Abstract
A striking characteristic of liver cancer is its extensive heterogeneity, particularly with regard to its varied response to immunotherapy. In this study, we employed multimodal sequencing approaches to explore the various aspects of neoadjuvant nivolumab treatment in liver cancer patients. We used spatially-resolved transcriptomics, single- and bulk-cell transcriptomics, and TCR clonotype analyses to examine the spatiotemporal dynamics of the effects of nivolumab. We observed a significantly higher clonal expansion of T cells in the tumors of patients who responded to the treatment, while lipid accumulation was detected in those of non-responders, likely due to inherent differences in lipid metabolic processes. Furthermore, we found a preferential enrichment of T cells, which was associated with a better drug response. Our results also indicate a functional antagonism between tumor-associated macrophages (TAMs) and CD8 cells and their spatial separation. Notably, we identified a UBASH3B/NR1I2/CEACAM1/HAVCR2 signaling axis, highlighting the intense communication among TAMs, tumor cells, and T-cells that leads to pro-tumorigenic outcomes resulting in poorer nivolumab response. In summary, using integrative multimodal sequencing investigations, combined with the multi-faceted exploration of pre- and post-treatment samples of neoadjuvant nivolumab-treated HCC patients, we identified useful mechanistic determinants of therapeutic response. We also reconstructed the spatiotemporal model that recapitulates the physiological restoration of T cell cytotoxicity by anti-PD1 blockade. Our findings could provide important biomarkers and explain the mechanistic basis differentiating the responders and non-responders.
Supplementary Information
The online version contains supplementary material available at 10.1186/s12943-025-02314-w.
Keywords: Immunotherapy, Spatial transcriptomics, Liver cancer
Introduction
Liver cancer presents a significant global health challenge, as it ranks as the sixth most common cancer and the third leading cause of cancer-related deaths worldwide [1]. Hepatocellular carcinoma (HCC) accounts for 90% of liver cancer cases and is known for its aggressive nature. Often, HCC is detected at an advanced stage, resulting in a high tumor recurrence rate (up to 70% within five years after resection) and a low survival rate (only 18% at five-year survival) [2, 3]. For advanced HCCs, multikinase inhibitors (MKIs) have been used to improve survival rates in patients. However, limitations such as inadequate tumor reduction and high toxicity have prompted the development of new treatment options, including molecular targeted agents (MTAs), immune checkpoint inhibitors (ICIs), and combined therapies. These emerging treatments aim to expand and enhance the available options for managing advanced HCC.
Immune checkpoint inhibitors (ICIs) have emerged as a crucial component of cancer therapy due to their ability to substantially improve outcomes and long-term survival for cancer patients. ICIs are now being utilized in various cancer types, including HCC, with expanded applications in adjuvant or neoadjuvant settings [4]. In particular, advancements in this research area are highlighted by antibodies targeting the programmed cell death- 1 receptor (PD- 1) or its ligand (PD-L1). Nivolumab, a monoclonal antibody to PD- 1, is being developed for the treatment of multiple cancer types. Clinical trials have demonstrated its promising activity in HCC at diverse stages [5, 6]. Despite substantial evidence supporting ICI treatment benefits, reports on its use as a neoadjuvant therapy are limited [7–9], and the landscape of its tumor microenvironment (TME) remains largely unexplored.
Traditional single-cell RNA sequencing techniques can reveal the heterogeneity within cell populations and uncover alterations that distinguish individual cell types. However, the status of cancer cells heavily depends on their precise spatial positioning and interplay with adjacent cells. Since the necessary dissociation process for single-cell RNA sequencing (scRNA-seq) results in a loss of location data, it fails to inform the functional associations among distinct tissue regions. The advent of spatial transcriptomics sequencing enables the simultaneous assessment of gene expression levels and spatial distribution within tissue samples at spot or cellular resolution. In this study, we obtained tissue specimens from 16 HCC patients who underwent neoadjuvant nivolumab therapy and subsequent tumor resection. We employed a range of techniques, including spatial transcriptomics, single-cell and bulk-cell transcriptomics, and TCR sequencing. Our study aimed to investigate the intrinsic differences between patients displaying varying therapeutic responsiveness, identify relevant biomarkers, and reconstruct the spatiotemporal dynamics of nivolumab response.
Results
Neoadjuvant nivolumab-treated HCC patient cohort and multimodal sequencing
We randomly selected a subset of sixteen HCC patients with intermediate and locally advanced tumors who received preoperative nivolumab at a dosage of 3 mg/kg for 3 cycles prior to surgical resection [10]. Their resected tumor tissues were collected for sequencing analyses. All samples were subjected to bulk RNA sequencing (bulk RNA-seq) and spatial transcriptomic sequencing (ST-seq), while 4 of them underwent additional single-cell RNA sequencing (scRNA-seq) and T-cell receptor sequencing (TCR-seq) (Fig. 1A). The demographic and clinical characteristics of the patients are described in Table S1. Following our established definition of major pathologic tumor necrosis (≥ 60%) for classifying the nivolumab response [10], our study consisted of six responders (R) and ten non-responders (NR).
Fig. 1.
ScRNA-seq profiling of HCC upon nivolumab treatment. A A schematic diagram of the study and experimental strategy. B t-SNE of scRNA-seq data from all samples (n = 4) colored by cell type, patient, and treatment response. R, responder; NR, non-responders. C Bubble chart of gene ontology enrichment for the cell type markers as indicated. D Odds ratios of cell type proportion in R and NR groups. E t-SNE visualization of T cell subsets. F t-SNE visualization of clonotype expansion levels among T cell subsets. G Bar plot of clonal expansion levels across different samples, with expansion rate represented in a line plot. H Volcano plot showing the DEGs between expansion and non-expansion T cells
ScRNA-seq and TCR-seq profiling of HCC ecosystem revealed unique treatment-related cellular stratification and clonal expansion
For the scRNA-seq, after the removal of low-quality cells, potential doublets, and batch effects, we obtained more than 133 million unique transcripts from 16,848 cells, capturing 23,143 genes. Using unsupervised clustering and the t-distributed stochastic neighbor embedding (tSNE) algorithm, we created a single-cell atlas categorized by cell types, samples, and patient groups by responsiveness to nivolumab (Responder [R] group: R1, R6; Non-responder [NR] group: NR3, NR6) (Fig. 1B). Based on the canonical marker expression, we identified eight major cell types, including tumor cells, CD8 cells, CD4 cells, tumor-associated macrophages (TAMs), B cells, plasma B cells, endothelial cells and cancer-associated fibroblasts (CAFs). The distribution and quantity of each cell type varied across the different samples (Figure S1 A-D). Gene Ontology (GO) enrichment analysis further supported the precision of the classification (Fig. 1C). Essentially, pathways related to fatty acid metabolism were exclusively enriched in tumor cells, while T-cell activation and differentiation pathways were observed in CD8 and CD4 cells. Similarly, B cell activation pathways were found in both B cells and plasma B cells. When comparing the R and NR groups, we found increased quantities of CD4, CD8 and plasma B cells in the R group, indicating significant immune activation following nivolumab treatment (Fig. 1D).
To further validate the immune activation observed in the R group, we employed TCR-seq to analyze the T cell clonotypes. We categorized the T cells into various subtypes, with CD8 cells divided into 4 distinct subgroups, while only one single group of CD4 cells was detected (Fig. 1E). A corresponding TCR was detected in 79% of the T cells analyzed, representing a considerably high detection rate (Figure S1E). We observed significant clonal expansion of CD8 cells but rarely CD4 cells, and the most pronounced clonal expansion occurred in the c1 - 3 subtype of CD8 cells (Fig. 1F). We noted that patients with more favorable treatment response exhibited an increased level of clonal expansion in their CD8 cells (Fig. 1G). Specifically, the tumor of the patient (R6), who had the best therapeutic outcome with pathologic tumor necrosis of 96%, displayed the highest level of clonal expansion (Fig. 1G). When comparing expanded CD8 cells with their non-expanded counterparts (i.e. those with a unique clonotype), genes involved in cytotoxicity and clonal expansion function (such as NKG7, CREM, PRF1, BATF, and SOCS3) were upregulated in the expanded T cells. In contrast, genes associated with T cell exhaustion and suppression (like FOXO3, CCL3, TNFRSF18) tended to be expressed in the non-expanded T cells (Fig. 1H).
In-depth spatial exploration of intricate HCC architecture with ST-seq
We examined the ST-seq data to comprehensively analyze the spatial architecture of the HCC tumor microenvironment following neoadjuvant nivolumab therapy. Since a single spot in ST-seq using 10X Genomics Visium platform may encompass multiple cells, deconvolution was essential before further analysis. We deconvoluted the determined cell type abundance of spots in the ST-seq data by BayesPrism [11] with the aforementioned scRNA-seq data as a reference. We were able to determine the spatial abundance of the eight major cell types, with the results consistently aligning with those from the histology with hematoxylin and eosin (H&E) staining (Fig. 2A). Moreover, we also verified the results using representative markers corresponding to the cell types (Figure S2).
Fig. 2.
Spatial multi-omics profiling of HCC upon nivolumab treatment. A Characterization of ST-seq data by cell-type deconvolution, H&E staining, and unsupervised clustering. B t-SNE visualization of ST-seq spots colored by regions. C Dot plot of average expression for selected marker genes in each region. D Heatmap of Hallmark pathway (from MSigDB) activities scored per region by ssGSEA. E Spatially mapped feature-level ssGSEA scores for selected Hallmark gene sets. Color scale represents enrichment scores
We discovered that using conventional methods to combine all samples and then performing clustering resulted in inconsistencies with the histological analysis, which were primarily due to resolution and heterogeneity concerns. To address this, we first conducted clustering on individual samples before merging them. We then utilized a hierarchical clustering approach that initially divided all clusters into two categories: tumor region and non-tumor region (Figure S3 A-B). Subsequently, we applied a similar method to further subdivide these two categories into two Tumor Regions (TR1 and TR2), a Fibrotic Region (FR), an Immune Region (IR), and a Mixed Region (MR) (Figure S3 C-D), with different regions enriched with the respective cell types. Therefore, IR was enriched with immune cells, FR with CAFs, TR with mainly tumor cells and minimal non-immune cells, while MR had equal admixtures of different cell types (Figure S4 A). The MR was situated at the interface between the tumor and non-tumor compartments, comprising a more heterogeneous mixture of tumor cells, immune cells, and CAFs. Similar to the scRNA-seq analysis, we utilized the t-SNE algorithm for ST-seq data visualization and selected matched samples that corresponded to the same set of scRNA-seq cases for illustration (Fig. 2B). The results of differentially expressed genes (DEGs) revealed the distinct gene signature for each region, with APOH and LCN2 exhibiting elevated expression in the TR1 and TR2, respectively; PTPRC was enriched in the IR, and collagen-associated genes were enhanced in the FR (Fig. 2C and Figure S4B). These findings supported the accuracy of the clustering.
Next, we analyzed the activities of hallmark pathways by single-sample gene set enrichment analysis (ssGSEA) [12] for each spot. In general, hierarchical clustering showed that MR exhibited a substantial degree of similarity with those of TR1 and TR2, further supporting their similar intrinsic properties and closely connected spatial localization (Fig. 2D). The IR and FR formed a separate cluster, indicating a degree of connectivity between these two regions. Furthermore, the heatmap revealed that, apart from fatty acid and cholesterol metabolism as well as other metabolic processes, both two TRs displayed considerable enrichment in the Myc target v1 and oxidative phosphorylation pathways. Consistently, the IR demonstrated a marked activation in the interferon-α/γ response pathways. Additionally, the genes in the FR were intricately involved in the angiogenesis pathway (Fig. 2D). To enhance the visualization of pathway enrichment results, representative samples illustrating the spatial mapping of the corresponding specific molecular pathways are shown (Fig. 2E). Together, these ST-seq results facilitated the reconstruction of the action paths of neoadjuvant nivolumab treatment and visualization of the spatial heterogeneity in the HCC microenvironment concerning different regions.
Spatial heterogeneity of tumor regions revealed redistribution dynamics pinpointing nivolumab response in HCC
To explore the spatial heterogeneity of the individual tumor regions, we carefully examined the TR1 and TR2 and, upon a re-clustering analysis, revealed notable inter-patient tumor heterogeneity. For each patient, the tumor cells formed separate clusters, encompassing both TR1 and TR2 (Fig. 3A), but their proportions varied (Fig. 3B, S5 A). Furthermore, the GO enrichment results revealed distinctions between the two regions (Fig. 3C), with genes predominantly enriched in pathways associated with steroid metabolic process, xenobiotic metabolic process and other metabolic processes in TR1, whereas the up-regulated pathways in TR2 appeared more diverse (Fig. 3C). In addition, the genes upregulated in the R group, as compared to NR group, were closely associated with a higher humoral immune response and acute inflammatory response (Fig. 3D, left panel; Figure S5B). In contrast, the upregulated genes in the NR group were primarily involved in the cholesterol biosynthetic process and carboxylic acid biosynthetic process (Fig. 3D, right panel; Figure S5B), suggesting a potential pro-tumor maintenance effect by these lipid-related molecules.
Fig. 3.
Characteristics of tumor cells. A t-SNE visualization of two tumor regions TR1 and TR2 colored by distribution and patient tumors. R, responder; NR, non-responders. B Stacked barplot of the proportion of regions in each of the tumors. C Bubble chart of gene ontology enrichment for the tumor region markers. D Violin plots of the humoral immune, inflammatory, cholesterol biosynthetic process and carboxylic acid biosynthetic process scores of tumor cells between R and NR groups. E Bar plot showing the top 10 enrichment scores of significantly enriched pathways for up-regulated genes in pre-treatment bulk-seq data. F Heatmap showing the genes associated with humoral immune response pathway in the pre-treatment bulk-seq (liver biopsies) and ST-seq data. G Violin plot depicting the proliferation and hepatic level for tumor region. H Violin plot depicting the stemness level for each patient (upper panel), accompanied by a heatmap displaying the expression levels of liver cancer stem cell markers (lower panel). I. Correlation analysis between necrosis rate and stemness score in HCC
To explore whether patients in the R and NR groups could be differentiated prior to treatment, we examined the Bulk-seq data of the pre-treatment liver biopsies (Fig. 3E) (part of the data were presented in our previous study [10]) and identified significant phagocytosis pathways within the tumors in the R group. Furthermore, activation of the complement system and activation of pathways related to tumor-infiltrating B cells (TIBs) were observed. All of the above pertained to activation of both innate and adaptive immunities. Additionally, genes enriched in the humoral immune pathway were highly expressed in the R group (Fig. 3F, left panel). Heatmap analysis of the data from the ST-seq, although pertaining to post-treatment samples, demonstrated that most of the selected genes associated with humoral immune response pathway exhibited higher expression levels in the TR of the R group than NR group (Fig. 3F, right panel). The findings suggest that macrophages and other natural immune precursors were closely associated with tumor cells within the tumor microenvironment, both prior to and following the treatment.
Furthermore, we also assessed the tumor cell proliferation and hepatic features in the ST-seq analysis [13] (Fig. 3G). We found that in both TRs, the tumor proliferation levels in the NR group were higher than those in the R group, indicating that the tumors in the NR group exhibited higher proliferation potential. We also observed that the hepatic score was significantly higher in the R group than NR group in TR1, while it was lower in the R group in TR2. The findings suggest that there could be differential remnant hepatic functions in the two regions.
Furthermore, we examined a panel of representative liver cancer stem cell (LCSC) marker genes [14] and observed no particular difference in their expression between the NR and R groups (Fig. 3H). There was also no significant correlation between the expression of LCSC marker genes and the post-treatment pathologic tumor necrosis (Fig. 3I). These findings suggest that the stemness of HCC tumors may not be a major determinant of the effectiveness of nivolumab. However, we observed a positive correlation between the expression of the CD47 gene and degree of tumor necrosis in these tumors, suggesting an inherently higher immuno-evasive capability of CD47+ tumor cells (Figure S5 C). The action of Nivolumab may enable the immune system to more effectively attack these endogenously difficult-to-target tumor cells.
Spatial heterogeneity of non-tumor cells in the HCC tumor microenvironment
Unlike the tumors, many of the non-tumor regions, displayed a trend for homogeneity with no substantial stratification of spots (Fig. 4A). Most tumors consisted of FR, IR, and MR in varied proportions (Fig. 4B). The GO enrichment results revealed that the MR had enrichment of pathways such as steroid metabolic process and carboxylic acid biosynthetic process (Fig. 4C), possibly due to the presence of a small proportion of tumor cells. In contrast, the IR had clear enrichment of pathways related to immune cells, particularly T cells. The pathways enriched in the FR were related to wound healing and extracellular matrix (ECM), likely due to the presence of CAFs (Fig. 4C).
Fig. 4.
Characteristics of the immune cells. A t-SNE visualization of three stromal regions (MR, IR, and FR), colored by distribution and patient tumors. B Stacked bar plot of the proportion of different stromal regions of each tumor. C Bubble chart of gene ontology enrichment for the stromal region markers. D Network plots of gene ontology enrichment for the up-regulated genes in the IR of the responder group. E Heatmap showing the expression levels of immune-related genes in post-treatment bulk-seq data (left panel) and ST stromal regions data (right panel), respectively. F Correlation analysis between MinDistance and various features in HCC (upper panel), accompanied by violin plots illustrating the M1/M2/CD8_native/CD8_Exhausted/CD8_Cytotoxic levels in the TRs (lower panel)
Characteristically, the IR had a high fraction of T cells. In the GO enrichment results, the highly expressed genes in the R group (as compared to the NR group) showed elevated lymphocyte activation and adaptive immunity, exemplifying the anticipated effect of nivolumab on T cell activation (Fig. 4D). Subsequently, the post-treatment bulk-seq data showed that a series of marker genes of immune cells were highly expressed in the R group, while ST-seq data also indicated similar enrichment in the R group, particularly those with the most favorable treatment outcomes (Fig. 4E).
Making use of the fact that ST-seq allows derivation of the spatial proximity between distinct spot types and elucidation of the relationships between spatial separation distance and the associated molecular features, we compared the R and NR groups with respect to the proximity of various cell types to tumor cells, as well as their corresponding enrichment scores. In particular, we found that in the R group, the M1-like TAMs were closer to the tumor cells, while the M2-like TAMs were more distant. Moreover, the M1-like TAMs were more abundant in the R group than NR group, while the M2-like TAM had the opposite trend (Fig. 4F).
Regarding the three distinct functional states of CD8 cells (naïve, exhausted, and cytotoxic), their levels were all elevated to varying degrees in the R group compared to the NR group (Fig. 4F). This suggests the R group exhibits a more robust immune response.
Precise deconvolution of cell type identity suggested preferential enrichment of cell types in relation to nivolumab response
It was crucial to further substantiate the cell type identity of each spot. Based on BayesPrism [11] that performed cell type deconvolution primarily based on abundance, we simultaneously employed MISTY [15], a machine-learning-based, annotation-free inference of spatial relationships of cell types. It modeled marker relationships at different views, and its results could complement BayesPrism to better define the identity and heterogeneity of cell types for each spot. Collectively, spots with concordant classification by both algorithms (Figure S6 A-B) were defined as cell type-enriched spots (8 major classes: tumor, CAFs, endothelial cells, TAMs, CD4, CD8, plasma B cells and B cells), whereas the discordant ones were named as mixed spots (Fig. 5A). We were able to sub-divide CAFs into 3 different subtypes (inflammatory CAF [iCAF], myofibroblastic CAF [mCAF], and antigen-presenting CAF [apCAF]), according to their corresponding signature scores [16] (Fig. 5B). Mixed spots and tumor-enriched spots were most commonly detected (Fig. 5C).
Fig. 5.
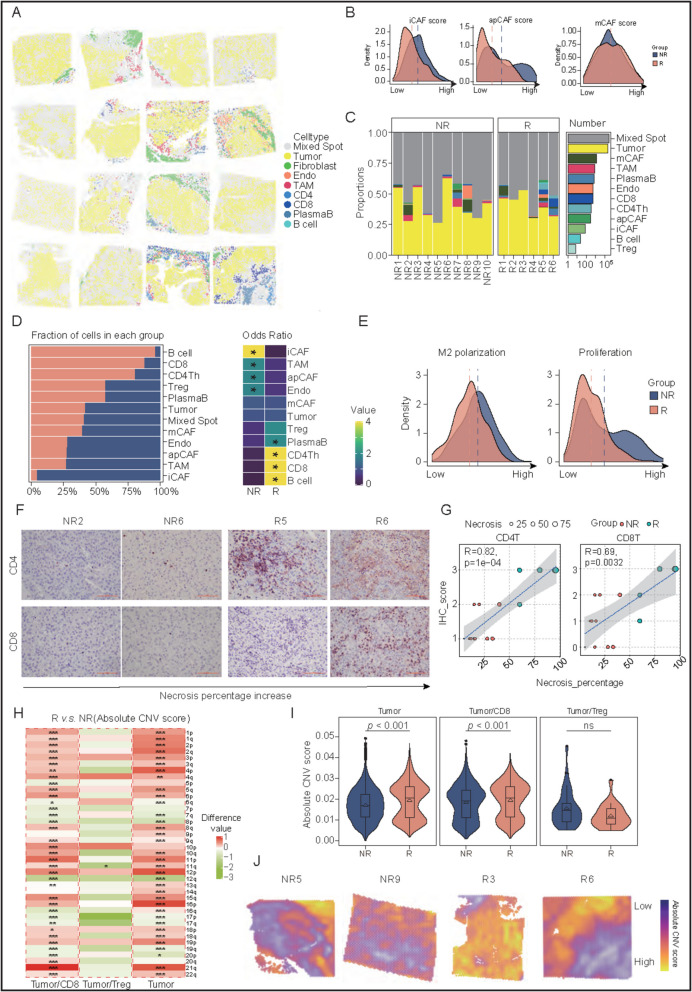
Spatial transcriptome reveals the immune microenvironment and CNV landscape after neoadjuvant nivolumab therapy. A Spatial visualization with definitions of the spots in the 16 samples. B Density plot of subtype score of CAF between R and NR groups. C Bar chart showing the proportion of the various cell types (left) and the total number of spots (right) in all samples. D The proportion of the various cell types in the R and NR groups as according to the degree of pathologic tumor necrosis (left panel) and heatmap showing the Odds Ratio (OR) of every spot between the R and NR groups (right panel). E Density plot of M2 polarization and proliferation of TAM between R and NR groups. F Immunohistochemical stains for CD4 and CD8 T cells in representative cases of the R and NR groups. G Correlation between immunohistochemical positivity rate (score) of CD4 and CD8 T cells and the degree of pathologic tumor necrosis. H Heatmap showing the difference in the CNV absolute scores between the R and NR groups. Each row represents a chromosomal arm, and each column represents a cell-type spot. I Boxplot showing the statistical comparison of CNV absolute score in the Tumor/T cell spot between R and NR groups. J Spatial visualization of CNV absolute scores in representative cases of the R and NR groups
Intriguingly, although spots of individual cell types varied in their proportions among patients, specific cell types were preferentially enriched in R (B cells, CD8 and CD4) and NR (iCAF, TAMs and apCAF) patients (Fig. 5D). Cytotoxic CD8 tended to be enriched in the R group, whereas exhausted CD8 tended to be enriched in the NR group (Figure S7 A). For CD4 cells, we assessed their three subtypes based on their respective scores, with Th1 cells determined to be significantly concentrated in the R group (Figure S7B), which plays a vital role in regulating the anti-tumor activity of tumor-infiltrating immune cells [9]. Regarding the TAM-enriched spots, those identified in NR patients tended to be more M2-like and proliferative (Fig. 5E). The results showed that the drug responsiveness might likely be associated with the abundance and intrinsic properties of individual cell types. We further made use of immunohistochemistry to validate the higher abundance of CD8 and CD4 cells in the R patients, as compared to the NR counterparts (Fig. 5F). The infiltration of CD4 and CD8 significantly and positively correlated with the pathologic tumor necrosis percentage, indicating an immune-stimulatory microenvironment in the tumors of the R patients (Fig. 5G).
Lipid metabolic process could represent intrinsic differences indicating nivolumab response
We investigated if copy number variation (CNV) could represent a major intrinsic difference predisposing to drug responsiveness. To this end, upon extracting the tumor-related spots to perform genome-wide CNV analysis (Figure S7 C), we identified frequent and substantial CNV events among the tumors in the R group in general, suggesting possible intrinsic differences rendering drug responsiveness (Fig. 5H). To further examine this, since the action of nivolumab likely primarily exerted on CD8 and Treg cells, we isolated the tumor/CD8 and tumor/Treg spots, which represented the primary sites of anti-tumor immunity and pro-tumor immunosuppression that was restored and inhibited by nivolumab, respectively. In line with our expectation, tumor/CD8 spots were detected with a significantly higher degree of CNV in R group, supporting our belief that CNV alterations might serve as signals to trigger CD8-mediated anti-tumor immunity. In contrast, Treg cells demonstrated no preferential enrichment between R and NR groups (Fig. 5I), suggesting that they might not be the major site responsible for drug response. The disparity in the intrinsic extent of CNV could help to stratify HCC patients according to drug response (Fig. 5J).
We then analyzed the genes that displayed concordance in the differential gene expression and CNV status between R and NR groups (Figure S8 A-B). Importantly, the majority (> 50%) of the different gene expression events were supported by underlying CNV alterations (Figure S8 C). CNV-supported DEGs were subjected to functional and pathway enrichment analyses. While NR tumors were accompanied by upregulation of myc target signaling, xenobiotic, bile acid metabolism, and cholesterol homeostasis, R tumors were enriched with upregulation of gene sets related to complement, coagulation, and interferon γ response (Figure S8D). When we examined the protein–protein networks related to these upregulated gene sets in R tumors, PPARG was found to be common in all (Figure S8E). More importantly, the expression of PPARG was positively correlated with the presence of its CNV amplification (Figure S8 F). This suggests that PPARG might serve as a key modulating factor affecting drug response. Moreover, in our in-house cohort, PPARG was overexpressed in HCCs as compared with the corresponding non-tumorous livers, with 55% of HCCs showing more than twofold upregulation (Figure S8G) and upregulation of PPARG expression was significantly associated with poorer disease-free and overall survival (Figure S8H).
In addition, HCC tumors with higher PPARG expression were associated with more aggressive tumor behavior, in terms of more frequent venous invasion and tumor microsatellites, as well as larger tumor size and more advanced tumor stages (Figure S8I). This association of poorer overall survival was confirmed in TCGA LIHC cohort (Figure S8 J). In summary, high expression of PPARG could be a potentially useful biomarker for anti-PD1 drug response and prognostication of HCC patients. Susceptibility to nivolumab treatment in PPARG-high patients may possibly be explained by the PPARG-mediated stimulation of interferon γ response, resulting in the activation of PD-L1 expression and related addiction to PD- 1-related immunosuppression [17].
Based on the results of enrichment analysis of the TRs (Fig. 3D and Figure S4B), we further focused particularly on exploring the lipid metabolism in R and NR patients. Here, we identified a significant enrichment of CNV-supported differential gene expression events in lipogenesis in the NR tumors, while lipolysis was enriched in the R tumors (Fig. 6A). The results of up-regulation of lipid biosynthesis in the NR tumors and down-regulation in the R tumors were consistently observed using Oil Red O stain (Fig. 6B). Meanwhile, qPCR analysis also consistently supported the upregulated expression of the different representative genes involved in lipid metabolism in the NR tumors (Fig. 6C). In addition, the degree of pathologic tumor necrosis showed a negative correlation with the expression of lipid-associated genes (Fig. 6D). These data suggest that high lipid content might render the patients less responsive to nivolumab. Although our observation was obtained in HBV-associated HCC patients, our findings were somehow in line with the previous report [18] that metabolic-related HCCs (steatosis) were less susceptible to immunotherapy.
Fig. 6.
Lipid accumulation, spatial distance between TAM and CD8, and drug response. A Dot plot showing pathway enrichment of lipogenesis and lipolysis in the NR group. B Oil red O staining in representative cases of the R (lower panel; 60% and 96% of tumor necrosis) and NR (upper panel; 15% and 30% of tumor necrosis) groups. C Scatter plot of the expression of genes related to lipid synthesis using qPCR analysis. D Correlation diagram between the expression of genes related to lipid synthesis and percentage of tumor necrosis. E Spatial visualization of CD8Rel spots and TAMRel spots in representative cases of the R and NR groups. F Box plot showing the statistical comparison of the spatial distance between CD8Rel and TAMRel spots between the R and NR groups. G Alluvial diagram showing the composition of enriched TAM, enriched CD8, and TAM/CD8 spots in the R and NR groups. H Conceptual diagram showing the derivation of heterotypic and homotypic scores. I Box plot showing the statistical comparison of heterotypic score (TAMRel to CD8Rel, with TAMRel serving as the center point) between the R and NR groups. J Spatial visualization of TAMRel and CD8Rel as spatial neighborhoods in representative cases of the R and NR groups. K Box plot showing the statistical comparison of the functional scores of CD8Rel surrounding TAMRel between the R and NR groups. L. Box plot showing the statistical comparison of homotypic scores of TAMRel and CD8Rel between the R and NR groups. M Spatial visualization of the homotypic scores of TAMRel and CD8Rel between R and NR groups. N Ripley's L diagram showing TAMRel and CD8Rel distribution patterns (dispersed or clustered) between R and NR groups
Functional opposition between TAM and CD8 and their spatial exclusion
From the spatial architecture in the HCC tumors, we identified an intriguing and potentially implicative exclusive feature between pro-tumorigenic TAM and anti-tumorigenic CD8 cells, in that they displayed significantly different spatial localization (Fig. 6E). There was a significantly greater distance between the TAM- and CD8-related spots in the R tumors, indicating more intense functional opposition (Fig. 6F). This implies that the restoration of cytotoxicity in CD8 cells upon nivolumab treatment would impose more profound functional opposition between TAM and CD8 cells. This was also supported by the respective enrichment of CD8-enriched spots and TAM-enriched spots in the R and NR tumors, together with a smaller proportion of TAM/CD8 spots (mixing of TAM and CD8 in the same spot) in the R tumors (Fig. 6G).
We applied the concept of homotypic [19] and heterotypic [20] arrangements of cell types to further investigate the divergence of spatial organization of TAM and CD8 cells in relation to drug response (Fig. 6H, S9 A). Upon calculating the heterotypic scoreTAM to CD8, which reflected the extent of TAMs being surrounded by CD8 cells, we found the score was significantly higher in the R tumors, indicating CD8 cells tended to surround TAMs. Such pattern was not observed in the NR tumors. CD8 cells, with restored cytotoxicity by nivolumab, seemingly more actively surrounded the TAMs to counteract its pro-tumorigenic effect regionally (Fig. 6I-J). Subsequently, we extracted those CD8 spots in proximity surrounding the TAM spots. As expected, CD8 cells were more cytotoxic in the R tumors, suggesting a significantly better restoration of CD8 cytotoxicity by nivolumab (Fig. 6K, S9B). Such T cell cytotoxicity was possibly mediated by conversion from memory T cells and originally exhausted T cells (Fig. 6K), which were previously implicated [21].
In line with the results of the heterotypic score, the homotypic score consistently indicated higher accumulation of CD8 cells in the R group and TAMs in the NR group (Fig. 6L). This was supported by the spatial localization of spots in the tissue (Fig. 6M) and Ripley’s L metric indication (higher value indicates a high degree of aggregation) [22]. CD8-related spots in the R tumors exhibited a higher Ripley’s L value than those in NR ones, whereas the TAM-related spots demonstrated the reverse trend (Fig. 6N). Collectively, the findings indicate that divergent accumulation of CD8 cells and TAMs were associated with the drug response.
Signaling axis pinpointing an intense communication among TAMs, tumor cells and T cells to elicit pro-tumorigenic consequences in relation to nivolumab response
We investigated if alternative activation of immune checkpoint inhibition could be induced, alleviating the action of nivolumab. To this end, we performed cell–cell communication exploration pinpointing antigen presenting cells and T cells, focusing the analysis on different immune checkpoint ligand-receptor (L-R) pairs (Fig. 7A). By reconstructing the spatiotemporal trajectory of L-R interactions, CD8 and CD4 Th were the major cell types participating in the drug response. Significant interactions via PVR-CD226 were detected between CD8 and tumor cells, implicating that this co-stimulatory checkpoint interaction might be crucial in leading to better drug response. Importantly, we identified a prominent CEACAM1-HAVCR2 interaction between the tumor cells and CD8 cells, which likely shed light on the potential role of this co-inhibitory immune checkpoint axis in modulating the action of nivolumab. Moreover, we also observed the CD48-CD244 interaction between apCAF and CD8 cells and the interaction between apCAF and CD4 Th cells via multiple immune checkpoint axes (Fig. 7A).
Fig. 7.
Intense communication among TAM, tumor cells and T cells to elicit pro-tumorigenic consequences in relation to nivolumab response. A Dot plot showing the interactions of immune checkpoint ligand-receptor pairs (left panel). Each row represents a cell–cell interaction pair, and each column represents a ligand-receptor pair. Chord diagram showing significant immune checkpoint ligand-receptor pairs between R and NR groups (right panel). B Chord diagram showing significant ligand-receptor pairs via CellphoneDB, with enriched tumor spots as the receiver between R and NR groups. C Dot plot showing significant ligand-receptor pair between the R and NR groups, with enriched tumor spots as the receiver. Each row represents a ligand-receptor pair and each column represents a sender-receiver pair. D A schematic representation of the molecular and cellular axis in the R group. E Spatial visualization of cell type distribution (column in far left), monocular expression, and communication signal strength, with the multiple small black arrows showing the directions of the communication signal trend
Given the potential involvement of CEACAM1-HAVCR2 axis, we confined the CEACAM1 to be the target gene (expressed in tumor cells) in our search and identified UBASH3B, TIMP1, RNASET2 and MIF as upstream ligand candidates through NicheNet analysis [23] (Figure S9 C-D). The tumor cells frequently interacted with CD4 Th, TAMs and iCAF (Figure S9E). We further identified the comprehensive interactome profile between multiple cells (contributing ligands) with tumor cells (contributing receptors) (Figure S10 A-B). Encouragingly, those upstream ligand candidates suggested by NicheNet analysis were all identified and confirmed by the interactome profiling.
Next, based on these upstream ligand candidates, we found the differentially expressing cell types and the corresponding receptor molecules that were differentially expressed on the tumor cells (Fig. 7B). UBASH3B-NR1I2, TIMP1-FGFR2, RNASET2-TLR8 and MIF-EGFR were determined to be the major L-R axes potentially affecting drug response outcome (Fig. 7C), and among them, the UBASH3B-NR1I2 axis demonstrated the most contrasting difference between the R and NR tumors. The ligand and receptor were expressed in TAMs (UBASH3B) and tumor cells (NR1I2), respectively (Fig. 7D). Taken together, our findings suggest that the UBASH3B-NR1I2-CEACAM1-HAVCR2 axis involving TAMs, tumor cells, and CD8 cells likely have important differential inter-cellular communication between the R and NR groups, leading to drug response. Consequently, the gene expression signals and directional inference also supported our proposed model of cell–cell communication (Fig. 7E). With multicolor immunofluorescence, we found that the respective expressions of the signal axis in TAMs (CD68 & UBASH3B [encoding STS1 protein]), tumor cells (CEACAM1), and T cells (CD8 A & HAVCR2 [encoding TIM3 protein]) increased in the R group, which confirmed our model of multiple cell type endeavor (Fig. 8A-B). Comparing the R and NR tumors, we observed a more prominent expression of the UBASH3B- CEACAM1-HAVCR2 axis. In addition, the expression of HAVCR2 and communication signal strength were significantly and positively correlated with the degree of pathologic tumor necrosis (Figure S10 C), suggesting that HAVCR2 might serve as a useful biomarker for drug response. Targeting both PD- 1 and HAVCR2 could potentially deliver more complete treatment efficacy.
Fig. 8.
In vitro and in vivo experiments demonstrating the UBASH3B-NR1I2-CEACAM1-HAVCR2 cell–cell communication axis. A Multiplex immunofluorescence staining of representative cases of the R and NR groups. B Histogram comparing the proportion of positive cells for the respective genes between the R and NR groups. C Bar plot of CEACAM1 expression between the Huh7 control cells and Huh7 cells treated with recombinant UBASH3B protein at 20ug/ml for 24 h. D Bar plot of CEACAM1 expression in Huh7 cells treated with conditioned media collected from the THP- 1 of non-target control (NTC) and shUBASH3B cells, respectively. E Flowchart of orthotopic mouse HCC model using luciferase-labelled, shCEACAM1 Hepa1 - 6 HCC cell line. F In vivo bioluminescence images of the orthotopic injection mouse model using luciferase-labelled Hepa1 - 6 NTC and shCEACAM1 cells. G Flow cytometry and scatter plot showing changes in the percentages of CD8+ and TIM3+ T cells in the Hepa1 - 6-derived HCCs in the NTC and shCEACAM1 groups. H Relative expression of the genes of study in the HCCs of the orthotopic injection model between the NTC and shCEACAM1 groups
To validate the communication axis, we first conducted multiple in vitro experiments, including a knockdown approach, a co-culture experiment, and the use of specific recombinant protein. First, we found that treatment with recombinant UBASH3B protein at 20 µg/ml on Huh7 cells increased CEACAM expression (Fig. 8C). Considering the suggestion that TAMs are the source of UBASH3B protein, we employed THP- 1, a cell line frequently used to represent TAMs [24], to determine if the conditioned medium from UBASH3B-knockdown THP- 1 cells would impact CEACAM1 expression in the target HCC cells. Intriguingly, we found that the conditioned medium from shUBASH3B THP- 1 cells (Figure S10D) significantly suppressed the CEACAM1 expression of Huh7 cells (Fig. 8D). With the suggested importance of CEACAM1-mediated HAVCR2 expression in our nivolumab-treated HCC patients, we further investigated whether inhibiting CEACAM1 would inhibit the immunosuppression elicited by the HAVCR2 checkpoint. We employed the immunocompetent orthotopic mouse HCC model to study this (Fig. 8E) and found that stable CEACAM1 knockdown (shCEACAM1) in the Hepa1 - 6 mouse HCC cells (Figure S10E) significantly impeded tumor initiation (Figs. 8F and S10 F). We also observed a trend of reduction in TIM3+ (TIM3 is encoded by HAVCR2 gene) CD8 T cells in the shCEACAM1 tumors (Fig. 8G). Notably, the CD8 T cells also demonstrated higher T cell activation markers (GRAMB and CD69) in shCEACAM1 tumors as compared to the non-target control (NTC) (Fig. 8H). This implies that knockdown of CEACAM1 leads to an increase in cytotoxic T cell activation.
Discussion
In this study, based on a wealth of multimodal sequencing data pinpointing different unique aspects of neoadjuvant nivolumab treatment in HCC patients, we carried out spatial investigation of the cellular and molecular determinants for drug response, complemented by single-cell and bulk-cell transcriptomics and TCR clonotype analyses. With the combination of spatial expression profiling and solid pathological understanding of cytotoxicity action of nivolumab, we were able to reconstruct the spatiotemporal process of nivolumab. Importantly, we were able to delineate the essential elements and biomarkers that stratify HCC patients according to their anticipated drug response. On the other hand, given the limited sample size of our study, further confirmation with larger sample cohorts is awaited in future studies. Besides, some statistical analysis e.g. Ripley’s L metric calculation for the spatial distribution was only calculated using default parameters without comprehensive optimization procedures. We acknowledge these shortcomings and therefore complemented our findings with the corresponding experimental confirmation e.g. multicolor immunofluorescence to strengthen our spatial pattern observation.
First, we identified that clonal expansion of CD8 cells could exert a durable cytotoxic action, leading to very significant drug response and persistent HCC tumor clearance after curative resection. This finding is in line with our recent report [10] in which we found that cases subjected to pre-operative nivolumab treatment with almost complete (> 90%) pathologic tumor necrosis could indicate long-lasting drug action, and all of them were free from tumor relapse at 35–46 months after tumor resection. Our current study further provides the mechanistic basis with evidence from TCR clonotype analysis, showing that a substantial extent of clonal expansion of cytotoxic T cells was observed in extreme responders (with pathologic tumor necrosis > 90%). It is very likely that such prominent clonal restoration of T cell cytotoxicity could elicit a marked difference in cellular anti-tumor action, and it could render additional effect to provide long-lasting tumor surveillance and clearance even after the tumor resection. This finding implicates the potential remission of HCC tumors in a specific subset of extreme responders that exhibit clonal restoration of T cell cytotoxicity and may provide important prognostic prediction for HCC patients that receive neoadjuvant nivolumab treatment.
Moreover, by retaining the necessary spatial information and deconvolution of cell type abundance and correlation, we confidently reconstructed the spatiotemporal action path of nivolumab treatment. In our multi-stage spatiotemporal reconstruction model, there were traces of regional aggregates of tumor cells and immune infiltrates (namely the MRs) that represented the frontline of cytotoxic action towards tumor cells. MRs were located between the early-stage TRs (tumor areas depleted with immune cells) and IRs (immune areas with depleted tumor cells), indicating their functional role as interface for cytotoxic anti-tumor activities, whereas the FRs probably indicate the end-stage fibroblast replacement as a consequence to the tissue necrosis.
More importantly, we were able to combine the spatiotemporal reconstruction of cytotoxicity action with the regional profiling comparison of nivolumab response. We identified evidence pinpointing the importance of humoral immune response in the responders. Together with the implications of phagocytosis and complement activation in the pre-treatment data, both innate and adaptive immune activation were suggested to govern nivolumab response consequence. Regarding useful biomarkers to predict nivolumab susceptibility, our pre-treatment data already indicated the prognostic value of innate immune response [10]. This is in concordance with the study by Carroll et al. [25], which reported that the status of innate immunity could predict the favorable clinical benefit of anti-PD1 related immunotherapy. The complement system is a crucial component of the innate immune system and also serves as a functional bridge between innate and adaptive immune responses, allowing an integrated host defense to pathogenic challenges [26]. Its function in HCC research is multifaceted and can both promote and inhibit tumor development [27]. Given that the innate immune system can modulate adaptive immune responses, it is therefore possible to assess the responsiveness of patients based on their status of natural immunity.
On the other hand, there are multiple post-treatment biomarkers that suggest susceptibility of HCC patients towards nivolumab treatment. For instance, lipid-related metabolic processes are generally enriched in non-responders. This may particularly relate to the elevated lipid biosynthesis in those cases. In contrast, the corresponding lipid catabolic processes are suggested to be enriched in responders. On the other hand, susceptibility to nivolumab treatment in patients with PPARG-high tumors may possibly be explained by the PPARG-mediated stimulation of interferon γ response, which results in the activation of PD-L1 expression and the related addiction to PD- 1 related immunosuppression [17].
In our previous investigations, we proposed the negative correlation between CD8 cells and TAMs in HCC [28, 29]. Focusing on these two cell types, we observed a plausible functional opposition between them in relation to nivolumab response. In the responders, CD8 cells and TAMs tended to locate further apart. Such spatial exclusion could also be exemplified by the difference in the regional aggregation between CD8 cells and TAMs in responders and non-responders. Our current study provides useful evidence supporting previous hypothesis proposing TAM to be an important determinant for the establishment of a phenotypic consequence of T cell-excluded tumor. In the study by Peranzoni et al. on lung cancer [30], they found that CD8 cells poorly migrated and invaded tumors due to long-lasting interactions with TAMs. The depletion of TAMs restored T cell migration and infiltration into tumor islets and improved the efficacy of anti-PD1 immunotherapy. Taken together, our preliminary evidence highlights the importance of co-administration of immune checkpoint blockade and TAM inhibition for delivering better anti-tumor immune restoration and therapeutic value.
Although single-agent immune checkpoint blockade, such as the setting with pre-operative treatment currently used in our study, could provide favorable treatment outcomes in some HCC patients, we detected signs of residual pro-tumorigenic signaling i.e. the expression CEACAM1-HAVCR2 axis, that potentially supported HCC growth and limited anti-tumor immunity. Anti-CEACAM1 and anti-HAVCR2 (or anti-TIM3 as HAVCR2 gene encodes for TIM3 protein) antibodies have been individually investigated preclinically in human cancers. The results showed that they could significantly enhance T cell activities and cytotoxicity on tumor cells to inhibit tumor cell migration and enhance response to anti-PD1 immunotherapy [31–34]. Therefore, there are many ongoing clinical trials use single- or multi-agent blockades targeting the CEACAM1-HAVCR2 axis in combination with anti-PD1 blockade. We anticipate precise patient stratification with biomarkers and relevant immune checkpoint blockade would provide a better hope for patients. In fact, our findings also support the possibility of dual targeting of PD1 and TIM3 (protein encoded by HAVCR2). Prior studies have shown that co-expression of PD1 and TIM3 was associated with more severe CD8 cell exhaustion [35–37]. Therefore, future drug development may take advantage of the potentially synergistic effect of dual targeting of PD1 and TIM3. This may also lend support to the application of anti-PD1/TIM3 bispecific antibodies (e.g. RO7121661, LB1410, AZD7789) that are currently undergoing clinical trials on solid tumors. Taken together, our integrative multimodal sequencing investigations, combined with the multi-faceted exploration of pre- and post-treatment samples of neoadjuvant nivolumab-treated HCC patients, identified useful mechanistic determinants of therapeutic response. With the exploitation of spatial transcriptomics, single-cell and bulk-cell transcriptomics, as well as TCR clonotype profiling, we reconstructed the spatiotemporal model that recapitulates the physiological restoration of T cell cytotoxicity by anti-PD1 blockade. Our findings could also possibly provide important biomarkers and explain the mechanistic basis differentiating the responders and non-responders.
Materials and methods
Patient samples
Sample collection
Informed consent was obtained from all patients. HCC tissues were obtained from patients who underwent surgical resections at the Queen Mary Hospital. All protocols and experiments were approved by the Institutional Review Board of the University of Hong Kong/Hospital Authority Hong Kong West Cluster (UW 17–056).
Spatial transcriptome
We performed the spatial transcriptome experiments using 10 × Genomics'Visium Spatial for FFPE Gene Expression Starter Kit, Human Transcriptome (PN- 1000334), following the manufacturer's procedures. First, we conducted an RNA quality assessment on selected formalin-fixed, paraffin-embedded (FFPE) tissue blocks according to 10 × Genomics'protocols (CG000408 Rev C). All the FFPE tissue blocks passed the test, with DV200 ≥ 50%. We cut 5 μm-thick sections and trimmed the selected areas of around 6.5 mm × 6.5 mm, and mounted them on the provided Visium Spatial Gene Expression Slides (PN2000233). Next, we performed deparaffinization and H&E staining according to the manufacturer's procedures (CG000409), and captured the image using a Nikon Eclipse Ti2E inverted microscope. We assembled the slides onto the Visium Cassette and performed decrosslinking, followed by probe hybridization, ligation, probe release, and extension according to the manufacturer's procedures (CG000407 Rev D). Finally, we sent the generated libraries for QC and sequencing by Illumina Novaseq platform with adequate coverage.
Oil red O, immunohistochemistry and multicolor immunofluorescence staining
Oil red staining was performed as previously described [38, 39]. For immunohistochemical staining FFPE sections, it was performed as previously described [28]. For multicolor immunofluorescence staining, we used Opal Polaris 7 Color Kit (NEL861001 KT) according to the manufacturer's recommendations. All antibodies used in this study are listed in supplementary Table S2.
Cell culture
HCC cell line Huh7 (JCRB0403), murine hepatoma cell line Hepa1 - 6, and human embryonic kidney cell line HEK293 FT (R70007) were cultured in Dulbecco’s Modified Eagle Medium (DMEM-HG, Invitrogen Gibco, Massachusetts, USA) supplemented with 10% FBS, 1% penicillin and 1% streptomycin. THP- 1 cells were cultured in the RPMI- 1640 medium (Invitrogen Gibco) supplemented with 10% FBS (v/v), 1% penicillin and 1% streptomycin. Cell lines were cultured in 37 °C and 5% CO2 incubator. Huh7 and HEK293 FT were obtained from JCRB Cell Bank and Invitrogen (Carlsbad, USA), respectively. Hepa1 - 6 and THP- 1 were obtained from American Type Culture Collection (ATCC). All cell cultures were tested negative for Mycoplasma contamination. For the recombinant UBASH3B protein-treatment experiment, Huh7 cells seeded at 3.2 × 10 [5] cells per well in a 6-well plate were treated on the next day with 20 µg/ml recombinant UBASH3B protein (cat. No.: HY-P74484), which was purchased from MedChemExpress (New Jersey, NJ) and dissolved in 0.22 µm-filterd 0.1% bovine serum albumin (BSA), for 24 h before examining the CEACEM1 mRNA expression by qPCR.
Establishment of luciferase-labeled cell lines
For viral packaging, pCDH-CMV-Luc-EF1a-Hygro plasmid containing the luciferase gene for overexpression was co-transfected with viral packaging mix (Invitrogen) into HEK293 FT cells using Lipofectamine 2000 reagent (Invitrogen). The viral supernatant was collected to transduce Hepa1 - 6 cells, followed by selection with 500 μg/mL Hygromycin B to generate luciferase-labeled Hepa1 - 6 cell line for in vivo experiments. Luciferase reporter assay was used to confirm successful overexpression of the luciferase gene.
Establishment of stable knockdown
Stable knockdown cells for respective genes in THP- 1, Huh7 and luciferase-labelled Hepa1 - 6 cells were established as previously described [29, 40, 41]. pLKO.1-Puro short hairpin (shRNA) expression constructs for targeting the various gene mRNA were from Sigma-Aldrich (St Louis, MO). According to the manufacturer’s protocol for the MISSION® Lentiviral Packaging System (Sigma-Aldrich, St Louis, MO), viral packaging for the lentiviruses containing the shRNA vectors occurred in HEK293 FT and the viral particles containing shRNA were then transduced into HCC cells or THP- 1 cell line to establish the shRNA stably expressing cells, which were subjected to selection by puromycin.
The lentiviral-based vector pLKO.1 was used to express shRNA targeting in mouse Hepa1 - 6 cells. The oligonucleotides (5’- CAACTCAATCAAGCTGGACAT- 3’) were annealed and subcloned into pLKO.1. To stably silence the gene expression in cells, the pLKO.1 shRNA was transfected with lentiviral packaging plasmids into HEK293 T cells to generate lentiviruses. The lentiviral supernatants were collected after 48 h transfection. Cells were then infected with lentiviruses for 24 h with polybrene and selected with puromycin for 72 h. The knockdown efficiency was assessed by quantitative real-time PCR. All shRNA sequences used in this study are provided in supplementary Table S2.
Conditioned medium collection and treatment
For the study using THP- 1 cell conditioned medium, the THP- 1 cells were seeded at 2 × 104 cells in 1 ml per well in 12-well plate and the conditioned media were collected 4 days later, with cells spun down and the supernatant filtered by 0.22 µm syringe filter (Sartorius, Germany). The media were then used to treat Huh7 cells seeded at 1 × 104 cells per well in 96-well plate one day before. The 24 h-treated Huh7 cells were subjected to lysis for collection of the mRNA to make cDNA by using the Cell-to-CT kit (Thermo Fisher, Massachusetts, MA) for qPCR for CEACAM1.
RNA extraction, reverse transcription and quantitative polymerase chain reaction (qPCR)
Total RNA was extracted by Trizol (Invitrogen) and cDNA was synthesized by reverse transcription kit (Invitrogen). qRT-PCR was performed as described [29, 40, 41], using ABI Power SYBR® Green master mix and detected by ABI QuantStudio 5 Real-Time PCR System (Applied Biosystems, Foster City, CA). All primer sequences used in this study are provided in supplementary Table S3.
Animal experiments
Orthotopic liver injection assay
For the orthotopic liver injection model, 3 × 106 luciferase-labeled Hepa1 - 6 cells were injected into the left lobe of the liver of each C57BL/6 mouse. Two weeks post-injection, the mice were injected with 100 mg/kg D-luciferin (PerkinElmer) and then subjected to bioluminescence imaging by the IVIS Spectrum Imaging System (Perkin Elmer). Then, the livers were dissected and imaged as previously described [42]. The intensity of corresponding bioluminescence signals detected was used to indicate the burden of the liver tumor xenografts. The animal experiment was approved by the Committee on the Use of Live Animals in Teaching and Research (CULATR 5688–21), Li Ka Shing Faculty of Medicine, University of Hong Kong.
Flow cytometry assay
For flow cytometry-based analysis of tumor-infiltrating T cells, single cells derived from the dissociated orthotopic tumor tissues were blocked with anti-mouse CD16/32 antibody (101,320, Biolegend, San Diego, USA) at room temperature for 10 min and then incubated with fluorochrome-conjugated primary antibodies at 4 °C for 30 min before detection by BioRad ZE5 TM Cell Analyzer (Bio-Rad Laboratories, California, USA). Primary antibodies and their concentrations are listed in Table S2. Software FlowJo was used for data analysis. Gating strategies for tumor-infiltrating lymphocyte analysis are provided in Fig. 8. For sorting-based measurement of the expression of genes, CD45 +/CD8 + positive T cells were sorted and collected by a Cell Sorter BD FACSAria SORP (BD Biosciences, USA). All antibodies used in this study are provided in supplementary Table S2.
Quantification and statistical analysis
ScRNA-seq data processing
Raw data alignment and quality control
The sequenced data were subjected to processing using the Cell Ranger Software Suite (v5.0.1, 10X Genomics). In brief, Illumina base call (BCL) sequence files were demultiplexed to obtain FASTQ format, followed by the alignment of FASTQ files from each sample to the human GRCh38 reference genome “refdata-gex-GRCh38 - 2020-A” (https://cf.10xgenomics.com/supp/spatial-exp/refdata-gex-GRCh38-2020-A.tar.gz). Upon completion of the default quality control filtering for individual samples, a unique molecular identifier (UMI) count matrix, accompanied by feature barcodes, was generated for further analysis.
Unsupervised clustering and marker identification
The UMI count matrix was converted to a Seurat object using the R package Seurat (v4.4.0) [43]. A procedure was employed to exclude low-quality cells from each sample, wherein cells with fewer than 200 detected genes or more than 5,000 genes and cells with over 20% of UMIs derived from mitochondrial genes were omitted. The FindIntegrationAnchors and IntegrateData functions were used to integrate cells from different datasets into a shared space for anchor identification and eliminate batch effects. To account for disparities in sequencing depth across cells, the datasets were normalized using regularized negative binomial regression (SCTransform) with default parameters and the first 30 principal components were selected through principal component analysis (PCA) for unsupervised clustering via the t-Distributed Stochastic Neighbor Embedding (t-SNE) algorithms at a resolution of 1.6, and cell types were annotated based on prior knowledge of marker genes.
Differential expression and functional enrichment analysis
To characterize each cluster, differentially expressed genes (DEGs) for each subset were identified using the non-parametric Wilcoxon Rank Sum test through the “FindMarkers” function in the Seurat package. DEGs were filtered using a minimum log2 FC of 0.25 and a maximum p-value of 0.05. For enrichment analysis, DEGs were employed using the ClusterProfiler package (v4.2.2) [44]. In brief, gene symbols were converted to ENTREZ IDs with the"org.Hs.eg.db"package (v3.14.0), followed by the GO enrichment analysis via the “enrichGO” function by the BH method.
Odds ratio
We used a previously published article to calculate the odd ratio in cell-type composition between the two groups [45]. A Wilcox test was used to compare the mean values between the two groups.
T cells sub-clustering
The dimensional reduction was specifically re-performed on the T cells. A sub-clustering analysis of the T cells was performed, analogous to the previously described clustering methodology. Following this, clusters were merged into major cell classes by combining related/highly similar clusters according to the expression of marker genes.
TCR-seq data processing
Raw data alignment and quality control
The droplet-based TCR-seq data was processed using Cell Ranger (version 3.0.1, 10 × Genomics) against the human GRCh38 vdj reference provided by 10 × Genomics (https://cf.10xgenomics.com/supp/cell-vdj/refdata-cellranger-vdj-GRCh38-alts-ensembl-7.1.0.tar.gz). we used scRepertoire (v1.10.1) package [46] to integrate the results with the aforementioned Seurat object (extracted T cell populations). During this procedure, individual clonotypes were determined using the"CTstrict"criteria, which stipulate that a clonotype must possess an identical nucleotide sequence within the TCR.
Differential expression and functional enrichment analysis
The method employed in this section is consistent with the previously described scRNA-seq analysis.
ST-seq data processing
Raw data alignment and quality control
Raw FASTQ files and histology images were aligned to the hg38 reference genome “refdata-gex-GRCh38 - 2020-A” by Space Ranger (v1.3.1) using default parameters. For the generated spot matrixes, the basic QCs were applied to the data. In detail, the spots with extremely low nUMI or nGene (outliers) were eliminated, as well as spots where more than 2% of UMIs originated from mitochondrial genes.
Dimension reduction and unsupervised clustering
After QC, the raw Unique Molecular Identifiers (UMI) count spot matrices, pictures, spot-image coordinates were imported to R (v4.3.2). We used the R package Seurat package (v4.4.0) [43] to perform the basic dimension reduction, unsupervised clustering as well as visualization and harmony (v1.0) [47] to integrate the expression data from different sections of each patient. Due to the possibility of each spot carrying genes from 2–10 cells, we observed that integrating all samples and using Seurat’s conventional methods for spatial clustering of specific cell types did not yield satisfactory outcomes. Therefore, we conducted clustering analysis on each individual sample. Additionally, we integrated the data using Harmony by setting the section source as the batch factor and facilitate TSNE visualization, but the spatial representation primarily relied on the clustering of individual samples. In detail, the datasets were individually normalized by the comprehensive regularized negative binomial regression (SCTransform) with default parameters to remove the differences in sequencing depth across cells, and the top 3000 highly variable genes were used for dimensionality reduction for each sample. The top 30 dimensions were obtained by performing principal component analysis (PCA) to project the spots into a low-dimensional space. To ensure a comprehensive identification of all potential subpopulations, unsupervised clustering was conducted on the basis of shared nearest neighbor graphs with resolution parameters varying from 0.2 to 3, with intervals of 0.2. After manually integrated similar clusters, the t-Distributed Stochastic Neighbor Embedding (t-SNE) algorithms were used for visualization with resolution = 3 and the cell types were annotated on the basis of the prior knowledge of marker genes.
Deconvolution
We first calculated each spot's cell abundance and spatial dependence using “BayesPrism” [11] and “mistyR” [15], respectively. Default values were used for all other code parameters unless otherwise stated. We then calculate the top 1 cell abundance and the top 1 spatial dependence of the associated cell type for each spot. If both identities are the same, the spot is an enriched cell type. If the spatial dependence of all cell types is less than 0, the primary cell type of the spot is the cell type of the top 1 cell abundance, which is also an enriched cell type. If the cell type of the top 1 cell abundance and the top 1 cell spatial dependence for each spot are inconsistent and the spatial dependence is greater than 0, the spot is defined as a mixed spot.
Region classification
After performing deconvolution, a normalized matrix of cell composition was obtained for each ST spot and computed the mean value within each cluster as identified by Seurat. To merge similar clusters, we initially employed the clValid (v0.7) R package [48] to assess three clustering techniques: hierarchical, k-means, and Partitioning Around Medoids (PAM) with the number of clusters ranging from 2 to 8, and ultimately the matrix subjected to hierarchical clustering with optimal K (k = 2) determined according to the Shiloutte method implemented in the fviz_nbclust function of the factoextra (v1.0.7) package (http://www.sthda.com/english/rpkgs/factoextra). A comparable methodology was adopted for the refined categorization of both tumor and stromal regions, implementing k = 2 and k = 3 for each region, respectively.
Differential expression and functional enrichment analysis
The method employed in this section is consistent with the previously described scRNA-seq analysis. In addition, the STRING database (https://cytoscape.org/) was used to construct the protein–protein interaction network.
Gene set enrichment analysis
Single-cell gene set enrichment analysis (ssGSEA) was conducted utilizing the escape R package (v1.4.0) [12]. ssGSEA scores were computed on this normalized matrix using MSigDB hallmark gene sets for each spot, followed by Wilcox test analysis.
Definition of cellular scores and signature attributes
To define the features of humoral immune, inflammatory, cholesterol biosynthetic process, and carboxylic acid biosynthetic process, the gene enrolled in these pathways was taken from GO enrichment results, and gene set scoring was performed using the R package UCell (v2.6.2) [49]. The features of proliferation, hepatic function (Table S4), M1/M2 signature, and CD8 naive/exhausted/cytotoxic signature were delineated utilizing UCell (v2.6.2). Corresponding gene sets associated with these functions were described by Sun et al. [13], Azizi et al [50]. and Ma et al. [51], respectively.
Copy Number Variation (CNV) calling
We used a method based on the"SPATA2"package [52] to infer copy number variation (CNV) status. We extracted all tumor-containing spots for copy number variation inference. The reference dataset includes all spots that do not contain tumors, and all other code parameters use defaults unless otherwise noted.
Homotypic and heterotypic score calculations
Homotypic and heterotypic scores were calculated based on previous studies [19, 20], respectively. The geneset for CD8 functional scores was obtained from a previous study [51, 53]. Squidpy was performed to obtain Ripley's L results [22].
Cell–cell interaction analysis
We used the “stLearn” [50] to calculate the communication of immune checkpoint LRs within spots. The ligand-target network was calculated using Nichenet's R package [23]. We retrieved ligand and receptor information from the “CellPhoneDB” algorithm [54] to explore inter-spot interactions and performed calculations. In addition, gene communication strength and signal flow direction were calculated from the “COMMOT” [55] and “SPATA2” [52] packages, respectively.
Statistical analysis
For the scRNA-seq, TCR-seq, and ST-seq data, most statistical analyses and graphics production were performed using R (v4.1.0), while a minor proportion of incompatible packages were analyzed employing v4.3.2 (Foundation for Statistical Computing). Statistical analyses and graphics production for the experimental data were performed using GraphPad Prism 7 (GraphPad Software).
Supplementary Information
Supplementary Material 1. Fig. S1. ScRNA-seq of HCC and distribution of clonal expansion levels in T cell clusters in relation to Figure 1. A. t-SNE visualization of markers for the cell types as indicated. B. Stacked bar plot of the proportion of cell types for the tumors in R and NR groups. C. Heatmap of the expression of the top 10 marker genes for each cell cluster. D. Pie chart showing the overall proportion of each cell type in the tumors of N and NR groups. E. t-SNE visualization of T cell with TCR repertoires. Each cell identified with a TCR clonotype is labeled blue; otherwise it is labeled grey (no TCRα/β or TCRγ/δ identified).
Supplementary Material 2. Fig. S2. Spatial representation of normalized markers for the cell types in HCC in relation to Figure 2.
Supplementary Material 3. Fig. S3. Procedures of clustering in different regions and corresponding spatial distribution in relation to Figure 2. A. Procedure of clustering for all clusters. B. Spatial mapping of two distinct regions. C. Procedure of clustering for tumor region (upper panel) and stromal region (lower panel). D. Spatial mapping of five distinct regions.
Supplementary Material 4. Fig. S4. Analysis of spatial contextualized views in relation to Figure 3. A. Stacked bar plot of the proportion of cell types for each region. B. Spatial representation of normalized markers for each region in HCC.
Supplementary Material 5. Fig. S5. Differential characteristics and stemness gene expression of tumor cells across groups in relation to Figure 4. A. Relative changes in cell ratios in different regions across the two groups. B. Bar chart of gene ontology enrichment for upregulated genes in R and NR groups. C. Correlation analysis between the degree of pathologic tumor necrosis and established liver CSC markers.
Supplementary Material 6. Fig. S6. Landscape of cell abundance and spatial dependence in spatial transcriptome. A. Spatial visualization of the cell abundance of the 16 samples. Each color represents one cell type, and the color shades represent the cell abundance. B. Spatial visualization of the spatial dependence of the 16 samples. Each color represents one cell type, and the color shades represent the size of the spatial dependence.
Supplementary Material 7. Fig. S7. Immune and CNV landscape with neoadjuvant nivolumab treatment. A. Heatmap showing the Odds Ratio (OR) of CD8 related spots between the R and NR groups. B. Heatmap showing the Odds Ratio (OR) of CD4 related spots between the R and NR groups. C.Landscape of the copy number variation in tumors with neoadjuvant nivolumab treatment.
Supplementary Material 8. Fig. S8. CNV landscape is a key factor for neoadjuvant nivolumab efficacy. A. The ranking plot showing the differential CNV genes between R and NR groups. B. MversusA plot shows differential expression genes between R and NR groups. C. The donut chart showing the composition of the DEGs in the R (left panel) and NR (right panel) groups. D. Dot plot showing the pathway enrichment of DEGs with CNV occurred. E. PPARG-associated protein-protein interaction network in the R group. F. Boxplot showing the statistical comparison of PPARG expression between CNV gain and CNV loss groups. G. The expression of PPARG between HCCs and the corresponding non-tumor liver (NTL) in the HKU-QMH cohort. H. Kaplan-Meier analysis for disease-free and overall survival rates in relation to PPARG expression in HCC tumors in the HKU-QMH cohort. I. The clinicopathological correlation of PPARG expression in HCCs in HKU-QMH cohort. Higher PPARG expression was associated with more aggressive tumor behavior. J. Kaplan-Meier analysis for overall survival rates in relation to PPARG expression in TCGA database.
Supplementary Material 9. Fig. S9. TAM and CD8 are key cell types for the efficacy of neoadjuvant nivolumab treatment. A. Box plot showing the statistical comparison of heterotypic scores (CD8Rel to TAMRel, with CD8Rel serving as center point) between R and NR groups. B. Box plot showing the statistical comparison of the functional scores of CD8Rel surrounding TAMRel between R and NR groups. C. Heatmap showing the ligand-target network of the enriched tumor via NiecheNet. D. Heatmap showing the ligand-target network of the enriched apCAF via NiecheNet. E. Heatmap showing summarized numbers of ligand-receptor pairs among different cell-cell interaction pairs using CellphoneDB.
Supplementary Material 10. Fig. S10. Cell-cell communication between R and NR groups. A.Dot plot showing the expression of ligand-receptor pair in different cell types. B. Correlation of NR1I2 and CEACAM1 with the percentage of pathologic tumor necrosis. C. Correlation between HAVCR2 expression (left panel) and HAVCR2 communication signal strength (right panel) and degree of pathologic tumor necrosis. D. Bar plot showing the knockdown efficiency of UBASH3B in THP-1 cell line. E. Bar plot showing the knockdown efficiency of CEACAM1 in Hepa1-6 cells for orthotopic liver injection HCC model. F. Orthotopic tumors for the non-target control (shNTC) and shCEACAM1 Hepa1-6 cell groups.
Acknowledgements
We thank the Centre for PanorOmic Sciences (CPOS) of the LKS Faculty of Medicine of the University of Hong Kong for providing the facilities and services for scRNA-seq, TCR-seq, and ST-seq analysis.
Abbreviations
- TAMs
Tumor-associated macrophages
- HCC
Hepatocellular carcinoma
- MKIs
Multikinase inhibitors
- MTAs
Molecular targeted agents
- ICIs
Immune checkpoint inhibitors
- PD- 1
Programmed cell death- 1
- PD-L1
Programmed cell death-ligand 1
- TME
Tumor microenvironment
- scRNA-seq
Single-cell RNA sequencing
- bulk RNA-seq
Bulk RNA sequencing
- ST-seq
Spatial transcriptomic sequencing
- TCR-seq
T-cell receptor sequencing
- R
Responders
- NR
Non-responders
- tSNE
T-distributed stochastic neighbor embedding
- CAFs
Cancer-associated fibroblasts
- GO
Gene Ontology
- H&E
Histology with hematoxylin and eosin staining
- TR
Tumor Regions
- FR
Fibrotic Region
- IR
Immune Region
- MR
Mixed Region
- DEGs
Differentially expressed genes
- ssGSEA
Single-sample gene set enrichment analysis
- TIBs
Tumor-infiltrating B cells
- LCSC
Liver cancer stem cell
- ECM
Extracellular matrix
- iCAF
Inflammatory CAF
- mCAF
Myofibroblastic CAF
- apCAF
Antigen-presenting CAF
- CNV
Copy number variation
- L-R
Ligand-receptor pairs
- shCEACAM1
CEACAM1 knockdown
- NTC
Non-target control
Authors’ contributions
Study concept and design: DWH, ION; Acquisition of data: YT, GCC, PL, JML, HM, LT, AH, JL, VXZ, ION; Analysis and interpretation of data: FZ, QZ, DWH, ION; Acquisition of clinical samples: TC, ION; Drafting of the manuscript: FZ, QZ, DWH, ION; All authors reviewed and approved the final draft of the manuscript.
Funding
The study was supported by the Hong Kong Research Grants Council Theme-based Research Scheme (T12-704/16-R and T12-716/22-R), Innovation and Technology Commission grant to State Key Laboratory of Liver Research (ITC PD/17-9), Health and Medical Research Fund (10212956 & 07182546), RGC General Research Fund (17100021 & 17117019), and University Development Fund of The University of Hong Kong. I.O.L. Ng is Loke Yew Professor in Pathology.
Data availability
No datasets were generated or analysed during the current study.
Declarations
Ethics approval and consent to participate
All human experiments were approved by the Institutional Review Board of the University of Hong Kong/Hospital Authority Hong Kong West Cluster (UW 17–056). The animal experiment was approved by the Committee on the Use of Live Animals in Teaching and Research (CULATR 5688–21), Li Ka Shing Faculty of Medicine, University of Hong Kong.
Competing interests
The authors declare no competing interests.
Footnotes
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Fanhong Zeng and Qingyang Zhang contributed equally to this work.
Contributor Information
Daniel Wai-Hung Ho, Email: dwhho@hku.hk.
Irene Oi-Lin Ng, Email: iolng@hku.hk.
References
- 1.Llovet JM, et al. Hepatocellular carcinoma. Nat Rev Dis Primers. 2021;7(6):151–72. [DOI] [PubMed] [Google Scholar]
- 2.Hasegawa K, et al. Comparison of resection and ablation for hepatocellular carcinoma: a cohort study based on a Japanese nationwide survey. J Hepatol. 2013;58:724–9. [DOI] [PubMed] [Google Scholar]
- 3.Villanueva A. Hepatocellular carcinoma. N Engl J Med. 2019;380:1450–62. [DOI] [PubMed] [Google Scholar]
- 4.Haslam A, Prasad V. Estimation of the percentage of US patients with cancer who are eligible for and respond to checkpoint inhibitor immunotherapy drugs. JAMA Netw Open. 2019;2: e192535. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Ho WJ, et al. Neoadjuvant cabozantinib and nivolumab converts locally advanced HCC into resectable disease with enhanced antitumor immunity. Nat Cancer. 2021;2:891–903. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Zhu XD, et al. Downstaging and resection of initially unresectable hepatocellular carcinoma with tyrosine kinase inhibitor and anti-PD-1 antibody combinations. Liver Cancer. 2021;10:320–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.LaPelusa M, et al. Tissue and imaging biomarkers of response to neoadjuvant nivolumab or nivolumab plus ipilimumab in patients with resectable hepatocellular carcinoma. Oncology. 2024:1-8. [DOI] [PMC free article] [PubMed]
- 8.Rossi AJ, Khan TM, Saif A, Marron TU, Hernandez JM. Treatment of hepatocellular carcinoma with neoadjuvant nivolumab alone versus in combination with a CCR2/5 inhibitor or an anti-IL-8 antibody. Ann Surg Oncol. 2022;29:30–2. [DOI] [PubMed] [Google Scholar]
- 9.Pinato DJ, et al. PRIME-HCC: phase Ib study of neoadjuvant ipilimumab and nivolumab prior to liver resection for hepatocellular carcinoma. BMC Cancer. 2021;21:301. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Cheung TT, et al. Multimodal integrative genomics and pathology analyses in neoadjuvant nivolumab treatment for intermediate and locally-advanced HCC. Liver Cancer. 2024;13:70–88. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Chu T, Wang Z, Pe’er D, Danko CG. Cell type and gene expression deconvolution with BayesPrism enables Bayesian integrative analysis across bulk and single-cell RNA sequencing in oncology. Nat Cancer. 2022;3:505–17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Borcherding N, et al. Mapping the immune environment in clear cell renal carcinoma by single-cell genomics. Commun Biol. 2021;4:122. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Sun Y, et al. Single-cell landscape of the ecosystem in early-relapse hepatocellular carcinoma. Cell. 2021;184:404–421 e416. [DOI] [PubMed] [Google Scholar]
- 14.Lee TK, Guan XY, Ma S. Cancer stem cells in hepatocellular carcinoma - from origin to clinical implications. Nat Rev Gastroenterol Hepatol. 2022;19:26–44. [DOI] [PubMed] [Google Scholar]
- 15.Tanevski J, Flores ROR, Gabor A, Schapiro D, Saez-Rodriguez J. Explainable multiview framework for dissecting spatial relationships from highly multiplexed data. Genome Biol. 2022;23:97. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Zhang M, et al. Single-cell transcriptomic architecture and intercellular crosstalk of human intrahepatic cholangiocarcinoma. J Hepatol. 2020;73:1118–30. [DOI] [PubMed] [Google Scholar]
- 17.Mimura K, et al. PD-L1 expression is mainly regulated by interferon gamma associated with JAK-STAT pathway in gastric cancer. Cancer Sci. 2018;109:43–53. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Pfister D, et al. NASH limits anti-tumour surveillance in immunotherapy-treated HCC. Nature. 2021;592:450–6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Backdahl J, et al. Spatial mapping reveals human adipocyte subpopulations with distinct sensitivities to insulin. Cell Metab. 2021;33:1869–1882 e1866. [DOI] [PubMed] [Google Scholar]
- 20.Bergenstrahle J, Larsson L, Lundeberg J. Seamless integration of image and molecular analysis for spatial transcriptomics workflows. BMC Genomics. 2020;21:482. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Gebhardt T, Park SL, Parish IA. Stem-like exhausted and memory CD8(+) T cells in cancer. Nat Rev Cancer. 2023;23:780–98. [DOI] [PubMed] [Google Scholar]
- 22.Palla G, et al. Squidpy: a scalable framework for spatial omics analysis. Nat Methods. 2022;19:171–8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Guilliams M, et al. Spatial proteogenomics reveals distinct and evolutionarily conserved hepatic macrophage niches. Cell. 2022;185:379–396 e338. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Chen A, et al. Targeting the oncogenic m6A demethylase FTO suppresses tumourigenesis and potentiates immune response in hepatocellular carcinoma. Gut. 2024;74:90–102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Carroll TM, et al. Tumor monocyte content predicts immunochemotherapy outcomes in esophageal adenocarcinoma. Cancer Cell. 2023;41:1222–1241 e1227. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Dunkelberger JR, Song WC. Complement and its role in innate and adaptive immune responses. Cell Res. 2010;20:34–50. [DOI] [PubMed] [Google Scholar]
- 27.Reis ES, Mastellos DC, Ricklin D, Mantovani A, Lambris JD. Complement in cancer: untangling an intricate relationship. Nat Rev Immunol. 2018;18:5–18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Ho DW, et al. Single-cell RNA sequencing shows the immunosuppressive landscape and tumor heterogeneity of HBV-associated hepatocellular carcinoma. Nat Commun. 2021;12:3684. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Zhang Q, et al. Reciprocal interactions between malignant cells and macrophages enhance cancer stemness and M2 polarization in HBV-associated hepatocellular carcinoma. Theranostics. 2024;14:892–910. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Peranzoni E, et al. Macrophages impede CD8 T cells from reaching tumor cells and limit the efficacy of anti-PD-1 treatment. Proc Natl Acad Sci U S A. 2018;115:E4041–50. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Acharya N, Sabatos-Peyton C, Anderson AC. Tim-3 finds its place in the cancer immunotherapy landscape. J Immunother Cancer. 2020;8:e000911. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Sauer N, et al. TIM-3 as a promising target for cancer immunotherapy in a wide range of tumors. Cancer Immunol Immunother. 2023;72:3405–25. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Zhang J, et al. The role of Tim-3 blockade in the tumor immune microenvironment beyond T cells. Pharmacol Res. 2024;209: 107458. [DOI] [PubMed] [Google Scholar]
- 34.Zhao L, Li T, Zhou Y, Wang P, Luo L. Monoclonal antibody targeting CEACAM1 enhanced the response to anti-PD1 immunotherapy in non-small cell lung cancer. Int Immunopharmacol. 2024;143: 113395. [DOI] [PubMed] [Google Scholar]
- 35.Granier C, et al. Tim-3 expression on tumor-infiltrating PD-1(+)CD8(+) T cells correlates with poor clinical outcome in renal cell carcinoma. Cancer Res. 2017;77:1075–82. [DOI] [PubMed] [Google Scholar]
- 36.Zhou Q, et al. Coexpression of Tim-3 and PD-1 identifies a CD8+ T-cell exhaustion phenotype in mice with disseminated acute myelogenous leukemia. Blood. 2011;117:4501–10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Jin HT, et al. Cooperation of Tim-3 and PD-1 in CD8 T-cell exhaustion during chronic viral infection. Proc Natl Acad Sci U S A. 2010;107:14733–8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Husain A, et al. Ephrin-A3/EphA2 axis regulates cellular metabolic plasticity to enhance cancer stemness in hypoxic hepatocellular carcinoma. J Hepatol. 2022;77:383–96. [DOI] [PubMed] [Google Scholar]
- 39.Lee D, et al. Folate cycle enzyme MTHFD1L confers metabolic advantages in hepatocellular carcinoma. J Clin Invest. 2017;127:1856–72. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Tsui YM, et al. Sorted-cell sequencing on HCC specimens reveals EPS8L3 as a key player in CD24/CD13/EpCAM-triple positive, stemness-related HCC cells. Cell Mol Gastroenterol Hepatol. 2024;18: 101358. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Huang H, et al. LANCL1, a cell surface protein, promotes liver tumor initiation through FAM49B-Rac1 axis to suppress oxidative stress. Hepatology. 2024;79:323–40. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Wang X, et al. S100A10 promotes HCC development and progression via transfer in extracellular vesicles and regulating their protein cargos. Gut. 2023;72:1370–84. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Stuart T, et al. Comprehensive integration of single-cell data. Cell. 2019;177:1888–1902 e1821. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Yu GC, Wang LG, Han YY, He QY. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS. 2012;16:284–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Zheng L, et al. Pan-cancer single-cell landscape of tumor-infiltrating T cells. Science. 2021;374:abe6474. [DOI] [PubMed] [Google Scholar]
- 46.Borcherding N, Bormann NL, Kraus G. scRepertoire: an R-based toolkit for single-cell immune receptor analysis. F1000Res. 2020;9:47. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Korsunsky I, et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat Methods. 2019;16:1289–96. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Brock G, Datta S, Pihur V, Datta S. clValid: an R package for cluster validation. J Stat Softw. 2008;25:1–22. [Google Scholar]
- 49.Andreatta M, Carmona SJ. UCell: Robust and scalable single-cell gene signature scoring. Comput Struct Biotechnol J. 2021;19:3796–8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Azizi E, et al. Single-cell map of diverse immune phenotypes in the breast tumor microenvironment. Cell. 2018;174:1293–1308 e1236. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Ma L, et al. Single-cell atlas of tumor cell evolution in response to therapy in hepatocellular carcinoma and intrahepatic cholangiocarcinoma. J Hepatol. 2021;75:1397–408. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Ravi VM, et al. Spatially resolved multi-omics deciphers bidirectional tumor-host interdependence in glioblastoma. Cancer Cell. 2022;40:639–655 e613. [DOI] [PubMed] [Google Scholar]
- 53.Bi K, et al. Tumor and immune reprogramming during immunotherapy in advanced renal cell carcinoma. Cancer Cell. 2021;39:649-+. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Garcia-Alonso L, et al. Single-cell roadmap of human gonadal development. Nature. 2022;607:540–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Cang Z, et al. Screening cell-cell communication in spatial transcriptomics via collective optimal transport. Nat Methods. 2023;20:218–28. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Supplementary Material 1. Fig. S1. ScRNA-seq of HCC and distribution of clonal expansion levels in T cell clusters in relation to Figure 1. A. t-SNE visualization of markers for the cell types as indicated. B. Stacked bar plot of the proportion of cell types for the tumors in R and NR groups. C. Heatmap of the expression of the top 10 marker genes for each cell cluster. D. Pie chart showing the overall proportion of each cell type in the tumors of N and NR groups. E. t-SNE visualization of T cell with TCR repertoires. Each cell identified with a TCR clonotype is labeled blue; otherwise it is labeled grey (no TCRα/β or TCRγ/δ identified).
Supplementary Material 2. Fig. S2. Spatial representation of normalized markers for the cell types in HCC in relation to Figure 2.
Supplementary Material 3. Fig. S3. Procedures of clustering in different regions and corresponding spatial distribution in relation to Figure 2. A. Procedure of clustering for all clusters. B. Spatial mapping of two distinct regions. C. Procedure of clustering for tumor region (upper panel) and stromal region (lower panel). D. Spatial mapping of five distinct regions.
Supplementary Material 4. Fig. S4. Analysis of spatial contextualized views in relation to Figure 3. A. Stacked bar plot of the proportion of cell types for each region. B. Spatial representation of normalized markers for each region in HCC.
Supplementary Material 5. Fig. S5. Differential characteristics and stemness gene expression of tumor cells across groups in relation to Figure 4. A. Relative changes in cell ratios in different regions across the two groups. B. Bar chart of gene ontology enrichment for upregulated genes in R and NR groups. C. Correlation analysis between the degree of pathologic tumor necrosis and established liver CSC markers.
Supplementary Material 6. Fig. S6. Landscape of cell abundance and spatial dependence in spatial transcriptome. A. Spatial visualization of the cell abundance of the 16 samples. Each color represents one cell type, and the color shades represent the cell abundance. B. Spatial visualization of the spatial dependence of the 16 samples. Each color represents one cell type, and the color shades represent the size of the spatial dependence.
Supplementary Material 7. Fig. S7. Immune and CNV landscape with neoadjuvant nivolumab treatment. A. Heatmap showing the Odds Ratio (OR) of CD8 related spots between the R and NR groups. B. Heatmap showing the Odds Ratio (OR) of CD4 related spots between the R and NR groups. C.Landscape of the copy number variation in tumors with neoadjuvant nivolumab treatment.
Supplementary Material 8. Fig. S8. CNV landscape is a key factor for neoadjuvant nivolumab efficacy. A. The ranking plot showing the differential CNV genes between R and NR groups. B. MversusA plot shows differential expression genes between R and NR groups. C. The donut chart showing the composition of the DEGs in the R (left panel) and NR (right panel) groups. D. Dot plot showing the pathway enrichment of DEGs with CNV occurred. E. PPARG-associated protein-protein interaction network in the R group. F. Boxplot showing the statistical comparison of PPARG expression between CNV gain and CNV loss groups. G. The expression of PPARG between HCCs and the corresponding non-tumor liver (NTL) in the HKU-QMH cohort. H. Kaplan-Meier analysis for disease-free and overall survival rates in relation to PPARG expression in HCC tumors in the HKU-QMH cohort. I. The clinicopathological correlation of PPARG expression in HCCs in HKU-QMH cohort. Higher PPARG expression was associated with more aggressive tumor behavior. J. Kaplan-Meier analysis for overall survival rates in relation to PPARG expression in TCGA database.
Supplementary Material 9. Fig. S9. TAM and CD8 are key cell types for the efficacy of neoadjuvant nivolumab treatment. A. Box plot showing the statistical comparison of heterotypic scores (CD8Rel to TAMRel, with CD8Rel serving as center point) between R and NR groups. B. Box plot showing the statistical comparison of the functional scores of CD8Rel surrounding TAMRel between R and NR groups. C. Heatmap showing the ligand-target network of the enriched tumor via NiecheNet. D. Heatmap showing the ligand-target network of the enriched apCAF via NiecheNet. E. Heatmap showing summarized numbers of ligand-receptor pairs among different cell-cell interaction pairs using CellphoneDB.
Supplementary Material 10. Fig. S10. Cell-cell communication between R and NR groups. A.Dot plot showing the expression of ligand-receptor pair in different cell types. B. Correlation of NR1I2 and CEACAM1 with the percentage of pathologic tumor necrosis. C. Correlation between HAVCR2 expression (left panel) and HAVCR2 communication signal strength (right panel) and degree of pathologic tumor necrosis. D. Bar plot showing the knockdown efficiency of UBASH3B in THP-1 cell line. E. Bar plot showing the knockdown efficiency of CEACAM1 in Hepa1-6 cells for orthotopic liver injection HCC model. F. Orthotopic tumors for the non-target control (shNTC) and shCEACAM1 Hepa1-6 cell groups.
Data Availability Statement
No datasets were generated or analysed during the current study.