Abstract
Introduction
Serologic characterization of antihuman platelet antigen (HPA) alloantibodies is crucial in fetal neonatal alloimmune thrombocytopenia. The gold standard MAIPA assay requires fresh platelets from HPA-genotyped donors, which is challenging for some laboratories. Megakaryocytes express HPA epitopes and offer an alternative source for detecting anti-HPA antibodies. The objective of this study was to assess the efficacy of a novel assay called monoclonal antibody immobilization of megakaryocyte antigens (MAIMA) for detecting anti-HPA antibodies.
Methods
CD34+ cells from buffy coats were differentiated into megakaryocytes in vitro. The performance of the MAIMA assay was evaluated using WHO reference reagents for HPA-1a, HPA-3a, and HPA-5b, along with sera samples from patients who had well-characterized anti-HPA antibodies.
Results
The WHO anti-HPA-1a reference reagent showed similar binding to megakaryocytes and platelets in MAIMA and MAIPA, respectively. On the other hand, optical density (OD) values for the WHO anti-HPA-3a reference reagent were lower in MAIMA than in MAIPA. Anti-HPA-5b antibodies were not detectable in MAIMA. Patients’ sera containing anti-HPA-1a antibodies were successfully detected in MAIMA in all clinical samples. Moreover, OD values in MAIPA and MAIMA showed high correlation (r = 0.96, p < 0.001). MAIMA was reactive for samples with anti-HPA-3a as well as anti-HPA-3b; however, OD values were lower compared to MAIPA. Interestingly, all patient samples with anti-HPA-5b antibodies were tested negative in MAIMA.
Conclusion
In vitro generated megakaryocytes can be used to detect anti-HPA-1a alloantibodies. However, despite this potential, they may be less suitable for the detection of alloantibodies against other HPAs such as HPA-5b.
Keywords: Antibodies, Buffy coat, Fetal alloimmune thrombocytopenia, Platelet antigens
Plain Language Summary
The identification of antibodies to human platelet antigens (HPA) is crucial. The standard test (MAIPA assay) is difficult because it requires fresh platelets from specific donors. This study investigated a new method, monoclonal antibody immobilization of megakaryocyte antigens (MAIMA), using in vitro generated megakaryocytes from CD34+ cells. The assay was tested against WHO reference reagents and patient samples with known anti-HPA antibodies. Results showed that MAIMA successfully detected anti-HPA-1a antibodies and correlated well with MAIPA. However, MAIMA had lower sensitivity for anti-HPA-3a and anti-HPA-5b antibodies. Notably, anti-HPA-5b antibodies were not detectable by MAIMA. In conclusion, in vitro generated megakaryocytes can effectively detect anti-HPA-1a antibodies. However, MAIMA may be less suitable for antibodies against other HPAs, such as HPA-5b. This study highlights the potential of MAIMA as an alternative assay and provides insight into its strengths and limitations in detecting specific anti-HPA antibodies.
Introduction
Fetal and neonatal alloimmune thrombocytopenia (FNAIT) is a rare bleeding disorder caused by maternal alloantibodies that react to human platelet antigens (HPAs) [1]. Maternal IgG class alloantibodies directed to fetal HPAs can be transported across the placenta and cause thrombocytopenia in the fetus or newborn [1]. Approximately 10% of affected cases may have life-threatening intracranial bleeding with severe morbidity [2, 3]. The incidence of FNAIT in Caucasian population is estimated to be 1 in 800–1,500 pregnancies [4].
To date, 35 HPAs have been identified on platelets [5]. They are located on 6 glycoproteins (GPs) that play an important role in platelet function [6]. In addition to FNAIT, anti-HPA alloantibodies are involved in several immune-related platelet disorders, including post-transfusion purpura, and platelet transfusion refractoriness [7].
Serological detection of anti-HPA antibodies is essential for the correct diagnosis and appropriate treatment of FNAIT. Several methods have been developed, such as the platelet immunofluorescence assay, the monoclonal antibody immobilization of platelet antigens (MAIPA) assay, the modified antigen capture enzyme immunoassay, and beads-based flow cytometric assays [8, 9]. In Europe, the MAIPA assay is considered to be the gold standard for the detection of platelet-specific antibodies. One commercial kit with test cells is also available for the diagnosis of platelet antibodies. However, due to its lower sensitivity for certain antigens and high cost, several laboratories prefer in house MAIPA. However, the MAIPA assay requires a panel of well-characterized, HPA-genotyped fresh platelets obtained from blood donors [8, 9], which is a limitation for the use of MAIPA in laboratories that do not have direct access to blood donors. Additionally, the availability of donors with rare antigens for testing poses additional limitations for MAIPA with fresh donor blood.
Recently, Zhang et al. [10] demonstrated a novel approach by utilizing induced pluripotent stem cell-derived human megakaryocytes engineered to express rare platelet antigens, enabling the detection of HPA-specific alloantibodies. Additionally, the idea of cryopreserving test cells for convenient off-the-shelf testing has been introduced [11]. As preservation techniques advance, the prospect of storing in vitro generated cells for extended periods becomes feasible, opening up opportunities for prolonged use in testing scenarios. The in vitro generation of genetically engineered megakaryocytes for platelet alloantibodies against HPA antigens holds promise in overcoming the current limitations of gold standard testing.
Our group has previously shown that CD34+ cells isolated from either peripheral blood or buffy coat can be differentiated into megakaryocytes in vitro [12]. Indeed, buffy coat-derived megakaryocytes have a higher yield of platelets with better in vitro functionality than cells derived from peripheral blood stem cells. In vitro generated cells can be used to overcome the demand for cells with certain antigens for the diagnosis of anti-HPA alloantibodies [10, 13]. In the current study, we hypothesized that buffy coat derived in vitro generated megakaryocytes can be used as a substitute for platelets in the monoclonal antibody immobilization assay for the detection of HPA alloantibodies.
Study Design and Methods
WHO Reference Reagents and Patients’ Samples
The reference reagents, suggested by the World Health Organization (WHO), for HPA-1a (NIBSC Code 03/152, NIBSC, Hertfordshire, UK), HPA-3a (NIBSC Code 03/190, NIBSC, Hertfordshire, UK) and HPA-5b (NIBSC Code 99/666, NIBSC, Hertfordshire, UK) were used in this study. In addition, serum samples containing well-characterized HPA antibodies were obtained from clinical samples referred to our platelet immunology laboratory for serological investigation (online suppl. Table 1; for all online suppl. material, see https://doi.org/10.1159/000539617).
In vitro Generation of Megakaryocytes
Human CD34+ cells were isolated from buffy coat samples as previously described with minor modifications [12]. In brief, Buffy coats were obtained from HPA-genotyped blood donors. The CD34 MicroBead Ultra-Pure Kit (Miltenyi Biotec, Bergisch Gladbach, Germany) was used to isolate CD34+ cells from buffy coat according to the manufacturer’s instructions. Briefly, samples were diluted (1:1) with phosphate-buffered saline (PBS) with 2% fetal bovine serum (FBS; PBS-FE) buffer (Sigma Aldrich, St. Louis, MO, USA and 0.5% EDTA [Invitrogen]), layered on density gradient medium (Ficoll-Paque, GE Healthcare, Uppsala, Sweden) and centrifugated (400 g, 40 min at RT, without brake). Next, the peripheral blood mononuclear cells fraction was collected and after three washing steps with PBS-FE (200 g, 12 min at RT), the cell number was determined using the Neubauer chamber. Thereafter, FcR blocking reagent and CD34 MicroBeads were incubated with the cells (30 min at 4°C). After a washing step (300 g, 10 min at RT), cells were collected and applied to the column on a magnetic separator and flushed out. This step was repeated with a second column to increase the purity of the isolated CD34+ cells. Finally, cells were cultured for 6 days in StemSpan SFEM medium (Stem Cell Technology, Vancouver, BC, Canada) containing 20 ng/mL stem cell factor, 20 ng/mL interleukin 3 and 30 ng/mL TPO (all from Miltenyi Biotec, Bergisch Gladbach, Germany).
CD34+ cells were then seeded in 8 wells chamber slide (Corning, NY, USA; 50,000 cells/well) and cultured in StemSpan SFEM medium in the presence of 50 ng/mL of TPO. After 14 days, cells were collected by gently pipetting, washed once with PBS (650 g, 7 min at RT) and used for further investigations.
Flow Cytometer Characterization of MKs
The characterization of the megakaryocytes was performed by flow cytometry as previously described with minor modifications [12]. In brief, antihuman CD41-PC5, CD61-FITC, CD42a-FITC and CD42b-PE (all from Beckman Coulter, Brea, CA, USA) antibodies were used to analyze the cell differentiation status. For some experiments, we also used and CD49b-FITC for HPA-5 (GP Ia). The flow cytometric characterization of the megakaryocytes is presented in the online supplementary Figure 1. Furthermore, the polyploidy of the megakaryocytes was assessed performing propidium iodide (Sigma Aldrich, St. Louis, MO, USA) staining [12]. In brief, cells were fixed with 4% paraformaldehyde (PFA 15 min at RT; Morphisto, Frankfurt, Germany), washed with PBS (650 g, 7 min at RT) and incubated with 100 µg/mL RNAse (Sigma Aldrich, 10 min at RT) prior to the incubation with 50 µg/mL of propidium iodide for 15 min at RT and immediately analyzed.
HPA Genotyping
Blood donors were genotyped for 6 HPA systems (HPA-1 to 5 and HPA-15) by polymerase chain reaction with sequence-specific primers method using a commercially available kit (HPA-Ready Gene Kit, Inno-Train, Germany) according to the manufacturer’s instructions.
Antibody Detection Using MAIPA and Monoclonal Antibody Immobilization of Megakaryocyte Antigens
The MAIPA assay was performed as previously described [14]. The same protocol was used for antibody detection with buffy coat-derived megakaryocytes in monoclonal antibody immobilization of megakaryocyte antigen (MAIMA) assay. In brief, aliquots of test platelets (40 × 106 cells) or in vitro generated megakaryocytes (100–200 × 103 cells) were resuspended in 50 μL of PBS/bovine serum albumin (BSA) supplemented with 0.2% w/v BSA (SERVA Electrophoresis GmbH, Heidelberg, Germany). The resuspended cells were incubated with 50 μL of patient serum, WHO reference reagents or control serum for 30 min at 37°C. Serum samples without HPA and HLA antibodies were used as a negative control. After 2 washing steps, 30 μL of capture antibodies (1:10 diluted in BSA) were added and incubated for 30 min at 37°C. In this study, we used monoclonal antibodies P2 and Gi9 (both from Beckman Coulter, Brea, CA, USA) specific for GPIIb/IIIa and GPIa/IIa, respectively, as capture antibodies. Following 3 washing steps with PBS/BSA, cells were solubilized by incubation with 0.5% Triton-X-100 (BDH Laboratory Supplies, Poole, UK) for 30 min at 4°C. After centrifugation at 12,000 g for 20 min at 7°C, 50 μL of supernatant containing GPs was taken into new tube prefilled with 200 μL of Tris-buffered saline (TBS). A 96-well microtiter plate (Greiner Bio-One, Frickenhausen, Germany) was coated with goat anti-mouse IgG (1:2,500, Jackson Immunoresearch Lab., West Grove, PA, USA) washed with wash buffer. 100 µL of either platelet or megakaryocyte suspension were transferred into the corresponding coated wells and incubated 90 min at 4°C. The plate was washed four times with 200 μL of TBS. Thereafter, 100 μL of horseradish peroxidase-conjugated goat antihuman IgG (1:5,000, Jackson Immunoresearch Labs., West Grove, PA, USA) was added and the plate was incubated for 120 min at 4°C. Next, the plate was washed five times followed by addition of 100 μL of TMB One ECO-TEK solution (3, 3′, 5, 5′-tetramethylbenzidine, Kementec Solutions, Taastrup, Denmark). The enzymatic reaction was stopped by the addition of 50 μL of 0.5 m H2SO4 to each well. The absorbance (optical density [OD]) was measured at 450 nm with a reference filter of 620 nm on a microtiter plate reader (Tekan, Greilsheim, Germany).
Statistics
Statistical analyses were performed using GraphPad Prism 9.4.1 (GraphPad Software, La Jolla, USA). A t test was used to analyze normally distributed results. If the data did not follow a normal distribution as assessed by D’Agostino and Pearson omnibus normality test, a nonparametric test (Mann-Whitney U test) was used. Data were presented as either n (%) or mean ± standard error of mean (SEM). A Pearson correlation test was used to compare OD values. A p value <0.05 was assumed to represent statistical significance.
Results
Clinical Samples
We used sera from 21 patients (all female) with a median age of 31 years (range: 22–66) in this study. Most of the patients had FNAIT. A list of the characteristics of patients and antibodies can be found in online supplementary Table 1.
The Reactivity of WHO Anti-HPA Standards in MAIMA
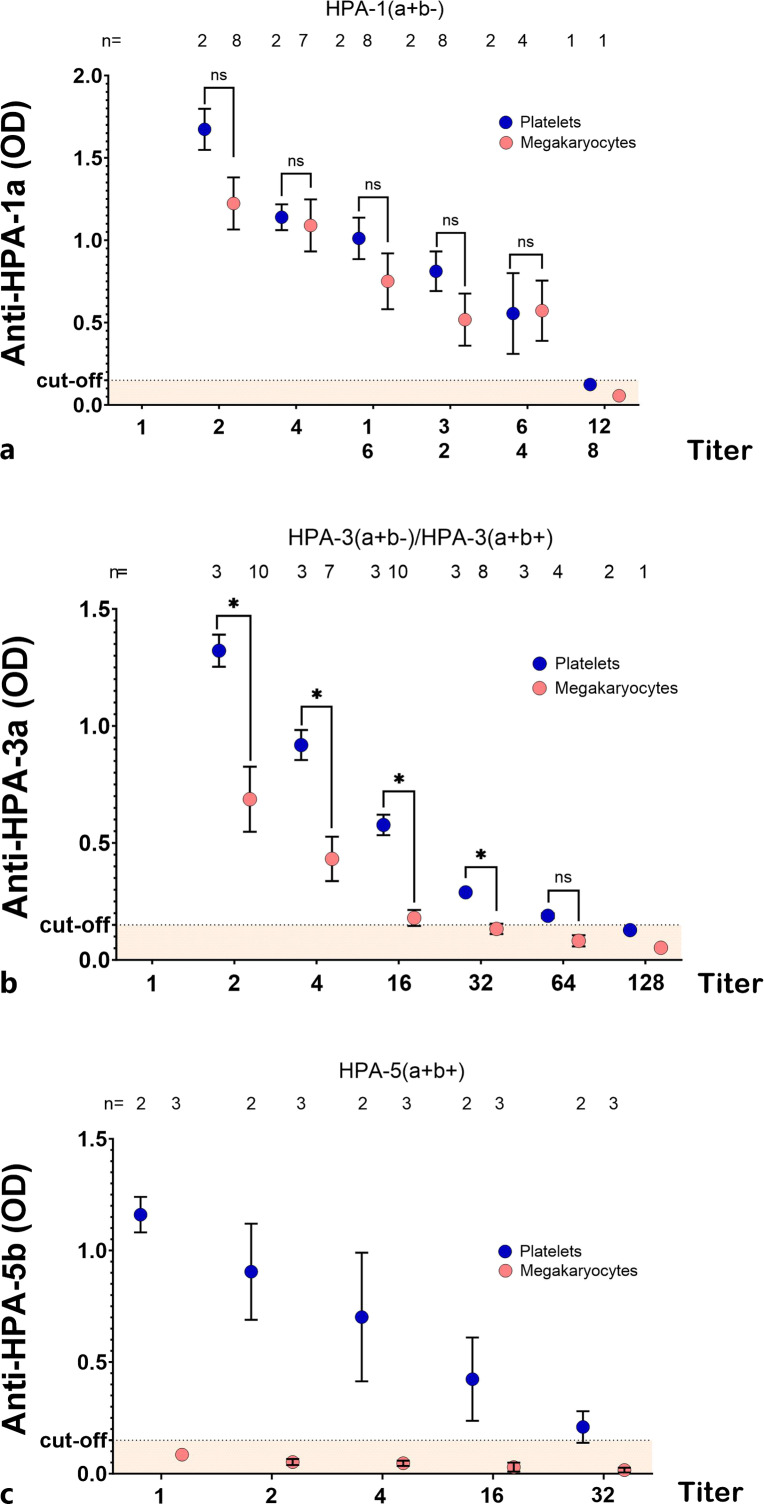
To test the sensitivity of MAIMA, WHO standard antibodies for HPA-1a, HPA-3a, and HPA-5b were used at different titrations. For anti-HPA-1a, OD values measured with HPA-1(a+b−) megakaryocytes and platelets were similar in all titrations up to 1:128 titration (p > 0.05, Fig. 1a). On the other hand, OD values measured, when anti-HPA-3a was incubated with HPA-3(a+b−) and HPA-3(a+b+) megakaryocytes, were significantly lower than OD values measured with platelets in MAIPA (Fig. 1b). MAIMA with HPA-5(a+b+) megakaryocytes showed negative reactions and failed to detect anti-HPA-5b antibodies in WHO standard (Fig. 1c).
Fig. 1.
Comparison of MAIMA and MAIPA for anti-HPA antibody detection. The optical density (OD) was compared in monoclonal antibody immobilization of megakaryocyte antigens (MAIMA) assay and monoclonal antibody immobilization of platelet antigens (MAIPA) assay using WHO reference reagents for anti-HPA-1a, anti-HPA-3a, and anti-HPA-5b at different titrations. While OD values for anti-HPA-1a were similar between MAIMA and MAIPA for all tested titrations (a), they were significantly lower for anti-HPA-3a in MAIMA compared to MAIPA for most of the tested titrations (b). On the other hand, MAIMA failed to detect anti-HPA-5b antibodies (c). Platelets are represented with blue circles, and megakaryocytes with red circles. An OD of greater than 0.15 was considered positive (dashed line).
Detection of Anti-HPA Alloantibodies in Patients’ Sera
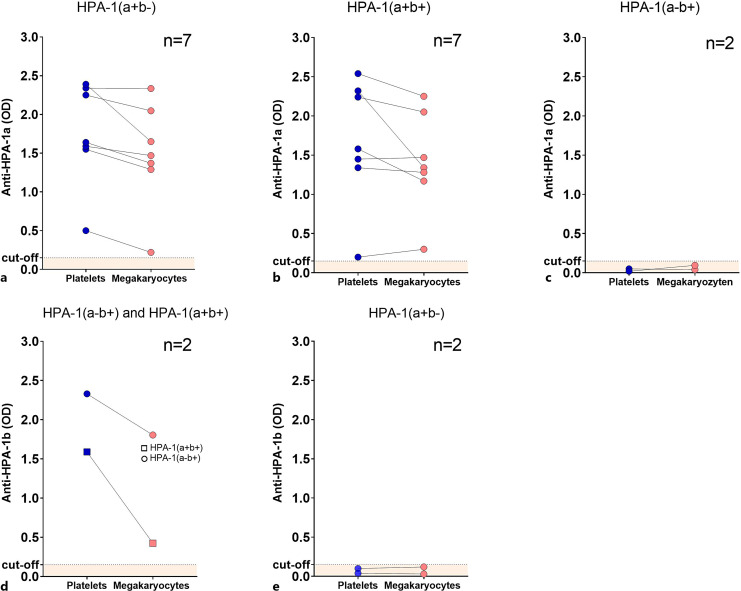
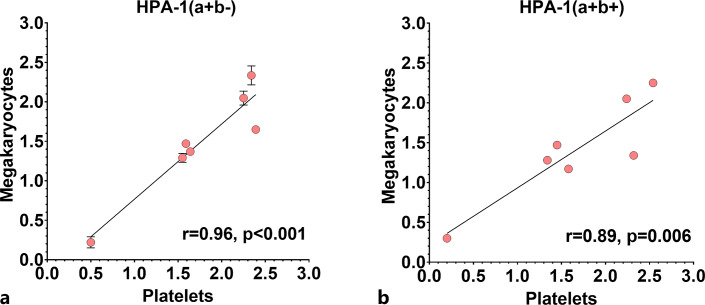
To test the specificity of MAIMA, we used well-characterized clinical samples with anti-HPA-1a, -1b, -3a, -3b, and -5b antibodies. MAIMA with HPA-1(a+b−) or HPA-1(a+b+) megakaryocytes successfully detected anti-HPA-1a antibodies in all samples (Fig. 2a–b). Furthermore, HPA-1(a−b+) megakaryocytes were negative with sera of HPA-1a (Fig. 2c). Although MAIMA with HPA-1(a−b+) and HPA-1(a+b+) megakaryocytes was reactive with patient sera containing HPA-1b alloantibody, OD values were lower than those measured with HPA-1(a−b+) and HPA-1 (a+b+) platelets in MAIPA (Fig. 2d). HPA-1(a+b−) megakaryocytes were negative with sera of HPA-1b (Fig. 2e). OD values in MAIPA and MAIMA showed a high correlation (r = 0.96, p < 0.001 with HPA-1(a+b−) cells; r = 0.89, p = 0.006 with HPA-1(a+b+) cells, Fig. 3a, b).
Fig. 2.
Reactivity of sera with anti-HPA-1a alloantibodies in MAIPA and MAIMA. The reactivity of sera with anti-HPA-1a and anti-HPA-1b alloantibodies was compared in monoclonal antibody immobilization of megakaryocyte antigens (MAIMA) assay and in monoclonal antibody immobilization of platelet antigens (MAIPA) assay. The optical density (OD) values were similar between MAIMA and MAIPA for all tested sera in the HPA-1(a+b−) cells (a) for HPA-1(a+b+) cells (b). HPA-1(a−b+) megakaryocytes were negative with sera of HPA-1a (c). OD values of anti-HPA-1b alloantibodies were lower in MAIMA using HPA-1 (a−b+) or HPA-1 (a+b+) megakaryocytes (d). HPA-1(a+b−) megakaryocytes were negative with sera of HPA-1b (e). Platelets are represented with blue circles, and megakaryocytes with red circles. n refers to number of sera tested. An OD of greater than 0.15 was considered positive (dashed line).
Fig. 3.
Correlation between monoclonal antibody immobilization of megakaryocyte antigens (MAIMA) assay and monoclonal antibody immobilization of platelet antigens (MAIPAs) assay for anti-HPA-1a alloantibody. The OD values in MAIPA and MAIMA showed a high correlation with HPA-1(a+b−) cells (a) as well as with HPA-1(a+b+) cells (b).
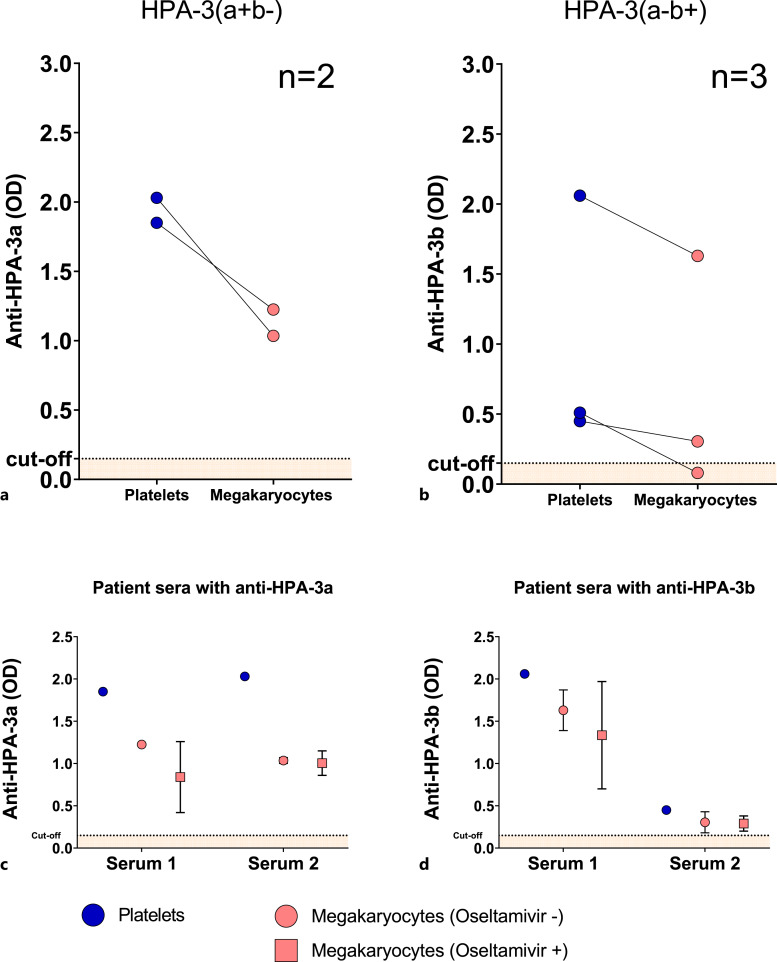
Two patients’ sera with HPA-3a antibody were tested in MAIMA with HPA-3(a+b−) megakaryocytes. MAIMA was reactive for both samples, but the OD values were 33–50% lower in MAIMA then MAIPA (Fig. 4a). We tested 3 patients’ sera with anti-HPA-3b antibody in MAIMA with HPA-3(a−b+) megakaryocytes. One serum was negative in MAIMA and OD values were lower in MAIMA than those measured in MAIPA (Fig. 4b).
Fig. 4.
Reactivity of Sera with anti-HPA-3a and Anti-HPA-3b alloantibodies in MAIPA and MAIMA The reactivity of sera with anti-HPA-3a or anti-HPA-3b alloantibodies was compared in monoclonal antibody immobilization of megakaryocyte antigens (MAIMA) assay and in monoclonal antibody immobilization of platelet antigens (MAIPA) assay. The optical density (OD) values were 33–50% lower in MAIMA then MAIPA for sera with HPA-3a alloantibody (a). Similarly, the OD values were 5–60% lower in MAIMA than those measured in MAIPA for sera with HPA-3b alloantibody (b). We investigated whether the variable reactivity of HPA-3a antibodies in MAIMA assay is due to the loss of sialic acid from platelet receptors on megakaryocytes during cultivation. To test our hypothesis, we treated megakaryocytes with oseltamivir, a neuraminidase inhibitor, and compared the reactivity of HPA-3a (c) and HPA-3b (d) patient sera in MAIMA assay with oseltamivir-treated and untreated megakaryocytes. n refers to number of sera tested. An OD of greater than 0.15 was considered positive (dashed line).
We hypothesized that the loss of sialic acid from GPs on megakaryocytes during cultivation may explain the heterogenic reactivity of HPA-3a antibodies in MAIPA. To verify this hypothesis, megakaryocytes were treated with oseltamivir, a neuraminidase inhibitor, during the in vitro differentiation steps. However, in MAIMA with HPA-3a and HPA-3b patient sera, the reactivity was similar between oseltamivir-treated and untreated megakaryocytes (Fig. 4c, d).
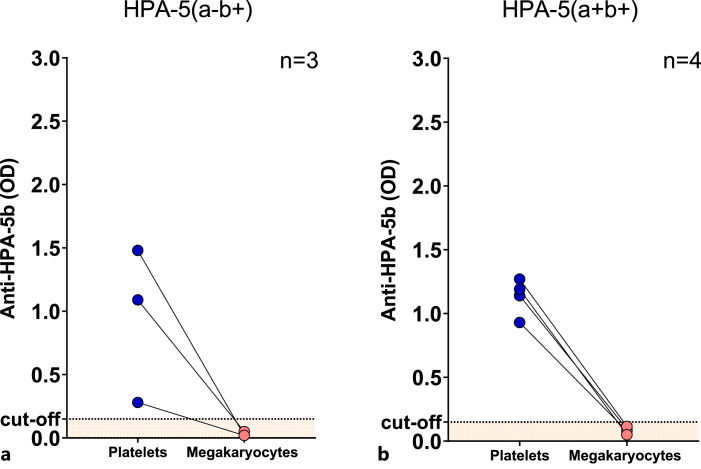
Finally, 3 patients’ sera with anti-HPA-5b antibodies were tested with HPA-5(a−b+) megakaryocytes and 4 patients’ sera were tested with HPA-5(a+b+) megakaryocytes. MAIMA failed to detect any of these anti-HPA-5b antibodies (Fig. 5a, b). In the same line, in vitro generated megakaryocytes showed in a subsequent experiment low expression of CD49b/CD41 compared to platelets from healthy donors (% marker expression, mean ± SEM, platelets vs. megakaryocytes, 4.6 ± 1.6% vs. 98.6 ± 0.3%; p < 0.001, n = 3, Fig. 6). On the other hand, CD41/CD42a expression was comparable between megakaryocyte and platelets (% marker expression, mean ± SEM, platelets vs. megakaryocytes, 88.3 ± 2.2% vs. 95.6 ± 2.8%; p = 0.110; n = 3, Fig. 6).
Fig. 5.
Reactivity of sera with anti-HPA-5b Alloantibodies in MAIPA and MAIMA. We compared the reactivity of sera with anti-HPA-5b alloantibodies in monoclonal antibody immobilization of megakaryocyte antigens (MAIMA) assay and in monoclonal antibody immobilization of platelet antigens (MAIPA) assay using HPA-5(a−b+) (a) and HPA-1(a+b+) (b) cells. MAIMA has failed to detect anti-HPA-5b alloantibodies in clinical samples. Platelets are represented with blue circles, and megakaryocytes represented with red circles. n refers to number of sera tested. An optical density (OD) of greater than 0.15 was considered positive (dashed line).
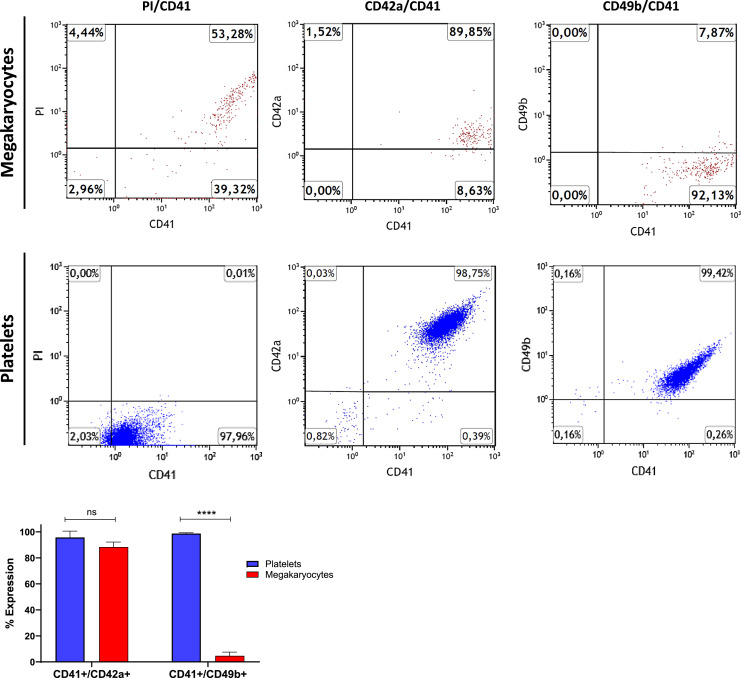
Fig. 6.
CD49b expression on in vitro generated megakaryocytes analyzed by flow cytometry. Expression of CD49b on in vitro generated megakaryocytes and peripheral blood platelets was analyzed using flow cytometry. Megakaryocytes were generated from CD34+ hematopoietic stem cells isolated from buffy coat using a standardized protocol, while platelets were isolated from peripheral blood. Cells were stained with antibodies against CD41, CD42a, and CD49b. Representative dot plots showing CD41 expression on the x-axis and CD42a or CD49b expression on the y-axis seen. Cells were stained with propidium iodide (PI), a DNA marker. The percentage of CD41/CD42a positive cells was similar between platelets and in vitro generated megakaryocytes. In comparison to peripheral blood platelets, in vitro generated megakaryocytes showed significantly lower levels of CD41/CD49b double positive cells.
Discussion
Detection of alloantibodies against HPAs is an essential step in the diagnosis of FNAIT, post-transfusion purpura, and platelet transfusion refractoriness [8]. The need for fresh platelets for the current gold standard MAIPA assay is a limitation for laboratories with no immediate access to HPA-typed blood donors [10, 13]. In vitro generation of platelets is a novel area of investigation to overcome this limitation for the generation of cells with certain antigens [10, 13]. In the current study, we investigated the use of buffy coat-derived megakaryocytes in the detection of anti-HPA alloantibodies. While megakaryocytes showed a similar performance in the detection of both WHO standard antibodies and patients’ antibodies against HPA-1a, a variable reactivity was observed for anti-HPA-3a antibodies and anti-HPA-5b antibodies could not be detected using the new assay.
Alloantibodies against HPA-1a are detected in almost 75% of cases with FNAIT [15, 16]. The HPA-1 antigen system is located on the GP IIIa subunit of the GP complex IIb/IIIa (αIIbβ3 integrin, CD41/CD61), which is the most abundant receptor on platelet surface. A single nucleotide polymorphism (rs5918; T176>C) leads to a leucine (HPA-1a) to proline (HPA-1b) exchange at residue 33 from the NH2-terminus of the protein [17]. In the current study, anti-HPA-1a antibodies could be detected in all clinical samples using megakaryocytes expressing HPA-1a. This result suggests that the use of buffy coat-derived megakaryocytes for the detection of anti-HPA-1a alloantibodies in patient samples is feasible. However, further studies are needed to determine the true sensitivity and specificity of MAIMA for HPA-1a-specific antibodies.
The biallelic HPA-3 antigen system is also located on GP complex IIb/IIIa. The single nucleotide variation on (GPIIb, CD41) leads to a isoleucine (HPA-3a) to serine (HPA-3b) exchange on C-terminus of GPIIb heavy chain [18]. Although anti-HPA-3a antibodies have low frequency in FNAIT [15, 16], it might be associated with a severe clinical course [19]. In the current study, OD values in MAIMA assay were significantly lower with anti-HPA-3a WHO reference reagent as well as with HPA-3a patient sera, compared to MAIPA. Similar to our findings, Hayashi et al. [20] showed that K562 cells, transfected with cDNA encoding specific HPA, could detect anti-HPA-3a alloantibodies in only 4 of 6 clinical samples. On the other hand, Zhang et al. [10] used megakaryocytes expressing either HPA-3a or HPA-3b derived from iPSCs to detect HPA-3 antibodies. HPA-3a and HPA-3b sera reacted with iPSC-derived megakaryocytes expressing HPA-3a or HPA-3b, respectively, in flow cytometry [10]. A number of factors could explain the discrepancy between the findings of our study and previous studies. It has been shown that whole platelet assays such as flow cytometry are more sensitive than GP-specific assays such as MAIPA, probably because the HPA-3 epitopes are affected by the solubilization steps [21, 22]. While Zhang et al. [10] used flow cytometry, Hayashi et al. [20] and we used MAIPA and MAIMA, respectively, for the detection of HPA antibodies. Flow cytometry may be a more sensitive technique than MAIMA for the detection of specific antibodies, including those directed against HPA-3a. Our preliminary results (data not shown) suggest the feasibility of using buffy coat-derived megakaryocytes in flow cytometry assays for HPA-1a as well as HPA-3a alloantibody detection. Further research is required to fully explore this promising approach.
Another possible explanation for the decreased sensitivity for anti-HPA-3 antibodies could be the loss of sialic acids on the megakaryocyte surface during cultivation. In fact, the epitopes recognized by anti-HPA-3 antibodies has been shown to be a complex [21–23]. n-acetylneuraminic acid also contributes to the formation of HPA-3a epitopes [24]. Platelets loose sialic acids on their surface during prolonged storage [25]. Socher et al. [22] showed that the reactivity of some anti-HPA-3a and anti-HPA-3b alloantibodies in MAIPA decreases when test platelets are stored at 4°C for several days. Furthermore, they showed that cold-storage sensitive HPA-3a antibodies are also sensitive for neuraminidase treatment suggesting the loss of sialic acid on platelet surface as an explanation for the variable reactivity of HPA-3 antibodies with stored test platelets [22]. However, in the current study, treatment of megakaryocytes with a neuraminidase inhibitor, which inhibits desialylation, did not increase the reactivity of HPA-3 alloantibodies in MAIMA (Fig. 3). This suggests that a mechanism other than desialylation may be important for the variable reactivity of HPA-3 antibodies in MAIMA.
Antibodies against HPA-5 are the second most commonly detected antibodies in patients with FNAIT [15, 16, 26]. The HPA-5 antigen system is located on GPIa (CD49b) [24]. Single nucleotide variation results in a lysine (HPA-5a) to glutamate (HPA-5b) exchange at amino acid residue 505 on GPIa [24]. Alternative sources of cells have been experimented for the detection of HPA-5 alloantibodies [27]. For example, K562 cells expressing HPA-5a or HPA-5b were able to detect anti-HPA-5a and anti-HPA-5b antibodies, respectively [27]. However, buffy coat-derived megakaryocytes in the current study failed to detect anti-HPA-5b antibodies (Fig. 5). Expression of CD49b on megakaryocytes shows a great variation between individuals. Furthermore, CD49b is weakly expressed on immature megakaryocytes [28, 29]. Moussuz et al. [29] demonstrated that the expression of CD49b on megakaryocytes generated from bone marrow stem cells reached up to only 14% after 10 days of differentiation. In our study, we also found a low CD49b expression on megakaryocytes (Fig. 6). This finding explains the lack of reactivity with HPA-5b antibodies in MAIMA.
To our knowledge, this is the first study to investigate the use of buffy coat-derived megakaryocytes for the detection of anti-HPA alloantibodies. Our findings suggest that while buffy coat-derived megakaryocytes can effectively detect anti-HPA-1a antibodies, their reactivity is reduced in the case of HPA-3a antibodies and they are unable to detect HPA-5b antibodies. Overall, our results indicate that while buffy coat-derived megakaryocytes hold promise as a potential alternative to donor platelets, their suitability for the detection of certain HPA antibodies, such as HPA-5b, may be limited.
Acknowledgments
G.U. (CS 3014-0-0) is supported by the MINT-Clinician Scientist program of the Medical Faculty Tübingen, funded by the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation) – 493665037.
Statement of Ethics
The study was conducted in accordance with the Declaration of Helsinki. Written informed consent was obtained from all volunteers prior to any study-related procedure. The study protocol was approved by the Institutional Review Board of the University of Tübingen (134/2019B02).
Conflict of Interest Statement
Tamam Bakchoul has received research funding from Stiftung Transfusionsmedizin und Immunhämatologie e.V. Other authors declare no competing financial interests.
Funding Sources
This work was supported by grants from the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation), from the Stiftung Transfusionsmedizin und Immunhämatologie e.V. and Blood Donation Services of German Red Cross to TB.
Author Contributions
G.U., J.L., I.M., S.N., K.A., and T.B. designed the study. G.U., J.L., I.M., F.R., F.L., O.H., and N.W. performed the experiments. G.U., I.M., F.R., O.H., N.W., S.N., K.A., U.J.S., and T.B. analyzed the data and interpreted the results. G.U., I.M., and T.B. wrote the manuscript. All authors read and approved the manuscript.
Funding Statement
This work was supported by grants from the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation), from the Stiftung Transfusionsmedizin und Immunhämatologie e.V. and Blood Donation Services of German Red Cross to TB.
Data Availability Statement
Data may be requested for academic collaboration from the corresponding author.
Supplementary Material.
References
- 1. Bussel JB, Vander Haar EL, Berkowitz RL. New developments in fetal and neonatal alloimmune thrombocytopenia. Am J Obstet Gynecol. 2021;225(2):120–7. [DOI] [PubMed] [Google Scholar]
- 2. Kovanlikaya A, Tiwari P, Bussel JB. Imaging and management of fetuses and neonates with alloimmune thrombocytopenia. Pediatr Blood Cancer. 2017;64(12). [DOI] [PubMed] [Google Scholar]
- 3. de Vos TW, Winkelhorst D, Árnadóttir V, van der Bom JG, Canals Surís C, Caram-Deelder C, et al. Postnatal treatment for children with fetal and neonatal alloimmune thrombocytopenia: a multicentre, retrospective, cohort study. Lancet Haematol. 2022;9(11):e844–53. [DOI] [PubMed] [Google Scholar]
- 4. Kamphuis MM, Paridaans N, Porcelijn L, De Haas M, van der Schoot CE, Brand A, et al. Screening in pregnancy for fetal or neonatal alloimmune thrombocytopenia: systematic review. BJOG. 2010;117(11):1335–43. [DOI] [PubMed] [Google Scholar]
- 5. Versiti: human platelet antigen (HPA) database. Versiti. https://www.versiti.org/products-services/human-platelet-antigen-hpa-database (accessed January 19, 2023). [Google Scholar]
- 6. Curtis BR, McFarland JG. Human platelet antigens: 2013. Vox Sang. 2014;106(2):93–102. [DOI] [PubMed] [Google Scholar]
- 7. Wen YH, Chen DP. Human platelet antigens in disease. Clin Chim Acta. 2018;484:87–90. [DOI] [PubMed] [Google Scholar]
- 8. Kiefel V. Platelet antibodies in immune thrombocytopenia and related conditions. J Lab Med. 2020;44(5):273–84. [Google Scholar]
- 9. Porcelijn L, Huiskes E, de Haas M. Progress and development of platelet antibody detection. Transfus Apher Sci. 2020;59(1):102705. [DOI] [PubMed] [Google Scholar]
- 10. Zhang N, Santoso S, Aster RH, Curtis BR, Newman PJ. Bioengineered iPSC-derived megakaryocytes for the detection of platelet-specific patient alloantibodies. Blood. 2019;134(22):e1–8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Kanack AJ, Jones CG, Singh B, Leger RR, Splinter NP, Heikal NM, et al. Off-the-shelf cryopreserved platelets for the detection of HIT and VITT antibodies. Blood. 2022;140(25):2722–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Marini I, Rigoni F, Zlamal J, Pelzl L, Althaus K, Nowak-Harnau S, et al. Blood donor-derived buffy coat to produce platelets in vitro. Vox Sang. 2020;115(1):94–102. [DOI] [PubMed] [Google Scholar]
- 13. Hayashi T, Hirayama F. Advances in alloimmune thrombocytopenia: perspectives on current concepts of human platelet antigens, antibody detection strategies, and genotyping. Blood Transfus. 2015;13(3):380–90. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Kiefel V, Santoso S, Weisheit M, Mueller-Eckhardt C. Monoclonal antibody: specific immobilization of platelet antigens (MAIPA): a new tool for the identification of platelet-reactive antibodies. Blood. 1987;70(6):1722–6. [PubMed] [Google Scholar]
- 15. Kroll H, Yates J, Santoso S. Immunization against a low-frequency human platelet alloantigen in fetal alloimmune thrombocytopenia is not a single event: characterization by the combined use of reference DNA and novel allele-specific cell lines expressing recombinant antigens. Transfusion. 2005;45(3):353–8. [DOI] [PubMed] [Google Scholar]
- 16. Davoren A, Curtis BR, Aster RH, McFarland JG. Human platelet antigen-specific alloantibodies implicated in 1162 cases of neonatal alloimmune thrombocytopenia. Transfusion. 2004;44(8):1220–5. [DOI] [PubMed] [Google Scholar]
- 17. Newman PJ, Derbes RS, Aster RH. The human platelet alloantigens, PlA1 and PlA2, are associated with a leucine33/proline33 amino acid polymorphism in membrane glycoprotein IIIa, and are distinguishable by DNA typing. J Clin Invest. 1989;83(5):1778–81. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Lyman S, Aster RH, Visentin GP, Newman PJ. Polymorphism of human platelet membrane glycoprotein IIb associated with the Baka/Bakb alloantigen system. Blood. 1990;75(12):2343–8. [PubMed] [Google Scholar]
- 19. Glade-Bender J, McFarland JG, Kaplan C, Porcelijn L, Bussel JB. Anti-HPA-3A induces severe neonatal alloimmune thrombocytopenia. J Pediatr. 2001;138(6):862–7. [DOI] [PubMed] [Google Scholar]
- 20. Hayashi T, Amakishi E, Matsuyama N, Yasui K, Furuta RA, Hori Y, et al. Establishment of a cell line panel as an alternative source of platelet antigens for a screening assay of anti-human platelet antibodies. Transfus Med. 2011;21(3):199–204. [DOI] [PubMed] [Google Scholar]
- 21. Harrison CR, Curtis BR, McFarland JG, Huff RW, Aster RH. Severe neonatal alloimmune thrombocytopenia caused by antibodies to human platelet antigen 3a (Baka) detectable only in whole platelet assays. Transfusion. 2003;43(10):1398–402. [DOI] [PubMed] [Google Scholar]
- 22. Socher I, Zwingel C, Santoso S, Kroll H. Heterogeneity of HPA-3 alloantibodies: consequences for the diagnosis of alloimmune thrombocytopenic syndromes. Transfusion. 2008;48(3):463–72. [DOI] [PubMed] [Google Scholar]
- 23. Kataoka S, Kobayashi H, Chiba K, Nakamura M, Shinada S, Morita S, et al. Neonatal alloimmune thrombocytopenia due to an antibody against a labile component of human platelet antigen-3b (Bak b). Transfus Med. 2004;14(6):419–23. [DOI] [PubMed] [Google Scholar]
- 24. Santoso S, Kalb R, Walka M, Kiefel V, Mueller-Eckhardt C, Newman PJ. The human platelet alloantigens Br(a) and Brb are associated with a single amino acid polymorphism on glycoprotein Ia (integrin subunit alpha 2). J Clin Invest. 1993;92(5):2427–32. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Cho J, Kim H, Song J, Cheong J-W, Shin JW, Yang WI, et al. Platelet storage induces accelerated desialylation of platelets and increases hepatic thrombopoietin production. J Transl Med. 2018;16(1):199. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Alm J, Duong Y, Wienzek-Lischka S, Cooper N, Santoso S, Sachs UJ, et al. Anti-human platelet antigen-5b antibodies and fetal and neonatal alloimmune thrombocytopenia; incidental association or cause and effect? Br J Haematol. 2022;198(1):14–23. [DOI] [PubMed] [Google Scholar]
- 27. Hayashi T, Amakishi E, Matsuyama N, Yasui K, Furuta RA, Hori Y, et al. Detection of anti-human platelet antibodies against integrin α2β1 using cell lines. Blood Transfus. 2014;12(Suppl 1):s273–80. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Lagrue-Lak-Hal AH, Debili N, Kingbury G, Lecut C, Le Couedic JP, Villeval JL, et al. Expression and function of the collagen receptor GPVI during megakaryocyte maturation. J Biol Chem. 2001;276(18):15316–25. [DOI] [PubMed] [Google Scholar]
- 29. Mossuz P, Schweitzer A, Molla A, Berthier R. Expression and function of receptors for extracellular matrix molecules in the differentiation of human megakaryocytes in vitro. Br J Haematol. 1997;98(4):819–27. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
Data may be requested for academic collaboration from the corresponding author.