ABSTRACT
The initial microbial colonization of the infant gut during birth plays a critical role in shaping both immediate and long-term health outcomes. While mode of delivery is a known determinant of this colonization process, the potential impacts of infant sex and birth order remain underexplored. This study investigates the influence of delivery mode, infant sex, and birth order (maternal parity) on the microbial communities in first-pass meconium samples from neonates, using 16S rRNA gene sequencing. We found that delivery mode impacted the presence of detectable microbial communities. Specifically, only 17% of samples from neonates delivered by elective Cesarean section showed any microbial presence, compared to approximately two-thirds of samples from neonates exposed to maternal vaginal microbes (emergency C-section or vaginal delivery). Among vaginally delivered neonates without antibiotic exposure, birth order was associated with taxonomic shifts. Neonates born to primiparous mothers had a lower abundance of Bifidobacterium, a keystone species in the infant gut microbiome. Unexpectedly, the gut microbiota differed by infant sex, with males having lower alpha diversity and shifts in microbial community composition (PERMANOVA p = 0.008), characterized by elevated levels of Enterobacteriales, which was both less prevalent and less abundant in female neonates. These findings highlight the intricate interplay between delivery mode, infant sex, and birth order in shaping the early gut microbiome.
KEYWORDS: Gut microbiota, first-pass meconium, birth order, parity, delivery mode, birth mode, infant sex, neonatal colonization
Background
The infant gut microbiome plays a crucial role in nutrient absorption, immune education, and neurodevelopment. First direct contact with microbes occurs during birth and is a critical step in the process of microbial colonization, influencing both immediate and long-term health outcomes.1 There is a general consensus in the current literature that mode of delivery is a key factor shaping these early-life patterns of colonization. Shortly after birth, maternal microbes can be detected in infant meconium – the first stool passed.2 Maternal skin microbes are more abundant in the meconium of infants born by Cesarean section (C-section),2 whereas maternal vaginal and gut microbes are initially enriched in the meconium of vaginally-delivered infants.3 As only gut-adapted maternal microbes such as Bifidobacteria and Bacteroides persistently colonize the infant gut4 the impact of delivery mode on the infant gut microbiome gradually decreases,5 but its effect can be detected up to at least 2 years of age.6 Even temporary differences in microbial exposures during critical developmental windows in early life can influence long-term health trajectories. Differences in microbial exposure due to delivery mode have been linked to health outcomes, including the risk of developing allergies and autoimmune diseases later in life.1,7
Much less is known about the potential impacts of infant sex and birth order on the gut microbiome generally or on meconium-associated microbes in particular. Data from animal models suggest that sex hormones may play a role in shaping gut microbiome community composition.8 However, few studies have investigated sex differences in the human microbiome before puberty despite known differences in sex hormones early in life.9 Birth order may also impact neonatal sex hormones, with one study showing higher levels of progesterone and estrogen in umbilical cord blood of first-borns of both sexes.10 Recently, we reported that parity impacts the maternal gut microbiome during pregnancy and the infant microbiome at 6 months of age.11 Others have found that sibling number is associated with the composition of the infant gut microbiota.12 As the maternal microbiota is the main source of infant microbes it is possible that the influence of birth-order (or maternal parity) on the infant microbiome is due to differential seeding of the microbiome during birth. However, whether our earliest microbial colonizers differ by birth-order and sex is unknown. To understand the intersection of these factors on neonatal gut colonization, we investigated the effects of infant sex, birth order, and delivery mode on the presence and composition of microbes in first-pass meconium in this study.
Results
First-pass meconium samples from 70 neonates (first-born/primiparous, n = 41; later-born/multiparous, n = 29) were collected within 48 hours of birth. Male and female neonates were similar across all participant characteristics including delivery mode, exposure to peripartum antibiotics, maternal age, gestational age, birth weight, and premature rupture of membrranes (PROM) prevalence (all p > 0.3 when not stratified by birth order; data and p-values stratified by birth order shown in Table 1).
Table 1.
Participant characteristics by infant sex.
| First-born (primiparous) |
Later-born (multiparous) |
|||||
|---|---|---|---|---|---|---|
| Female | Male | p | Female | Male | p | |
| n | 17 | 24 | 14 | 15 | ||
| Maternal age1 | 28.9 (3.7) | 30.4 (4.6) | 0.2 | 32.6 (3.9) | 33.0 (3.0) | 0.8 |
| Gestational age2 | 40+0 (38+5,40+4) |
40+1 (39+3,40+5) |
0.7 | 39+6 (39+1,40+3) |
39+2 (38+4,39+6) |
0.11 |
| Mode of delivery | 0.5 | 0.6 | ||||
| Elective C-section | 1 (5.9%) | 3 (13%) | 1 (7.1%) | 1 (6.7%) | ||
| Emergency C-section | 1 (5.9%) | 2 (8.3%) | 1 (7.1%) | 4 (27%) | ||
| Spontaneous vaginal | 12 (71%) | 11 (46%) | 12 (86%) | 10 (67%) | ||
| Vaginal extraction | 2 (18%) | 8 (33%) | 0 (0%) | 0 (0%) | ||
| Peripartum antibiotics | 3 (18%) | 6 (25%) | 0.7 | 4 (29%) | 5 (33%) | >0.9 |
| Birth weight (g)1 | 3,178 (514) | 3,443 (459) | 0.2 | 3,502 (462) | 3,543 (391) | >0.9 |
| PROM | 3 (18%) | 5 (21%) | >0.9 | 4 (29%) | 2 (13%) | 0.4 |
1Mean (sd); 2median (IQR); Continuous variables analyzed by Mann-Whitney U-test and categorical variables by Fisher’s exact test; Premature rupture of membranes (PROM).
Impact of delivery mode
Consistent with in utero sterility,13 only 44 samples (62.9%) produced any sequencing reads and the proportion of samples with any reads varied significantly by delivery mode, but not by sex or birth order. Elective C-section neonates had the lowest proportion of samples with >0 sequencing reads (17%, Table 2). In contrast, other delivery modes had higher proportions of samples with sequencing reads (63–69%). Of the samples with >0 sequencing reads, the number of reads also varied by delivery mode with the fewest sequencing reads in the one remaining elective C-section neonate and the highest in neonates born by spontaneous vaginal delivery (Table 2).
Figure 1.
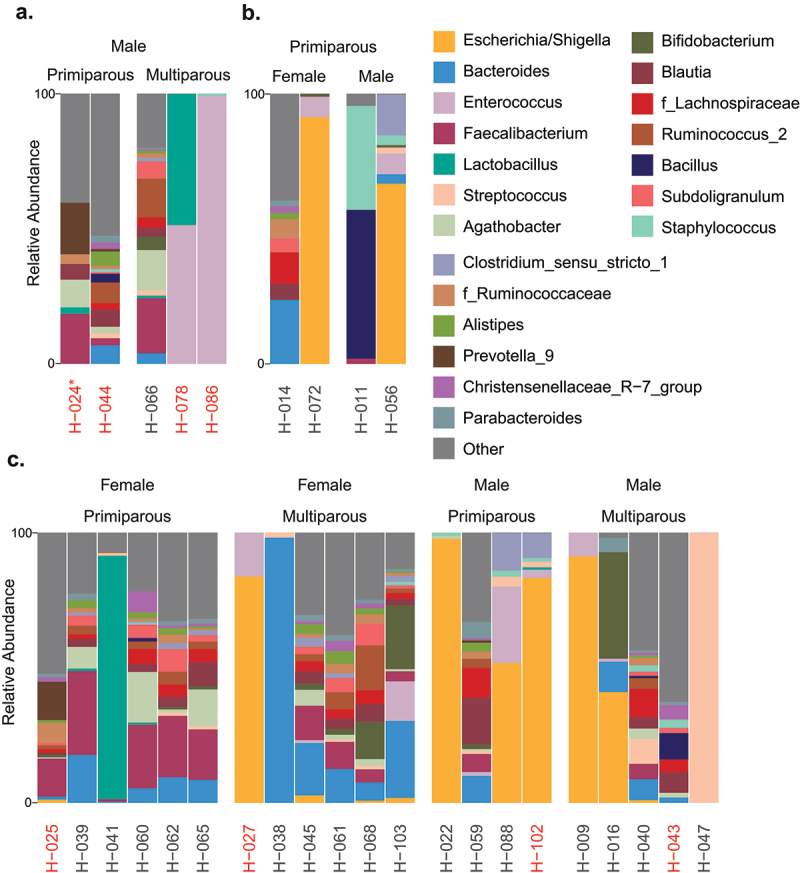
Relative abundance of 20 most abundant bacterial genera in first-pass meconium samples with >100 sequencing reads. (a) Cesarean section (C-section), (b) vacuum extraction, and c) spontaneous vaginal delivery. Neonates exposed to peripartum antibiotics are indicated by red text. One neonate delivered by elective C-section is indicated by an asterisk, all other C-sections are emergency.
Table 2.
Sequencing read count by delivery mode.
| Delivery mode | total n | Samples with >0 sequencing reads |
||
|---|---|---|---|---|
| n (%) | median | mean | ||
| Elective C-section | 6 | 1 (17%) | 172 | 172 |
| Emergency C-section | 8 | 5 (63%) | 21906 | 27744 |
| Spontaneous vaginal | 45 | 31 (69%) | 11353 | 31848 |
| Vaginal extraction | 11 | 7 (64%) | 397 | 14681 |
In samples with read counts >100 (total n = 30; elective C-section, n = 1; emergency C-section; n = 4, normal spontaneous vaginal delivery, n = 21; vacuum extraction, n = 4), taxonomic composition was highly variable (Figure 1). A large proportion of samples (14 of 30 total) were dominated by one or two genera. Of 244 genera detected, only 20 were present in more than half of first-pass meconium samples.
Impact of infant sex and birth order
To avoid the potential confounding effects of delivery mode and intrapartum antibiotic exposure, further analysis was performed using only first-pass meconium samples from spontaneous vaginal deliveries without antibiotic exposure (n = 17). Most females were dominated by taxa belonging to the order Eubacteriales (aka Clostridiales; seven of ten with a relative abundance greater than 65%), while males were more commonly dominated by Enterobacterales (aka Enterobacteriales, four of seven with a relative abundance greater than 41%), which was rare among females (detected in three of ten females; maximum relative abundance 2.1%; Figure 2). Specifically, the Enterobacterales genus Escherichia/Shigella was detected in only three of ten females (mean relative abundance, 0.34%; maximum abundance, 2.1%) but in five of seven males (mean relative abundance, 40.6%) and was the dominant genus in four of the males (minimum, 41.3%; maximum, 98.4%). The mean relative abundance of Bifidobacterium was greater in neonates of multiparous participants (9.2%) compared to those of primiparous participants (0.30%). This trend was also observed for Bacteroides, particularly in females (multiparous, 33.7%; primiparous, 7.8%), who had a greater mean relative abundance compared to males (female, 20.8%; male 4.3%).
Figure 2.
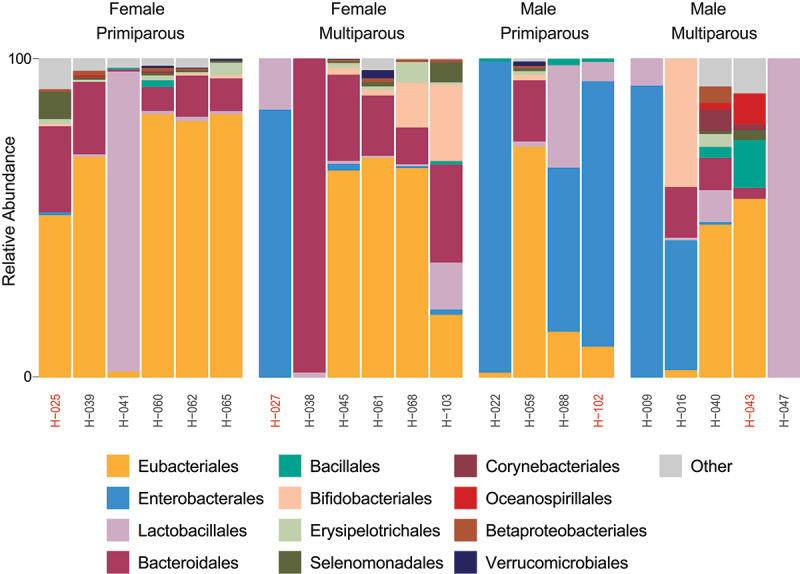
Relative abundance of 20 most abundant bacterial orders in meconium samples of spontaneously vaginally delivered neonates differ by parity and sex. Individual sample identifiers are noted below taxonomic bars. Neonates with peripartum antibiotic exposure are indicated by red text.
In terms of alpha diversity, the number of observed ASVs (species richness) was lower in male neonates (mean = 73.4, SD = 109.4) compared to females (mean = 197.1, SD = 120.3; p = 0.047; Figure 3) but Shannon diversity (a measure of both species richness and evenness) did not differ significantly by infant sex (p = 0.10). Birth order did not significantly impact either the number of observed ASVs (p = 0.91; first-born, mean = 150.0, SD = 113.8; later born mean = 142.8, SD = 147.6) or Shannon diversity (p = 0.75; first-born, mean = 2.86, SD = 1.61; later born mean = 2.57, SD = 1.97). Overall community composition (Beta diversity: Bray-Curtis dissimilarity) differed by infant sex (R2 = 14.9, p = 0.008) but was similar between birth order groups (p = 0.70, Figure 3). Together, these data suggest that, in addition to the large effect of delivery mode, parity and infant sex may also impact the initial colonization of the infant gut microbiota.
Figure 3.
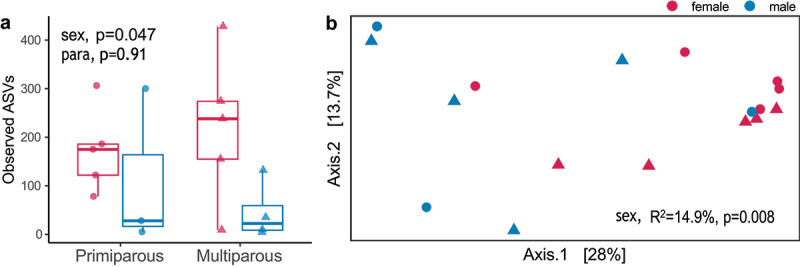
Overall community composition varies by infant sex. (a) Alpha diversity, measured by the total number of ASVs detected in each sample, was significantly lower in male (n = 7) neonates compared to females (n = 10). Mean ± SD for first-born females (173.4 ± 85.9, n = 5), first-born males (111.0 ± 164.1, n = 3), later born females (220.8 ± 154.3, n = 5), and later born males (45.25 ± 59.4, n = 3). Significance assessed by linear regression (lm function in the stats package in R). (b) Beta diversity differed significantly by infant sex (R2 = 14.9%, p = 0.008) but not by parity (R2 = 0.05, p = 0.70). Significance assessed by PERMANOVA (adonis2 function in the vegan package in R). Females are noted as red and males as blue, primiparous samples as closed circles and multiparous as closed triangles.
Discussion
This study investigated the impact of delivery mode, birth order, and infant sex on the initial colonization of the infant gut by examining 70 first-pass meconium samples. We observed significant variations in microbial presence and composition based on these factors. Specifically, we found the lowest microbial presence in neonates delivered by elective C-section, an increased abundance of Bifidobacterium in neonates born to multiparous participants, and distinct differences in bacterial taxa between male and female infants delivered vaginally. These data underscore the importance of delivery mode in neonatal microbial colonization and suggest that together, both maternal and infant factors influence the early development of the gut microbiome.
We previously found that fetal meconium does not have a microbiome.13 Confirming this, the majority of first-pass meconium samples produced few or no reads and the proportion of neonatal meconium samples with any sequencing reads varied by delivery mode. C-section neonates had both the greatest proportion of meconium samples without any reads (i.e. low microbial presence) and the lowest number of reads per sample, consistent with previous findings that meconium from vaginally delivered infants has significantly higher bacterial counts than that from C-section infants.12 As C-section delivered infants are not exposed to the maternal vaginal or fecal microbiota during labor or delivery, delivery mode is associated with altered mother-to-infant transmission of key bacterial taxa, such as Bifidobacterium.14
Here, we found that Bifidobacterium was less abundant in first-born neonates. The protective role of bifidobacteria against pathogenic bacteria is well-documented,15 highlighting its importance in early gut health. While others have hypothesized inter-sibling microbial transfer may contribute to Bifidobacteria abundance in infants,12,16 we previously reported Bifidobacterium was more abundant in the multiparous mothers of these neonates compared to primiparous mothers.11 This suggests that the impact of birth order on the neonatal microbiome may be initiated prior to contact with older sibling and be partially driven by altered maternal microbiome composition, with multiparous mothers having a greater abundance of Bifidobacteria during pregnancy11 resulting in higher levels of Bifidobacteria being transmitted to infants during birth.
Unexpectedly, we found that infant sex also impacted the microbiome of vaginally delivered neonates. Females were dominated by Eubacteriales, and males were dominated by Enterobacterales. These findings are consistent with one previous report in preterm neonates17 and we hypothesize that sex-specific differences in intestinal development may play a role. Indeed, studies have shown sex-specific gene expression patterns in organ development18 and fetal growth.19 Estrogens are known to have impacts on intestinal function postnatally20,21 and in vitro, including increased mucin production.22 High concentrations of estrogens in female fetuses may similarly impact prenatal intestinal development, leading to enhanced intestinal maturation which could explain our finding that anaerobes were more abundant in females than males. Sex-specific differences in early life intestinal development may therefore confer changes in the postnatal intestinal niche and in patterns of microbial colonization.
The influence of parity on infant colonization is a novel observation that suggests that some ecological memory of previous pregnancies in the maternal microbiome may be transmitted to the next generation and could be sex specific. The greater mean abundance of Bifidobacterium and Bacteroides in female neonates born to multiparous women indicates a possible maternal influence on the neonatal gut microbiota. Moreover, the significant presence of Escherichia/Shigella in male neonates, but not females, underscores the potential for sex-specific microbial colonization patterns, which may have implications for immune education. Indeed, significantly higher rates of allergies are found both in boys compared to girls23 and in first-born children compared to later-born children,24 consistent with a protective effect of early Bifidobacterium and Bacteroides colonization.25,26
These findings highlight the complex interplay between delivery mode, birth order, and infant sex in shaping the neonatal gut microbiome. As expected, a large proportion of included samples did not produce any sequencing reads and the proportion of samples with sufficient reads for microbial community analysis varied by delivery mode. Therefore, our conclusions regarding the impacts of infant sex and birth order are limited by the modest sample size included in these analyses and are restricted to spontaneous vaginal deliveries. As delivery mode is known to alter the infant microbiome the impacts of sex and birth order may also vary by delivery mode. Future studies should continue to examine the combined effects of birth order and infant sex on early-life colonization across delivery modes and over time to better understand the mechanisms driving these differences and their potential long-term health implications.
Methods
Study design
The study protocol was approved by the Charité ethics committee (EA 4/059/16). All participants provided written informed consent. Participants were healthy females >18 years old with singleton pregnancies recruited as part of an ongoing research program at Charité University. Exclusion criteria included severe chronic gastrointestinal diseases, significant heart, kidney, liver, or pancreatic diseases, preexisting diabetes, depression, and delivery of infants with severe malformations. First-pass meconium samples from 70 participants were collected within 48 hours of birth and stored at −80°C. We found no significant differences in maternal age, gestational age, birthweight, or PROM between groups. Our analysis of microbial community composition was limited to samples from spontaneous vaginal deliveries without antibiotic exposure to avoid the potential confounding effects of delivery mode and intrapartum antibiotic exposure.
DNA extraction and amplification
Genomic DNA was extracted using a MagMAX Express semi-automatic robot with the MagMAX-96 DNA Multi-Sample Kit (Invitrogen, cat# 4413022), incorporating a mechanical lysis step with 2.8 mm ceramic beads and 0.1 mm glass beads for improved efficiency. Each extraction plate included four negative controls. The V3-V4 regions of the 16S rRNA gene were amplified (341F, CCTACGGGNGGCWGCAG; 806 R, GGACTACNVGGGTWTC-TAAT) modified with Illumina adapter sequences and 6-base pair multiplex barcodes. Absence of amplification was confirmed in all negative controls via gel electrophoresis, and one negative control per extraction plate was sequenced.
16S rRNA gene sequencing
Sequenced used the Illumina MiSeq platform (2×300bp) at the Farncombe Genomics Facility, McMaster University. Primers were trimmed from FASTQ files using Cutadapt (RRID: SCR_011841), amplicon sequence variants (ASVs) were derived using DADA2, and taxonomy was assigned using the Silva 132 reference database. Non-bacterial ASVs and those with < 2 sequencing reads were excluded.
Data and code availability
Sequencing data have been deposited at the NCBI Sequence Read Archive (SRA; PRJNA878704) and are publicly available as of the data of publication. Any additional information required to reanalyze the data reported in this paper is available from the lead contact upon request.
Statistical analysis
Participant characteristics shown in Table 1 were analyzed using the gtsummary package (RRID: SCR_021319) in R. Significance was assessed by Mann-Whitney U test for continuous variable and Fisher’s exact test for categorical variables. The impact of delivery mode, birth-order, and infant sex on the proportion of samples with >0 reads was assessed by generalized linear model. 16S data were analyzed using phyloseq27 (RRID: SCR_013080). Significance of alpha diversity (observed ASVs and Shannon diversity) was analyzed by independent linear regression models for each of infant sex, birth order, premature rupture of membranes, and gestational age. Significance of beta diversity (Bray-Curtis dissimilarity) was analyzed using vegan’s28 (RRID: SCR_011950) implementation of permutational multivariate analysis of variance (PERMANOVA) in the adonis2 command with infant sex and birth order as interacting effects. These results were visualized via Principal Coordinate Analysis (PCoA) ordination using R’s ggplot2 package (RRID: SCR_014601).29
Acknowledgments
We thank all the participants who were recruited in this study. We would like to thank Alexandra Kühn, Dr. Hanna Brinkmann, Laura Pasura, Laura Maschirow, Alexander Schwickert, and Sonja Entringer for assisting with patient recruitment and Loreen Ehrlich, Kerstin Melchior, and Thomas Ziska for assisting with sample preparation. We thank Michelle Shah for performing genomic DNA extractions and Laura Rossi for preparing amplicons for sequencing.
Funding Statement
M.G.S. and D.M.S. are supported by the Canada Research Chairs Program. T.B. is supported by Deutsche Forschungsgemeinschaft grant no. [BR2925/10-1]. A.P. is supported by Deutsche Forschungsgemeinschaft grant no. [PL241/16-1].
Disclosure statement
No potential conflict of interest was reported by the author(s).
Author contributions
Conceptualization, K.M.K., T.B., A.P., and D.M.S.; Investigation, K.M.K, J.S., M.H., and T.B.; Data Curation, K.M.K; Formal Analysis, K.M.K.; Visualization, K.M.K; Writing – Original Draft, K.M.K; Writing – Review & Editing, K.M.K, M.G.S., T.B., and D.M.S; Supervision, T.B., A.P., W.H., and D.M.S.; Project Administration, T.B., A.P., W.H., and D.M.S.; Resources, T.B., A.P., W.H., M.G.S., and D.M.S.; Funding Acquisition, T.B. and D.M.S.
References
- 1.Tamburini S, Shen N, Wu HC, Clemente JC.. The microbiome in early life: implications for health outcomes. Nat Med. 2016;22(7):713–9. doi: 10.1038/nm.4142. [DOI] [PubMed] [Google Scholar]
- 2.Dominguez-Bello MG, Costello EK, Contreras M, Magris M, Hidalgo G, Fierer N, Knight R. Delivery mode shapes the acquisition and structure of the initial microbiota across multiple body habitats in newborns. Proc Natl Acad Sci USA. 2010;107(26):11971–11975. doi: 10.1073/pnas.1002601107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Ferretti P, Pasolli E, Tett A, Asnicar F, Gorfer V, Fedi S, Armanini F, Truong DT, Manara S, Zolfo M, et al. Mother-to-infant microbial transmission from different body sites shapes the developing infant gut microbiome. Cell Host & Microbe. 2018;24(1):133–145.e5. doi: 10.1016/j.chom.2018.06.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Korpela K, Costea P, Coelho LP, Kandels-Lewis S, Willemsen G, Boomsma DI, Segata N, Bork P. Selective maternal seeding and environment shape the human gut microbiome. Genome Res. 2018;28(4):561–568. doi: 10.1101/gr.233940.117. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Bäckhed F, Roswall J, Peng Y, Feng Q, Jia H, Kovatcheva-Datchary P, Li Y, Xia Y, Xie H, Zhong H, et al. Dynamics and stabilization of the human gut microbiome during the first year of life. Cell Host & Microbe. 2015;17(6):852. doi: 10.1016/j.chom.2015.05.012. [DOI] [PubMed] [Google Scholar]
- 6.Jakobsson HE, Abrahamsson TR, Jenmalm MC, Harris K, Quince C, Jernberg C, Björkstén B, Engstrand L, Andersson AF. Decreased gut microbiota diversity, delayed bacteroidetes colonisation and reduced Th1 responses in infants delivered by caesarean section. Gut. 2014;63(4):559–566. doi: 10.1136/gutjnl-2012-303249. [DOI] [PubMed] [Google Scholar]
- 7.Nunez H, Nieto PA, Mars RA, Ghavami M, Sew Hoy C, Sukhum K. Early life gut microbiome and its impact on childhood health and chronic conditions. Gut Microb. 2025;17(1):2463567. doi: 10.1080/19490976.2025.2463567. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Sisk-Hackworth L, Kelley ST, Thackray VG. Sex, puberty, and the gut microbiome. Reproduction. 2023;165(2):R61–R74. doi: 10.1530/REP-22-0303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Corbier P, Dehennin L, Castanier M, Mebazaa A, Edwards DA, Roffi J. Sex differences in serum luteinizing hormone and testosterone in the human neonate during the first few hours after birth. J Clin Endocrinol Metab. 1990;71(5):1344–1348. doi: 10.1210/jcem-71-5-1344. [DOI] [PubMed] [Google Scholar]
- 10.Maccoby EE, Doering CH, Jacklin CN, Kraemer H. Concentrations of sex hormones in umbilical-cord blood: their relation to sex and birth order of infants. Child Dev. 1979;50(3):632–642. doi: 10.2307/1128928. [DOI] [PubMed] [Google Scholar]
- 11.Kennedy KM, Plagemann A, Sommer J, Hofmann M, Henrich W, Barrett JFR, Surette MG, Atkinson S, Braun T, Sloboda DM, et al. Parity modulates impact of BMI and gestational weight gain on gut microbiota in human pregnancy. Gut Microb. 2023;15(2):2259316. doi: 10.1080/19490976.2023.2259316. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Martin R, Makino H, Cetinyurek Yavuz A, Ben-Amor K, Roelofs M, Ishikawa E, Kubota H, Swinkels S, Sakai T, Oishi K, et al. Early-life events, including mode of delivery and type of feeding, siblings and gender, shape the developing gut microbiota. PLOS ONE. 2016;11(6):e0158498. doi: 10.1371/journal.pone.0158498. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Kennedy KM, Gerlach MJ, Adam T, Heimesaat MM, Rossi L, Surette MG, Sloboda DM, Braun T. Fetal meconium does not have a detectable microbiota before birth. Nat Microbiol. 2021;6(7):865–873. doi: 10.1038/s41564-021-00904-0. [DOI] [PubMed] [Google Scholar]
- 14.Makino H, Kushiro A, Ishikawa E, Kubota H, Gawad A, Sakai T, Oishi K, Martin R, Ben-Amor K, Knol J, et al. Mother-to-infant transmission of intestinal bifidobacterial strains has an impact on the early development of vaginally delivered infant’s microbiota. PLOS ONE. 2013;8:e78331. doi: 10.1371/journal.pone.0078331. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Fukuda S, Toh H, Hase K, Oshima K, Nakanishi Y, Yoshimura K, Tobe T, Clarke JM, Topping DL, Suzuki T, et al. Bifidobacteria can protect from enteropathogenic infection through production of acetate. Nature. 2011;469(7331):543–547. doi: 10.1038/nature09646. [DOI] [PubMed] [Google Scholar]
- 16.Imoto N, Kano C, Aoyagi Y, Morita H, Amanuma F, Maruyama H, Nojiri S, Hashiguchi N, Watanabe S. Administration of β-lactam antibiotics and delivery method correlate with intestinal abundances of Bifidobacteria and Bacteroides in early infancy, in Japan. Sci Rep. 2021;11(1):6231. doi: 10.1038/s41598-021-85670-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Cong X, Xu W, Janton S, Henderson WA, Matson A, McGrath JM, Maas K, Graf J. Gut microbiome developmental patterns in early life of preterm infants: impacts of feeding and gender. PLOS ONE. 2016;11(4):e0152751. doi: 10.1371/journal.pone.0152751. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Dewing P, Shi T, Horvath S, Vilain E. Sexually dimorphic gene expression in mouse brain precedes gonadal differentiation. Brain Res Mol Brain Res. 2003;118(1–2):82–90. doi: 10.1016/S0169-328X(03)00339-5. [DOI] [PubMed] [Google Scholar]
- 19.Broere-Brown ZA, Baan E, Schalekamp-Timmermans S, Verburg BO, Jaddoe VWV, Steegers EAP. Sex-specific differences in fetal and infant growth patterns: a prospective population-based cohort study. Biol Sex Differ. 2016;7(1):65. doi: 10.1186/s13293-016-0119-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Sankaran-Walters S, Macal M, Grishina I, Nagy L, Goulart L, Coolidge K, Li J, Fenton A, Williams T, Miller MK, et al. Sex differences matter in the gut: effect on mucosal immune activation and inflammation. Biol Sex Differ. 2013;4(1):10. doi: 10.1186/2042-6410-4-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.O’Mahony F, Thomas W, Harvey BJ. Novel female sex-dependent actions of oestrogen in the intestine. J Physiol. 2009;587(21):5039–5044. doi: 10.1113/jphysiol.2009.177972. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Diebel ME, Diebel LN, Manke CW, Liberati DM. Estrogen modulates intestinal mucus physiochemical properties and protects against oxidant injury. J Trauma Acute Care Surg. 2015;78(1):94–99. doi: 10.1097/TA.0000000000000499. [DOI] [PubMed] [Google Scholar]
- 23.Pinart M, Keller T, Reich A, Fröhlich M, Cabieses B, Hohmann C, Postma DS, Bousquet J, Antó JM, Keil T, et al. Sex-related allergic rhinitis prevalence switch from childhood to adulthood: a systematic review and meta-analysis. Int Arch Allergy Immunol. 2017;172(4):224–235. doi: 10.1159/000464324. [DOI] [PubMed] [Google Scholar]
- 24.Lisik D, Ermis SSÖ, Ioannidou A, Milani GP, Nyassi S, Spolidoro GCI, Kankaanranta H, Goksör E, Wennergren G, Nwaru BI, et al. Siblings and risk of allergic rhinitis: a systematic review and meta-analysis. Pediatr Allergy Immunol. 2023;34(7):e13991. doi: 10.1111/pai.13991. [DOI] [PubMed] [Google Scholar]
- 25.Sjögren YM, Tomicic S, Lundberg A, Böttcher MF, Björkstén B, Sverremark‐Ekström E, Jenmalm MC. Influence of early gut microbiota on the maturation of childhood mucosal and systemic immune responses. Clin Exp Allergy. 2009;39(12):1842–1851. doi: 10.1111/j.1365-2222.2009.03326.x. [DOI] [PubMed] [Google Scholar]
- 26.Björkstén B, Sepp E, Julge K, Voor T, Mikelsaar M. Allergy development and the intestinal microflora during the first year of life. J Allergy Clin Immun. 2001;108(4):516–520. doi: 10.1067/mai.2001.118130. [DOI] [PubMed] [Google Scholar]
- 27.McMurdie PJ, Holmes S, Watson M. Phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLOS ONE. 2013;8(4):e61217. doi: 10.1371/journal.pone.0061217. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Oksanen J. Vegan: community ecology package. 2013. R-package version 2.0-10. Preprint at 2013.
- 29.Wickham H. Ggplot2: elegant graphics for data analysis. Verlag New York: Springer; 2016. 978-3-319-24277-4. https://ggplot2.tidyverse.org/. [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
Sequencing data have been deposited at the NCBI Sequence Read Archive (SRA; PRJNA878704) and are publicly available as of the data of publication. Any additional information required to reanalyze the data reported in this paper is available from the lead contact upon request.


