Abstract
Odorants and pheromones as well as sweet- and bitter-tasting small molecules are perceived through activation of G protein-coupled chemosensory receptors. In contrast, gustatory detection of salty and sour tastes may involve direct gating of sodium channels of the DEG/ENaC family by sodium and hydrogen ions, respectively. We have found that ppk25, a Drosophila melanogaster gene encoding a DEG/ENaC channel subunit, is expressed at highest levels in the male appendages responsible for gustatory and olfactory detection of female pheromones: the legs, wings, and antennae. Mutations in the ppk25 gene reduce or even abolish male courtship response to females in the dark, conditions under which detection of female pheromones is an essential courtship-activating sensory input. In contrast, the same mutations have no effect on other behaviors tested. Importantly, ppk25 mutant males that show no response to females in the dark execute all of the normal steps of courtship behavior in the presence of visible light, suggesting that ppk25 is required for activation of courtship behavior by chemosensory perception of female pheromones. Finally, a ppk25 mutant allele predicted to encode a truncated protein has dominant-negative properties, suggesting that the normal Ppk25 protein acts as part of a multiprotein complex. Together, these results indicate that ppk25 is necessary for response to female pheromones by D. melanogaster males, and suggest that members of the DEG/ENaC family of genes play a wider role in chemical senses than previously suspected.
Keywords: courtship, behavior, olfaction, taste
As in most other animals, pheromones play key roles in the regulation of sexual behaviors of Drosophila melanogaster (1-3). In particular, several pheromones modulate male courtship of the female, which involves a stereotyped series of behaviors. By analogy with olfactory and gustatory perception of organic molecules in both insects and vertebrates (4), perception of these pheromones most likely involves interactions with seven-transmembrane receptors and subsequent activation of a G protein-coupled signal transduction pathway. Indeed, a male-specific member of the seven-transmembrane gustatory receptor family has been identified as a putative receptor for female courtship-stimulating pheromones (5). In contrast, gustatory perception of hydrogen and sodium ions, perceived as sour and salty tastes, respectively, has been suggested to involve direct gating of sodium channels of the DEG/ENaC family (6, 7). In support of this possibility, inactivation of ppk11 or ppk19, two Drosophila DEG/ENaC subunit genes, results in loss of behavioral and electrophysiological responses to salt (8). Here we report the unexpected finding that another Drosophila DEG/ENaC subunit gene, ppk25, is specifically required for male response to courtship-activating female pheromones. This observation suggests that members of this protein family play more diverse roles in chemical senses than previously suspected.
Experimental Procedures
Mutant and Transgenic Flies. Deletions used in this report were generated by imprecise excision of the KG05881 P element (9), scored by loss of the w+ marker and CheB42a expression (10), and sequenced. G7 isogenic control males were generated in the same screen but carry a precise excision of the P element. In Exelixis line e04217, a homozygous lethal mutation closely linked to the Piggyback insertion within ppk25 was separated by meiotic recombination for the generation of homozygous viable flies. The Tg1 and Tg2 rescuing constructs contain genomic fragments starting ≈3.5 kb upstream of the CheB42a transcription initiation site and ending 50 nt downstream of the predicted ppk25 stop codon or immediately downstream of the CheB42a transcription unit, respectively, and were used for generation of transgenic animals by standard methods (11).
Expression Analysis. Analysis of mRNA and protein levels in various tissues was essentially as described (10). Mass separation of body parts results in three fractions: appendages (legs, wings, and third antennal segments), heads (without the third antennal segment), and bodies (without heads, legs, or wings) (12). In Fig. 3b, heads were manually separated from bodies and each fraction was frozen and sieved separately, yielding one sample with third antennal segments and another with legs and wings. Real-time PCR was performed on a DNA Engine Opticon cycler (MJ Research, Waltham, MA), using TaqMan primers that hybridize specifically to ppk25 or rp49 cDNA sequences, as well as appropriate amplification primers. The specificity of the assays was confirmed by the amplification of a single reverse transcriptase-dependent band of the correct size and, for ppk25, by the absence of amplification product or fluorescent signal from Δ5-22 homozygous males. For each sample, the concentration of ppk25 mRNA was obtained by comparison with a standard curve and normalized to that of rp49 mRNA. Sequencing of the cDNA product corresponding to the largest ppk25 hybrid mRNA expressed in Δ5-22 homozygous males showed that it includes intron 3 and lacks intron 4.
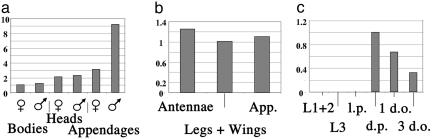
Fig. 3.
ppk25 is expressed in adult appendages involved in taste and smell. Real-time PCR was performed on cDNA prepared from RNA as follows. (a) RNA extracted from male or female body parts as in Fig. 2. In three independent experiments, expression of ppk25 was higher in males than female appendages with an average ratio of 2.4 ± 0.46 (standard error). (b) RNA extracted from single types of male appendages. In three independent experiments, expression in antennae was within a factor of two of that found in combined legs and wings. App., appendages. (c) RNA extracted from whole animals at specific developmental stages. In two independent experiments, ppk25 expression was observed in dark pupae and young adults but not larvae or light pupae. L1 + 2, first and second instar larvae; L3, third instar larvae; l.p., light pupae; d.p., dark pupae; 1 d.o. and 3 d.o., 1- and 3-day-old adults, respectively. In a and b, the relative concentration of ppk25 mRNA is obtained by dividing the normalized value for each sample (see Experimental Procedures) by the lowest value observed in the same experiment (for example, in a, female bodies are set at 1). In c, because the expression level of ppk25 in larvae and light pupae is below detection, the highest sample (dark pupae) was set at 1.
Behavioral Analysis. Flies were raised at 25°C, 50% relative humidity, and courtship behavior was recorded and analyzed essentially as described (13). For courtship analysis, virgin yw female flies were aged for 2-5 days and decapitated 1-2 h before the experiment to eliminate female behavior as a source of variation (14). Virgin males of each genotype aged in isolation for 2-5 days were placed in the presence of a decapitated female inside a solid Plexiglas chamber (7 mm diameter × 7 mm deep), and their behavior was recorded for 10 min by using a digital 8-mm camera with infrared capturing capability. Behaviors were scored blind and analyzed by using a recent version of the lifesong software (15): lifesong x (version 0.51-r2). Statistical significance was calculated by using ANOVA. Geotactic behavior of flies of each genotype was scored by using a geotaxis maze apparatus as described (16). Response to 0.2 mM sucrose was measured in a preference assay that compares ingestion of 0.2 mM sucrose and water (17).
Results
ppk25, a Drosophila Member of the DEG/ENaC Sodium Channel Subunit Family, Is Preferentially Expressed in Male Appendages Rich in Chemosensory Sensilla. We have previously reported that CheB42a, a member of a recently discovered family of Drosophila proteins, is only expressed in a small subset of gustatory sensilla on the front legs of males, suggesting that it may be involved in male-specific gustatory perception (10). Subsequently, Gr68a, a gustatory receptor gene, was found to be expressed in a similar pattern (5). Excitingly, the loss of male response to female pheromones upon either inactivation of Gr68a-expressing neurons or knock-down of Gr68a expression suggests that Gr68a may be a receptor for female pheromones that activate male courtship behavior. More recently, we have found that CheB42a and Gr68a are expressed in the same subset of gustatory sensilla on male front legs (unpublished data), suggesting that CheB42a also plays a role in this process. Intriguingly, ppk25, a gene predicted to encode another protein with a function in chemical senses, is found only 103 nt downstream of the 3′ end of CheB42a (Fig. 1a). The ≈30 members of the Drosophila ppk family of genes (8, 18-21) are part of the large family of DEG/ENaC sodium channel subunits that is found in all animals, from Caenorhabditis elegans to humans, and is involved in a wide variety of functions (22, 23). Several Drosophila ppks are expressed in gustatory neurons, and ppk11 and ppk19 are required for gustatory response to salt (8).
Fig. 1.
The CheB42a and ppk25 genes: transposon insertions and deletions. (a) Map showing the genomic region that includes the CheB42a and ppk25 genes [modified from Flybase (46), http://flybase.org]. The insertion sites for two transposable elements used in this study, a P-element in line KG05881 (9) and a Piggyback element in line e04217 (28), are indicated by arrows. The CheB42a transcription unit is based on the sequence of the corresponding cDNA (10). The predicted transcription unit of ppk25 is supported by the sequence similarity of the predicted protein to other Drosophila Ppks (8), and the position of the introns confirmed by direct sequence of RT-PCR products (data not shown). (b) The endpoints of the Δ5-68, Δ5-2, and Δ5-22 deletions as well as the presence of a partial P element remaining in Δ5-22 are indicated. (c) Cartoon comparing the structural domains present in a wild-type Ppk25 protein and the truncated Ppk25PB predicted to result from insertion of a Piggyback transposable element in the second intron of the ppk25 gene (see text). TM1 and TM2, transmembrane domains 1 and 2; ICD, intracellular domain; ECD, extracellular domain.
To evaluate a possible involvement of ppk25 in male response to pheromones, we tested ppk25 expression in pooled adult appendages that are highly enriched for gustatory (legs and wings) and olfactory (third antennal segment) sensory hairs (24), as well as in body parts that have much fewer chemosensory cells relative to their total mass: heads (without third antennal segment) and bodies (without heads or appendages, see Experimental Procedures). mRNA was isolated from all three types of body parts and analyzed by Northern blot using a full-length ppk25 cDNA probe (Fig. 2). Remarkably, hybridization is by far the strongest to mRNAs from the appendages fraction, yielding a set of bands between 2.1 and 2.4 kb in size, consistent with the predicted ppk25 transcript. Upon longer exposure of the filter, mRNAs of identical sizes, but much lower abundance, are detected in both head and body fractions. Probing the same filter with CheB42a sequences reveals a much smaller mRNA of ≈700 nt that is present only in the appendages fraction, as reported (10).
Fig. 2.
ppk25 mRNA is preferentially expressed in appendages highly enriched in chemosensory organs. mRNA extracted from appendages (third antennal segment, legs and wings), heads (without third antennal segment), and bodies (without heads, legs or antennae) was analyzed on a Northern blot that was hybridized with a full-length ppk25 cDNA probe and exposed for 48 h (Left) or six days (Center Left). The same filter was subsequently boiled and rehybridized with a CheB42a probe. A third hybridization with a probe for the ubiquitous rp49 mRNA (47) shows that somewhat less “Appendages” mRNA was loaded compared to the other body parts so that preferential ppk25 expression in appendages is underrepresented. A, appendages; B, bodies; H, heads.
To determine which appendages express ppk25 mRNA, we used quantitative real-time RT-PCR (Fig. 3) on total RNA extracted from different types of male appendages. This analysis confirms that, in adults, ppk25 mRNA is most abundant in appendages and also shows that ppk25 expression is approximately three times higher in male than female appendages (Fig. 3a). Interestingly, however, ppk25 mRNA is present at equivalent levels in male legs and wings, appendages that carry many gustatory sensilla, and in the third antennal segment, the main olfactory organ of the fly (Fig. 3b). Finally, ppk25 mRNA is not detectable in larvae or at early pupal stages (light pupae), but first appears at late pupal stages (dark pupae), and persists for at least 3 days after eclosion, by which time males are sexually mature (1) (Fig. 3c).
Together, these data show that ppk25 expression is highest in olfactory and gustatory appendages of sexually mature males, a distribution consistent with a role in response to female pheromones. In addition, two observations argue that, despite their proximity, CheB42a and ppk25 are indeed two separate genes that are independently transcribed into two separate mRNAs. First, we find no evidence of any transcript containing both CheB42a and ppk25 sequences (Fig. 2). Second, the two mRNAs have related, but not identical, tissue distributions. Both transcripts are present at highest levels in male appendages. However, whereas CheB42a expression is only detectable in male front legs (10), ppk25 mRNA is expressed equally in male legs and antennae and at lower, but significant levels in female appendages as well as bodies and heads of either sex.
Deletion of ppk25, but Not CheB42a, Dramatically Decreases Male Response to Female Pheromones. To test the possible involvement of CheB42a and ppk25 in male response to female pheromones, we generated three homozygous viable deletions in the region by imprecise excision of a P element inserted ≈1 kb upstream of CheB42a (9) (Fig. 1b). All three deletions remove part or all of the CheB42a gene, leading to the complete absence of CheB42a mRNA (not shown) and CheB42a protein (Fig. 4a). In contrast, these three deletions have very different effects on ppk25. Males homozygous for Δ5-68, a deletion removing all sequences between the P insertion site and roughly the middle of the CheB42a gene, have normal or even slightly increased levels of ppk25 mRNA in their appendages (Fig. 4b). The Δ5-2 deletion completely removes the CheB42a gene and terminates only 59 bp before the ppk25 ATG initiation codon. Although this deletion preserves the predicted ppk25 coding region in its entirety, it significantly impairs transcription of ppk25, such that ppk25 mRNA in male appendages is undetectable by Northern blot. Finally, in addition to deleting all sequences between the P-element insertion site and the midpoint of ppk25, the Δ5-22 deletion retains part of the original transposon, resulting in a series of hybrid transcripts that originate in P-element sequences but retain the 3′ half of the normal ppk25 mRNA. Characterization of the corresponding cDNAs indicates that these aberrant transcripts are unlikely to produce any Ppk25-related polypeptide and suggest that Δ5-22 is a null mutant for ppk25 (see Fig. 4).
Fig. 4.
Three deletions that remove part or all of the CheB42a gene have differential effects on ppk25 expression. (a) Western blot of extracts from the front legs of males of each genotype using an anti-CheB42a antibody (10). The CheB42a protein is absent in all three deletions. (b) Δ5-68, Δ5-2, and Δ5-22 have differential effects on ppk25 mRNA. Poly(A)+ mRNA was extracted from the appendages of male flies homozygous for each of the deletions and a control, G7, and analyzed on a Northern blot that was sequentially probed with radiolabelled full-length ppk25 (Upper) and rp49 (Lower) cDNAs. RT-PCR experiments confirm that the mRNAs expressed in Δ5-22 males initiate within the P-element sequences that remain in that deletion and proceed through the remaining ppk25 sequences (not shown). Given that Δ5-22 retains normal sequences up to 70 bp upstream of the 5′ splice site for the third intron of ppk25, we were surprised to find that this deletion specifically disrupts splicing of intron 3 but not that of intron 4 (Experimental Procedures). Importantly, although these hybrid mRNAs contain sequences encoding ppk25 C-terminal residues, retention of intron 3 disrupts all but 23 aa of the remaining ppk25 ORF within a poorly conserved stretch of the Ppk25 extracellular domain. Furthermore, the ATG that initiates this residual ppk25 ORF is unlikely to function as an initiation of translation because it follows, by 25 nt, another ATG that has a better match to the Kozak consensus translation initiation site (48) (data not shown). Together, these results suggest that no Ppk25-related peptide is produced in Δ5-22 homozygous flies.
Is the function of either CheB42a or ppk25 required for male response to female pheromones? When placed in the presence of a female, a D. melanogaster male quickly initiates a striking series of stereotyped steps that include following the female, tapping her with his front legs, generating a courtship song by vibrating one of his wings, licking her genitalia, attempting copulation, and copulating (1). Both visual and chemosensory perception of the female stimulate male courtship behavior. Therefore, we observed the response of males carrying deletions in the CheB42a/ppk25 region to females under infrared lights, which D. melanogaster cannot detect (25), to enhance the contribution of pheromone detection to male behavior. For each male, a courtship index is calculated (26), which represents the fraction of the total observation time spent performing any courtship behavior multiplied by 100 (Fig. 5a). Males homozygous for the Δ5-68 deletion display normal levels of overall courtship. In contrast, males homozygous for either Δ5-2 or Δ5-22 have a much reduced courtship index relative to the G7 controls (P < 9 × 10-4 and P < 2 ×10-4 for Δ5-2 and Δ5-22, respectively), suggesting unexpectedly that males require ppk25, but not CheB42a, to achieve normal overall levels of courtship behavior in response to a female. In addition, introduction of a transgenic copy of the genomic region that spans both CheB42a and ppk25 genes rescues the courtship behavior of Δ5-22 homozygous males, whereas an almost identical transgene that lacks ppk25 does not (Tg1 and Tg2, respectively, in Fig. 5b). This result indicates that the courtship deficit of Δ5-22 homozygous males is indeed caused by the loss of ppk25. Importantly, ppk25 is not required for two behaviors unrelated to courtship: walking and preening (Fig. 5). In fact, males homozygous for Δ5-22 walk more than controls or those carrying a transgenic ppk25, whereas Δ5-2 homozygous males display normal levels of this behavior. To further test whether Δ5-2 and Δ5-22 cause generalized brain dysfunction, we measured two other complex behavioral responses to sensory stimuli. Neither the typical climbing response of Drosophila to mechanosensory detection of gravity nor stimulation of food intake by gustatory detection of sucrose is affected by any of the deletions in the region (Fig. 8, which is published as supporting information on the PNAS web site). Together, these results suggest that ppk25 is required specifically for male response to females.
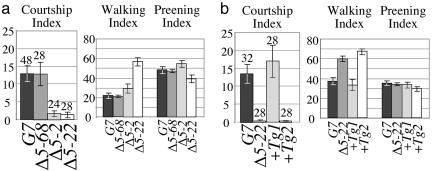
Fig. 5.
Male response to females is debilitated by deletions that remove or prevent expression of ppk25, and is restored by a ppk25-carrying transgene. The response of males of different genotypes to females was quantitated by a courtship index: the fraction of the observation time spent performing any step in the courtship sequence multiplied by 100 (26), and similar indices measure the time spent walking and preening. (a) Courtship response is dramatically reduced in males homozygous for Δ5-2 or Δ5-22, but not Δ5-68, relative to G7 isogenic control males (P < 9 × 10-4 and P < 2 × 10-4 for Δ5-2 and Δ5-22, respectively). (b) Introduction of Tg1, a transgene carrying the genomic region that includes CheB42a and ppk25, rescues the courtship response of Δ5-22 homozygous males (+Tg1), whereas Tg2, an almost identical transgene that lacks ppk25 (+Tg2), does not. Error bars indicate standard error of the mean and n for each genotype is indicated immediately above.
Insertion of a Transposable Element into the Second Intron of ppk25 Causes a Dominant-Negative Decrease of Male Response to Females. Analysis of our deletion lines suggests that the ppk25 gene is required for normal male response to females. As a further test of this possibility, we used males with an independent mutation in ppk25 (27, 28). In this ppk25 mutant, a transposable element is inserted in the second intron of the ppk25 gene, resulting in what we will refer to as the ppk25PB allele (Fig. 1 a and c). The presence of 4 kb of extraneous sequences, including a termination site from the miniwhite gene (28), make it unlikely that this modified ppk25 intron 2 can be spliced properly to produce functional ppk25 mRNA. Instead, transcription from the normal ppk25 promoter can be expected to result in an mRNA that retains exons 1 and 2 followed by part of intron 2, which contains multiple in-frame stop codons. Alternatively, the 5′ splice junction of intron 2 may be spliced aberrantly to a cryptic 3′ splice site within Piggyback sequences. The protein product of ppk25PB should therefore be limited to the first transmembrane domain and part of the extracellular domain of Ppk25, perhaps fused to Piggyback sequences (Fig. 1c). Interestingly, for several other DEG/ENaC genes, similarly truncated or fused proteins that retain the first transmembrane domain have dominant-negative properties likely caused by the formation of nonfunctional complexes with other DEG/ENaC subunits or other interacting proteins (8, 21, 29, 30, 45).
To test the effect of the ppk25PB allele on male response to females, we generated males that carry the following mutations: (i) ppk25PB, (ii) CPB a similar Piggyback insertion in an unrelated site on the second chromosome in an otherwise isogenic background to ppk25PB, (iii) Δ42E, a deletion of the ppk25 genomic region spanning ≈100 kb and 20 genes, or (iv) CΔ, a deletion of similar size in an unrelated area of the second chromosome (Fig. 6). Remarkably, none of the 28 ppk25PB/Δ42E males that we tested displayed any detectable courtship behavior during the 10-min observation period under infrared lights, a highly significant decrease relative to control males (compare the courtship index for ppk25PB/Δ42E and CPB/Δ42E males in Fig. 6, P = 3 × 10-5). This result confirms that ppk25 is required for male response to females. In addition, because ppk25PB homozygous males have normal levels of CheB42a mRNA (data not shown), the result indicates that the requirement for ppk25 is independent of CheB42a. Finally, the complete loss of male response to females in ppk25PB/Δ42E males is a significantly more severe phenotype than the reduced courtship observed for Δ5-22 homozygous males, suggesting that ppk25PB is indeed a dominant-negative allele. This conclusion is validated by the significantly reduced levels of courtship behavior exhibited by males that carry a single copy of ppk25PB in the presence of a wild-type ppk25 gene compared to males that only carry one wild-type copy of ppk25 (compare ppk25PB/CΔ to CPB/Δ42E in Fig. 6, P = 0.012).
Fig. 6.
The ppk25PB allele has a dominant-negative effect on male response to female pheromones. The male response to females was measured as in Fig. 5. The males tested carry the following mutations: ppk25PB, Piggyback insertion into ppk25 (line e04217); CPB, control with a normal ppk25 gene and the same Piggyback element inserted at an unrelated site on the second chromosome (line e00673); Δ42E, a deletion spanning 20 genes in the ppk25 region [Df(2R)Exel6051]; CΔ, a control deletion in an unrelated region on the second chromosome that retains a normal ppk25 gene (Df(2R)ED1552). *1 and *2, P = 3 × 10-5 and P = 0.012 for comparisons of the control CPB/Δ42E to ppk25PB/Δ42E and ppk25PB/CΔ, respectively.
Visible Light Completely Alleviates the Block of Mutant Males Carrying the ppk25PB Allele on the Initiation, but Not Maintenance of Courtship Behavior. The deficient male response to females observed for ppk25 loss-of-function and dominant-negative alleles under infrared light could be due either to a lack of sensory detection of females or to a more general inability to perform courtship behaviors, regardless of sensory stimulus. To distinguish between these two possibilities, we analyzed the effect of visible light on the response of ppk25PB mutant males. The two types of males we compared in this experiment carry a single copy of either the wild-type ppk25 gene or the dominant-negative ppk25PB allele in an otherwise isogenic background that includes the Δ5-22 deletion. As in Fig. 6, under infrared light and in the absence of any wild-type ppk25, a single copy of the dominant-negative ppk25PB allele results in the complete loss of male response to females under infrared light, but no decrease in walking or preening (not shown). In sharp contrast, in the presence of visible light, males of the same genotype perform all of the normal steps of courtship, albeit at a significantly reduced rate (Fig. 9, which is published as supporting information on the PNAS web site). This result suggests that the complete inability of males carrying the dominant-negative ppk25PB allele to respond to females under infrared lights is due to a lack of sensory input rather than an inability to perform courtship behaviors. Furthermore, this experiment provides an indirect test of whether the dominant-negative ppk25PB mutation blocks pheromone perception through olfaction, gustation, or both chemical senses. Both visual and olfactory inputs can initiate courtship behavior. In contrast, gustatory perception of pheromones may only be required for efficient performance of subsequent steps (5). Because the lag to initiation of courtship behavior and the number of courtship bouts per second displayed by ppk25PB/Δ5-22 males are similar to controls in the presence of visible light, the lack of response of the same males under infrared lights likely results at least in part from their inability to initiate courtship in response to pheromones as would be expected for an olfactory defect. On the other hand, despite the presence of visible lights, the average length of a courtship bout for mutant males is less than half that of controls, suggesting that the ppk25PB mutation also affects a subsequent step, perhaps gustatory detection of pheromones. ppk25PB's dominant negative properties in the absence of wild-type ppk25 are most likely due to interactions between the truncated protein and other factors involved in pheromone perception. However, the decreased levels of courtship behavior displayed by males homozygous for the Δ5-2 or Δ5-22 deletions (Fig. 5) also result from a combination of increased lags to courtship, decreased numbers of bouts initiated per seconds, and shorter bout lengths (data not shown). Together, these results suggest that ppk25 itself is required for both initiation and maintenance of courtship bouts in response to female pheromones.
Discussion
A Member of the Drosophila Family of DEG/ENaC Sodium Channel Subunits Is Required for Male Response to Females. We have found that ppk25, a member of the Drosophila family of DEG/ENaC sodium channel subunits, is required for male response to females. First, we have generated two deletions that inactivate both CheB42a and ppk25: Δ5-2 and Δ5-22. Males homozygous for either deletion display a much reduced response to females but no similar decrease in other behaviors. In contrast, another deletion that results in complete loss of CheB42a expression but has no effect on ppk25 does not reduce male courtship behavior. Second, a genomic fragment that includes both CheB42a and ppk25 rescues the response of Δ5-22 homozygous males to females, whereas an almost identical fragment lacking ppk25 does not. Third, ppk25PB, an independent mutation resulting from insertion of a transposable element into the second intron of ppk25, affects male response to females even more severely than Δ5-22, even though this allele has no detectable effect on CheB42a expression. Indeed, ppk25PB has dominant-negative effects on male response to females, observable both in the presence or absence of a wild-type copy of ppk25. Fourth, the dominant-negative properties of ppk25PB are readily interpreted in light of the predicted generation in this mutant of a truncated Ppk25 protein retaining the N-terminal cytoplasmic domain, the first transmembrane domain, and part of the extracellular domain of the normal Ppk25. Similarly truncated variants of various members of the DEG/ENaC family, including several Drosophila ppks, also have dominant-negative properties (8, 21, 29, 30, 45).
Our discovery of a role for ppk25 in male response to female pheromones was the unexpected result of our interest in the neighboring CheB42a. The data in this report show that deletion of CheB42a does not decrease overall male response to courtship-activating pheromones. However, the restricted expression of CheB42a in the same subset of gustatory sensilla that express Gr68a (unpublished data) and are required for response to female courtship-activating pheromones (5) suggest that CheB42a's requirement may be obscured by functional redundancy with one or more the other 11 Drosophila CheB genes (10) or, alternatively, that CheB42a has a different role in male-specific chemical senses.
Is it a coincidence that two genes implicated in male-specific chemical senses are within <103 nt of each other? These two genes produce mRNAs of different sizes with related, albeit different, expression patterns. Both are preferentially expressed in male gustatory appendages starting late in pupal development and remaining through at least sexual maturity of the adult males. However, whereas CheB42a is only expressed in male front legs (10), ppk25 mRNA is present at similar levels in legs and in the third antennal segment, and at much lower but detectable levels in heads and bodies. The proximity of these two genes may therefore reflect a shared dependence on regulatory elements important for overlapping spatial and/or temporal characteristics of their expression. Indeed, the lack of detectable ppk25 mRNA in males homozygous for Δ5-2 suggests the presence of a regulatory element essential for ppk25 expression within or immediately downstream of the 3′ half of CheB42a. Alternatively, the proximity between these two genes may be more a reflection of their involvement in evolutionarily important and related aspects of sexual behavior.
ppk25 Is Required for Chemosensory Activation of Male Courtship Behavior by Female Pheromones. Why can't ppk25 mutant males respond to females normally? Vision and pheromone detection have both been implicated in the response of Drosophila melanogaster males to females (1, 2). Absence of visible light or mutations that cause partial or complete blindness reduce, but do not eliminate, male response to females. In addition, a number of studies suggest that males detect courtship-stimulating female pheromones by using either gustation, olfaction, or both chemical senses (5, 31-40). Although both vision and olfactory detection of pheromones are important for initiation of courtship behavior, gustatory perception of the same or other pheromones may be required for efficient progression to later steps in the courtship sequence (5). Because both initiation and maintenance of courtship bouts are affected in dominant-negative (Figs. 6 and 7) as well as null mutations in ppk25 (not shown), this gene may be required for detection of pheromones by both sensory modalities, a possibility supported by the expression of ppk25 in both olfactory (antennae) and gustatory (wings and legs) appendages.
Fig. 7.
Visible light enables courtship behavior in males carrying the dominant-negative ppk25PB allele. Male response to females was measured as in Fig. 5 except for the presence of visible light. For this experiment, three separate parameters of male behavior are shown to demonstrate the differential effect of the dominant-negative ppk25PB allele: lag to courtship, number of courtship bouts per minute, and length of courtship bouts (see text). The males tested carry a Δ5-22 deletion on one copy of the second chromosome and are completely isogenic except for the presence on their other second chromosome of either (i) the dominant-negative ppk25PB allele, or (ii) the normal ppk25 gene and another Piggyback insertion at an unrelated site. In the presence of visible light, replacement of the normal ppk25 gene by the ppk25PB allele causes a statistically significant decrease in the average length of a courtship bout (P < 0.02) but no change in the lag to courtship or in the number of courtship bouts per minute.
Is a Ppk25-Containing Sodium Channel Involved in the Peripheral Detection of Female Pheromones? Our data strongly support the requirement for ppk25 in the male's ability to respond to female courtship-activating pheromones. In addition, mutations in ppk25 do not similarly impair other behaviors that are either largely independent of sensory inputs, such as walking and preening, or sensory-driven such as geotaxis and chemosensory response to sugars. Most importantly, these mutations have no effect on the initiation of courtship behavior in the presence of visible light. Therefore, ppk25's requirement for male response to pheromones likely reflects a specific role in the sensory detection of pheromones or subsequent processing within the central nervous system rather than a more general requirement for neural function or even for performance of courtship behavior. Finally, ppk25 expression is first detectable during late pupal stages, after determination of all of the various types of chemosensory cells and as they undergo the final stages of differentiation (41, 42), suggesting that ppk25 is required for the function, rather than the development of chemosensory organs.
Is ppk25 required in peripheral olfactory or gustatory neurons that sense and respond to female pheromones in the environment, or in central nervous system neurons that receive and process the information coming from the periphery? Although these alternatives remain to be tested, the former hypothesis is supported by ppk25's preferential expression in male chemosensory appendages as well as by the established roles of other DEG/ENaC subunits in peripheral sensory responses to mechanical stimuli (43) and salt (8). ppk25's putative role in pheromone detection may not involve direct participation in the primary molecular response to pheromones. However, recent imaging of the electrophysiological response in mechanosensory neurons indicate that the C. elegans DEG/ENaC gene mec-4 is specifically required for the mechanosensory function rather than the general physiology of the neurons in which it is expressed (44). Similar questions arise regarding the role ppk25 plays in male detection of female pheromones and in particular, whether it interacts, directly or indirectly, with the G protein-coupled signal transduction pathways that underlie chemical senses in Drosophila as in other animals (4).
Finally, the dominant-negative properties of the ppk25PB allele most likely reflect the participation of the Ppk25 protein in a multisubunit protein complex. Proteins of the DEG/ENaC family are thought to interact in the formation of heteromeric sodium channels (22, 23). Several truncated versions of DEG/ENaC proteins have dominant-negative properties that most likely result from their ability to form partial and inactive complexes with other DEG/ENaC subunits (8, 29, 30, 45). By analogy, our results suggest that one or more of the ≈30 other Ppk proteins encoded in the Drosophila genome (8) interacts with Ppk25 within a heteromeric sodium channel.
In conclusion, our data demonstrate a role for a member of the DEG/ENaC family of sodium channel subunits in the peripheral detection or central processing of a pheromonal signal. This finding opens the door to the dissection of ppk25's role in pheromone response and its relationship with other proteins involved in pheromone response. Finally, this work suggests that members of the Drosophila ppk family, as well as DEG/ENaC subunits in other organisms, play more complex roles in chemical senses than previously suspected.
Supplementary Material
Acknowledgments
We are very grateful to Jeff Hall, John Rieffel, and Adriana Villella (Brandeis University, Waltham, MA) for generously providing and assisting in the use of the lifesong x software; to Yashi Ahmed for numerous invaluable suggestions during the completion of this research; and to Adriana Villella, Stephen Goodwin, and Yashi Ahmed for thoughtful comments on the manuscript. Fly lines were made freely available by the Exelixis collection at Harvard Medical School, the Drosophila Genome Project, the Drosdel Project, and the Bloomington Drosophila Stock Center at Indiana University. This work was supported by National Institute on Deafness and Other Communication Disorders/National Institutes of Health Grant RO1DC04284 (to C.W.P.).
Author contributions: H.L., K.J.M., E.S., R.D.K., and C.W.P. designed research; H.L., K.J.M., E.S., and R.D.K. performed research; H.L., K.J.M., E.S., and R.D.K. contributed new reagents/analytic tools; H.L., K.J.M., E.S., R.D.K., and C.W.P. analyzed data; and C.W.P. wrote the paper.
References
- 1.Hall, J. C. (1994) Science 264, 1702-1714. [DOI] [PubMed] [Google Scholar]
- 2.Greenspan, R. J. & Ferveur, J. F. (2000) Annu. Rev. Genet. 34, 205-232. [DOI] [PubMed] [Google Scholar]
- 3.Amrein, H. (2004) Curr. Opin. Neurobiol. 14, 435-442. [DOI] [PubMed] [Google Scholar]
- 4.Mombaerts, P. (2004) Nat. Rev. Neurosci. 5, 263-278. [DOI] [PubMed] [Google Scholar]
- 5.Bray, S. & Amrein, H. (2003) Neuron 39, 1019-1029. [DOI] [PubMed] [Google Scholar]
- 6.Lin, W., Finger, T. E., Rossier, B. C. & Kinnamon, S. C. (1999) J. Comp. Neurol. 405, 406-420. [DOI] [PubMed] [Google Scholar]
- 7.Lin, W., Ogura, T. & Kinnamon, S. C. (2002) J. Neurophysiol. 88, 133-141. [DOI] [PubMed] [Google Scholar]
- 8.Liu, L., Leonard, A. S., Motto, D. G., Feller, M. A., Price, M. P., Johnson, W. A. & Welsh, M. J. (2003) Neuron 39, 133-146. [DOI] [PubMed] [Google Scholar]
- 9.Bellen, H. J., Levis, R. W., Liao, G., He, Y., Carlson, J. W., Tsang, G., Evans-Holm, M., Hiesinger, P. R., Schulze, K. L., Rubin, G. M., et al. (2004) Genetics 167, 761-781. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Xu, A., Park, S. K., D'Mello, S., Kim, E., Wang, Q. & Pikielny, C. W. (2002) Cell Tissue Res. 307, 381-392. [DOI] [PubMed] [Google Scholar]
- 11.Rubin, G. M. & Spradling, A. C. (1982) Science 218, 348-353. [DOI] [PubMed] [Google Scholar]
- 12.Oliver, D. V. & Philips, J. P. (1970) Dros. Inf. Serv. 45, 58. [Google Scholar]
- 13.Villela, A. & Hall, J. C. (1996) Genetics 143, 331-344. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Grossfield, J. (1972) Anim. Behav. 20, 243-251. [Google Scholar]
- 15.Bernstein, A. S., Neumann, E. K. & Hall, J. C. (1992) J. Insect Behav. 5, 15-35. [Google Scholar]
- 16.Toma, D. P., White, K. P., Hirsch, J. & Greenspan, R. J. (2002) Nat. Genet. 31, 349-353. [DOI] [PubMed] [Google Scholar]
- 17.Tanimura, T., Isono, K., Takamura, T. & Shimada, I. (1982) J. Comp. Phys. 147, 433-437. [Google Scholar]
- 18.Darboux, I., Lingueglia, E., Champigny, G., Coscoy, S., Barbry, P. & Lazdunski, M. (1998) J. Biol. Chem. 273, 9424-9429. [DOI] [PubMed] [Google Scholar]
- 19.Darboux, I., Lingueglia, E., Pauron, D., Barbry, P. & Lazdunski, M. (1998) Biochem. Biophys. Res. Commun. 246, 210-216. [DOI] [PubMed] [Google Scholar]
- 20.Adams, C. M., Anderson, M. G., Motto, D. G., Price, M. P., Johnson, W. A. & Welsh, M. J. (1998) J. Cell Biol. 140, 143-152. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Liu, L., Johnson, W. A. & Welsh, M. J. (2003) Proc. Natl. Acad. Sci. USA 100, 2128-2133. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Bianchi, L. & Driscoll, M. (2002) Neuron 34, 337-340. [DOI] [PubMed] [Google Scholar]
- 23.Kellenberger, S. & Schild, L. (2002) Physiol. Rev. 82, 735-767. [DOI] [PubMed] [Google Scholar]
- 24.Stocker, R. F. (1994) Cell Tissue Res. 275, 3-26. [DOI] [PubMed] [Google Scholar]
- 25.Frank, K. D. & Zimmerman, W. F. (1969) Science 163, 688-689. [DOI] [PubMed] [Google Scholar]
- 26.Hall, J. C. (1977) Behav. Genet. 7, 291-312. [DOI] [PubMed] [Google Scholar]
- 27.Parks, A. L., Cook, K. R., Belvin, M., Dompe, N. A., Fawcett, R., Huppert, K., Tan, L. R., Winter, C. G., Bogart, K. P., Deal, J. E., et al. (2004) Nat. Genet. 36, 288-292. [DOI] [PubMed] [Google Scholar]
- 28.Thibault, S. T., Singer, M. A., Miyazaki, W. Y., Milash, B., Dompe, N. A., Singh, C. M., Buchholz, R., Demsky, M., Fawcett, R., Francis-Lang, H. L., et al. (2004) Nat. Genet. 36, 283-287. [DOI] [PubMed] [Google Scholar]
- 29.Hong, K., Mano, I. & Driscoll, M. (2000) J. Neurosci. 20, 2575-2588. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Bruns, J. B., Hu, B., Ahn, Y. J., Sheng, S., Hughey, R. P. & Kleyman, T. R. (2003) Am. J. Physiol. 285, F600-F609. [DOI] [PubMed] [Google Scholar]
- 31.Shorey, H. & Bartell, R. (1970) Anim. Behav. 18, 159-164. [DOI] [PubMed] [Google Scholar]
- 32.Averhoff, W. & Richardson, R. (1976) Proc. Natl. Acad. Sci. USA 73, 591-593. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Tompkins, L., Hall, J. C. & Hall, L. M. (1980) J. Insect Physiol. 26, 689-697. [Google Scholar]
- 34.Tompkins, L. & Hall, J. C. (1981) J. Insect Physiol. 27, 17-21. [Google Scholar]
- 35.Anthony, C. & Jallon, J.-M. (1982) J. Insect Physiol. 28, 873-880. [Google Scholar]
- 36.Robertson, H. M. (1983) Experientia 39, 333-335. [Google Scholar]
- 37.Gailey, D. A., Lacaillade, R. C. & Hall, J. C. (1986) Behav. Genet. 16, 375-405. [DOI] [PubMed] [Google Scholar]
- 38.Boll, W. & Noll, M. (2002) Development (Cambridge, U.K.) 129, 5667-5681. [DOI] [PubMed] [Google Scholar]
- 39.Heimbeck, G., Bugnon, V., Gendre, N., Keller, A. & Stocker, R. F. (2001) Proc. Natl. Acad. Sci. USA 98, 15336-15341. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Ejima, A., Smith, B. P., Lucas, C., Levine, J. D. & Griffith, L. C. (2005) Curr. Biol. 15, 194-206. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Ray, K., Hartenstein, V. & Rodrigues, V. (1993) Dev. Biol. 155, 26-37. [DOI] [PubMed] [Google Scholar]
- 42.Ray, K. & Rodrigues, V. (1995) Dev. Biol. 167, 426-438. [DOI] [PubMed] [Google Scholar]
- 43.Tavernarakis, N. & Driscoll, M. (2001) Cell Biochem. Biophys. 35, 1-18. [DOI] [PubMed] [Google Scholar]
- 44.Suzuki, H., Kerr, R., Bianchi, L., Frokjaer-Jensen, C., Slone, D., Xue, J., Gerstbrein, B., Driscoll, M. & Schafer, W. R. (2003) Neuron 39, 1005-1017. [DOI] [PubMed] [Google Scholar]
- 45.Adams, C. M., Snyder, P. M. & Welsh, M. J. (1997) J. Biol. Chem. 272, 27295-27300. [DOI] [PubMed] [Google Scholar]
- 46.Drysdale, R. A. & Crosby, M. A. (2005) Nucleic Acids Res. 33, D390-D395. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.O'Connell, P. O. & Rosbash, M. (1984) Nucleic Acids Res. 12, 5495-5513. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Kozak, M. (1999) Gene 234, 187-208. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.