Abstract
Summary
The design and optimization of antibodies requires an intricate balance across multiple properties. Protein inverse folding models, capable of generating diverse sequences folding into the same structure, are promising tools for maintaining structural integrity during antibody design. Here, we present AntiFold, an antibody-specific inverse folding model, fine-tuned from ESM-IF1 on solved and predicted antibody structures. AntiFold outperforms existing inverse folding tools on sequence recovery across complementarity-determining regions, with designed sequences showing high structural similarity to their solved counterpart. It additionally achieves stronger correlations when predicting antibody-antigen binding affinity in a zero-shot manner. AntiFold assigns low probabilities to mutations that disrupt antigen binding, synergizing with protein language model residue probabilities, and demonstrates promise for guiding antibody optimization while retaining structure-related properties.
Availability and implementation
AntiFold is freely available under the BSD 3-Clause as a web server (https://opig.stats.ox.ac.uk/webapps/antifold/) and pip-installable package (https://github.com/oxpig/AntiFold).
1 Introduction
Antibodies are one of the largest classes of therapeutics, used to treat diseases including cancers, autoimmune conditions, and viral infections (Lu et al. 2020). Therapeutic antibody design is complex, requiring the optimization of numerous properties related to efficacy, manufacturability, and safety (Rabia et al. 2018).
Machine learning-based methods have shown promise in accelerating multiple steps in the antibody development pipeline (Hummer et al. 2022) by reducing liabilities such as immunogenicity (Marks et al. 2021, Prihoda et al. 2022, Tennenhouse et al. 2024) and aggregation (Makowski et al. 2023) or rationally optimizing for desirable properties such as binding affinity and developability (Makowski et al. 2022).
A guiding consideration in antibody optimization is selecting mutations that maintain the structure and therefore structure-mediated properties, ranging from stability to antigen binding mode. Protein inverse folding models are trained to predict sequence given structure (Li et al. 2014, O’Connell et al. 2018, Ingraham et al. 2019, Strokach et al. 2020, Jing et al. 2021, Anand et al. 2022, Hsu et al. 2022, Dauparas et al. 2022) and can therefore be used to design novel sequences without altering the antibody backbone structure. Backbone-constrained design could enable the optimization of individual properties without disrupting others.
Antibodies demonstrate distinct structure and sequence properties as compared to general proteins (Stanfield and Wilson 2014, Regep et al. 2017) (Supplementary Fig. S1). While the framework (FR) regions are germline-encoded and relatively conserved, the complementarity-determining region (CDR) loops are hypervariable and form most of the antigen-binding contacts. Over two-thirds of CDRH3 loops have distinct structures not found in other general protein structures (Regep et al. 2017).
Multiple tools for antibody inverse folding design have recently been released, including AbMPNN (Dreyer et al. 2023) and IgMPNN (Shanehsazzadeh et al. 2023), based on the ProteinMPNN architecture (Dauparas et al. 2022). Furthermore, recent work has shown the promise of protein language and inverse folding models for guiding antibody affinity maturation through the identification of high-fitness, structurally constrained regions of the mutational landscape (Hie et al. 2023, Shanker et al. 2024, Outeiral and Deane 2024). However, the sequence recovery of existing tools for the CDR regions has been limited. Additionally, ProteinMPNN-based tools have features such as the occasional re-ordering of antibody chains or CDRH3 position 112 insertions, or the introduction of gaps into IMGT-numbered antibodies, incompatible with antibody structures.
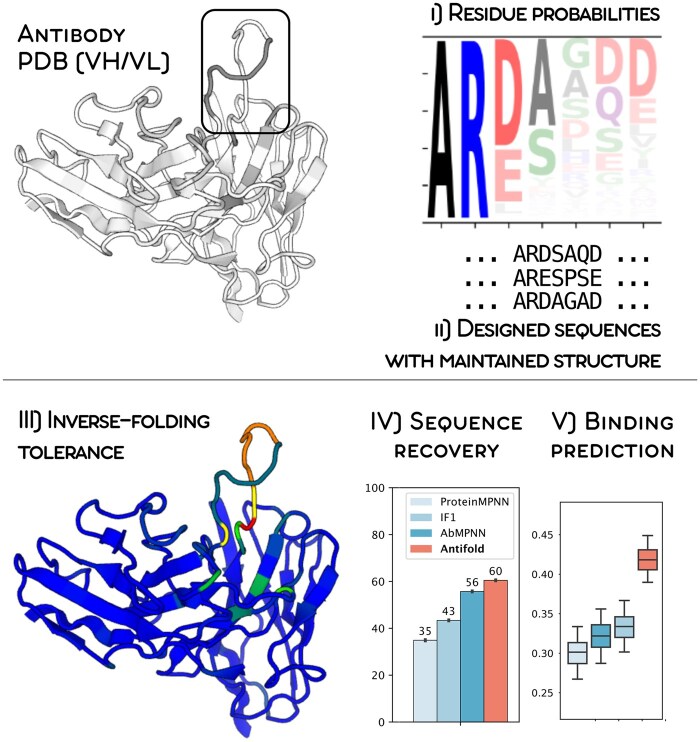
Here, we present AntiFold, an antibody-specific inverse folding model fine-tuned from ESM-IF1 (Hsu et al. 2022), which significantly improves upon CDR sequence recovery and zero-shot affinity prediction. AntiFold accepts a solved or predicted antibody variable domain structure as input. For each residue position, the tool outputs the overall tolerance to mutations without altering the backbone structure (perplexity) and individual amino acid probabilities (Fig. 1). Optionally, the user may specify regions to sample new sequences for and a temperature parameter to control sequence diversity. Designed sequences show high structural similarity to the original structure when re-folded. The use of AntiFold in tandem with other property prediction tools could therefore guide antibody optimization campaigns by prioritizing experimental validation to a smaller search space.
Figure 1.
Structure-constrained antibody design with AntiFold. The user inputs an antibody variable domain PDB structure (heavy and light chain) and specifies an IMGT region to design. AntiFold outputs for each residue position in the PDB: (i) residue probabilities, (ii) a number of designed sequences (default 10) for the selected region, predicted to maintain its structural fold, and (iii) structural tolerance to mutations without altering the backbone structure. The diversity of the generated sequences may be controlled with a temperature parameter. AntiFold achieved (iv) state-of-the-art CDRH3 sequence recovery and (v) inverse folding zero-shot binding affinity prediction on an anti-lysozyme antibody dataset (Warszawski et al. 2019).
2 Methods
2.1 Data
AntiFold leverages ESM-IF1’s pre-training on >12M protein structures by fine-tuning the model further on solved and predicted antibody structures. To enable a direct comparison with AbMPNN, we trained, validated, and tested our model on the same data: 2074 solved complexes from the Structural Antibody Database (SAbDab) (Dunbar et al. 2014, Schneider et al. 2021) and 147,458 structures of sequences from the Observed Antibody Space (OAS) paired database (Kovaltsuk et al. 2018, Olsen et al. 2022) modeled with ABodyBuilder2 (Abanades et al. 2023). Each dataset was split using a 90% concatenated CDR sequence identity cutoff (80/10/10 train/validation/test) (Dreyer et al. 2023). We did not directly compare against IgMPNN as the model weights and training data were not available.
2.2 Training AntiFold
During fine-tuning, we applied several strategies to improve performance on the validation set, specifically amino acid recovery (AAR) of the heavy chain CDR3 loop (CDRH3), which forms most of the antigen binding site interactions. CDRH3 AAR was improved through the use of a span-masking scheme (36.4%), additional masking of random residues (53.2%), weighting masking toward CDR residues with a 3:1 weight (53.3%), layer-wise learning-rate decay (54.4%), and including predicted structures from OAS (58.4%) (Supplementary Tables S1–S3). These augmentations were all included in the final AntiFold model. Training AntiFold without the ESM-IF1 pre-training yielded a CDRH3 AAR of 31.5% on the validation set (Supplementary Table S4), highlighting the value of ESM-IF1’s large pre-training dataset. For more details on these results and our fine-tuning strategy, see the Supplementary Information.
3 Results
3.1 Fine-tuning improves amino acid recovery on antibody sequences
AntiFold demonstrated a substantial improvement in AAR on the experimental structure test set as compared to the original ESM-IF1 model (60% vs 43% AAR for CDRH3, p < 0.005; Table 1, Supplementary Fig. S2). AntiFold also outperformed AbMPNN for the CDRH3 loop (60% vs 56%), all remaining CDR regions (75%–84% vs 63%–76%), and most FR regions (87%–94% vs 85%–89%, Supplementary Fig. S3). On a subset of the test dataset limited to antibodies with <70% length-matched CDR sequence identity cutoff with any sequence in the training or validation dataset (81%, 178 structures), performance was maintained (AAR < 2.6% for any CDR or FR region). When evaluating trained models, full backbone and sequence context were provided.
Table 1.
Summary of AntiFold performance on sequence design and binding affinity prediction.a
| Evaluation | Dataset | Model |
|||
|---|---|---|---|---|---|
| ProteinMPNN | ESM-IF1 | AbMPNN | AntiFold | ||
| Amino acid recovery, CDRH3 (%, ) | AbMPNN test-set | 35% | 43% | 56% | 60% |
| Sequence design, sampled CDR loops (RMSD, ) | AbMPNN test-set | 1.03 | 1.01 | 0.98 | 0.95 |
| AbAg binding affinity (, ) | Warszawski, anti-lysozyme DMS | 0.30 | 0.32 | 0.33 | 0.42 |
| AbAg improved variants (Rank %, ) | Hie et al., 7x Ab affinity-maturation | 73% | 57% | 55% | 80% |
Full backbone and sequence context were provided for all models. For detailed methods and results, see the Supplementary Information. Arrows indicate where higher/lower values are better; the best results are shown in bold. Evaluations from top to bottom: AbMPNN test-set amino acid recovery (Supplementary Fig. S2), backbone RMSD of inverse folding-designed CDR loops (Supplementary Fig. S5), affinity prediction on the Warszawski et al. (2019) antibody-antigen deep mutational scan (Supplementary Fig. S8), separation of Hie et al. (2023) affinity maturation-improved antibody variants (fold-change > 1.25) (Supplementary Fig. S11); AbAg: antibody-antigen.
We confirmed that AntiFold can be accurately applied to modeled structure inputs by testing it on the test set structures predicted with ABodyBuilder2. AntiFold achieved similar AAR for solved and predicted structures (AAR −0.5%), unlike AbMPNN which performed slightly worse on experimental structures (AAR −2.7%, Supplementary Fig. S2C). AntiFold also maintained performance when applied to an antibody structure predicted with AlphaFold [PDB 7M3N, Supplementary Table S5 (Mirdita et al. 2022)].
3.2 Predicted sequences show high structural agreement with experimental structures
To assess whether mutations suggested by AntiFold preserve the backbone structure, we sampled and refolded CDR sequences for a set of high-quality structures. We identified 56 antibody structures in the test set that were solved using X-ray crystallography with a resolution below 2.5 Å. Next, we sampled 20 sequences for each antibody using AntiFold, AbMPNN, ESM-IF1, and ProteinMPNN using a residue sampling temperature of 0.20 (for more details, see Supplementary Methods).
We modeled these sequences using ABodyBuilder2, aligned them with the FR backbone of their experimentally solved counterpart, then calculated the root-mean-square deviation (RMSD) over the CDR residues (for more details, see Supplementary Methods). As a baseline, we modeled the true sequences with ABodyBuilder2 (native, Supplementary Fig. S5). AntiFold generated sequences with high structural similarity to the original backbone, with a mean CDR region RMSD of 0.95 Å, versus AbMPNN (0.98 Å), ESM-IF1 (1.01 Å), and ProteinMPNN (1.03 Å). Structure predictions of the native sequences made using ABodyBuilder2 demonstrated a mean CDR region RMSD of 0.63 Å compared to the experimental CDR backbone.
3.3 Inverse folding predicts antibody-antigen binding affinity
We assessed the ability of AntiFold and other inverse folding models to predict antibody-antigen binding affinity by applying them to a deep mutational scan of an anti-lysozyme antibody (Warszawski et al. 2019). We calculated the log-likelihoods of the 2209 variable domain variants of Fab D44.1 (PDB 1MLC, present in AntiFold’s test set) (Supplementary Fig. S6). As a sequence-only baseline, we included the sequence-based ESM-2 model (650M parameters). AntiFold significantly outperformed the other models with a Spearman’s rank correlation of 0.418, versus ESM-IF1 (0.334), AbMPNN (0.322), ProteinMPNN (0.301), and ESM-2 (0.264) (significance assessed by comparing the 95% confidence interval overlaps). Including the antigen as additional context for the inverse folding models did not result in a statistically significant difference in affinity prediction (Supplementary Figs S9 and S10).
To further explore the ability of AntiFold to guide antibody design by predicting antibody-antigen binding affinity, we applied the model to 124 variants across 7 antibodies generated in protein language model-guided affinity maturation experiments (Hie et al. 2023) (see Supplementary Methods).
We first rank-normalized across all possible single amino acid variant scores for each antibody and each model. We then rank-normalized again across this set to assess the rankings of the 124 experimentally measured variants. Using the experimental binding affinity values, variants were separated into lower (fold-change < 0.75), maintained (0.75–1.25), and improved (>1.25) binding affinity groups (Supplementary Fig. S11).
AntiFold achieved significantly improved separation of these groups, scoring the improved variants with a median rank score of 80% (p < 0.005), versus ProteinMPNN (73%), ESM-IF1 (57%), and AbMPNN (55%).
3.4 Availability and implementation
AntiFold can be accessed as a web server or freely downloaded as a pip-installable package.
Web server: https://opig.stats.ox.ac.uk/webapps/antifold/
Code repository: https://github.com/oxpig/AntiFold
4 Discussion
AntiFold, fine-tuned from the general protein inverse folding model ESM-IF1, achieved state-of-the-art performance on antibody sequence recovery and refolding of designed sequences. The use of pre-trained weights substantially improved AntiFold’s performance versus training from scratch, while including predicted structures and weighting masking toward individual CDR residues increased sequence recovery further.
AntiFold’s inverse folding probabilities correlate with antibody-antigen binding affinity across multiple independent experiments, likely by identifying mutations that disrupt the structure and antigen binding. These probabilities also synergized with protein language model-suggested variants, supporting that the models learn orthogonal information. Consistent with previous results (Hie et al. 2023, Shanker et al. 2024), our findings indicate that AntiFold identifies structurally constrained, high-fitness regions of the mutational landscape, predicted to preserve binding.
Supplementary Material
Contributor Information
Magnus Haraldson Høie, Section for Bioinformatics, Department of Health Technology, Technical University of Denmark, Lyngby DK-2800, Denmark.
Alissa M Hummer, Department of Statistics, University of Oxford, Oxford OX1 3LB, United Kingdom.
Tobias H Olsen, Department of Statistics, University of Oxford, Oxford OX1 3LB, United Kingdom.
Broncio Aguilar-Sanjuan, Department of Statistics, University of Oxford, Oxford OX1 3LB, United Kingdom.
Morten Nielsen, Section for Bioinformatics, Department of Health Technology, Technical University of Denmark, Lyngby DK-2800, Denmark.
Charlotte M Deane, Department of Statistics, University of Oxford, Oxford OX1 3LB, United Kingdom.
Supplementary data
Supplementary data are available at Bioinformatics Advances online.
Conflict of interest
No competing interest is declared.
Funding
This work was supported by the Sino-Danish Center [2021, awarded to M.H.H.], the UKRI Medical Research Council [MR/N013468/1 awarded to A.M.H.], the UKRI Engineering and Physical Sciences Research Council [EP/S024093/1 awarded to T.H.O.] and GlaxoSmithKline plc.
References
- Abanades B, Wong WK, Boyles F et al. ImmuneBuilder: deep-learning models for predicting the structures of immune proteins. Commun Biol 2023;6:575–8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Anand N, Eguchi R, Mathews II et al. Protein sequence design with a learned potential. Nat Commun 2022;13:746. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dauparas J, Anishchenko I, Bennett N et al. Robust deep learning-based protein sequence design using proteinmpnn. Science 2022;378:49–56. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dreyer FA, Cutting D, Schneider C et al. Inverse folding for antibody sequence design using deep learning. In: The 2023 ICML Workshop on Computational Biology. 2023.
- Dunbar J, Krawczyk K, Leem J et al. SAbDab: the structural antibody database. Nucleic Acids Res 2014;42:D1140–6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hie BL, Shanker VR, Xu D et al. Efficient evolution of human antibodies from general protein language models. Nat Biotechnol 2023;42:275–83. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hsu C, Verkuil R, Liu J et al. Learning inverse folding from millions of predicted structures. bioRxiv, 2022, preprint: not peer reviewed.
- Hummer AM, Abanades B, Deane CM. Advances in computational structure-based antibody design. Curr Opin Struct Biol 2022;74:102379. [DOI] [PubMed] [Google Scholar]
- Ingraham J, Garg V, Barzilay R et al. Generative models for graph-based protein design. In: Advances in Neural Information Processing Systems, Vol. 32. 2019.
- Jing B, Eismann S, Suriana P et al. Learning from protein structure with geometric vector perceptrons. In: International Conference on Learning Representations. 2021.
- Kovaltsuk A, Leem J, Kelm S et al. Observed antibody space: a resource for data mining next-generation sequencing of antibody repertoires. J Immunol 2018;201:2502–9. [DOI] [PubMed] [Google Scholar]
- Li Z, Yang Y, Faraggi E et al. Direct prediction of profiles of sequences compatible with a protein structure by neural networks with fragment-based local and energy-based nonlocal profiles. Proteins 2014;82:2565–73. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lu R-M, Hwang Y-C, Liu I-J et al. Development of therapeutic antibodies for the treatment of diseases. J Biomed Sci 2020;27:1–30. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Makowski EK, Kinnunen PC, Huang J et al. Co-optimization of therapeutic antibody affinity and specificity using machine learning models that generalize to novel mutational space. Nat Commun 2022;13:3788. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Makowski EK, Wang T, Zupancic JM et al. Optimization of therapeutic antibodies for reduced self-association and non-specific binding via interpretable machine learning. Nat Biomed Eng 2023;8:45–56. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marks C, Hummer AM, Chin M et al. Humanization of antibodies using a machine learning approach on large-scale repertoire data. Bioinformatics 2021;37:4041–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mirdita M, Schütze K, Moriwaki Y et al. Colabfold: making protein folding accessible to all. Nat Methods 2022;19:679–82. [DOI] [PMC free article] [PubMed] [Google Scholar]
- O’Connell J, Li Z, Hanson J et al. Spin2: predicting sequence profiles from protein structures using deep neural networks. Proteins 2018;86:629–33. [DOI] [PubMed] [Google Scholar]
- Olsen TH, Boyles F, Deane CM. Observed antibody space: a diverse database of cleaned, annotated, and translated unpaired and paired antibody sequences. Protein Sci 2022;31:141–6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Outeiral C, Deane CM. Perfecting antibodies with language models. Nat Biotechnol 2024;42:185–6. [DOI] [PubMed] [Google Scholar]
- Prihoda D, Maamary J, Waight A et al. Biophi: a platform for antibody design, humanization, and humanness evaluation. MAbs 2022;14:2020203. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rabia LA, Desai AA, Jhajj HS et al. Understanding and overcoming trade-offs between antibody affinity, specificity, stability and solubility. Biochem Eng J 2018;137:365–74. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Regep C, Georges G, Shi J et al. The h3 loop of antibodies shows unique structural characteristics. Proteins 2017;85:1311–8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schneider C, Raybould MIJ, Deane CM. SAbDab in the age of biotherapeutics: updates including SAbDab-nano, the nanobody structure tracker. Nucleic Acids Res 2021;50:D1368–72. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shanehsazzadeh A, Alverio J, Kasun G et al. In vitro validated antibody design against multiple therapeutic antigens using generative inverse folding. bioRxiv, 2023, preprint: not peer reviewed.
- Shanker VR, Bruun TUJ, Hie BL et al. Unsupervised evolution of protein and antibody complexes with a structure-informed language model. Science 2024;385:46–53. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stanfield RL, Wilson IA. Antibody structure. Microbiol Spectr 2014;2. doi: 10.1128/microbiolspec.aid-0012-2013. [DOI] [PubMed] [Google Scholar]
- Strokach A, Becerra D, Corbi-Verge C et al. Fast and flexible protein design using deep graph neural networks. Cell Syst 2020;11:402–11.e4. [DOI] [PubMed] [Google Scholar]
- Tennenhouse A, Khmelnitsky LEV, Khalaila R et al. Computational optimization of antibody humanness and stability by systematic energy-based ranking. Nat Biomed Eng 2024;8:30–44. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Warszawski S, Borenstein Katz A, Lipsh R et al. Optimizing antibody affinity and stability by the automated design of the variable light-heavy chain interfaces. PLoS Comput Biol 2019;15:e1007207. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.