Abstract
The biofilm formation capacity of Staphylococcus aureus clinical isolates is considered an important virulence factor for the establishment of chronic infections. Environmental conditions affect the biofilm formation capacity of S. aureus, indicating the existence of positive and negative regulators of the process. The majority of the screening procedures for identifying genes involved in biofilm development have been focused on genes whose presence is essential for the process. In this report, we have used random transposon mutagenesis and systematic disruption of all S. aureus two-component systems to identify negative regulators of S. aureus biofilm development in a chemically defined medium (Hussain-Hastings-White modified medium [HHWm]). The results of both approaches coincided in that they identified arlRS as a repressor of biofilm development under both steady-state and flow conditions. The arlRS mutant exhibited an increased initial attachment as well as increased accumulation of poly-N-acetylglucosamine (PNAG). However, the biofilm formation of the arlRS mutant was not affected when the icaADBC operon was deleted, indicating that PNAG is not an essential compound of the biofilm matrix produced in HHWm. Disruption of the major autolysin gene, atl, did not produce any effect on the biofilm phenotype of an arlRS mutant. Epistatic experiments with global regulators involved in staphylococcal-biofilm formation indicated that sarA deletion abolished, whereas agr deletion reinforced, the biofilm development promoted by the arlRS mutation.
Staphylococcus aureus is a gram-positive bacterium responsible for many infections ranging from folliculitis and food-borne intoxications to severe endocarditis, osteomyelitis, or septicemia. S. aureus can live harmlessly on many skin surfaces, especially around the nose, mouth, genitals, and rectum. This organism shares with other pathogens the ability to adhere to catheters and other indwelling devices and form multicellular communities embedded in an exopolysaccharidic matrix known as a biofilm (for reviews, see references 22 and 11). Inside the biofilm, S. aureus becomes more resistant to antibiotic treatments and the actions of the immune system (2, 23). As a consequence, staphylococcal-biofilm-associated infections of this type are difficult to eradicate, and most of them can be eliminated only by the removal and substitution of the contaminated implant.
With the exception of the icaADBC operon, discovered by heterologous complementation of a biofilm-negative Staphylococcus carnosus isolate with a genomic library of Staphylococcus epidermidis RP62A (29), early genetic studies of the staphylococcal-biofilm formation process have been performed using transposon mutagenesis on a biofilm-positive isolate and the ensuing selection of biofilm-deficient mutants by a microtiter plate assay (6, 13, 24, 27, 28, 37, 38, 42, 46, 49). These studies have been crucial for identifying S. aureus genes whose presence is essential for primary attachment and/or biofilm development under the environmental conditions used during the screening process. However, this approach, based on the selection of biofilm-deficient mutants, biased the screening in favor of those genes that encode products that are required for the biofilm development process, either because their enzymatic activities are required for the synthesis of compounds of the biofilm matrix, because they are themselves compounds of the matrix, or simply because they play a positive regulatory role in the process. As regards the regulatory genes, a lot of evidence suggests the presence of negative regulatory mechanisms in the expression of the product of the icaADBC operon, the extracellular poly-N-acetylglucosamine (PNAG) polysaccharide, and consequently on the formation of a biofilm. First, icaR, located adjacent to the ica operon, encodes a transcriptional repressor involved in the environmental regulation of ica operon expression both in S. aureus (33) and in S. epidermidis (10). Second, TcaR, a transcriptional regulator of the teicoplanin-associated locus, has very recently been described as a negative regulator of ica operon transcription, though deletion of tcaR alone did not induce changes in PNAG production and adherence to polystyrene (33). Third, strong biofilm development can be induced when Trypticase soy broth (TSB) medium is supplemented with 0.5 M of NaCl, indicating that NaCl either induces or derepresses the expression of biofilm genes (45, 49). In this regard, the capacity of S. aureus to produce a biofilm is dependent on the environmental conditions, which suggests that there is a mechanism that senses and responds to environmental signals. Two-component systems (TCS) are common signal transduction mechanisms used by bacteria to modulate gene expression in response to a variety of stimuli. To date, only two TCS of S. aureus, agr and arlRS, have been related to biofilm formation. As regards agr, an epidemiological study demonstrated that agr mutants of S. aureus can frequently be isolated and that these mutants exhibit an enhanced attachment to polystyrene compared to that of agr-positive isolates (51). In the case of arlRS, it has been reported that an arlS transposon mutant, isolated according to its increased resistance to tetraphenylphosphonium bromide, displayed an enhanced capacity to form biofilms, but the mechanism responsible for this enhancement was not determined (18).
This study has identified those genes whose expression repressed the biofilm formation process in a chemically defined medium (Hussain-Hastings-White modified medium [HHWm]).For that identification, we have used systematic mutagenesis of two-component systems and random transposon mutagenesis in combination with screening for biofilm-positive mutants. The results revealed that the deletion of the arlRS two-component system bestows on S. aureus the capacity to form a biofilm in HHWm by a mechanism independent of the presence of the PNAG exopolysaccharide.
MATERIALS AND METHODS
Bacterial strains, culture conditions, and plasmids.
The most relevant bacterial strains and plasmids used in this study are listed in Table 1. Escherichia coli XL1-Blue cells were grown in Luria-Bertani (LB) broth or on LB agar (Pronadisa, Madrid, Spain) with appropriate antibiotics. Staphylococcal strains were cultured using different media as indicated: Trypticase soy agar (TSA), Trypticase soy broth supplemented with glucose (TSBg) (0.25%, wt/vol), B2 medium (1% casein hydrolysate, 2.5% yeast extract, 2.5% NaCl, 0.1% K2HPO4, and 0.5% glucose [wt/vol]), and HHWm (Table 2). Biofilm-positive mutants were screened on microtiter polystyrene plates using HHWm. Media were supplemented with appropriate antibiotics at the following concentrations: erythromycin (Erm), 20 μg ml−1 or 1.5 μg ml−1; ampicillin, 100 μg ml−1; chloramphenicol, 20 μg ml−1; kanamycin (Km), 50 μg ml−1; and tetracycline (Tet), 10 μg ml−1. When required, TSA was supplemented with X-Gal (Bioline, London, United Kingdom).
TABLE 1.
Strains and plasmids used in this study
| Strain(s) or plasmid | Relevant characteristic(s) | Source or referencea |
|---|---|---|
| S. aureus strains | ||
| 15981 | Clinical strain isolate; biofilm positive in TSBg | 49 |
| 12313 | Clinical strain isolate; biofilm positive | CUN |
| V329 | Bovine subclinical mastitis isolate; biofilm positive | 13 |
| 7879 | Clinical strain isolate; biofilm positive in TSBg | CUN |
| 49 | Clinical strain isolate; biofilm positive in TSBg | CUN |
| 1246 | Clinical strain isolate; biofilm positive in TSBg | CUN |
| G-3320 | Clinical strain isolate; biofilm positive in TSBg | CUN |
| MM4, -7, -13, and -14 | 15981 arlS::Tn917 | This study |
| MM11 and MM19 | 15981 arlR::Tn917 | This study |
| 15981 ΩTCS-3 | 15981 pBT2 SA0214 gene insertional mutant | This study |
| 15981 ΩTCS-4 | 15981 pBT2 lytS gene insertional mutant | This study |
| 15981 ΩTCS-5 | 15981 pBT2 SA0614 gene insertional mutant | This study |
| 15981 ΔsaeRS | 15981 with deletion of the saeRS genes | This study |
| 15981 ΔTCS-7 | 15981 with deletion of the SA1158-SA1159 sequence | This study |
| 15981 ΔarlRS | 15981 with deletion of the arlRS operon | This study |
| 15981 ΔsrrAB | 15981 with deletion of the srrAB genes | This study |
| 15981 ΔTCS-10 | 15981 with deletion of the SA1515-SA1516 sequence | This study |
| 15981 ΩTCS-11 | 15981 pBT2 SA1667 insertional mutant | This study |
| 15981 ΔTCS-12 | 15981 with deletion of the vraSR genes | This study |
| 15981 Δagr | 15981 with deletion of the agrBDCA genes | 49 |
| 15981 ΩTCS-14 | 15981 pBT2 SA1882 insertional mutant | This study |
| 15981 ΔTCS-15 | 15981 with deletion of the SA2151-SA2152 sequence | This study |
| 15981 ΩTCS-16 | 15981 pBT2 SA2180 insertional mutant | This study |
| 15981 ΩTCS-17 | 15981 pBT2 SA2418 insertional mutant | This study |
| 15981 pCU1 | 15981 complemented with pCU1 | This study |
| 15981 ΔarlRS pCU1 | ΔarlRS complemented with pCU1 | This study |
| 15981 ΔarlRS parlRS | ΔarlRS complemented with the arlRS operon | This study |
| 15981 pSB2019 | 15981 complemented with the pSB2019 plasmid, which expresses GFP | This study |
| 15981 ΔarlRS pSB2019 | ΔarlRS complemented with the pSB2019 plasmid, which expresses GFP | This study |
| 15981 Δica | 15981 with deletion of the icaADBC operon | This study |
| 15981 ΔarlRS Δica | ΔarlRS with deletion of the icaADBC operon | This study |
| 15981 ΔsarA | 15981 with deletion of the sarA gene | 49 |
| 15981 ΔarlRS ΔsarA | ΔarlRS with deletion of the sarA gene | This study |
| ISP479c | ISP479 with rsbU deficiency. Cured of the pID258 plasmid | 49 |
| ISP479r | ISP479c with rsbU restored | This study |
| ISP479r ΔarlRS | ISP479r with a deletion of the arlRS operon | This study |
| ISP479-sarA | ISP479 sarA::Km | 49 |
| ISP479r-sarA | ISP479r sarA::Km | This study |
| ISP479r ΔarlRS-sarA | ISP479r ΔarlRS sarA::Km | This study |
| ISP479-agr | ISP479 agr::tet | 49 |
| ISP479r-agr | ISP479r agr::tet | This study |
| ISP479r-agr ΔarlRS | ISP479r agr::tet with deletion of the arlRS operon | This study |
| SH108 | 8325-4 atl::LacZ (Erm) | 17 |
| ISP479r-atl | ISP479r atl::LacZ (Erm) | This study |
| ISP479r ΔarlRS-atl | ISP479r ΔarlRS atl::LacZ (Erm) | This study |
| ISP479r-agr-atl | ISP479r agr::tet atl::LacZ (Erm) | This study |
| ISP479r-agr ΔarlRS-atl | ISP479r agr::tet ΔarlRS atl::LacZ (Erm) | This study |
| Plasmids | ||
| pMAD | E. coli-S. aureus (Listeria) shuttle vector with the bgaB gene encoding a β-galactosidase. Apr/Err | 3 |
| pMAD TCS6AD | pMAD plasmid contained the mutant allele for the deletion of the saeRS genes | This study |
| pMAD TCS7AD | pMAD plasmid containing the mutant allele for deletion of the SA1158-SA1159 sequence | This study |
| pMAD TCS8AD | pMAD plasmid containing the mutant allele for deletion of the arlRS operon | This study |
| pMAD TCS9AD | pMAD plasmid containing the mutant allele for deletion of the srrAB genes | This study |
| pMAD TCS10AD | pMAD plasmid containing the mutant allele for deletion of the SA1515-SA1516 sequence | This study |
| pMAD TCS12AD | pMAD plasmid containing the mutant allele for deletion of the vraSR genes | This study |
| pMAD TCS13AD | pMAD plasmid containing the mutant allele for deletion of the agrBDCA genes | 49 |
| pMAD TCS15AD | pMAD plasmid containing the mutant allele for deletion of the SA2151-SA2152 sequence | This study |
| pBT2 | E. coli-S. aureus shuttle vector with thermosensitive origin of replication for S. aureus | 8 |
| pBT2 TCS-1 | pBT2 plasmid with an internal fragment of the yycF gene | This study |
| pBT2 TCS-2 | pBT2 plasmid with an internal fragment of the SA0067 sequence | This study |
| pBT2 TCS-3 | pBT2 plasmid with an internal fragment of the SA0214 sequence | This study |
| pBT2 TCS-4 | pBT2 plasmid with an internal fragment of the lytS gene | This study |
| pBT2 TCS-5 | pBT2 plasmid with an internal fragment of the SA0614 sequence | This study |
| pBT2 TCS-11 | pBT2 plasmid with an internal fragment of the SA1667 sequence | This study |
| pBT2 TCS-14 | pBT2 plasmid with an internal fragment of the SA1882 sequence | This study |
| pBT2 TCS-16 | pBT2 plasmid with an internal fragment of the SA2180 sequence | This study |
| pBT2 TCS-17 | pBT2 plasmid with an internal fragment of the SA2418 sequence | This study |
| pCU1 | Vector for complementation experiments. Apr Cmr | 4 |
| parlRS | pCU1 plasmid containing arlRS operon | This study |
| pMAL30 | 3-kb OL-80-OL-81 rsbU rsbV rsbW sigB PCR fragment in pAZ106 | 30 |
| pMAD rsbU | pMAD plasmid containing the 3-kb fragment encompassing the complete rsbU-rsbV-rsbW region and part of sigB from plasmid pMAL30 | This study |
| pMAD ica | pMAD plasmid containing the mutant allele for deletion of the icaADBC operon | This study |
| pMAD sarAD | pMAD plasmid containing the mutant allele for deletion of the sarA gene | 49 |
| pSB2019 | Translationally enhanced gfp-3 under the control of PxylA, with the xylR repressor removed to give xylose-independent expression. Apr Cmr | 44 |
CUN, Clínica Universitaria de Navarra (University Clinic of Navarre, Spain).
TABLE 2.
Compositions of chemically defined HHWm
| Ingredient(s)a | Volb | Final concn (mg/liter) |
|---|---|---|
| Sterile MilliQ water | 710 | |
| Solution A (10×) | 100 | |
| Na2HPO4 · 2H2O | 8,950 | |
| KH2PO4 | 3,000 | |
| (NH4)2SO4 | 2,000 | |
| Solution B (50×) | 20 | |
| MgSO4 · 7H2O | 50 | |
| MnSO4 · H2O | 3.8 | |
| FeSO4 · 7H2O | 2.8 | |
| Solution C (50×) | 20 | |
| Glucose | 10,000 | |
| Solution D (100×) | 10 | |
| Biotin | 0.1 | |
| Nicotinic acid | 2 | |
| d-Panthothenic acid, Ca salt | 2 | |
| Pyridoxyne hydrochloride (vitamin B6) | 4 | |
| Riboflavin | 2 | |
| Thiamine hydrochloride (vitamin B1) | 2 | |
| Solution adenine (100×) | 50 | |
| Adenine · 1/2 H2SO4 | 3 | |
| Solution guanine (100×) | 50 | |
| Guanine · 1/2 H2SO4 | 4 | |
| Solution trace elements (20×) | 10 | |
| CaCl2 · 2H2O | 7.28 | |
| ZnCl2 | 1.70 | |
| CuCl2 · 2H2O | 0.43 | |
| CoCl2 · 6H2O | 0.60 | |
| Na2MoO4 · 2H2O | 0.60 | |
| Solution amino acids (50×)c | 20 | |
| Solution glutamine (100×) | 10 | |
| Glutamine | 292 | |
| Solution Ca (100×) | 0.14 | |
| CaCl2 · 2H2O | 735 |
The solutions are listed in the order they are aggregated to the sterile water.
Milliliters of each solution to prepare 1 liter of HHWm.
RPMI 1640 amino acids solution without l-glutamine (Sigma; catalog no. R7131).
DNA manipulations.
DNA plasmids were isolated from the E. coli strain using the Bio-Rad plasmid miniprep kit (Bio-Rad Laboratories, Inc.) according to the manufacturer's protocol. Plasmids were transformed into staphylococci by electroporation using a previously described protocol (13). Restriction enzymes were purchased from Takara Shuzo Co. Ltd. or New England Biolabs and used according to the manufacturer's instructions. Oligonucleotides were obtained from Life Technologies (see Table S1 in the supplemental material). For Southern hybridization, chromosomal DNA was purified, digested, and analyzed by agarose gel electrophoresis. DNA fragments were transferred by alkaline capillary blotting onto nylon membranes (Schleicher and Schuell) using standard methods (5). Labeling of the probe and DNA hybridization were performed according to the protocol supplied with the PCR-DIG DNA-labeling and chemiluminescence detection kit (Boehringer Mannheim). The agr, sarA, and atl genes were inactivated in S. aureus ISP479r by transferring agr::tet, sarA::Km, and atl::LacZ (Erm) from strains ISP479-agr, ISP479-sar (49), and SH108 (17), respectively, by phage transduction using Φ85 (43).
Transposon mutagenesis and gene identification.
The collection of 10,000 transposon mutants of S. aureus 15981 was generated from three independent mutagenesis experiments in a previous work (49). To identify genes disrupted by Tn917 insertions, chromosomal DNA of each mutant was digested with EcoRI and religated in 200 μl of ligation buffer (Promega) for 12 h at 14°C as previously described (41). The ligation products were transformed into E. coli XL1-Blue and selected on LB agar containing ampicillin. Plasmid DNA was extracted using a Bio-Rad plasmid miniprep kit. Chromosomal DNA sequences flanking the transposon were obtained using the primer pseq (see Table S1 in the supplemental material), which corresponds to the inverted repeat region located 70 bp from the Erm-proximal end of Tn917. Nucleotide sequences were determined in an ABI PRISM 310 genetic analyzer (Applied Biosystems). Homology searches were carried out using the BLAST 2.0 program (1) at the NCBI server.
Disruption of TCS operons by insertion of the pBT2 plasmid.
For disruption of TCS1, -3, -4, -5, -11, -14, -16, and -17 in strain 15981 (Table 3), a PCR fragment within the N-terminal regions of the different TCS amplified with primers A and B (see Table S1 in the supplemental material) was cloned in the pGEM-T Easy vector (Promega). The corresponding TCS fragment was then cloned into the EcoRI site of the shuttle vector pBT2, and the resulting plasmid (Table 1) was transformed into S. aureus by electroporation. After electroporation, bacterial strains were incubated for 24 to 48 h at 30°C on TSA with chloramphenicol. Ten milliliters of TSBg-chloramphenicol was subsequently inoculated with a single colony which had been previously resuspended in 100 μl of TSA broth, and the culture was incubated for 24 h at 44.5°C without shaking. Tenfold serial dilutions of this culture in sterile TSBg were plated on TSA-chloramphenicol and incubated for 24 h at 44.5°C. After overnight incubation, the disruption of the respective TCS was confirmed by Southern blot analysis.
TABLE 3.
Schematic representation of all the TCS of S. aureus and the biofilm formation ability of the corresponding mutants in HHWm
Numbers are based on the Staphylococcus aureus N315 sequence (36).
Names of genes are based on S. aureus N315 (36) or according to the first reference where the TCS is described in the text.
Schematic representation of the TCS, including the genes that belong to the same operon. Black arrows represent a response regulator gene, gray arrows represent a histidine kinase sensor protein, and white arrows represent any gene in the operon.
NO, no obtained mutant; Ω, pBT2 insertional mutation; Δ, deletion mutant.
Biofilm formation ability in HHWm in microtiter plates. ND, not determined; +, positive biofilm; −, negative biofilm.
Essential genes.
The TCS is absent in the S. aureus 15981, NCTC8325, COL, MW2, and MSSA476 strains.
Allelic exchange of chromosomal genes.
To construct the deletions of TCS6, -7, -8, -9, -10, -12, -13, and -15 (Table 3), we amplified by PCR two fragments of 500 bp that flanked the sequences to the left (oligonucleotides A and B) and the right (oligonucleotides C and D) of the sequence targeted for deletion (Table S1 in the supplemental material). Oligonucleotide C has at least a 16-base complementary region with oligonucleotide B to allow the products of the first PCR to anneal. A second PCR was performed with primers A and D to obtain a single fragment. Specifically, 1 μl of each of the first PCR products was mixed with 10 pm of the outside primers and PCR amplified. The fusion products were purified and cloned in the pGEM-T Easy vector (Promega). The fragments were then cloned into the EcoRI site of the shuttle plasmid pMAD. To construct the mutant allele to produce the ica deletion, the icaAB and icaCD PCR fragments were cloned separately in the pGEM-T Easy vector (Promega), the plasmids were digested with EcoRI and BamHI to purify both fragments, and finally, the fragments were fused by ligation into plasmid pMAD. The resulting plasmids (Table 1) were transformed into S. aureus by electroporation. pMAD contains a temperature-sensitive origin of replication and an erythromycin resistance gene (3). Homologous recombination experiments were performed as previously described (49). Erythromycin-sensitive white colonies, which no longer contained the pMAD plasmid, were tested by PCR using oligonucleotides A and D to confirm the gene replacement. For the construction of a functional rsbU strain, ISP479r (ISP479 rsbU+), a 3-kb fragment encompassing the complete rsbU-rsbV-rsbW region and part of sigB from plasmid pMAL30 (30), was digested with EcoRI and BamHI and cloned into pMAD. The plasmid was integrated into the chromosome through homology with the parental copy as previously described (49), and the replacement was confirmed by PCR with primers rsbU-3 and rsbU-4 (Table S1 in the supplemental material).
Complementation experiments.
The arlRS operon was amplified with thermophilic DNA polymerase (Certamp long amplification kit; Biotools, Spain) from S. aureus 15981 with primers E and F (Table S1 in the supplemental material). The PCR product was cloned into pCU1 (4), and the resulting plasmid, parlRS, (Table 1) was transformed by electroporation into 15981 ΔarlRS.
Biofilm formation assays.
A biofilm formation assay in microtiter wells was performed as described previously (29). To analyze biofilm formation under flow conditions, we used 60-ml microfermentors (Pasteur Institute) (www.pasteur.fr/recherche/unites/Ggb/biofilmfermenter.html) with a continuous flow of 40 ml of chemically defined medium h−1 and constant aeration with sterile pressed air. Submerged Pyrex slides served as the growth substratum. Bacteria from an overnight culture grown at 37°C with aeration in chemically defined media were diluted to an optical density (OD) at 600 nm of 1. For the inoculation, the Pyrex slides were introduced in the bacterial solution for 2 min and placed back in the microfermentors. The microfermenters were cultivated for 24 h at 37°C. Biofilm development was recorded with a Nikon Coolpix 950 digital camera. To quantify the 24-h biofilm, bacteria that adhered to the surfaces of the Pyrex slides were resuspended in 10 ml chemically defined medium. The OD of the suspension was measured at 650 nm. Biofilm detachment assays with proteases and sodium metaperiodate were carried out as previously described (34) except that 24-well microtiter plates (Sarstedt; catalog no. 1836300) were used and the bacteria that remained attached to the surface were stained with crystal violet, rewashed, and destained with 1 ml of ethanol-acetone (80:20, vol/vol). A total of 200 μl of the ethanol-acetone-dye solution was transferred to a 96-well microtiter plate for spectrophotometric analysis at 595 nm.
Confocal laser scanning microscopy.
Overnight cultures of the 15981 and ΔarlRS strains carrying pSB2019, a plasmid that expresses green fluorescent protein (GFP) (Table 1), grown at 37°C with aeration were diluted 1:40 in a four-well Lab-Tek II sterile-chambered borosilicate coverglass (Nalge Nunc International). After 24 h of incubation at 37°C, the biofilm was directly monitored using a Nikon D-Eclipse C1 confocal microscope with a Plan Apo 60×A/1.40 Oil Dic H objective. Images (512 by 512 pixels) from nine different microscopic fields for each strain were acquired and analyzed with EZ-C1 software (Nikon). The experiment was repeated three times.
PNAG detection.
PNAG production in S. aureus strains was detected as described previously (12). Overnight cultures in chemically defined media were diluted 1:100 and grown for 3, 4.5, 6, 7.5, and 24 h. Two milliliters of each culture was centrifuged, the weights of the pellets were determined, and the pellets were resuspended in a proportional quantity of 0.5 M EDTA (pH 8.0) to obtain the same density of each sample. Fifty microliters of cells was boiled for 5 min and centrifuged to pellet them. Forty microliters of the supernatant was incubated with 10 μl of proteinase K (20 mg/ml; Sigma) for 30 min at 37°C. After the addition of 10 μl of Tris-buffered saline (20 mM Tris-HCl, 150 mM NaCl [pH 7.4]) containing 0.01% bromophenol blue, 5-μl aliquots of 1/2, 1/5, and 1/25 dilutions were spotted on a nitrocellulose filter using a Bio-Dot microfiltration apparatus (Bio-Rad), blocked overnight with 5% skim milk in phosphate-buffered saline with 0.1% Tween 20, and incubated for 2 hours with an anti-S. aureus PNAG antibody diluted 1:20,000 (40). Bound antibodies were detected with a peroxidase-conjugated goat anti-rabbit immunoglobulin G antibody (Jackson ImmunoResearch Laboratories, Inc., PA) diluted 1:10,000 and the Amersham ECL Western blotting system using a Chemi-Doc apparatus (Bio-Rad).
Statistical analysis.
The SPSS 12.0 program was used for the statistical analysis. A two-tailed Student t test was used to determine the differences in biofilm thickness between the wild type and the ΔarlRS mutant as determined by EZ-C1 software and also in the primary attachment assay. A nonparametric test (Mann-Whitney U test) was used to assess significant differences in bacterial density between the wild-type and ΔarlRS strains' recovery from the glass spatula of the microfermentors. Differences were considered statistically significant when P was <0.05 in all cases.
RESULTS
Development of a chemically defined medium to study the S. aureus biofilm formation process.
It has been previously described that the clinical S. aureus 15981 strain is capable of producing strong biofilms in TSBg or B2 medium, indicating that the machinery required for biofilm formation is fully functional in this strain (49). To get further insight into the effect of the growing media on the biofilm development of 15981, we decided to study the biofilm development of this strain in a modified version of the chemically defined medium previously developed by Hussain et al. (31) (Table 2). We named the medium HHWm. Unexpectedly, we found that S. aureus 15981 was unable to produce a biofilm in HHWm, suggesting either that HHWm has nutritional restrictions or that HHWm lacks an activating factor to support biofilm formation or that biofilm formation is repressed in this medium. To answer this question, the biofilm-forming capacity of eight genetically unrelated S. aureus strains in HHWm was assayed in the classical microtiter plate assay. Figure 1 shows that only two strains, 12313 and V329, were able to produce biofilms after 24 h, despite the fact that all selected strains produced biofilms in TSBg (Fig. 1A and B) and exhibited similar growth rates in HHWm (data not shown). These results strongly suggest that the biofilm-negative phenotype of most S. aureus strains in HHWm, including the 15981 strain, is not due to nutritional restrictions.
FIG. 1.
Biofilm formation ability of natural isolates of S. aureus in HHWm. (A) Biofilm formation by S. aureus strains on microtiter plates in HHWm and TSBg after 24 h of incubation at 37°C. The results of a representative experiment are shown. (B) Quantification of the biofilm formation capacity. The bacterial cells were stained with crystal violet and were quantified by dissolving the dye in ethanol-acetone (80:20), and the absorbance was determined at 595 nm. Data represent the averages of results from 12 wells from three different plates. The vertical line at the top of the each bar represents a standard deviation.
Biofilm formation capacity in HHWm of mutants in two-component systems.
Two-component systems are principal signal transducers involved in environmental sensing. We hypothesized that any of the 17 two-component systems identified in the genome of S. aureus N315 might be responsible for the repression of the biofilm formation of strain 15981 in HHWm. The SA0066-SA0067 TCS present in the genome of strain N315 is contained in the SCCmec (the staphylococcal chromosomal cassette involved in methicillin resistance) and, consequently, is not present in the methicillin-susceptible NCTC8325, COL, MW2, or MSSA467 strain. Clinical strain 15981 is methicillin susceptible, and the absence of the SA0066-SA0067 TCS was verified by PCR and Southern blot analysis. Thus, we generated isogenic derivatives of 15 out of the 16 TCS of S. aureus either by insertional inactivation or gene replacement by allelic exchange in strain 15981 (see Materials and Methods) (Table 3). We were unable to generate strains with mutations in yycFG, which has been shown to be essential for the viability of S. aureus (39). All the resulting TCS mutants exhibited growth rates in HHWm that were indistinguishable from those of the wild type. The analysis of the biofilm formation capacity of the TCS mutants in HHWm revealed that deletion of arlRS provoked an increase in biofilm development (Fig. 2A). These results support our hypothesis that the biofilm formation phenotype is repressed in HHWm and that at least one TCS is involved in the process.
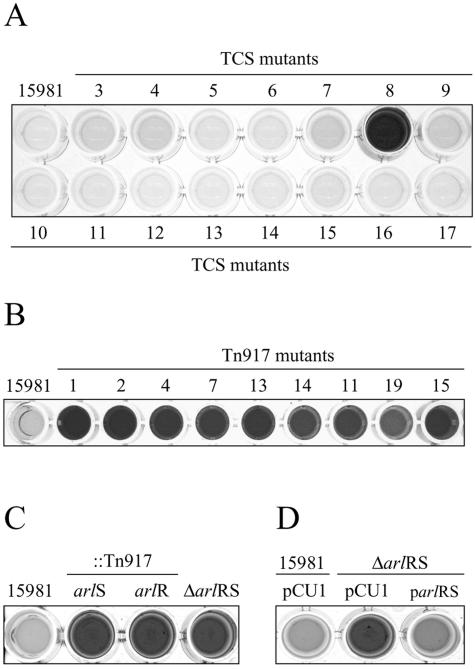
FIG. 2.
Biofilm formation phenotype of biofilm-positive mutants on microtiter plates in HHWm. (A) Biofilm formation phenotype of wild-type 15981 and isogenic mutants in all the TCS on microtiter plates grown in HHWm at 37°C for 24 h (Table 3). (B) Biofilm formation phenotype of wild-type and selected biofilm-positive mutants on microtiter plates grown in HHWm at 37°C for 24 h. Tn917 was inserted in different positions of the arlS gene in mutants MM4, -7, -13, and -14 and two different positions of the arlR gene in MM11 and MM19. (C) Biofilm formation phenotype of the ΔarlRS mutant compared to the wild-type strain and insertional transposon mutants. (D) Biofilm formation phenotype of the wild-type and ΔarlRS mutant strains complemented with either the parlRS plasmid carrying the arlRS operon under the control of its own promoter or the pCU1 plasmid. HHWm was supplemented with 20 μg/ml of chloramphenicol. The results of a representative experiment are shown.
Random screening for repressors of the biofilm formation process of S. aureus 15981 in HHWm.
A second approach was used to identify repressors of the biofilm formation process of S. aureus 15981 in HHWm, based on the screening of a collection of strains with approximately 10,000 transposon mutations (49) to look for mutants that had gained the capacity to produce a biofilm on microtiter polystyrene plates in HHWm. Nine transposon mutants, exhibiting growth rates indistinguishable from the wild type, were found to acquire biofilm formation capacity in HHWm (Fig. 2B). Sequencing of regions adjacent to the transposon insertion site and homology searches using the BLAST 2.0 program at the NCBI server showed that, in six out of nine mutants, the transposon insertion site was localized in four different positions of the arlS gene (MM4, -7, -13, and -14) and in two different positions of the arlR gene (MM11 and MM19). The arlR/arlS transposon mutants, akin to the ΔarlRS strain, showed an increase in their biofilm formation capacity in HHWm (Fig. 2C). The loci interrupted by the transposon in the remaining three mutants were lysA (MM1), odhA (MM15), and the intergenic region SA1243-SA1244 (MM2). The role of these genes in the repression of biofilm development in HHWm is under study.
Although the finding that both approaches coincided in the arlRS two-component system is strong evidence to conclude that arlRS negatively affects biofilm formation in HHWm, we complemented the ΔarlRS strain with a plasmid carrying a copy of the arlRS operon, including its own promoter region (see Materials and Methods for details). This plasmid (parlRS) and the vector control (pCU1) were introduced into the wild-type and arlRS mutant strains, and their biofilm formation capacity was analyzed. The results of the complementation analysis are shown in Fig. 2D. When parlRS was introduced into the arlRS mutants, inhibition of biofilm formation was observed, whereas the vector control (pCU1) had no effect on biofilm formation in either the wild type or the arlRS mutant.
arlRS is involved in an early stage of biofilm development.
An initial adhesion defect could have been the underlying cause of the wild-type strain's failure to form a biofilm in HHWm. Therefore, we tested whether 15981 ΔarlRS adhered to polystyrene petri dishes under static conditions more efficiently than the corresponding wild-type strain. Quantification of adherent bacterial cells on a phase-contrast microscope showed that the numbers of attached cells were significantly higher in the case of the ΔarlRS strains than that of the wild-type strain (n = 30; P < 0.001, Student's t test) (data not shown). This result indicates that genes regulated by arlRS are at least involved in the early attachment of S. aureus to abiotic surfaces.
Biofilm formation of the arlRS mutant under flow cell conditions.
Since no fresh medium is provided during biofilm development on a microtiter plate, nutrient concentration declines and the concentration of waste products increases throughout the incubation period. Since any or both of these factors could represent the input signal for the activation of the arlRS two-component system and the ensuing inhibition of biofilm development in HHWm, we used microfermentors where nutrients were continually replenished to compare the biofilm development of wild-type and arlRS mutant strains. Both macroscopic biofilm development in microfermentors and biofilm cell density on the removable glass slides of the fermentors were examined. The data in Fig. 3 show that 15981 ΔarlRS displayed a robust biofilm phenotype. It is worth noting that wild-type bacteria displayed substantial biofilm development in the microfermentors in comparison to that of the phenotype shown on microtiter plates, suggesting that limited aeration and/or nutrient depletion and/or accumulation of waste products during steady-state incubation conditions also have a negative effect on strain 15981's biofilm development. However, the fact that the absence of arlRS still enhanced biofilm formation in microfermentors (n = 3; P <0.05) suggests that arlRS-mediated repression of biofilm development is also effective under these conditions.
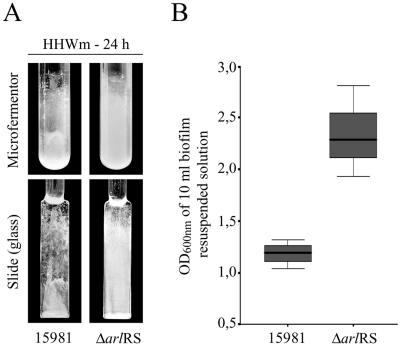
FIG. 3.
Biofilm formation phenotype of the wild-type and ΔarlRS strains in microfermentors. (A) Biofilm development of bacteria grown under continuous-flow conditions with HHWm after 24 h at 37°C. The microfermentors (upper panels) contain the glass slides where bacteria form the biofilm (lower panels). The results of a representative experiment are shown. (B) Quantification of the biofilms adhered to the glass slides. The cells were removed from the glass slides into 10 ml of HHWm by using a vortex, and the OD of the resulting solution was measured at 650 nm. Significant differences were detected between the wild-type and the ΔarlRS mutant (n = 3; P < 0.05, Mann-Whitney U test).
Biofilm architecture of arlRS mutants.
Biofilms formed by the 15981 and ΔarlRS strains in HHWm were examined by confocal laser scanning microscopy. Cells expressing the GFP were incubated overnight at 37°C as a static culture in a four-well Lab-Tek II chamber slide system (Nalge Nunc International). Since the biofilms produced by arlRS mutants did not tightly attach to the polystyrene surface and were easily disrupted when rinsed, bacterial structures attached to the coverslip were directly observed by confocal laser scanning microscopy (Fig. 4A). The biofilm formed by the ΔarlRS strain appeared to consist of microcolonies of bacterial cells separated by interstitial voids (medium-filled channels). This was in contrast to what occurred with the wild-type strain, which covered the surface uniformly and lacked a three-dimensional architecture, strongly suggesting that these bacteria correspond to the normal sedimentation process that occurred during the incubation period at the bottom of the chamber. We used the EZ-C1 software to quantify the thickness of the biofilm. In agreement with the microtiter dish assays and the visual impression, the value for total thickness was significantly increased (n = 27; P < 0.0001) in the case of the arlRS mutant strain compared with that of the wild-type strain (Fig. 4B).
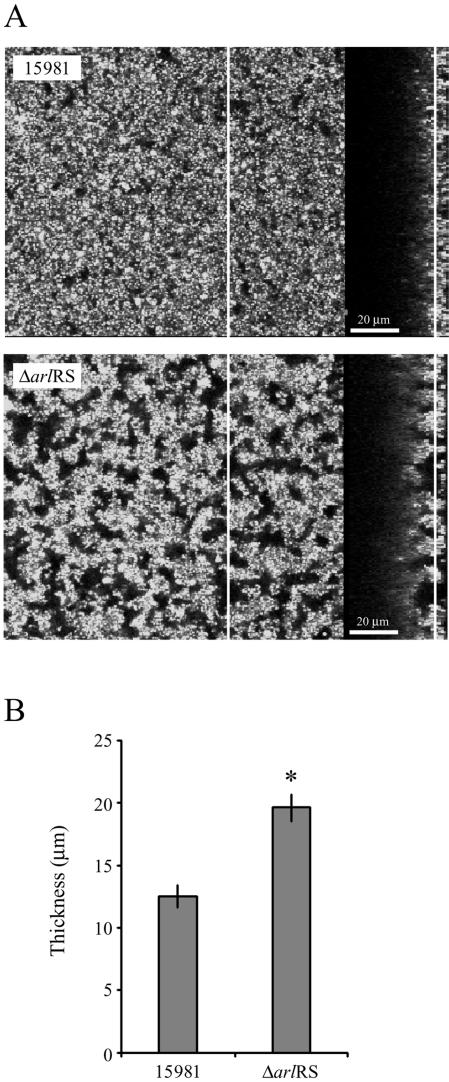
FIG. 4.
Effect of an arlRS mutation on biofilm architecture. S. aureus 15981 and 15981 ΔarlRS strains harboring plasmid pSB2019 to provide constitutive GFP expression were grown in a four-well Lab-Tek II chamber with HHWm at 37°C for 24 h. (A) Biofilm development was monitored by scanning confocal laser microscope. The panels on the left are an overlay of multiple slices, and the side frames of the panels on the right are the z planes, which show the thickness of the biofilm and the architecture as viewed from the cross section. The line on the z plane indicates the level at which the photograph of the x-y plane was taken. Photographs were taken at a magnification of ×600. The results of a representative experiment are shown. (B) Quantification of the biofilm thickness using EZ-C1 software. Data represent the averages of results from 27 different fields from three independent experiments. The vertical line at the top of each bar represents the standard deviation. The asterisk represents a statistically significant difference (P < 0.0001, Student's t test).
PNAG is not required for the biofilm development promoted by the arlRS mutation.
It has been shown that PNAG production is essential for the biofilm development of S. aureus. To investigate whether PNAG production was altered in 15981 ΔarlRS, the level of PNAG was monitored by dot blotting using specific anti-PNAG polyclonal antisera. The results showed that PNAG production in 15981 ΔarlRS was enhanced at all phases of growth in HHWm compared to the PNAG production of the parental strain (Fig. 5A). The fact that 15981 produced detectable amounts of PNAG indicates that arlRS is relevant but not essential for PNAG production. These results suggest that, in addition to a higher level of primary attachment, the enhancement in biofilm formation presented by the ΔarlRS mutant may be attributable to an increase in PNAG levels.
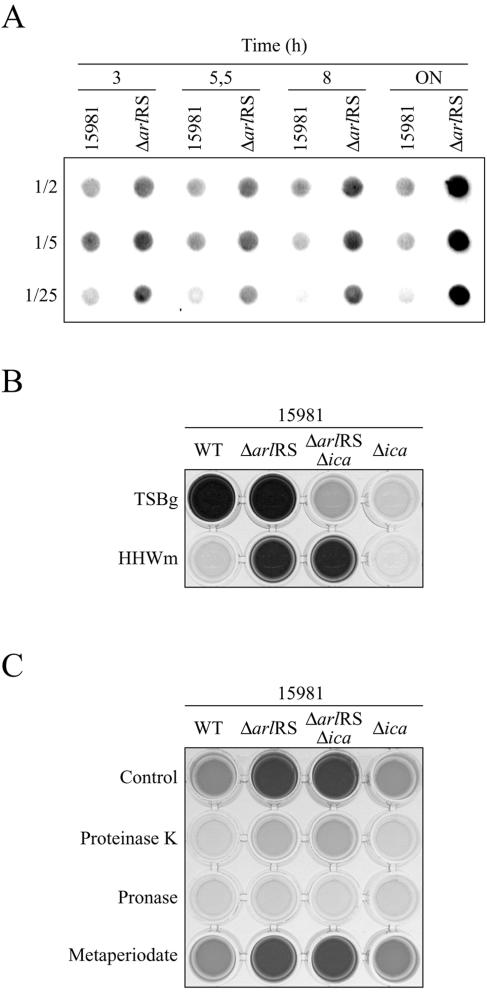
FIG. 5.
(A) Dot blot analysis of PNAG accumulation in the wild-type strain and its corresponding ΔarlRS mutant at different points of the growth curve. Cell surface extracts at different times were treated with proteinase K, and dilutions of the extracts (1/2, 1/5, and 1/25) were spotted onto nitrocellulose membranes. PNAG production was detected with an anti-S. aureus PNAG antiserum. The ΔarlRS mutant produced higher levels of PNAG product at all points of the growth curve. The results of a representative experiment are shown. (B) Biofilm formation phenotype of the 15981, ΔarlRS, Δica, and ΔarlRS Δica strains grown in TSBg and HHWm on microtiter plates. After 24 h of incubation, the microplates were washed and stained with crystal violet. The dye was dissolved by the addition of 200 μl of ethanol-acetone (80:20). The results of a representative experiment are shown. (C) Biofilm detachment assays. Biofilms of the 15981, ΔarlRS, Δica, and ΔarlRS Δica strains grown in HHWm for 16 h were treated for 2 h at 37°C with 10 mM sodium metaperiodate or with 100 μg of proteinase K or pronase. The bacteria that remained attached to the surface were stained with crystal violet. The dye was dissolved by the addition of 200 μl of ethanol-acetone (80:20). The results of a representative experiment are shown. ON, overnight; WT, wild type.
To unambiguously demonstrate that enhanced accumulation of PNAG was directly related to the enhanced biofilm development of arlRS mutants, we produced a nonpolar deletion of the icaADBC operon in the arlRS mutant. Surprisingly, the resulting double arlRS icaADBC mutant retained the capacity to produce biofilm in HHWm, whereas the ability to produce biofilm in TSBg was completely abolished (Fig. 5B). These results indicated that PNAG, in contrast to its essential role in the biofilm matrix produced in TSBg, is dispensable for the biofilm promoted by the arlRS mutation in HHWm.
To determine whether the biofilm produced by the arlRS mutant was mediated by another exopolysaccharide or a surface adhesin, biofilm detachment experiments were carried out with proteinase K, pronase, and sodium metaperiodate (a carbohydrate-modifying agent). The results showed that the biofilm induced by the ΔarlRS strain was sensitive to detachment by proteases and resistant to sodium metaperiodate (Fig. 5C). These results are consistent with our finding that PNAG is not required for the arlRS mutant biofilm formation process and strongly suggest that a proteinaceous compound might be responsible for biofilm development in the arlRS mutant.
SarA is essential for the biofilm development promoted by the arlRS mutation.
Previous studies have described that the arl mutation decreased sarA transcription and increased the synthesis of RNA III (19). In addition, others and we have found that SarA is essential for the biofilm development of S. aureus (6, 49) and S. epidermidis (48). To investigate the role of SarA in the biofilm produced by the arlRS mutant, the sarA gene was deleted in the arlRS strain. Interestingly, the double arlRS sarA mutant lost the capacity to produce biofilm in both HHWm and TSBg, indicating that SarA is epistatic to the arlRS mutation in that the double mutant resembled the sarA single mutant with respect to its inability to form biofilm (Fig. 6A). To confirm the independence of the biofilm-deficient phenotype with respect to the strain background, we also produced nonpolar mutations of arlRS, sarA, and arlRS sarA in strain ISP479r, a derivative of 8325-4 in which the 11-nucleotide deletion of the rsbU gene has been restored by allelic exchange. Consistent with the results observed in strain 15981, the mutation of arlRS in strain ISP479r promoted biofilm development in HHWm (Fig. 6A), whereas the arlRS sarA double mutant displayed a phenotype indistinguishable from that of the wild-type strain.
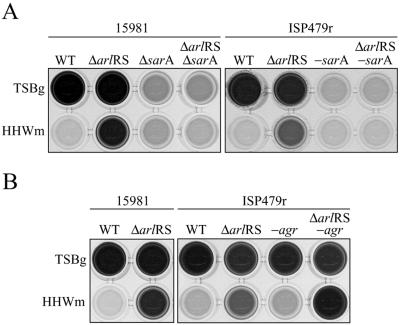
FIG. 6.
Effects of the mutation of sarA and agr regulators on the biofilm promoted by an arlRS mutation. Bacteria were inoculated onto microtiter plates containing TSBg and HHWm. After 24 h of incubation, the microplates were washed and stained with crystal violet. The dye was dissolved in 200 μl of ethanol-acetone (80:20). The results of a representative experiment are shown. (A) Comparison of biofilm formations by S. aureus 15981 and ISP479r and their corresponding ΔarlRS, ΔsarA, and ΔarlRS ΔsarA mutants in HHWm and TSBg. (B) Comparison of biofilm formations by S. aureus strains 15981, 15981 ΔarlRS, ISP479r, and ISP479r's corresponding mutant derivatives ISP479r ΔarlRS, ISP479r-agr, and ISP479r-agr ΔarlRS in HHWm and TSBg. WT, wild type.
Several studies with contradictory results have related the agr quorum-sensing system with the biofilm formation process of S. aureus. The discrepancies of the results might be explained by the possibility that the role of agr expression in biofilm development is dependent upon the conditions under which the biofilm was grown (52). To determine whether agr was involved in the biofilm produced by the arlRS mutant in HHWm, we produced the deletion of agr and arlRS agr in S. aureus ISP479r. We used this strain because during the course of this study, we noticed that the clinical strain 15981 has a natural mutation of the agr system (M. Otto, personal communication). The ISP479r-agr strain, which would be equivalent to the 15981 strain, displayed a biofilm formation phenotype indistinguishable from that of the wild-type strain ISP479r in HHWm and TSBg (Fig. 6B). Interestingly, disruption of arlRS in the agr mutant reproducibly resulted in increased biofilm development in comparison to that of the single arlRS mutant. These results suggest that agr mutation alone has no effect on the biofilm formation capacity of S. aureus in HHWm. However, since the combination of agr and arlRS mutations reinforced biofilm development in HHWm, the results also suggest that both arlRS and agr play a negative role in biofilm development in HHWm.
Increased autolysin activity is not responsible for biofilm development in HHWm.
It has been described previously that an arlS mutant exhibited increased autolysis and altered peptidoglycan hydrolase activity (18). To rule out the possibility that the increased biofilm development of the arlRS mutant in HHWm was due to an increased autolysin activity, the atl gene was deleted in wild-type ISP479r and the corresponding ISP479r ΔarlRS, ISP479r-agr, and ISP479r agr ΔarlRS strains. No differences between the wild-type and the corresponding mutants were observed (Fig. 7), indicating that the activity of the major autolysin was not responsible for the biofilm enhancement of the arlRS mutant.
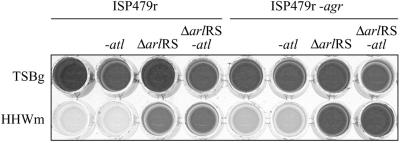
FIG. 7.
Effects of the mutation of atl on the biofilm promoted by an arlRS mutation. Comparison of biofilm formations by strain ISP479r and its corresponding ISP479r ΔarlRS, ISP479r-atl, and ISP479r ΔarlRS-atl mutant derivatives and by the same mutants constructed in the ISP479r agr null background. Bacteria were inoculated onto microtiter plates containing TSBg and HHWm. After 24 h of incubation, the microplates were washed and stained with crystal violet. The dye was dissolved in 200 μl of ethanol-acetone (80:20). The results of a representative experiment are shown.
DISCUSSION
S. aureus is a ubiquitous bacterium, able to exist on inanimate objects as well as in a variety of niches of the human or animal host, such as the skin, mucous membranes, blood, bones, and mammary glands. These locations are not hermetic, and during its lifetime, a single bacterium can travel from one location to another. These transitions are very likely to require quick switches from a unicellular planktonic existence to the settled multicellular communities commonly referred to as biofilms. This versatility depends on a complex regulatory network responsible for sensing and responding to the environmental conditions that favor biofilm formation or the release of bacteria from the biofilm to colonize new niches.
In this study, we present evidence that the two-component regulatory system arlRS negatively affects the biofilm formation of S. aureus. These results are consistent with a previous study showing that arlS mutation induced biofilm formation on a polystyrene surface in a complex TSB medium (18). In the same report, it was shown that biofilm formation was associated with an enhanced primary attachment probably owing to an increased autolysin activity. However, in contrast to this hypothesis, our results indicate that autolysin activity is not responsible for the biofilm enhancement observed in an arlRS mutant, because a mutation in the atl gene, which encodes the main autolysin of S. aureus, did not restore the phenotypes of wild-type bacteria.
Apart from Atl, another candidate that we needed to take into consideration in relation to the biofilm development of S. aureus was the exopolysaccharide PNAG. It has been demonstrated that disruption of the icaADBC operon responsible for the synthesis of PNAG impairs the biofilm development of most of the S. aureus strains analyzed to date (21, 49). Based on that demonstration, PNAG has been considered an essential compound of the biofilm matrix of S. aureus. On the contrary, the results of the present study indicate that arlRS regulates a mechanism of biofilm development independent of PNAG accumulation. This alternative mechanism is being supported by an increasing amount of evidence (7, 14). First, in an earlier work, we showed that disruption of the ica operon in a bovine mastitis bap-positive strain had no effect on in vitro biofilm formation, a finding which strongly suggested that the Bap protein could compensate for the deficiency of PNAG (14). It is important to note that this bovine strain, V329, is one of the strains capable of producing biofilm in HHWm (see Fig. 1). Second, Beenken et al. also demonstrated that deletion of the icaADBC operon in the clinical S. aureus UAMS-1 strain had little impact on biofilm formation both in vivo and in vitro (7), though the mechanism for biofilm formation of this strain was not determined. In an attempt to explain the PNAG-independent mechanism shown in the current study, we established that it was not dependent on the presence of Bap, since we were not able to detect any Bap-homologous protein in the 15981 or ISP479 strain (data not shown). Fournier et al. determined that one of the most remarkable consequences of the arlS mutation was the dramatic decrease in extracellular proteolytic activity (18). Accordingly, and given that surface proteins could initiate primary attachment to the surface or participate in intercellular adhesion, it is likely that the absence of extracellular proteases in 15981 ΔarlRS and ISP479r ΔarlRS could facilitate the accumulation of one or more proteins that would mediate the attachment to surfaces. In this scenario, disruption of sarA in such mutants abolished biofilm development. This could be the result of the restoration of the secretion of the main extracellular proteases and promotion of bacterial dissemination by cleavage of bacterial attachment proteins. Another consequence of the absence of extracellular proteases in arlRS mutants might be the preservation of Ica proteins from degradation, which would allow the accumulation of PNAG. Our data indicated that the arlRS mutant accumulated higher levels of PNAG than the wild-type strain, whereas the transcriptional levels of the icaADBC operon in the mutant were similar to or were up-regulated at the postexponential phase of growth in comparison to the corresponding levels of the wild-type strain (our unpublished results). A similar discrepancy between the accumulation of polysaccharide intercellular adhesin/PNAG and the transcription of the icaADBC operon has also been described for S. epidermidis growing in the presence of glucose (15). This is due to the fact that under these conditions, S. epidermidis exhibited polysaccharide intercellular adhesin and a biofilm-positive phenotype, whereas ica transcription was down-regulated. Altogether, it seems as though as soon as enough PNAG is accumulated, bacteria may repress the synthesis of Ica proteins by some kind of negative-feedback regulatory mechanism. Besides the effect of arl on the accumulation of extracellular proteases, we cannot rule out the possibility that arlRS could directly repress the expression of an adhesin or some other effector protein that is required for biofilm formation. Alternatively, arlRS could also activate the expression of a factor whose presence could be deleterious for biofilm formation.
In this study, we have also performed the systematic disruption of 15 two-component regulatory systems in the same strain of S. aureus. yycGF was the only TCS in which we were unable to generate an insertional mutation, confirming that this TCS is essential for S. aureus (16, 39). The remnant TCS were not essential for the survival of S. aureus, and examination of the growth curves of TCS mutants in HHWm did not show significant differences among them. It is noteworthy that we have unintentionally constructed all our isogenic mutants in the two-component systems in an agr null genetic background, which occurs quite frequently in natural isolates of S. aureus (51). Since the functionality of 9 out of 16 of the TCS of S. aureus is completely unknown, it is impossible to predict how the absence of a functional agr gene might affect their expression. The finding that a single mutation of agr has no effect on the biofilm formation of strains 15981 and ISP479r indicates that agr is not responsible for the biofilm enhancement of the arlRS mutant. The strength of the biofilm observed in the double mutant ISP479r arlRS agr could be due to the detergent-like physicochemical properties of the delta-hemolysin (50) whose gene is carried within the gene for RNA III, and its expression is therefore directly linked to agr activity (32). Nevertheless, the roles of agr and quorum sensing in biofilm formation remain elusive. Recently, Yarwood et al. observed that the influence of agr on biofilm formation ranged from an enhanced effect or no discernible effect to impaired biofilm development, depending on the conditions of the experiments (52). They interpreted these results as indicating that the wide range of genes regulated by agr could influence S. aureus attachment and biofilm development under some conditions but not others.
Recently, in an effort to identify the potential function of each TCS in the biology of a clinical isolate of Enterococcus faecalis, systematic inactivation of 17 out of 18 TCS was carried out (26). Only one of the TCS, fsrABC, displayed a deficiency in biofilm formation (25). The fsrABC locus is a quorum-sensing system that belongs to the so-called agr-like subfamily of two-component signal transduction systems characterized by the presence of a third component, a secreted signaling peptide. Hancock and Perego (25) concluded that disruption of fsr provokes a deficiency in a zinc-metalloprotease gelatinase which is required for the formation of biofilm by a still-unknown mechanism. None of the TCS mutants we constructed showed any deficiency on biofilm formation in rich media, and apparently there is no resemblance between the role of fsr in E. faecalis and the role of the corresponding TCS of S. aureus in the regulation of biofilm development, suggesting the existence of a species-specific biofilm formation process in gram-positive bacteria.
While there is little doubt that there exist other S. aureus genes critical for biofilm development yet to be discovered, our results suggest that the regulatory network for biofilm formation is more complex than first anticipated and includes positive and negative regulators. In this regard, this work also indicates that screening for mutants that gain biofilm formation capacity in microtiter dishes could be useful for the identification of those genes that have negative consequences in the bacterial-biofilm formation process.
Supplementary Material
Acknowledgments
We express our gratitude to T. Maira-Litrán for providing us with the anti-PNAG antiserum, P. Hill for plasmid pSB2019, S. Foster for plasmid pMAL30 and strain SH108, B. Fournier for strains BF16 and ISP794, and Joelle Bignon for technical assistance in the establishment of the HHWm. We thank Michael Otto for determining that strain 15981 is agr negative and C. Solano and J. M. Ghigo for critical reading of the manuscript. We also thank the reviewers of the manuscript for their helpful comments.
Alejandro Toledo-Arana and Marta Vergara-Irigaray are predoctoral fellows (FPU) from the Ministerio de Educación y Ciencia, Spain. Nekane Merino is a predoctoral fellow from the Basque Government, Spain. This work was supported by the BIO2002-04542-C02 and GEN2003-20234-C06-05 grants from the Comisión Interministerial de Ciencia y Tecnología and grant 17/2004 from the Departamento de Educación y Cultura del Gobierno de Navarra, Spain.
Footnotes
Supplemental material for this article may be found at http://jb.asm.org/.
REFERENCES
- 1.Altschul, S. F., T. L. Madden, A. A. Schaffer, J. Zhang, Z. Zhang, W. Miller, and D. J. Lipman. 1997. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25:3389-3402. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Amorena, B., E. Gracia, M. Monzón, J. Leiva, C. Oteiza, M. Pérez, J. Alabart, and J. Hernandez-Yago. 1999. Antibiotic susceptibility of Staphylococcus aureus in biofilms developed in vitro. J. Antimicrob. Chemother. 44:43-55. [DOI] [PubMed] [Google Scholar]
- 3.Arnaud, M., A. Chastanet, and M. Débarbouillé. 2004. New vector for efficient allelic replacement in naturally nontransformable, low-GC-content gram-positive bacteria. Appl. Environ. Microbiol. 70:6887-6891. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Augustin, J., R. Rosenstein, B. Wieland, U. Schneider, N. Schnell, G. Engelke, K. D. Entian, and F. Gotz. 1992. Genetic analysis of epidermin biosynthetic genes and epidermin-negative mutants of Staphylococcus epidermidis. Eur. J. Biochem. 204:1149-1154. [DOI] [PubMed] [Google Scholar]
- 5.Ausubel, F. M., R. Brent, R. E. Kingston, D. D. Moore, J. G. Seidman, J. A. Smith, and K. Struhl (ed.). 1990. Current protocols in molecular biology. John Wiley & Sons, New York, N.Y.
- 6.Beenken, K. E., J. S. Blevins, and M. S. Smeltzer. 2003. Mutation of sarA in Staphylococcus aureus limits biofilm formation. Infect. Immun. 71:4206-4211. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Beenken, K. E., P. M. Dunman, F. McAleese, D. Macapagal, E. Murphy, S. J. Projan, J. S. Blevins, and M. S. Smeltzer. 2004. Global gene expression in Staphylococcus aureus biofilms. J. Bacteriol. 186:4665-4684. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Bruckner, R. 1997. Gene replacement in Staphylococcus carnosus and Staphylococcus xylosus. FEMS Microbiol. Lett. 151:1-8. [DOI] [PubMed] [Google Scholar]
- 9.Brunskill, E. W., and K. W. Bayles. 1996. Identification and molecular characterization of a putative regulatory locus that affects autolysis in Staphylococcus aureus. J. Bacteriol. 178:611-618. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Conlon, K. M., H. Humphreys, and J. P. O'Gara. 2002. icaR encodes a transcriptional repressor involved in environmental regulation of ica operon expression and biofilm formation in Staphylococcus epidermidis. J. Bacteriol. 184:4400-4408. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Costerton, J. W., and P. S. Stewart. 2000. Biofilms and device-related infections, p. 423-439. In J. P. Nataro, M. J. Blaser, and S. Cunningham-Rundles (ed.), Persistent bacterial infections. American Society for Microbiology, Washington, D.C.
- 12.Cramton, S. E., C. Gerke, N. F. Schnell, W. W. Nichols, and F. Götz. 1999. The intercellular adhesion (ica) locus is present in Staphylococcus aureus and is required for biofilm formation. Infect. Immun. 67:5427-5433. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Cucarella, C., C. Solano, J. Valle, B. Amorena, Í. Lasa, and J. R. Penadés. 2001. Bap, a Staphylococcus aureus surface protein involved in biofilm formation. J. Bacteriol. 183:2888-2896. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Cucarella, C., M. A. Tormo, C. Úbeda, M. P. Trotonda, M. Monzón, C. Peris, B. Amorena, Í. Lasa, and J. R. Penadés. 2004. Role of biofilm-associated protein Bap in the pathogenesis of bovine Staphylococcus aureus. Infect. Immun. 72:2177-2185. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Dobinsky, S., K. Kiel, H. Rohde, K. Bartscht, J. K.-M. Knobloch, M. A. Horstkotte, and D. Mack. 2003. Glucose-related dissociation between icaADBC transcription and biofilm expression by Staphylococcus epidermidis: evidence for an additional factor required for polysaccharide intercellular adhesin synthesis. J. Bacteriol. 185:2879-2886. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Dubrac, S., and T. Msadek. 2004. Identification of genes controlled by the essential YycG/YycF two-component system of Staphylococcus aureus. J. Bacteriol. 186:1175-1181. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Foster, S. J. 1995. Molecular characterization and functional analysis of the major autolysin of Staphylococcus aureus 8325/4. J. Bacteriol. 177:5723-5725. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Fournier, B., and D. C. Hooper. 2000. A new two-component regulatory system involved in adhesion, autolysis, and extracellular proteolytic activity of Staphylococcus aureus. J. Bacteriol. 182:3955-3964. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Fournier, B., A. Klier, and G. Rapoport. 2001. The two-component system ArlS-ArlR is a regulator of virulence gene expression in Staphylococcus aureus. Mol. Microbiol. 41:247-261. [DOI] [PubMed] [Google Scholar]
- 20.Giraudo, A. T., C. G. Raspanti, A. Calzolari, and R. Nagel. 1994. Characterization of a Tn551-mutant of Staphylococcus aureus defective in the production of several exoproteins. Can. J. Microbiol. 40:677-681. [DOI] [PubMed] [Google Scholar]
- 21.Gotz, F. 2002. Staphylococcus and biofilms. Mol. Microbiol. 43:1367-1378. [DOI] [PubMed] [Google Scholar]
- 22.Götz, F., and G. Peters. 2000. Colonization of medical devices by coagulase-negative staphylococci, p. 55-88. In F. A. Waldvogel and A. L. Bisno (ed.), Infections associated with indwelling medical devices. ASM Press, Washington, D.C.
- 23.Gracia, E., A. Lacleriga, M. Monzon, J. Leiva, C. Oteiza, and B. Amorena. 1998. Application of a rat osteomyelitis model to compare in vivo and in vitro the antibiotic efficacy against bacteria with high capacity to form biofilms. J. Surg. Res. 79:146-153. [DOI] [PubMed] [Google Scholar]
- 24.Gross, M., S. E. Cramton, F. Götz, and A. Peschel. 2001. Key role of teichoic acid net charge in Staphylococcus aureus colonization of artificial surfaces. Infect. Immun. 69:3423-3426. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Hancock, L. E., and M. Perego. 2004. The Enterococcus faecalis fsr two-component system controls biofilm development through production of gelatinase. J. Bacteriol. 186:5629-5639. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Hancock, L. E., and M. Perego. 2004. Systematic inactivation and phenotypic characterization of two-component signal transduction systems of Enterococcus faecalis V583. J. Bacteriol. 186:7951-7958. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Heilmann, C., C. Gerke, F. Perdreau-Remington, and F. Götz. 1996. Characterization of Tn917 insertion mutants of Staphylococcus epidermidis affected in biofilm formation. Infect. Immun. 64:277-282. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Heilmann, C., M. Hussain, G. Peters, and F. Gotz. 1997. Evidence for autolysin-mediated primary attachment of Staphylococcus epidermidis to a polystyrene surface. Mol. Microbiol. 24:1013-1024. [DOI] [PubMed] [Google Scholar]
- 29.Heilmann, C., O. Schweitzer, C. Gerke, N. Vanittanakom, D. Mack, and F. Gotz. 1996. Molecular basis of intercellular adhesion in the biofilm-forming Staphylococcus epidermidis. Mol. Microbiol. 20:1083-1091. [DOI] [PubMed] [Google Scholar]
- 30.Horsburgh, M. J., J. L. Aish, I. J. White, L. Shaw, J. K. Lithgow, and S. J. Foster. 2002. σB modulates virulence determinant expression and stress resistance: characterization of a functional rsbU strain derived from Staphylococcus aureus 8325-4. J. Bacteriol. 184:5457-5467. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Hussain, M., J. G. Hastings, and P. J. White. 1991. A chemically defined medium for slime production by coagulase-negative staphylococci. J. Med. Microbiol. 34:143-147. [DOI] [PubMed] [Google Scholar]
- 32.Janzon, L., S. Lofdahl, and S. Arvidson. 1989. Identification and nucleotide sequence of the delta-lysin gene, hld, adjacent to the accessory gene regulator (agr) of Staphylococcus aureus. Mol. Gen. Genet. 219:480-485. [DOI] [PubMed] [Google Scholar]
- 33.Jefferson, K. K., D. B. Pier, D. A. Goldmann, and G. B. Pier. 2004. The teicoplanin-associated locus regulator (TcaR) and the intercellular adhesin locus regulator (IcaR) are transcriptional inhibitors of the ica locus in Staphylococcus aureus. J. Bacteriol. 186:2449-2456. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Kaplan, J. B., K. Velliyagounder, C. Ragunath, H. Rohde, D. Mack, J. K.-M. Knobloch, and N. Ramasubbu. 2004. Genes involved in the synthesis and degradation of matrix polysaccharide in Actinobacillus actinomycetemcomitans and Actinobacillus pleuropneumoniae biofilms. J. Bacteriol. 186:8213-8220. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Kuroda, M., K. Kuwahara-Arai, and K. Hiramatsu. 2000. Identification of the up- and down-regulated genes in vancomycin-resistant Staphylococcus aureus strains Mu3 and Mu50 by cDNA differential hybridization method. Biochem. Biophys. Res. Commun. 269:485-490. [DOI] [PubMed] [Google Scholar]
- 36.Kuroda, M., T. Ohta, I. Uchiyama, T. Baba, H. Yuzawa, I. Kobayashi, L. Cui, A. Oguchi, K. Aoki, Y. Nagai, J. Lian, T. Ito, M. Kanamori, H. Matsumaru, A. Maruyama, H. Murakami, A. Hosoyama, Y. Mizutani-Ui, N. K. Takahashi, T. Sawano, R. Inoue, C. Kaito, K. Sekimizu, H. Hirakawa, S. Kuhara, S. Goto, J. Yabuzaki, M. Kanehisa, A. Yamashita, K. Oshima, K. Furuya, C. Yoshino, T. Shiba, M. Hattori, N. Ogasawara, H. Hayashi, and K. Hiramatsu. 2001. Whole genome sequencing of methicillin-resistant Staphylococcus aureus. Lancet 357:1225-1240. [DOI] [PubMed] [Google Scholar]
- 37.Mack, D., M. Nedelmann, A. Krokotsch, A. Schwarzkopf, J. Heesemann, and R. Laufs. 1994. Characterization of transposon mutants of biofilm-producing Staphylococcus epidermidis impaired in the accumulative phase of biofilm production: genetic identification of a hexosamine-containing polysaccharide intercellular adhesin. Infect. Immun. 62:3244-3253. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Mack, D., H. Rohde, S. Dobinsky, J. Riedewald, M. Nedelmann, J. K.-M. Knobloch, H.-A. Elsner, and H. H. Feucht. 2000. Identification of three essential regulatory gene loci governing expression of Staphylococcus epidermidis polysaccharide intercellular adhesin and biofilm formation. Infect. Immun. 68:3799-3807. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Martin, P. K., T. Li, D. Sun, D. P. Biek, and M. B. Schmid. 1999. Role in cell permeability of an essential two-component system in Staphylococcus aureus. J. Bacteriol. 181:3666-3673. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Mckenney, D., K. Pouliot, V. Wang, V. Murthy, M. Ulrich, G. Döring, J. C. Lee, D. A. Goldmann, and G. B. Pier. 1999. Broadly protective vaccine for Staphylococcus aureus based on an in vivo-expressed antigen. Science 284:1523-1527. [DOI] [PubMed] [Google Scholar]
- 41.Mei, J. M., F. Nourbakhsh, C. W. Ford, and D. W. Holden. 1997. Identification of Staphylococcus aureus virulence genes in a murine model of bacteraemia using signature-tagged mutagenesis. Mol. Microbiol. 26:399-407. [DOI] [PubMed] [Google Scholar]
- 42.Muller, E., J. Hübner, N. Gutierrez, S. Takeda, D. A. Goldmann, and G. B. Pier. 1993. Isolation and characterization of transposon mutants of Staphylococcus epidermidis deficient in capsular polysaccharide/adhesin and slime. Infect. Immun. 61:551-558. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Novick, R. P. 1991. Genetic systems in staphylococci. Methods Enzymol. 204:587-636. [DOI] [PubMed] [Google Scholar]
- 44.Qazi, S. N., C. E. Rees, K. H. Mellits, and P. J. Hill. 2001. Development of gfp vectors for expression in Listeria monocytogenes and other low G+C gram positive bacteria. Microb. Ecol. 41:301-309. [DOI] [PubMed] [Google Scholar]
- 45.Rachid, S., K. Ohlsen, U. Wallner, J. Hacker, M. Hecker, and W. Ziebuhr. 2000. Alternative transcription factor σB is involved in regulation of biofilm expression in a Staphylococcus aureus mucosal isolate. J. Bacteriol. 182:6824-6826. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Shiro, H., E. Muller, N. Gutierrez, S. Boisot, M. Grout, T. D. Tosteson, D. Goldmann, and G. B. Pier. 1994. Transposon mutants of Staphylococcus epidermidis deficient in elaboration of capsular polysaccharide/adhesin and slime are avirulent in a rabbit model of endocarditis. J. Infect. Dis. 169:1042-1049. [DOI] [PubMed] [Google Scholar]
- 47.Throup, J. P., F. Zappacosta, R. D. Lunsford, R. S. Annan, S. A. Carr, J. T. Lonsdale, A. P. Bryant, D. McDevitt, M. Rosenberg, and M. K. Burnham. 2001. The srhSR gene pair from Staphylococcus aureus: genomic and proteomic approaches to the identification and characterization of gene function. Biochemistry 40:10392-10401. [DOI] [PubMed] [Google Scholar]
- 48.Tormo, M. A., M. Martí, J. Valle, A. C. Manna, A. L. Cheung, I. Lasa, and J. R. Penadés. 2005. SarA is an essential positive regulator of Staphylococcus epidermidis biofilm development. J. Bacteriol. 187:2348-2356. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Valle, J., A. Toledo-Arana, C. Berasain, J. M. Ghigo, B. Amorena, J. R. Penades, and I. Lasa. 2003. SarA and not σB is essential for biofilm development by Staphylococcus aureus. Mol. Microbiol. 48:1075-1087. [DOI] [PubMed] [Google Scholar]
- 50.Vuong, C., C. Gerke, G. A. Somerville, E. R. Fischer, and M. Otto. 2003. Quorum-sensing control of biofilm factors in Staphylococcus epidermidis. J. Infect. Dis. 188:706-718. [DOI] [PubMed] [Google Scholar]
- 51.Vuong, C., H. L. Saenz, F. Gotz, and M. Otto. 2000. Impact of the agr quorum-sensing system on adherence to polystyrene in Staphylococcus aureus. J. Infect. Dis. 182:1688-1693. [DOI] [PubMed] [Google Scholar]
- 52.Yarwood, J. M., D. J. Bartels, E. M. Volper, and E. P. Greenberg. 2004. Quorum sensing in Staphylococcus aureus biofilms. J. Bacteriol. 186:1838-1850. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Yarwood, J. M., J. K. McCormick, and P. M. Schlievert. 2001. Identification of a novel two-component regulatory system that acts in global regulation of virulence factors of Staphylococcus aureus. J. Bacteriol. 183:1113-1123. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.