Abstract
Background and aims
Hepatocellular carcinoma (HCC) is a leading cause of cancer-related mortality worldwide, and its etiology involves a complex interplay of genetic and environmental factors. Despite advancements in our understanding of HCC biology and the development of novel therapeutic strategies, the molecular mechanisms underlying its onset, progression, and resistance to therapy remain largely vague. This study aimed to investigate the role of mechanosensitive ion channel-related genes (MICRGs) in HCC, focusing on their potential as prognostic biomarkers and their involvement in immune modulation and drug resistance.
Methods
A comprehensive analysis was conducted using The Cancer Genome Atlas database to identify MICRGs that are upregulated in HCC. Gene expression profiling, bioinformatics tools, and functional experiments were employed to elucidate the role of these channels. In addition, protein–protein interaction (PPI) network analyses and enrichment analyses were performed to explore the biological significance of these genes. An immune cell infiltration analysis was also conducted to understand MICRG-related immune landscape. Single-cell RNA sequencing (scRNA-seq) data were utilized to identify MICRGs in different cell types within the HCC tissue. Deep-learning neural network analysis across patient cohorts was conducted to identify genes associated with sorafenib resistance. Knockdown experiments, cell viability assays, and apoptosis assays on HCC cell lines were performed to examine the role of Piezo-type mechanosensitive ion channel component 1 (PIEZO1) in sorafenib resistance.
Results
The analysis identified a subset of MICRGs, including PIEZO1, that were significantly upregulated in HCC and associated with poor prognosis. The PPI network analysis revealed complex interactions among these genes. Gene Ontology and Kyoto Encyclopedia of Genes and Genomes pathway enrichment analyses proposed the involvement of these genes in calcium signaling pathways. Immune cell infiltration analysis demonstrated distinct associations between MICRGs and various immune subpopulations, highlighting their potential roles in immune modulation. scRNA-seq data indicated the upregulation of MICRGs in various cell types in HCC tissues, particularly in endothelial cells and tumor-associated macrophages. Deep-learning neural network analysis across different patient cohorts identified PIEZO1 as a crucial regulator of sorafenib resistance in HCC, which was further validated by functional assays on HCC cell lines.
Conclusions
This study provides evidence that MICRGs, particularly PIEZO1, take on crucial roles in HCC progression and drug resistance. The upregulation of PIEZO1 in HCC cells is associated with poor prognosis and resistance to sorafenib. These findings indicate that PIEZO1 could serve as a potential therapeutic target for overcoming drug resistance and a prognostic biomarker in HCC. Future studies should focus on validating these findings in larger patient cohorts and exploring the functional implications of targeting PIEZO1 in preclinical models.
Keywords: Hepatocellular carcinoma (HCC), Mechanosensitive ion channel, Piezo-type mechanosensitive ion channel component 1 (PIEZO1), Drug resistance, Prognosis, Immune infiltration
1. Introduction
Hepatocellular carcinoma (HCC) is a leading cause of cancer-related mortality worldwide, and its etiology involves a complex interplay of genetic and environmental factors that promote its development and metastasis.1 Despite considerable advancements in our understanding of HCC biology and the development of novel therapeutic strategies, the molecular mechanisms underlying its onset, progression, and resistance to therapy remain largely unclear.2 Sorafenib, a multikinase inhibitor, is commonly used as a first-line treatment for patients with advanced HCC who are not eligible for surgery or liver transplantation.3 However, sorafenib resistance often develops in patients with HCC, reducing its effectiveness as first-line therapy and presenting a significant challenge in HCC treatment.4 In recent years, immunotherapies such as immune checkpoint inhibitors have emerged as a promising strategy for HCC treatment. However, a poor response to immunotherapies is observed in HCC, which is attributed to the immunosuppressive tumor microenvironment (TME).5 Understanding the underlying mechanisms of resistance to chemotherapy and immunotherapy is crucial for the development of strategies to overcome resistance and improve treatment outcomes for patients with HCC.
The mechanical microenvironment of solid tumors is characterized by high extracellular matrix (ECM) stiffness, solid stress induced by rapid tumor growth, and high interstitial fluid pressure caused by solid stress and fluid accumulation in the interstitial space. These mechanical forces in the TME play a critical role in the development and progression of various cancers,6 including HCC.7 High ECM stiffness promotes the proliferation, migration, and invasion of cancer cells.8 It was also found to induce chemotherapy resistance by activating signaling pathways that enhance cell survival.8
Mechanosensitive ion channels (MICs) are a type of ion channels that are activated by mechanical forces such as stretching, compression, and fluid shear stress.9 These channels are found in various cells and take on a crucial role in converting mechanical stimuli into electrical signals, which can then trigger cellular responses. Eukaryotic MICs include Piezo-type MIC component (PIEZO) family, transient receptor potential (TRP) superfamily, transmembrane channel-like (TMC) family, TWIK-related potassium (TREK)-1/2, TRAAK, and OSCA/TMEM63 channels. Among these MICs, PIEZO1/2, TREK-1/2, TRAAK, and OSCA/TMEM63 are primary mechanotransducers, and others are considered to function downstream of primary mechanotransducers.10 MICs are critical regulators of cellular processes including proliferation, migration, and differentiation.11 Targeting MICs may be a promising strategy for solid tumor treatment. In HCC, the dysregulation of these channels may result in abnormal cellular behaviors that promote tumor growth.12 Previous studies have shown that PIEZO1 promotes HCC tumor growth, epithelial–mesenchymal transition (EMT), and angiogenesis.13 However, whether other MICs and their related genes also contribute to HCC development and drug resistance remains unclear. In this study, gene expression profiling and bioinformatics tools were employed to investigate the role of MICs and MIC-related genes (MICRGs) in HCC and examine their potential contribution to prognosis, TME modulation, and drug resistance.
2. Materials and methods
2.1. Ethical approval
This study was approved by the Medical Ethics Committee of The Third Affiliated Hospital of Sun Yat-sen University (Approval No. RG2023-291-02). This study complied with the guidelines of the Declaration of Helsinki of the World Medical Association. Participation was voluntary, and written informed consent was obtained from each patient.
2.2. Data collection and preprocessing
RNA sequencing (RNA-seq) data and associated clinical information for this study were obtained from The Cancer Genome Atlas (TCGA) (https://portal.gdc.cancer.gov/), comprising 374 patients with HCC and 50 normal tissue samples. The data included gene expression profiles, immune infiltration metrics, and clinical characteristics. For subsequent analyses, RNA-seq data were converted from the fragments per kilobase of transcript per million mapped reads (FPKM) to the transcripts per million (TPM) format, with clinical data retained and aligned with TCGA’s publication guidelines.
2.3. Identifying MICRGs
Genes associated with “MICs” were identified by querying the GeneCards database (https://www.genecards.org/) and selecting the top 62 genes with a relevance score >20. A heatmap was then generated to visualize the expression patterns of these genes across the HCC and normal tissue samples.
2.4. Protein–protein interaction (PPI) network and enrichment analyses
A PPI network of coregulated MICRGs was constructed using the Search Tool for the Retrieval of Interacting Genes (STRING) database (https://cn.string-db.org/), and visualization was performed using R software version 4.3.1 (https://www.r-project.org/).14 An interaction confidence score threshold of 0.4 was applied, where higher scores indicate more reliable PPI relationships. To explore the potential biological functions and pathways of the MICRGs, Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analyses were performed using the “clusterProfiler” package in R.15, 16, 17 GO analysis encompassed the biological process (BP), molecular function (MF), and cellular component (CC). Both GO and KEGG analyses utilized a hypergeometric test to assess enrichment significance, with P-value adjusted for multiple testing using the Benjamini–Hochberg method.
2.5. Univariate analysis
To evaluate the relationship between the 62 candidate genes and the prognosis of patients with HCC, univariate Cox proportional hazards regression analysis was initially conducted for each gene. In the univariate analysis, the independent variable was the expression level of each gene, whereas the dependent variable represented the patient’s survival time. For each gene, hazard ratios (HRs) were calculated along with their corresponding 95% confidence intervals (CIs). Subsequently, for genes exhibiting significance in the univariate Cox regression analysis, Kaplan–Meier survival analysis was performed. The expression levels of these genes were classified into high and low-expression groups according to the median value. Kaplan–Meier survival curves were constructed, and the log-rank test was employed to compare survival differences between the high- and low-expression groups. Kaplan–Meier curves were employed to assess the effect of gene expression on the prognosis of patients with HCC.
2.6. Immune cell infiltration analysis
To explore the immune landscape associated with the selected genes, an in-depth immune infiltration analysis was conducted using single-sample gene set enrichment analysis (ssGSEA) and CIBERSORT algorithms.18, 19, 20 These methods enabled us to investigate the relationship between the seven genes of interest and the immune cell types within the TME. Specifically, ssGSEA was employed to infer immune cell infiltration levels based on gene expression data, providing a comprehensive view of the immune context surrounding the studied genes.21 Concurrently, CIBERSORT was used to estimate the relative abundance of immune cell subsets, providing a quantitative perspective on immune cell composition.
Furthermore, to understand the functional implications of the seven genes within the immune context, Spearman correlation analysis was conducted. This analysis assessed the association between the expression levels of the seven genes and the infiltration levels of immune cells, as inferred by ssGSEA and CIBERSORT. The Spearman correlation coefficient provided a measure of the strength and direction of these relationships, revealing potential immune-related functions of the studied genes. In addition, our analysis was expanded to include the correlation between the examined genes and the expression of immune checkpoint genes. This aimed to explore the potential immunomodulatory roles of the studied genes, which could have implications for immunotherapy strategies. The Spearman correlation analysis helped quantify these relationships, providing insights into the potential interactions between the studied genes and the immune checkpoint molecules.
2.7. Basic analysis workflow of single-cell RNA-seq (scRNA-seq) data
In scRNA-seq data quality control process, cells that expressed at least 200 genes and had mitochondrial gene counts <10% of the total gene counts were retained. Dimensionality reduction and clustering analysis were performed using the Seurat package. To address batch effects that might compromise the precision of the single-cell analysis, the harmony package was employed for batch effect correction, focusing on the top 2000 variable genes with default harmony parameters. Subsequently, cell cluster partitioning with an optimal resolution set at 0.5 was conducted to visualize the clustering results. Liver marker genes were then downloaded from the ACT database (http://xteam.xbio.top/ACT/index.jsp) for manual annotation. The cluster annotation information was calculated for each cell using UCell algorithm, based on the marker gene set of each cell.
2.8. Deep-learning neural network analysis
In this study, TensorFlow was utilized to develop a deep-learning model designed to identify genes associated with sorafenib resistance in HCC. Gene expression profiles were curated and normalized from six cohorts using GSE109211 as the foundational training set. The model was configured with various parameters including batch sizes of 5 and 10; learning rates of 0.001, 0.005, and 0.010; dropout rates at 0.25 and 0.50; classification thresholds at 0.25, 0.50, and 0.75; and layer neuron configurations of 32, 16, 8, and 1. L2 regularization with lambda values of 0 and 0.001 was employed, and the model was trained for 50 epochs. To evaluate the model, SHAP analysis was used to understand gene contributions, performance metrics and loss curves were monitored, and receiver operating characteristic (ROC) curves were generated to assess the model’s predictive accuracy in distinguishing sorafenib-resistant cases.
2.9. Collection of human specimens
Between January 2022 and December 2023, 22 paired tumor and adjacent nontumoral liver tissues were collected from patients who underwent curative surgical resection for HCC at The Third Affiliated Hospital of Sun Yat-sen University, Guangzhou, China. Samples were snap-frozen in liquid nitrogen and stored at −80 °C. Pathological diagnosis of HCC was confirmed by histology.
2.10. Cell lines and cell culture
Hep3B, PLC/PRF/5, and HEK-293T cells were obtained from the American Type Culture Collection (ATCC, Manassas, VA, USA) and cultured in DMEM (Gibco, Grand Island, NY, USA) supplemented with 10% fetal bovine serum (ExCell Bio, Suzhou, Jiangsu, China) and 100 IU/mL penicillin/streptomycin (Gibco, Grand Island, NY, USA). All cells were incubated at 37 °C with 5% CO2 and regularly tested for mycoplasma contamination.
2.11. PIEZO1 knockdown with lentivirus-mediated short hairpin RNA (shRNA)
The transfection was performed using Lipofectamine 3000 (Invitrogen, Carlsbad, CA, USA). For gene knockdown, viruses were generated in HEK-293T cells through cotransfection of the expression vector (pLKO.1-puro vector, Addgene, Watertown, MA, USA) with the lentiviral packaging system (pMD2.G and psPAS2). shRNA sequences targeting PIEZO1 (shPIEZO1#1: 5′-CTCACCAAGAAGTACAATCAT-3′ or shPIEZO1#2: 5′-GCTGCTCTGCTACTTCATCAT-3′) were inserted into the multiple cloning sites of pLKO.1-puro vector. Sixteen hours after transfection, the medium was replaced with a fresh HEK-293T cell growth medium, and the virus-containing supernatant was collected and filtered through 0.45 μm polyethersulfone (PES) filters 48 and 72 h after transfection. For transduction, target cells were seeded 16 h earlier and then infected with lentivirus. Forty-eight hours after transduction, puromycin was added to the culture medium for the selection of transduced cells.
2.12. Total RNA isolation and reverse transcription-quantitative polymerase chain reaction (RT-qPCR) assay
Total RNA was isolated using the RNA Quick Purification kit (Esscience Biotech, Shanghai, China) and quantified by the NanoDrop One (Thermo Fisher Scientific, Waltham, MA, USA). Complemetary DNA (cDNA) was synthesized from 1 μg of total RNA using the RT reagent Kit (Vazyme, Nanjing, China) following the manufacturer’s instruction. qPCR was performed with ChamQ Universal SYBR qPCR Master Mix (Vazyme, Nanjing, China) on the LightCycler 480 system (Roche, Switzerland). ΔΔCt method was used to determine the relative amounts of PCR products generated from each primer set. The primer sequences used for qPCR are listed in Table 1.
Table 1.
List of primers used in qPCR.
| Gene name | Forward primer sequence (5′ - 3′) | Reverse primer sequence (5′ - 3′) |
|---|---|---|
| GAPDH | ACAGTTGCCATGTAGACC | TTTTTGGTTGAGCACAGG |
| PIEZO1 | ATCGCCATCATCTGGTTCCC | TGGTGAACAGCGGCTCATAG |
| CLIC1 | CAGTCCCAGCAACCCAGAAT | GAACTGGGAGGGGAGCTTG |
| CALM1 | TTGACTTCCCCGAATTTTTGACT | GGAATGCCTCACGGATTTCTT |
| VDAC2 | GCTACAGGACTGGGGACTTC | AATGCCAAAACGAGTGCAGTT |
Abbreviations: CALM1, calmodulin 1; CLIC1, chloride intracellular channel 1; PIEZO1, Piezo-type mechanosensitive ion channel component 1; qPCR, quantitative polymerase chain reaction; VDAC2, voltage-dependent anion channel 2.
2.13. Western blotting
Cells were lysed in radioimmunoprecipitation assay (RIPA) lysis buffer (Beyotime, Nanjing, China) containing protease and phosphatase inhibitors. Protein concentration was determined using a BCA protein assay kit (Beyotime, Nanjing, China). Equal amounts of protein (20 μg) were loaded onto a 6%–10% sodium dodecyl sulfate-polyacrylamide gel electrophoresis and electrophoresed at 150 V for 90 min. Proteins were then transferred onto a polyvinylidene fluoride membrane (Merck Millipore, Darmstadt, Germany) at 100 V for 1.5 h using a wet transfer system (Bio-Rad, Hercules, CA, USA). The membrane was blocked with 5% non-fat milk in TBST (TBS containing Tween-20) for 1 h at room temperature and then incubated with primary antibodies overnight at 4 °C. β-Actin (1:5000, #66009-1-Ig, Proteintech, Wuhan, China) and PIEZO1 (1:500, #NBP1-78537, Novus International, St. Louis, MO, USA) were used as primary antibodies in the study. The membrane was washed three times with TBST and incubated with horseradish peroxidase-conjugated secondary antibodies (Cell Signaling Technology, Danvers, MA, USA) for 1 h at room temperature. Protein bands were detected using a chemiluminescent substrate (#P10300, NCM Biotech, Suzhou, Jiangsu, China) and imaged on Tanon 5200 Chemiluminescent Imaging System (Shanghai, China). Quantitative analysis of protein expression was performed using densitometric analysis of band intensities normalized to a loading control.
2.14. Half-maximal inhibitory concentration (IC50) assay
The IC50 of Hep3B and PLC/PRF/5 cells to sorafenib (#S7397, Selleck, Houston, TX, USA) were evaluated using the Cell Counting Kit-8 (CCK-8) kit (#100-106-500, GOONIE, Guangzhou, Guangdong, China). Cells were seeded at a density of 2000 cells per well in 96-well plates. Subsequently, the cells were treated with 0, 2, 4, 6, 8, 10, 12, 15, and 20 μM sorafenib (#S7397, Selleck). After 48 h of incubation, 10 μL of CCK-8 reagent (#100-106-500, GOONIE) was added to each well containing 100 μL of complete medium. The plates were then incubated for an additional 2 h, and absorbance at 450 nm was measured. The A450 values for each concentration group were normalized and fitted using the nonlinear regression algorithm implemented in GraphPad Prism version 8 (GraphPad Software Inc., La Jolla, CA, USA).
2.15. Analysis of apoptosis by flow cytometry
The Annexin V-fluorescein isothiocyanate (FITC)/propidium iodide (PI) apoptosis detection kit (#KGA1102-100, KeyGEN, Nanjing, Jiangsu, China) was used to assess cell apoptosis. Cells were seeded at a density of 3–4 × 106 cells per well in 6-well plates, treated with 12 μM sorafenib (#S7397, Selleck) for 24 h, harvested, washed with cold phosphate-buffered saline, and resuspended in binding buffer. Annexin V-FITC (5 μL) and PI (5 μL) were added to the cell suspension and then incubated in the dark for 15 min at room temperature. The samples were then analyzed by flow cytometry (Cytoflex LX, Beckman Coulter, Brea, CA, USA), where Annexin V-FITC positivity indicated early apoptosis, and PI staining indicated late apoptosis or necrosis. Results were analyzed using FlowJo data analysis software (TreeStar, USA) to distinguish between viable, early apoptotic, and late apoptotic/necrotic cells.
2.16. Statistical analysis
Statistical analyses for bioinformatic analyses were conducted using R version 4.3.2 (https://www.r-project.org/).22 The Wilcoxon rank-sum test was used to compare the expression levels of MICRGs between the HCC and normal groups. Welch’s one-way analysis of variance (ANOVA) and t-tests were utilized to examine the association between MICRG expression and clinicopathological factors, with subsequent Bonferroni correction. The effect of clinicopathological factors on MICRG expression was analyzed through univariate logistic regression, Fisher’s exact test, and Pearson’s χ2 test. Kaplan–Meier curves were plotted to evaluate the prognostic significance of MICRGs, and individual risk for overall survival (OS) was determined using univariate Cox proportional hazards regression. The pROC package was employed to perform ROC analysis of MICRGs, with the area under the curve (AUC) values ranging from 0.5 to 1.0. This reflects the model’s ability to distinguish between different conditions, where values closer to 1.0 indicate a higher predictive accuracy, and values approximately 0.5 indicate no discriminative power.
GraphPad Prism 8.0 (GraphPad Software Inc., La Jolla, CA, USA) was used for statistical analysis of experimental results. Unpaired two-tailed t-test was used to compare the differences between the two groups, and the one-way ANOVA was used to compare differences among three groups or more. P < 0.05 was considered significant.
3. Results
3.1. MICRG expression levels are significantly high in HCC
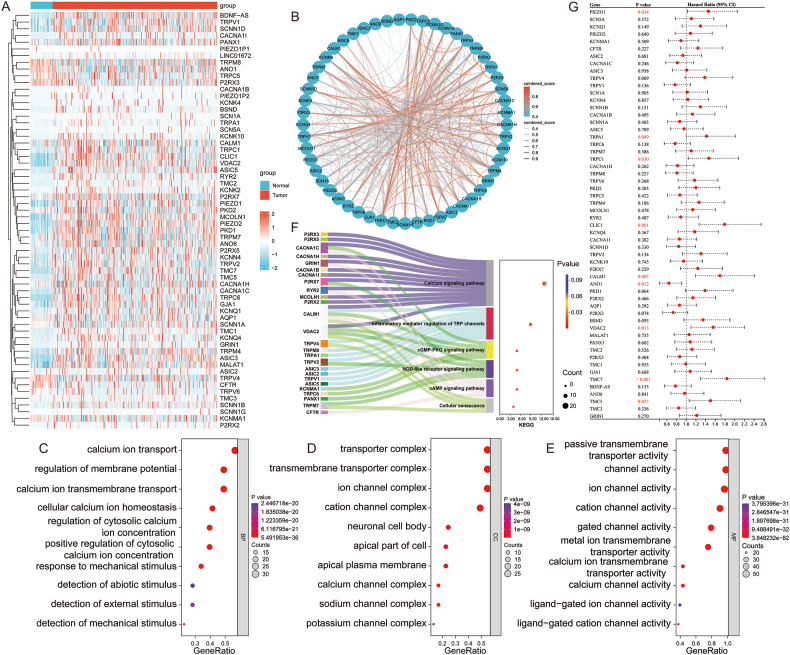
The workflow of this study is presented in Fig. 1. Genes associated with “mechanosensitive ion channels” were identified by querying the GeneCards database, and the top 62 genes with a relevance score >20 were selected, which we defined as MICRGs. The heatmap presented in Fig. 2A illustrates that among these 62 genes, 38 were significantly upregulated in HCC. The box plots depicting the expression patterns of these 62 genes are shown in Supplemental Fig. 1. In addition, a PPI network of the differentially expressed proteins was constructed using STRING. In this network, 53 MICRGs were closely connected, suggesting that they were coregulated in HCC (Fig. 2B). The GO enrichment analysis of these MICRGs revealed that key BPs included calcium ion transport, regulation of membrane transport, and cellular calcium ion homeostasis (Fig. 2C). The CCs were predominantly enriched in transporter complexes, cation channel complexes, and specific membrane regions (Fig. 2D). MFs involved activities such as ion channel and transmembrane transporter activity (Fig. 2E). In addition, KEGG pathway enrichment analysis primarily indicated involvement in the calcium signaling pathway, inflammatory regulation of TRP channels, and cellular senescence pathways (Fig. 2F). To evaluate the relationship among the 62 candidate genes and patient prognosis, a univariate Cox proportional hazards regression model was initially applied to analyze each gene. All candidate genes demonstrated a significant association with OS in HCC, with HRs >1 and P-values <0.05. Following this, Kaplan–Meier survival analysis was conducted on the genes that were significant in the univariate Cox regression analysis, ultimately identifying nine prognostic genes with HRs >1 and P-values <0.05 (Fig. 2G).
Fig. 1.
Flow diagram of the study. In this study, 62 MICRGs were initially screened from GeneCards and the TCGA, leading to the identification of seven notable MICRGs through thermography, enrichment, PPI network, prognostic, and clinical correlation analyses. Subsequently, these genes were subjected to immune infiltration analysis, single-cell analysis, and deep-learning techniques. Finally, in vitro experiments were conducted to verify the role of the PIEZO1 gene in hepatocellular carcinoma cells. Abbreviations: GO, Gene Ontology; KEGG, Kyoto Encyclopedia of Genes and Genomes; MICRGs, mechanosensitive ion channel-related genes; PIEZO1, Piezo-type mechanosensitive ion channel component 1; PPI, protein–protein interaction; TCGA, The Cancer Genome Atlas.
Fig. 2.
Differential analysis of mechanosensitive ion channel-related genes (MICRGs). (A) Heatmap of MICRGs in tumor tissues versus normal tissues. (B) PPI network analysis. (C–F) GO (C–E) and KEGG (F) analyses of MICRGs. (G) Univariate Cox regression analysis of MICRGs. Abbreviations: ANO1, anoctamin 1; BP, biological process; CALM1, calmodulin 1; CC, cellular component; CI, confidence interval; CLIC1, chloride intracellular channel 1; GO, Gene Ontology; KEGG, Kyoto Encyclopedia of Genes and Genomes; MF, molecular function; PIEZO1, Piezo-type mechanosensitive ion channel component 1; PPI, protein–protein interaction; TMC, transmembrane channel-like; TRPA1, transient receptor potential ankyrin 1; TRPC1, transient receptor potential C1; VDAC2, voltage-dependent anion channel 2.
3.2. Association between MICRG expression and disease progression and patient outcomes
Kaplan–Meier survival plots were generated for the nine MICRGs to investigate the association of their expression and the clinical outcomes of patients with HCC (Fig. 3A–I). The results revealed that high expression levels of calmodulin 1 (CALM1), chloride intracellular channel 1 (CLIC1), PIEZO1, TMC5, TMC7, TRPA1, TRPC1, and VDAC2 were linked to poor OS in patients with HCC (Fig. 3B–I). Conversely, ANO1 exhibited an opposite trend, indicating that its high expression is associated with better OS (Fig. 3A).
Fig. 3.
Analysis of the prognostic significance of MICRGs in HCC. (A) A low ANO1 expression was related to worse OS in HCC. (B–I) High CALM1 (B), CLIC1 (C), PIEZO1 (D), TMC5 (E), TMC7 (F), TRPA1 (G), TRPC1 (H), and VDAC2 (I) expression levels were related to worse OS in HCC. Abbreviations: ANO1, anoctamin 1; CALM1, calmodulin 1; CI, confidence interval; CLIC1, chloride intracellular channel 1; HCC, hepatocellular carcinoma; HR, hazard ratio; MICRGs, mechanosensitive ion channel-related genes; OS, overall survival; PIEZO1, Piezo-type mechanosensitive ion channel component 1; TMC, transmembrane channel-like; TRPA1, transient receptor potential ankyrin 1; TRPC1, transient receptor potential C1; VDAC2, voltage-dependent anion channel 2.
Subsequently, the expression patterns of the nine prognostic genes were analyzed in patients with HCC at different clinical stages. The results indicated that ANO1 was highly expressed in stage T1 but exhibited a gradual decrease in expression through stages T2–T4 (Fig. 4A). Conversely, CALM1, CLIC1, PIEZO1, TMC5, and TMC7 showed a pattern of gradually increasing expression as the tumor stages advanced (Fig. 4B–F). TRPA1 maintained consistently low-expression levels across all clinical stages (Fig. 4G). TRPC1 and VDAC2 increased up to stage T3 (Fig. 4H and I). This analysis highlights the differential expression of these genes in relation to disease progression, which could have implications for understanding their roles in HCC.
Fig. 4.
Correlation between MICRGs and tumor stage in HCC. (A)ANO1, (B)CALM1, (C)CLIC1, (D)PIEZO1, (E)TMC5, (F)TMC7, (G)TRPA1, (H)TRPC1, and (I)VDAC2 expressions were correlated with clinical stages. *P < 0.05, **P < 0.01. Abbreviations: ANO1, anoctamin 1; CALM1, calmodulin 1; CLIC1, chloride intracellular channel 1; HCC, hepatocellular carcinoma; MICRGs, mechanosensitive ion channel-related genes; PIEZO1, Piezo-type mechanosensitive ion channel component 1; TMC, transmembrane channel-like; TRPA1, transient receptor potential ankyrin 1; TRPC1, transient receptor potential C1; VDAC2, voltage-dependent anion channel 2.
Based on these findings, this study aimed to identify genes that exhibited significant expression patterns and were strongly associated with poor prognosis in patients with HCC. By intersecting the 38 upregulated genes in HCC with the genes associated with poor prognosis, seven MICRGs were identified, including PIEZO1, TRPC1, CLIC1, CALM1, VDAC2, TMC7, and TMC5 (Supplemental Fig. 2A). Correlation analysis revealed strong positive correlations among the seven hub genes (Supplemental Fig. 2B). The overall performance of these genes as prognostic markers was measured using ROC analysis. The results showed that PIEZO1, CLIC1, VDAC2, and TMC7 had AUC values > 0.8, suggesting their good performance in prognosis (Supplemental Fig. 2C).
3.3. Immune cell correlation analysis of the seven MICRGs reveals distinct associations with various immune subpopulations
Immune cell infiltration, which shapes the tumor immune microenvironment (TIME), is closely related to cancer development, drug resistance, and effect of immunotherapy. An immune infiltration analysis was performed using CIBERSORT and ssGSEA. CIBERSORT revealed significant correlations between specific MICRGs and various immune cell subtypes. Most of the seven MICRGs were negatively correlated with γδ T cells, natural killer (NK) cells, and M1 macrophages, but were positively correlated with M0 macrophages and dendritic cells. PIEZO1 was negatively correlated with CD8+ T, γδ T, and NK cells, whereas positively correlated with M0 macrophage (Fig. 5A). The ssGSEA algorithm analysis showed that most of the seven MICRGs were negatively correlated with cytotoxic, dendritic and Th17 cells, while positively correlated with macrophages, CD56bright NK cells, and Th1 and Th2 cells (Fig. 5B).
Fig. 5.
Correlation of key genes with immune cell infiltration and tumor mutation burden in patients with HCC. (A) The correlation analysis of seven MICRGs with the content of immune cells in patients with HCC by the CIBERSORT algorithm. (B) The correlation analysis of seven MICRGs with the content of immune cells in patients with HCC by the ssGSEA algorithm. *P < 0.05, **P < 0.01, ***P < 0.001. (C) Correlation analysis of seven MICRG expression with immune checkpoint genes. Abbreviations: CALM1, calmodulin 1; CLIC1, chloride intracellular channel 1; HCC, hepatocellular carcinoma; MICRG, mechanosensitive ion channel-related gene; PIEZO1, Piezo-type mechanosensitive ion channel component 1; ssGSEA, single-sample gene set enrichment analysis; TMC, transmembrane channel-like; TRPC1, transient receptor potential C1; VDAC2, voltage-dependent anion channel 2.
A set of 39 immune checkpoint genes based on previous literature for correlation analysis was collected. Notably, PIEZO1 exhibited a positive correlation with all 39 immune checkpoints, highlighting its potential as a central regulator in immune checkpoint pathways. In addition, CALM1 was positively correlated with 38 immune checkpoints, CLIC1 with 37 immune checkpoints, TMC7 and TRPC1 with 38 immune checkpoints, and VDAC2 with 34 immune checkpoints (Fig. 5C). TMC5 was positively correlated with 33 immune checkpoints (Fig. 5C). These results underscore the potential involvement of these genes, particularly PIEZO1, in modulating immune responses within the TME.
3.4. scRNA-seq identifies the upregulated MICRGs in HCC tissues
To further investigate the cell types in which MICRGs contribute to HCC development, the expression of MICRGs across different cell populations in HCC was assessed using scRNA-seq data from peritumoral and HCC tissues. Following quality control procedures to remove suspected doublet cells and low-activity cells (Supplemental Fig. 3), 15,717 cells were included for further analysis. By employing principal component analysis (PCA) for dimension reduction and selecting a resolution of 0.5 based on the total resolution cluster tree (Supplemental Fig. 4), 23 clusters were identified, which were annotated as endothelial cells, hepatocytes, periportal hepatocytes, midzonal hepatocytes, myofibroblasts, mature NK cells, circulating NK cells, macrophages, tumor-associated macrophages, tissue-resident CD8+ αβ memory T cells, CD8+ αβ memory T cells, proliferating CD8+ αβ T cells, lymphoid tissue-inducer cells, B cells, plasma cells, monocytes, hematopoietic stem cells, neutrophils, CD1c-positive myeloid dendritic cells, plasmacytoid dendritic cells, and CD141-positive myeloid dendritic cells, based on the analysis of marker genes (Fig. 6A). The expression profiles of marker genes across different cell clusters, identified through scRNA-seq analysis, are displayed in the heatmap (Supplemental Fig. 5).
Fig. 6.
Identification of the cell type associated with MICRGs in HCC tissues. (A) Visualizing the distribution of 21 cell types in the TME by the UMAP plot. (B) Correlation analysis of MICRGs with cell types in HCC and normal tissues by the ssGSEA algorithm. (C–I) The UMAP plot visualized the expression levels of CALM1 (C), CLIC1 (D), VDAC2 (E), PIEZO1 (F), TRPC1 (G), TMC7 (H), and TMC5 (I) in 21 different cell types. Abbreviations: CALM1, calmodulin 1; CLIC1, chloride intracellular channel 1; HCC, hepatocellular carcinoma; MICRGs, mechanosensitive ion channel-related genes; PIEZO1, Piezo-type mechanosensitive ion channel component 1; ssGSEA, single-sample gene set enrichment analysis; TMC, transmembrane channel-like; TME, tumor microenvironment; TRPC1, transient receptor potential C1; VDAC2, voltage-dependent anion channel 2.
Subsequent analysis of the proportion of cells in HCC and normal tissues revealed that, in comparison to normal tissues, the proportions of tumor-associated macrophages, periportal hepatocytes, midzonal hepatocytes, macrophages, hepatocytes, hematopoietic stem cells, and endothelial cells were increased in HCC (Fig. 6B). Conversely, the proportions of tissue-resident CD8+ αβ memory T cells, proliferating CD8+ αβ T cells, plasma cells, neutrophils, monocytes, mature NK cells, circulating NK cells, CD8+ αβ memory T cells, CD1c-positive myeloid dendritic cells, CD141-positive myeloid dendritic cells, lymphoid tissue-inducer cells, and B cells were all found to be decreased in HCC compared with normal tissues (Fig. 6B). The examination of the expressions of the seven MICRGs in HCC tissues indicated that CALM1, CLIC1, VDAC2, and PIEZO1 showed high expression levels across nearly all cell types (Fig. 6C–F), whereas TRPC1, TMC7, and TMC5 exhibited lower expression specifically in endothelial cells (Fig. 6G–I).
3.5. Remodeling of the intercellular communication in the TME of HCC
In dot plots, distinct expression patterns of signaling pathways were observed across various cellular clusters (Fig. 7A). UMAP plot was used to visualize the distribution of all signaling pathways involved in cell–cell communication across different cell clusters identified the single-cell analysis (Supplemental Fig. 6). Interestingly, MICs, transforming growth factor (TGF)-β signaling pathway, and ECM were highly expressed in endothelial cells and tumor-associated macrophages, suggesting a role of MIC in mediating these signaling pathways in HCC (Fig. 7B–D). Then, the results of intercellular communication analysis show that compared with normal tissues, the interactions between certain cells in the tumor tissues are significantly increased or become more complex, particularly the connections between endothelial cells, B cells, and tumor-associated macrophages (Fig. 7E and F), which appear denser. This change indicates remodeling of cellular communication within the TME, potentially involving tumor growth, immune evasion, and metastasis.
Fig. 7.
Single-cell analysis of intercellular communication networks in tumor microenvironmentof HCC. (A) Dot plots of different signaling pathways for cellular clusters. (B–D) Feature plots showing the expression of mechanosensitive ion channel (B), transforming growth factor-beta (TGF-β) (C), extracellular matrix (ECM) (D) in different cell clusters. (E, F) Intercellular communication between different cell clusters in the normal (E) and tumor (F) tissues.
3.6. Deep-learning neural network analysis identifies PIEZO1 as a central regulator of sorafenib resistance in HCC
Sorafenib, a targeted therapy, is widely used for treating HCC. However, the emergence of drug resistance poses a significant challenge to its therapeutic efficacy. To understand the molecular mechanisms underlying this resistance, the expression of seven candidate genes was analyzed across six distinct sorafenib-resistant datasets (GSE158458, GSE192912, GSE151412, GSE109211, GSE182593, and GSE186280) (Fig. 8A–F). Our findings indicated that PIEZO1 was overexpressed in four datasets, whereas CALM1 showed increased expression in three datasets. CLIC1 exhibited upregulation in one dataset and downregulation in two others, and TMC7 was downregulated in two datasets (Fig. 8A–F). Utilizing a sophisticated deep-learning neural network algorithm, PIEZO1 was identified as the primary contributor to sorafenib resistance among the genes analyzed (Fig. 8G), suggesting its critical role in conferring drug resistance of HCC.
Fig. 8.
Analysis ofMICRGs’ sensitivity to sorafenib. (A–F) The box diagram shows the expression of seven MICRGs across six datasets, including four datasets from cell lines (GSE158458 (A), GSE192912 (B), GSE151412 (C), and GSE186280 (F)) and two datasets from patients (GSE109211 (D) and GSE182593 (E)) resistant to sorafenib. *P< 0.05, **P< 0.01, ***P< 0.001. ns, not significant. (G) The SHAP summary diagram shows that PIEZO1 has the greatest influence on the model output, and shows how the level of the seven genes affects the prediction direction of the model. Abbreviations: CALM1, calmodulin 1; CLIC1, chloride intracellular channel 1; MICRGs, mechanosensitive ion channel-related genes; PIEZO1, Piezo-type mechanosensitive ion channel component 1; SHAP, SHapley Additive exPlanations; TMC, transmembrane channel-like; TRPC1, transient receptor potential C1; VDAC2, voltage-dependent anion channel 2.
3.7. The pivotal role of PIEZO1 in modulating drug resistance and regulating cellular apoptosis in HCC
RT-qPCR validations on specimens of patients with HCC substantiated the heightened expressions of PIEZO1, CLIC1, CALM1, and VDAC2 within the tumorigenic milieu (Fig. 9A–D). To elucidate the functional role of PIEZO1 in HCC, stable PIEZO1-knockdown Hep3B and PLC/PRF/5 cell lines were established. The knockdown efficiency of PIEZO1 was confirmed through RT-qPCR and Western blot analyses (Fig. 9E–I). Drug susceptibility assays were conducted on control and PIEZO1-knockdown Hep3B and PLC/PRF/5 cells treated with sorafenib at a series of concentrations. A decrease in IC50 indicated enhanced sensitivity to sorafenib in PIEZO1-knockdown cells (Fig. 9J–M). The apoptosis assay showed that PIEZO1 knockdown led to increased basal and sorafenib-induced apoptosis in both Hep3B and PLC/PRF/5, suggesting a proapoptotic role of PIEZO1 knockdown (Fig. 9N–Q). These results indicated PIEZO1 as a critical determinant for drug resistance in HCC and a potential therapeutic target for augmenting chemotherapeutic responses.
Fig. 9.
The PIEZO1 expression in HCC and its clinical significance. (A–D) The mRNA expressions of PIEZO1 (A), CLIC1 (B), CALM1 (C), and VDAC2 (D) in adjacent and HCC tissues (n = 22). (E–I)PIEZO1 knockdown in Hep3B and PLC/PRF/5 cells was detected by RT-qPCR (E and F) and Western blot (G–I). (J–M) Validation of IC50 values of sorafenib using a concentration range in Hep3B (J and K) and PLC/PRF/5 (L and M) cell lines. (N–P) Apoptosis of Hep3B (N and O) and PLC/PRF/5 (P and Q) cells after PIEZO1 knockdown was detected using the Annexin-V-FITC/PI apoptosis detection kit. *P < 0.05, **P < 0.01, ∗∗∗P < 0.001. Abbreviations: CALM1, calmodulin 1; CLIC1, chloride intracellular channel 1; FITC, fluorescein isothiocyanate; HCC, hepatocellular carcinoma; PI, propidium iodide; PIEZO1, Piezo-type mechanosensitive ion channel component 1; RT-qPCR, reverse transcription-quantitative polymerase chain reaction; VDAC2, voltage-dependent anion channel 2.
4. Discussion
Identifying novel biomarkers and therapeutic targets is important in HCC research.23 MICs transduce mechanical stimuli into biochemical responses and play a crucial role in modulating cell functions such as proliferation, migration, and apoptosis.24 MICs have been implicated in tumor growth, metastasis, and resistance to therapy.25 In this study, a comprehensive analysis of MICs and their related genes in HCC were conducted, providing insights into their role in disease progression and potential as therapeutic targets.
Seven MICRGs were identified, namely PIEZO1, TRPC1, CLIC1, CALM1, VDAC2, TMC7, and TMC5, which were upregulated in HCC tumor tissue, indicating their involvement in oncogenic processes. PIEZO1, a mechanosensitive cation channel, can mediate Ca2+ influx upon various types of mechanical stimulation.26 A previous research indicated that PIEZO1 could influence the proliferation, migration, and invasion of HCC cells.27 It was hypothesized that overexpression of PIEZO1 may be associated with alterations in the mechanical microenvironment of HCC.13 TRPC1 is a calcium channel and its upregulation in HCC is associated with reduced patient survival rates.28 CLIC1 is an intracellular chloride ion channel that plays a significant role in regulating cell volume, proliferation, and migration.29 CALM1, a key modulator in the calcium signaling pathway, influences tumor cell proliferation, migration, and metabolism.30 VDAC2 regulates cellular metabolism and drug resistance.31 TMC7 and TMC5 participate in ion transport and cellular signal transduction.32,33 Among these seven MICRGs, PIEZO1, TRPC1, and CLIC1 influence key processes in HCC cells, such as proliferation, migration, invasion, and apoptosis, and their abnormal expression is often linked to poorer patient outcomes,27,34 whereas the role of the other four MICRGs in HCC remains to be elucidated. This study suggests that these MICRGS may represent promising targets for HCC prognosis and therapy.
The TIME of HCC is highly immunosuppressive, and many patients with HCC have poor responses to immunotherapies such as immune checkpoint inhibitors.5 This unveils the mechanism that regulating HCC TIME is crucial to improve the effect of immunotherapies. A recent study showed an immunomodulatory role of ECM stiffness in cancer.35 High ECM stiffness not only impairs immune signaling in tumor cells but also inhibits T-cell infiltration.36,37 High ECM stiffness also promotes the exhaustion of tumor-specific CD8+ T cells through the PIEZO1/CaMKII/CREB pathway.38 This study found a significant correlation between MICRGs and immune cell infiltration in HCC, proposing their potential roles in modulating the TIME. Moreover, PIEZO1 was positively correlated with all the immune checkpoints, indicating its role in modulating immune cell functions. Our findings propose that PIEZO1 and other MICRGs might mediate the effect of the ECM on the TIME, and intervention with MICRGs might boost the effect of immunotherapy.
scRNA-seq data identified the upregulation of MICRGs across 21 different cell types in HCC, including hepatocytes, immune cells, and endothelial cells, with particularly high expression observed in tumor-associated macrophages and endothelial cells, signifying that MICRGs might be involved not only in immune modulation but also angiogenesis in HCC. MICRGs, along with TGF-β and ECM-related proteins, were highly expressed in tumor-associated macrophages and endothelial cells. These concur with the knowledge that high ECM stiffness and TGF-β promote immunosuppressive TIME and tumor angiogenesis.35,39, 40, 41, 42 The simultaneous upregulation of MICRGs, TGF-β, and ECM-related proteins in specific cell clusters recommends that MICRGs may play a role in mediating the effect of the ECM and TGF-β on the TIME and angiogenesis in tumors.
Stiff ECM can contribute to drug resistance in cancer. Therefore, targeting ECM stiffness and mechanotransducers is a way to enhance cancer therapy.43 Higher ECM stiffness decreased the sensitivity of HCC cells to cisplatin.8 MICs contribute to cancer drug resistance by allowing tumor cells to adapt to mechanical stresses in the TME, thereby forming mechanical memory through epigenetic changes.44 This study also suggested that MICRGs play a role in drug resistance in HCC. Particularly, PIEZO1 affected the response to sorafenib of HCC cells. PIEZO1 was significantly upregulated in sorafenib-resistant HCC cells, and in vitro experiments showed that its suppression increased cell sensitivity to sorafenib and promoted apoptosis. This indicates that PIEZO1 not only plays a critical role in HCC development and progression but also serves as a potential therapeutic target for overcoming drug resistance and enhancing chemotherapy efficacy. Several possible mechanisms exist by which PIEZO1 affects drug resistance. First, PIEZO1 regulates the Wnt signaling pathway in both normal and cancerous tissues.45,46 The Wnt pathway is crucial to the stemness and EMT of cancer cells, which promote drug resistance.47 Second, PIEZO1 was also reported to promote proliferation and EMT through the Yes-associated protein (YAP) and TGF-β pathways.12,13 As shown in our study, PIEZO1 might influence the TIME, which is an important factor that contributes to drug resistance.13 Therefore, PIEZO1 may affect drug resistance by promoting stemness, EMT, and modulating TIME.
Overall, a comprehensive analysis was performed to explore the role of MICRGs in HCC. However, this study has certain limitations. First, while our bioinformatic analysis provides a robust foundation for identifying MICRGs in HCC, the retrospective design limits our ability to establish causal relationships. Second, functional experiments were performed in vitro, which might not capture the complexity of the TME in vivo. Although PIEZO1 was identified as a potential therapeutic target, the validation of this finding in preclinical models and clinical samples is limited. Future studies with larger patient cohorts and multicenter collaborations are necessary to validate our findings. Lastly, while the role of MICRGs in drug resistance and immune modulation was explored, the underlying molecular mechanisms require further investigation, particularly in the context of combinatorial therapies. Addressing these limitations will be crucial for advancing the understanding of MICRGs in HCC and development of targeted therapies.
5. Conclusions
In conclusion, this study presents a comprehensive analysis of MICRGs, particularly PIEZO1, in HCC progression and drug resistance. The upregulation of PIEZO1 in HCC cells is associated with poor prognosis and resistance to sorafenib, revealing its potential as prognostic marker and therapeutic target in HCC. Future studies should validate these findings in larger cohorts and explore the functional consequences of targeting these channels in preclinical models. In addition, understanding the precise molecular mechanisms by which these channels exert effects in HCC will be essential for developing effective therapeutic strategies.
Data availability statement
All the data generated and analyzed in this study are available from the corresponding authors upon reasonable request. Some data can be obtained without restriction from online databases including TCGA, STRING, and GEO.
Authors’ contributions
Xinyan Huo: Writing – review & editing, Writing – original draft, Investigation. Shiyu Jiang: Writing – review & editing, Writing – original draft, Investigation. Sihuang Wu: Writing – review & editing, Writing – original draft, Software, Data curation. Qinghai Lian: Writing – review & editing, Writing – original draft, Visualization, Validation, Software, Resources, Formal analysis. Hui Chen: Writing – review & editing, Writing – original draft, Supervision, Project administration, Methodology, Funding acquisition.
Declaration of competing interest
The authors declare that they have no conflict of interest.
Acknowledgements
This study was supported by the Natural Science Foundation of Guangdong Province, China (No. 2023A1515012686 and No. 2024A1515012988), Natural Science Foundation of China (No. 82470904), Science and Technology Program of Guangzhou, China (No. 202102010289), and Fostering Fund of The Third Affiliated Hospital of Sun Yat-Sen University for NSFC Program (No. 2022GZRPYMS01).
Footnotes
Peer review under the responsibility of Editorial Office of Liver Research.
Supplementary data to this article can be found online at https://doi.org/10.1016/j.livres.2025.01.002.
Contributor Information
Qinghai Lian, Email: lianqh5@mail.sysu.edu.cn.
Hui Chen, Email: chenh567@mail.sysu.edu.cn.
Appendix A. Supplementary data
The following are the Supplementary data to this article:
Supplemental Fig. 1.
mRNA expression levels of mechanosensitive ion channel-related genes (MICRGs) in tumor tissues and normal tissues.∗P< 0.05, ∗∗P< 0.01, ∗∗∗P< 0.001.
Supplemental Fig. 2.
Analysis of MICRGs associated with prognosis in hepatocellular carcinoma (HCC). (A) Venn diagram displaying seven genes shared by upregulated genes and unfavorable prognostic genes. (B) Correlation between the seven hub genes. (C) Receiver operating characteristic (ROC) analysis of PIEZO1, TRPC1, CLIC1, CALM1, VDAC2, TMC7, and TMC5 in HCC. Abbreviations: CALM1, calmodulin 1; CLIC1, chloride intracellular channel 1; MICRGs, mechanosensitive ion channel-related genes; PIEZO1, Piezo-type mechanosensitive ion channel component 1; TMC, transmembrane channel-like; TRPC1, transient receptor potential C1; VDAC2, voltage-dependent anion channel 2.
Supplemental Fig. 3.
Quality control results of single cell analysis. (A, B) Quality control of non-tumor (A) and tumor (B) tissue single cells in patients with hepatocellular carcinoma (HCC).
Supplemental Fig. 4.
Cluster tree of single cell transcriptome data set of hepatocellular carcinoma (HCC) tissues.
Supplemental Fig. 5.
The heatmap displays the expression profiles of marker genes across different cell clusters identified by single-cell RNA sequencing analysis.
Supplemental Fig. 6.
The UMAP plot visualizes the distribution of all signaling pathways involved in cell-cell communication across different cell clusters identified by the single-cell analysis.
References
- 1.Lee T.K., Guan X.Y., Ma S. Cancer stem cells in hepatocellular carcinoma - from origin to clinical implications. Nat Rev Gastroenterol Hepatol. 2022;19:26–44. doi: 10.1038/s41575-021-00508-3. [DOI] [PubMed] [Google Scholar]
- 2.Shen P., Jia Y., Zhou W., et al. A biomimetic liver cancer on-a-chip reveals a critical role of LIPOCALIN-2 in promoting hepatocellular carcinoma progression. Acta Pharm Sin B. 2023;13:4621–4637. doi: 10.1016/j.apsb.2023.04.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Chen L.T., Martinelli E., Cheng A.L., et al. Pan-Asian adapted ESMO Clinical Practice Guidelines for the management of patients with intermediate and advanced/relapsed hepatocellular carcinoma: a TOS-ESMO initiative endorsed by CSCO, ISMPO, JSMO, KSMO, MOS and SSO. Ann Oncol. 2020;31:334–351. doi: 10.1016/j.annonc.2019.12.001. [DOI] [PubMed] [Google Scholar]
- 4.Tang W., Chen Z., Zhang W., et al. The mechanisms of sorafenib resistance in hepatocellular carcinoma: theoretical basis and therapeutic aspects. Signal Transduct Target Ther. 2020;5:87. doi: 10.1038/s41392-020-0187-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Kurebayashi Y., Ojima H., Tsujikawa H., et al. Landscape of immune microenvironment in hepatocellular carcinoma and its additional impact on histological and molecular classification. Hepatology. 2018;68:1025–1041. doi: 10.1002/hep.29904. [DOI] [PubMed] [Google Scholar]
- 6.Zhou J., Valentini E., Boutros M. Microenvironmental innate immune signaling and cell mechanical responses promote tumor growth. Dev Cell. 2021;56:1884–1899 (e5). doi: 10.1016/j.devcel.2021.06.007. [DOI] [PubMed] [Google Scholar]
- 7.Roy A.M., Iyer R., Chakraborty S. The extracellular matrix in hepatocellular carcinoma: mechanisms and therapeutic vulnerability. Cell Rep Med. 2023;4 doi: 10.1016/j.xcrm.2023.101170. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Schrader J., Gordon-Walker T.T., Aucott R.L., et al. Matrix stiffness modulates proliferation, chemotherapeutic response, and dormancy in hepatocellular carcinoma cells. Hepatology. 2011;53:1192–1205. doi: 10.1002/hep.24108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Karska J., Kowalski S., Saczko J., Moisescu M.G., Kulbacka J. Mechanosensitive ion channels and their role in cancer cells. Membranes (Basel) 2023;13:167. doi: 10.3390/membranes13020167. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Cox C.D., Poole K., Martinac B. Re-evaluating TRP channel mechanosensitivity. Trends Biochem Sci. 2024;49:693–702. doi: 10.1016/j.tibs.2024.05.004. [DOI] [PubMed] [Google Scholar]
- 11.Jin P., Jan L.Y., Jan Y.N. Mechanosensitive ion channels: structural features relevant to mechanotransduction mechanisms. Annu Rev Neurosci. 2020;43:207–229. doi: 10.1146/annurev-neuro-070918-050509. [DOI] [PubMed] [Google Scholar]
- 12.Liu S., Xu X., Fang Z., et al. Piezo1 impairs hepatocellular tumor growth via deregulation of the MAPK-mediated YAP signaling pathway. Cell Calcium. 2021;95 doi: 10.1016/j.ceca.2021.102367. [DOI] [PubMed] [Google Scholar]
- 13.Li Y.M., Xu C., Sun B., Zhong F.J., Cao M., Yang L.Y. Piezo1 promoted hepatocellular carcinoma progression and EMT through activating TGF-β signaling by recruiting Rab5c. Cancer Cell Int. 2022;22:162. doi: 10.1186/s12935-022-02574-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Tan S., Chao R. An exploration of osteosarcoma metastasis diagnostic markers based on tumor-associated neutrophils. Discov Med. 2023;35:300–311. doi: 10.24976/Discov.Med.202335176.31. [DOI] [PubMed] [Google Scholar]
- 15.Zhao X., Zhang L., Wang J., et al. Identification of key biomarkers and immune infiltration in systemic lupus erythematosus by integrated bioinformatics analysis. J Transl Med. 2021;19:35. doi: 10.1186/s12967-020-02698-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Kanehisa M., Goto S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 2000;28:27–30. doi: 10.1093/nar/28.1.27. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Wang J., Chen X., Wu D., et al. Single-cell and machine learning approaches uncover intrinsic immune-evasion genes in the prognosis of hepatocellular carcinoma. Liver Res. 2024;8:282–294. doi: 10.1016/j.livres.2024.11.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Kalbasi A., Ribas A. Tumour-intrinsic resistance to immune checkpoint blockade. Nat Rev Immunol. 2020;20:25–39. doi: 10.1038/s41577-019-0218-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Li H., Guo L., Cai Z. TCN1 is a potential prognostic biomarker and correlates with immune infiltrates in lung adenocarcinoma. World J Surg Oncol. 2022;20:83. doi: 10.1186/s12957-022-02556-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Danaher P., Warren S., Lu R., et al. Pan-cancer adaptive immune resistance as defined by the tumor inflammation signature (TIS): results from The Cancer Genome Atlas (TCGA) J Immunother Cancer. 2018;6:63. doi: 10.1186/s40425-018-0367-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Zhou X., Du J., Liu C., et al. A pan-cancer analysis of CD161, a potential new immune checkpoint. Front Immunol. 2021;12 doi: 10.3389/fimmu.2021.688215. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Clemente M.R.C., Felix N., Navalha D.D.P., et al. Long-term impact of home-based monitoring after an admission for acute decompensated heart failure: a systematic review and meta-analysis of randomised controlled trials. EClinicalMedicine. 2024;71 doi: 10.1016/j.eclinm.2024.102541. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Zheng W., Rawson S., Shen Z., et al. TMEM63 proteins function as monomeric high-threshold mechanosensitive ion channels. Neuron. 2023;111:3195–3210 (e7). doi: 10.1016/j.neuron.2023.07.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Affo S., Yu L.X., Schwabe R.F. The role of cancer-associated fibroblasts and fibrosis in liver cancer. Annu Rev Pathol. 2017;12:153–186. doi: 10.1146/annurev-pathol-052016-100322. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Hamill O.P., Martinac B. Molecular basis of mechanotransduction in living cells. Physiol Rev. 2001;81:685–740. doi: 10.1152/physrev.2001.81.2.685. [DOI] [PubMed] [Google Scholar]
- 26.Xiao B. Mechanisms of mechanotransduction and physiological roles of PIEZO channels. Nat Rev Mol Cell Biol. 2024;25:886–903. doi: 10.1038/s41580-024-00773-5. [DOI] [PubMed] [Google Scholar]
- 27.Zhang X., Zhao Y., Li M., et al. A synergistic regulation works in matrix stiffness-driven invadopodia formation in HCC. Cancer Lett. 2024;582 doi: 10.1016/j.canlet.2023.216597. [DOI] [PubMed] [Google Scholar]
- 28.Guan S.H., Hu W.J., Wang X.Y., Gu Y.X., Zhou D.H. New perspectives in prognostication of hepatocellular carcinoma: the role and clinical implications of transient receptor potential family genes. World J Gastrointest Oncol. 2024;16:2862–2864. doi: 10.4251/wjgo.v16.i6.2862. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Wei X., Li J., Xie H., et al. Chloride intracellular channel 1 participates in migration and invasion of hepatocellular carcinoma by targeting maspin. J Gastroenterol Hepatol. 2015;30:208–216. doi: 10.1111/jgh.12668. [DOI] [PubMed] [Google Scholar]
- 30.Liu T., Han X., Zheng S., et al. CALM1 promotes progression and dampens chemosensitivity to EGFR inhibitor in esophageal squamous cell carcinoma. Cancer Cell Int. 2021;21:121. doi: 10.1186/s12935-021-01801-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Rosenberg P. VDAC2 as a novel target for heart failure: Ca2+ at the sarcomere, mitochondria and SR. Cell Calcium. 2022;104 doi: 10.1016/j.ceca.2022.102586. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Zhang H., Zhang X., Xu W., Wang J. TMC5 is highly expressed in human cancers and corelates to prognosis and immune cell infiltration: a comprehensive bioinformatics analysis. Front Mol Biosci. 2022;8 doi: 10.3389/fmolb.2021.810864. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Zhang X., Shao J., Wang C., et al. TMC7 functions as a suppressor of Piezo2 in primary sensory neurons blunting peripheral mechanotransduction. Cell Rep. 2024;43 doi: 10.1016/j.celrep.2024.114014. [DOI] [PubMed] [Google Scholar]
- 34.Li M., Zhang X., Wang M., et al. Activation of Piezo1 contributes to matrix stiffness-induced angiogenesis in hepatocellular carcinoma. Cancer Commun (Lond) 2022;42:1162–1184. doi: 10.1002/cac2.12364. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Yuan Z., Li Y., Zhang S., et al. Extracellular matrix remodeling in tumor progression and immune escape: from mechanisms to treatments. Mol Cancer. 2023;22:48. doi: 10.1186/s12943-023-01744-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Liu Y., Yao X., Zhao Y., et al. Mechanotransduction in response to ECM stiffening impairs cGAS immune signaling in tumor cells. Cell Rep. 2023;42 doi: 10.1016/j.celrep.2023.113213. [DOI] [PubMed] [Google Scholar]
- 37.Nicolas-Boluda A., Vaquero J., Vimeux L., et al. Tumor stiffening reversion through collagen crosslinking inhibition improves T cell migration and anti-PD-1 treatment. Elife. 2021;10 doi: 10.7554/eLife.58688. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Zhang J., Li J., Hou Y., et al. Osr2 functions as a biomechanical checkpoint to aggravate CD8+ T cell exhaustion in tumor. Cell. 2024;187:3409–3426 (e24). doi: 10.1016/j.cell.2024.04.023. [DOI] [PubMed] [Google Scholar]
- 39.Yu K.X., Yuan W.J., Wang H.Z., Li Y.X. Extracellular matrix stiffness and tumor-associated macrophage polarization: new fields affecting immune exclusion. Cancer Immunol Immunother. 2024;73:115. doi: 10.1007/s00262-024-03675-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Batlle E., Massagué J. Transforming growth factor-β signaling in immunity and cancer. Immunity. 2019;50:924–940. doi: 10.1016/j.immuni.2019.03.024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Bordeleau F., Califano J.P., Negrón Abril Y.L., et al. Tissue stiffness regulates serine/arginine-rich protein-mediated splicing of the extra domain B-fibronectin isoform in tumors. Proc Natl Acad Sci U S A. 2015;112:8314–8319. doi: 10.1073/pnas.1505421112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Safina A., Vandette E., Bakin A.V. ALK5 promotes tumor angiogenesis by upregulating matrix metalloproteinase-9 in tumor cells. Oncogene. 2007;26:2407–2422. doi: 10.1038/sj.onc.1210046. [DOI] [PubMed] [Google Scholar]
- 43.Jiang Y., Zhang H., Wang J., Liu Y., Luo T., Hua H. Targeting extracellular matrix stiffness and mechanotransducers to improve cancer therapy. J Hematol Oncol. 2022;15:34. doi: 10.1186/s13045-022-01252-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Cambria E., Coughlin M.F., Floryan M.A., Offeddu G.S., Shelton S.E., Kamm R.D. Linking cell mechanical memory and cancer metastasis. Nat Rev Cancer. 2024;24:216–228. doi: 10.1038/s41568-023-00656-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Hu Y., Tian H., Chen W., et al. The critical role of the Piezo1/β-catenin/ATF4 axis on the stemness of Gli1+ BMSCs during simulated microgravity-induced bone loss. Adv Sci (Weinh) 2023;10 doi: 10.1002/advs.202303375. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Bo H., Wu Q., Zhu C., Zheng Y., Cheng G., Cui L. PIEZO1 acts as a cancer suppressor by regulating the ROS/Wnt/β-catenin axis. Thorac Cancer. 2024;15:1007–1016. doi: 10.1111/1759-7714.15278. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Lei Z.N., Tian Q., Teng Q.X., et al. Understanding and targeting resistance mechanisms in cancer. MedComm (2020) 2023;4 doi: 10.1002/mco2.265. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
All the data generated and analyzed in this study are available from the corresponding authors upon reasonable request. Some data can be obtained without restriction from online databases including TCGA, STRING, and GEO.