genetics analysis
Principal Component Analysis (PCA) using Genome-wide Complex Trait Analysis (GCTA) software (v1.93.2) was employed to explore the genetic structure and reduce genotype data dimensionality, revealing major axes of variation and genetic relatedness among individuals (Yang et al., 2011). Population structure and admixture proportions were assessed using ADMIXTURE (v1.3) to delineate ancestral components (Alexander et al., 2009). Additionally, genetic relationships were constructed using the neighbor-joining (NJ) algorithm based on pairwise genetic distances between populations according to the polymorphic SNPs across the whole genome. The resulting relationships were visualized using Molecular Evolutionary Genetics Analysis (MEGA) software (v7) (Kumar et al., 2008).
Identification of runs of homozygosity (ROH) and selection sweep
ROH for each line were estimated using PLINK software (v1.9) with specific parameters (Purcell et al., 2007): –homozyg-density 50, –homozyg-gap 1,000, –homozyg-kb 500, –homozyg-window-het 1, –homozyg-window-snp 50, –homozyg-window-missing 5. The analysis revealed 1,737 ROHs in the HRFI line and 1,276 in the LRFI line, with chromosome 1 containing the highest number of ROHs and more than half located on chromosomes 1-4 (Supplementary Fig. S1A). Most ROHs were under 1,000 kb in length (> 88 %) (Supplementary Fig. S1B). The prevalence of SNPs within ROHs was quantified by counting the occurrences of specific SNPs across a diverse set of individuals, and the top 1 % of SNPs with the highest prevalence were selected. Then, the adjacent SNPs were combined into genomic regions, termed ROH islands.
Fixation index (Fst) values of SNPs across the genomes were calculated using VCFtools (v0.1.16) (Danecek et al., 2011), with a window size of 50 kb and a window step size of 25 kb. Converting the Fst values into Z-scores (Z-Fst values), and selecting the top 5 % of Z-Fst values indicated the most significant genetic differentiation between populations. The putative selected genes (PSGs) were annotated based on the reference genome (GRCg7b).
RNA sequencing and data analysis
Fifteen samples from each group were randomly selected from the 30 slaughtered samples for RNA-seq analysis. Total RNA extraction was carried out from the liver samples using Trizol reagent (15596018, Thermofisher, Waltham, MA, USA) following the manufacturer's guidelines. RNA quantity and purity were assessed using Bioanalyzer 2100 and RNA 6000 Nano LabChip Kit (5067-1511, Agilent, CA, USA). Transcriptome sequencing libraries were prepared from high-quality RNA samples with RIN numbers > 7.0, according to the standard Illumina RNAseq protocol. Subsequently, 150 bp paired-end sequencing (PE150) was conducted on an Illumina Novaseq 6000 (LC-Bio Technology CO., Ltd., Hangzhou, China) following the vendor's recommended protocol.
Reads containing adaptor contamination, low-quality bases, and undetermined bases were filtered out using Fastp (v0.20.1) (Chen et al., 2018). After data filtration, each sample yielded an average of more than 5 Gb of clean reads, which were aligned to the chicken reference genome (GRCg7b) using HISAT2 (v2.0.5) (Kim et al., 2015) with default parameters. SAM files were then converted to the BAM format using SAMtools (v1.11) (Li et al., 2009). Subsequently, count matrices for each gene were generated using HTSeq-count (v0.12.3) (Anders et al., 2015). Differentially expressed genes (DEGs) were identified using DESeq2 (v1.32.0) (Love et al., 2014), employing thresholds of baseMean (mean of normalized counts of all samples) > 10, adjusted P value (padj) < 0.05 and |Log2 (Fold change)| > 1.
Quantitative real-time PCR (qRT-PCR)
Total RNA was converted to complementary DNA (cDNA) using a PrimeScript™ RT Reagent Kit (RR047A, TaKaRa, Tokyo, Japan). qRT-PCR was conducted using an iTaq™ Universal SYBR® Green Supermix (1725124, Bio-Rad, Hercules, CA, USA) in accordance with the manufacturer's instructions. The β-actin gene served as the internal control, and the primer sequences for all candidate genes are listed in Supplementary Table S3.
Enrichment analysis
The Metascape online tool (http://metascape.org) was used to annotate genes. The Gene Ontology (GO) annotations were based on categories of biological process, cellular component, and molecular function. The Kyoto Encyclopedia of Genes and Genomes (KEGG) database was used to annotate pathways. A P value < 0.05 was set as the threshold for significant enrichment.
Lipid biochemical assays
Lipid biochemical assays were performed according to the manufacturer's instructions, with all assay kits provided by Nanjing Jiancheng Bioengineering Institute (Nanjing, China). The concentrations of TG (A110-2-1, 510 nm), FFA (A042-2-1, 546 (primary) / 600 nm (auxiliary)), total bile acid (TBA, E003-2-1, 505 nm), cholesterol (A111-1-1, 510 nm), low-density lipoproteins (LDL, A113-1-1, 546 nm), high-density lipoproteins (HDL, A112-1-1, 546 nm) and Malondialdehyde (MDA, A003-1, 532 nm) in livers were measured by colorimetric methods based on protein quantification (A045-2, 595 nm). Additionally, the concentration of very low-density lipoproteins (VLDL, H249-1) and 4-Hydroxynonenal (4-HNE, H268-1) were measured using an ELISA kit. Twenty biological replicates (the same as the genome sequencing samples) were performed for each assay.
Widely targeted metabolomic and lipidomic assays
Fifteen replicates from each group of liver biochemical assay samples were randomly selected, and metabolites and lipid contents were extracted and analyzed by MetWare (http://www.metware.cn/) for metabolomic and lipidomic analysis (see published reports for detailed methodology (Chen et al., 2013; Guo et al., 2023)). Metabolites and lipids were identified, and pathway mapping was performed using the KEGG database (http://www.kegg.jp/kegg). Significantly differences between groups were determined based on variable importance of the projection (VIP > 1) and a P value < 0.05, with VIP values calculated from OPLS-DA results using the R package MetaboAnalystR (v3.2) (Pang et al., 2020). To prevent overfitting, a permutation test with 200 permutations was conducted.
Statistical analysis
The data are presented as mean ± SD (standard deviation) and were subjected to analysis using a two-tailed Student's t-test. Each experiment was performed with a minimum of three replicates in multiple independent trials, and statistical significance was defined as a P value < 0.05. Pearson correlation analysis was used to estimate the relationship between genes and lipids. Bioinformatic analysis and visualization were performed using R (v4.3.1) and GraphPad Prism Software 9.
Results
Divergent RFI selection leads to genetic divergence in Huiyang Bearded chickens
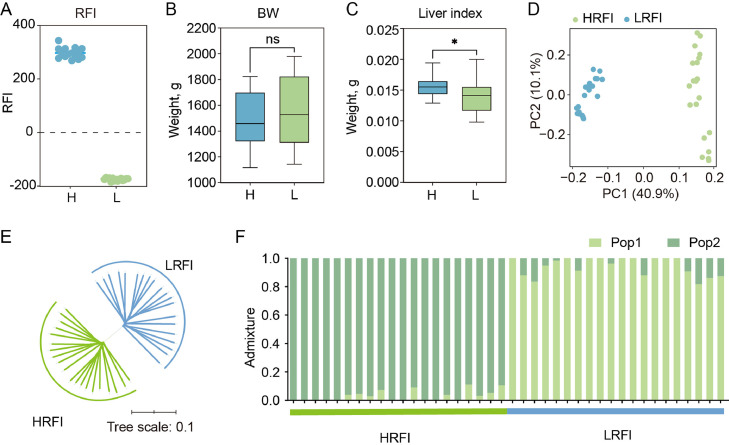
Thirty hens were randomly selected from a 15-generation RFI divergent selection population of HB chickens for sample collection. Significant differences in RFI were observed between the high and low RFI lines (Fig. 1A), while no significant difference was found in body weight (Fig. 1B), indicating that RFI selection did not affect body weight. However, the liver weight index was significantly lower in LRFI chickens compared to HRFI chickens (P value < 0.05) (Fig. 1C). To compare the genetic differences, chickens from each of the high and low RFI lines underwent whole-genome resequencing. A phylogenetic tree was constructed and revealed that individuals from the same line tended to cluster into a single group, which is corroborated by PCA (Fig. 1D and E). In the analysis of population structure, distinct clustering of high and low RFI lines was observed when K = 2 (Fig. 1F). These results underscores the successful breeding strategy that has led to the independent grouping of the high and low RFI lines across multiple generations of selection.
Fig. 1.
Differences in the genetic structure between high and low RFI chickens. (A) RFI values of individuals from the high and low RFI lines. (B) Body weights of individuals from the high and low RFI lines. (C) Liver weight index (liver weight/body weight) of individuals from the high and low RFI lines. The values are the mean ± SEM, * represents P < 0.05. (D) PCA of the RFI population. (E) Neighbor-joining tree analysis of the RFI population. (F) Admixture analysis of the RFI population.
Genetic selection and expression profiles drive RFI divergence in chickens
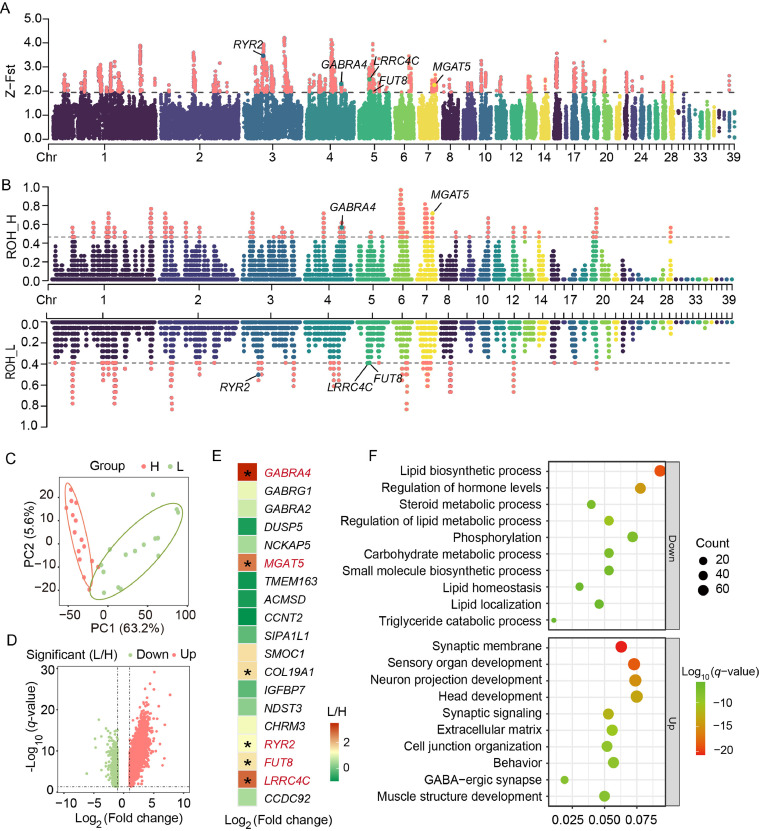
Next, we performed selective sweep analysis to investigate the genetic basis of the differences in RFI divergent chickens. Normalized window-based Fst values, expressed as z-scores, identified the top 5 % as potential selection regions, including 1,489 annotated genes (Fig. 2A, Supplementary Table S4). These genes were primarily involved in pathways such as "regulation of MAPK cascade", "regulation of cell growth", "muscle structure development", "protein kinase activity", and "GABAergic synapse" (Supplementary Fig. S2). We also performed a genome-wide ROH analysis, to pinpoint ROH islands, the top 1 % regions with the highest SNP density were selected. In the HRFI line, 29 ROH islands were identified, averaging 733.5 kb in length and containing an average of 2,809 SNPs each. In the LRFI line, 26 ROH islands were identified, averaging 727.0 kb in length and containing an average of 2,710 SNPs each. The islands identified included 591 annotated genes in the HRFI line (Fig. 2B, Supplementary Table S5) and 526 annotated genes in the LRFI line (Fig. 2B, Supplementary Table S6). By overlapping the top 5 % Z-Fst genes with the ROH islands, we identified the positively selected genes (PSGs), resulting in 11 genes in the HRFI line and 8 genes in the LRFI line (Supplementary Table S7).
Fig. 2.
Genetic selection and differential expression analysis identify important genes. (A) Manhattan plot of Z-Fst for the RFI population, highlighting the top 5 % as significant regions. (B) Manhattan plot of ROH islands for high and low RFI lines, highlighting the top 1 % as significant regions. (C) PCA of samples from high (red) and low (green) RFI lines. (D) Volcano plot of DEGs between high and low RFI lines. Red indicates up-regulated genes, green indicates down-regulated genes. (E) Expression of PSGs in the RFI population (LRFI/HRFI). Asterisks indicate P value < 0.05, and genes in red are both PSGs and DEGs. (F) Enrichment analysis for upregulated and downregulated DEGs between high and low RFI lines.
Using the liver as the target tissue, differentially expressed genes (DEGs) and regulatory pathways were obtained by RNA-seq to elucidate the underlying mechanisms of RFI differences. Principal component analysis (PCA) confirmed the clear separation between the two groups (Fig. 2C), reflecting the high reproducibility of the transcriptomic profiles within different RFI groups, with 63.2 % of the RFI variation explained along the PC1 direction. Pairwise comparative analysis was performed to identify DEGs. Specifically, 2,117 up-regulated genes and 846 down-regulated genes were identified between LRFI and HRFI lines (Fig. 2D, Supplementary Table S8). Subsequently, we analyzed the expression differences of PSGs between the high and low lines and identified six genes that were significantly differentially expressed between the two groups. After further filtering using baseMean (> 10), five key RFI-related functional genes were highlighted, including GABRA4 and MGAT5 in the HRFI line, as well as RYR2, FUT8, and LRRC4C in the LRFI line (Fig. 2E).
Functional enrichment analysis revealed that up-regulated DEGs in the LRFI line were primarily involved in neural and developmental processes such as "synaptic signal transduction", "neuron development", and "tissue morphogenesis". Conversely, down-regulated DEGs were mainly associated with lipid metabolism, including "lipid biosynthesis", "lipid localization", "lipid homeostasis", and "steroid metabolism" (Fig. 2F). These results indicate significant differences in the expression of neural regulation and lipid metabolism-related genes in liver tissues between the HRFI and LRFI lines.
Significant differences in fatty acid and cholesterol metabolism across RFI divergent chickens
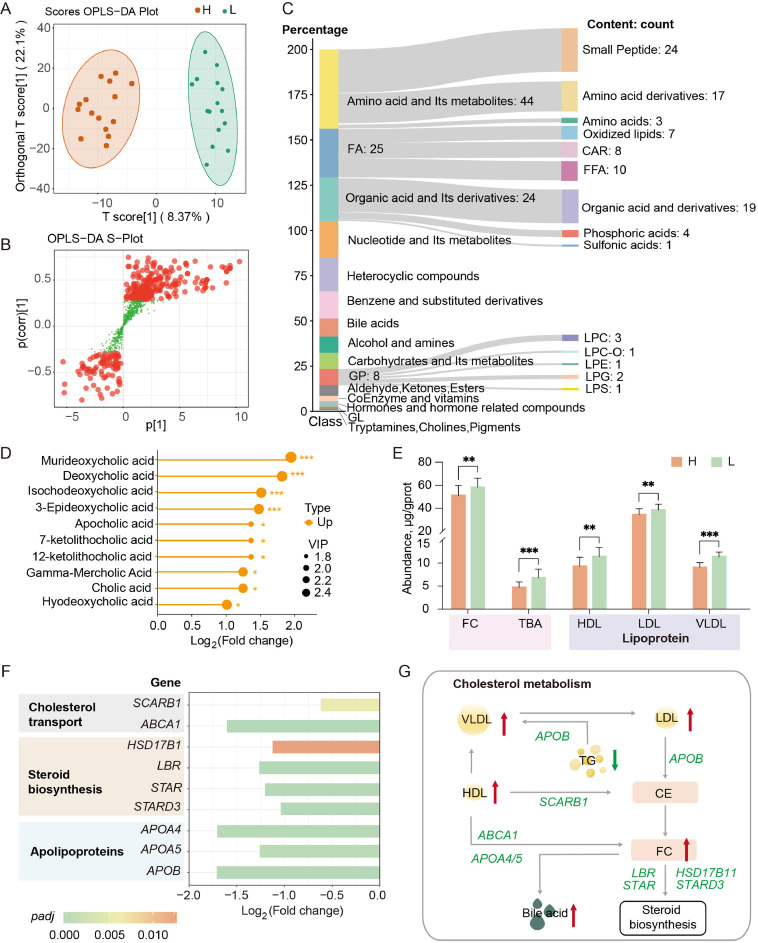
To identify the liver metabolic differences between HRFI and LRFI individuals, a metabolomic analysis was conducted on their liver samples. A total of 1,206 metabolites were identified, organized into 15 categories. Using the OPLS-DA model to analyze the data, a clear distinction between the two sets of data was observed, with significant differences in metabolite levels (Fig. 3A and B). The differential metabolites were predominantly in the categories of "amino acid and its metabolites", "fatty acid", and "organic acid and its derivatives" (Fig. 3C). Additionally, several glycerophospholipids (GPs) showed significant differences. These differential metabolites were significantly enriched in multiple lipid metabolism pathways, including "fatty acid biosynthesis", "fatty acid degradation", and "biosynthesis of unsaturated fatty acids" (Supplementary Fig. S3). These results highlight significant differences in liver fatty acid metabolism between high and low RFI chickens.
Fig. 3.
Differences in liver metabolites between high and low RFI lines. (A) OPLS-DA score plot of metabolites from high and low RFI line samples. (B) OPLS-DA S-plot. Red points indicate metabolites with VIP > 1, green points denote VIP ≤ 1. (C) Classification and quantity of differential metabolites. (D) Variations in bile acid content between high and low RFI lines (LRFI/HRFI). (E) Levels of free cholesterol (FC), total bile acid (TBA), high-density lipoprotein (HDL), low-density lipoprotein (LDL), and very low-density lipoprotein (VLDL) by lipid biochemical analysis. The values are the mean ± SEM, * represents P < 0.05, ** represents P < 0.01 and *** represents P < 0.001. (F) Differential gene expression related to cholesterol transport, steroid biosynthesis and apolipoproteins (LRFI/HRFI). (G) Schematic representation of lipid content and gene expression changes in cholesterol metabolism pathways in the liver of the LRFI line.
Given the crucial role of bile acids in lipid metabolism, we examined changes in bile acid content and found that all ten identified bile acids were increased in the LRFI line (Fig. 3D). To validate these findings, we used lipid biochemical assays to measure the levels of total bile acid and its essential precursor, cholesterol. The results were consistent with those of sequencing data, showing significantly higher levels of total bile acids and cholesterol in the LRFI line (Fig. 3E). Moreover, the levels of lipoproteins involved in triglyceride and cholesterol transport, such as HDL, LDL, and VLDL, were significantly elevated in the LRFI line (Fig. 3E). At the transcription level, genes involved in cholesterol transport (e.g., ABCA1) and apolipoproteins (APOA and APOB) showed significant changes in expression (Fig. 3F). Furthermore, cholesterol is a vital precursor for steroid hormone synthesis. We observed significant changes in the expression of genes involved in steroid biosynthesis, such as HSD17B11, LBR, STAR, and STARD3 (Fig. 3F). These findings indicate substantial differences in cholesterol synthesis, transport, and steroid hormone synthesis in the liver between the RFI divergent lines (Fig. 3G), suggesting that variations in feed efficiency may be linked to cholesterol metabolism.
Lipidomic analysis reveals reduced fat deposition and increased oxidative stress in LRFI chickens
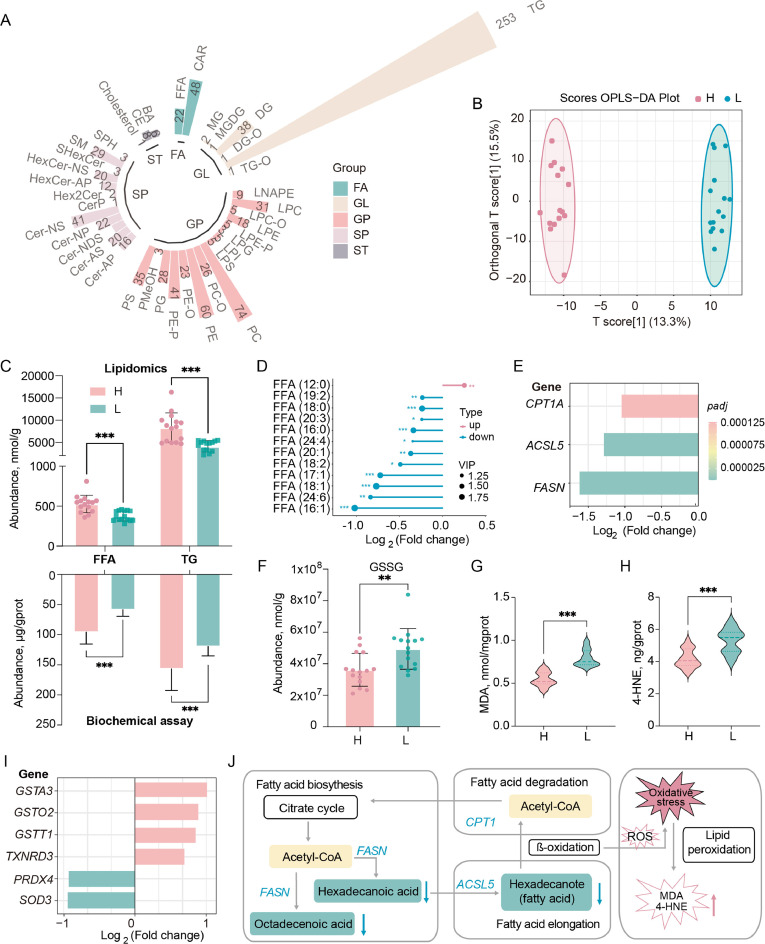
To further determine the specific lipids associated with RFI, the lipid composition and quantity in the high and low RFI lines were determined by lipidomics. A total of 934 lipids were identified, organized into 5 categories comprising 40 lipid classes (Fig. 4A). The lipid profiles of the high and low RFI lines were clearly distinguishable (Fig. 4B), with 358 lipids showing significant differences in content between the two groups (Supplementary Table S9). Notably, the contents of FFA and TG were markedly lower in the LRFI line compared to the HRFI line (Fig. 4C, upper). Complementary biochemical assays of hepatic FFA and TG contents corroborated the lipidomics findings (Fig. 4C, lower). Specifically, among the 12 detected FFAs, 11 long-chain FFAs exhibited significant reductions in the LRFI line, with the exception of FFA (12:0) (Fig. 4D). Furthermore, the expression levels of key genes involved in lipid synthesis, elongation, and degradation were significantly downregulated in the LRFI line (Fig. 4E, Supplementary Table S10).
Fig. 4.
Differences in liver lipid profiles across RFI divergent chickens. (A) Lipid class and quantity detected by lipidomic analysis. (B) OPLS-DA score plot of metabolites in high and low RFI samples. (C) Free fatty acid (FFA) and triglyceride (TG) contents in lipidomic and lipid biochemical analysis. (D) Differential content of FFA between high and low RFI groups (LRFI/HRFI). (E) Differential expression of key genes in lipid metabolism pathways (LRFI/HRFI). (F) GSSG levels in high and low RFI chicken livers. (G) MDA levels in high and low RFI chicken livers. (H) 4-HNE levels in high and low RFI chicken livers. (I) Differential expression of oxidative stress-related genes (LRFI/HRFI). (J) Schematic representation of changes in lipid content and gene expression in fatty acid metabolism pathways in the LRFI chicken livers. The values are the mean ± SEM, * represents P < 0.05, ** represents P < 0.01 and *** represents P < 0.001.
Interestingly, oxidized glutathione (GSSG), a crucial antioxidant, was significantly elevated in the LRFI line (Fig. 4F). Lipid peroxidation typically produces malondialdehyde (MDA) and 4-hydroxynonenal (4-HNE). Upon measuring their concentrations, both were found to be significantly increased in the LRFI line (Fig. 4G and H). Correspondingly, the expression levels of oxidative stress-related genes also exhibited significant changes. Specifically, the expression of antioxidant enzymes such as glutathione S-transferase (GSTA3, GSTO2, GSTT1) and thioredoxin reductase (TXNRD3) were significantly upregulated, whereas the expression of superoxide dismutase (SOD3) and peroxiredoxin (PRDX4) was significantly downregulated (Fig. 4I). These findings suggest that LRFI chickens exhibit reduced hepatic fatty acid synthesis, elongation, and deposition, but experience an imbalance in reactive oxygen species (ROS) production and antioxidant capacity, leading to hepatic oxidative stress (Fig. 4J).
Correlation analysis reveals key genes and their associations with lipids in chickens with differing RFI
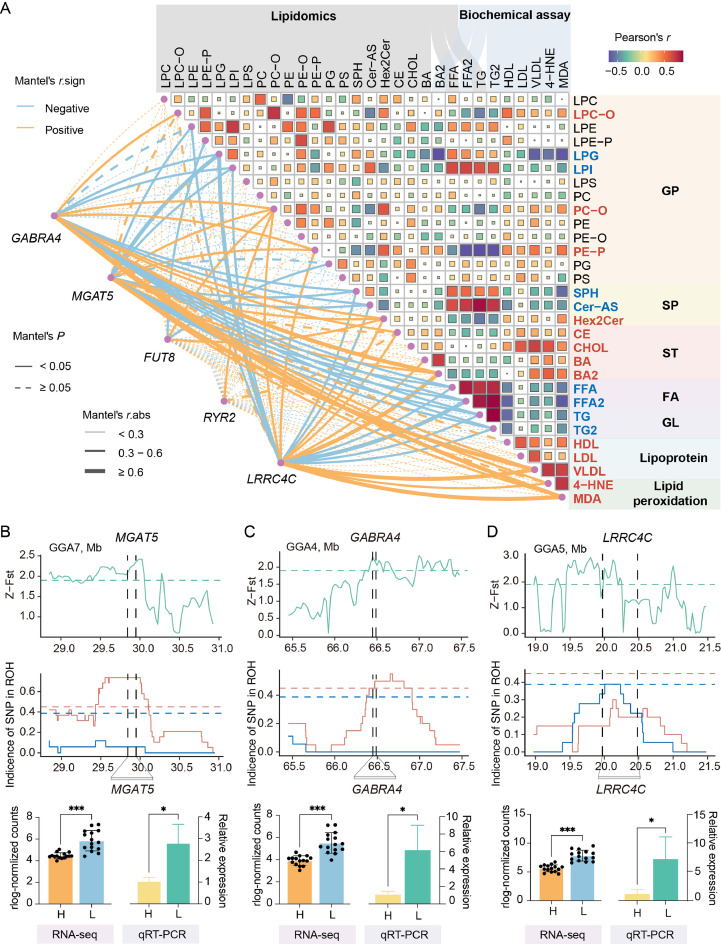
To identify lipids associated with the differences in hepatic fatty acid deposition between high and low RFI chickens, a correlation analysis was performed on lipid classes. The analysis revealed that FFA was not only significantly positively correlated with TG but also significantly correlated with several GPs (Fig. 5A). Specifically, FFA was positively correlated with lysophosphatidylglycerol (LPG) and lysophosphatidylinositol (LPI), while it exhibited a negative correlation with phosphatidylethanolamine (PE-P) and phosphatidylcholine (PC-O) (Fig. 5A). Furthermore, LPG and LPI showed significant negative correlations with indicators of lipid transport (VLDL), digestion (BA), and oxidative stress (4-HNE, MDA), while PC-O and PE-P were significantly positively correlated with these indicators. Moreover, some sphingolipids (SPs), such as sphingosine (SPH) and ceramide (Cer-AS) also had significant positive correlations with FFA and TG, and negative correlations with lipid transport (HDL, VLDL) and oxidative stress indicators (4-HNE, MDA) (Fig. 5A). Additionally, correlation analysis indicated that apolipoproteins (HDL, LDL, VLDL) and lipid peroxides (4-HNE, MDA) were positively correlated with sterol lipids (STs) such as cholesterol and bile acids, but negatively correlated with FFA and TG (Fig. 5A). Apolipoproteins are crucial for lipid transport, while oxidative stress is linked to the overload of lipids such as FFA, TG, and cholesterol (Arroyave-Ospina et al., 2021). These findings suggest that the difference in RFI may be related to lipid metabolism processes, such as hepatic lipid deposition, transport, digestion and oxidative stress, in which various GPs and SPs potentially play a role.
Fig. 5.
Important candidate genes and lipids associated with RFI. (A) The relationship between lipid profiles and key candidate genes. Lipid data were obtained from lipidomics and biochemical assays. The heatmap on the right displays the Pearson correlation coefficients between lipid pairs, where red and blue squares represent positive and negative correlations, respectively. The size of each square reflects the absolute value of the correlation coefficient. Lipid names in red or blue indicate positive or negative correlations, respectively, with lipid transport or oxidative stress indicators. The lines on the left represent the correlation between candidate gene expression and lipid content. Orange and blue lines indicate positive and negative correlations, respectively, with line thickness representing the strength of the correlation. Solid lines indicate significant correlations (P value < 0.05), while dashed lines indicate non-significant correlations (P value ≥ 0.05). (B-D) Selection sweep regions near the MGAT5 (B), GABRA4 (C), and LRRC4C (D) genes, and their expression levels in high and low RFI lines using RNA-seq and qRT-PCR. The values are the mean ± SEM, * represents P < 0.05, ** represents P < 0.01 and *** represents P < 0.001.
Correlation analysis was also performed on lipid classes and the 5 key RFI-related functional genes to further identify candidate genes that play a role in lipid metabolism. Notably, FUT8 and RYR2 showed relatively low correlations with various lipids (Fig. 5A), suggesting their involvement in other biological processes influencing RFI. The remaining three genes, GABRA4, MGAT5, LRRC4C, showed significant negative correlations with lipid deposition indicators (FFA and TG), as well as certain GPs (LPG and LPI) and SPs (SPH and Cer-AS). Conversely, they exhibited significant positive correlations with two GPs (PE-P and PC-O), and indicators related to lipid digestion (BA), transport (VLDL), and oxidative stress (MDA) (Fig. 5A). Since these highly correlated GPs and SPs were previously identified as important lipids, it is hypothesized that GABRA4, MGAT5, and LRRC4C play a role in hepatic lipid metabolism. Therefore, as the MGAT5, GABRA4, and LRRC4C genes have been subject to selection in the RFI selection lines, exhibit significant differential expression between the high and low lines (Fig. 5B–D and Supplementary Fig. S4A and B), and show strong correlations with lipid metabolism, they are considered important candidate genes involved in hepatic lipid metabolism and feed efficiency.
Discussion
This study constructed multiomics profiles to investigate the genetic and metabolic differences between RFI divergent lines, identifying significant candidate genes and lipids influencing hepatic lipid metabolism and RFI. Our findings provide unique insights into the regulatory mechanisms of feed efficiency in chickens from the perspective of genetics and lipid metabolism.
Differences in hepatic lipid metabolism between HRFI and LRFI chickens
RFI, independent of body weight and other production traits, reflects differences in basal metabolism caused by genetic background by partitioning animal energy into growth and maintenance energy. Studies on divergent selection populations of Angus steers for RFI found that metabolic processes, body composition, and energy expenditure explained two-thirds of RFI variation (Herd and Arthur, 2009). Among these, lipid metabolism is a crucial factor, as fat has the highest energy density. Animals with lower feed efficiency tend to accumulate more body fat, which increases energy expenditure and reduces feed efficiency (Bai et al., 2022; Zhang et al., 2023). Our previous work with divergent RFI chicken lines found that LRFI chickens had lower abdominal fat and higher breast muscle rates than HRFI chickens (Zou et al., 2023), and this study further confirmed their significantly lower lipid deposition. These findings suggest that LRFI animals are capable of optimizing energy allocation, leading to a reduction in lipid deposition and an enhancement in feed efficiency.
Cholesterol is a component of biological membranes and plays a crucial role in the integrity and regulation of signaling pathways (Luo et al., 2020). Cholesterol mainly maintains homeostasis through esterification to cholesterol ester (CE), an essential component of lipid droplets besides triglycerides. Furthermore, cholesterol is a precursor for bile acid synthesis, which aids in lipid digestion and hydrolysis, promoting fat transport and absorption in the intestine (Hofmaenner et al., 2022). In this context, we propose a link between feed efficiency and cholesterol metabolism, wherein increased bile acid synthesis may enhance lipid digestion and transport, thereby reducing liver lipid deposition to some extent.
Increased feed utilization may elevate oxidative stress risk
Oxidative stress refers to the imbalance between the production of reactive oxygen species (ROS) and antioxidant defenses, where excessive ROS cannot be neutralized by the antioxidant defense mechanisms (Forman and Zhang, 2021). The primary source of ROS is the mitochondrial electron transport chain (ETC), and it may also arise from inadequate fatty acid β-oxidation (Arroyave-Ospina et al., 2021), leading to the formation of lipid peroxides such as malondialdehyde (MDA) and 4-hydroxy-2-nonenal (4-HNE), which cause cellular damage. The strict balance between antioxidant defenses is crucial for preventing the accumulation of oxidants and free radicals (Goicoechea et al., 2023). The occurrence of oxidative stress in the LRFI line is attributed to increased metabolic activity and nutrient processing in the liver, which leads to ROS production. Previous reports in broilers also found a correlation between high feed efficiency and increased hepatic oxidative stress markers (Bottje and Carstens, 2009), and similar results have been observed in other livestock, where animals with fast growth rates or high feed conversion rates tend to have higher oxidative stress levels (Pamplona and Costantini, 2011; Russell et al., 2016). Therefore, while selective breeding for improved feed efficiency benefits growth and productivity, it also increases the risk of oxidative stress, especially in the liver. This highlights the importance of balancing growth and feed efficiency with oxidative stress mitigation measures, such as antioxidant supplementation and dietary management.
MGAT5, GABRA4, and LRRC4C are dey candidate genes
MGAT5, a key glycosyltransferase involved in protein glycosylation, plays crucial roles in maintaining cell structure, regulating cell adhesion, and contributing to immune and metabolic homeostasis (de-Souza-Ferreira et al., 2023). Studies show that Mgat5 deficiency in mice disrupts metabolic processes, leading to impaired insulin receptor function and increased glycogen storage in hepatocytes (Johswich et al., 2014). Despite high-energy feeding, MGAT5 deficiency can inhibit weight gain and hepatic fat accumulation, while increased MGAT5 activity accelerates triglyceride synthesis and VLDL secretion (Cheung et al., 2007; Kamada et al., 2016). These correlations suggest that MGAT5 may modulate liver lipid metabolism, with further research needed to clarify its role in chicken feed efficiency.
Additionally, GABRA4 and LRRC4C were found to be under selection during the divergent selection for RFI, showing differential expression between high and low RFI lines. Though these genes have not been directly linked to lipid metabolism or feed efficiency, they encode proteins crucial for neural processes in the hypothalamus, a key regulator of food intake and energy balance (Choi et al., 2019; Gupta and Sharma, 2019; Sajan et al., 2024). The hypothalamus integrates signals related to lipid levels and regulates metabolism through complex neural pathways(Geller et al., 2019; Goldstein et al., 2021). Sphingolipid signaling in the hypothalamus, particularly via S1P, influences food intake and energy homeostasis, with disruptions potentially leading to obesity and metabolic disorders (Silva et al., 2014; Vozella et al., 2019). These findings support the role of sphingolipid-mediated signaling in hypothalamic control of food intake. Thus, GABRA4 and LRRC4C may participate in regulating hepatic lipid metabolism related to feed efficiency by altering neural development and function.
RYR2 and FUT8 are two genes that cannot be overlooked. RYR2 is primarily associated with cardiac function and calcium regulation (Steinberg et al., 2023), while FUT8 is renowned for its role in fucosylation (Bastian et al., 2021). This study identifies them as potential candidate genes, despite their low correlation with lipids, they may be involved in influencing feed efficiency through alternative pathways.
Conclusions
In summary, this study systematically compared the genomic, transcriptomic, metabolomic, and lipidomic differences between RFI divergent lines. The findings revealed that LRFI chickens have lower liver lipid deposition, likely due to reduced lipid synthesis and increased lipid transport and bile acid synthesis; however, improving feed efficiency may increase the risk of oxidative stress. Furthermore, this study identified certain lipids (cholesterol, PE-P, PC-O, and Cer) and genes (MGAT5, GABRA4, and LRRC4C) as important candidates influencing liver lipid metabolism and feed efficiency. These results indicate a complex relationship between liver lipid metabolism and feed utilization efficiency, providing valuable insights for improving livestock feed efficiency and productivity.
Supplementary information
Supplementary Table S1. Ingredients and nutrient composition of diets.
Supplementary Table S2. Summary of resequencing data generated in this study.
Supplementary Table S3. The primer sequences used in this study.
Supplementary Table S4. List of genes in the selected regions by top 5 % highest Z-Fst.
Supplementary Table S5. List of genes in the selected regions by ROH islands in the HRFI group.
Supplementary Table S6. List of genes in the selected regions by ROH islands in the LRFI group.
Supplementary Table S7. List of genes in the selected regions by top 5 % highest Z-Fst and ROH islands.
Supplementary Table S8. Differentially expressed genes identified by RNA-seq.
Supplementary Table S9. Differentially lipids identified by lipidomics.
Supplementary Table S10. Enrichment of DEGs.
Supplementary Fig. S1. ROH information in high and low RFI lines. (A) Number of ROH segments in high and low RFI lines across different chromosomes. (B) Distribution of ROH lengths in high and low RFI lines. Short: 500 -1,000 kb, Medium: 1,000-3,000 kb, Long: > 3,000 kb.
Supplementary Fig. S2. Enrichment analysis of differential metabolites.
Supplementary Fig. S3. Enrichment analysis of top 5 % Z-Fst genes.
Supplementary Fig. S4. Selection sweep regions near the RYR2 (A) and FUT8 (B) genes and the expression levels of these genes in high and low RFI lines.
Declaration of competing interest
The authors declare no competing interests.
Acknowledgements
Fundings: This work was supported by the Special Fund for the Promotion of Breeding Industry in the 2024 Guangdong Provincial Rural Revitalization Strategy (2024-XPY-00-002), the 2023 Independent Research Projects of the State Key Laboratory of Swine and Poultry Breeding Industry (2023QZ-NK01), the 2023 Provincial Agricultural Science and Technology Innovation and Extension System Construction Project (2023KJ106), the Earmarked Fund for Modern Agri-industry Technology Research System (CARS-41), and the China Postdoctoral Science Foundation (2024M760600).
Author contributions: CL and XG conceived and designed the project. CL, DS, HQ, JJ, and XZ established and managed the population resources. CL and DS provided financial and technical support. XG and JL collected samples. XG, JL and JS performed experiments. XG and XL analyzed the data. XG created visualizations and wrote the manuscript. CL revised the manuscript. All authors read and approved the final draft.
Footnotes
Section: Genetics and Molecular Biology
Supplementary material associated with this article can be found, in the online version, at doi:10.1016/j.psj.2025.104885.
Appendix. Supplementary materials
Data availability
All of the sequencing data in this study have been deposited into NCBI and can be accessed via accession PRJNA1149694.
References
- Alexander D.H., Novembre J., Lange K. Fast model-based estimation of ancestry in unrelated individuals. Genom. Res. 2009;19:1655–1664. doi: 10.1101/gr.094052.109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Alexandre P.A., Kogelman L.J., Santana M.H., Passarelli D., Pulz L.H., Fantinato-Neto P., Silva P.L., Leme P.R., Strefezzi R.F., Coutinho L.L., Ferraz J.B., Eler J.P., Kadarmideen H.N., Fukumasu H. Liver transcriptomic networks reveal main biological processes associated with feed efficiency in beef cattle. BMC Genom. 2015;16:1073. doi: 10.1186/s12864-015-2292-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Anders S., Pyl P.T., Huber W. HTSeq–a Python framework to work with high-throughput sequencing data. Bioinformatics. 2015;31:166–169. doi: 10.1093/bioinformatics/btu638. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Arroyave-Ospina J.C., Wu Z., Geng Y., Moshage H. Role of oxidative stress in the pathogenesis of non-alcoholic fatty liver disease: implications for prevention and therapy. Antioxid. (Basel) 2021;10:174. doi: 10.3390/antiox10020174. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Badmus O.O., Hillhouse S.A., Anderson C.D., Hinds T.D., Stec D.E. Molecular mechanisms of metabolic associated fatty liver disease (MAFLD): functional analysis of lipid metabolism pathways. Clin. Sci. (Lond.) 2022;136:1347–1366. doi: 10.1042/CS20220572. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bai H., Guo Q., Yang B., Dong Z., Li X., Song Q., Jiang Y., Wang Z., Chang G., Chen G. Effects of residual feed intake divergence on growth performance, carcass traits, meat quality, and blood biochemical parameters in small-sized meat ducks. Poult. Sci. 2022;101 doi: 10.1016/j.psj.2022.101990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bastian K., Scott E., Elliott D.J., Munkley J. FUT8 alpha-(1,6)-fucosyltransferase in cancer. Int. J. Mol. Sci. 2021;22:455. doi: 10.3390/ijms22010455. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bottje W.G., Carstens G.E. Association of mitochondrial function and feed efficiency in poultry and livestock species. J. Anim. Sci. 2009;87:E48–E63. doi: 10.2527/jas.2008-1379. [DOI] [PubMed] [Google Scholar]
- Browning B.L., Tian X., Zhou Y., Browning S.R. Fast two-stage phasing of large-scale sequence data. Am. J. Hum. Genet. 2021;108:1880–1890. doi: 10.1016/j.ajhg.2021.08.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen S., Zhou Y., Chen Y., Gu J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics. 2018;34:i884–i890. doi: 10.1093/bioinformatics/bty560. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen W., Gong L., Guo Z., Wang W., Zhang H., Liu X., Yu S., Xiong L., Luo J. A novel integrated method for large-scale detection, identification, and quantification of widely targeted metabolites: application in the study of rice metabolomics. Mol. Plant. 2013;6:1769–1780. doi: 10.1093/mp/sst080. [DOI] [PubMed] [Google Scholar]
- Cheung P., Pawling J., Partridge E.A., Sukhu B., Grynpas M., Dennis J.W. Metabolic homeostasis and tissue renewal are dependent on beta1,6GlcNAc-branched N-glycans. Glycobiology. 2007;17:828–837. doi: 10.1093/glycob/cwm048. [DOI] [PubMed] [Google Scholar]
- Choi Y., Park H., Jung H., Kweon H., Kim S., Lee S.Y., Han H., Cho Y., Kim S., Sim W.S., Kim J., Bae Y., Kim E. NGL-1/LRRC4C deletion moderately suppresses hippocampal excitatory synapse development and function in an input-independent manner. Front. Mol. Neurosci. 2019;12:119. doi: 10.3389/fnmol.2019.00119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cingolani P., Platts A., Wang le L., Coon M., Nguyen T., Wang L., Land S.J., Lu X., Ruden D.M. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly (Austin) 2012;6:80–92. doi: 10.4161/fly.19695. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Crespo N., Esteve-Garcia E. Dietary fatty acid profile modifies abdominal fat deposition in broiler chickens. Poult. Sci. 2001;80:71–78. doi: 10.1093/ps/80.1.71. [DOI] [PubMed] [Google Scholar]
- Danecek P., Auton A., Abecasis G., Albers C.A., Banks E., DePristo M.A., Handsaker R.E., Lunter G., Marth G.T., Sherry S.T., McVean G., Durbin R., G. Genomes Project Analysis The variant call format and VCFtools. Bioinformatics. 2011;27:2156–2158. doi: 10.1093/bioinformatics/btr330. [DOI] [PMC free article] [PubMed] [Google Scholar]
- de-Souza-Ferreira M., Ferreira E.E., de-Freitas-Junior J.C.M. Aberrant N-glycosylation in cancer: MGAT5 and beta1,6-GlcNAc branched N-glycans as critical regulators of tumor development and progression. Cell Oncol. (Dordr.) 2023;46:481–501. doi: 10.1007/s13402-023-00770-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fan Z., Li J., Zhang Y., Wu D., Zheng X., Wang C., Wang L. Excessive dietary lipid affecting growth performance, feed utilization, lipid deposition, and hepatopancreas lipometabolism of large-sized common carp (cyprinus carpio) Front. Nutr. 2021;8 doi: 10.3389/fnut.2021.694426. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Feingold K.R. Lipid and lipoprotein metabolism. Endocrinol. Metab. Clin. North Am. 2022;51:437–458. doi: 10.1016/j.ecl.2022.02.008. [DOI] [PubMed] [Google Scholar]
- Forman H.J., Zhang H. Targeting oxidative stress in disease: promise and limitations of antioxidant therapy. Nat. Rev. Drug. Discov. 2021;20:689–709. doi: 10.1038/s41573-021-00233-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Friedman S.L., Neuschwander-Tetri B.A., Rinella M., Sanyal A.J. Mechanisms of NAFLD development and therapeutic strategies. Nat. Med. 2018;24:908–922. doi: 10.1038/s41591-018-0104-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Geller S., Arribat Y., Netzahualcoyotzi C., Lagarrigue S., Carneiro L., Zhang L., Amati F., Lopez-Mejia I.C., Pellerin L. Tanycytes regulate lipid homeostasis by sensing free fatty acids and signaling to key hypothalamic neuronal populations via FGF21 secretion. Cell. Metab. 2019;30:833–844. doi: 10.1016/j.cmet.2019.08.004. e837. [DOI] [PubMed] [Google Scholar]
- Goicoechea L., Conde de la Rosa L., Torres S., Garcia-Ruiz C., Fernandez-Checa J.C. Mitochondrial cholesterol: metabolism and impact on redox biology and disease. Redox. Biol. 2023;61 doi: 10.1016/j.redox.2023.102643. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goldstein N., McKnight A.D., Carty J.R.E., Arnold M., Betley J.N., Alhadeff A.L. Hypothalamic detection of macronutrients via multiple gut-brain pathways. Cell. Metab. 2021;33:676–687.e675. doi: 10.1016/j.cmet.2020.12.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guo X., Zhou Q., Jin J., Lan F., Wen C., Li J., Yang N., Sun C. Hepatic steatosis is associated with dysregulated cholesterol metabolism and altered protein acetylation dynamics in chickens. J. Anim. Sci. Biotechnol. 2023;14:108. doi: 10.1186/s40104-023-00910-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gupta G.L., Sharma L. Bacopa monnieri abrogates alcohol abstinence-induced anxiety-like behavior by regulating biochemical and Gabra1, Gabra4, Gabra5 gene expression of GABA(A) receptor signaling pathway in rats. Biomed. Pharmacother. 2019;111:1417–1428. doi: 10.1016/j.biopha.2019.01.048. [DOI] [PubMed] [Google Scholar]
- Herd R.M., Arthur P.F. Physiological basis for residual feed intake. J. Anim. Sci. 2009;87:E64–E71. doi: 10.2527/jas.2008-1345. [DOI] [PubMed] [Google Scholar]
- Hofmaenner D.A., Kleyman A., Press A., Bauer M., Singer M. The many roles of cholesterol in sepsis: a review. Am. J. Respir Crit. Care Med. 2022;205:388–396. doi: 10.1164/rccm.202105-1197TR. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jin S., Xu Y., Zang H., Yang L., Lin Z., Li Y., Geng Z. Expression of genes related to lipid transport in meat-type ducks divergent for low or high residual feed intake. Asian-Austral. J. Anim. Sci. 2020;33:416–423. doi: 10.5713/ajas.19.0284. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Johswich A., Longuet C., Pawling J., Abdel Rahman A., Ryczko M., Drucker D.J., Dennis J.W. N-glycan remodeling on glucagon receptor is an effector of nutrient sensing by the hexosamine biosynthesis pathway. J. Biol. Chem. 2014;289:15927–15941. doi: 10.1074/jbc.M114.563734. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kamada Y., Ebisutani Y., Kida S., Mizutani K., Akita M., Yamamoto A., Fujii H., Sobajima T., Terao N., Takamatsu S., Yoshida Y., Takehara T., Miyoshi E. Ectopic expression of N-acetylglucosaminyltransferase V accelerates hepatic triglyceride synthesis. Hepatol. Res. 2016;46:E118–E129. doi: 10.1111/hepr.12541. [DOI] [PubMed] [Google Scholar]
- Karimi P., Bakhtiarizadeh M.R., Salehi A., Izadnia H.R. Transcriptome analysis reveals the potential roles of long non-coding RNAs in feed efficiency of chicken. Sci. Rep. 2022;12:2558. doi: 10.1038/s41598-022-06528-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim D., Langmead B., Salzberg S.L. HISAT: a fast spliced aligner with low memory requirements. Nat. Method. 2015;12:357–360. doi: 10.1038/nmeth.3317. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Koch R.M., Swiger L.A., Chambers D., Gregory K.E. Efficiency of feed use in beef cattle. J. Anim. Sci. 1963;22:486–494. [Google Scholar]
- Kumar S., Nei M., Dudley J., Tamura K. MEGA: a biologist-centric software for evolutionary analysis of DNA and protein sequences. Brief. Bioinform. 2008;9:299–306. doi: 10.1093/bib/bbn017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li H., Durbin R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics. 2009;25:1754–1760. doi: 10.1093/bioinformatics/btp324. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li H., Handsaker B., Wysoker A., Fennell T., Ruan J., Homer N., Marth G., Abecasis G., Durbin R., S. Genome Project Data Processing The sequence alignment/map format and SAMtools. Bioinformatics. 2009;25:2078–2079. doi: 10.1093/bioinformatics/btp352. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li W., Zheng M., Zhao G., Wang J., Liu J., Wang S., Feng F., Liu D., Zhu D., Li Q., Guo L., Guo Y., Liu R., Wen J. Identification of QTL regions and candidate genes for growth and feed efficiency in broilers. Genet. Sel. Evol. 2021;53:13. doi: 10.1186/s12711-021-00608-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Love M.I., Huber W., Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genom. Biol. 2014;15:550. doi: 10.1186/s13059-014-0550-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Luo J., Yang H., Song B.L. Mechanisms and regulation of cholesterol homeostasis. Nat. Rev. Mol. Cell. Biol. 2020;21:225–245. doi: 10.1038/s41580-019-0190-7. [DOI] [PubMed] [Google Scholar]
- McKenna A., Hanna M., Banks E., Sivachenko A., Cibulskis K., Kernytsky A., Garimella K., Altshuler D., Gabriel S., Daly M., DePristo M.A. The genome analysis toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genom. Res. 2010;20:1297–1303. doi: 10.1101/gr.107524.110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nafikov R.A., Beitz D.C. Carbohydrate and lipid metabolism in farm animals. J. Nutr. 2007;137:702–705. doi: 10.1093/jn/137.3.702. [DOI] [PubMed] [Google Scholar]
- Nkrumah J.D., Okine E.K., Mathison G.W., Schmid K., Li C., Basarab J.A., Price M.A., Wang Z., Moore S.S. Relationships of feedlot feed efficiency, performance, and feeding behavior with metabolic rate, methane production, and energy partitioning in beef cattle. J. Anim. Sci. 2006;84:145–153. doi: 10.2527/2006.841145x. [DOI] [PubMed] [Google Scholar]
- Pamplona R., Costantini D. Molecular and structural antioxidant defenses against oxidative stress in animals. Am. J. Physiol. Regul. Integr. Comp. Physiol. 2011;301:R843–R863. doi: 10.1152/ajpregu.00034.2011. [DOI] [PubMed] [Google Scholar]
- Pang Z., Chong J., Li S., Xia J. MetaboAnalystR 3.0: toward an optimized workflow for global metabolomics. Metabolites. 2020;10:186. doi: 10.3390/metabo10050186. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Perino A., Demagny H., Velazquez-Villegas L., Schoonjans K. Molecular physiology of bile acid signaling in health, disease, and aging. Physiol. Rev. 2021;101:683–731. doi: 10.1152/physrev.00049.2019. [DOI] [PubMed] [Google Scholar]
- Purcell S., Neale B., Todd-Brown K., Thomas L., Ferreira M.A., Bender D., Maller J., Sklar P., de Bakker P.I., Daly M.J., Sham P.C. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 2007;81:559–575. doi: 10.1086/519795. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reyer H., Oster M., Magowan E., Dannenberger D., Ponsuksili S., Wimmers K. Strategies towards improved feed efficiency in pigs comprise molecular shifts in hepatic lipid and carbohydrate metabolism. Int. J. Mol. Sci. 2017;18:1674. doi: 10.3390/ijms18081674. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rezen T., Rozman D., Kovacs T., Kovacs P., Sipos A., Bai P., Miko E. The role of bile acids in carcinogenesis. Cell Mol. Life Sci. 2022;79:243. doi: 10.1007/s00018-022-04278-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Russell J.R., Sexten W.J., Kerley M.S., Hansen S.L. Relationship between antioxidant capacity, oxidative stress, and feed efficiency in beef steers. J. Anim. Sci. 2016;94:2942–2953. doi: 10.2527/jas.2016-0271. [DOI] [PubMed] [Google Scholar]
- Sajan S.A., Gradisch R., Vogel F.D., Coffey A.J., Salyakina D., Soler D., Jayakar P., Jayakar A., Bianconi S.E., Cooper A.H., Liu S., William N., Benkel-Herrenbruck I., Maiwald R., Heller C., Biskup S., Leiz S., Westphal D.S., Wagner M., Clarke A., Stockner T., Ernst M., Kesari A., Krenn M. De novo variants in GABRA4 are associated with a neurological phenotype including developmental delay, behavioral abnormalities and epilepsy. Eur. J. Hum. Genet. 2024;32:912–919. doi: 10.1038/s41431-024-01600-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schulze R.J., Drizyte K., Casey C.A., McNiven M.A. Hepatic lipophagy: new insights into autophagic catabolism of lipid droplets in the liver. Hepatol. Commun. 2017;1:359–369. doi: 10.1002/hep4.1056. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Silva V.R., Micheletti T.O., Pimentel G.D., Katashima C.K., Lenhare L., Morari J., Mendes M.C., Razolli D.S., Rocha G.Z., de Souza C.T., Ryu D., Prada P.O., Velloso L.A., Carvalheira J.B., Pauli J.R., Cintra D.E., Ropelle E.R. Hypothalamic S1P/S1PR1 axis controls energy homeostasis. Nat. Commun. 2014;5:4859. doi: 10.1038/ncomms5859. [DOI] [PubMed] [Google Scholar]
- Steinberg C., Roston T.M., van der Werf C., Sanatani S., Chen S.R.W., Wilde A.A.M., Krahn A.D. RYR2-ryanodinopathies: from calcium overload to calcium deficiency. Europace. 2023;25:euad156. doi: 10.1093/europace/euad156. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sun H., Xu W., Gu T., Sun J., Li C., Chen L., Tian Y., Li G., Lu L., Zeng T. Association of residual feed intake with intestinal microbiome and metabolome in laying period of ducks. Front. Microbiol. 2023;14 doi: 10.3389/fmicb.2023.1138914. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vozella V., Realini N., Misto A., Piomelli D. Feeding stimulates sphingosine-1-phosphate mobilization in mouse hypothalamus. Int. J. Mol. Sci. 2019;20:4008. doi: 10.3390/ijms20164008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wen C., Yan W., Mai C., Duan Z., Zheng J., Sun C., Yang N. Joint contributions of the gut microbiota and host genetics to feed efficiency in chickens. Microbiome. 2021;9:126. doi: 10.1186/s40168-021-01040-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Willems O.W., Miller S.P., Wood B.J. Aspects of selection for feed efficiency in meat producing poultry. World. Poult. Sci. J. 2019;69:77–88. [Google Scholar]
- Xie C., Liang Q., Cheng J., Yuan Y., Xie L., Ji J. Transplantation of fecal microbiota from low to high residual feed intake chickens: impacts on RFI, microbial community and metabolites profiles. Poult. Sci. 2024;104 doi: 10.1016/j.psj.2024.104567. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang J., Lee S.H., Goddard M.E., Visscher P.M. GCTA: a tool for genome-wide complex trait analysis. Am. J. Hum. Genet. 2011;88:76–82. doi: 10.1016/j.ajhg.2010.11.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang L., He T., Xiong F., Chen X., Fan X., Jin S., Geng Z. Identification of key genes and pathways associated with feed efficiency of native chickens based on transcriptome data via bioinformatics analysis. BMC Genom. 2020;21:292. doi: 10.1186/s12864-020-6713-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yi Z., Li X., Luo W., Xu Z., Ji C., Zhang Y., Nie Q., Zhang D., Zhang X. Feed conversion ratio, residual feed intake and cholecystokinin type A receptor gene polymorphisms are associated with feed intake and average daily gain in a Chinese local chicken population. J. Anim. Sci. Biotechnol. 2018;9:50. doi: 10.1186/s40104-018-0261-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang X., Li G., Li F., Zhang D., Yuan L., Zhao Y., Zhang Y., Li X., Song Q., Wang W. Effect of feed efficiency on growth performance, body composition, and fat deposition in growing Hu lambs. Anim. Biotechnol. 2023;34:183–198. doi: 10.1080/10495398.2021.1951747. [DOI] [PubMed] [Google Scholar]
- Zou X., Liu T., Li Y., Chen P., Yan X., Ma J., Ji J., Qu H., Cai M., He Y., He J., Xu X., Lin C., Zhou G., Shu D., Luo C. Long-term divergent selection for residual feed intake in Chinese broiler chickens. Poult. Sci. 2023;102 doi: 10.1016/j.psj.2022.102298. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
All of the sequencing data in this study have been deposited into NCBI and can be accessed via accession PRJNA1149694.