Abstract
Proliferative hepatocellular carcinoma (HCC) is an aggressive phenotype associated with unfavorable clinical outcomes. Predicting the preoperative subtype of HCC can aid in the development of individualized treatment. We retrospectively recruited 180 HCC patients who underwent hepatic resection and established a CT-based radiomics model for predicting proliferative HCCs. The evaluation of tumor response to transarterial chemoembolization therapy and progression-free survival (PFS) according to the radiomics model was further performed in internal (n = 54) and external (n = 80) outcome cohorts. In our study, 98 of 180 (54%) patients were confirmed to have proliferative HCCs. The radiomics model comprising 9 radiomic features and exhibited good performance for predicting proliferative HCCs. The nomogram integrated radiomics and serum α-fetoprotein level showed good calibration and discrimination in both the training cohort (AUC = 0.848) and the validation cohort (AUC = 0.825). Predicted proliferative HCCs (high radiomics scores) were associated with lower response rate (P < 0.05) and worse PFS (P < 0.05) compared to predicted non-proliferative HCCs in outcomes cohorts. We linked radiomics model to gene expression, unveiling that activated/immature B cells and tertiary lymphoid structures were downregulated in the high radiomics group. The proposed CT radiomics model exhibited good performance for identifying proliferative HCCs, which may facilitate clinical decision-making. Our findings suggest a potential correlation between proliferative HCC and immunosuppressive tumor microenvironment.
Supplementary Information
The online version contains supplementary material available at 10.1038/s41598-025-94684-w.
Keywords: Hepatocellular carcinoma, Proliferative subtype, Radiomics, Transarterial chemoembolization, Immune infiltration
Introduction
Hepatocellular carcinoma (HCC) is one of the most common cause of cancer-related death worldwide, characterized by highly molecular and histological heterogeneity1. The poor outcome associated with HCC, partly due to the limited therapeutic strategies based on molecular or histologic classification for HCC2,3. Thus, identifying patient-specific molecular, imaging, and pathological prognostic features that can aid in individualized treatment decision-making is needed4.
A recently proposed integrated classification has categorized HCC into proliferative and non-proliferative subtypes3. Proliferative HCC associated with chromosomal instability and unfavorable clinical outcomes and includes macrotrabecular-massive, scirrhous, sarcomatoid, neutrophilrich and cytokeratin 19 (CK19)–positive HCC3,5,6. The diagnosis of proliferative HCC conventionally requires access to biopsy samples. However, most patients present with advanced disease not eligible for resection. For patients with intermediate-stage HCC, conventional transarterial chemoembolization (cTACE) remains the standard therapy7. Moreover, small biopsy may subject to sampling bias due to intratumour spatial heterogeneity8. Bao et al. found that proliferative HCCs have worse overall survival and progression-free survival than non-proliferative HCCs5. Therefore, noninvasive manner, such as imaging procedure, for the prediction of histological subtype before treatment is crucial.
Previous studies have demonstrated some distinctive computer tomography (CT) and magnetic resonance imaging (MRI) features in proliferative HCCs5,6,9,10. Even so, the bias of interobserver among radiologist may lead to inconsistent evaluation of these imaging features. Radiomics can automatically obtain innumerable quantitative features from digitally encrypted medical images11,12. Notably, CT or MRI based radiomics model have demonstrated promising performance in prediction of macrotrabecular-massive subtype, which belong to the proliferative class of HCC12,13. Studies on radiogenomics in the context of HCC have also concentrated on discovering imaging-based signatures linked to genetic changes and molecular pathways12. However, studies on the CT based-radiomics approach for the prediction of proliferative HCC are still lacking in the literature.
In this study, we aim to construct a radiomics model to predict proliferative HCC by utilizing enhanced CT radiomic features and compare therapeutic outcomes after cTACE between predicted proliferative and non-proliferative HCCs. The underlying pathway and immune infiltration patterns were also investigate using public database.
Materials and methods
Study design
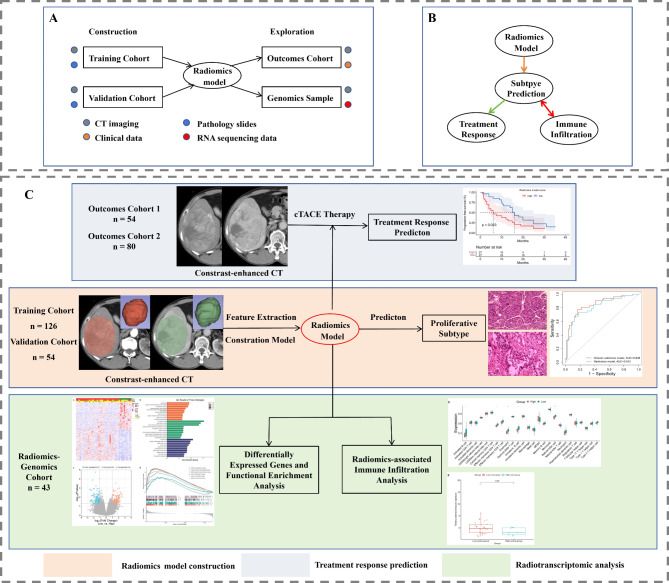
This study include three cohorts, radiomics-construction cohort and outcomes cohort were enrolled from the The Third Xiangya Hospital Central South University and Hunan Cancer Hospital, the radiomics-genomics cohort was collected from The Cancer Imaging Archive (TCIA) database. The overall study design is shown in Fig. 1. The institutional review board of The Third Xiangya Hospital of Central South University and Hunan Cancer Hospital approved this study and waived the requirement for written informed consent owing to the retrospective study design. All methods were performed in accordance with the relevant guidelines and regulations.
Fig. 1.
Overview of the study. (A) Schematic diagram of the four datasets or cohorts. (B) Overall design circuit of the study. (C) Flowchart of the study in three steps. cTACE, conventional transarterial chemoembolization.
First, we constructed radiomics model by using retrospective radiologic and electronic medical data of patients who underwent hepatic resection for HCC from January 2016 to May 2021 at Third Xiangya Hospital (Fig. 2A). The inclusion criteria were as follow: (a) undergoing diagnostically acceptable contrast-enhanced liver CT within 1 month before surgery; (b) available pathology slides and histologically confirmed HCC. The exclusion criteria were as follows: (a) receiving intervention therapy, such as systemic chemotherapy, before surgery; (b) suboptimal image quality of CT or pathologic images. Clinical data, including age, sex, hepatitis virus infection, presence or absence of liver cirrhosis, serum α-fetoprotein (AFP) levels, and Barcelona Clinic Liver Cancer (or BCLC) stage were retrospectively collected. Second, we enrolled consecutive adult patients with unresectable HCC who undergoing conventional cTACE as the primary therapy between March 2018 and August 2021 at Third Xiangya Hospital (outcomes cohort 1) and between January 2021 and October 2023 at Hunan Cancer Hospital (outcomes cohort 2) (Fig. 2B). The inclusion criteria were as follow: (a) undergoing diagnostically acceptable contrast-enhanced liver CT within 1 month before cTACE; (b) at least one measurable target lesion to evaluate tumor response; (c) at least 2 cTACE sessions with 6 week-interval. The exclusion criteria were as follows: (a) receiving intervention therapy, such as surgery, systemic chemotherapy, local ablation before cTACE; (b) absence of imaging follow-up data; (c) suboptimal image quality of CT. Clinical data, including age, sex, serum AFP levels, tumor burden (i.e, six-and-twelve score)14 and cTACE sessions were retrospectively collected. Finally, HCC patients from TCIA databases with available baseline contrast-enhanced CT (both arterial and portal venous phases) and transcriptome sequencing data were constituted radiomics-genomics cohort (Fig. 2C). The detailed patient recruitment process is shown in Fig. 2.
Fig. 2.
Organization flowchart of three cohorts. (A) Radiomics-construction cohort. (B) Outcomes cohorts. (C) Radiomics-genomics cohort. HCC, hepatocellular carcinoma; cTACE, conventional transarterial chemoembolization; TCIA, The Cancer Imaging Archive; TCGA, The Cancer Genome Atlas.
An experienced pathologist reviewed all resected specimens and recorded the information of size, presence or absence of microvascular invasion, and the histologic subtype of the tumor. The histomorphologic subtype of HCC was classified based on the 2019 WHO classification15. Additionally, immunohistochemistry for CK19 was conducted, and HCCs expressed CK19 in greater than 5% of tumor cells were defined as CK19-positive HCCs. In summary, macrotrabecular-massive, neutrophil-rich, scirrhous, sarcomatoid, and CK19-positive HCCs were designated proliferative HCCs, whereas steatohepatitic, lymphocyte-rich, clear cell, and CK19-negative HCCs were categorized as non-proliferative HCCs.
Tumor response assessment and follow-up
For outcomes cohorts, radiologic follow-up consisted of dynamic CT or MRI every 1–2 months within 6 months after cTACE treatment and every 3–6 months after 6 months. Treatment response was assessed 1–2 months after the second cTACE treatment according to modified Response Evaluation Criteria (mRECIST) in Solid Tumors16. The primary outcome was progression-free survival (PFS), defined as the interval between first cTACE treatment and disease progression or death. The last follow-up was the last visit to the hospital.
CT examination and radiomic feature extraction
All patients underwent contrast-enhanced liver CT within 2 weeks before surgery or TACE treatment. CT was performed in the axial plane with 5-mm-thick sections using a 256-section (GE Revolution; GE Healthcare) or 64-section (Philips Brilliance; Philips) multidetector CT scanner. The detailed parameters of the CT examination are shown in Appendix S1.
The segmentation of the tumors was performed by two radiologists by using 3D Slicer software (version 4.11.2; Boston, MA, USA). Volumes of interest were manually delineated defining the tumor contour in a slice- by- slice manner on arterial phase and portal venous phase images, as shown in Fig. S1. A senior radiologist confirmed all segmentations and resolved the disagreements with consensus. Radiomics features extraction and image preprocessing were performed with the PyRadiomics software (https://github.com/Radiomics/pyradiomics). All images were resampled to a resolution of 1 × 1 × 1 mm by linear interpolation. In total, we retrieved 2632 features for each patient: 1316 features each from the arterial phase image and portal venous phase image. The features were then normalized using the Z-score method. Feature extraction algorithms are summarized in Appendix S1.
Radiomics model construction
The two radiologists repeated the segmentation in 30 randomly selected patients after 2 weeks and calculate intraclass correlation coefficient (ICC), evaluating intra- and interobserver reproducibility. Only features with inter- and intraobserver ICCs > 0.75 indicated satisfactory repeatability and were selected for further analysis. Additionally, some redundant features were remove based on the Pearson correlation coefficient.
The least absolute shrinkage and selection operator (LASSO) regression algorithm can prevent model overfitting and was used to select the best features to determine the radiomics model score using 10-fold cross-validation. The clinical-radiomics model of the proliferative HCCs subtype was constructed by uni- and multivariable logistic regression analysis in the training cohort. Using receiver operating characteristic curve and calibration curve to test and evaluate model performance. Decision curve analysis was used to evaluate the clinical value. The Delong test was used to evaluate the difference in area under the receiver operating characteristic curve (AUC) between the two models.
Gene ontology (GO) analysis and gene set enrichment analysis (GSEA)
Patients in the radiomics-genomics cohort were stratified into high- or low-radiomics score groups according to the optimal cutoff value of the constructed model. Differentially expressed genes (DEGs) between groups were identified according to the criteria of a |log2 (fold change)|> 2 and false-discovery rate (FDR) < 0.05. Our FDR analysis followed the Benjamini–Hochberg method. GO analysis was carried out to using clusterProfiler R package to explore biological functions and molecular functions. GSEA analysis was conducted using fgsea package to determine the associated pathways. Single-sample GSEA was used to analyze the immune infiltration characteristics.
Statistical analysis
In this study, statistical analysis and graphic production were performed with R software (Version 4.2.3 https://www.Rproject.org). Normality and variance homogeneity of clinical data from the two groups were assessed by the Shapiro–Wilk test and Bartlett test, respectively. The chi-square test or Fisher's exact test for categorical variables and T test or Wilcoxon rank-sum test for continuous variables. For outcomes cohort, patients were divided into high- or low- radiomics score groups based on the optimal cutoff value of the radiomics model. The optimal cutoff value (0.015) was determined by the surv_cutpoint function from the ‘survminer’ R package as previous described17. Kaplan–Meier survival curves and log-rank tests were conducted using the "survival" and "survminer" R packages to illustrate the survival differences between the two groups. Uni- and multivariable Cox regression (stepwise model) analyses were performed to identify the identify potential independent prognostic factors of PFS.
Results
Patient clinical characteristics
A total of 180 HCC patients (158 men, 22 women) were included in radiomics-construction cohort, with a mean age of 55 ± 10 years (standard deviation). The patients in radiomics-construction cohort were randomly divided into training cohort (n = 126) and validation cohort (n = 54) at a ratio of 7:3. The clinical and pathologic characteristics of patients are summarized in Table 1. No statistical difference was found in clinical factors between the training and validation cohort (all P > 0.05). In the training cohort, 53 of 126 patients (42%) were categorized as proliferative HCC. Proliferative HCCs were more frequently correlated with larger tumor size (median, [interquartile range]) (6.2mm [4.8–8.0mm] vs 5.0mm [3.0–7.5mm]; P = 0.011), AFP level higher than 200 ng/mL (30 of 53[57%] vs 6 of 73 [8%], P < 0.001) and positive MVI status (32 of 53[60%] vs 21 of 73 [29%], P < 0.001) when comparing with those nonproliferative HCC. Detail characteristics in the training cohort based on the histologic HCC subtype are presented in Table S1.
Table 1.
Clinical and pathologic characteristics of patients for radiomics model construction.
| Characteristic | Training set (n = 126) | Validation set (n = 54) | P value |
|---|---|---|---|
| Age (y)* | 55 ± 10 | 54 ± 9 | 0.696 |
| Male sex (%) | 110(87) | 48(89) | 0.766 |
| HBV or HCV infection (%) | 111(88) | 45(85) | 0.560 |
| Liver cirrhosis (%) | 69(55) | 27(50) | 0.557 |
| Serum AFP level > 200 ng/mL (%) | 36 (28) | 12 (22) | 0.377 |
| Tumor size (cm)† | 5.50 (3.5–8.0) | 5.55 (4.0–9.0) | 0.347 |
| BCLC stage (%) | 0.474 | ||
| 0 or A | 68(54) | 26(48) | |
| B or C | 58(46) | 28(52) | |
| Edmonson-Steiner grade (%) | 0.308 | ||
| I–II | 69 (55) | 34 (63) | |
| III–IV | 57 (45) | 20 (37) | |
| MVI (YES) (%) | 53 (42) | 28 (52) | 0.226 |
| Proliferative HCCs (%) | 0.151 | ||
| Absent | 73(58) | 25(46) | |
| Present | 53(42) | 29(54) | |
Unless indicated otherwise, data are number of patients, and data in parentheses are percentages. AFP, α-fetoprotein; BCLC, Barcelona Clinic Liver Cancer; HBV, hepatitis B virus; HCV, hepatitis C virus; HCC, hepatocellular carcinoma.
*Data are means ± standard deviation.
†Data are medians, with interquartile ranges in parentheses.
Construction of the radiomics model
Of the 2036 extracted radiomics features with high stability (inter- and intraobserver ICCs > 0.75), 343 features were selected after redundancy analysis. In training cohorts, 9 features with non-zero coefficients in the LASSO logistic regression model were identified (Fig. 3A, B). A radiomics model for predicting proliferative HCC was constructed and the calculation formula is shown in Appendix S1. The distribution of the radiomics score in the training and validation cohorts is showed in Fig. 3C, D. The radiomics model score was higher for the proliferative HCC group than for the nonproliferative HCC group in the training cohort (P < 0.001) and validation cohort (P < 0.001).
Fig. 3.
Construction and evaluation of the radiomics models for predicting proliferative hepatocellular carcinoma (HCC) subtype. (A, B) Using the least absolute shrinkage and selection operator (LASSO) regression, radiomic feature selection is accomplished. (C, D) Plots showing the distribution of radiomics model scores in training set (C) and validation set (D). (E) Forest plot of the multivariable analysis for proliferative HCC in the training set. (F) The constructed clinical-radiomics nomogram for proliferative HCC. (G, H) The receiver operating characteristic curve of the radiomics model and the clinical-radiomics model in the training set (G) and validation set (H). AFP, α-fetoprotein; HBV, hepatitis B virus; HCV, hepatitis C virus; BCLC, Barcelona Clinic Liver Cancer; OR, odds ratio.
In the multivariable logistic regression analysis, radiomics model (odds ratio [OR], 9.15 [95% CI 2.52, 33.24]; P < 0.001) and AFP level higher than 200 ng/mL (OR, 6.72 [95% CI 2.24, 20.15]; P < 0.001) were identified as independent predictors of proliferative HCCs (Fig. 3E). The radiomics model yielded an AUC of 0.81 in the training cohort and 0.80 in the validation cohort, indicating favorable predictive efficacy for proliferative HCCs (Fig. 3G, H). A clinical-radiomics model that integrated the clinical variable (AFP) and radiomics was developed (Fig. 3F) and the calculation formula is as follows: Y = − 1.092 + 1.904 × high serum AFP level + 2.214 × radiomics model score. The predictive performance of clinical-radiomics model improved compared to radiomics model, with AUCs of 0.85 in the training cohort (P = 0.10) and 0.83 in the validation cohort (P = 0.22) (Fig. 3G, H, Table 2), though statistical significance was not reached. The calibration curves showed that the clinical-radiomics model had good calibration in both the training and validation cohort (Fig. S2a–S2b). The decision curve (Fig. S2c–S2d) indicates that clinical-radiomics model has a better clinical net benefit than radiomics model.
Table 2.
Performance of the radiomics models for predicting the proliferative HCC subtype.
| Data set | AUC | Sensitivity (%) | Specificity (%) | PPV (%) | NPV (%) | P Value* |
|---|---|---|---|---|---|---|
| Radiomics model | ||||||
| Training set | 0.81 (0.73, 0.89) |
73.6 (62.7, 85.5) [39/53] |
82.2 (73.4, 91.0) [60/73] |
75.0 (63.2, 86.8) [39/52] |
81.1 (72.2, 90.0) [60/74] |
– |
| Validation set | 0.78 (0.66, 0.90) |
62.1 (44.4, 79.7) [18/29] |
84.0 (69.6, 98.4) [21/25] |
81.8 (65.7, 97.9) [18/22] |
65.5 (49.2, 82.1) [21/32] |
– |
| Clinical-radiomics model | ||||||
| Training set | 0.85 (0.78, 0.92) |
77.4 (66.1, 88.6) [41/53] |
83.6 (81.4, 92.1) [61/73] |
77.4 (66.1, 88.8) [41/53] |
83.6 (75.1, 92.1) [61/73] |
0.10 |
| Validation set | 0.83 (0.71, 0.94) |
65.5 (48.2, 82.8) [19/29] |
92.0 (81.4, 100.0) [23/25] |
90.5 (77.9, 103.0) [19/21] |
69.7 (54.0, 85.4) [23/33] |
0.22 |
Data in parentheses are 95% CIs, and data in brackets are numbers of patients. AUC, area under the receiver operating characteristics curve; HCC, hepatocellular carcinoma; NPV, negative predictive value; PPV, positive predictive value. The sensitivity, specificity, PPV, and NPV of the radiomics model and clinical-radiomics model in the two data sets were calculated with the cutoff values of 0.015 or lower and − 0.883, respectively, which maximize the Youden index in the training set.
*P value indicates the significance level of the comparison of AUCs with the radiomics model as the reference in the corresponding data set.
Predictive value of the radiomics model for tumor response and survival
A total of 54 patients from outcomes cohort 1 and 80 patients from outcomes cohort 2 were used to evaluate prognostic value of the radiomics model. The baseline characteristics of outcomes cohorts are listed in Table S2-3. The median follow-up period was 11 months (range, 2–35 months) for outcomes cohort 1 and was 10 months (range, 1.5–45 months) for outcomes cohort 2. In outcomes cohort 1, 54% (29 of 54) patients were cTACE responders, and 46% (25 of 54) patients were non-responders. In outcomes cohort 2, 53% (42 of 80) patients were cTACE responders, and 47% (38 of 80) patients were non-responders. Comparing with non-responders, responders had lower radiomics score (outcomes cohort 1: P < 0.005; outcomes cohort 2: P = 0.013), indicating that the radiomics score could efficiently distinguish between two groups (Fig. 4A, C). Based on the radiomics score, patients were categorized as having predicted proliferative (high-score) and non-proliferative HCCs (low-score), respectively. Patients with predicted proliferative HCC showed a worse PFS than patients with predicted non-proliferative HCCs (outcomes cohort 1: P = 0.023; outcomes cohort 2: P = 0.043) (Fig. 4B, D). Multivariable Cox Regression analysis revealed that high six-and-twelve score (outcomes cohort 1: hazard ratio [HR], 1.9, 95% CI 1.1–3.6, P = 0.034; outcomes cohort 2: HR 1.9, 95% CI 1.1–3.1, P = 0.016) and predicted proliferative HCC (outcomes cohort 1: HR 2.0, 95% CI 1.1–3.6, P = 0.031; outcomes cohort 2: HR 1.7, 95% CI 1.0–2.8; P = 0.036) were independent prognostic predictors for PFS after cTACE treatment (Table 3).
Fig. 4.
Ability of radiomics models in predicting tumor response in the outcomes cohorts. Association between radiomics model score and tumor response to conventional transarterial chemoembolization (cTACE) treatment in outcomes cohort 1 (A) and cohort 2 (C). Survival curves of progression-free survival based on radiomics model score (low or high defined by cutoff value) in outcomes cohort 1 (B) and cohort 2 (D).
Table 3.
Uni- and multivariable cox regression analyses for PFS in the outcomes cohorts.
| Variables | Univariable HR | P Value | Multivariable HR | P Value |
|---|---|---|---|---|
| Outcomes cohort 1 | ||||
| Age (> 50 y vs ≤ 50 y) | 1.0 (0.6–1.9) | 0.930 | – | – |
| Men | 0.58 (0.2–1.4) | 0.217 | – | – |
| Serum AFP level > 200 ng/mL | 1.24 (0.7–2.3) | 0.507 | – | – |
| Six-and-twelve score (high vs low) intermediate)* | 1.95 (1.1–3.6) | 0.032 | 1.93 (1.1–3.6) | 0.034 |
| cTACE sessions (≥ 3 vs 2) | 1.74 (0.9–3.3) | 0.084 | – | – |
| Radiomics model score (high vs low) | 1.98 (1.1–3.7) | 0.029 | 1.96 (1.1- 3.6) | 0.031 |
| Outcomes cohort 2 | ||||
| Age (> 50 y vs ≤ 50 y) | 1.30 (0.7–2.2) | 0.336 | – | – |
| Men | 1.00 (0.5–1.8) | 0.988 | – | – |
| Serum AFP level > 200 ng/mL | 1.41 (0.8–2.3) | 0.199 | – | – |
| Six-and-twelve score (high vs low) intermediate)* | 1.80 (1.1–2.9) | 0.023 | 1.87 (1.1–3.1) | 0.016 |
| cTACE sessions (≥ 3 vs 2) | 1.12 (0.6–1.8) | 0.660 | – | – |
| Radiomics model score (high vs low) | 1.62 (1.0–2.7) | 0.056 | 1.70 (1.0–2.8) | 0.036 |
Data in parentheses are 95% CIs. AFP, α-fetoprotein; HR, hazard ratio; PFS, progression-free survival; cTACE, conventional transarterial chemoembolization.
*The six-and-twelve score is defined as the summation of the largest tumor diameter and the numbers of tumors and included three strata according to the summation of scores of 6 or less (low), more than 6 to 12 (intermediate), and greater than 12 (high).
Immune infiltration patterns associated with the radiomics model
A total of 45 patients were included for transcription analysis. The detail characteristics in the radiomics-genomics cohort are shown in Table S4. We identified 283 DEGs between the low- and high-radiomics score groups (Fig. 5A and Fig. S3, Table S5). The GO analysis revealed that these DEGs were associated with immune-related molecular function or biological processes, such as immune response, immunoglobulin production and circulating, immunoglobulin receptor binding, antigen binding, and cytokine receptor activity (Fig. 5B). The GSEA analysis showed that the immunoglobulin production, immunoglobulin complex, immune receptor activity, T cell receptor complex, and natural killer cell mediated immunity-related pathways were significantly downregulation in the high-score group (Fig. 5C). The ssGSEA analysis showed the activated B cell, immature B cell, effector memeory CD4 T cell, natural killer (NK) cell and eosinophil levels were lower in the high-score group (Fig. 5D). Moreover, 39-gene tertiary lymphoid structure (TLS) signature levels18 were also lower in the high-score group (Fig. 5E). Kaplan–Meier curve analysis revealed that patients with high-score had a shorter survival time than those with low-score (Fig. 5F), indicating that the predicted non-proliferating HCC associated with higher degree of immune infiltration and a better prognosis.
Fig. 5.
Exploration of radiomics model molecular patterns and immune infiltration statuses. (A) Heatmap of the expression patterns of 284 differentially expressed genes (DEGs) between low and high radiomics model score groups in the genomics sample. (B) Gene Ontology (GO) analysis of the DEGs. (C) The six representative GO pathways enriched by GSEA between the low radiomics model score group and high radiomics model score group. (D, E) Immune cell infiltration levels (D) and tertiary lymphoid structure signature levels (E) between the low radiomics model score group and high radiomics model score group. (F) Survival curves of overall survival based on radiomics model score.
Discussion
Proliferative HCC are more frequent poorly differentiated and associated with adverse clinical outcome3,5. Therefore, the noninvasive identification of proliferative HCC is essential for the treatment planning and accurate prognostication. In this study, we developed a CT-based radiomics model to predict proliferative HCC for the first time, with AUC of 0.81 in the training cohort and 0.78 in the validation cohort. Specifically, clinical-radiomics model provided better predictive ability, with AUC of 0.85 and 0.83 in the training and validation cohort, respectively. Our study demonstrated that a high radiomics score, likely indicative of proliferative HCC, was correlated with lower tumor response rate and worse PFS among patients who underwent cTACE. Of note, bulk RNA sequencing uncovered that predicted proliferating HCC were associated with low degree of immune infiltration and worse prognosis.
Previous studies have highlighted the distinct imaging features, such as arterial phase hyperenhancement, tumor shape, and mosaic architecture, with potential implications for predicting proliferative HCC5,6. Radiomics contains high-dimensional imaging features with additional information and may indicate the biologic features of tumors19,20. Radiomics have been successfully used to assesses microvascular invasion21, distinguish histological subtype12,22, improve diagnostic accuracy23, evaluate tumor response to TACE therapy or chemotherapy24, and preoperative predict prognosis20,25 in HCC. Our radiomics model achieved comparable predictive performance when compared to the CT imaging features model in predicting proliferative HCC6. Meanwhile, radiomics may reduce the bias of the radiologist, suggesting its better generalizability and clinical application.
We demonstrated that proliferative HCC was associated with larger tumor size and higher serum AFP level than non-proliferative HCC, consistent with previous study6. Moreover, AFP level higher than 200 ng/mL is an independent predictor of the proliferative subtype and could improve the predictive performance of radiomics model. The evaluation of tumor response to the cTACE is essential because not all intermediate-stage HCC patients benefit from this therapy. Our study provided evidence of correlations between worse prognosis and predicted proliferating HCC compared to predicted non-proliferating HCC after cTACE. The radiomics model provides a non-invasive tool with significant potential for clinical application. For example, patients identified as having proliferative HCC might need undergo a biopsy before cTACE to confirm the subtype. Therapeutic intensification through combination regimens incorporating tyrosine kinase inhibitors or immune checkpoint inhibitors may optimize clinical outcomes for these patient. Additionally, the model facilitates risk stratification, offering clinicians a basis for more precise prognostic evaluations and tailored follow-up strategies.
Linking radiomics to transcriptional changes unveils biologically significant associations. The upregulation of EPO could increase HCC-associated angiogenesis and decrease antitumor activity, relating to poor response to TACE treatment26–28. PITX2 overexpression promotes HCC metastasis, self-renewal and chemoresistance and associated with poor overall survival time29. High expression of LHB, HHLA2 and HMGA2 promoted the cell proliferation and progress of HCC, resulting in shorter overall survival30–33. The liver is an immunologic organ, previous studies have confirmed that the immune cell composition of HCC tumors is clearly associated with overall prognosis as well as responses to therapy34. The GO and GSEA analysis indicate the dysregulation of immune activation and response are implicated in the pathology of proliferative HCC. We further analyzed the underlying immune infiltration patterns based our radiomics model, and the results revealed that activated/immature B cell, NK cell, and memory CD4 T cell were lower in high-radiomics score group. Activated/immature B cells and B-cell-rich tertiary lymphoid structures are major components of adaptive antitumor immune response, downregulation if which are typically associated with poor clinical outcomes and drive resistance to immunotherapy35,36. NK cells play a critical role in the innate antitumor immune response, the low expression of NK cell associated with immunosuppression and influence the response to immunotherapy in HCC patients37. CD4+ T cells are essential for anti-tumor surveillance, and decreased CD4+ T cells could promote HCC development38. This study observed reduced TLS-related gene expression in predicted proliferative HCC, suggesting impaired TLS formation or function, which may contribute to immune evasion. Radiomic features might correlate with microvascular density or cellular heterogeneity, potentially disrupting TLS development and exacerbating immunosuppression. Taken together, proliferative HCC may contribute to immunosuppressive tumor microenvironment, consistent with previous studies12,34. Our study provided insight into the underlying mechanisms and immune infiltration in proliferative HCC, which might benefit from exploring novel intervention strategies.
This study has some limitations. Its retrospective design introduces the possibility of selection bias, which may compromise the objectivity of the findings, even though strict inclusion and exclusion criteria were employed. Moreover, the study relies on data from two centers, differences in imaging equipment, scanning protocols, and patient demographics across institutions could affect the consistency of radiomic features and the model’s predictive accuracy. Additionally, the relatively small sample size raises concerns about overfitting, especially given the high-dimensional feature of radiomic data. While techniques like LASSO regression were used to reduce this risk, the limited sample size may still undermine the robustness of the results. Validation with larger, multicenter datasets is thus essential to assess the model's real-world performance.
In conclusion, we constructed a CT-based radiomics model to predict proliferative HCC, which showed worse prognosis to cTACE treatment. Of note, dysregulation of immune infiltration was observed based on our radiomics model, revealing the association between the immunosuppressive tumor microenvironment and the predicted proliferative HCC. The radiomics model may serve as a noninvasive tool for clinical utilizing.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Author contributions
Conceptualization: H.H., Q.L and Y.X.; Data curation: H.H. and Q.L.; Formal analysis: H.H.,Z.F. and Y.H; Investigation: H.H. and J.D.; Methodology: H.H. and W.D.; Software: H.H. and Z.W.; Supervision: Y.H. and Y.X.; Visualization: H.H. and Z.F.; Writing – original draft: H.H.; Writing-review & editing: H.H., Q.L., Z.F. J.D., W.D. and Y.X.
Data availability
The data generated for this study are available on request to the corresponding author.
Declarations
Competing interests
The authors declare no competing interests.
Footnotes
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Contributor Information
Qi Liang, Email: csuliangqi10@163.com.
Yongzhi Xie, Email: yzxiexy3@163.com.
References
- 1.Vogel, A., Meyer, T., Sapisochin, G., Salem, R. & Saborowski, A. Hepatocellular carcinoma. Lancet (London, England)400, 1345–1362. 10.1016/s0140-6736(22)01200-4 (2022). [DOI] [PubMed] [Google Scholar]
- 2.Llovet, J. M. et al. Molecular pathogenesis and systemic therapies for hepatocellular carcinoma. Nat. Cancer3, 386–401. 10.1038/s43018-022-00357-2 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Calderaro, J., Ziol, M., Paradis, V. & Zucman-Rossi, J. Molecular and histological correlations in liver cancer. J. Hepatol.71, 616–630. 10.1016/j.jhep.2019.06.001 (2019). [DOI] [PubMed] [Google Scholar]
- 4.Li, Y.-K. et al. Portal venous and hepatic arterial coefficients predict post-hepatectomy overall and recurrence-free survival in patients with hepatocellular carcinoma: A retrospective study. J. Hepatocell Carcinoma11, 1389–1402. 10.2147/JHC.S462168 (2024). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Kang, H. J. et al. Gadoxetate-enhanced MRI features of proliferative hepatocellular carcinoma are prognostic after surgery. Radiology300, 572–582. 10.1148/radiol.2021204352 (2021). [DOI] [PubMed] [Google Scholar]
- 6.Bao, Y. et al. Identifying proliferative hepatocellular carcinoma at pretreatment CT: Implications for therapeutic outcomes after transarterial chemoembolization. Radiology308, e230457. 10.1148/radiol.230457 (2023). [DOI] [PubMed] [Google Scholar]
- 7.Raoul, J. L. et al. Updated use of TACE for hepatocellular carcinoma treatment: How and when to use it based on clinical evidence. Cancer Treat. Rev.72, 28–36. 10.1016/j.ctrv.2018.11.002 (2019). [DOI] [PubMed] [Google Scholar]
- 8.Brown, Z. J. et al. Management of hepatocellular carcinoma: A review. JAMA Surg.158, 410–420. 10.1001/jamasurg.2022.7989 (2023). [DOI] [PubMed] [Google Scholar]
- 9.Heo, S. et al. Proliferative hepatocellular carcinomas in cirrhosis: Patient outcomes of LI-RADS category 4/5 and category M. Eur. Radiol.10.1007/s00330-023-10305-y (2023). [DOI] [PubMed] [Google Scholar]
- 10.Feng, Z. et al. Preoperative CT for characterization of aggressive macrotrabecular-massive subtype and vessels that encapsulate tumor clusters pattern in hepatocellular carcinoma. Radiology300, 219–229. 10.1148/radiol.2021203614 (2021). [DOI] [PubMed] [Google Scholar]
- 11.Gillies, R. J., Kinahan, P. E. & Hricak, H. Radiomics: Images are more than pictures, they are data. Radiology278, 563–577. 10.1148/radiol.2015151169 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Feng, Z. et al. CT radiomics to predict macrotrabecular-massive subtype and immune status in hepatocellular carcinoma. Radiology307, e221291. 10.1148/radiol.221291 (2023). [DOI] [PubMed] [Google Scholar]
- 13.Zhu, Y. et al. A radiomics nomogram based on contrast-enhanced MRI for preoperative prediction of macrotrabecular-massive hepatocellular carcinoma. Abdominal Radiol. (New York)46, 3139–3148. 10.1007/s00261-021-02989-x (2021). [DOI] [PubMed] [Google Scholar]
- 14.Wang, Q. et al. Development of a prognostic score for recommended TACE candidates with hepatocellular carcinoma: A multicentre observational study. J. Hepatol.70, 893–903. 10.1016/j.jhep.2019.01.013 (2019). [DOI] [PubMed] [Google Scholar]
- 15.Nagtegaal, I. D. et al. The 2019 WHO classification of tumours of the digestive system. Histopathology76, 182–188. 10.1111/his.13975 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Llovet, J. M. & Lencioni, R. mRECIST for HCC: Performance and novel refinements. J. Hepatol.72, 288–306. 10.1016/j.jhep.2019.09.026 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Zhong, M. E. et al. CT-based radiogenomic analysis dissects intratumor heterogeneity and predicts prognosis of colorectal cancer: A multi-institutional retrospective study. J. Transl. Med.20, 574. 10.1186/s12967-022-03788-8 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Sautès-Fridman, C., Petitprez, F., Calderaro, J. & Fridman, W. H. Tertiary lymphoid structures in the era of cancer immunotherapy. Nat. Rev. Cancer19, 307–325. 10.1038/s41568-019-0144-6 (2019). [DOI] [PubMed] [Google Scholar]
- 19.Lambin, P. et al. Radiomics: The bridge between medical imaging and personalized medicine. Nat. Rev. Clin. Oncol.14, 749–762. 10.1038/nrclinonc.2017.141 (2017). [DOI] [PubMed] [Google Scholar]
- 20.Bera, K., Braman, N., Gupta, A., Velcheti, V. & Madabhushi, A. Predicting cancer outcomes with radiomics and artificial intelligence in radiology. Nat. Rev. Clin. Oncol.19, 132–146. 10.1038/s41571-021-00560-7 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Xia, T. Y. et al. Predicting microvascular invasion in hepatocellular carcinoma using CT-based radiomics model. Radiology307, e222729. 10.1148/radiol.222729 (2023). [DOI] [PubMed] [Google Scholar]
- 22.Wang, X., Wang, S., Yin, X. & Zheng, Y. MRI-based radiomics distinguish different pathological types of hepatocellular carcinoma. Comput. Biol. Med.141, 105058. 10.1016/j.compbiomed.2021.105058 (2022). [DOI] [PubMed] [Google Scholar]
- 23.Nie, P. et al. CT-based radiomics nomogram: A potential tool for differentiating hepatocellular adenoma from hepatocellular carcinoma in the noncirrhotic liver. Acad. Radiol.28, 799–807. 10.1016/j.acra.2020.04.027 (2021). [DOI] [PubMed] [Google Scholar]
- 24.Kong, C. et al. Prediction of tumor response via a pretreatment MRI radiomics-based nomogram in HCC treated with TACE. Eur. Radiol.31, 7500–7511. 10.1007/s00330-021-07910-0 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Wang, X. H. et al. MRI-based radiomics model for preoperative prediction of 5-year survival in patients with hepatocellular carcinoma. Br. J. Cancer122, 978–985. 10.1038/s41416-019-0706-0 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Wen, Y. et al. Bclaf1 promotes angiogenesis by regulating HIF-1α transcription in hepatocellular carcinoma. Oncogene38, 1845–1859. 10.1038/s41388-018-0552-1 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Zhang, S. et al. Angiotensin-converting enzyme inhibitors have adverse effects in anti-angiogenesis therapy for hepatocellular carcinoma. Cancer Lett.501, 147–161. 10.1016/j.canlet.2020.12.031 (2021). [DOI] [PubMed] [Google Scholar]
- 28.Wei, X. et al. MiR-125b loss activated HIF1α/pAKT loop, leading to transarterial chemoembolization resistance in hepatocellular carcinoma. Hepatology (Baltimore, Md.)73, 1381–1398. 10.1002/hep.31448 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Jiang, L. et al. PITX2C increases the stemness features of hepatocellular carcinoma cells by up-regulating key developmental factors in liver progenitor. J. Exp. Clin. Cancer Res. CR41, 211. 10.1186/s13046-022-02424-z (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Guo, Y. et al. Large HBV surface protein-induced unfolded protein response dynamically regulates p27 degradation in hepatocellular carcinoma progression. Int. J. Mol. Sci.24, 13825. 10.3390/ijms241813825 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Lee, S. A. et al. Male-specific hepatitis B virus large surface protein variant W4P potentiates tumorigenicity and induces gender disparity. Mol. Cancer14, 23. 10.1186/s12943-015-0303-7 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Ding, L. et al. Comprehensive analysis of HHLA2 as a prognostic biomarker and its association with immune infiltrates in hepatocellular carcinoma. Front. Immunol.13, 831101. 10.3389/fimmu.2022.831101 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Rong, D. et al. m6A modification of circHPS5 and hepatocellular carcinoma progression through HMGA2 expression. Mol. Therapy Nucleic acids26, 637–648. 10.1016/j.omtn.2021.09.001 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Ruf, B., Heinrich, B. & Greten, T. F. Immunobiology and immunotherapy of HCC: Spotlight on innate and innate-like immune cells. Cell. Mol. Immunol.18, 112–127. 10.1038/s41423-020-00572-w (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Laumont, C. M., Banville, A. C., Gilardi, M., Hollern, D. P. & Nelson, B. H. Tumour-infiltrating B cells: Immunological mechanisms, clinical impact and therapeutic opportunities. Nat. Rev. Cancer22, 414–430. 10.1038/s41568-022-00466-1 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Fridman, W. H. et al. B cells and tertiary lymphoid structures as determinants of tumour immune contexture and clinical outcome. Nat. Rev. Clin. Oncol.19, 441–457. 10.1038/s41571-022-00619-z (2022). [DOI] [PubMed] [Google Scholar]
- 37.Zhang, P. F. et al. Cancer cell-derived exosomal circUHRF1 induces natural killer cell exhaustion and may cause resistance to anti-PD1 therapy in hepatocellular carcinoma. Mol. Cancer19, 110. 10.1186/s12943-020-01222-5 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Brown, Z. J. et al. Carnitine palmitoyltransferase gene upregulation by linoleic acid induces CD4(+) T cell apoptosis promoting HCC development. Cell Death Dis.9, 620. 10.1038/s41419-018-0687-6 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The data generated for this study are available on request to the corresponding author.