Abstract
Objective: Our understanding of the pathophysiology of pancreatic neuroendocrine tumors (PanNETs) remains incomplete, largely due to their historically underestimated incidence and the perception of these tumors as rare and slow-growing cancers. Additionally, conventional reliance on histological examination alone is gradually being supplemented by the exploration and introduction of molecular biomarkers, such as microRNAs (miRNAs). As miRNAs modulate the expression of multiple genes and pathways involved in the tumorigenesis of PanNETs, these biomarkers hold considerable promise for diagnosis and prognosis applications. In this study, we aimed to identify miRNAs as tissue markers associated with the diagnosis of PanNETs. Methods: We conducted a case-control study including: 7 PanNETs and 19 nontumoral pancreatic tissues obtained from Romanian patients. The samples underwent miRNA profiling via quantitative RT-PCR to assess the expression of 84 miRNAs. Our results were compared with those obtained by reanalyzing a public dataset. Furthermore, we structured our miRNA expression data according to their targeted mRNAs and their roles in signaling pathways. Results: Fourteen miRNAs (miR-1, miR-133a-3p, miR-210-3p, miR-7-5p, miR-10a-5p, miR-92b-3p, miR-132-3p, miR-221-3p, miR-29b-3p, miR-107, miR-103a-3p, let-7b-5p, miR-148a-3p, and miR-202-3p) were identified as differentially expressed by comparing PanNETs with pancreatic nontumoral tissues, with six miRNAs (miR-7-5p, miR-92b-3p, miR-29b-3p, miR-107, miR-103a-3p, and miR-148a-3p) also found in the public dataset analyzed. Bioinformatic analysis revealed that the 14 identified miRNAs target 17 genes. Reanalyzing two public gene expression datasets, five of these genes have been found differentially expressed in PanNET compared to controls. Conclusions: Our preliminary results, albeit limited by a small sample size, highlighted a specific miRNA expression pattern able to distinguish tumoral from normal pancreatic tissue. The diagnostic performance of these miRNAs, matching with circulating miRNAs and validated in more homogeneous and large cohorts, could represent a starting point for improving the diagnostic accuracy of PanNETs.
Keywords: Pancreatic neuroendocrine tumors, miRNA profile, biomarkers, diagnosis, molecular oncology
Introduction
Neuroendocrine neoplasms (NENs) encompass a diverse range of tumors that can develop from cells dispersed throughout the neuroendocrine system. They can occur in various locations within the body, especially in the gastro-entero-pancreatic and bronchopulmonary tract. 1 Using histopathological techniques, NENs can be categorized into well-differentiated pancreatic neuroendocrine tumors (NETs) and poorly differentiated neuroendocrine carcinomas (NECs). 2
Historically, there was a misperception that NETs were rare and benign, however their tumor morphology is highly heterogeneous and can evolve over time from indolent to aggressive behavior. 3 Additionally, recent evidence indicates that NETs are often underdiagnosed, and their incidence appears to be increasing. Recently, the incidence of NETs has been comparable to that of other cancers, such as multiple myeloma, Hodgkin's lymphoma, testicular cancer, and cervical cancer.4,5 Moreover, the prevalence of gastro-entero-pancreatic NETs (GEP-NETs) is higher than that of most gastrointestinal cancers, including pancreatic, gastric, esophageal, and hepatobiliary carcinomas, and is surpassed only by colorectal neoplasia. 4 Globally, the estimated incidence of NENs is 6 per 100,000 individuals, with the prevalence estimated to be 35 per 100,000 individuals.6–8.
Pancreatic NENs (PanNENs) constitute approximately 11% of the total NEN cases. 9 The number of patients diagnosed with PanNENs has risen nearly fivefold in recent decades, with incidence rates being higher in the USA than Europe and showing the most significant increase among NENs.4,10–14 Their rising incidence is also due to advancements in diagnostic technologies, particularly in the fields of ultrasound endoscopy (EUS), which is considered the diagnostic gold standard for PanNENs. 15 More importantly, patients with PanNENs have a worse prognosis than patients with other NENs, as metastases are present at the time of diagnosis in 60% to 80% of PanNETs cases.10,16,17 Consequently, the 5-year survival rates for patients who underwent surgical interventions for PanNETs between 1985 and 2004 ranged from 61% at stage 1 to 16% at stage 4, when surgical removal is not feasible. Therefore, there is a compelling need for new biomarkers and follow-up modalities designed to enhance the outcomes of PanNENs patients. 18
However, current plasma monoanalytes (e.g. chromogranin A) are only indicative of the pathological status of PanNENs, and their positivity typically prompts further diagnostic evaluation through imaging techniques. New diagnostic and prognostic biomarkers are necessary to enhance the stratification of patients and expand treatment possibilities. Recent research has highlighted the significant correlation between poor prognosis and mutations in death domain-associated protein (DAXX) and alpha-thalassemia/mental retardation X-linked chromatin remodeler (ATRX) genes in PanNETs, but also for diagnostic purposes when differentiating PanNETs from pancreatic NECs. 19 Since mutations in both ATRX and DAXX result in the loss of nuclear expression of their proteins, immunohistochemistry (IHC) could be a sensitive and specific tool for detecting these alterations.20,21
In the last two decades, the “omics” approach has been at the forefront of PanNENs diagnosis and prognosis as the sensitivity of technologies has continued to improve, enabling the detection of smaller amounts of biomarkers through noninvasive procedures. This advancement has facilitated earlier diagnosis and a more accurate assessment of minimal residual disease post-treatment. However, the role of markers such as cell-free DNA and circulating tumor cells remains contentious and requires expensive equipment and well-trained personnel for analysis. In contrast, the detection of noncoding RNAs, such as microRNAs (miRNAs) and long noncoding RNAs (lncRNAs), is more cost-effective and accessible, both economically and in terms of expertise.
miRNAs are small noncoding RNAs, composed of approximately 22 nucleotides, that regulate target mRNA, influencing downstream effector protein expression and post-transcriptional gene regulation. Dysregulation of miRNAs is considered a hallmark of human disease, including tumorigenesis.22,23
In the context of NENs, sample data has demonstrated miRNA dysregulation.24–26 Several miRNAs have been examined as candidates for diagnostic purposes in retrospective studies using formalin-fixed paraffin-embedded (FFPE) tissues or serum as PanNET specimens.27–31 However, working with older FFPE samples containing extensively degraded RNA presents a particularly challenging technical concern. 32 Currently, the question of the extent to which formalin crosslinks are reversed during sample preparation remains unresolved. 33 On the other hand, statements about experimental data regarding the characterization of miRNA expression in freshly obtained surgical PanNETs samples are sparse, possibly due to the rarity of the disease. Consequently, there is currently no endorsed miRNA profiling method for clinical applications to enhance our understanding of PanNETs.
The aim of this study is to identify the miRNA expression profile of PanNETs samples using an 84-miRNA panel through a case-control study. The results are compared with those retrieved by reanalyzing a public dataset. Additionally, our miRNA expression data is further organized based on their target mRNAs and their involvement in signaling pathways. The identification of these miRNAs and the related pathways could represent a starting point for better understanding the pathophysiology of PanNETs.
Materials and methods
Sample collection
In this study, 7 PanNETs tissues and 19 nontumoral pancreatic tissues were analyzed. Seven patients with a diagnosis of PanNETs were enrolled in the Fundeni Clinical Institute in Bucharest, Romania, between December 2023 and April 2024. Patient data was collected from hospital records. Nontumoral tissue was selected from three of the seven PanNETs patients, because three others had insufficient peritumoral tissue containing nontumoral cells for molecular analysis, and the remaining patient had nontumoral tissue collected, but the RNA quality was poor. The pancreatic nontumoral control group encompassed these three samples along with 16 nontumoral tissues. The absence of tumoral cells or other pathological diseases in the nontumoral pancreatic tissue was established by a pathologist based on a histological examination.
Only adults aged 18 years or older were included in the study and none of the patients underwent preoperative or postoperative radiotherapy or chemotherapy. During the surgical intervention, the tissue specimens were collected and preserved in RNAprotect Tissue Reagent (Qiagen, Hilden, Germany) for 48 to 72 hours to avoid RNA degradation. After the RNAprotect Tissue Reagent was removed, the samples were stored in the Victor Babes National Institute of Pathology biobank at −80°C until total RNA isolation.
The study was approved by the Ethic Committee of the Fundeni Clinical Institute (Bucharest, approval no. 66032, 15 December 2023), and the Ethic Committee of the Victor Babes National Institute of Pathology (Bucharest, approval no. 78, 3 December 2019), where miRNAs expression analyses were conducted. All the individuals involved in the study signed informed consent forms, in agreement with the Helsinki Declaration of 1975 as revised in 2013.
Immunohistochemistry
Part of the tumoral specimens were processed to perform immunostaining of chromogranin A, synaptophysin, NSE (neuron specific enolase), DAXX, and ATRX for the diagnosis of PanNETs. Paraffin-embedded tissue blocks were cut into 4-microns sections. Immunohistochemical analysis was conducted on tissue sections using the MD Stainer automatized platform (Vitro/Master Diagnostica, Spain) for sample processing for histopathological and immunohistochemical investigations of surgical specimens. Anti-ATRX (polyclonal 782QD-3/V, Cell Signaling Technology, USA, ready-to-use), anti-DAXX (CL3580, Novus Biologicals, USA, dilution of 1:300), antichromogranin A (LK2H10, Vitro Spain, ready-to-use), antisynaptophysin (EP158, Vitro Spain, ready-to-use), and anti-NSE (polyclonal, Vitro Spain, ready-to-use), antibodies were used. PanNETs were graded according to the 2022 WHO classification system 2 based on their mitotic rate and Ki67 IHC. Specifically, grade 1 (G1) was defined as less than 2 mitoses/2 mm2 and Ki67 of <3%; grade 2 (G2) was 2-20 mitoses /2 mm2 or Ki67 of 3% to 20%; or grade 3 (G3) was more than 20 mitoses/2 mm2 or Ki67 of more than 20%. The number of mitoses/2 mm2 was evaluated by pathologist under the Olympus microscope BX43 with a field diameter of 0.55 mm, where mitoses were counted in 42 high power fields (10 mm2) with the resulting number of mitoses divided by 5 to determine the number of mitoses per 2 mm2, which is necessary for assigning the tumor grade. For cases with discordant mitotic rate and Ki67 measurements, the highest grade was assigned. Tumor staging was assessed according to the eighth edition of the American Joint Committee on Cancer Staging Manual. 34
miRNAs expression and gene expression omnibus dataset analysis
The total RNA was isolated using the miRNeasy Mini Kit (Qiagen, Hilden, Germany) according to the manufacturer's instructions, with 25 mg of tissue. Total RNA was assessed for quality and quantity using a spectrophotometric method (NanoDrop 2000, Thermo Scientific, Wilmington, NC, USA). A miRCURY LNA RT Kit (Qiagen, Hilden, Germany) was used to reverse transcribe 10 ng of total RNA. The miRCURY LNA SYBR Green PCR Kit (Qiagen, Hilden, Germany) and the miRCURY LNA miRNA Focus PCR Panel Human Cancer YAHS-102 were used to assess the expression of 84 miRNAs (see Supplemental Table I). Any miRNAs with Ct values above 35 were excluded from the analysis. The Ct values of the target miRNAs were normalized against the geometric mean of two reference RNAs (SNORD38B and SNORD49A). These two RNA were selected after analyzing the stability of three putative reference RNAs (SNORD38B, SNORD49A, and U6 snRNA) on both tumoral and nontumoral tissue, using the RefFinder algorithm (https://www.ciidirsinaloa.com.mx/RefFinder-master/). The miRNA expression data are shown as fold regulation (FR) and as 2−ΔCt values.
A search of the NCBI GEO database was conducted to identify datasets reporting miRNAs and mRNA expression changes in PanNETs compared to nontumoral pancreatic tissue. One dataset was identified (GSE43797 35 ) that analyzed the miRNA expression profiles in solid-pseudopapillary neoplasm of the pancreas, ductal adenocarcinoma, and PanNETs, with the authors using the Agilent-031181 Human miRNAV16.0 Microarray system. Two datasets on mRNA expression levels were identified: GSE43795 35 (expression profiling by array, Illumina HumanHT-12 V4.0 expression beadchip) and GSE73338 36 (expression profiling by array, 18.5 K human oligo microarrays obtained from the Ohio State University Cancer Center). These datasets were reanalyzed using GEO2R for differential expression, considering only the comparison between PanNETs versus controls (GSE43797: 6 PanNETs vs 5 CTRL; GSE43795: 6 PanNETs vs 5 CTRL; GSE73338: 63 PanNETs vs 5 CTRL).
Identification of targeted mRNA and pathway analysis
The miRNAs found differentially expressed in our study were analyzed using miENTURNET (http://userver.bio.uniroma1.it/apps/mienturnet/), a web tool devised for inferring possible evidences of their regulation on target genes, based on a statistical analysis for overrepresentation of miRNA-target interactions. 37 The miENTURNET analysis was based on miRTarbase, with the following filters set: (i) a minimum of five miRNA-target interactions and (ii) a False Discovery Rate (FDR) of <0.05. A pathway analysis of the identified targets was performed using Reactom 2022 through Enrichr, a comprehensive gene set enrichment analysis web server (https://maayanlab.cloud/Enrichr/).
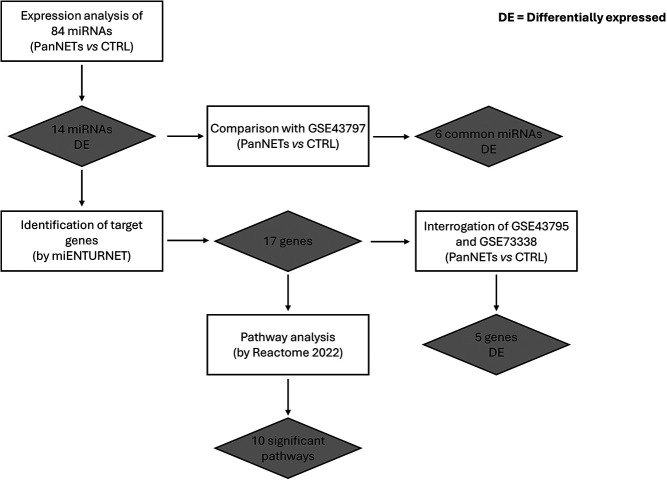
For a better understanding of the workflow a schematic representation was included (Figure 1).
Figure 1.
Schematic representation of the workflow.
Statistical analysis
To ascertain if the distribution of the miRNA expression data was normal, the Shapiro-Wilk test was run. Nonparametric tests were employed for the data analysis due to the nonnormal distribution of the miRNA levels (Shapiro-Wilk p < 0.05). The expression of miRNAs was compared between the PanNETs and nontumoral pancreatic tissue control (CTRL) groups using the Mann-Whitney test. The differences in miRNA expression were considered statistically significant with a p-value < 0.05 and −2 ≥ FR ≥ 2. The categorical and continuous variables related to the sociodemographic features of the groups were compared using a chi-squared test and an unpaired Student's t-test, respectively. Correlations between miRNAs expression and continuous variable were performed using Pearson test.
Results
The PanNET cohort and tumoral samples are described in Table 1.
Table 1.
Characteristics of the seven PanNETs patients included in the study.
| Features | |
|---|---|
| Age (mean ± SD) (years) | 55.14 ± 12.39 years |
| Gender (female/male) | 3 Females, 4 males |
| Grade (G1, G2, G3) | 4 G1, 3 G2 |
| Tumor location (head, body-tail) | 3 Head, 4 body-tail |
| Tumor size (cm) | 2.0 ± 1.31 cm |
| Hepatic metastases | 2 |
| Surgical procedure | 2 Whipple pancreaticoduodenectomies 2 Partial resections 2 Distal pancreatectomies and splenectomies 1 Central resection |
| Immunohistochemistry results | 6 Tumors with SYN+/CHGA+/NSE+ 1 Tumor with SYN+/ CHGA + |
| 4 Tumors with DAXX-/ATRX- 2 Tumors with DAXX+/ATRX- 1 Tumor with DAXX-/ATRX + |
ATRX: alpha-thalassemia/mental retardation X-linked chromatin remodeler; DAXX: death domain-associated protein; NSE: neuron specific enolase; SYN: synaptophysin.
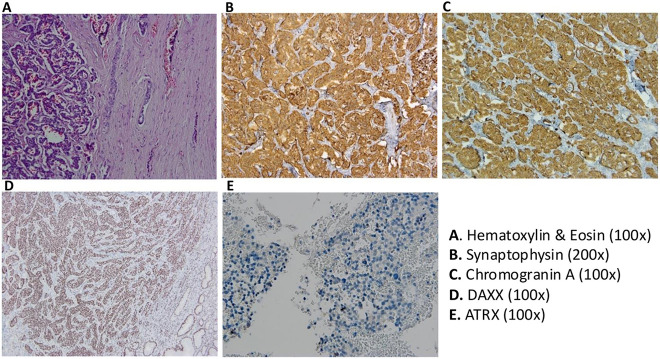
Furthermore, representative examples of nonfunctional pancreatic neuroendocrine tumors that have been assessed by immunolabelling for ATRX and DAXX are represented in Figure 2.
Figure 2.
Representative images of nonfunctional-PanNETs with (a) H&E staining (100×), (b) positive expression for SYN (200×), (c) CgA (100×), (d) positive expression for DAXX (100×), and (e) positive expression for ATRX (100×) which exhibited preserved nuclear expression. ATRX: alpha-thalassemia/mental retardation X-linked; CgA: chromogranin A; DAXX: death domain-associated protein; H&E: hematoxylin & eosin; SYN: synaptophysin.
The patient and CTRL groups were similar in terms of age (PanNET: mean 55.14 ± 12.39, CTRL: mean 59.68 ± 11.03, p = 0.376) and sex distribution (p = 0.780, χ2 = 0.078). All patients were diagnosed with nonfunctional PanNETs. Among them, four patients had small (<2 cm) PanNETs tumors, and all underwent surgical resection. Moreover, these four cases required surgery owing to an overestimation of tumor diameters on imaging. Loss of DAXX/ATRX (negative expression on IHC of both DAXX/ATRX) was observed in three G2 tumors larger than 2 cm, and in one G1 tumor with a tumor size smaller than 2 cm.
The expression analysis revealed 23 miRNAs differentially expressed between the PanNETs and CTRL group (p < 0.05), but only 14 (miR-1, miR-133a-3p, miR-210-3p, miR-7-5p, miR-10a-5p, miR-92b-3p, miR-132-3p, miR-221-3p, miR-29b-3p, miR-107, miR-103a-3p, Let-7b-5p, miR-148a-3p, and miR-202-3p) were significant in terms of FR (three downregulated, and 11 up regulated).
A positive correlation between miR-210-3p and tumor size was found (p = 0.012, r = 0.864). However, the sample size is too small to draw a conclusion based on this correlation. No difference in any miRNA expression was found comparing the DAXX-/ATRX- (n = 4) vs DAXX+/ATRX- and DAXX-/ATRX + (n = 3) and comparing G2 (n = 3) vs G1 (n = 4).
We further compared our miRNA expression results with those obtained by reanalyzing the GSE43797 35 dataset, including only the PanNETs and the nontumoral tissues. All six miRNAs identified as common in the two datasets (miR-7-5p, miR-92b-3p, miR-29b-3p, miR-107, miR-103a-3p, and miR-148a-3p) showed the same expression trend in terms of FR between the groups (up- and downregulated). Five of these were significant in term of their nominal p-value, but only two showed statistical significance after correcting for multiple comparisons (adjusted p-value) (Table 2).
Table 2.
miRNAs found statistically differentially expressed in PanNET versus CTRL: comparison between the results of our study and GSE43797 results.
| Our study 7 PanNETs vs 19 CTRL | GSE43797 6 PanNETs versus 5 CTRL | ||||
|---|---|---|---|---|---|
| miRNA | FR | p-Value | FR | p-Value | Adj p-Value |
| miR-1 | 14.52 | 0.010 | N/A* | N/A | N/A |
| miR-133a-3p | 13.94 | 0.013 | N/A | N/A | N/A |
| miR-210-3p | 6.62 | 0.022 | N/A | N/A | N/A |
| miR-7-5p | 5.64 | 0.010 | 68.89 | <0.00001 | 0.0002 |
| miR-10a-5p | 4.50 | <0.001 | N/A | N/A | N/A |
| miR-92b-3p | 3.64 | 0.030 | 1.15 | 0.0027 | N/S** |
| miR-132-3p | 3.24 | 0.010 | N/A | N/A | N/A |
| miR-221-3p | 2.78 | 0.013 | N/A | N/A | N/A |
| miR-29b-3p | 2.46 | 0.001 | 2.9 | 0.0005 | 0.03 |
| miR-107 | 2.31 | 0.010 | 2.48 | 0.0259 | N/S |
| miR-103a-3p | 2.22 | 0.025 | 1.07 | 0.0479 | N/S |
| Let-7b-5p | −2.41 | 0.002 | N/A | N/A | N/A |
| miR-148a-3p | −7.43 | 0.025 | −8.00; | 0.0095 | N/S |
| miR-202-3p | −41.38 | 0.0189 | N/A | N/A | N/A |
CTRL, control; *N/A, not applicable; ** N/S, not significant; PanNETs, pancreatic neuroendocrine tumors.
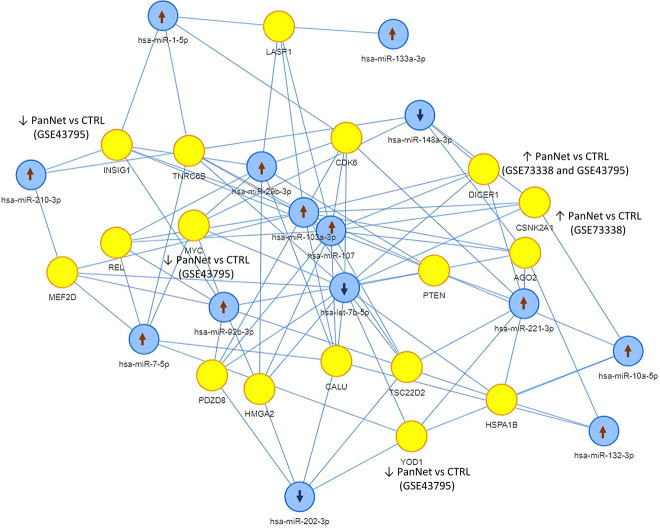
Using the miENTURNET tool we identified a panel of 17 mRNAs (MEF2D, REL, INSIG1, TNRC85, MYC, PDZD8, HMGA2, LASP1, CAL1, CDK6, DICER1, PTEN, TSC22D2, YOD1, HSPA1B, AGO2, and CSNK2A1) targeted by the 14 miRNAs identified in our study (as shown in Table 2). The 17 mRNA targets and their interactions with our 14 miRNAs are depicted in Figure 3. The expression of the identified mRNA was investigated in GSE43795 35 and GSE73338 36 public datasets finding that DICER1 was significantly upregulated in PanNET in both studies. YOD1 and CSNK2A1 were upregulated in the PanNETs included in GSE43795 35 and GSE73338, 36 respectively. INSIG1 and MYC were found downregulated in the PanNETs included in GSE43795 35 compared to controls (Figure 3 and Supplemental Tables II and III).
Figure 3.
The 17 mRNAs targeted by the 14 microRNAs differentially expressed in our PanNETs cohort. Blue and yellow dots represent miRNAs and mRNAs, respectively. The arrows ↑ and ↓ indicate the up and the down regulation of the miRNAs and mRNAs in PanNETs versus controls identified in our study (miRNA) and public datasets GSE43795 and GSE73338 (mRNA).
To highlight the mechanisms through which the identified miRNAs and their mRNAs targets could be potentially involved, a pathway analysis was performed, and the results are reported in Table 3.
Table 3.
The 10 most representative pathways considering the 22 identified mRNA targets.
| Reactome 2022 pathway | Adj p-value | Genes |
|---|---|---|
| Regulation of PTEN mRNA translation R-HSA-8943723 | 1.15E-05 | AGO2;PTEN;TNRC6B |
| Regulation of RUNX1 expression and activity R-HSA-8934593 | 2.36E-05 | CDK6;AGO2;TNRC6B |
| Post-transcriptional silencing by small RNAs R-HSA-426496 | 4.16E-04 | AGO2;TNRC6B |
| Transcriptional regulation By MECP2 R-HSA-8986944 | 4.16E-04 | AGO2;PTEN;TNRC6B |
| Ca2+ pathway R-HSA-4086398 | 4.16E-04 | MYC;AGO2;TNRC6B |
| Cellular responses to stress R-HSA-2262752 | 4.16E-04 | CDK6;MYC;HMGA2;MEF2D;HSPA1B;TNRC6B |
| Cellular responses to stimuli R-HSA-8953897 | 4.16E-04 | CDK6;MYC;HMGA2;MEF2D;HSPA1B;TNRC6B |
| Small interfering RNA (siRNA) biogenesis R-HSA-426486 | 4.18E-04 | AGO2;DICER1 |
| Competing endogenous RNAs (ceRNAs) regulate PTEN translation R-HSA-8948700 | 4.64E-04 | AGO2;TNRC6B |
| TP53 regulates metabolic genes R-HSA-5628897 | 5.72E-04 | AGO2;PTEN;TNRC6B |
Discussion
In this study we compared the miRNA expression profile of PanNETs samples to those from nontumoral pancreatic tissues using an 84-miRNA panel and identified 14 miRNAs differentially expressed between the groups (miR-1, miR-133a-3p, miR-210-3p, miR-7-5p, miR-10a-5p, miR-92b-3p, miR-132-3p, miR-221-3p, miR-29b-3p, miR-107, miR-103a-3p, Let-7b-5p, miR-148a-3p, and miR-202-3p). To provide a more comprehensive description of our findings, our results were further compared with those identified by reanalyzing the GSE43797 dataset, which consisted of six PanNETs and five controls. Specifically, six miRNAs were identified as common between our study and the public dataset (miR-7-5p, miR-92b-3p, miR-29b-3p, miR-107 miR-103a-3p as upregulated and miR-148a-3p as downregulated). After identifying a panel of 17 genes regulated by the 14 identified miRNAs through bioinformatic analysis, we established an association between the 11 upregulated and three downregulated miRNAs and the activation of the 10 most representative signaling pathways.
In recent years, there has been a growing interest in better understanding tumorigenesis mechanisms from the molecular point of view, with miRNA expression emerging as a promising research strategy. In pancreatic ductal adenocarcinoma (PDAC), the most frequent neoplasms in the pancreas, alterations in the expression levels of several miRNAs have been documented in many studies.38–40 Clinical trials are now being conducted to investigate the potential of miRNAs as diagnostic and prognostic biomarkers in PDAC (ClinicalTrials.gov identifiers: NCT06388967, NCT06139042, NCT06760741) and in other types of cancer such as nonmuscle-invasive bladder cancer, breast cancer, non-Hodgkin's lymphoma and acute leukemia, thyroid and colorectal cancer (ClinicalTrials.gov identifiers: NCT03591367, NCT04516330, NCT05477667, NCT04285476, NCT05346757). Although a few clinical trials have been recently completed, the results are not reported. 41
Compared to PDAC, there is relatively limited knowledge regarding the involvement of miRNAs in PanNET tumorigenesis, and their role as diagnostic, prognostic, and therapy response biomarkers.
Disease-associated miRNA profiles have been detected in PanNETs, with distinct pattern correlating with tumor grade, presence of metastasis, and elevated Ki-67 index. Notably, high expression of miR-196a is significantly associated with decreased overall survival in PanNETs. 42 Additionally, miR-21, miR-642, and miR-210 have been identified as prognostic markers, indicating more aggressive tumor behavior correlated with higher Ki-67 index, advanced stages, and spread of the metastases.29,43,44 While these pivotal studies are important, further research is needed.
The most significantly upregulated miRNA in PanNETs of our study and confirmed in the GSE43797 35 datasets was miR-7-5p. In line with our results, this miRNA has been found overexpressed in other neuroendocrine tumors, such as neoplasms of the small intestine, both in tumor and serum 45 and GEP-NETs, 46 while in other type of tumors, such as colorectal, lung, bladder, breast cancers, and glioblastoma it acts as tumor suppressor. 47
miR-92b-3p identified as upregulated in our study compared to the CTRL group, has been reported to play a diagnostic role in distinguishing PanNETs from ileal, appendicular, and rectal NETs. 48 Moreover, high levels of this miRNA have been observed in colon cancer tissues 49 and in prostate cancer, where it operates as an oncomiR and predictor for poor prognosis. 50
An association between miR-29b and neuroendocrine tumors has been suggested since a study has shown that the level of this miRNA is able to discriminate between invasive and noninvasive pituitary neuroendocrine tumors. 51 Moreover, another study has reported an upregulation in gastrointestinal NET and pulmonary carcinoid. 52 Additionally, Özdirik and collaborators found a significantly downregulation of miR-29b in the serum of patients with NETs compared to controls, without finding association with tumor stage and patient's outcome. 53
miR-107, another miRNA found upregulated in our study, is of special interest, since its dysregulation has been observed in other cancers, such as those affecting the colon, pancreas, and stomach.54,55 Furthermore, the diagnostic significance of miR-107 in PDAC or PanNETs is evident from analogous observations, in which its expression was notably elevated compared to nontumoral tissue. Of note, in patients with PDAC, miR-107 expression correlated with worse clinicopathological parameters and prognosis.29,55 Unfortunately, the sample size, the duration of follow-up and the occurrence of events in our study were insufficient to perform this analysis.
To our knowledge, no study has investigated the role in NETs of miR-103a-3p and miR-148a-3p, found up- and downregulated, respectively, both in our study and GSE43797. 35 miR-103a-3p has been found to promote the progression of gastric cancer 56 and nonsmall-cell lung cancer cells via Akt signaling by targeting PTEN, 57 while miR-148a-3p acts as a tumor suppressor in several tumors, including hepatocellular carcinoma, oral, breast, and bladder cancer. 58
Besides the six common differentially expressed miRNAs found in our study and GSE43797, 35 our study revealed upregulation of miR-132-3p in PanNETs samples. Interestingly, PanNETs cases with vascular invasion have been found with high expression levels of miRNA-132-3p, while downregulated in high-graded PanNETs compared to low-graded tumors. 59
Notably, comparing our results with those obtained in a study investigating the circulating miRNAs, we found that let-7b-5p was downregulated also in the serum of patients with PanNETs compared to controls. Kovesdi et al. 60 discovered that the integration of let-7b-5p and other miRNAs into a panel, along with the chromogranin A, could improve the diagnostic accuracy for distinguishing PanNET samples from controls.
Our study, through a bioinformatic analysis also revealed a panel of 17 genes regulated by the miRNAs found differentially expressed in the enrolled patients. The level of these genes has been investigated in two datasets, finding that only YOD1, CSNK2A1, and DICER1 were significantly upregulated in PanNET compared to controls, while a downregulation of INSIG1 and MYC was identified. These transcripts are not key elements of the significant pathways identified in our study, suggesting that the enrichment analysis, in this case, does not allow to draw a conclusion on the biological mechanism involved in PanNETs.
Several observations have provided further insight into the association between the loss of ATRX/DAXX and poorer survival and more aggressive behavior in patients with PanNETs.61,62 Current guidelines 8 recommend active surveillance or local surgical procedure based on tumor size, specifically in NF-PanNETs of 2 cm or less. Still, 15% of these small tumors already present lymph node metastases at diagnosis and can become aggressive.62,63 The loss of ATRX/DAXX could differentiate between patient outcomes, as their prognosis role is under investigation. In our study, we found three distinct immunophenotypic profiles: ATRX-/DAXX- in four cases, ATRX-/DAXX + in two cases and ATRX+/DAXX- in the remaining case. Although in our cases, the loss of ATRX/DAXX expression was predominantly found in G2 tumors with a diameter greater than 2 cm, our data did not allow for a longer follow up to correlate these findings.
Currently, the preoperative diagnosis of PanNETs relies on a limited number of biomarkers, such as tumor size determined through imaging and tumor grade assessed by the Ki67 index on EUS samples. 64 Hence, the discovery of additional pathological markers, such as miRNAs combined with ATRX/DAXX expression, capable of improving the current diagnostic process and stratifying the risk of aggressive lesions would be advantageous for treatment planning. One of the strengths of our study lies in our assessment of miRNA expression profiles in freshly acquired samples obtained immediately after resection. This method minimizes susceptibility to RNA degradation, which occurs in FFPE tissues and enables a more accurate application of miRNA profiling in preoperative samples to distinguish between pancreatic cancer and benign lesions.
The limitations of our study include its small sample size, the heterogeneity of the study population, and the absence of gene expression analysis of the mRNAs targeted by the miRNAs found differentially expressed in our cohort. While this study serves as an initial step, further examination involving an analysis of miRNA via EUS-fine needle aspiration as a diagnosis parameter would be helpful in analyzing their utility in a preoperative manner. In addition, a larger sample size would be necessary to extrapolate the findings presented here.
Conclusions
In conclusion, our preliminary results show a panel of six miRNAs differentially expressed in PanNETs compared to nontumoral pancreatic tissues. Although PanNETs are rare, understanding their molecular mechanisms remains crucial. These data offer compelling exploratory insight in understanding the mechanisms underlying PanNET tumorigenesis and diagnosis.
Supplemental Material
Supplemental material, sj-pdf-1-sci-10.1177_00368504251326864 for miRNA profile in pancreatic neuroendocrine tumors: Preliminary results by Oana A Ciobanu, Vlad Herlea, Elena Milanesi, Maria Dobre and Simona Fica in Science Progress
Supplemental material, sj-pdf-2-sci-10.1177_00368504251326864 for miRNA profile in pancreatic neuroendocrine tumors: Preliminary results by Oana A Ciobanu, Vlad Herlea, Elena Milanesi, Maria Dobre and Simona Fica in Science Progress
Supplemental material, sj-pdf-3-sci-10.1177_00368504251326864 for miRNA profile in pancreatic neuroendocrine tumors: Preliminary results by Oana A Ciobanu, Vlad Herlea, Elena Milanesi, Maria Dobre and Simona Fica in Science Progress
List of abbreviations
- ATRX
alpha-thalassemia/mental retardation X-linked chromatin remodeler
- CTRL
control group
- DAXX
death domain-associated protein
- EUS
Ultrasound endoscopy
- FFPE
formalin-fixed paraffin-embedded
- FR
fold regulation
- G
grade
- GEP-NETs
gastro-entero-pancreatic NETs
- GEO
gene expression omnibus
- IHC
immunohistochemistry
- LncRNAs
longnoncodingRNAs
- miRNAs
microRNAs
- NEC
poorly differentiated neuroendocrine carcinomas
- NENs
neuroendocrine neoplasms
- NETs
neuroendocrine tumors
- NSE
neuron enolase specific
- PanNENs
pancreatic neuroendocrine neoplasms
- PanNETs
pancreatic neuroendocrine tumors
- PDAC
pancreatic ductal adenocarcinoma
Footnotes
Authors’ contributions: OAC, VH, EM, and MD were involved in conceptualization; EM and MD in formal analysis; MD, EM, and VH in investigation; MD and VH in resources; OAC, EM, and MD in drafting of the manuscript; EM, MD, VH, and SF in critical revision of the manuscript for important intellectual content; SF in supervision; and MD in funding acquisition. All authors have read and agreed to the published version of the manuscript.
Consent to participate: Written informed consent was obtained from all the participants involved in the study.
Data availability: Data are contained within this article and the Supplemental materials. Raw data can be obtained from the corresponding author upon reasonable request.
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical considerations: The study was conducted in accordance with the Declaration of Helsinki of 1975 as revised in 2013 and approved by the Ethic Committee of the Fundeni Clinical Institute (Bucharest, approval no. 66032, 15 December 2023), and the Ethic Committee of the Victor Babes National Institute of Pathology (Bucharest, approval no. 78, 3 December 2019), where miRNAs expression analyses were conducted.
Funding: The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was funded by the Romanian Ministry of Research, Innovation and Digitization under grant no. PN 23.16.02.04.
ORCID iDs: Oana A Ciobanu https://orcid.org/0000-0003-2997-4485
Maria Dobre https://orcid.org/0000-0002-1376-4021
Supplementary materials: Supplemental material for this article is available online.
References
- 1.Dasari A, Shen C, Halperin D, et al. Trends in the incidence, prevalence, and survival outcomes in patients with neuroendocrine tumors in the United States. JAMA Oncol 2017; 3: 1335–1342. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Rindi G, Mete O, Uccella S, et al. Overview of the 2022 WHO classification of neuroendocrine neoplasms. Endocr Pathol 2022; 33: 115–154. [DOI] [PubMed] [Google Scholar]
- 3.Lamberti G, La Salvia A. Neuroendocrine tumors: challenges and future perspectives. J Clin Med 2022; 11: 4351. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Lawrence B, Gustafsson BI, Chan A, et al. The epidemiology of gastroenteropancreatic neuroendocrine tumors. Endocrinol Metab Clin North Am 2011; 40: 1–18. [DOI] [PubMed] [Google Scholar]
- 5. Surveillance, Epidemiology, and End Results (SEER) Program . SEER*Stat Data- base: Incidence - SEER 17 Regs Research Data 1 Hurricane Katrina Impacted Louisiana Cases, Nov 2009 Sub (1973-2007 varying) - Linked To County Attri- butes - Total U.S., 1969-2007 Counties, National Cancer Institute, DCCPS, Surveillance Research Program, Cancer Statistics Branch, released April 2010, based on the November 2009 submission. Available at: www.seer.cancer.gov. Accessed May 12, 2009. - Căutare Google. Accessed May 10, 2024. https://www.google.com/search?q=Surveillance%2C+Epidemiology%2C+and+End+Results+(SEER)+Program.+SEER*Stat+Data-+base%3A+Incidence+-+SEER+17+Regs+Research+Data+1+Hurricane+Katrina+Impacted+Louisiana+Cases%2C+Nov+2009+Sub+(1973-2007+varying)+-+Linked+To+County+Attri-+butes+-+Total+U.S.%2C+1969-2007+Counties%2C+National+Cancer+Institute%2C+DCCPS%2C+Surveillance+Research+Program%2C+Cancer+Statistics+Branch%2C+released+April+2010%2C+based+on+the+November+2009+submission.+Available+at%3A+www.seer.cancer.gov.+Accessed+May+12%2C+2009.&oq=Surveillance%2C+Epidemiology%2C+and+End+Results+(SEER)+Program.+SEER*Stat+Data-+base%3A+Incidence+-+SEER+17+Regs+Research+Data+1+Hurricane+Katrina+Impacted+Louisiana+Cases%2C+Nov+2009+Sub+(1973-2007+varying)+-+Linked+To+County+Attri-+butes+-+Total+U.S.%2C+1969-2007+Counties%2C+National+Cancer+Institute%2C+DCCPS%2C+Surveillance+Research+Program%2C+Cancer+Statistics+Branch%2C+released+April+2010%2C+based+on+the+November+2009+submission.+Available+at%3A+www.seer.cancer.gov.+Accessed+May+12%2C+2009.&gs_lcrp=EgZjaHJvbWUqBggAEEUYOzIGCAAQRRg70gEHMzY3ajBqN6gCALACAA&sourceid=chrome&ie=UTF-8.
- 6.Ramage JK, De Herder WW, Delle Fave G, et al. ENETS consensus guidelines update for colorectal neuroendocrine neoplasms. Neuroendocrinology 2016; 103: 139–143. [DOI] [PubMed] [Google Scholar]
- 7.Öberg K, Castellano D. Current knowledge on diagnosis and staging of neuroendocrine tumors. Cancer Metastasis Rev 2011; 30: 3–7. [DOI] [PubMed] [Google Scholar]
- 8.Pavel M, Öberg K, Falconi M, et al. Gastroenteropancreatic neuroendocrine neoplasms: ESMO clinical practice guidelines for diagnosis, treatment and follow-up. Ann Oncol 2020; 31: 844–860. [DOI] [PubMed] [Google Scholar]
- 9.Man D, Wu J, Shen Zet al. et al. Prognosis of patients with neuroendocrine tumor: a SEER database analysis. Cancer Manag Res 2018; 10: 5629–5638. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Wyld D, Wan MH, Moore J, et al. Epidemiological trends of neuroendocrine tumours over three decades in Queensland, Australia. Cancer Epidemiol 2019; 63: 101598. [DOI] [PubMed] [Google Scholar]
- 11.Heetfeld M, Chougnet CN, Olsen IH, et al. Characteristics and treatment of patients with G3 gastroenteropancreatic neuroendocrine neoplasms. Endocr Relat Cancer 2015; 22: 657–664. [DOI] [PubMed] [Google Scholar]
- 12.Lee MR, Harris C, Baeg KJ, et al. Incidence trends of gastroenteropancreatic neuroendocrine tumors in the United States. Clin Gastroenterol Hepatol 2019; 17: 2212–2217.e1. [DOI] [PubMed] [Google Scholar]
- 13.Yao JC, Hassan M, Phan A, et al. One hundred years after “carcinoid”: epidemiology of and prognostic factors for neuroendocrine tumors in 35,825 cases in the United States. J Clin Oncol 2008; 26: 3063–3072. [DOI] [PubMed] [Google Scholar]
- 14.Fraenkel M, Kim M, Faggiano A, et al. Incidence of gastroenteropancreatic neuroendocrine tumours: a systematic review of the literature. Endocr Relat Cancer 2014; 21: R153–R163. [DOI] [PubMed] [Google Scholar]
- 15.Rossi RE, Massironi S. The increasing incidence of neuroendocrine neoplasms worldwide: current knowledge and open issues. J Clin Med 2022; 11: 3794. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Gudmundsdottir H, Möller PH, Jonasson JGet al. et al. Gastroenteropancreatic neuroendocrine tumors in Iceland: a population-based study. Scand J Gastroenterol 2019; 54: 69–75. [DOI] [PubMed] [Google Scholar]
- 17.Modlin IM, Oberg K, Chung DC, et al. Gastroenteropancreatic neuroendocrine tumours. Lancet Oncol 2008; 9: 61–72. [DOI] [PubMed] [Google Scholar]
- 18. Survival Rates for Pancreatic Cancer . American Cancer Society. Accessed May 9, 2024. https://www.cancer.org/cancer/types/pancreatic-cancer/detection-diagnosis-staging/survival-rates.html.
- 19.Heaphy CM, Singhi AD. The diagnostic and prognostic utility of incorporating DAXX, ATRX, and alternative lengthening of telomeres to the evaluation of pancreatic neuroendocrine tumors. Hum Pathol 2022; 129: 11–20. [DOI] [PubMed] [Google Scholar]
- 20.Hechtman JF, Klimstra DS, Nanjangud G, et al. Performance of DAXX immunohistochemistry as a screen for DAXX mutations in pancreatic neuroendocrine tumors. Pancreas 2019; 48: 396–399. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Couvelard A, Cazes A, Cros J. Updates in histopathological classification and tissue biomarkers of digestive neuroendocrine neoplasms: what the clinician should know. Best Pract Res Clin Endocrinol Metab 2023; 37: 101795. [DOI] [PubMed] [Google Scholar]
- 22.Lin S, Gregory RI. MicroRNA biogenesis pathways in cancer. Nat Rev Cancer 2015; 15: 321–333. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Havasi A, Sur D, Cainap SS, et al. Current and new challenges in the management of pancreatic neuroendocrine tumors: the role of miRNA-based approaches as new reliable biomarkers. Int J Mol Sci 2022; 23: 1109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Bowden M, Zhou CW, Zhang S, et al. Profiling of metastatic small intestine neuroendocrine tumors reveals characteristic miRNAs detectable in plasma. Accessed May 9, 2024. www.impactjournals.com/oncotarget.
- 25.Arvidsson Y, Rehammar A, Bergström A, et al. miRNA profiling of small intestinal neuroendocrine tumors defines novel molecular subtypes and identifies miR-375 as a biomarker of patient survival. Mod Pathol 2018; 31: 1302–1317. [DOI] [PubMed] [Google Scholar]
- 26.Vicentini C, Fassan M, D’Angelo E, et al. Clinical application of microRNA testing in neuroendocrine tumors of the gastrointestinal tract. Molecules 2014; 19: 2458–2468. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Vicentini C, Calore F, Nigita G, et al. Exosomal miRNA signatures of pancreatic lesions. BMC Gastroenterol 2020; 20: 137. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Gill P, Kim E, Chua TC, et al. MiRNA-3653 is a potential tissue biomarker for increased metastatic risk in pancreatic neuroendocrine tumours. Endocr Pathol 2019; 30: 128–133. [DOI] [PubMed] [Google Scholar]
- 29.Roldo C, Missiaglia E, Hagan JP, et al. MicroRNA expression abnormalities in pancreatic endocrine and acinar tumors are associated with distinctive pathologic features and clinical behavior. J Clin Oncol 2006; 24: 4677–4684. [DOI] [PubMed] [Google Scholar]
- 30.Lee YS, Kim H, Kim HW, et al. High expression of microRNA-196a indicates poor prognosis in resected pancreatic neuroendocrine tumor. Medicine (Baltimore) 2015; 94: e2224. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Liu Z, Sandhu S, Gambardella G, et al. Altered miRNAs expression correlates with gastroenteropancreatic neuroendocrine tumors grades. Front Oncol 2020; 1: 1187. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Cannizzo MD, Wood CE, Hester SDet al. et al. Case study: targeted RNA-sequencing of aged formalin-fixed paraffin-embedded samples for understanding chemical mode of action. Toxicol Rep 2022; 9: 883–894. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Obi EN, Tellock DA, Thomas GJet al. et al. Biomarker analysis of formalin-fixed paraffin-embedded clinical tissues using proteomics. Biomolecules 2023; 13: 96. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Amin MB, Greene FL, Edge SB, et al. The eighth edition AJCC cancer staging manual: continuing to build a bridge from a population-based to a more “personalized” approach to cancer staging. CA Cancer J Clin 2017; 67: 93–99. [DOI] [PubMed] [Google Scholar]
- 35.Park M, Kim M, Hwang D, et al. Characterization of gene expression and activated signaling pathways in solid-pseudopapillary neoplasm of pancreas. Mod Pathol 2014; 27: 580–593. [DOI] [PubMed] [Google Scholar]
- 36.Sadanandam A, Wullschleger S, Lyssiotis CA, et al. A cross-species analysis in pancreatic neuroendocrine tumors reveals molecular subtypes with distinctive clinical, metastatic, developmental, and metabolic characteristics. Cancer Discov 2015; 5: 1296–1313. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Licursi V, Conte F, Fiscon Get al. et al. MIENTURNET: an interactive web tool for microRNA-target enrichment and network-based analysis. BMC Bioinform 2019; 20: 1–10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Tesfaye AA, Azmi AS, Philip PA. Pancreatic cancer theme issue REVIEW miRNA and gene expression in pancreatic ductal adenocarcinoma. Am J Pathol 2019; 189: 58–70. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Mok ETY, Chitty JL, Cox TR. miRNAs in pancreatic cancer progression and metastasis. Clin Exp Metastasis 2024; 41: 163–186. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Dobre M, Poenaru RC, Niculae AM, et al. Increased levels of miR-15b-5p and miR-20b-5p in pancreatic ductal adenocarcinoma with hepatic metastases. Genes (Basel) 2023; 14: 1577. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Kim T, Croce CM. MicroRNA: trends in clinical trials of cancer diagnosis and therapy strategies. Exp Mol Med 2023; 55: 1314–1321. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Lee YS, Kim H, Kim HW, et al. High expression of microRNA-196a indicates poor prognosis in resected pancreatic neuroendocrine tumor. Medicine (Baltimore) 2015; 94: e2224. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Zatelli MC, Grossrubatscher EM, Guadagno E, et al. Circulating tumor cells and miRNAs as prognostic markers in neuroendocrine neoplasms. Endocr Relat Cancer 2017; 24: R223–R237. [DOI] [PubMed] [Google Scholar]
- 44.McMurry HS, Rivero JD, Chen EY, et al. Gastroenteropancreatic neuroendocrine tumors: epigenetic landscape and clinical implications. Curr Probl Cancer 2024; 52: 101131. [DOI] [PubMed] [Google Scholar]
- 45.Heverhagen AE, Legrand N, Wagner V, et al. Overexpression of microRNA miR-7-5p is a potential biomarker in neuroendocrine neoplasms of the small intestine. Neuroendocrinology 2018; 106: 312–317. [DOI] [PubMed] [Google Scholar]
- 46.Cavalcanti E, Galleggiante V, Coletta S, et al. Altered miRNAs expression correlates with gastroenteropancreatic neuroendocrine tumors grades. Front Oncol 2020; 10: 1187. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Mirzaei Z, Barati T, Ebrahimi A, et al. The role of mir-7-5p in cancer: function, prognosis, diagnosis, and therapeutic implications. Mol Biol Rep 2024; 52: 12. [DOI] [PubMed] [Google Scholar]
- 48.Panarelli N, Tyryshkin K, Mun Wong JJ, et al. Evaluating gastroenteropancreatic neuroendocrine tumors through microRNA sequencing. Endocr Relat Cancer 2019; 26: 47–57. [DOI] [PubMed] [Google Scholar]
- 49.Liu X, Zhang L. microRNA-92b-3p augments colon cancer development through inhibiting KLF3. J Biochem Mol Toxicol 2023; 37: e23488. [DOI] [PubMed] [Google Scholar]
- 50.Wang G, Cheng B, Jia R, et al. Altered expression of microRNA-92b-3p predicts survival outcomes of patients with prostate cancer and functions as an oncogene in tumor progression. Oncol Lett 2021; 21: 1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Nikolova E, Nankova A, Kalenderova S, et al. MiRNA signatures related to invasiveness and recurrence in patients with non-functioning pituitary neuroendocrine tumors. Exp Clin Endocrinol Diabetes 2024; 132: 240–248. [DOI] [PubMed] [Google Scholar]
- 52.Yoshimoto T, Motoi N, Yamamoto N, et al. Pulmonary carcinoids and low-grade gastrointestinal neuroendocrine tumors show common microRNA expression profiles, different from adenocarcinomas and small cell carcinomas. Neuroendocrinology 2018; 106: 47–57. [DOI] [PubMed] [Google Scholar]
- 53.Özdirik B, Stueven AK, Mohr R, et al. Analysis of miR-29 serum levels in patients with neuroendocrine tumors—results from an exploratory study. J Clin Med 2020; 9: 2881. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Volinia S, Calin GA, Liu CG, et al. A microRNA expression signature of human solid tumors defines cancer gene targets. Published online 2006. Accessed May 10, 2024. https://www.pnas.org.
- 55.Xiong J, Wang D, Wei A, et al. Deregulated expression of miR-107 inhibits metastasis of PDAC through inhibition PI3 K/Akt signaling via caveolin-1 and PTEN. Exp Cell Res 2017; 361: 316–323. [DOI] [PubMed] [Google Scholar]
- 56.Hu X, Miao J, Zhang M, et al. miRNA-103a-3p promotes human gastric cancer cell proliferation by targeting and suppressing ATF7 in vitro. Mol Cells 2018; 41: 390–400. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Li H, Huhe M, Lou J. MicroRNA-103a-3p promotes cell proliferation and invasion in non-small-cell lung cancer cells through Akt pathway by targeting PTEN. Biomed Res Int 2021; 2021: 1–10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Yuan X, Wen Y, Shi Q, et al. MicroRNA-148a-3p suppresses the glycolysis and cell proliferation by targeting transmembrane protein 54 in liver cancer. Biochem Biophys Res Commun 2024; 695: 149424. [DOI] [PubMed] [Google Scholar]
- 59.Klieser E, Urbas R, Swierczynski S, et al. HDAC-linked “proliferative” miRNA expression pattern in pancreatic neuroendocrine tumors. Int J Mol Sci 2018; 19: 2781. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Kövesdi A, Kurucz PA, Nyír G, et al. Circulating miRNA increases the diagnostic accuracy of chromogranin A in metastatic pancreatic neuroendocrine tumors. Cancers (Basel) 2020; 12: 2488. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Marinoni I, Kurrer AS, Vassella E, et al. Loss of DAXX and ATRX are associated with chromosome instability and reduced survival of patients with pancreatic neuroendocrine tumors. Gastroenterology 2014; 146: 453–460.e5. [DOI] [PubMed] [Google Scholar]
- 62.Wm H, Laa B, Jy K, et al. Non-functional pancreatic neuroendocrine tumours: ATRX/DAXX and alternative lengthening of telomeres (ALT) are prognostically independent from ARX/PDX1 expression and tumour size. Gut 2022; 71: 961–973. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Park JK, Paik WH, Lee K, et al. DAXX/ATRX and MEN1 genes are strong prognostic markers in pancreatic neuroendocrine tumors. Oncotarget 2017; 8: 49796–49806. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Mastrosimini MG, Manfrin E, Remo A, et al. Endoscopic ultrasound fine-needle biopsy to assess DAXX/ATRX expression and alternative lengthening of telomeres status in non-functional pancreatic neuroendocrine tumors. Pancreatology 2023; 23: 429–436. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Supplemental material, sj-pdf-1-sci-10.1177_00368504251326864 for miRNA profile in pancreatic neuroendocrine tumors: Preliminary results by Oana A Ciobanu, Vlad Herlea, Elena Milanesi, Maria Dobre and Simona Fica in Science Progress
Supplemental material, sj-pdf-2-sci-10.1177_00368504251326864 for miRNA profile in pancreatic neuroendocrine tumors: Preliminary results by Oana A Ciobanu, Vlad Herlea, Elena Milanesi, Maria Dobre and Simona Fica in Science Progress
Supplemental material, sj-pdf-3-sci-10.1177_00368504251326864 for miRNA profile in pancreatic neuroendocrine tumors: Preliminary results by Oana A Ciobanu, Vlad Herlea, Elena Milanesi, Maria Dobre and Simona Fica in Science Progress