Abstract
We study a linear evolutionary model based on the two-dimensional distribution of protocells by total enantiomeric excess and the amount of stored information, which they can pass from generation to generation, and without any mutual inhibition. We show that the evolution of such systems occurs in four distinct stages. The first stage is an exponential growth of the concentration of protocells near the point  and it should take negligible time on a geological scale. The second stage is a diffusion-like process in both dimensions. This process can also be accompanied by a drift in the direction of increased information passed from generation to generation, provided that the appropriate linear coefficient in the information storage subspace is large enough. The third stage is a rapid symmetry breaking and formation of the species near
and it should take negligible time on a geological scale. The second stage is a diffusion-like process in both dimensions. This process can also be accompanied by a drift in the direction of increased information passed from generation to generation, provided that the appropriate linear coefficient in the information storage subspace is large enough. The third stage is a rapid symmetry breaking and formation of the species near  value of enantiomeric excess (assuming a small positive global enantiomeric asymmetry factor). The fourth stage is a relaxation toward a global stationary point, which is a narrow peak located near
value of enantiomeric excess (assuming a small positive global enantiomeric asymmetry factor). The fourth stage is a relaxation toward a global stationary point, which is a narrow peak located near  value of enantiomeric excess and some optimal value of the amount of stored information.
value of enantiomeric excess and some optimal value of the amount of stored information.
Supplementary Information
The online version contains supplementary material available at 10.1038/s41598-025-97319-2.
Keywords: Molecular evolution, Chiral symmetry breaking, Biological information storage
Subject terms: Origin of life, Evolutionary theory
Introduction
The emergence of life on Earth is a question that has captivated scientists, philosophers, and thinkers for centuries. Among the critical stages in the origins of life is the formation and evolution of protocells, the rudimentary biological compartments that preceded modern cells. While theories surrounding abiogenesis and the RNA world have garnered significant attention, the focus on protocells provides a lens through which to understand the specific transitions that must have occurred for inanimate molecules to give rise to living, self-replicating systems. This article aims to consider two highly significant facets of protocell evolution: chiral symmetry breaking and the increase in informational complexity passed down through generations. Both aspects are crucially influenced by thermodynamics, specifically by the availability of low entropy energy inflow (e.g., energy from the Sun) and the ability to dissipate extra entropy.
Chiral molecules are those that exist in non-superimposable mirror image forms, known as enantiomers. In modern biology, only one type of organic enantiomer—either the “left-handed” (L) or the “right-handed” (D) version—is overwhelmingly favored for each class of biomolecules like amino acids and sugars. This phenomenon, known as “homochirality” presents an intriguing puzzle: how did the early Earth, presumably awash in a racemic mixture of both L- and D-enantiomers, give rise to life that prefers one type over the other? The breaking of chiral symmetry, wherein one enantiomer is favored over its mirror image, is thought to be a crucial milestone in the transition from chemistry to biology1–13.
Related to the question of chiral symmetry breaking is the requirement for an increase in informational complexity. For life to evolve, each successive generation of protocells must not only replicate but also adapt to its environment and optimize its performance to persist. Thus, the process would involve an increased amount of information being passed from one generation to the next, allowing for the emergence of more complex structures and functions14–16. This increase in inherited complexity serves as a cornerstone for Darwinian evolution and eventually leads to the complex web of life as we know it today17.
However, neither chiral symmetry breaking nor the increase in inherited complexity can occur in a thermodynamic vacuum. Life is an inherently non-equilibrium process, requiring a constant influx of low entropy energy to maintain its ordered state14,18,19. This energy source, typically in the form of sunlight for modern biology, is crucial for driving the endothermic reactions required for life. Moreover, the ability of a system to dissipate the extra entropy it generates – by exporting it to its surroundings – is a defining characteristic of living systems and likely played a vital role in the emergence and evolution of protocells14,20–22.
This article aims to expand upon the foundational work in protocell evolution, particularly focusing on chiral symmetry breaking and increasing informational complexity as pivotal milestones. Drawing inspiration from the one-dimensional model considered by23, which considered the distribution of protocells based on enantiomeric excess, we extend this model into a two-dimensional framework. The extended model not only accounts for distribution by enantiomeric excess but also incorporates another crucial dimension—the amount of information inherited from one generation to the next.
The move to a two-dimensional model aims to encapsulate a more comprehensive picture of the evolutionary dynamics at play in early life forms. By considering both the chiral composition and inherited information, we hope to explore the interplay between these two essential facets and their combined influence on protocell survival and evolution. This dual consideration is consistent with the broader understanding that these processes are governed by thermodynamic factors, specifically the need for low entropy energy sources and mechanisms for effective dissipation of extra entropy21,22.
While previous models have often focused on either the chiral aspects7,13 or information complexity15,16, rarely have both dimensions been considered simultaneously. This article aims to bridge that gap and offer a more synthesized view of how these critical factors co-evolved in shaping the first rudimentary biological systems on Earth.
It is also known that modern life utilizes various error correction mechanisms24, which allow correcting replication errors. In this work we do not want to take that into account, as that ability requires some advanced cellular machinery, which could not exist before life.
Evolution of protocells in two-dimensional space
The model considered here is essentially the model considered by23 extended further into two-dimensional space of chiral composition (total enantiomeric excess of the protocell) and inherited information (the amount of information passed from generation to generation). Apart from finding the stationary points, we also wanted to solve the dynamic evolution problem because it is expected to be much more complex and richer than in one-dimensional space.
As the number of organic molecules on prebiotic Earth was very large, we first start from a continuous representation of chemical systems, where the evolution of the system can be described using differential (or in this case integrodifferential) equations. This allows us to use differential and integral machinery, which, in turn allows us to make some interesting conclusions without solving the equations. However, once we get to a time evolution, the continuous representation breaks down and results in some non-existent artefacts. We will consider that in detail below when we talk about solution method.
The exact chemical reactions, which occurred in protocells on a prebiotic Earth are not known yet, and therefore we would like to concentrate on mathematical aspects of protocell evolution, rather than to speculate on what chemical reactions could have been a driving force at that time. One of the converging ideas is that the transition from non-living matter to living matter is largely due to so called pre-biological compartmentalization25, of which the transition from zero to just one compartment is the first crucial step. It is those, as they are called in25, “self-propagating, chemically simple compartmentalized mainly organic systems” that we are after in this research but without trying to get into a discussion about the unknown chemical reactions of that time. From that point of view, a protocell, which we would like to consider here, is a simple, single-compartment unit with at least some reactions inside a compartment driven by an external energy source. This single-compartment protocell requires neither a symmetry breaking to occur first, nor an advanced replication machinery as we have in modern life. Rather, a set of coupled reactions, which under the energy inflow, could increase the amounts of its reagents is what’s sufficient to start the process. A relatively simple experiment showcasing such compartmentalization was performed back in 196326 and then repeated recently27.
Subsequently, we would like to consider a protocell as a black box, which consumes some “food” and energy out of the environment and creates another protocell. As protocells are not alive yet, we also would like to ignore the consumption of food to support a “life” of a protocell, because there is no life yet. A protocell is a relatively complex structure consisting of many molecules and so it needs more than one molecule of food to replicate. Nevertheless, it is convenient to express the models in protocells, rather than in the number of food molecules. The rate of collision between a protocell and a “molecule” of food is proportional to the concentration of protocells and a concentration of food. And food molecules are consumed one by one, not all at a time. Therefore, the reaction rate should be proportional to the first power of concentration of food and the first power of concentration of protocell. And the fact that many molecules of food are needed to replicate a protocell simply results in a replication time proportional to the number of needed molecules of food. That new protocell may have some small random mutations in comparison to the original protocell. This can be expressed as:
 |
1 |
where  is an amount of food “molecules”,
is an amount of food “molecules”,  is original protocell,
is original protocell,  is a new protocell. We shall stress that
is a new protocell. We shall stress that  here is a concentration of food molecules expressed in protocells, that is the actual concentration of food molecules divided by a number of food molecules needed to replicate a protocell, say
here is a concentration of food molecules expressed in protocells, that is the actual concentration of food molecules divided by a number of food molecules needed to replicate a protocell, say  . We will not use this parameter
. We will not use this parameter  in any of the calculations. It also seems interesting to consider a nonlinear scenario where some
in any of the calculations. It also seems interesting to consider a nonlinear scenario where some  molecules of food are simultaneously needed to create a protocell:
molecules of food are simultaneously needed to create a protocell:
 |
2 |
We have run a few simulations using this equation (and with relevant changes to all other equations) and the only noticeable changes were to the first stage of evolution of the system. Larger than one values of  resulted in faster evolution time of that stage. We will talk about that phenomenon in more detail below. Subsequently, we have used
resulted in faster evolution time of that stage. We will talk about that phenomenon in more detail below. Subsequently, we have used  for all further calculations due to increased numerical stability and smaller calculation time in comparison to larger values of
for all further calculations due to increased numerical stability and smaller calculation time in comparison to larger values of  .
.
We further consider that the protocells can “die off” without any mutual inhibition:
 |
3 |
where  is some waste “molecule” and by waste “molecule” we mean that we also express waste in protocells, the same as food.
is some waste “molecule” and by waste “molecule” we mean that we also express waste in protocells, the same as food.
Finally, we need to close the model, and this is achieved by recycling the waste.
 |
4 |
The alternative is to introduce a pass-through model, where the food flows in at some constant rate and the waste is discarded. It can be shown that these models are dynamically nearly equivalent up to some time-dependent transformations. However, pass-through models don’t have a well-defined integral of motion, which makes them harder to model and account for errors. In addition, as the total amount of matter in a pass-through model linearly increases over time that makes such models less stable than the models with recycling. In other words, a pass-through model is substantially more likely to blow up numerically if some coefficients of the model are not chosen correctly.
Given the equations above, the changes in protocell concentration  can be written as:
can be written as:
 |
5 |
where  is a concentration of protocells with total enantioselectivity
is a concentration of protocells with total enantioselectivity  and the amount of stored information
and the amount of stored information  at time
at time  ,
,  is the concentration of food molecules at time
is the concentration of food molecules at time  ,
,  is the rate at which protocells
is the rate at which protocells  can produce protocells
can produce protocells  ,
,  is a normalization constant, so that
is a normalization constant, so that  , and
, and  is the decay rate of protocells
is the decay rate of protocells  . The range of enantioselectivity is naturally the interval
. The range of enantioselectivity is naturally the interval  with the initial value located near
with the initial value located near  (nearly racemic mixture). The range in the information space depends on how we define the amount of information passed from generation to generation and what we mean as that information. However, we can always change the variables so that the domain in
(nearly racemic mixture). The range in the information space depends on how we define the amount of information passed from generation to generation and what we mean as that information. However, we can always change the variables so that the domain in  starts from 0 and we keep the upper boundary as some value
starts from 0 and we keep the upper boundary as some value  . We can further rescale
. We can further rescale  so that to make that
so that to make that  any value we want, e.g., 1, but we find it more convenient to keep it without rescaling in case we’d want to compare some models with different values of
any value we want, e.g., 1, but we find it more convenient to keep it without rescaling in case we’d want to compare some models with different values of  in the future. Subsequently, that makes the initial state of protocells as some small narrow peak near
in the future. Subsequently, that makes the initial state of protocells as some small narrow peak near  in
in  space.
space.
The changes in  and
and  can be written as:
can be written as:
 |
6 |
where  is the concentration of waste “molecules” at time
is the concentration of waste “molecules” at time  and
and
 |
7 |
where  is a recycling rate.
is a recycling rate.
We can first look at the stationary point of these equations. Equation (5) leads to the following condition:
 |
8 |
where  ,
,  and
and  is
is  normalized so that:
normalized so that:
 |
9 |
This is a two-dimensional Fredholm integral operator of the first kind for the kernel  . If
. If  and
and  subspaces are separable:
subspaces are separable:  then we can express
then we can express  and then Eq. (8) also splits into one-dimensional equations.
and then Eq. (8) also splits into one-dimensional equations.
Kernel normalization
Before we proceed further, it is convenient to perform some transformations of the original kernel  . First, we can extract a normalization constant
. First, we can extract a normalization constant  :
:
 |
10 |
which is a total production rate at a point  , and a total normalized replication rate
, and a total normalized replication rate  :
:
 |
11 |
Then we can rewrite the kernel as:
 |
12 |
where  is defined as:
is defined as:
 |
13 |
from which it follows that:
 |
14 |
which means that  is the probability that species with enantiomeric excess
is the probability that species with enantiomeric excess  and amount of stored information
and amount of stored information  would produce species with total enantiomeric excess
would produce species with total enantiomeric excess  and amount of stored information
and amount of stored information  . As mutations are small, that probability should be a narrow peak centered near the point
. As mutations are small, that probability should be a narrow peak centered near the point  .
.
Assuming that mutations in  and
and  spaces are independent and taking into account that the number of molecules and protocells was very large, allows us to utilize Central Limit Theorem. That means that we can use normal distributions to model
spaces are independent and taking into account that the number of molecules and protocells was very large, allows us to utilize Central Limit Theorem. That means that we can use normal distributions to model  as some narrow peak near
as some narrow peak near  :
:
 |
15 |
where  ,
,  are some small parameters (in general functions of
are some small parameters (in general functions of  and
and  ) defining mutation rates,
) defining mutation rates,  is the error function, and the normalization coefficient follows from normalization condition Eq. (14). This probability is nearly identical to the two-dimensional normal probability distribution inside the domain:
is the error function, and the normalization coefficient follows from normalization condition Eq. (14). This probability is nearly identical to the two-dimensional normal probability distribution inside the domain:  and has some bias at the edges of the full domain
and has some bias at the edges of the full domain  . This bias is irrelevant because the starting point
. This bias is irrelevant because the starting point  is in the middle of the domain in
is in the middle of the domain in  space and as we are interested in the system moving away from
space and as we are interested in the system moving away from  some irregularities near that point at the beginning of the time evolution are insignificant.
some irregularities near that point at the beginning of the time evolution are insignificant.
After all these transformations, the Eq. (8) becomes:
 |
16 |
where  . We can estimate the largest eigenvalue and approximate location of its eigenvector under reasonable assumptions that the mutations are small. Consider that the first eigenvector is a narrow “bump” function of some height
. We can estimate the largest eigenvalue and approximate location of its eigenvector under reasonable assumptions that the mutations are small. Consider that the first eigenvector is a narrow “bump” function of some height  with the widths (standard deviations)
with the widths (standard deviations)  and
and  and it is centered near point
and it is centered near point  Then, we can integrate Eq. (16) by
Then, we can integrate Eq. (16) by  and approximate it. Then:
and approximate it. Then:
 |
17 |
where  is some numerical factor, which depends on the actual shape of
is some numerical factor, which depends on the actual shape of  and:
and:
 |
18 |
where the first approximate transition can be made because  is non-zero when
is non-zero when  is close to
is close to  and
and  is close to
is close to  (the mutations are considered small) and
(the mutations are considered small) and  is a slow function in the range where
is a slow function in the range where  is non-zero (the decay rate for new protocells is not very different from the one of the original protocells), and the second approximation follows from the fact that we considered
is non-zero (the decay rate for new protocells is not very different from the one of the original protocells), and the second approximation follows from the fact that we considered  as a narrow peak near
as a narrow peak near  . This leads to an estimate:
. This leads to an estimate:
 |
19 |
And since we are interested in the largest eigenvalue, that means that the point  is where the function:
is where the function:
 |
20 |
has a global maximum on the domain:  .
.
Diffusion
As the mutations are considered small, then  is a narrow peak centered around the point
is a narrow peak centered around the point  . In this case we can perform a Taylor expansion of
. In this case we can perform a Taylor expansion of  near
near  when
when  is substantially wider than
is substantially wider than  :
:
 |
21 |
and substituting into Eq. (5) we can obtain:
 |
22 |
where:
 |
23 |
This means that Eq. (5) can be interpreted as time dependent convection-diffusion equation with a sink in the scenarios when the mutations are small and shape of  is substantially wider than the width of mutation probability, though the form is slightly different from what is usually considered as convection-diffusion equation. It is important to note that both conditions: small mutation rate and abundance of species (which is a manifestation of the fact that
is substantially wider than the width of mutation probability, though the form is slightly different from what is usually considered as convection-diffusion equation. It is important to note that both conditions: small mutation rate and abundance of species (which is a manifestation of the fact that  is substantially wider than mutation probability) seem to hold in life and in the considered model for all values of time except some small amount of time at the beginning of the evolution of the system. The term
is substantially wider than mutation probability) seem to hold in life and in the considered model for all values of time except some small amount of time at the beginning of the evolution of the system. The term  is responsible for creation
is responsible for creation  or destruction
or destruction  of protocells, vector
of protocells, vector  is the “speed” at which the species drift away, and matrix
is the “speed” at which the species drift away, and matrix  determines the “diffusion”. The drift is the most interesting factor here. We can perform a Taylor expansion of
determines the “diffusion”. The drift is the most interesting factor here. We can perform a Taylor expansion of  near the point
near the point  :
:
 |
24 |
and then consider that  is a nearly symmetric function near that point
is a nearly symmetric function near that point  inside
inside  . Then:
. Then:
 |
25 |
and where:
 |
26 |
and we ignored higher order terms by  and
and  . That means that the drift is determined by a vector
. That means that the drift is determined by a vector  , which is a local gradient of
, which is a local gradient of  up to some coefficient of proportionality. We also note that
up to some coefficient of proportionality. We also note that  and
and  with a very high precision within
with a very high precision within  .
.
Stages of evolution with time
We note that  must be an even function of
must be an even function of  due to
due to  symmetry and therefore all odd derivatives of
symmetry and therefore all odd derivatives of  by
by  at the point
at the point  must be zeros. Therefore, second order Taylor expansion of
must be zeros. Therefore, second order Taylor expansion of  near point
near point  should not have terms linear in
should not have terms linear in  :
:
 |
27 |
or, if we want to keep  in a separable form then:
in a separable form then:
 |
28 |
The latter form has some extra higher than level 2 terms in comparison to the Taylor expansion. This only affects the evolution in diagonal directions and only when cross terms become substantial. If both quadratic terms are positive (which is the case in life, except probably near some catastrophic events, which we are not considering here), then value of  in separable form is larger in diagonal directions than when using the Taylor expansion. However, we have not seen the system going in diagonal directions even though we used a separable version of
in separable form is larger in diagonal directions than when using the Taylor expansion. However, we have not seen the system going in diagonal directions even though we used a separable version of  in most of our calculations.
in most of our calculations.
Assuming that the mutations are small, we can consider various stages of evolution of the system. The initial state of the system is a very narrow peak near point  in
in  space. If we replace
space. If we replace  where
where  is the Dirac delta function, then we can perform integrations in Eqs. (5–7) to obtain:
is the Dirac delta function, then we can perform integrations in Eqs. (5–7) to obtain:
 |
29 |
 |
30 |
 |
31 |
Provided that initially the food is abundant, and the initial number of protocells is very small, the first stage of evolution is an exponential growth of the total number of protocells with the initial growth rate:  . The time interval of that period is when the initial number of protocells:
. The time interval of that period is when the initial number of protocells:  would exponentially grow to the order of
would exponentially grow to the order of  :
:
 |
32 |
from which it follows that:
 |
33 |
When the food is abundant (as it is in the initial stage), then  and so we can ignore
and so we can ignore  in the denominator. Taking just one initial protocell and the total number of organic molecules or even all atoms on Earth as the hard limit gives an estimate that the logarithm in numerator is naturally limited to somewhere between
in the denominator. Taking just one initial protocell and the total number of organic molecules or even all atoms on Earth as the hard limit gives an estimate that the logarithm in numerator is naturally limited to somewhere between  . The actual value is irrelevant as the process is very quick. The value
. The actual value is irrelevant as the process is very quick. The value  is essentially a doubling rate (to be more precise the time to increase the population in
is essentially a doubling rate (to be more precise the time to increase the population in  times) and so using even the upper (unrealistic) boundary of the nominator as
times) and so using even the upper (unrealistic) boundary of the nominator as  and unrealistically slow doubling rate of protocells of, say, one year gives
and unrealistically slow doubling rate of protocells of, say, one year gives  years, which is negligible on a geological time scale. This time period,
years, which is negligible on a geological time scale. This time period,  is the amount of time, which would take the most inefficient protocells (located near point
is the amount of time, which would take the most inefficient protocells (located near point  in
in  space) to consume nearly all food. If we look at Eq. (22), then
space) to consume nearly all food. If we look at Eq. (22), then  can be interpreted as the period of time during which the value
can be interpreted as the period of time during which the value  reaches 0 at point
reaches 0 at point  :
:
 |
34 |
Once this point in time is reached, then further evolution of the system is only possible by increasing the efficiency of protocells. The less efficient protocells then will die off whereas the more efficient protocells will continue to evolve thus making it possible to sustain their replication at smaller and smaller concentrations of food. That means that the boundary:
 |
35 |
should move away from point  as the system evolves.
as the system evolves.
The second stage of evolution substantially depends on the values of the drift vector,  . The value
. The value  due to symmetry, however
due to symmetry, however  does not have to be zero. If it is zero or close to zero, then the second stage of evolution is a diffusion in
does not have to be zero. If it is zero or close to zero, then the second stage of evolution is a diffusion in  space until the time when quadratic coefficients in
space until the time when quadratic coefficients in  start to play a role. However, if
start to play a role. However, if  is large enough, then the initial bump near
is large enough, then the initial bump near  will move in
will move in  space faster than it diffuses. Biologically this means that in the first case the system produces a variety of species that are diverse both in total enantiomeric excess and the amount of stored information, whereas in the second case the species form a compact “bump” of not yet separated into left or right species, which quickly advance in the ability to pass information from generation to generation.
space faster than it diffuses. Biologically this means that in the first case the system produces a variety of species that are diverse both in total enantiomeric excess and the amount of stored information, whereas in the second case the species form a compact “bump” of not yet separated into left or right species, which quickly advance in the ability to pass information from generation to generation.
The third stage of evolution starts when quadratic coefficients in Eq. (22) become substantial. There is an extensive discussion in23 regarding how replication with errors helps to transition replicated structures toward homochirality. And while it is intuitively clear that passing more information from generation to generation is beneficial for the evolution of life, we will not dwell in theoretical considerations of this matter in the current work. Rather, we note, that if any of the quadratic coefficients were negative, then the diffusion in that direction would have been suppressed and subsequently life, as we know it, would not exist. Therefore, both quadratic coefficients must be positive. The time evolution in that stage is complicated enough so that it is not possible to deduce how the system should evolve without actually solving the dynamical problem. Before we get to that, we shall note the following. The drift vector,  is proportional to the gradient of
is proportional to the gradient of  at a given point. Therefore, the further is the point from
at a given point. Therefore, the further is the point from  in any direction, the larger is that gradient and therefore the faster the protocells that are at that point move in the direction of that gradient because
in any direction, the larger is that gradient and therefore the faster the protocells that are at that point move in the direction of that gradient because  has positive quadratic coefficients in the Taylor expansion in both directions. If we note that the evolution equation resembles the convection-diffusion equation, then we can talk about the speed at which the width of the initial spike increases. And it is a well-known fact that a standard diffusion equation with constant diffusion coefficient has a Gaussian solution with the width
has positive quadratic coefficients in the Taylor expansion in both directions. If we note that the evolution equation resembles the convection-diffusion equation, then we can talk about the speed at which the width of the initial spike increases. And it is a well-known fact that a standard diffusion equation with constant diffusion coefficient has a Gaussian solution with the width  where
where  is the diffusion coefficient and
is the diffusion coefficient and  is evolution time. Therefore the “speed” at which the edge of the bump moves is
is evolution time. Therefore the “speed” at which the edge of the bump moves is  (as a derivative of the width by time). This can be estimated in the region where quadratic coefficients give substantial contribution to
(as a derivative of the width by time). This can be estimated in the region where quadratic coefficients give substantial contribution to  (for example for large enough values of
(for example for large enough values of  ) as
) as  whereas
whereas  . Another critical fact comes from stochasticity related to chemical evolution. A stochasticity in relation to the current model means that there is a probability that several favorable mutations happen in a row, and which would place some protocells in the region with higher amplification and subsequently drift speed. When that happens and because the speed of the edge of the bump decreases with time, then it could be possible for the protocells with favorable mutations to get separated from the rest of the pack. That’s the scenario that we are the most interested in when solving dynamical evolution problem.
. Another critical fact comes from stochasticity related to chemical evolution. A stochasticity in relation to the current model means that there is a probability that several favorable mutations happen in a row, and which would place some protocells in the region with higher amplification and subsequently drift speed. When that happens and because the speed of the edge of the bump decreases with time, then it could be possible for the protocells with favorable mutations to get separated from the rest of the pack. That’s the scenario that we are the most interested in when solving dynamical evolution problem.
Solution of dynamical problems
The continuous representation is well-suitable for finding the stationary points of the considered system and performing some simplified analysis. However, it breaks down when we want to consider a dynamical evolution, that is, how does the system get to that stationary point from the initial state? This is mostly due to machine noise. As the number of calculations on each step is very large, the machine noise produces some very small yet non-zero values of  far from the initial state near
far from the initial state near  . That machine noise is also likely to appear in the regions of
. That machine noise is also likely to appear in the regions of  with large values and subsequently with larger “amplification”. As a result, the machine noise will experience exponential growth and that results in some artefacts appearing where they are not supposed to be.
with large values and subsequently with larger “amplification”. As a result, the machine noise will experience exponential growth and that results in some artefacts appearing where they are not supposed to be.
As chemical systems evolve in integer numbers rather than in real numbers, we could try using some version of chemical master equation to calculate the evolution of the system in the number of molecules. Then very small real numbers cannot appear in the regions where they are not supposed to be and so the issue of unrealistic artefacts is removed. Since we want to use a large number of molecules and protocells in our calculations, Gillespie tau-leaping algorithm28 is well suitable for such a problem. We have used a custom version of this algorithm to ensure stability near zero values. It is interesting to note that the Gillespie tau-leaping algorithm is essentially a stochastic version of a simple Euler algorithm for solving differential equations. In fact, it is straightforward to show that as the number of molecules goes to infinity, the Gillespie tau-leaping algorithm converges to the Euler algorithm. This means that the Gillespie tau-leaping algorithm should be plagued by the same issues as the Euler algorithm. In particular, it means that the Gillespie tau-leaping algorithm should undershoot for exponential growth and overshoot for exponential decay, of which the exponential decay is the most dangerous. The algorithm can easily overshoot below zero, after which all the calculations break down. This is why chemical equations are considered stiff. When solving differential equations this is resolved either by substantially reducing the time step, thus drastically increasing the solution time, or using higher-order direct methods, or using indirect methods, which require inverting matrices. Apart from the need to adapt indirect methods for the Gillespie tau-leaping algorithm, this is simply not suitable for the current problem because, for example, a 500 × 500 grid has 250,000 variables and using indirect method would require inverting 250,000 × 250,000 matrices, where machine noise, calculation time, and memory requirements would make it prohibitive. Using higher order direct methods with the tau-leaping algorithm would also require adaptation.
The solution to this problem in case of chemical equations is simple. We let the system overshoot into negative values but treat all negative values as exact zeros in subsequent calculations. This is the approach that we used in some previous works, and it showed results consistent with the ones obtained using more advanced but slower algorithms. Out of all the variables in the system considered in the current work it is food that could easily become negative at some steps, especially if some of the parameters are large enough. Even though the approach described above works even in such extreme cases, we fine-tune the algorithm further by adjusting the step so that to consume all food but no more than is available on a given step. The solution is stable and since the system has an invariant of motion, we can monitor that it is conserved exactly at each step.
Results
We used the following separable approximations of  and
and  in the current work:
in the current work:
 |
36 |
 |
37 |
 |
38 |
Parameter  is a small linear global asymmetry factor and our view is that it should be introduced in the decay rate
is a small linear global asymmetry factor and our view is that it should be introduced in the decay rate  rather than in replication rate
rather than in replication rate  because the global asymmetry factor influences the stability of the left vs. right amino acids and sugars, which subsequently affects the lifespan of protocells. Parameter
because the global asymmetry factor influences the stability of the left vs. right amino acids and sugars, which subsequently affects the lifespan of protocells. Parameter  is an artificially introduced limiting factor to keep the stationary point off the boundary
is an artificially introduced limiting factor to keep the stationary point off the boundary  . If this factor is not introduced, then the stationary point in
. If this factor is not introduced, then the stationary point in  space becomes near point
space becomes near point  . Another view is that any model should have limitations, and we wanted to limit how far our model could evolve. Parameter
. Another view is that any model should have limitations, and we wanted to limit how far our model could evolve. Parameter  is a scaling coefficient, which we introduced for convenience.
is a scaling coefficient, which we introduced for convenience.
One of the most important parameters in the Gillespie tau-leaping algorithm is the step size when all other parameters are fixed. However, we can always rescale the parameters so that to set step size to some desired value. We used fixed step size  and then set other parameters as necessary. This allows us to treat
and then set other parameters as necessary. This allows us to treat  as evolution epochs and it makes it easier to interpret the results, e.g., if
as evolution epochs and it makes it easier to interpret the results, e.g., if  day, then
day, then  means that average protocell lifespan is:
means that average protocell lifespan is:  days.
days.
We used the following parameter values in calculations:
| Parameter | Value | Description |
|---|---|---|

|
500 | Size of the grid used in calculations. We used a  grid. grid. |

|
25 | Upper boundary of the domain in  space. Currently the choice is arbitrary as we can always rescale the space. Currently the choice is arbitrary as we can always rescale the  variable. However, if we consider it as the natural logarithm of bits of information passed from generation to generation, then that value translates into variable. However, if we consider it as the natural logarithm of bits of information passed from generation to generation, then that value translates into  bytes, which by far exceeds reasonable estimates of how much information the protocells could have passed from generation to generation. bytes, which by far exceeds reasonable estimates of how much information the protocells could have passed from generation to generation. |

|

|
Scaling coefficient to make it more convenient to set values of parameters in  dimension. This choice of dimension. This choice of  was determined by our desire to scale was determined by our desire to scale  into 1 in scaled coordinate into 1 in scaled coordinate  . . |

|
0.1 | The parameter  , as determined by Eq. (10), can be used as is when using ODE evolution. The value of , as determined by Eq. (10), can be used as is when using ODE evolution. The value of  is used instead when using Gillespie tau-leaping evolution in integer numbers. This adjustment is necessary because the term with is used instead when using Gillespie tau-leaping evolution in integer numbers. This adjustment is necessary because the term with  in the original differential Eq. (5) is in the original differential Eq. (5) is  and therefore and therefore  has a dimension has a dimension  . Therefore, we need to account for that when transitioning to the equations expressed in the number of molecules. . Therefore, we need to account for that when transitioning to the equations expressed in the number of molecules. |

|
0.01 | Decay rate of protocells at point  . . |

|
1 | Recycling rate. The value of that parameter determines how much residual waste is near the stationary point. |

|
1 | The value of  at point at point  . . |

|
 or or 
|
The value of  at point at point  . . |

|
1 | The value of  at point at point  . . |

|
0.005 or 0.01 | Mutation rate in  space. space. |

|
0.005 or 0.01 | Mutation rate in  space. space. |

|

|
Threshold parameter to control the values of probability Eq. (15) to treat as exact zero. When  we treat it as exact zero and remove it from the kernel function (but adjust total probability to sum up to 1). This allows us to consider a four-dimensional kernel as a sparse array resulting in a very substantial speed increase of the algorithm. As typical number of such non-zero points in we treat it as exact zero and remove it from the kernel function (but adjust total probability to sum up to 1). This allows us to consider a four-dimensional kernel as a sparse array resulting in a very substantial speed increase of the algorithm. As typical number of such non-zero points in  is somewhere between 10 to 100 instead of is somewhere between 10 to 100 instead of  , which results in a 1,000- to 10,000-fold speed increase of the algorithm. , which results in a 1,000- to 10,000-fold speed increase of the algorithm. |

|
 or or 
|
Global asymmetry factor. The values smaller than  are beyond the resolution of the grid used in our calculations. are beyond the resolution of the grid used in our calculations. |

|
1000 | This is an artificial parameter to keep the stationary point off the upper boundary in  space and model the scenario when further increase of stored information results in faster decay rate. If this parameter is not introduced, the system will run toward the space and model the scenario when further increase of stored information results in faster decay rate. If this parameter is not introduced, the system will run toward the  edge of the domain, which produces harder to visualize results. edge of the domain, which produces harder to visualize results. |

|

|
Total number of molecules in the system. This is an invariant, and this number does not change over the course of evolution of the system. The bigger is  the more the dynamical evolution starts to resemble evolution of the system in real numbers. the more the dynamical evolution starts to resemble evolution of the system in real numbers. |

|

|
Initial number of protocells located at point  at a time at a time  . . |
The following scenarios were chosen for this article:  ,
,  ,
,  ,
,  ,
,  or 1,
or 1,  or
or  and we summarized them in the Table 1.
and we summarized them in the Table 1.
Table 1.
Parameters of the models considered in the current work.
| No | Model parameters | Model code |
|---|---|---|
| 1 (a) |
 , ,  , ,  , ,  , ,  , , 
|
d500k1e005g01a002f1E |
| 2 (b) |
 , ,  , ,  , ,  , ,  , , 
|
d500k1e01g01a002f1E |
| 3 (c) |
 , ,  , ,  , ,  , ,  , , 
|
d500k1e005g01a002i10f1E |
| 4 (d) |
 , ,  , ,  , ,  , ,  , , 
|
d500k1e01g01a002i10f1E |
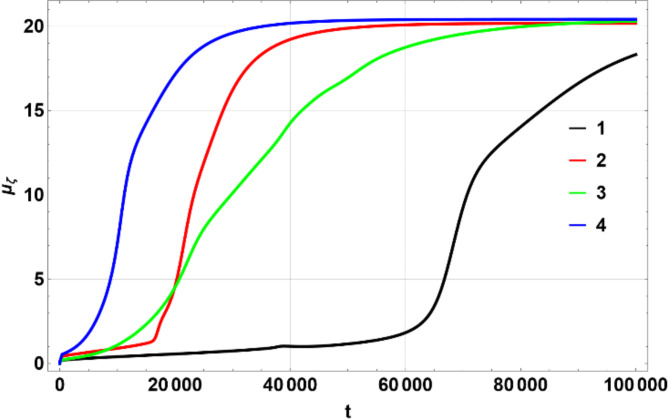
The colors matching a number in the table above were used on multi-line 2D figures and 3D figures use (a) – (d) marking. 2D charts show means and widths (standard deviation) of  over time of which the mean in
over time of which the mean in  space,
space,  (Fig. 1) and standard deviation
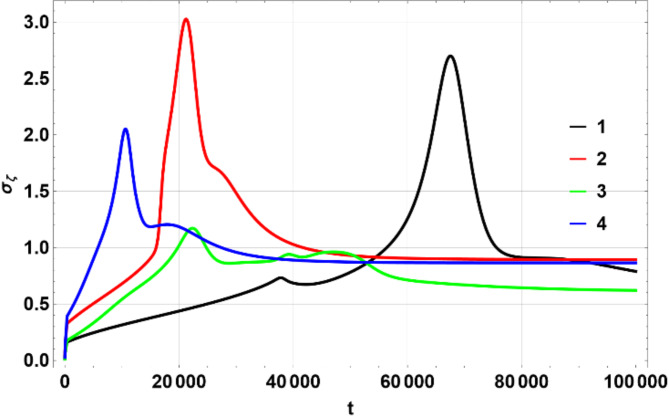
(Fig. 1) and standard deviation  , Fig. 2 are the most important. The 3D figures (distributions of
, Fig. 2 are the most important. The 3D figures (distributions of  ) are shown for some chosen values of time to illustrate the most interesting effects. All values of
) are shown for some chosen values of time to illustrate the most interesting effects. All values of  on 3D figures are in percent.
on 3D figures are in percent.
Fig. 1.
Dependence of  on
on  . Produced using Wolfram Mathematica 13.
. Produced using Wolfram Mathematica 13.
Fig. 2.
Dependence of  on
on  . Produced using Wolfram Mathematica 13.
. Produced using Wolfram Mathematica 13.
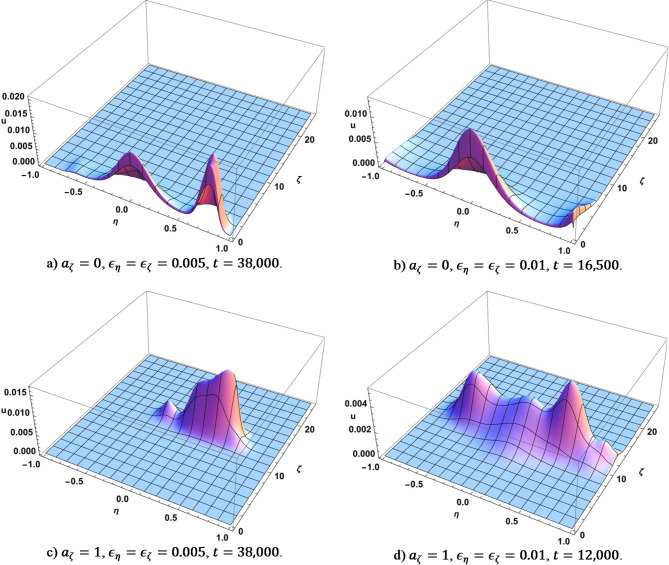
The first two stages of evolution (very quick exponential growth of the initial narrow bump followed by a diffusion-like spreading) were consistently observed among all runs that we performed. Figure 3 shows  for some chosen values of time when the second stage of evolution has not completed yet. The time is
for some chosen values of time when the second stage of evolution has not completed yet. The time is  for the case when
for the case when  and
and  when
when  . The width of Gaussian solution of the diffusion equation with constant diffusion coefficient grows
. The width of Gaussian solution of the diffusion equation with constant diffusion coefficient grows  (
( is either
is either  or
or  ). The widths of the bumps for
). The widths of the bumps for  vs.
vs.  are nearly the same on the Figure 3 even though the evolution time is 4x faster, which is consistent with the estimate. There is a drift toward higher information content when
are nearly the same on the Figure 3 even though the evolution time is 4x faster, which is consistent with the estimate. There is a drift toward higher information content when  . The widths of the bumps are approximately the same as in the case of no drift, though the drift is more significant in case when
. The widths of the bumps are approximately the same as in the case of no drift, though the drift is more significant in case when  than when
than when  . This is expected as the run time is 4x longer in the first case.
. This is expected as the run time is 4x longer in the first case.
Fig. 3.
Dependence of  on
on  and
and  when the second stage of evolution (diffusion) has not completed yet. Produced using Wolfram Mathematica 13.
when the second stage of evolution (diffusion) has not completed yet. Produced using Wolfram Mathematica 13.
Figure 4 shows some points in time for the four considered scenarios where each of the corresponding diffusion-like evolution runs break down and experience rapid transformation. We will talk about that in more detail when we discuss Fig. 2. Here we would like to note that the process is essentially random and using different random seed values will produce different intermediate shapes until the process settles down near stationary points.
Fig. 4.
Dependence of  on
on  and
and  for some values of time when the system experiences rapid transformations. Produced using Wolfram Mathematica 13.
for some values of time when the system experiences rapid transformations. Produced using Wolfram Mathematica 13.
Figure 5 shows the point in time ( ) when the stationary point has been reached. The stationary point looks nearly the same for all considered variants: it is a narrow peak near
) when the stationary point has been reached. The stationary point looks nearly the same for all considered variants: it is a narrow peak near  and
and  , which is a point in
, which is a point in  space where
space where  experiences a rapid increase due to a limiting factor
experiences a rapid increase due to a limiting factor  . Without that limiting factor the system would just run toward
. Without that limiting factor the system would just run toward  . The stationary point for
. The stationary point for  is a narrower peak than for
is a narrower peak than for  , as expected, and the value of
, as expected, and the value of  does not affect the position and the width of the peak.
does not affect the position and the width of the peak.
Fig. 5.
Dependence of  on
on  and
and  near stationary point (
near stationary point ( ). Produced using Wolfram Mathematica 13.
). Produced using Wolfram Mathematica 13.
We present animation of the evolution of the system for these 4 scenarios in supplementary materials.
We also want to look at some cumulative characteristics of the evolution because it is hard to present changes to a 3D chart over time in a compact form. Means and standard deviations of  in
in  and
and  spaces are a convenient choice. In integral representation they can be expressed as follows (note that the normalization here is different in comparison to Eq. (9)):
spaces are a convenient choice. In integral representation they can be expressed as follows (note that the normalization here is different in comparison to Eq. (9)):
 |
39 |
Figures 1, 2 and 6, and 7 show  ,
,  ,
,  ,
,  for the four scenarios considered above. All figures have lines 1 through 4 corresponding to the parameters shown in the Table 1 above.
for the four scenarios considered above. All figures have lines 1 through 4 corresponding to the parameters shown in the Table 1 above.
Fig. 6.
Dependence of  on
on  . Produced using Wolfram Mathematica 13.
. Produced using Wolfram Mathematica 13.
Fig. 7.
Dependence of  on
on  . Produced using Wolfram Mathematica 13.
. Produced using Wolfram Mathematica 13.
The results presented above show that a two-dimensional model is much more complex and richer than a one-dimensional one23. In particular, as food becomes scarce, the boundary between efficient and inefficient protocells is just two points in one-dimensional model. And so, these points are not connected. That boundary becomes a connected curve in the two-dimensional model. Those protocells that are inside the boundary “die off” from starvation because they are not efficient enough to sustain their population at a given concentration of food. And the boundary extends from  over time due to diffusion and appearance of more efficient species. The most interesting part here is that many species are possible. Second, random fluctuations may create several favorable mutations in a row. A favorable mutation is a mutation that makes some species of protocells more efficient. That slight increase in efficiency results in placing some small number of protocells into the place in
over time due to diffusion and appearance of more efficient species. The most interesting part here is that many species are possible. Second, random fluctuations may create several favorable mutations in a row. A favorable mutation is a mutation that makes some species of protocells more efficient. That slight increase in efficiency results in placing some small number of protocells into the place in  space where they have a positive exponential growth factor in comparison to the boundary where the bulk of the protocells exist at some moment in time. Subsequently these more efficient protocells can temporarily experience exponential growth. This may result in splitting of the original “bump” distribution of protocells into several non-connected groups, which we can call species. The greater the total number of molecules in the system the more likely this is to occur.
space where they have a positive exponential growth factor in comparison to the boundary where the bulk of the protocells exist at some moment in time. Subsequently these more efficient protocells can temporarily experience exponential growth. This may result in splitting of the original “bump” distribution of protocells into several non-connected groups, which we can call species. The greater the total number of molecules in the system the more likely this is to occur.
The easiest way to illustrate four stages of evolution of the system is to look at the concentration of food molecules. As the time scale of the first stage is much faster than that of the other three stages, we need two charts to showcase all four stages.
Figure 8 illustrates the first stage of evolution. It shows an exponential, or more correctly, a hyperbolic tangent-like shape. Models 1 and 2 are indistinguishable from each other during this stage (that’s why model 1’s chart is not visible in the figure). Model 3 is slightly faster, and model 4 is even faster at consuming most of the food. Models 3 and 4 have a drift coefficient, which moves protocells into the region of higher efficiency (resulting in faster food consumption). Model 4 also has a larger diffusion coefficient (larger values of  and
and  ), which further accelerates some protocells into the region of higher efficiency.
), which further accelerates some protocells into the region of higher efficiency.
Fig. 8.
Dependence of the amount of food in percent on  during stage 1. Produced using Wolfram Mathematica 13.
during stage 1. Produced using Wolfram Mathematica 13.
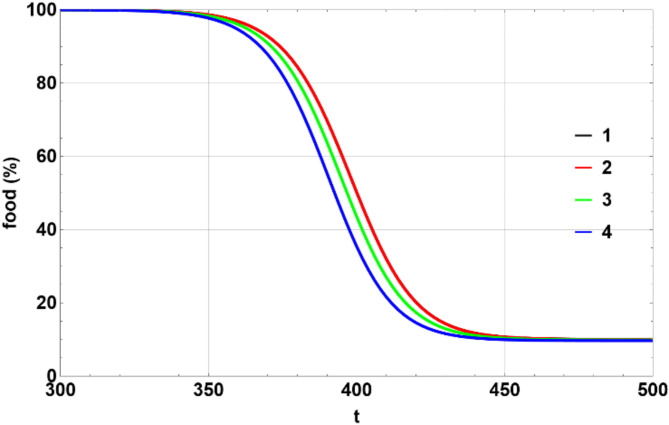
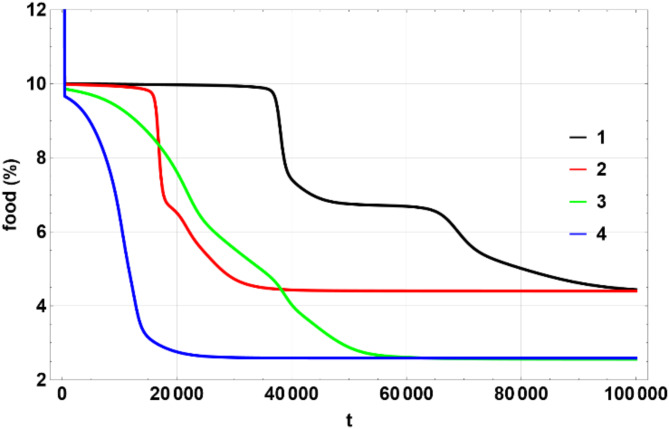
Figure 9 illustrates stages 2–4 of the evolution. Of all the models considered here, only model 1 shows four clear stages of evolution, where stages 2 and 3 appear as temporary plateaus, followed by a sharp transition to the next stage. Model 2, which has a zero-drift coefficient like model 1 but a larger diffusion coefficient, also exhibits a clear stage 2, while stages 3 and 4 are much faster and not clearly visible. Model 3 has a drift coefficient but a smaller diffusion coefficient. The drift coefficient moves the protocells into the region of higher efficiency, making the plateaus almost disappear. Finally, model 4 has both a drift coefficient and a larger diffusion coefficient, combining stages 2–4 into a single continuous slide.
Fig. 9.
Dependence of the amount of food in percent on  during stages 2 – 4. Produced using Wolfram Mathematica 13.
during stages 2 – 4. Produced using Wolfram Mathematica 13.
Conclusions
We consider an evolution of protocells without competition in a two-dimensional space: total enantioselectivity,  , and the amount of information passed from generation to generation,
, and the amount of information passed from generation to generation,  , and we present a simple two-dimensional model, which describes such an evolution. We show that such a model is described by a system of integrodifferential equations, which under the hood is powered by a protocell replication kernel,
, and we present a simple two-dimensional model, which describes such an evolution. We show that such a model is described by a system of integrodifferential equations, which under the hood is powered by a protocell replication kernel,  , and a mortality rate,
, and a mortality rate,  . We show that it is convenient to split a replication kernel into a multiplication of the pieces: a total replication rate at a point
. We show that it is convenient to split a replication kernel into a multiplication of the pieces: a total replication rate at a point  in
in  space,
space,  , a total normalized replication rate,
, a total normalized replication rate,  , which is the normalized rate at which the species with total enantioselectivity
, which is the normalized rate at which the species with total enantioselectivity  and the amount of stored information
and the amount of stored information  can produce any species, and the probability that species with total enantiomeric excess
can produce any species, and the probability that species with total enantiomeric excess  and amount of stored information
and amount of stored information  would produce species with total enantiomeric excess
would produce species with total enantiomeric excess  and amount of stored information
and amount of stored information  ,
,  .
.
We show that observation of life, as it exists, leads to a conclusion that a Taylor expansion of total normalized replication rate  near point
near point  should be a function with zero linear coefficient in
should be a function with zero linear coefficient in  space, non-negative linear coefficient in
space, non-negative linear coefficient in  space, and positive quadratic coefficients both in
space, and positive quadratic coefficients both in  and
and  spaces, provided that we limit the considerations to the terms not higher than quadratic ones.
spaces, provided that we limit the considerations to the terms not higher than quadratic ones.
We show that under reasonable assumptions the mutations from generation to generation of protocells are small (the probability  is a narrow peak centered near
is a narrow peak centered near  and
and  ), the evolution of the system can be described in four stages.
), the evolution of the system can be described in four stages.
The first stage is a very quick exponential growth of the population of the nearly racemic species near the point  in
in  space. Since this is an exponential process, it should happen in a negligible amount time on a geological scale, and we show that the time estimate for that process is on the order of
space. Since this is an exponential process, it should happen in a negligible amount time on a geological scale, and we show that the time estimate for that process is on the order of  years for the closed model considered here.
years for the closed model considered here.
The second stage is a relatively slow diffusion-like process, when the system spreads out in both  and
and  directions. The system at that stage can be very well approximated by a convection-diffusion equation with a drift. If a linear coefficient in the Taylor expansion of total normalized replication rate
directions. The system at that stage can be very well approximated by a convection-diffusion equation with a drift. If a linear coefficient in the Taylor expansion of total normalized replication rate  near point
near point  in
in  space is zero or small enough, then the system experiences just a diffusion (spearing out of the initial delta function like peak), and if that linear coefficient is large enough then that peak travels in
space is zero or small enough, then the system experiences just a diffusion (spearing out of the initial delta function like peak), and if that linear coefficient is large enough then that peak travels in  space while spreading out, thus, gaining information storage capacity able to be passed from generation to generation. The time of that stage depends on the combination of factors: larger mutation rates in
space while spreading out, thus, gaining information storage capacity able to be passed from generation to generation. The time of that stage depends on the combination of factors: larger mutation rates in  and larger quadratic coefficients in
and larger quadratic coefficients in  make it faster and the other way around.
make it faster and the other way around.
The third stage starts when diffusion powered widening of the initial narrow peak slows down enough in comparison to local drift at the edges of the peak. That’s the point in time when the process becomes stochastic in nature. One or several “species” could be formed, and they run away in the direction of increased efficiency of the species. This process is quick as it is also exponential in nature, though not as quick as the first stage. It is exponential because a small number of more efficient protocells find the food abundant whereas the bulk of the protocells balance between replication and extinction. When the grid step is sufficiently small to resolve a global asymmetry factor, the system consistently produces more efficient species. As we used a grid with  points in both
points in both  directions, that corresponds to global asymmetry factor
directions, that corresponds to global asymmetry factor  . We performed multiple runs, and the results consistently show directed symmetry breaking in the system. Smaller values of
. We performed multiple runs, and the results consistently show directed symmetry breaking in the system. Smaller values of  may result in two species form near
may result in two species form near  , and less efficient species (
, and less efficient species ( in our case) may dominate in the system for some prolonged period, after which more efficient species may still overcome less efficient ones. If the total amount of food in the system
in our case) may dominate in the system for some prolonged period, after which more efficient species may still overcome less efficient ones. If the total amount of food in the system  is larger, then such event is more likely to happen. This is because for this event to occur there must be at least some protocells near
is larger, then such event is more likely to happen. This is because for this event to occur there must be at least some protocells near  and some large enough value of
and some large enough value of  . However, if
. However, if  is small, thenе this may not statistically happen and symmetry breaking becomes random when a grid cannot resolve the global asymmetry factor.
is small, thenе this may not statistically happen and symmetry breaking becomes random when a grid cannot resolve the global asymmetry factor.
The fourth stage is a relaxation toward a stationary point, which is a narrow peak near  ,
,  in case of the models considered here. When
in case of the models considered here. When  , then symmetry breaking occurs first and the system transitions toward a peak
, then symmetry breaking occurs first and the system transitions toward a peak  and some small value of
and some small value of  first, then slides toward
first, then slides toward  either as a whole bump or forming some runaway species. When
either as a whole bump or forming some runaway species. When  is large enough (we considered
is large enough (we considered  and it is large enough for the model considered here), then the system first accumulates substantial ability to pass information from generation to generation without experiencing any symmetry breaking, and symmetry breaking occurs only when the system already possesses substantial ability to pass information from generation to generation.
and it is large enough for the model considered here), then the system first accumulates substantial ability to pass information from generation to generation without experiencing any symmetry breaking, and symmetry breaking occurs only when the system already possesses substantial ability to pass information from generation to generation.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Author contributions
A.K. and K.K. developed the model. K.K. provided coding of the model. A.K. provided structure of the manuscript and charts. A.K. and K. K. wrote the main manuscript and K.K. generated charts and animations. All authors reviewed the manuscript.
Data availability
The datasets generated during the current study are available in a “frozen” GitHub repository branch: https://github.com/kkkmail/CoreClm/tree/clm700-Fredholm-frozen-V2.
Declarations
Competing interests
The authors declare no competing interests.
Footnotes
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
- 1.Frank, F. C. On spontaneous asymmetric synthesis. Biochim. Biophys. Acta. 11, 459–463 (1953). [DOI] [PubMed] [Google Scholar]
- 2.Kondepudi, D. K. & Nelson, G. W. Chiral symmetry breaking in nonequilibrium systems. Phys. Rev. Lett.50 (14), 1023–1026 (1983). [Google Scholar]
- 3.Steel, M. The emergence of a Self-Catalysing structure in abstract Origin-Of-Life models. Appl. Math. Lett.13, 91–95 (2000). [Google Scholar]
- 4.Sandars, P. G. H. A toy model for the generation of homochirality during polymerization. Orig Life Evol. Biosph. 33, 575–587 (2003). [DOI] [PubMed] [Google Scholar]
- 5.Plasson, R., Bersini, H. & Commeyras, A. Recycling Frank: spontaneous emergence of homochirality in noncatalytic systems. PNAS101 (48), 16733–16738 (2004). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Pizzarello, S. & Weber, A. L. Prebiotic amino acids as asymmetric catalysts. Science303 (5661), 1151 (2004). [DOI] [PubMed] [Google Scholar]
- 7.Joyce, G. F. Directed evolution of nucleic acid enzymes. Annu. Rev. Biochem.73, 791–836 (2004). [DOI] [PubMed] [Google Scholar]
- 8.Wattis, J. A. D. & Coveney, P. V. Symmetry-breaking in chiral polymerisation. Orig Life Evol. Biosph. 35, 243 (2005). [DOI] [PubMed] [Google Scholar]
- 9.Weissbuch, I., Leiserowitz, L. & Lahav, M. Stochastic mirror symmetry breaking via Self-Assembly, reactivity and amplification of chirality: relevance to abiotic conditions. Top. Curr. Chem.259, 123–165 (2005). [Google Scholar]
- 10.Brandenburg, A., Lehto, H. J. & Lehto, K. M. Homochirality in an early peptide world. Astrobiology7 (5), 725–732 (2007). [DOI] [PubMed] [Google Scholar]
- 11.Gleiser, M., Thorarinson, J. & Walker, S. I. Punctuated chirality. Orig Life Evol. Biosph. 38, 499–508 (2008). [DOI] [PubMed] [Google Scholar]
- 12.Kafri, R., Markovitch, O. & Lancet, D. Spontaneous chiral symmetry breaking in early molecular networks. Biol. Direct. 5, 38 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Blackmond, D. G. The origin of biological homochirality. Cold Spring Harb. Perspect. Biol.2 (5), a002147 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Eigen, M. Selforganization of matter and the evolution of biological macromolecules. Naturwissenschaften58, 465–523 (1971). [DOI] [PubMed] [Google Scholar]
- 15.Kauffman, S. A. Autocatalytic sets of proteins. J. Theor. Biol.119 (1), 1–24 (1986). [DOI] [PubMed] [Google Scholar]
- 16.Szostak, J. W. An optimal degree of physical and chemical heterogeneity for the origin of life? Proc. of the National Academy of Science. 108 (44), 17676–17682 (2011). [DOI] [PMC free article] [PubMed]
- 17.Smith, E. & Morowitz, H. J. Universality in intermediary metabolism. PNAS101 (36), 13168–13173 (2004). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Schrödinger, E. What Is Life? The Physical Aspect of the Living Cell (Cambridge University Press, 1944).
- 19.England, J. L. Dissipative adaptation in driven self-assembly. Nat. Nanotechnol.10 (11), 919–923 (2015). [DOI] [PubMed] [Google Scholar]
- 20.Lane, N. & Martin, W. The energetics of genome complexity. Nature467 (7318), 929–934 (2010). [DOI] [PubMed] [Google Scholar]
- 21.Pross, A. What Is Life? How Chemistry Becomes Biology (Oxford University Press, 2016).
- 22.Lanier, K. A. & Williams, L. D. The origin of life: models and data. J. Mol. Evol.86 (4–5), 249–260 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Konstantinov, K. K. & Konstantinova, A. F. Evolutionary approach to biological homochirality. Origins Life Evol. Biospheres. 52, 205–232 (2022). [DOI] [PubMed] [Google Scholar]
- 24.Robertson, M. P. & Joyce, G. F. The origins of the RNA world. Cold Spring Harb Perspect. Biol.4 (5), a003608 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Monnard, P-A., Walde, P. & Current Ideas about Prebiological Compartmentalization Life5, 1239–1263 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Bahadur, K. & Ranganayaki, S. Synthesis of Jeewanu, the units capable of growth, multiplication and metabolic activity. I. Preparation of units capable of growth and division and having metabolic activity. Zentr Bakteriol Parasitenk. 117 (11), 567–574 (1964). [Google Scholar]
- 27.Gupta, V. K. & Rai, R. K. Histochemical localisation of RNA-like material in photochemically formed self-sustaining, abiogenic supramolecular assemblies ‘jeewanu’. Int. Res. J. Sci. Eng.1 (1), 1–4 (2013). [Google Scholar]
- 28.Gillespie, D. T. Approximate accelerated stochastic simulation of chemically reacting systems. J. Chem. Phys.115, 1716–1733 (2001). [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The datasets generated during the current study are available in a “frozen” GitHub repository branch: https://github.com/kkkmail/CoreClm/tree/clm700-Fredholm-frozen-V2.