Abstract
The molecular determinants of stimulation of –1 programmed ribosomal frameshifting (–1 PRF) by RNA pseudoknots are poorly understood. Sugarcane yellow leaf virus (ScYLV) encodes a 28-nt mRNA pseudoknot that promotes –1 PRF between the P1 (protease) and P2 (polymerase) genes in plant luteoviruses. The solution structure of the ScYLV pseudoknot reveals a well ordered loop 2 (L2) that exhibits continuous stacking of A20 through C27 in the minor groove of the upper stem 1 (S1), with C25 flipped out of the triple-stranded stack. Five consecutive triple base pairs flank the helical junction where the 3′ nucleotide of L2, C27, adopts a cytidine 27 N3-cytidine 14 2′-OH hydrogen bonding interaction with the C14-G7 base pair. This interaction is isosteric with the adenosine N1–2′-OH interaction in the related mRNA from beet western yellows virus (BWYV); however, the ScYLV and BWYV mRNA structures differ in their detailed L2–S1 hydrogen bonding and L2 stacking interactions. Functional analyses of ScYLV/BWYV chimeric pseudoknots reveal that the ScYLV RNA stimulates a higher level of –1 PRF (15 ± 2%) relative to the BWYV pseudoknot (6 ± 1%), a difference traced largely to the identity of the 3′ nucleotide of L2 (C27 vs. A25 in BWYV). Strikingly, C27A ScYLV RNA is a poor frameshift stimulator (2.0%) and is destabilized by ≈1.5 kcal·mol–1 (pH 7.0, 37°C) with respect to the wild-type pseudoknot. These studies establish that the precise network of weak interactions nearest the helical junction in structurally similar pseudoknots make an important contribution to setting the frameshift efficiency in mRNAs.
Keywords: base triple, RNA pseudoknot, translational recoding
The Luteoviridae family encompasses a large class of single-stranded positive sense RNA viruses that infect a wide range of agriculturally important plants (1, 2). Assembly and replication of these and other vertebrate RNA viruses, e.g., retroviruses and coronaviruses, is regulated by maintaining optimal ratios of gene products (3). The molar ratio of two gene products, the P1-encoded viral protease and P2-encoded RNA-dependent RNA polymerase, is controlled at the level of translation with –1 programmed ribosomal frameshifting (–1 PRF) mediating the production of a P1-P2 fusion protein. The efficiency of this frameshifting event varies between 2% and 15% among different viruses, with the majority of elongating ribosomes terminating translation normally at a P1 stop codon (4–6). Although it is unknown to what extent the frameshifting efficiency in plant luteoviruses influences viral replication and infectivity, studies in the human retrovirus HIV-1, Moloney murine leukemia virus, and the yeast L-A double-stranded RNA virus reveal that an optimal level of frameshift stimulation is essential for maintaining high levels of virus infectivity and propagation (7–10).
–1 PRF has been discovered in many classes of eukaryotic RNA viruses from yeast to plants to humans, including pathogenic retroviruses, coronaviruses (severe acute respiratory syndrome and mouse hepatitis virus), and many plant viruses (11). Although it is relatively rare, –1 PRF has also been documented to occur in prokaryotes (Escherichia coli dnaX gene), with just one instance of –1 PRF clearly shown to occur in mammalian cells (Edr gene) (12). Recent findings suggest that –1 PRF might be used in Saccharomyces cerevisiae to endogenously posttranscriptionally regulate gene expression by targeting mRNAs containing premature termination codons for nonsense-mediated mRNA decay (13).
The key mRNA-encoded determinants that are necessary and sufficient for frameshift stimulation are a heptanucleotide slips-ite (XXXYYYZ), a downstream RNA structural motif, which is typically an RNA pseudoknot, and a connecting 6- to 8-nt spacer (11, 14). The identity of two nucleotides 5′ to the slip-site heptamer also appear to influence frameshifting efficiencies in yeast (15). A modeling experiment with the beet western yellows virus (BWYV) pseudoknot suggests that the pseudoknot lies just outside the narrow mRNA entry tunnel of the ribosome (16), allowing for RNA–protein interactions with the ribosome, which may contribute to ribosomal pausing and/or facilitate the frameshifting process in some unknown way (17). It has been suggested that the RNA pseudoknot provides a torsional restraint to the movement of the single-stranded mRNA during translocation (16). To release this tension, the pseudoknot could be unwound or the mRNA can shift by one nucleotide producing a –1 frameshift. An alternate model suggests that proteins S3, S4, and S5 of the ribosome function as an RNA helicase to unfold mRNA structures (18, 19). A stable three-stranded structure such as a pseudoknot would provide a barrier for the helicase, inhibiting the unwinding of the pseudoknot and preventing forward motion of the elongating ribosome in the reference frame.
Hairpin-type luteoviral mRNA pseudoknots from BWYV, pea enation mosaic virus 1 (PEMV-1) and potato leaf roll virus (PLRV) have been previously studied by NMR, x-ray crystallographic, and thermodynamic methods (4–6, 20–25). Each of these pseudoknots adopts a unique triple helical architecture that is characterized by an adenosine stack at the 3′ end of L2, which forms hydrogen bonding interactions into the minor groove of the base of S1. These pseudoknots also contain a protonated C+·(G-C) L1-S2 major groove Hoogsteen-type base triple at the helical junction of S1 and S2, with protonation strongly stabilizing these RNAs by 2–3 kcal·mol–1 at 37°C (23, 24). The closing base pair of S1 stacks on the L1-S2 C+·G Hoogsteen base pair, creating a strongly over-rotated and horizontally displaced helical junction, allowing the 3′ adenosine base of L2 to stack directly on S2.
Previous studies on BWYV and PLRV P1-P2 mRNA pseudoknots showed that substitutions and deletions in the 3′ end of L2 reduced frameshifting efficiency to a level below 1% (4, 5). Adenosine-to-cytidine substitutions in this region of either RNA were not reported, despite the fact that several proposed P1-P2 pseudoknots, e.g., sugarcane yellow leaf virus (ScYLV) (2), cereal yellow dwarf virus–RPS (CYDV-RPS) and beet chlorosis virus (BCV), contain naturally occurring L2 cytidines (26). Cytidine is isosteric with adenosine in forming cis-Watson–Crick/sugar edge hydrogen bonding interactions found in the BWYV and PEMV-1 pseudoknots (27). The ScYLV pseudoknot is of particular interest because this RNA contains two naturally occurring L2 adenosine to cytidine substitutions (C25 and C27; see Fig. 1).
Fig. 1.
Nucleotide sequences of the ScYLV and BWYV RNA pseudoknots. (A) The genomic ScYLV RNA sequence (GenBank entry NC_000874) that corresponds to the P1-P2 –1 mRNA frameshift signal. Nucleotide sequence and secondary structural representations of the ScYLV (B) and BWYV (C) P1-P2 frameshift-stimulating RNAs used in this study. Nonnative nucleotides are in lowercase. Symbols represent non-Watson–Crick base pair interactions (27) identified in the ScYLV (this work) and BWYV (20) RNA tertiary structures.
Here, we present the NMR-derived solution structure of the P1-P2 frameshift-stimulating pseudoknot from ScYLV. The structure reveals that C27 interacts with the minor groove of S1, where it has a significant impact on setting the frameshifting efficiency. A mutational analysis of ScYLV/BWYV chimeric pseudoknots, coupled with structural and thermodynamic studies, provides significant insight into the functional differences between the structurally similar ScYLV and BWYV mRNA pseudoknots. These studies provide significant support for the hypothesis (28) that small decreases in the stability of the helical junction (29) due to small changes in local structure significantly modulate frameshifting efficiency by mRNA pseudoknots.
Methods
Preparation of RNA Samples. All RNA constructs were prepared by runoff transcription using SP6 RNA polymerase and purified as described in ref. 24. The final NMR buffer was 10 mM potassium phosphate, 100 mM KCl, and 5 mM MgCl2 (pH 6.0), with RNA concentrations ranging from ≈0.9 to 2.0 mM. For residual dipolar coupling (RDC) experiments, 12.5 mg/ml Pf1 phage (ASLA Biotech, Riga, Latvia) was added, producing a 2H2O splitting of ≈9 Hz.
Resonance Assignments. Nonexchangeable and exchangeable resonance assignments (BMRB deposition 6509) were obtained by using standard methodologies (see Supporting Methods, which is published as supporting information on the PNAS web site) (30) at 25°C and 10°C, respectively. NMR experiments were acquired on Varian Inova 500 MHz (Texas A&M University), Varian Inova 600 MHz (Texas A&M University/Scripps Research Institute), or Bruker 900 MHz (Scripps Research Institute) spectrometers. A 3D 13C-separated NOESY spectrum (τm = 120 ms, 1H frequency 900 MHz) provided the majority of NOE-derived distance restraints used for structure calculations. NMR data were processed by using NMRPIPE (31) and analyzed by using SPARKY (32).
RDC Experiments. Nucleobase 1DHC RDCs were derived by analysis of peak positions in constant time–transverse relaxation optimized spectroscopy (CT-TROSY) and CT-antiTROSY, acquired in the absence (isotropic) and the presence (anisotropic) of Pf1 phage (33). One-bond 1DHC ribose RDCs were obtained with a series of 20 J-modulated heteronuclear single quantum coherence (HSQC) experiments (34). Couplings were obtained from a simultaneous nonlinear least squares analysis of the isotropic and anisotropic data by constraining 1DHC to zero for the isotropic data set and allowing the value 1JHC to be a globally fitted parameter as implemented in IGOR (WaveMetrics, Lake Oswego, OR). One hundred and two RDCs (57 sugar, 43 base) were obtained from these experiments.
NMR Restraints and Structure Calculation Protocols. A total of 629 NOE distance restraints were obtained from a combination of a 13C-separated NOESY, short and long mixing time watergate NOESY, and 15N Carr–Purcell–Meiboom–Gill (CPMG)N-OESY spectra. The distribution of distance restraints throughout the molecule was even (≈25 per residue) and included 68 L2-S1 restraints. NOEs were categorized as strong (2.5 ± 0.5 Å), medium (3.3 ± 0.7 Å), weak (4.0 ± 1.0 Å), and very weak (5.0 ± 1.0 Å). Seventy-two NOE-type distance and base pair planarity restraints were used to constrain the eight Watson–Crick base pairs as well as the trans-Watson–Crick/Hoogsteen base pair formed by C8+·G12, consistent with nonselective 1H-15N-15N-correlation spectroscopy (HNN-COSY) and a selective 2Jnn HNN-COSY experiment (35, 36). No hydrogen bonding restraints were included for L2 nucleotides, with all reported noncanonical hydrogen bonds inferred from the final structure bundle. Loose backbone dihedral angle restraints (±60°) based on standard A-form RNA geometry were used for S1 and S2, consistent with the qualitative inspection of the 1H-31P crosspeak intensities in proton-phosphorus correlation experiments. Sugar pucker geometries were defined by the large number of intra-ribose NOE distance restraints.
Structure calculations were performed by using XPLOR-NIH 2.10 (37). The initial ScYLV RNA NOE-derived structure bundle of the 20 lowest-energy structures with no NOE violations >0.5 Å was derived from 100 random structures by using a simulated annealing protocol like that described in ref. 6. Each of the 20 structures was then subjected to an RDC-based refinement protocol, in which local structure is first refined, and then global structure (38) (see Supporting Methods). The convergence of the final 20 structures was excellent (global rms deviation of 1.85 Å) with structure statistics compiled in Table 1. A comparison of the experimental and calculated RDCs derived from the structure bundle revealed an excellent correlation (R = 0.983, slope = 1.008, intercept = –0.062), revealing self-consistency of the refinement. The atomic coordinates of the ScYLV RNA structure bundle (ID code 1YG3) and the average structure (ID code 1YG4) have been deposited in the Protein Data Bank.
Table 1. Experimental restraints and structure statistics for the ScYLV pseudoknot.
| NOE only | NOE+RDC | |
|---|---|---|
| Number of structures | 20 | 20 |
| Distance restraints | 629 | 629 |
| Intranucleotide | 380 | 380 |
| Internucleotide | 249 | 249 |
| Backbone dihedral angle restraints | 65 | 65 |
| Residual dipolar couplings | n/a | 102 |
| Sugar | n/a | 57 |
| Base | n/a | 45 |
| rms from experimental restraints | ||
| Distance restraints,* Å | 0.061 ± 0.005 | 0.065 ± 0.005 |
| Dihedral restraints,† ° | 0.949 ± 0.753 | 0.041 ± 0.122 |
| Dipolar couplings, Hz | n/a | 1.60 ± 0.04 |
| Deviations from idealized geometry | ||
| Bonds, Å | 0.018 ± 0.001 | 0.018 ± 0.001 |
| Angle, ° | 2.35 ± 0.10 | 2.84 ± 0.11 |
| Impropers, ° | 1.48 ± 0.07 | 1.87 ± 0.11 |
| Heavy-atom rms deviation, Å | ||
| Overall (residues 3-8, 10-12, 14-24, 26-30) | 2.20 ± 0.40 | 1.85 ± 0.42 |
| S1 | 1.47 ± 0.47 | 1.33 ± 0.36 |
| S2 | 1.60 ± 0.57 | 1.34 ± 0.60 |
No violations >0.5 Å.
No violations >5°.
Frameshifting Assays. The dual luciferase reporter assay system (Promega) was used to measure frameshifting efficiencies as described in refs. 6 and 39. Pseudoknot constructs were cloned into p2luc and contained the wild-type ScYLV slippery site (GGGAAAC), a 6-nt linker containing a stop codon in the reference frame, and the appropriate sequence for the RNA pseudoknot (Fig. 1). All frameshifting data were normalized to an in-frame control (6).
Thermal Denaturation Experiments. Optical melting profiles were collected in 10 mM buffer/0.5 M KCl at the indicated pH as described in ref. 23. Both sets of derivative data (dA260/dT and dA280/dT) were subjected to a simultaneous nonlinear least-squares fit to a multiple (i) sequential interacting two-state transition model that optimizes ΔHi and tm,i for the unfolding of each ith unfolding transition implemented by the algorithm t-melt assuming ΔC°p = 0 (29, 40). A model invoking two sequential two-state unfolding transitions, assigned to folded RNA to S1 hairpin and S1 hairpin to unfolded RNA unfolding steps, was the simplest model consistent with the data, as judged by the criteria that ΣΔHvH,i ≈ ΔHcal (calorimetric enthalpy) and ΔHvH,2 and ΔHcal,2 for the unfolding of S1 (transition 2) was as expected from the nearest-neighbor model (41). Fitting the data to a single two-state transition results in nonrandom residuals and insufficient van't Hoff enthalpy of unfolding. The unfolding entropies were obtained from ΔS = ΔH/tm while ΔG37 = ΔH –310.15 × ΔS.
Results and Discussion
NMR Analysis and Overview of the Structure. The naturally occurring frameshift-stimulating signal that overlaps the P1 and P2 genes in ScYLV is shown in Fig. 1 A, as are secondary structural representations of the ScYLV and BWYV RNA pseudoknot constructs characterized in this study (Fig. 1 B and C). The NMR spectra of the ScYLV pseudoknot exhibit features characteristic of this family of pseudoknots (see Figs. 8 and 9, which are published as supporting information on the PNAS web site), revealing that the Watson–Crick faces of L2 adenosines and C27 make multiple hydrogen bonding interactions with the minor groove of S1, including the 2′-OH protons.
A global superposition of the 20 lowest-energy structures is displayed in Fig. 2A, with the average structure shown in Fig. 2B. Structure statistics are compiled in Table 1. The final structure bundle is characterized by a global [nucleotides 3–30, excepting extrahelical residues G9 (L1), A13 (at the S1-S2 junction), and C25 (L2)] heavy atom rms deviation of 1.85 Å. The helix–helix bend angle deviates from colinearity by ≈39° and is well defined by the structure bundle with little fraying at the helical termini. The local structure of the molecule is also very well determined, particularly in L2 (rms deviation of 1.39 Å, excluding C25) and at the helical junction (Fig. 4C). The S1-S2 helical junction is significantly over-rotated by 80° relative to A-form helical geometry, with the helices horizontally displaced by 7.6 Å. As a result, the junction base pair of S1, G7-C14, is stacked on the C8+·G12 Hoogsteen base pair in the C8+·(G12-C28) base triple (Fig. 2C), permitting pseudocontinuous stacking of the 3′ L2 nucleotide C27 on C28. G9 (3′ nucleotide in L1) is exposed to solvent, as is A13 (formally pseudoknot loop L3), which functions as a linker nucleotide between G12 and C14 (6, 20).
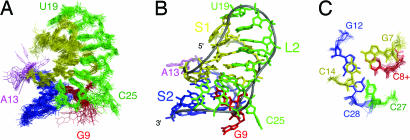
Fig. 2.
The structure of the ScYLV mRNA pseudoknot with stem S1 in yellow, stem S2 in blue, loop L1 in red, loop L2 in green, and A13 in violet. (A) A global superposition of the 20 lowest-energy structures. The minor groove side of S1 with the L2 nucleotides reaching into the groove is in front. (B) The average structure displayed in the same orientation as in A. Extrahelical residues are labeled. (C) The helical junction region showing the C27·(C14-G7) L2-S1 minor groove base triple stacked on the C8+·(G12-C28) L1-S2 Hoogsteen base triple.
Fig. 4.
Comparison of the helical junction regions of the ScYLV and BWYV RNA pseudoknots. (A) Superposition of the bottom two base pairs of stem S1 at the helical junction (shaded yellow) and the 3′ three nucleotides of loop L2 (ScYLV, green; BWYV residues A23, A24, and A25, blue) in the ScYLV and BWYV pseudoknots. (B) View of the two consecutive minor and major groove base triples that flank the helical junction in the ScYLV (Left) and BWYV (Right) pseudoknots (coloring scheme as in Fig. 2C).
A notable feature of the ScYLV molecule is the nearly continuous stacking of adenosines A20 to A26 down the minor groove of S1, with A26 stacked on C27, and C25 extruded from the triple-stranded stack (Fig. 2 A and B). The conformation of L2 is held into the S1 minor groove by four pairs of reciprocal cis-Watson–Crick/sugar edge L2 base-S1 ribose, L2 base-S1 base hydrogen bonds with the antiparallel strand of S1 (Fig. 3). C27 forms the base of the L2 stack with its Watson–Crick face pointing into the S1 minor groove where the N3 atom accepts a hydrogen bond from the 2′-OH group of C14, an arrangement that is isostructural with the other interactions (Fig. 3) as well as the analogous A25·C14 interaction in the BWYV pseudoknot. This run of four stacked L2-S1 minor groove base triples (Fig. 3) is further stacked on the L1-S2 C8+·(G12-C28) major groove triple base pair. Thus, the ScYLV RNA is characterized by five contiguous base triples, interrupted by only a single solvent-exposed nucleotide (C25) that crosses the helical junction of the molecule, thereby stabilizing the triple-helical fold of the RNA. The structures of the two related luteoviral RNA pseudoknots from BWYV (20) and PEMV-1 (6) also reveal stacked base triples that cross the helical junction; however, these RNAs contain just three contiguous triples. Like the ScYLV RNA, a run of five consecutive junction-crossing triple base pairs are found in human telomerase RNA pseudoknot (42).
Fig. 3.
Structures of the four consecutive isosteric cis-Watson–Crick/sugar edge interactions (27) that orient loop L2 in the minor groove of stem S1 in the ScYLV pseudoknot. Heavy atom to heavy atom distances are shown for each hydrogen bonding interaction and represent mean distances derived from the 20 lowest-energy structures. No hydrogen bonding restraints were used to constrain these distances during the structure calculations and RDC-based refinement. H-bonds are well supported by the NMR data (Figs. 8 and 9). Recalculation of the structure including these hydrogen bonding restraints (d = 2.8 ± 0.3 Å) yields an essentially identical average structure.
A comparison of the ScYLV and the related BWYV (21) pseudoknots (Fig. 4) reveals that despite similar global structures, there are distinct structural differences. L2 in the BWYV RNA is shorter by two nucleotides yet crosses the same length 5-bp S1, leading to an altered stacking arrangement of L2 nucleotides relative to ScYLV (21). The ScYLV RNA exhibits stacking in L2 from A20 to C27 (excluding C25) with all eight nucleotides pointing into the S1 minor groove, leading to distinct L2-S1 hydrogen bonding interactions (see Fig. 3) as well as the different helical junction architectures (Fig. 4 A and B). The stacking of C27 on C28 is altered in the ScYLV RNA with respect to A25 on C26 in the BWYV pseudoknot reflected by a shorter C27 C1′-C14 C1′ distance (7.5 Å vs. 9.2 Å), and a slightly larger degree of over-rotation and horizontal displacement of the upper stem S1 vs. S2 in the ScYLV RNA (Fig. 4B). The stacking of G7 on C8 also differs in the two RNAs, consistent with the distinct chemical shifts of the imino protons of both G7 and C8+ and the amino protons of C8+. This is also true for the C27A ScYLV RNA (vide infra; see Fig. 10, which is published as supporting information on the PNAS web site).
Functional Consequences of a C-Minor Groove Interaction. Although a cytidine cis-Watson–Crick/sugar edge interaction is notun precedented (27, 43–45), the isosteric adenosine cis-Watson–Crick/sugar edge interaction is more prevalent in RNA structures (46, 47). Indeed, the ScYLV RNA is the first example of a C·(C-G) minor groove base triple in a naturally occurring hairpin-type pseudoknot. It was therefore of interest to determine the extent to which a 3′ L2 cytidine (C27) vs. adenosine in the ScYLV RNA stimulates ribosomal frameshifting.
The in vitro frameshifting efficiency of the wild-type ScYLV pseudoknot in a heterologous dual luciferase system (39) was found to be 15.2%, revealing that the cytidine at the 3′ end of L2 yields a functional pseudoknot (Fig. 5A), in contrast with a recent report (15). Consistent with previous findings in other luteoviral pseudoknots (4–6), guanosine substitutions of C27, A26, and A24, i.e., the AAC-3′ stack in L2, reduces frameshifting to a level indistinguishable from the slip site alone or the S1 hairpin (Fig. 5A). Under identical conditions, the wild-type BWYV frameshifting pseudoknot exhibits a frameshifting efficiency of 6.6% or ≈2.5-fold lower than the ScYLV RNA, consistent with results previously obtained (Fig. 5C) (4).
Fig. 5.
In vitro frameshifting efficiencies determined for wild-type and substitution/deletion mutants of the ScYLV and BWYV pseudoknots (see Methods). (A) ScYLV mutations representative of those shown to abolish frameshift stimulation in other luteoviral systems (4–6). (B) Mutational analysis designed to functionally convert the ScYLV pseudoknot into the BWYV pseudoknot. (C) Direct comparison of frameshifting efficiencies of the wild-type ScYLV, C27A ScYLV, wild-type BWYV, and A25C BWYV pseudoknots.
A mutational study was carried out in an effort to functionally convert the higher frameshift-stimulating ScYLV pseudoknot into the less efficacious BWYV pseudoknot. Because S2 in BWYV and ScYLV RNAs is identical (see Fig. 1 B and C), unpaired, extrahelical residues G9, A13, and C25, as well as C27, were targeted for substitution, singly or in combination, with the corresponding residues in the BWYV RNA (Fig. 5B). The A13U substitution is functionally silent, consistent with its role as a linker nucleotide (Fig. 2). Interestingly, both G9A and ΔC25 mutants increase frameshifting by 1.5- and 2.0-fold, respectively, results difficult to rationalize from an isolated inspection of the ScYLV RNA structure alone. However, G9 might play a role in modulating the pKa of the adjacent C8 N3H+, and/or the structure of the Hoogsteen triple itself. Deletion of C25, on the other hand, might eliminate irregularities in the wild-type ScYLV triple helical architecture increasing the barrier for ribosome helicase-induced unwinding (19), or alternatively, reduce the loop-closing entropy of loop L2 thereby stabilizing the RNA. Another possible explanation is that deletion of C25 could lower the entropic cost of binding to the ribosome, leading to more efficient frameshift stimulation. Importantly, neither G9A and ΔC25 substitution lowers frameshifting efficiency.
In contrast, incorporation of a single BWYV-like C27A substitution into the ScYLV pseudoknot is strongly inhibitory (≈8-fold reduction), decreasing the frameshifting efficiency to a level just 2-fold above basal levels, a surprising finding given that this substitution is compatible with stable pseudoknot formation (vide infra). This finding makes the prediction that an A25C substitution in the BWYV RNA would significantly increase the frameshifting efficiency of the native BWYV pseudoknot; this is exactly what is observed (Fig. 5C).
Superposition of G9A and A13U extrahelical base substitutions on the C27A ScYLV RNA (triple; Fig. 5B) gives rise to a frameshifting efficiency 2-fold lower than the wild-type BWYV pseudoknot; subsequent reduction of the L2 length to that of the BWYV pseudoknot (nine to seven nucleotides; triple + ΔA24 + ΔC25) (see Fig. 1) results in a frameshifting efficiency equivalent to that of wild-type BWYV (Fig. 5C). Thus, three substitutions of G9, A13, and C27 to BWYV nucleotides, coupled with a reduction of L2 by two nucleotides, are sufficient to functionally convert the ScYLV pseudoknot to a frameshifting efficiency identical to that of the BWYV RNA. This analysis further suggests that disruption of the two triple base pairs unique to the ScYLV RNA and closest to the approaching ribosome, A21·(C17-G4) and A22·(A16-U5), significantly reduces frameshift stimulation because the 7-nt L2 of the triple + BWYV L2 RNA is less active than the RNA containing a 7-nt ScYLV L2 (triple + ΔA24 + ΔC25) packed against the native ScYLV S1 (Fig. 5B). This result is consistent with the functional effects of substitution of the middle S1 base pair in the BWYV RNA, G16-C5, with an ScYLV-like A16-U5 base pair, which reduces frameshift-stimulation ≈2-fold in the BWYV RNA (4).
C27A ScYLV and A25C BWYV RNAs Form Stable Folded Pseudoknots. The functional studies leave open the possibility that the C27A ScYLV RNA is partially folded, thus reducing frameshift-stimulation relative to the wild-type RNA. To address this, 1D NMR spectra of ScYLV C27A, wild-type BWYV, and BWYV A25C RNAs were compared with that of the wild-type ScYLV pseudoknot (Fig. 6). These downfield regions of the proton spectra reveal that the C27A ScYLV and A25C BWYV pseudoknots are stably folded at 10°C and characterized by a C+·(G-C) L1-S2 triple base pair at pH 6.0. However, the C14 2′-OH resonance in the C27A ScYLV RNA is shifted downfield to ≈10.0 ppm, relative to 8.57 ppm in the wild-type RNA (Fig. 9A). A comparison of the spectra of wild-type and A25C BWYV RNAs reveals the same trend: The C14 2′-OH proton resonates at ≈9.9 ppm when an adenosine and not a cytidine is present in the terminal 3′ L2 position. Thus, the network of hydrogen bonding and base stacking interactions around the helical junctions of the A-minor motif in the C27A ScYLV and wild-type BWYV pseudoknots are substantially similar but distinct from that of wild-type ScYLV and A25C BWYV RNAs with characteristic C-minor interactions; this has a significant impact on frameshift stimulation. A comparison of the assigned proton chemical shift data for wild-type and C27A ScYLV pseudoknots reveals that the 1H chemical shift changes are not strictly localized to the junction region (Fig. 10). Indeed, local structural changes are most pronounced for the A27·(C14-G7) L2-S1 and adjacent C8+·(G12-C28) L1-S2 base triples, but perturbations are detectable for the C15-G6, A16-U5, and C17-G4 base pairs in S1, as well as the minor groove penetrating L2 adenosine stack.
Fig. 6.
Exchangeable 1H regions of jump-return echo 1D NMR spectra (10°C) acquired for the wild-type ScYLV (ScYLV WT), C27A ScYLV, wild-type BWYV (BWYV WT), and A25C BWYV RNA with corresponding frameshifting efficiencies shown on the left. Insets shown on the C27A ScYLV and WT BWYV spectra above the C14 2′-OH resonances were obtained from watergate NOESY spectra (τ = 200 ms) and represent ribose (4.0–5.0 ppm) and adenosine H2 (8.0–8.5 ppm) regions diagnostic for N1–2′-OH hydrogen bonding (6). Note that the line widths of the BWYV and BWYV A25C samples are broader and potentially reporting on some self-association (24).
The C27A Substitution Thermodynamically Destabilizes the ScYLV Pseudoknot. The NMR data show that the poor frameshift-stimulator C27A ScYLV RNA is a stably folded pseudoknot and that the microenvironment of the helical junction region is perturbed relative to the wild-type ScYLV RNA. Thermal denaturation studies were next carried out to determine whether this substitution results in a stabilization or destabilization of the pseudoknot.
Representative equilibrium melting profiles (dA/dT) are shown for wild-type and C27A ScYLV RNA pseudoknots (Fig. 7A). The unfolding of both RNAs is well described by two unfolding transitions, the first assignable to the intact pseudoknot-to-S1 hairpin step, followed by melting of the thermodynamically most stable S1 hairpin (see Methods). Any destabilization of the helical junction or L2-S1 interactions will therefore manifest itself in the first unfolding transition. Visual inspection of the melting profiles reveals that this is indeed the case. Quantitative pH-dependent analysis reveals that the C27A substitution is enthalpically destabilizing leading to a destabilization in free energy (ΔΔG 377.0) of 1.5 (±0.2) kcal·mol–1. This destabilization is manifest over the entire pH range investigated (Fig. 7B), with a shift in the macroscopic unfolding pKa for C8 N3 protonation by 0.4 units (23, 24). These data reveal that at least part of the global destabilization must be localized to the helical junction given the downward shift in the pKa.
Fig. 7.
Thermal unfolding of the wild-type and C27A ScYLV pseudoknots. (A) Representative optically monitored thermal unfolding profiles of the wild-type (Left) and C27A (Right) ScYLV pseudoknots at 260 nm (○) and 280 nm (♦) acquired at pH 8.0, 0.5 M KCl. Nonlinear least-squares simultaneous composite fit to a two-transition unfolding model (black thin line) and component transition 1 (red solid lines) and transition 2 (blue solid lines) are shown. Results from four independent wild-type ScYLV RNA-unfolding experiments return the following thermodynamic parameters: transition 1, ΔH = 47.5 (±2.6) kcal·mol–1; tm = 67.5 (±0.7) °C; and transition 2, ΔH = 58.4 (±0.5) kcal·mol–1; tm = 77.9 (±0.2) °C. Six independent C27A ScYLV RNA melts yields transition 1, ΔH = 40.9 (±1.7) kcal·mol–1; tm = 63.0 (±0.3) °C; transition 2, ΔH = 59.2 (±0.5) kcal·mol–1; tm = 77.8 (±0.3) °C. (B) Plot of ΔG37 for transition 1 vs. pH for wild-type (red) and C27A (blue) ScYLV RNAs. The solid line represents a fit to a model that assumes a single titratable group on the RNA (24) with a pKa of 7.4 (±0.1) and 7.0 (±0.2), ΔGprotonated of 6.8 (±0.1) and 6.1 (±0.2) kcal·mol–1 and ΔGunprotonated of 3.5 (±0.2) and 2.6 (±0.2) for wild-type and C27A ScYLV RNAs, respectively.
Conclusions
Frameshift stimulation requires kinetic partitioning of the elongating ribosome into a new reading frame at a rate that must be fast relative to the time scale of translation elongation in the reference frame (14). To effect partitioning, the pseudoknot must be stable enough to remain largely folded when it encounters the force of the elongating ribosome, thereby stalling it over the slip-site (17). Our data are consistent with a scenario where a local destabilization allows the elongating ribosome to unfold the distal pseudoknot-forming stem S2 before significant partitioning into the –1 reading frame. In fact, although the data are limited, the destabilization separating wild-type and C27A ScYLV RNAs is similar in magnitude to the degree to which the helical junction regions are destabilized in other frameshifting systems (ΔΔG37 ≈ 1–2 kcal·mol–1) due to substitutions or deletions of junction nucleotides that negatively impact frameshift stimulation (24, 28, 29, 40, 48, 49). Indeed, if the entire ground-state destabilization of mutant pseudoknots observed here (ΔΔG37 ≈ 1.5 kcal·mol–1) is manifest as a decrease in the energy barrier to ribosome-promoted pseudoknot unwinding, ΔΔG‡, then this would translate into an ≈10-fold increase in the rate of unfolding in destabilized pseudoknots (14).
Our findings show that a major functional modulator of frameshift stimulation is the 3′ nucleotide of loop L2, particularly in the context of hairpin-type RNA pseudoknots characterized by short helical stems (14, 49, 50). This nucleotide sets the stage for defining the architecture of noncovalent interactions in the junction region and for anchoring additional interactions between L2 and S1. Key L2-S1 tertiary structural interactions can be envisioned to function as kinetic barriers to the unfolding of pseudoknot-forming stem S2, in the same way that Mg2+-dependent tertiary structural interactions function as kinetic barriers to the unwinding of component helices in larger RNAs (51). The related ScYLV and BWYV pseudoknotted RNAs characterized here constitute an excellent model system with which to further address mechanistic (52, 53) and kinetic (54) issues relevant for understanding mRNA pseudoknot-stimulated –1 PRF by structurally very similar mRNAs.
Supplementary Material
Acknowledgments
We thank Dr. Karl Koshlap (Texas A&M University) for acquiring some of the NMR data. This work was supported by National Institutes of Health (NIH) Grant AI040187 and Texas Higher Education Coordinating Board Advanced Research Program Grants 010361-0278-1999 and 010366-0172-2001. P.V.C. was supported by NIH Chemistry–Biology Interface Training Grant T32 GM08523 to Texas A&M University.
Author contributions: P.V.C., M.H., and D.P.G. designed research; P.V.C., M.H., and D.P.G. performed research; P.V.C. and D.P.G. analyzed data; and P.V.C., M.H., and D.P.G. wrote the paper.
Abbreviations: BWYV, beet western yellows virus; L1, loop 1; L2, loop 2; –1 PRF, –1 programmed ribosomal frameshifting; RDC, residual dipolar coupling; S1, stem 1; S2, stem 2; ScYLV, sugarcane yellow leaf virus.
Data deposition: The atomic coordinates have been deposited in the Protein Data Bank, www.pdb.org [PDB ID codes 1YG3 (ScYLV RNA structure bundle) and 1YG4 (average structure)].
References
- 1.Smith, H. G. & Barker, H. (1999) The Luteoviridae (CABI Publishing, New York).
- 2.Moonan, F., Molina, J. & Mirkov, T. E. (2000) Virology 269, 156–171. [DOI] [PubMed] [Google Scholar]
- 3.Miller, W. A., Dinesh-Kumar, S. P. & Paul, C. P. (1995) Crit. Rev. Plant Sci. 14, 179–211. [Google Scholar]
- 4.Kim, Y. G., Su, L., Maas, S., O'Neill, A. & Rich, A. (1999) Proc. Natl. Acad. Sci. USA 96, 14234–14239. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Kim, Y. G., Maas, S., Wang, S. C. & Rich, A. (2000) RNA 6, 1157–1165. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Nixon, P. L., Rangan, A., Kim, Y., Rich, A., Hoffman, D. W., Hennig, M. & Giedroc, D. P. (2002) J. Mol. Biol. 322, 621–633. [DOI] [PubMed] [Google Scholar]
- 7.Hung, M., Patel, P., Davis, S. & Green, S. R. (1998) J. Virol. 72, 4819–4824. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Felsenstein, K. M. & Goff, S. P. (1988) J. Virol. 62, 2179–2182. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Dinman, J. D. & Wickner, R. B. (1992) J. Virol. 66, 3669–3676. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Biswas, P., Jiang, X., Pacchia, A. L., Dougherty, J. P. & Peltz, S. W. (2004) J. Virol. 78, 2082–2087. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Baranov, P. V., Gesteland, R. F. & Atkins, J. F. (2002) Gene 286, 187–201. [DOI] [PubMed] [Google Scholar]
- 12.Manktelow, E., Shigemoto, K. & Brierley, I. (2005) Nucleic Acids Res. 33, 1553–1563. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Plant, E. P., Wang, P., Jacobs, J. L. & Dinman, J. D. (2004) Nucleic Acids Res. 32, 784–790. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Giedroc, D. P., Theimer, C. A. & Nixon, P. L. (2000) J. Mol. Biol. 298, 167–185. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Bekaert, M. & Rousset, J. P. (2005) Mol. Cell 17, 61–68. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Plant, E. P., Jacobs, K. L., Harger, J. W., Meskauskas, A., Jacobs, J. L., Baxter, J. L., Petrov, A. N. & Dinman, J. D. (2003) RNA 9, 168–174. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Kontos, H., Napthine, S. & Brierley, I. (2001) Mol. Cell. Biol. 21, 8657–8670. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Yusupova, G. Z., Yusupov, M. M., Cate, J. H. & Noller, H. F. (2001) Cell 106, 233–241. [DOI] [PubMed] [Google Scholar]
- 19.Takyar, S., Hickerson, R. P. & Noller, H. F. (2005) Cell 120, 49–58. [DOI] [PubMed] [Google Scholar]
- 20.Su, L., Chen, L., Egli, M., Berger, J. M. & Rich, A. (1999) Nat. Struct. Biol. 6, 285–292. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Egli, M., Minasov, G., Su, L. & Rich, A. (2002) Proc. Natl. Acad. Sci. USA 99, 4302–4307. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Egli, M., Sarkhel, S., Minasov, G. & Rich, A. (2003) Helv. Chim. Acta 86, 1709–1727. [Google Scholar]
- 23.Nixon, P. L. & Giedroc, D. P. (2000) J. Mol. Biol. 296, 659–671. [DOI] [PubMed] [Google Scholar]
- 24.Nixon, P. L., Cornish, P. V., Suram, S. V. & Giedroc, D. P. (2002) Biochemistry 41, 10665–10674. [DOI] [PubMed] [Google Scholar]
- 25.Giedroc, D. P., Cornish, P. V. & Hennig, M. (2003) J. Am. Chem. Soc. 125, 4676–4677. [DOI] [PubMed] [Google Scholar]
- 26.Smith, G. R., Borg, Z., Lockhart, B. E., Braithwaite, K. S. & Gibbs, M. J. (2000) J. Gen. Virol. 81, 1865–1869. [DOI] [PubMed] [Google Scholar]
- 27.Leontis, N. B., Stombaugh, J. & Westhof, E. (2002) Nucleic Acids Res. 30, 3497–3531. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Chen, X., Chamorro, M., Lee, S. I., Shen, L. X., Hines, J. V., Tinoco, I., Jr., & Varmus, H. E. (1995) EMBO J. 14, 842–852. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Theimer, C. A. & Giedroc, D. P. (2000) RNA 6, 409–421. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Furtig, B., Richter, C., Wohnert, J. & Schwalbe, H. (2003) Chembiochem 4, 936–962. [DOI] [PubMed] [Google Scholar]
- 31.Delaglio, F., Grzesiek, S., Vuister, G. W., Zhu, G., Pfeifer, J. & Bax, A. (1995) J. Biomol. NMR 6, 277–293. [DOI] [PubMed] [Google Scholar]
- 32.Goddard, T. D. & Kneller, D. G., sparky (Univ. of California, San Francisco).
- 33.Boisbouvier, J., Brutscher, B., Pardi, A., Marion, D. & Simorre, J. P. (2000) J. Am. Chem. Soc. 122, 6779–6780. [Google Scholar]
- 34.Ottiger, M., Delaglio, F., Marquardt, J. L., Tjandra, N. & Bax, A. (1998) J. Magn. Reson. 134, 365–369. [DOI] [PubMed] [Google Scholar]
- 35.Dingley, A. J. & Grzesiek, S. (1998) J. Am. Chem. Soc. 120, 8293–8297. [Google Scholar]
- 36.Pervushin, K., Ono, A., Fernandez, C., Szyperski, T., Kainosho, M. & Wuthrich, K. (1998) Proc. Natl. Acad. Sci. USA 95, 14147–14151. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Schwieters, C. D., Kuszewski, J. J., Tjandra, N. & Marius Clore, G. (2003) J. Magn. Reson. 160, 65–73. [DOI] [PubMed] [Google Scholar]
- 38.McCallum, S. A. & Pardi, A. (2003) J. Mol. Biol. 326, 1037–1050. [DOI] [PubMed] [Google Scholar]
- 39.Grentzmann, G., Ingram, J. A., Kelly, P. J., Gesteland, R. F. & Atkins, J. F. (1998) RNA 4, 479–486. [PMC free article] [PubMed] [Google Scholar]
- 40.Theimer, C. A. & Giedroc, D. P. (1999) J. Mol. Biol. 289, 1283–1299. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Xia, T., SantaLucia, J., Jr., Burkard, M. E., Kierzek, R., Schroeder, S. J., Jiao, X., Cox, C. & Turner, D. H. (1998) Biochemistry 37, 14719–14735. [DOI] [PubMed] [Google Scholar]
- 42.Theimer, C. A., Blois, C. A. & Feigon, J. (2005) Mol Cell 17, 671–682. [DOI] [PubMed] [Google Scholar]
- 43.Ban, N., Nissen, P., Hansen, J., Moore, P. B. & Steitz, T. A. (2000) Science 289, 905–920. [DOI] [PubMed] [Google Scholar]
- 44.Batey, R. T., Gilbert, S. D. & Montange, R. K. (2004) Nature 432, 411–415. [DOI] [PubMed] [Google Scholar]
- 45.Serganov, A., Yuan, Y. R., Pikovskaya, O., Polonskaia, A., Malinina, L., Phan, A. T., Hobartner, C., Micura, R., Breaker, R. R. & Patel, D. J. (2004) Chem. Biol. 11, 1729–1741. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Nissen, P., Ippolito, J. A., Ban, N., Moore, P. B. & Steitz, T. A. (2001) Proc. Natl. Acad. Sci. USA 98, 4899–4903. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Kieft, J. S., Zhou, K., Grech, A., Jubin, R. & Doudna, J. A. (2002) Nat. Struct. Biol. 9, 370–374. [DOI] [PubMed] [Google Scholar]
- 48.Michiels, P. J., Versleijen, A. A., Verlaan, P. W., Pleij, C. W., Hilbers, C. W. & Heus, H. A. (2001) J. Mol. Biol. 310, 1109–1123. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Liphardt, J., Napthine, S., Kontos, H. & Brierley, I. (1999) J. Mol. Biol. 288, 321–335. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Wang, Y., Wills, N. M., Du, Z., Rangan, A., Atkins, J. F., Gesteland, R. F. & Hoffman, D. W. (2002) RNA 8, 981–996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Onoa, B., Dumont, S., Liphardt, J., Smith, S. B., Tinoco, I., Jr., & Bustamante, C. (2003) Science 299, 1892–1895. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Sytnik, A., Vladimirov, S., Jia, Y., Li, L., Cooperman, B. S. & Hochstrasser, R. M. (1999) J. Mol. Biol. 285, 49–54. [DOI] [PubMed] [Google Scholar]
- 53.Blanchard, S. C., Gonzalez, R. L., Kim, H. D., Chu, S. & Puglisi, J. D. (2004) Nat. Struct. Mol. Biol. 11, 1008–1014. [DOI] [PubMed] [Google Scholar]
- 54.Tinoco, I., Jr., Collin, D. & Li, P. T. (2004) Biochem. Soc. Trans. 32, 757–760. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.