Abstract
The discovery of N6-methyladenine (6mA) in eukaryotic genomes, typically found in prokaryotic DNA, has revolutionized epigenetics. Here, we show that symmetric 6mA is essential in the early diverging fungus Rhizopus microsporus, as the absence of the MT-A70 complex (MTA1c) responsible for this modification results in a lethal phenotype. 6mA is present in 70% of the genes, correlating with the presence of H3K4me3 and H2A.Z in open euchromatic regions. This modification is found predominantly in nucleosome linker regions, influencing the nucleosome positioning around the transcription start sites of highly expressed genes. Controlled downregulation of MTA1c reduces symmetric 6mA sites affecting nucleosome positioning and histone modifications, leading to altered gene expression, which is likely the cause of the severe phenotypic changes observed. Our study highlights the indispensable role of the DNA 6mA in a multicellular organism and delineates the mechanisms through which this epigenetic mark regulates gene expression in a eukaryotic genome.
Subject terms: Fungal genetics, DNA methylation, Fungal genomics
Here, the authors characterize the epigenetic landscape of the human fungal pathogen Rhizopus microsporus with a focus on symmetric DNA N6-methyladenine, revealing its regulatory role in gene expression and its essentiality for viability.
Introduction
Phenotypic plasticity comprises alterations in morphology, physiology, and behavior in response to changes in environmental conditions or specific cell requirements1. Epigenetics often mediates these changes, mainly through DNA methylation, RNAi, non-coding RNAs, and histone modifications2, defining a puzzling yet compelling framework for gene expression regulation. Among these, DNA methylation has received considerable attention. Bacteria rely on N6-methyladenine (6mA) to distinguish between self and foreign DNA, but it also participates in cell cycle regulation, DNA replication and repair, and gene expression regulation3–5. Consequently, and given its specificity, inhibitors of the methylation machinery with antimicrobial activity have been developed6. Distinctly, eukaryotic organisms have shifted to 5-methylcytosine (5mC) as their main DNA epigenetic mark, participating in processes such as genomic imprinting, X-chromosome inactivation, transposon silencing, gene expression regulation, and maintenance of epigenetic memory7,8. As part of the complex epigenetic landscapes, DNA methylation engages in intricate interactions that are often reciprocally influenced by histone modifications9–13. With a remarkable diversity among fungal lineages, epigenetic modifications have progressively gained attention14. For instance, the functional implications of 5mC have been demonstrated in gene regulation and genome defense mechanisms of Candida albicans and Neurospora crassa9,15–22. Additionally, the effects of various histone modifications on fungal development and secondary metabolism have been characterized in the model yeast Saccharomyces cerevisiae and filamentous fungi (Aspergillus nidulans, Trichoderma ressei)23–27, which have collectively contributed to deciphering the significant implications of epigenetics in fungal biology, ecology, and evolution.
The early-diverging fungi (EDF) represent a rich collection of different lineages and an extraordinary repository of fungal diversity28–31. With a notable but scant number of studies32–37, epigenetics in EDF remains to be explored. Remarkably, contrasting patterns of 6mA and 5mC levels have been reported34,35,37, highlighting the epigenetic diversity in the fungal kingdom and within EDF. The presence of 6mA as the predominant DNA modification in many EDF is a feature shared with some protists and green algae38–41. In protists, an MT-A70 complex (MTA1c) formed by two DNA methyltransferases (Mta1 and Mta9) and two DNA-binding proteins (P1 and P2) has been reported to be involved in 6mA deposition38,42. Mutation of the components of this complex in the ciliate Oxytricha resulted in a viable mutant but with a loss of sexual reproductive ability due to the reduction in genomic 6mA levels that also alter chromatin structure38,42,43. We have studied this complex in fungi and demonstrated the role of its components complex in symmetric 6mA deposition (meaning that both DNA strands carried this modification) in Mucor lusitanicus, while a different DNA methyltransferase conserved in EDF is responsible for asymmetric adenine methylation37. However, the more comprehensive characterization of how symmetric 6mA impacts the biology of EDF has been hampered by the very limited number of symmetric 6mA sites found in this species37.
For this purpose, we chose Rhizopus microsporus. This fungus is one of the known causal agents of mucormycosis, an emerging and fatal fungal disease with mortality rates that reach up to 90% in some cases44–47. The versatility of R. microsporus is further underscored by its ability to establish mutualistic endosymbioses with bacteria, such as Mycetohabitans spp48–50. Mycetohabitans synthesize rhizoxin, a toxin that inhibits plant mitosis, thereby promoting plant co-infection51. This fungal-bacterial association has been established as an important model for the study of cross-kingdom symbioses48,50,52–56. In addition, the recent application of a CRISPR/Cas9 procedure has facilitated the genetic manipulation of this fungus57,58.
In this work, we survey the R. microsporus epigenome in the most complete and detailed approach to date in an EDF, including 6mA detection using PacBio SMRT (Single Molecule, Real-Time) sequencing, 5mC detection (whole-genome bisulfite sequencing (WGBS)), and chromatin studies, including nucleosome occupancy by MNase-seq (micrococcal nuclease digestion with deep sequencing) and ChIP-seq (chromatin immunoprecipitation sequencing) profiling of H3K4me3, H3K9me3, and the histone variant H2A.Z. We find that the R. microsporus genome is compartmentalized into constitutive heterochromatic regions and open euchromatin regions that can be in an active or inactive transcriptional state, depending on 6mA, H3K4me3, and H2A.Z occupancy. We detect a clear association between 6mA and gene expression regulation and chromatin organization. Moreover, we characterize the role of the MTA1c, leading to the discovery that symmetric 6mA is essential in the fungus.
Results
6mA dominates the genomic DNA methylation landscape in R. microsporus
To evaluate 6mA distribution across the R. microsporus (ATCC 11559) genome, we generated a genome assembly using PacBio SMRT sequencing (Supplementary Table 1). This assembly covered more regions (27.39 Mbp vs 24.08 Mbp) and exhibited higher contiguity (L50 = 6 vs 87) than the previously available assembly for this organism48. Subsequently, we used SMRT sequencing data to generate a single-site resolution map of 6mA distribution (Supplementary Table 2). This analysis revealed a widespread presence of this modification (Fig. 1a) at about 1.40% of adenines in the genome of this fungus, which is above the levels reported in most eukaryotic organisms, including Dikarya and higher eukaryotic organisms (0–0.35%)34,41,59. The methylation ratio (methylated reads/total reads) of all detected 6mA sites revealed that the majority were fully or nearly fully methylated (92.36% of sites showed a 0.9–1 methylation ratio) (Supplementary Fig. 1a and Supplementary Table 3). To further confirm the high 6mA levels, R. microsporus genomic DNA (gDNA) was analyzed through HPLC-MS/MS, showing that about 1–1.1% of adenines are methylated (Supplementary Fig. 1a), similarly to other representative Rhizopus species, such as R. oryzae and R. delemar, which also exhibit high (>0.7%) 6mA levels (Supplementary Fig. 1b). Moreover, an isoschizomer digestion assay (DpnI/DpnII) revealed a distinct degradation pattern in R. microsporus gDNA when treated with DpnI (which exclusively cleaves at methylated GATC sites), contrasting with the minimally methylated gDNA from M. lusitanicus (Supplementary Fig. 1c).
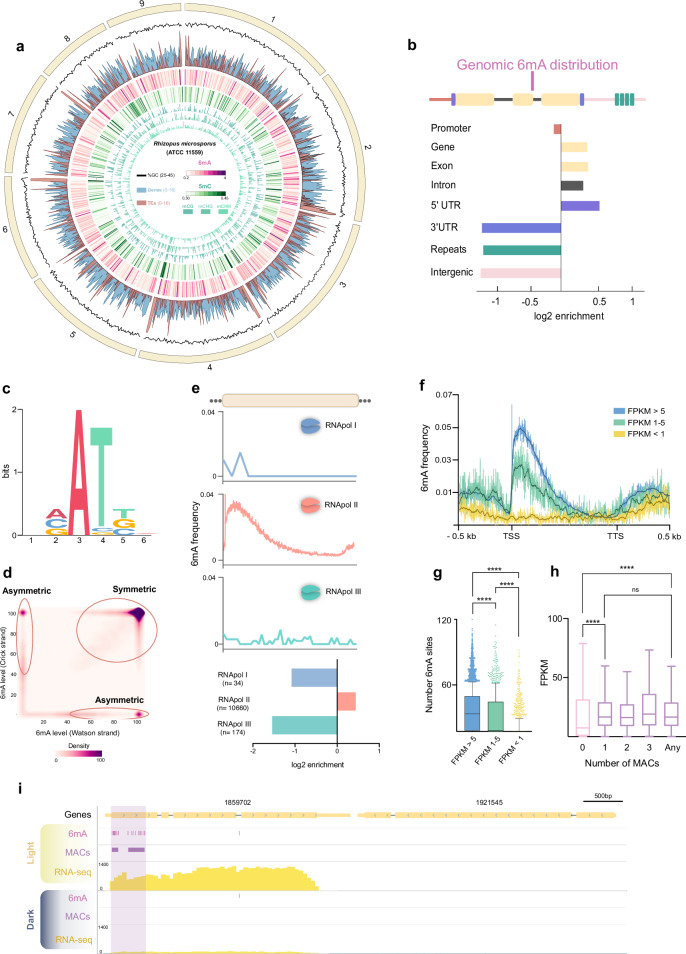
Fig. 1. Genome-wide implications and distribution of DNA epigenetic modifications in R microsporus.
a Gene (blue), TEs (red), 6mA (pink), and 5mC (green) density distribution across scaffolds 1–9 (20 kb bins). b 6mA enrichment by genomic features. c Logo of 6mA sites (bits indicate a measure of sequence conservation) d Density plot of 6mA methylation ratios on Watson (x-axis) and Crick strands (y-axis). e 6mA frequency over RNApol I (top), RNApol II (mid), and RNApol III (bottom) transcribed genes (bin size = 2.5% of scaled body length). The bottom part shows 6mA enrichment on RNApol I (blue), RNApol II (pink), and RNApol III (green). f 6mA frequency around TSS (bin size = 1) over highly (FPKM > 5, blue), intermediate (FPKM 1–5, green) and low (FPKM < 1, yellow) expressed genes. Wilcoxon rank was computed for each position and profile (P < 0.0001 for FPKM 1-5/FPKM < 1 and P > 0.0001 for FPKM 1-5/FPKM > 5). g Number of 6mA sites surrounding the TSS (−100nt + 400 nt) of highly (blue, n = 8279) intermediate (green, n = 924) and low (yellow, n = 1497) expressed genes. Boxplots indicate the median, first, and third quartile. Whiskers are drawn down to the 10th percentile and up to the 90th. Points above and below whiskers are drawn individually. A two-tailed student’s t-test showed significant differences (P < 0.0001 for all comparisons). h Expression levels (FPKM) of genes with different number of MACs surrounding the TSS (−100nt + 400 nt). Boxplots as in g. Whiskers were plotted according to Tukey method (75th and 25th percentile ±1.5 times interquartile distance). A student’s t-test (two-tailed) was performed to determine significant differences (P < 0.0001, ns = not significative). Non-significant correlation was found between any number of MACs (1, 2, 3, or any) and FPKM values (r = −0.01136, P = 0.3795) (n = 4718, 4331, 1447, 186, and 6982 genes for 0, 1, 2, 3, and any MACs). i Genome browser snapshot (scaffold_13: 373620-380092) indicating 6mA, MACs, and RNA-seq data under light and dark growth conditions. Purple shadow indicates MACs lost in dark versus light conditions. Two MACs upstream of gene ID:1859702 were lost in the dark, reducing expression. Source data are provided as a Source Data file.
6mA shows a symmetric distribution and it is concentrated in MACs
Genome-wide analysis of 6mA sites evidenced that this DNA modification was concentrated in specific regions and associated with specific genomic elements. 6mA marks were more frequent in gene-rich regions than in those populated by transposable elements (TEs) and repeats (Fig. 1a). Consistently, we found that 6mA was enriched in gene bodies, including exons and introns, rather than in intergenic regions or repeats (Fig. 1b). Particularly, 6mA marks tended to accumulate downstream the transcription start site (TSS) with a peak that decreases towards the transcription termination site (TTS) (Supplementary Fig. 1d). Such distribution closely mirrored the pattern previously observed in ciliates38,60, and differed from the bimodal distribution around the TSS detected in Chlamydomonas reinhardtii61. 6mA marks were preferentially located at ApT dinucleotides, with VATB (V: G, C or A - B: G, T or C) being the preferred motif (Fig. 1c). Noteworthy, ApT methylation accounted for more than 90% of the total sites although adenine methylation frequency did not correlate with ApT density (Supplementary Fig. 1e, P = 0.2933). Characteristically, 6mA sites in the R. microsporus genome were predominantly symmetric (Fig. 1d and Supplementary Table 4). The symmetry of 6mA sites in R. microsporus and other EDF34,35,37 suggests a mechanism for the propagation through DNA replication, as described in higher eukaryotes for CpG sites.
A substantial proportion of 6mA marks (90.31%) concentrated within methylated adenine clusters (MACs) (Supplementary Table 2), ranging from 15 to 20 bases up to more than 1 kb in length (Supplementary Fig. 1f) and were frequently located downstream of the TSS (Fig. 1f).
5mC and 6mA distributions diverge and play distinct roles
To generate a more complete picture of the R. microsporus methylome, we conducted WGBS. R. microsporus displayed low 5mC levels (0.34%), consistent with levels reported in other EDF34,37,62, with none of the three possible 5mC methylation contexts (CG, CHG, and CHG) being particularly enriched (Supplementary Fig. 2a). Contrary to 6mA, 5mC was less abundant in gene-dense regions and preferentially located in TE and repeat-rich regions (Fig. 1a and Supplementary Fig. 2b, c). Interestingly, 5mC was markedly enriched in DNA TEs, especially in the CG context, rather than retrotransposons (Supplementary Fig. 2d, e). Nevertheless, we found that 5mC might be relevant in LINE1 regulation, specifically in the CHG context (Supplementary Fig. 2d, e). Simple repeat regions also showed high 5mC levels (CG and CHG context), whereas MACs displayed low levels, indicating that 6mA and 5mC regulate different genomic elements (Supplementary Fig. 2d). These results show that both DNA modifications regulate different functions in the epigenomic landscape of R. microsporus.
6mA is associated with active RNApol II transcribed genes
The distribution of 6mA marks at the TSS prompted us to examine the relationship between this modification and gene expression. First, we found that more than 70% of genes were methylated (6mA) (Supplementary Fig. 3a). Second, consistent with the results in ciliates, 6mA in R. microsporus was enriched in RNApol II transcribed genes, but rare to nonexistent in RNApol I and RNApol III transcribed genes (Fig. 1e). The percentage of 6mA was higher in RNApol II transcribed genes compared to the whole genome (1.79% vs 1.32%), but remarkably lower in both RNApol I (0.62%) and RNApol III genes (0.45%) (Supplementary Table 5).
Next, we evaluated the association between 6mA presence and gene expression. A higher 6mA frequency around the TSS was detected in highly (FPKM > 5) and moderately (1 < FPKM < 5) expressed genes, but not in silent genes (FPKM < 1) (Fig. 1f). Accordingly, the number of 6mA sites around the TSS (TSS -100bp + 400 bp) was significantly larger in highly expressed genes than in the other two groups (Fig. 1g and Supplementary Fig. 3b). Remarkably, over 60% of silent genes lacked a 6mA mark surrounding the TSS, while a similar percentage of highly expressed genes had more than ten sites (Supplementary Fig. 3c). Genes lacking a MAC around the TSS (−100bp + 400 bp) showed a significantly lower expression level than those harboring a MAC (Fig. 1h). However, we did not find a correlation between the expression level and the number of MACs (Fig. 1h), suggesting that the presence or absence of a MAC is more important than the actual number of MACs. Furthermore, we performed Gene Ontology (GO) and Eukaryotic Orthologous Groups (KOG) enrichment analyses with MAC-marked genes. A broad variety of gene functions were enriched, such as RNA processing, translational and post-translational regulation, or vesicle transport (Supplementary Fig. 4), suggesting the wide implications of this modification on R. microsporus biology.
Given the association between 6mA and actively expressed genes, we analyzed the presence of 6mA in two different growth conditions. Since the initial R. microsporus samples were grown under light exposure, we performed SMRT sequencing on gDNA from cultures grown in the dark and coupled it with RNA-seq in the same conditions. In the dark, we detected a similar level of genomic 6mA and number of MACs as under light conditions (Supplementary Table 2), although there were some differences in their distribution. Dynamic changes in gene expression were accompanied by changes in 6mA as upregulated genes in light showed increased methylation levels compared to dark, and vice versa for downregulated genes (Supplementary Fig. 5a). Following the differences in 6mA distribution, we focused on genes that have lost a MAC in dark versus light conditions in the TSS surroundings. Among those, we detected 13 differentially expressed genes (DEGs) (p < 0.05) (Supplementary Table 6, Supplementary Data 1, and Supplementary Data 5) and 11 of them were significantly downregulated. However, the loss of MACs in upregulated genes also evidenced that this modification is not the only epigenetic component regulating gene expression in this organism (Fig. 1i, Supplementary Fig. 5b, c, and Supplementary Data 1).
6mA interplays with chromatin organization
To delve deeper into the impact of 6mA on the epigenomic landscape of R. microsporus, we evaluated the importance of this modification in chromatin configuration. We quantified 6mA levels by HPLC/MS in nucleosomal DNA from R. microsporus. Our analysis revealed a discernible decrease in 6mA levels within nucleosomal DNA when compared to gDNA (Supplementary Fig. 6a. One-way ANOVA). When using multi-nucleosomal DNA (bi-, tri-, and tetra-nucleosomal) containing linker regions, the 6mA levels were more similar to the total genomic levels (Supplementary Fig. 6a). To further characterize this distribution, we performed MNase-seq to map all the nucleosomes present in the R. microsporus genome (Supplementary Data 2). We detected 145,366 nucleosomes covering 21,322,704 nucleotides which represent ~78% of the genome. 6mA marks were more frequently located in linker regions, and nucleosome occupancy was clearly disadvantaged with respect to 6mA sites (Fig. 2a and Supplementary Fig. 6b). The 6mA levels increased towards the middle regions of the linker and not at the linker-nucleosome boundaries, resembling the 6mA distribution in Tetrahymena thermophila (Fig. 2b) and differing from that in C. reinhardtii63. The nucleosomes displayed a typical peak-to-valley pattern with respect to TSS, which was anti-correlated (opposite oscillating pattern downstream the TSS) with the oscillating-shape peak of the 6mA distribution towards the 3’ region of the gene (Fig. 2c, r = −0.2670, P < 0.0001).
Fig. 2. Association between 6mA and chromatin organization.
a 6mA is mostly found in linker regions. 6mA, and hence, MACs are frequent in nucleosome-free regions, as exemplified in this genome snapshot (scaffold 3:1062358-1068209). b 6mA frequency over linker and nucleosome regions. Both regions were scaled and divided into ten equal-sized bins. c 6mA and nucleosome dyads profiles with respect to TSS, including 0.5 kb upstream and downstream regions. d Methylated genes display a more consistent fixed distribution (left) than unmethylated genes (right). Normalized occupancy of nucleosome dyads was plotted in reference to TSS, including 1.5 kb flanking regions. A heatmap with the average normalized occupancy signal is also displayed below. Kolmogorov-Smirnov test (KS) was performed to evaluate differences between frequency distributions (P < 0.001) (n = 1732 genes and 1149 genes for methylated and unmethylated genes, respectively). Source data are provided as a Source Data file.
Next, we evaluated nucleosome occupancy profiles in methylated versus unmethylated genes, and although we did not detect dramatic changes in nucleosome occupancy, clear differences in peak-valley-peak profiles were noted (Fig. 2d). Methylated genes displayed more well-positioned nucleosomes, while in unmethylated genes, nucleosome positions were less fixed and showed a much fuzzier configuration (Fig. 2d and Supplementary Fig. 6c, Kolmogorov-Smirnov test P < 0.001). This pattern indicates that high levels of 6mA in linker regions are important for maintaining nucleosome positions, especially around the TSS, which could influence their favored transcription by RNApol II. Highly expressed genes showed a higher nucleosome occupancy and a less fuzzy nucleosome positioning (Supplementary Fig. 6d). Therefore, we hypothesize that 6mA in the linker regions of methylated genes, especially those marks clustered around the TSS, impacts nucleosome positioning, which regulates gene expression through RNA pol II accessibility.
6mA is enriched in euchromatin and H2A.Z-containing regions
To further investigate the interplay between 6mA and chromatin organization in R. microsporus, we conducted ChIP-seq to elucidate the distribution patterns of key histone modifications. In R. microsporus, H3K9me3 modification was concentrated in repeats and mostly absent in protein-coding genes (Fig. 3a, Supplementary Fig. 7, Supplementary Fig. 8a, and Supplementary Fig. 8b), aligning with previous findings in related species. In addition, it was specifically enriched in LTR retrotransposons (Class I) and DNA transposons (Class II, subclass 1), suggesting an important role in the maintenance of genome stability in this fungus (Supplementary Fig. 8c). Distinctly, H3K27me3, which typically marks facultative heterochromatic regions, was absent in the R. microsporus genome (Supplementary Fig. 7 and Supplementary Fig. 8a). This pattern can be explained by the lack of the enzymes responsible for this modification in R. microsporus as well as in several other EDF33.
Fig. 3. 6mA is condensed in H3K4me3 euchromatin regions and linked to H2A.Z occupancy.
a Genes (blue), repeats (red), expression (FPKM, yellow), MACs (pink), H3K4me3 and H3K9me enrichment (IP/Input ratio) and peaks (green and dark blue, respectively), and Input track (gray) across scaffold 11. b 6mA frequency over H3K4me3 (left, n = 4401) and H3K9me3 (right, n = 118) peaks. c H3K4me3 enrichment (IP/Input ratio) over MACs. Each MAC was extended to 2000 bp and fragmented into 200 bins (n = 7441). d Genome covered by H3K4me3 peaks, H3K9me3 peaks, and MACs (inside circles). Overlapping features are detailed outside the circles, with overlap percentages indicated. Overlapping percentages of random peaks generated by using average count and length of both H3K4me and H3K9me3 peaks are indicated in gray. A Chi-square test showed significant MAC enrichment in H4K3me3 peaks and H4K3me9 peaks compared to random regions (P < 0.0001) e Profile and heatmap of H3K4me3 enrichment (IP/Input ratio) per gene cluster, alongside 6mA (%), expression (log2 FPKM), H3K9me3 (IP/Input ratio), and H2A.Z occupancy (IP/Input ratio). f Scatter plot of gene expression (log2 FPKM, y-axis) vs H3K4me3 enrichment (x-axis). Genes were color-coded according to whether they harbor a MAC (dark pink) or not (light pink). MACs were filtered to a minimum of 20 bp in length and a 0.005 methylation rate. Differences in groups were tested using Mann-Whitney test (two-tailed) (P < 0.0001). g Correlation between H3K4me3, H3K9me3, and H2A.Z (IP/Input ratio) with 6mA levels (%). The genome was divided into 30 kb bins, and the enrichment for each ChIP-seq experiment and 6mA percentage was computed for each bin. Pearson’s correlation coefficient was computed for each comparison (top right corner of each plot). Source data are provided as a Source Data file.
For its part, H3K4me3, which is typically associated with euchromatin and frequently linked to active gene expression64–66, showed an opposite distribution to H3K9me3 and was mostly found over protein-coding genes. Moreover, it was enriched downstream of the TSS, in the 5’ region of the gene (Fig. 3a, Supplementary Fig. 7, and Supplementary Fig. 8a), following a similar distribution and pattern to 6mA. The frequency of 6mA was higher over H3K4me3 peaks, whereas H3K9me3 peaks were mostly devoid of 6mA (Fig. 3b and Supplementary Fig. 9a). In agreement with this pattern, MACs were enriched in H3K4me3 (Fig. 3c). Nearly 30% (28.42%) of the genome associated with MACs (7.66%) exhibited overlap with H3K4me3 peaks, while only 2.52% of the MACs coincide with H3K9me3 regions (Fig. 3d). This correlation was also found when we ranked genes into three groups based on their H3K4me3 levels, we found that those with higher H3K4me3 occupancy had higher 6mA levels and were more actively expressed (Fig. 3e and Supplementary Fig. 9b,c). Similar to genes harboring a MAC, genes with H3K4me3 showed significantly higher expression levels compared to those lacking H3K4me3 (Supplementary Fig. 9c). However, we did not observe a synergistic effect on the expression of genes carrying both 6mA and H3K4me3 (Supplementary Fig. 9d). When genes were plotted based on their expression level and H3K4me3 occupancy, two distinct groups of genes emerged: one characterized by high expression and H3K4me3 levels that overlapped with MAC-containing genes, and another with lower expression and reduced H3K4me3 levels (Fig. 3f, Mann-Whitney test P <0.0001).
Lastly, we focused on the histone variant H2A.Z, which followed a bimodal distribution with respect to the TSS, resembling those described in other organisms (Supplementary Fig. 10a)67–71. We also observed a very strong correlation between H2A.Z occupancy and H3K4me3 and an anti-correlation with H3K9me3 (Supplementary Fig. 10d), similar to what has been proposed in cases in which H2A.Z is associated with active gene expression67,72,73. Hence, we found that H2A.Z occupancy also correlated positively with 6mA presence (Fig. 3g). Interestingly, by analyzing nucleosomes flanked by either the 10,000 more or the 10,000 less dense 6mA linkers, we found that H2A.Z occupancy was remarkably higher in the nucleosomes spanning highly methylated regions than in those lacking 6mA (Supplementary Fig. 10b). Despite the bimodal distribution at the TSS in S. cerevisiae, H2A.Z is almost exclusively incorporated into the +1 nucleosome, and the occupancy at the -1 nucleosome is a consequence of bidirectional transcription in very compacted genomes74. When we analyzed H2A.Z occupancy, we found that 6mA-containing genes showed a much greater H2A.Z occupancy in the +1 nucleosome than unmethylated genes (Supplementary Fig. 10c). Altogether, our findings indicate the presence of two distinct chromatin states within the R. microsporus genome. 6mA is absent in constitutive heterochromatic regions crucial for genome stability. Conversely, this modification is prevalent in euchromatic regions marked by H3K4me3 and associated with nucleosomes containing the histone variant H2A.Z (Fig. 3g).
Components of the MTA1c is conserved in 6mA-rich eukaryotes
MT-A70 methyltransferases are known for their role in DNA and RNA methylation61,75,76. In EDF and other 6mA-rich eukaryotes, the MTA1c, which is responsible for symmetric 6mA deposition in the DNA, is formed by two MT-A70 methyltransferases (Mta1 and Mta9) and one or two proteins containing a Myb-like DNA-binding domain (P1 and P2)38,42. We investigated the conservation of MT-A70 proteins in representatives from the three groups of 6mA-rich organisms (green algae, protists, and EDF) as well as in representative species of plants, metazoans, and other fungal clades (Fig. 4a, Supplementary Fig. 11, and Supplementary Data 3). The MT-A70 proteins Mta1 and Mta9 were conserved only in the species displaying high 6mA levels, including R. microsporus and other EDF (Fig. 4a, b). Interestingly, Catenaria anguillulae, a known EDF with reduced 6mA levels34, lacked these proteins, as did Allomyces macrogynus (Supplementary Fig. 12), suggesting a likely loss of this trait in Blastocladiomycota. Notably, Chytridiomycota representatives possess both the 6mA RNA methylation system METTL3-14 and the 6mA DNA MTA1c, suggesting an intermediate state in fungal MT-A70 evolution with the retention (or partial retention) of both systems. Prokaryotic MT-A70 methyltransferases (PF05063) are clearly distinguishable from their eukaryotic counterparts, suggesting vertical inheritance and diversification after the transfer of the prokaryotic MT-A70 genes to eukaryotes, as previously reported77. Interestingly, Mta9 duplication was observed in Mortierellomycotina, Entomophthoromycotina, and Mucoromycotina, including most Mucorales (Supplementary Fig. 12, Supplementary Fig. 13, and Supplementary Data 4). The two R. microsporus Mta9 copies (PHZ11537.1, Mta9_A, and PHZ11228.1, Mta9_B) retained residues essential for interaction with Mta1 (Supplementary Fig. 12c)78. These Mta9 paralogs differ slightly in an important loop at the Mta1-Mta9 interface, with a proline instead of an arginine (K218 in Paramecium) and a serine/arginine instead of an asparagine (N221 in Paramecium), which might result in complex stability differences. In addition to Mta1 and Mta9, 6mA-rich eukaryotes, including R. microsporus, had the protein P1 containing a Myb-like DNA-binding domain, although some EDF had the second protein P2, similar to ciliates (Fig. 4a, b).
Fig. 4. The Mta1 complex in R microsporus.
a Phylogenetic tree of representative eukaryotic species coupled with a matrix indicating the presence (filled box) or absence (empty box) of each MTA-70 protein. Species are classified into color-coded groups. Fungal phyla clades are ordered in the same descending order they appear in the tree. b Schematic representation of the MTA1c present in R. microsporus (top) and the domain composition of each protein (bottom). Charge distribution is displayed under the schematic representations. c The interaction between Mta1, Mta9, and P1 was analyzed in a yeast two-hybrid assay. Mta1 shows both weak (-WLH) and strong (-WLA) interactions with Mta9 and P1. d Genome browser view (scaffold_4:1865593-1878823) showing 6mA presence and P1 enrichment (computed as IP/Input ratio). e The methylation complex binds preferentially to 6mA-rich regions. P1 enrichment from ChIP-seq experiments (top) and DAP-seq (bottom) was computer over MACs (divided into 50 bins). f Methylated genes (6mA) show a greater enrichment of P1 downstream of the TSS than unmethylated genes (No 6mA). Source data are provided as a Source Data file.
A yeast two-hybrid assay revealed the interaction between Mta1 and Mta9 as well as between Mta1 and P1 (Fig. 4c and Supplementary Fig. 14). This finding aligns with a recent structural analysis carried out on these proteins, suggesting that Mta1 acts as the catalytic subunit and the core component of the complex78. To characterize the interaction of this complex with DNA, we developed a strategy in R. microsporus for gene tagging at the 3’-end (Supplementary Fig. 15a, b) using the red fluorescent protein mCherry, which is the sole tag successfully employed in EDF79. Attempts to tag Mta1 were unsuccessful, as we were unable to isolate homokaryotic mutants, suggesting loss of gene function (Supplementary Fig. 15a). However, tagging of P1 following the same strategy was feasible, resulting in the isolation of homokaryotic mutants (Supplementary Fig. 15b), which were used in ChIP-seq experiments. In parallel, we conducted a DAP-seq experiment using the complete and four truncated versions of P180,81. Results from both ChIP-seq and DAP-seq revealed a stronger P1 binding signal over 6mA-rich regions (Fig. 4d, e). We also found greater enrichment at the 5’ region of 6mA-enriched genes compared to unmethylated genes (No 6mA) (Fig. 4f), with a peak that closely resembled the shape and location of the 6mA peak downstream of the TSS (Supplementary Fig. 16a). In addition, coinciding with 6mA-spanning linker regions, this complex bound preferentially to linker regions with increasing enrichment from nucleosome boundaries to linker centers and decreasing signal from nucleosome boundaries to nucleosome centers (Supplementary Fig. 16b). The shared AAC-containing motif identified in ChIP and DAP-seq experiments was coincident with the motif bound by the WER Myb-R2R3 transcription factor of Arabidopsis thaliana82 (Supplementary Fig. 16c). This motif is recognized by two asparagine residues (N106 and N110), which are highly conserved in all EDF P1 proteins, with the only exception of the Zoopagomycota representatives (Supplementary Fig. 17a, b).
MTA1c components are essential in R. microsporus
The inability to generate mta1-tagged strains prompted us to investigate whether mta1, mta9, and/or p1 are essential in R. microsporus. Although we obtained transformants for mta1, mta9, and p1 gene disruption (Fig. 5a), all of them were unstable heterokaryotic mutants (Fig. 5b) instead of homokaryons. When these heterokaryotic mutants were grown on selective media, very tiny and septate colonies were observed (Supplementary Fig. 18a). In addition, the fraction of mutant nuclei was almost lost after one growth cycle on non-selective media (Supplementary Fig. 18b), highlighting the deleterious effect caused by the loss of either of these genes. Although unstable, we were able to quantify a 20% reduction in genomic 6mA levels by HPLC-MS/MS in heterokaryotic mutants for mta1 (UM22) and p1 (UM24) (Supplementary Fig. 18c). To confirm the essential function of MTA1c and to facilitate the study of the mutants, we developed a zinc-regulated expression system. We utilized the promoter of the zrt1 gene, encoding a zinc permease, whose expression is strongly repressed by the presence of zinc in the media, as validated by qRT-PCR (Supplementary Fig. 19a). Using this promoter, we generated two mta1 constructs: construct A contained the entire mta1 gene, including the 3’ and 5’ UTR sequences and its own terminator, while construct B contained the mta1 coding sequence fused to the terminator sequence of the carRP gene, which had proven to be effective in the closely related species M. lusitanicus83 (Supplementary Fig. 19b). We selected homokaryotic mutants for the integration of each construct: UM45 (construct A) and UM46 (construct B) (Supplementary Fig. 19c). Next, we disrupted the endogenous mta1 gene in both UM45 and UM46 strains (Fig. 5c). The presence of the additional zinc-regulated copy of the mta1 gene, allowed us to isolate homokaryotic mutants, UM64 (construct A) and UM67 (construct B) (Fig. 5d and Supplementary Fig. 19c, d, e). These strains were able to grow normally on unsupplemented zinc media (promoter was activated), but showed a dramatic reduction in growth when zinc was added to the media (promoter was downregulated) that was not detected in the wild-type strain (Fig. 5e and Supplementary Fig. 19f, g). The prevalence of 6mA as the dominant DNA epigenetic modification in the R. microsporus, coupled with the inability to generate knock-out mutants for any of the genes comprising the MTA1c, and the manifestation of a severe phenotype upon the knockdown of the mta1 gene collectively demonstrated the indispensability of this gene, the associated complex, and consequently, 6mA, as an essential DNA modification in R. microsporus.
Fig. 5. Genetic engineering of R. microsporus reveals the essential role of the MTA1c.
a Schematic representation of the knockout strains generation. The double-strand break provoked by the Cas9+gRNA complex is repaired by microhomology regions flanking the auxotrophy marker. b Disruption of mta1 (UM22), mta9 (UM26), and p1 (UM24) genes results in the isolation of heterokaryotic mutants. The result of the PCR amplification of the targeted locus (hybridization sequences of the primers are indicated as red arrows in a reveals the presence of both WT nuclei (amplicon sizes: 1.5 kb mta1, 3.8 kb mta9, 3.1 kb p1) and mutant nuclei (amplicon sizes: 5.2 kb mta1, 7.3 kb mta9, 6.6 kb p1). This experiment was independently repeated three times with similar results. c Scheme describing the strategy to obtain the strain in which the mta1 gene is regulated by the Pzrt1 promoter. The double-strand break generated in the leuA locus was repaired by the construct carrying the pyrF marker, the Pzrt1 promoter, and the mta1 gene (see Fig S19). The endogenous copy of the mta1 gene was then disrupted using the leuA marker as a repair DNA template. d PCR amplification of the mta1 endogenous locus reveals that all the mutants (UM64, UM65, UM67, and UM68) analyzed are homokaryons, as only mutant nuclei are observed (band size: 5.2 kb, WT: 1.5 kb). Primers used for the amplification are indicated as red arrows in c. This experiment was independently repeated three times with similar results. e Growth of the UM64 and UM67 strains (two independent mutants carrying the regulated copy of mta1 with the endogenous mta1 copy disrupted) in the presence of zinc. 1000, 100, and 10 spores were plated in YNB and YNB supplemented with 20 mg/L ZnSO4. Source data are provided as a Source Data file.
Symmetric 6mA is a determinant of active chromatin regions
To characterize the dynamics of 6mA and its direct impact on gene expression and chromatin structure, the UM67 strain was grown initially on ZnSO4-unsupplemented medium (mta1 is expressed) and then transferred to medium supplemented with ZnSO4 and finally transferred back to unsupplemented medium to restore mta1 expression (Fig. 6a). Changes in 6mA levels were determined by SMRT sequencing (Fig. 6b), and confirmed by HPLC-MS/MS and 6mA dot blot (Fig. 6b and Supplementary Fig. 20a). Genomic 6mA levels were reduced from 1.3% to 1.15% (12.5% reduction) when the strain UM67 was transferred from unsupplemented medium to 10 mg/L ZnSO4 (Fig. 6b) and to 0.83% (37% reduction) when it was transferred to 20 mg/L ZnSO4 (Fig. 6b). HPLC-MS/MS quantification showed a similar trend in 6mA reduction, although the decreases were higher with a 38% and 46% reduction for medium and high Zn levels, respectively (Fig. 6b). This decline in genomic 6mA levels was ony detected in the strain UM67, in which the mta1 gene is under Pzrt1 control, and not in the wild-type strain when exposed to ZnSO4 (Supplementary Fig. 20b). Interestingly, genomic 6mA levels were restored when the strain UM67 was grown back without Zn supplementation (Fig. 6b), with 6mA distribution being essentially identical to the initial starting point (Supplementary Fig. 20c, d). The characteristic 6mA presence downstream of the TSS was reduced when mta1 was downregulated (Fig. 6c). Accordingly, we observed that the number of symmetric sites was reduced from 92.91% in the unsupplemented medium to 75.15% in 20 mg/L ZnSO4 medium (Fig. 6d). Altogether, these results evidenced a direct link between mta1 expression and the global genomic 6mA levels.
Fig. 6. Genome-wide effect of mta1 downregulation.
a Schematic diagram of the dynamic mta1 regulation experiment. From a starting point with 0 mg/L ZnSO4 and active Mta1 expression, it was downregulated by increasing ZnSO4 to 10 mg/L and 20 mg/L. Colonies growing in 20 mg/L ZnSO4-supplemented media were transferred again to media without ZnSO4. b Genomic 6mA levels measured by SMRT sequencing and HPLC-MS/MS. Data are represented as mean ± SD. Different letters indicate statistically significant differences, while the identical letters denote no significant differences, calculated using one-way ANOVA ( P < 0.001). (HPLC/MS n = 3 biological replicates for each growth condition analyzed) c 6mA frequency profile over protein-coding genes for the UM67 strain growing with 0 mg/L (blue), 10 mg/L (green), and 20 mg/L (yellow) ZnSO4. d Proportion of symmetric 6mA sites and motif logo for UM67 growing with 0 mg/L ZnSO4 (left) and 20 mg/L ZnSO4 (right). e Upregulated genes (red) and downregulated genes (blue) that have lost a MAC when growing in media supplemented with 20 mg/L (top panel) and with 10 mg/L (bottom panel). Two-tailed P values were obtained from DESeq2 Wald’s test. Multiple comparison adjustments were performed with the Benjamini-Hochberg false discovery rate (FDR) correction procedure. f 6mA frequency profiles over upregulated and downregulated genes (20 mg/L vs 0 mg/L). The blue line indicates the 6mA frequency for UM67—0 mg/L, and the yellow line indicates the 6mA frequency for UM67—20 mg/L. g H3K4me3 enrichment (IP/Input ratio) over MAC lost when UM67 is grown with 20 mg/L ZnSO4 (right panel) compared to 0 mg/L (left panel). Each MAC was extended to 2000 bp and fragmented into 200 equally sized bins. h Profile and heatmap of 6mA and H3K4me3 enrichment (IP/Input ratio) for each cluster of genes (upregulated, upregulated with MAC lost; downregulated, downregulated with MAC lost). Red and blue lines in the profiles indicated upregulated and downregulated genes in the 20 mg/L vs 0 mg/L, respectively. For both 6mA and H3K4me3 panels, the panel on the left is for UM67—0 mg/L, and the panel on the right is for UM67—20 mg/L. i Normalized occupancy of nucleosome dyads in DEGs (20 mg/L vs 0 mg/L) was plotted in reference to TSS, including 1.5 kb flanking regions. A heatmap with the average normalized occupancy signal is also displayed below. Kolmogorov-Smirnov test (KS) was performed to evaluate differences between frequency distributions (P < 0.001). j Nucleosome classification according to their fuzziness score for upregulated (top panel) and downregulated (bottom panel) genes. Nucleosomes bound to genes with MAC lost (20 mg/L vs 0 mg/L) are indicated in light pink. Kolmogorov-Smirnov test (KS) was performed to evaluate differences between frequency distributions (P = 0.0053 for upregulated and P < 0.0001 for downregulated genes). Source data are provided as a Source Data file.
Next, we evaluated how gene expression was affected by the loss of MACs (Supplementary Data 5), filtering out DEGs in response exclusively to zinc starvation/supplementation (Supplementary Data 1). Among all genes that have lost a MAC (481 genes), we found 134 out of the 163 downregulated genes comparing unsupplemented and high Zn growth conditions (28 were upregulated), and 29 out of the 42 downregulated genes (12 were upregulated) when comparing unsupplemented and medium Zn conditions (Fig. 6e). When we analyzed the number of genes that had lost a MAC among all DEGs detected when mta1 activity was reduced, we found that MAC losses were much more frequent in downregulated genes than in upregulated genes (Supplementary Fig. 20e). Following the dynamic association of 6mA and gene expression, we also detected a significantly different methylation distribution in genes that were upregulated and downregulated, with 6mA losses being especially notable in downregulated genes (Supplementary Fig. 20f). 6mA distribution downstream of the TSS did not show great differences in upregulated genes (Fig. 6f), whereas downregulated genes showed a clear reduction in methylation density (Fig. 6f). As symmetric 6mA deposition is compromised when mta1 expression is reduced (ZnSO4), upregulated genes under this condition must rely on different mechanisms to boost their expression, whereas the loss of 6mA seems to be one of the major determinants of the reduced expression in downregulated genes. These results reinforce the hypothesis that 6mA is associated with activation of transcription, although it does not have an exclusive role.
In constitutive heterochromatin regions, we found no noticeable changes when 6mA levels were reduced, with H3K9me3 showing an identical negative correlation with 6mA (Supplementary Fig. 20g) and a similar number and length of peaks and proportion of the genome occupied by this histone modification (Supplementary Fig. 20h, i). In the case of active chromatin regions, we observed that both H2A.Z and H3K4me3 still displayed a positive correlation in their distribution when compared to 6mA in the presence of Zn (Supplementary Fig. 20g). Nevertheless, while the positive correlation coefficient for H2A.Z and 6mA was not altered when mta1 expression was downregulated (0.36 and 0.35), we noticed a major change in the correlation coefficient for H3K4me3 and 6mA (0.52 and 0.39) (Supplementary Fig. 20g). Based on our previous observation that MACs were also rich in H3K4me3, we analyzed the occupancy of this histone modification in regions where MACs were lost as a result of reduced mta1 expression. This analysis showed that MAC loss provoked a reduction of H3K4me3 occupancy (Fig. 6g), indicating that the presence of symmetric 6mA sites could favor the deposition of this histone modification. We found that these upregulated genes displayed a higher H3K4me3 content in the presence of Zn, which was also noticeable in those that have lost a MAC, whereas no relevant differences in H3K4me3 occupancy were observed in the downregulated genes (Fig. 6h). These results suggest that H3K4me3 could act as a compensatory mechanism when gene upregulation is required and the symmetric 6mA machinery is compromised, whereas 6mA loss is sufficient to explain downregulation.
Finally, we performed MNase-seq on the UM67 strain at high Zn and unsupplemented Zn conditions to examine the effect of 6mA on the organization of chromatin structure. By plotting the distribution of nucleosome dyad with respect to the TSS of protein-coding genes, we observed that the overall reduction in genomic 6mA levels resulted in a shallower profile, indicating lower nucleosome positioning in the cell population (Fig. 6i, Kolmogorov-Smirnov test P < 0.001). This observation was supported by a higher fuzziness degree of nucleosomes bound to both upregulated and downregulated genes that have lost a MAC as a result of reduced mta1 expression (Fig. 6j, Kolmogorov-Smirnov test). These results suggest 6mA, rather than the gene expression itself, plays a crucial role in determining the positioning of nucleosomes.
Discussion
Epigenetic modifications significantly expand the information encoded by the DNA sequence, providing an additional layer of regulation that governs gene expression. These modifications play a crucial role in enabling cell specialization, driving differentiation in multicellular organisms, and allowing phenotypic adaptation to environmental changes84. In fungi, epigenetic regulation is decisive in development, pathogenesis, secondary metabolism, sexual reproduction, and antifungal drug resistance14,85–89. Most of the fungal genetics research, including epigenetics, has focused on higher fungi (Ascomycota and Basidiomycota), overlooking the broader group of EDF. In this study, we focus on the role of 6mA in the epigenome and regulation of gene expression in R. microsporus, as 6mA is proposed to be important in EDF biology. This modification is highly abundant in EDF, in contrast to the reduced presence in most eukaryotic organisms90,91. In the R. microsporus genome, 6mA is clustered in specific genomic regions (MACs) rather than randomly dispersed, and shows a mostly symmetric distribution as well as ApT enrichment. The clustering of 6mA resembles the distribution of 6mA in bacteria, deposited by the processive maintenance methyltransferases, such as Dam92. The processive mechanism (after binding, methyltransferases deposit several methylation sites before moving to other locations) could also be happening in the 6mA-rich eukaryotes (protists and algae).
We have detected a clear association between the presence of MACs downstream of TSS and active gene expression, in agreement with other EDF34,37. In other organisms, depending on the species and genomic context, 6mA is associated with either active or silencing gene expression39,93–98. 6mA is associated with active genes in C. reinhardtii39 and Arabidopsis99 and with the activation of TEs during embryonic stages in Drosophila95,100. In the case of vertebrates, 6mA is associated with active genes in Xenopus, zebrafish, and pig93,101. However, its role as a repressive mark has been reported in glioblastoma cells, young LINE-1 TEs silencing in mouse embryonic stem cells, and gene silencing of genes located on chromosome X96,102. In the case of T. thermophila, only a weak association with active genes has been described60. In contrast to 6mA, R. microsporus, as other EDF, displays reduced genomic levels of 5mC, the main DNA regulatory mark in eukaryotic genomes, including other fungi8,62. Levels of 5mC are significant over repetitive regions. We delineate a heightened presence of 5mC within LINE1 TEs (non-LTR). Intriguingly, in M. lusitanicus33, an organism closely related to the EDF, and in R. microsporus, our findings underscore the significance of repressive H3K9me3, especially in the regulation of LTR TEs. This suggests a potential complementary role of both 5mC and H3K9me3 in maintaining genome stability within EDFs.
The generation of a whole genome nucleosome map in an EDF representative revealed that 78% of the genome is covered by nucleosomes, which is similar to other fungal species such as S. cerevisiae (81%)103. In addition to the low frequency of 6mA sites in nucleosome-bound DNA, 6mA-rich linkers exhibit a higher degree of nucleosome positioning. These 6mA-rich linkers and their connection with actively expressed genes lead us to hypothesize that the 6mA dense regions located downstream of the TSS are determining nucleosome position, thus favoring RNA polymerase II accessibility and expression (Fig. 1f, Fig. 2b, and Supplementary Fig. 1d).
Furthermore, we found that 6mA is absent in H3K9me3-marked constitutive heterochromatin regions. As in several other species, H3K9 methylation is detected exclusively in transcriptional deserts and repeat-rich regions, indicating its importance for genome stability maintenance24,104–106. On the contrary, 6mA is mostly found in open chromatin regions and shows a significant enrichment of dense 6mA over H3K4me3 regions, a histone modification that has been associated with active gene expression in fungi64,65,107,108. Notably, 6mA and H3K4me3 follow a similar genomic distribution in the R. microsporus genome. Interestingly, we found that H3K27me3, an epigenetic modification associated with facultative heterochromatin regions71,109,110, is absent in R. microsporus. In conclusion, we propose an epigenetic landscape divided into constitutive heterochromatin regions marked by H3K9me3 and 5mC, and open chromatin regions dominated by H3K4me3 and 6mA presence (Fig. 7). We extended our epigenetic characterization by assessing the distribution of H2A.Z and linking this histone variant to the active compartment of open chromatin, resembling the connection between this variant and 6mA reported in ciliates60 (Fig. 7). We found that H2A.Z downstream the TSS (+1 nucleosome) in the R. microsporus genome, is associated with active 6mA-H3K4me3 marked chromatin regions (Fig. 3g and Supplementary Fig. 10c).
Fig. 7. Model of epigenetic modifications distribution and epigenomic compartmentalization of the R microsporus genome.
A schematic representation of chromatin configurations in a nucleus showing the epigenetic landscape described in this study. The distribution of nucleosomes, 6mA, H3K4me3, H3K9me3, and H2A.Z on euchromatin and constitutive heterochromatin regions is displayed.
This study delves into the characterization of the 6mA MT-A70 methylation complex responsible for symmetric methylation in EDF37, applying and developing molecular and genetic tools in R. microsporus. We identify the DNA binding targets by combining both ChIP-seq and DAP-seq and find a high degree of conservation in EDF of the two asparagine residues involved in the interaction with both A nucleotides from the AAC motif, as described for the WER Myb-R2R3 transcription factor of A. thaliana82. The presence of methyl groups in these adenine nucleotides inhibited this binding interaction which could suggest a possible mechanism of self-regulation for 6mA deposition in fungi. Knock-out of mta1, mta9 and p1 showed that the three components of this complex are essential in R. microsporus. This is further confirmed by fine-tuning modulation of mta1 expression, which encodes the catalytic component of this complex. Downregulation of mta1 results in a reduction in genomic 6mA levels of around 30–40%, which is comparable to the 30–50% reduction detected in the ciliates O. trifallax and T. thermophila (39, 167), suggesting that this could be the minimum threshold for 6mA reduction in 6mA rich eukaryotes. Distinctly, the generation of mta1 knockouts was feasible in ciliates, suggesting that other DNA methyltransferases could be responsible for 6mA deposition. However, in R. microsporus and probably other EDF37, only the MTA1c is responsible for 6mA deposition, underscoring its essential role in these organisms.
The reconstitution of the symmetric 6mA sites, by re-establishing mta1 expression, results in an almost identical 6mA distribution between the initial and final points of the dynamic experiment. This, together with Mta1 processivity, inferred from the distribution of 6mA in dense clusters, indicates that the MT-A70 may have an essential function in maintaining symmetric sites after DNA replication. Remarkably, the ability to regulate mta1 expression also allows for assessing the role of 6mA on chromatin structure and gene expression in R. microsporus. Our findings confirm that heterochromatic regions are independent of 6mA presence, whereas this modification plays a crucial role in the regulation of active chromatin, as its loss is directly associated with reduced gene expression and altered H3K4me3 distribution. Moreover, MNase-seq data under reduced and restored mta1 expression conditions indicate that the loss of symmetric 6mA results in more poorly positioned and overall fuzzier nucleosomes, affecting both upregulated and downregulated genes. This is consistent with the studies suggesting that 6mA provokes a physical impairment of nucleosomes, thereby restricting and fixing their distribution38,42,43,60. Depletion of 6mA disrupts this nucleosomal organization, potentially impeding the accessibility of RNA polymerase II and subsequently influencing gene expression.
On a practical note, considering that Rhizopus is the most frequent causal agent of mucormycosis45,111 and given that the lack of specificity is one of the major concerns in the antifungal drug development, we propose that 6mA and, in particular, MTA1c as an ideal antifungal drug target. This approach has the potential to overcome both the intrinsic high resistance of Mucorales to most of the antifungal drugs112,113 and the lack of specificity of the few active compounds available114,115 as symmetric 6mA is very uncommon in the human genome, and none of the MTA1c components is conserved in humans. While RNA 6mA methyltransferases have been reported to be crucial for humans116, future research is expected to provide information about the specificity of potential MTA1c inhibitors and to delve further into the DNA 6mA deposition mechanisms to ensure the efficient development of new antifungal drugs.
Methods
Genome assembly and annotation
DNA was extracted following a previously described phenol/chloroform extraction procedure57. The genome of R. microsporus ATCC 11559 was sequenced using PacBio Sequell IIe117. After a >10 kb Blue Pippin Size Selection, reads generated were filtered and processed with the JGI QC pipeline to remove artifacts. Mitochondrial DNA was assembled separately with the Circular Consensus Sequencing (CCS) reads using an in-house tool (assemblemito.py), used to filter the CCS reads and polished with two rounds of RACON (v. 1.4.13) racon [-u -t 36]118. The genome was annotated using the JGI Annotation pipeline and made available via the JGI fungal genome portal MycoCosm (https://mycocosm.jgi.doe.gov/Rhimi59_2/Rhimi59_2.home.html)119. Genome assembly completeness was assessed using BUSCO with Mucorales Odb10 dataset120 using Augustus gene prediction121 (Supplementary Table 1). The mitochondria-filtered CCS reads were then assembled with Flye (v. 2.9-b1768) (https://github.com/fenderglass/Flye) [-t 32 --pacbio-hifi] and subsequently polished with two rounds of RACON (v. 1.4.13) [-u -t 36]118.
6mA modification detection with PacBio SMRT Sequencing and data filtering
6mA base modifications were detected with the PacBio SMRT analysis platform (cromwell.workflows.pb_basemods). 10 ug of gDNA was sheared to >10 kb using Covaris g-Tubes. The sheared DNA was treated with exonuclease to remove single-stranded ends and DNA damage repair mix followed by end repair and ligation of blunt adapters using SMRTbell Template Prep Kit 1.0 (Pacific Biosciences). The library was purified with AMPure PB beads and size-selected with BluePippin (Sage Science) at >10 kb cutoff size. PacBio Sequencing primer was then annealed to the SMRTbell template library and sequencing polymerase was bound to them using Sequel Binding kit 3.0. The prepared SMRTbell template libraries were then sequenced on a Pacific Biosciences Sequel sequencer using v3 sequencing primer, 1 M v3 SMRT cells, and Version 3 sequencing chemistry with 1 × 600 sequencing movie run times. Raw reads were filtered using SFilter, to remove short reads and reads derived from sequencing adapters. Filtered reads were aligned to the reference genome of R. microsporus, using BLASR (1.5.0)122. Modified sites were then identified through kinetic analysis of the aligned DNA sequence data123. Detected 6mA sites were further filtered with a minimum 15x per-strand coverage to remove potential false positives and selecting only 6mA sites with >25 mQv.
6mA data analysis
Methylated Adenine Cluster (MACs) were defined as previously described34. Briefly, motifs are used to define MACs, where modification density within clusters and relative distance between methylated motifs are calculated. Here, the AT motif was used. Relative distance refers to how many unmodified motifs are permitted between modified motifs; lower relative distances are more efficient as they allow fewer unmethylated AT dinucleotides per cluster. The relationship between these two features (modification density and efficiency) is then used to determine optimal clustering. For each sample, relative distances from 1 to 40 were analyzed and the distance with the best score (density*efficiency) was used for genome-wide MAC identification. All MACs detected in all sequencing samples are indicated in Supplementary Data 5. Enrichment over genomic features was calculated as the ratio between the observed and expected 6mA. Promoters were determined using the transcriptional start site of each gene and adding ±200 nt of flanking sequence depending on the strand. Repeats were identified using RepeatModeler2124 (v2.0.4) with LTR structural identification enabled (-LTRstruct). The resulting output was used in RepeatMasker (v4.1.4) (http://www.repeatmasker.org) to detect every repeat in the genome with a slow search run (-s). Lower scoring matches whose domain partly (<80%) includes the domain of another match were removed from the final repeat list. To generate a more curated TE list, the RepeatModeler library was translated using EMBOSS transeq125 (v5.0.0) in all possible six frames with -clean option enabled and scanned using InterproScan126. Only those consensus sequences that reported hits with domains related to TE (reverse transcriptase, endonuclease, GAG protein, aspartic proteinase, integrase, RNAse) were used to construct a refined input library to run RepeatMasker (-s) with a strict cutoff value (4500) and with -no_is, -nolow and -norna options enabled. The rest of the genomic sequence not included in the gene body, promoters, or repeats were considered intergenic. 6mA profile graphs were generated by computing 6mA frequency over equal-size bins using deepTools (v3.1)127. Symmetry density plots were generated using the methylation ratio for each site and plotted using a scatter of points with kernel density estimations in two dimensions in R with the ggplot2 library128. To analyze the sequence context, we extracted ±6 nucleotides of each 6mA site. Those sequences were then analyzed using MEME-ChIP129. RNApol III genes (5S rRNAs, tRNAs, and U6 snRNAs) and RNApol I genes (rRNAs) were detected in the R. microsporus ATCC 11559 genome using tRNAscan-SE130 (v1.3.1), RNAmmer131 (v1.2), and Rfam (v14.8)132. For GO/KOG enrichment analysis, genes harboring at least one MAC (with at least 10 methylated sites) 100 nt upstream the TSS and 400 nt dowsntream the TSS. GO and KOG annotations were retrieved from the R. microsporus v2 annotation generated in this work and available at Mycocosm (https://mycocosm.jgi.doe.gov/Rhimi59_2/Rhimi59_2.home.html). A P value of 0.05 was used as a cutoff for the identification of enriched terms (Fisher’s exact test).
DNA extraction and library preparation for WGBS
Genomic DNA was extracted as described previously32. 1 µg of DNA was first sheared to ~500 bp using sonication (Covaris LE220), then subject to end repair, A-tailing, and ligation of Methylated Indexed Illumina Adaptor (IDT). Bisulfite conversion of unmethylated cytosine to uracil and clean-up of adapter-ligated DNA was done using Zymo EZ DNA Methylation-Lightening Kit (Zymo Research). Converted libraries were amplified using 10 cycles of PCR and purified using AMPure Purification (Beckman Coulter). Final libraries were sequenced with Illumina HiSeq-2500. Mitochondrial DNA was used as a negative control to determine the efficiency of the sodium bisulfite conversion rate.
WGBS data analysis
Raw fastq file reads were filtered and trimmed using the JGI QC pipeline. Using BBDuk (https://sourceforge.net/projects/bbmap/), raw reads were evaluated for artifact sequence by kmer matching (kmer = 23), allowing 1 mismatch, and the detected artifacts were trimmed from the 3’ end of the reads. Quality trimming was performed using the phred trimming method set at Q6. Reads under the length threshold were removed (minimum length 25 bases or 1/3 of the original read length). Finally, one base off the right of the reads was trimmed to prevent the creation of completely contained read pairs. Reads were mapped to the R. microsporus reference genome and methylation calling was done using Bismark133. For read mapping, bowtie1134 was used with seed length set to 50, maximum mismatches in the seed set to 1, and maximum insert size set to 1000. Reads were deduplicated and reads that mapped to multiple locations were removed. Methylation detection was performed using the bismark_methylation_extractor tool from the Bismark package. The average coverage was 69.70x and the mitochondrial genome was used to calculate the bisulfite conversion efficiency (99.85%). Weighted methylation levels by dividing the total number of methylated reads by the total number of methylated plus unmethylated reads for each sequence context135. Methylated sites were filtered by a minimum of 5x coverage and 10% methylation ratio. 5mC profile graphs metaplots were generated, including 1 kb flanking regions over equal size bins using deepTools (v3.1)127.
RNA extraction, library preparation, and RNA-seq analysis
Total RNA was isolated by using the RNeasy Mini kit (Qiagen) and subjected to a DNase treatment (Sigma, On-Column DNaseI treatment set). Three biological replicates were analyzed for each expression experiment. Plate-based RNA sample prep was performed on the PerkinElmer Sciclone NGS robotic liquid handling system using Illumina’s TruSeq Stranded mRNA HT sample prep kit utilizing poly-A selection of mRNA following the protocol outlined by Illumina in their user guide: support.illumina.com/sequencing/sequencing_kits/truseq-stranded-mrna.html, and with the following conditions: total RNA starting material was 1 ug per sample and 8 cycles of PCR was used for library amplification. Both types of libraries were quantified using KAPA Biosystem’s next-generation sequencing library qPCR kit (Roche) and run on a Roche LightCycler 480 real-time PCR instrument. The quantified library was then multiplexed with other libraries, and the pool of libraries was then prepared for sequencing on the Illumina NovaSeq 6000 sequencing platform using NovaSeq XP v1.5 reagent kits, S4 flow cell, following a 2×150 indexed run recipe. Raw fastq file reads were filtered and trimmed using the JGI QC pipeline. BBDuk (https://sourceforge.net/projects/bbmap/) was used for artifact sequence by kmer matching (kmer = 25), allowing 1 mismatch, and the detected artifacts were trimmed from the 3’ end of the reads. RNA spike-in reads PhiX reads and reads containing any Ns were removed. Quality trimming was performed using the phred trimming method set at Q6. Finally, following trimming, reads under the length threshold were removed (minimum length 25 bases or 1/3 of the original read length). For reference genome annotation, de novo transcriptome assembly was performed using Trinity (v2.11.0)136 with the --normalize_reads (In-silico normalization routine) and --jaccard_clip (Minimizing fusion transcripts derived from gene-dense genomes) options. For expression analysis, filtered reads from each library were aligned to the reference genome using HISAT2 (v2.2.0)137 Raw counts were calculated with featureCounts138. These raw counts were used to determine differentially expressed genes with DESeq2 (v1.30.0)139 (P < 0.05 as cutoff). Counts files were also used to generate FPKM and TPM information. Strand-specific coverage bigWigs were generated using deepTools (v3.1)127. All differentially expressed genes across all conditions are indicated in Supplementary Data 1.
Mononuclesomal DNA purification and MNase-seq data analysis
R. microsporus mycelium was crosslinked with formaldehyde at a final concentration of 1% for 20 min in a shaker (BRAUN BIOTECH Certomat R) at 80 rpm. Crosslinking was quenched by adding glycine at a final concentration of 135 mM for 10 min. Mycelium samples were washed twice with PBS 1X and flash-frozen with liquid nitrogen. 20 mg of mycelia were ground with mortar and pestle using liquid nitrogen until a fine powder was obtained. The powder was resuspended in 1 mL of MNase Lysis Buffer (15 mM Tris-HCl pH 7.5, 250 mM sucrose, 60 mM KCl, 15 mM Tris-HCl pH 7.5, 3 mM MgCl2, 2 mM CaCl2, and 1% Tritton X-100) and 5 µl of protease inhibitor cocktail (Merck). Tubes were placed on ice and vortexed for 10 s with 20 s rest for 3 min. For each treatment, we used 20 U of micrococcal nuclease (Thermo Scientific) for 15 min at 30 °C. Digestions were stopped with 100 µl of MNase Stop Buffer (80 mM EDT and 1.5% SDS). Tubes were centrifuged for 5 min at 2700 × g. The supernatant was transferred to a new tube and incubated for 10 min at 95 °C for reversal crosslinking. Samples were further treated with Proteinase K (Merck) at a final concentration of 1 mg/mL and RNAse A (Merck) at a final concentration of 1 mg/mL and NaCl at a final concentration of 0.1 M overnight at 65 °C. DNA was purified by adding 1/2 volume phenol and 1/2 volume of chloroform:isoamyl alcohol (24:1). After centrifugation (2 min at 16,000 × g), the aqueous phase was transferred to a new tube and 1 volume of chloroform:isoamyl alcohol (24:1) was added. After centrifugation (2 min at 16,000 × g), the aqueous phase was transferred to a new tube and 1/10 volume of sodium acetate 3 M and 1 volume of ethanol (96%) were added. Tubes were incubated at −20 °C for 2 h. DNA was precipitated by centrifugation at 4 °C for 10 min at 16,000 × g. DNA pellet was washed twice with 70% ethanol and dried at room temperature for 10 min. DNA was resuspended with 30 µl of ddH20 and submitted to electrophoresis at 100 V for 1 h in a 1% agarose gel. The band corresponding to mononucleosomic DNA was sliced from the gel and purified using the Gene Jet PCR Purification Kit (Thermo Scientific). Purity and DNA quantity were analyzed with Nanodrop 2000 (Thermo Scientific) and Qubit 4 (Thermo Scientific). DNA samples were sent to Novogene, which prepared and sequenced the libraries (Illumina Novaseq 6000, 150 bp paired-end reads). Raw reads were analyzed using FastQC (v0.11.8) (http://www.bioinformatics.babraham.ac.uk/projects/fastqc/) and adapters were trimmed using Cutadapt (v4.0)140. Filtered reads were then mapped to the R. microsporus v2 reference genome using Bowtie2 (v2.4.2)134. Sorted BAM files were then processed with DANPOS (v2.1.1)141 with dpos function (-m 1) to call nucleosomes across the genome, including nucleosome coordinates and center, and fuzziness degree for each of them (Supplementary Data 2). Coverage bigwig files were visualized using the Integrative Genomics Viewer (IGV) (v2.12.13)142. Nucleosome occupancy was normalized by average genome coverage and dyads (nucleosome center) were normalized by the genomic of dyads density. Profiles were generated from dyads information and plotted using computeMatrix, plotProfile, and plotHeatmap functions from deepTools (v3.1)127. For the nucleosome-linker plot, nucleosome and linker regions were divided into 20 bins of equal size, and 6mA frequency was calculated for each bin using deepTools (v3.1)127. Fuzziness analysis was performed by selecting nucleosomes located at -500 nt upstream of the TSS and 2000 nt downstream of the TSS. Nucleosomes were classified into 10 groups according to their fuzziness score.
ChIP-seq experiments
R. microsporus mycelium was crosslinked with formaldehyde (1%) and quenched with glycine in the same way as for mononucleosomal DNA purification and previously published ChIP procedures79. After flash-freezing with liquid nitrogen, 180 mg of frozen mycelium per ChIP experiment was ground until a fine powder was obtained and resuspended in 900 µl ChIP Lysis buffer (50 mM HEPES pH 7.4, 150 mM NaCl, 1% Tritton X-100, 0.1% DOC and 1 mM EDTA) and 10 µl of protease inhibitor cocktail. The lysates were sonicated using a Bioruptor Pico sonicator (Diogenode) with 30 s ON/OFF pulses for 25 cycles in a thermostated bath (4 °C). After sonication, samples were centrifuged at 12.000 × g for 8 min at 4 °C and transferred and divided into 3 tubes: 100 µl were frozen at −20 °C for Input DNA control, 400 µl were used for antibody immunoprecipitation (IP) and 400 µl were used for beads only control. Beads were equilibrated with Dilution Buffer (50 mM HEPES pH 7.4, 150 mM NaCl, and 1 mM EDTA). For H3K9me3, H3K4me3, H3K27me3 and H2A.Z experiments, 50 µl of Dynabeads protein A magnetic beads (ThermoFisher) equilibrated beads were added to mock beads only samples and another 50 µl were conjugated with the selected ChIP-grade antibodies (Anti-H3K9me3: Abcam #ab176916, clone EPR16601, lot number GR3218257-27; anti-H3K4me3: Abcam #ab213224, EPR20551-225, lot number GR3447206-1; anti-H3K27me3: Abcam #ab192985, clone EPR18607, lot number GR3306698-5; and anti-H2A.Z: Abcam #188314, clone EPR18090, lot number GR3448913-1) at 4 °C for 2 h. After washing again with dilution buffer, antibody-conjugated Dynabeads were added to the IP samples. Both IP and bead control were incubated overnight at 4°C. After incubation, the beads were washed twice with 1 mL of the Low Salt Wash Buffer (20 mM Tris pH 8.0, 150 mM NaCl, 0.1% SDS, 1% Triton X-100, and 2 mM EDTA), twice with 1 mL of High Salt Wash Buffer (20 mM Tris pH 8.0, 500 mM NaCl, 0.1% SDS, 1% Triton X-100 and 2 mM EDTA), once with LiCl Wash Buffer (10 mM Tris pH 8.0, 1 mM EDTA, 0.25 M LiCl, 1% NP40, and 1% DOC), and finally once with TE (10 mM Tris pH 8.0, and 1 mM EDTA). DNA was eluted from beads by adding 250 µl of TES Buffer (50 mM Tris pH 8.0, 10 mM EDTA, 1% SDS) and incubating for 10 min at 65 °C (twice). The final eluted 500 µl and the Input DNA were submitted to reversal crosslinking by incubating overnight at 65 °C with 0.2 M NaCl. Samples were treated with Proteinase K (Merck) at a final concentration of 1 mg/mL and RNAse A (Merck) at a final concentration of 1 mg/mL for 3 h at 65 °C. DNA was purified by adding 1/2 volume phenol and 1/2 volume of chloroform:isoamyl alcohol (24:1). After centrifugation (2 min at 16,000 × g), the aqueous phase was transferred to a new tube and 1 volume of chloroform:isoamyl alcohol (24:1) was added. After centrifugation (2 min at 16,000 × g), the aqueous phase was transferred to a new tube and 1/10 volume of sodium acetate 3 M, 1 volume of ethanol (96%), and 20 µg of glycogen (ThermoFisher) were added. The tubes were incubated at −20 °C overnight, and DNA was precipitated by centrifugation at 4 °C, 10 min at 16,000 × g. DNA was washed twice with 70% ethanol and dried at room temperature for 10 min. DNA was resuspended with 30 µl of ddH20. Purity and DNA quantity were analyzed with Nanodrop 2000 (Thermo Scientific) and Qubit (Thermo Scientific). For P1-mCherry ChIP experiments, the mycelia of the UM127 strain were processed as described above. 25 µl of Red Fluorescent Protein (RFP)-Trap Magnetic Agarose (MA) beads (Chromotek) were added to IP samples, and 25 µl of MA beads (Chromotek) were added to the beads-only control sample. The subsequent steps were performed as described above.
ChIP-seq data analysis
Purified DNA samples were used to construct libraries and to be sequenced (Illumina NovaSeq 6000, 150 bp paired-end reads) by Novogene. Raw reads were checked with FastQC (v0.11.8) (http://www.bioinformatics.babraham.ac.uk/projects/fastqc/) and adapters were trimmed using Cutadapt (v4.0)140. Trimmed reads were mapped to the R. microsporus v2 reference genome using bwa-mem for medium and long reads with default parameters (v0.7.17)143. BAM files were sorted and deepTools bamCoverage function127 (bin size = 10, CPM normalization) was used to visualize and manually inspect tracks on IGV (v2.12.13)142. bamCompare function (bin size = 10, CPM normalization) was used to compute ratios between IP and Input samples. The resulting bigwig or bedgraph files were used for downstream analysis. To assess the specificity of the immunoprecipitations, the mock beads-only signal was subtracted, and the resulting tracks were manually analyzed on IGV (v2.12.13)142. MACS2 software (v2.2.7.1)144 was used to call peaks for each ChIP-seq experiment (effective genome size = 27394811, FDR cut-off = 0.05) on narrow peaks mode for H3K4me3, H2A.Z, and P1 and in broad peaks (--broad) for H3K9me3. The MEME-ChIP suite129 was used to analyze the motifs of the enriched peaks.
DAP-seq
P1 binding sites were analyzed using DNA Affinity Purification sequencing (DAP-seq) following a previously established procedure80,81 with minor modifications145. Briefly, gDNA from R. microsporus was fragmented to an average size of 150 bp (Covaris LE220-Plus) and sequencing libraries were prepared using the KAPA HyperPrep kit (Roche). In parallel, Halo-tagged transcription factor proteins were cloned into pIX_HALO plasmid backbone, confirmed via sequencing, and used for PCR amplification to yield linear PCR fragments containing the Halo-TF fusion protein driven by a T7 promoter. The PCR product was purified with SPRI beads, and the correct size of each was verified using a Fragment Analyzer (Agilent Technologies). For each DAP-seq reaction, each expressed protein was used, along with 20 ng of the prepared fragment library and 2000 ng of salmon sperm DNA to mitigate non-specific binding. The resultant DAP-seq libraries were pooled for sequencing on a NovaSeq using the S4 flowcell (Illumina). Reads were mapped to the R. microsporus reference genome (https://mycocosm.jgi.doe.gov/Rhimi59_2/Rhimi59_2.home.html) using Bowtie2134. MACS3144 (v3.0.0a6) was used to call the peaks and motifs were generated using MEME129 with the 100 most significant peaks, as scored by signal value in column 7 of the narrowPeak files.
Phylogenetic analysis and ortholog search
Proteomes of representative species were retrieved from the Joint Genome Institute (JGI) MycoCosm genome portal119 and Uniprot146. Sequences of T. thermophila Mta1 (Q22GC0_TETTS), Mta9 (I7MIF9_TETTS), p1 (Q22VV9_TETTS), and p2 (I7M8B9_TETTS) proteins were queried against the selected proteomes using iterative HMMER jackhmmer searches (E value 1 × 10-3) (v3.3.2) (http://hmmer.org). Hits were collapsed using CD-HIT (Identity cut-off -c 0.97) to remove redundant sequences (v4.8.1)147. A reciprocal BLASTp search (v2.10.1)148 was conducted and sequences that failed to produce a hit were discarded. An additional search using Pfam-A database149 using HMMER hmmscan (v3.3.2) (http://hmmer.org) served to remove hits that lacked the MT-A70 (Mta1 and MTA9 searches) or the Homeobox/Myb-like domains (p1 and p2 searches) (Supplementary Table 7). The remaining MT-A70 candidates were aligned using MUSCLE (v3.8.31)150, and a phylogenetic tree was obtained using the approximate maximum-likelihood method with the JTT substitution model151 with a bootstrap procedure of 100 iterations with MEGA X (v10.2.4)152 (Supplementary Table 7 and Supplementary Fig. 11). The species tree was generated after analyzing the 36 proteomes with OrthoFinder (Inflation factor, -I 1.5)153. We identified 63502 protein orthogroups, of which 775 had all species present. Multiple sequence alignment was performed using MAFFT154 and the tree was obtained using RAxML155 (PROTGAMMAWAGF substitution model) with 100 bootstrap replicates. For P1 alignment, all the selected protein candidates were aligned using MUSCLE (v3.8.31)150, and the result of the alignment was visualized using Jalview (v2.11.2.6)156. The secondary structure of R. microsporus P1 protein was predicted using Phyre2 program (http://www.sbg.bio.ic.ac.uk/phyre2). Homolog search was performed with 65 eukaryotic MTA-70 sequences as queries (listed in Pan et al.40 supplementary materials) against NCBI non-redundant protein sequence database (nr). Sequences of identified homologs aligned using MAFFT154, trimmed manually down to the domain region, and clustered with cd-hit (70% sequence identity)147. Obtained representatives were subsequently used as queries for blastp157 searches against 183 fungal genomes (blastp, e value 0.001) (Supplementary Data 4). Sequence alignments were calculated with Mafft (v7) (Katoh and Standley 2013) (localpair, maxiterate 100), analyzed in Seaview158 and trimmed with TrimAl (gappyout)159. Phylogenetic trees were calculated in IqTree2160 with automated model selection (-B 1000 -alrt 1000, selected model according to BIC LG + R4). A phylogenomic tree was built with OrthoFinder153 (with mmseqs) for a set of 41 taxa with Drosophila melanogaster and Monosiga brevicolis as an outgroup. Phylogenetic trees were visualized in iTOL161. For sequence accessions, assembly IDs see Supplementary Data 4.
Protein structures of Ma1-Mta9a and Mta1-Mta9b were predicted using AlphaFold2 through ColabFold interface162, visualized in PyMol (https://pymol.org/). HMM logos of selected sequence positions were drawn with WebLogo 3163.
Yeast two-hybrid assay
To analyze the interactions between MTA1c proteins, total RNA from R. microsporus was extracted using the RNeasy Mini kit (Qiagen) and submitted to a DNase treatment (Sigma, On-Column DNaseI treatment set). cDNA was synthesized using 1 μg of total RNA with the iScriptTM cDNA synthesis kit (Biorad). The coding sequence of mta1, mta9, and p1 were amplified using specific primers (Suplementary Table 8 and Supplementary Fig. 14) and cloned in both pACTII (mta1) and pBD-Gal4 (mta1, mta9, and p1) plasmids (Supplementary Table 7). These plasmids were sequenced through Sanger sequencing to ensure the absence of mutations in the coding sequences. S. cerevisiae pJ69–4a strain was transformed with the plasmids generated. Yeast transformations were plated on SD without tryptophan and leucine (SD-WL) and grown for 72 h at 30 °C. Individual colonies from SD-WL plates were grown on SD without tryptophan, leucine, and histidine and supplemented with 2.5 mM of 3-amino-1,2,4-triazole (SD-WLH + 3AT) and SD without tryptophan, leucine, and adenine (SD-WLA) to detect weak and strong interaction, respectively. The plates to assay the interaction were incubated for 10 weeks at 30 °C.
HPLC-MS/MS analysis, 6mA dot blot, and DpnI/II digestion assay
Genomic 6mA levels were quantified with HPLC followed by mass spectrometry (MS) as we did previously32. Genomic DNA for dot blot assay was quantified using a Qubit 4 (Thermo Scientific) and denaturalized during 5 min at 95 °C. 2 µl drops containing 100, 50, and 10 ng were placed in an Amersham Hybond-N+ membrane (GE Healthcare, RPN203B). DNA was crosslinked with 1200 microjoules (50 s), and the membrane was washed with PBS + 0.002% Tween-20 to remove unbound DNA. The membrane was blocked for 1 h with 5% non-fat milk and subsequently incubated with the primary antibody anti-6mA (1:1000) (Synaptic Systems #202003, polyclonal. lot number 2116) overnight at 4°C. After washing with PBS + 0.002% Tween-20, HRP labeled secondary antibody was added (1:5000). The Amersham ECL detection kit (RPN2108) was used for peroxidase reaction and results visualization. For DpnI/II digestion assays, we added 5U of DpnI, 5U of DpnII, and 0.5 µl of ddH20 (Untreated) to 1 µg of gDNA. Digestions were incubated for 4 h at 37 °C and the product of this digestion was separated in a 1.5% agarose gel at 100 V for 3 h.
qRT-PCR
The expression of the zrt1 gene (ID: 1926170) was analyzed by qRT-PCR at different Zinc (ZnSO4) (0 mg/L, 1 mg/L, 10 mg/L, and 20 mg/L) with specific primers (qRT-PCR_zrt1_F and qRT-PCR_zrt1_R) (Supplementary Table 8). Total RNA was extracted using the RNeasy Mini kit (Qiagen) and submitted to a DNase treatment (Sigma, On-Column DNaseI treatment set) and cDNA was synthesized from 1 μg of total RNA using the iScriptTM cDNA synthesis kit (Biorad). cDNA amplification was carried out in triplicate using a reaction mixture containing 2x Power SYBR® Green Master Mix (Applied biosystem, Waltham, MS, USA), 150 nM of zrt1-specific primers, and 100 ng of cDNA. The real-time PCR was carried out using the QuantStudioTM 5 real-time PCR system (Applied biosystem) according to the established experiment template in the equipment. Briefly, 10 µl reactions were denatured at 95 °C during 2 min and amplified during 40 cycles (95 °C—15 s for denaturalization and 60 °C—1 min for hybridization, elongation, and fluorescence detection) The melting curve and non-template controls were also measured to discern non-specific amplifications. The relative expression of zrt1 was normalized with the amplification levels of the constitutively expressed Ribosomal protein L3 gene (ID:1909258) with qRT-PCR_rL3_F and qRT-PCR_rL3_R primers (Supplementary Table 8) and calculated using the 2−∆∆CTmethod.
Mutant strain generation and validation
R. microsporus mutant strains were generated following previously established procedures57,58. For mta1 (ID: 1822122), mta9 (ID: 1795109), and p1 (ID: 1926449) disruption, a template DNA containing the pyrF (ID: 1869696) gene and flanked by 38 nt homology regions was used to disrupt the targeted gene. Recombination was enabled through the directed double-strand break (DSB) provoked by Cas9 in a uracil-auxotrophic strain (UM1)57. Designed crRNAs (Alt-RTM CRISPR-Cas9 crRNA) used to target the CRISPR/Cas9 system were purchased from IDT (Supplementary Table 9). crRNAs were coupled with tracRNA (Alt-RTM CRISPR-Cas9 tracRNA, IDT) to form the gRNA, which was assembled in vitro with Cas9 (Alt-RTM S.p. Cas9 nuclease, IDT) to form the ribonucleoprotein complex. For the mta1 strain with regulated expression, this cassette contained the Pzrt1 promoter region and the mta1 gene. This construction was integrated into the leuA gene (ID: 1891741), generating a leucine auxotrophy that was complemented by disrupting the endogenous mta1 gene with a template DNA containing the leuA marker following the same strategy as described above. For Mta1 and P1 tagging experiments, the homologous recombination (HR) DNA template was generated using the 3’ regions of the gene (downstream the Cas9 cleavage site), the mCherry protein, and the pyrF marker. To avoid the gRNA/Cas9 complex used to provoke the DSB can recognize and cleave the DNA used as the template, we produced synonymous nucleotide substitutions in the seed sequence. The uracil-auxotrophic strain (UM1) was used as the recipient strain. Gene disruptions were validated by PCR using a primer that hybridizes in the marker gene and another primer upstream of the recombination region. Homo/heterokaryosis was validated by PCR using primers flanking the disruption site that amplifies both WT and mutant nuclei if present. All primers used for construct amplification and PCR validation are listed in Supplementary Table 8. DNA was extracted as described above, and all PCR amplifications were performed with the Herculase II Fusion DNA polymerase (Agilent), adapting the annealing temperature to each primer pair following the manufacturer’s recommendations.
Fungal growth conditions
All the fungal strains used and generated in this work derive from Rhizopus microsporus ATCC 11559. When specified, media was supplemented with uridine (200 mg/L). Transformants of the auxotrophic strain UM1 strain57 using the pyrF marker were grown in Minimal Media with Casamino acids (MMC)164. Transformants using the leuA marker were grown in Yeast Nitrogen Base (YNB)165. During transformation, electroporated protoplasts were resuspended in ice-cold Yeast Peptone Glucose (YPG) media + 0.5 M Sorbitol for 90 min and then centrifugated at 91 × g and resuspended in YNB media +0.5 M Sorbitol. All strains were grown at 30 °C under light or dark conditions. To asses the effect of ZnSO4 in growth, 6mA and gene expression regulation in R. microsporus ATCC 11559, as well to analyze the expression of the zrt1 gene and the growth of UM64 and UM67 strains YNB was supplemented with 20 mg/L or 10 mg/L of ZnSO4. Escherichia coli DH5α strain (Thermo Fisher Scientific) was used for all cloning experiments. Yeast cultures were grown at 30°C in YPAD complex medium or dropout Synthetic Defined (SD). Yeast transformations with PEG3350-lithium acetate were performed following previously established procedures166 with 250 ng of plasmid DNA.
Statistical analysis
Statistical details are detailed in the Results, Figures, and Figure legends, including the number of biological and technical replicates as well as the dispersion and precision measures (mean and SD). Statistical analyses were performed using GraphPad Prism 9 (https://www.graphpad.com). Data normality was analyzed using the Shapiro-Wilk normality test with a significance level (alpha) of 0.05. An ANOVA of a single factor was used to determine statistically significant differences between groups of data with normal distribution, assuming a significance level of 95% (P < 0.05), followed by Tuckey’s’ HSD post-hoc test. Direct comparisons between the expression levels of methylated and unmethylated genes were analyzed using Welch’s test. Pearson correlation factors, P values cut-off for Fisher exact tests, and Welch’s tests for direct comparisons are indicated in the respective figure legends. Fisher’s exact tests to determine whether two genomic intervals (ChIP-seq peaks and genomic features) were performed using BEDtools fisher tool (v2.32.1)167. Differences in the overlapping between MACs and H3K4me3 and H3K9me3 peaks compared to random regions we performed by generating random peaks considering the average number of peaks of each modification and their average length. A chi-square test was performed to test for statistically significant differences assuming a significance level of 95% (P < 0.05). 6mA profile differences were analyzed by calculating Wilcoxon rank test for each bin and the P value corrected for multiple comparisons was calculated using deepStats package (https://github.com/gtrichard/deepStats) (v0.3.31). Pearson correlation coefficient (r) was calculated to assess relationships between MACs presence and FPKM values for methylated genes a P < 0.05 was set as the cutoff to determine statistically significant differences. To compare differences between genes grouped by FPKM and H3K4m3 occupancy (log2) the ratio of these two parameters was computed for each gene and the two sets of genes (MAC and no MAC) were compared using a nonparametric Mann-Whitney test. a P < 0.05 was set as the cutoff to determine statistically significant differences. Nucleosome profile distributions and fuzziness frequency distribution differences were evaluated using the Kolmogorov-Smirnov test with a P < 0.05 was set as the cutoff to determine statistically significant differences.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Supplementary information
Description of Additional Supplementary Files
Source data
Acknowledgements
This research was supported by the grant PID2021-124674NB-I00 to F.E.N. and V.G., funded by MICIU/AEI/10.13039/501100011033 and by ERDF/EU. This work was also supported by the National Science Centre grant #2021/41/B/NZ2/02426 to A.M. The work (10.46936/10.25585/60001127) was conducted by the US Department of Energy Joint Genome Institute (https://ror.org/04xm1d337), a DOE Office of Science User Facility, was supported by the Office of Science of the US Department of Energy under Contract No. DE-AC02-05CH11231. We would like to thank Kamil Steczkiewicz for assistance with figure preparation.
Author contributions
C.L. conducted most of the R. microsporus experiments, prepared the R. microsporus material for sequencing, performed bioinformatic analysis of the results, prepared the figures and tables, designed and coordinated the project, and wrote the manuscript with significant input from S.J.M., A.M., I.V.G., T.E.P., F.E.N., and V.G. S.J.M. participated in the assembly of genomes and bioinformatic analyses. J.F.M. and P.C.-M. participated in the generation of R. microsporus mutant strains. L.B. participated in DAP-seq data analysis. J.A.P.-R. conducted yeast-two hybrid assays. A.M. analyzed the phylogeny. K.L. participated in the assembly of genomes. A.L. conducted transcriptomic sequencing. Y.Z. performed DAP-seq. J.G. conducted bisulfite sequencing. V.N. managed the project. E.N. managed the project and provided materials. I.V.G. supervised and coordinated the project. F.E.N. analyzed the results and supervised the study. V.G. analyzed the results and designed, supervised, and coordinated the project.
Peer review
Peer review information
Nature Communications thanks Rhys Farrer, who co-reviewed with Grace Paul and Diana Tamayo; Stefan Kusch, Soon-Jae Lee, and Frank Lyko for their contribution to the peer review of this work. A peer review file is available.
Data availability
The raw sequence data that support the findings of this study have been deposited in the Sequence Read Archive (SRA) under the accession numbers SRP496792, SRP496793, SRP496795, SRP496796, SRP496797, SRP496798, SRP496799, SRP496800, SRP496816, SRP496817, SRP496818, SRP496819, SRP496820, SRP496821, SRP496822, SRP496823, SRP496824, SRP496825, SRP496830, SRP496831, SRP496832, SRP496833, SRP497170, SRP497171, SRP497172, and SRP536317, and in the Gene Expression Omnibus (GEO) under the accession numbers GSE285378, GSE276938, and GSE276939. The Whole Genome Shotgun project for R. Microsporus ATCC 11559 has been deposited at DDBJ/ENA/GenBank under the accession JBFXLZ000000000. The version described in this paper is version JBFXLZ010000000 [https://www.ncbi.nlm.nih.gov/nuccore/JBFXLZ000000000.1/]. The HPLC/MS raw data has been deposited in Zenodo under the accession number 15194697 (10.5281/zenodo.15194696; https://zenodo.org/records/15194697). Source data are provided with this paper.
Competing interests
The authors declare no competing interests.
Footnotes
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Contributor Information
Francisco E. Nicolás, Email: fnicolas@um.es
Victoriano Garre, Email: vgarre@um.es.
Supplementary information
The online version contains supplementary material available at 10.1038/s41467-025-59170-x.
References
- 1.Fusco, G. & Minelli, A. Phenotypic plasticity in development and evolution: facts and concepts. Philos. Trans. R. Soc. B Biol. Sci.365, 547–556 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Duncan, E. J., Gluckman, P. D. & Dearden, P. K. Epigenetics, plasticity, and evolution: how do we link epigenetic change to phenotype? J. Exp. Zoöl. Part B Mol. Dev. Evol.322, 208–220 (2014). [DOI] [PubMed] [Google Scholar]
- 3.Adhikari, S. & Curtis, P. D. DNA methyltransferases and epigenetic regulation in bacteria. FEMS Microbiol. Rev.40, 575–591 (2016). [DOI] [PubMed] [Google Scholar]
- 4.Casadesús, J. & Low, D. Epigenetic gene regulation in the bacterial world. Microbiol. Mol. Biol. Rev.70, 830–856 (2006). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Zhao, J. et al. Roles of adenine methylation in the physiology of Lacticaseibacillus paracasei. Nat. Commun.14, 2635 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Wahnon, D. C., Shier, V. K. & Benkovic, S. J. Mechanism-based inhibition of an essential bacterial adenine DNA methyltransferase: rationally designed antibiotics. J. Am. Chem. Soc.123, 976–977 (2001). [DOI] [PubMed] [Google Scholar]
- 7.Smith, Z. D. & Meissner, A. DNA methylation: roles in mammalian development. Nat. Rev. Genet.14, 204–220 (2013). [DOI] [PubMed] [Google Scholar]
- 8.Zemach, A., McDaniel, I. E., Silva, P. & Zilberman, D. Genome-wide evolutionary analysis of eukaryotic DNA methylation. Science328, 916–919 (2010). [DOI] [PubMed] [Google Scholar]
- 9.Lewis, Z. A. et al. Relics of repeat-induced point mutation direct heterochromatin formation in Neurospora crassa. Genome Res.19, 427–437 (2009). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Eden, S., Hashimshony, T., Keshet, I., Cedar, H. & Thorne, A. W. DNA methylation models histone acetylation. Nature394, 842–842 (1998). [DOI] [PubMed] [Google Scholar]
- 11.Hashimshony, T., Zhang, J., Keshet, I., Bustin, M. & Cedar, H. The role of DNA methylation in setting up chromatin structure during development. Nat. Genet.34, 187–192 (2003). [DOI] [PubMed] [Google Scholar]
- 12.Ooi, S. K. T. et al. DNMT3L connects unmethylated lysine 4 of histone H3 to de novo methylation of DNA. Nature448, 714–717 (2007). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Tamaru, H. & Selker, E. U. A histone H3 methyltransferase controls DNA methylation in Neurospora crassa. Nature414, 277–283 (2001). [DOI] [PubMed] [Google Scholar]
- 14.Madhani, H. D. Unbelievable but true: epigenetics and chromatin in fungi. Trends Genet.37, 12–20 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Mishra, P. K., Baum, M. & Carbon, J. DNA methylation regulates phenotype-dependent transcriptional activity in Candida albicans. Proc. Natl. Acad. Sci. USA108, 11965–11970 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Singer, M. J., Marcotte, B. A. & Selker, E. U. DNA methylation associated with repeat-induced point mutation in Neurospora crassa. Mol. Cell. Biol.15, 5586–5597 (1995). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Lewis, Z. A. et al. DNA methylation and normal chromosome behavior in neurospora depend on five components of a histone methyltransferase complex, DCDC. PLoS Genet.6, e1001196 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Selker, E. U. et al. Induction and maintenance of nonsymmetrical DNA methylation in Neurospora. Proc. Natl. Acad. Sci. USA99, 16485–16490 (2002). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Kouzminova, E. & Selker, E. U. dim‐2 encodes a DNA methyltransferase responsible for all known cytosine methylation in Neurospora. EMBO J.20, 4309–4323 (2001). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Aramayo, R. & Selker, E. U. Neurospora crassa, a model system for epigenetics research. Cold Spring Harb. Perspect. Biol.5, a017921 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Selker, E. U. et al. DNA methylation and control of genome organization in Neurospora crassa. Gene74, 109–111 (1988). [DOI] [PubMed] [Google Scholar]
- 22.Selker, E. U. DNA methylation and chromatin structure: a view from below. Trends Biochem. Sci.15, 103–107 (1990). [DOI] [PubMed] [Google Scholar]
- 23.Brosch, G., Loidl, P. & Graessle, S. Histone modifications and chromatin dynamics: a focus on filamentous fungi. FEMS Microbiol. Rev.32, 409–439 (2008). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Wang, L. et al. A histone H3K9 methyltransferase Dim5 mediates repression of sorbicillinoid biosynthesis in Trichoderma reesei. Microb. Biotechnol.15, 2533–2546 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Osorio-Concepción, M., Cristóbal-Mondragón, G. R., Gutiérrez-Medina, B. & Casas-Flores, S. Histone deacetylase HDA-2 regulates Trichoderma atroviride growth, conidiation, blue light perception, and oxidative stress responses. Appl. Environ. Microbiol. 83, e02922-16 (2017). [DOI] [PMC free article] [PubMed]
- 26.Millar, C. B. & Grunstein, M. Genome-wide patterns of histone modifications in yeast. Nat. Rev. Mol. Cell Biol.7, 657–666 (2006). [DOI] [PubMed] [Google Scholar]
- 27.Colabardini, A. C. et al. Chromatin profiling reveals heterogeneity in clinical isolates of the human pathogen Aspergillus fumigatus. PLoS Genet.18, e1010001 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Wijayawardene, N. et al. Outline of Fungi and fungus-like taxa–2021. Mycosphere13, 53–453 (2022). [Google Scholar]
- 29.Amses, K. R. et al. Diploid-dominant life cycles characterize the early evolution of Fungi. Proc. Natl. Acad. Sci. USA119, e2116841119 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Spatafora, J. W. et al. A phylum-level phylogenetic classification of zygomycete fungi based on genome-scale data. Mycologia108, 1028–1046 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Galindo, L. J., López-García, P., Torruella, G., Karpov, S. & Moreira, D. Phylogenomics of a new fungal phylum reveals multiple waves of reductive evolution across Holomycota. Nat. Commun.12, 4973 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Osorio-Concepción, M., Lax, C., Navarro, E., Nicolás, F. E. & Garre, V. DNA methylation on N6-adenine regulates the hyphal development during dimorphism in the early-diverging fungus Mucor lusitanicus. J. Fungi7, 738 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Navarro-Mendoza, M. I., Pérez-Arques, C. & Heitman, J. Heterochromatin and RNAi act independently to ensure genome stability in Mucorales human fungal pathogens. Proc. Natl. Acad. Sci. USA120, e2220475120 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Mondo, S. J. et al. Widespread adenine N6-methylation of active genes in fungi. Nat. Genet.49, 964–968 (2017). [DOI] [PubMed] [Google Scholar]
- 35.Chaturvedi, A. et al. The methylome of the model arbuscular mycorrhizal fungus, Rhizophagus irregularis, shares characteristics with early diverging fungi and Dikarya. Commun. Biol.4, 901 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Dallaire, A. et al. Transcriptional activity and epigenetic regulation of transposable elements in the symbiotic fungus Rhizophagus irregularis. Genome Res.31, 2290–2302 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Lax, C. et al. Symmetric and asymmetric DNA N6-adenine methylation regulates different biological responses in Mucorales. Nat. Commun.15, 6066 (2024). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Beh, L. Y. et al. Identification of a DNA N6-adenine methyltransferase complex and its impact on chromatin organization. Cell177, 1781–1796.e25 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Fu, Y. et al. N 6-methyldeoxyadenosine marks active transcription start sites in Chlamydomonas. Cell161, 879–892 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Pan, B. et al. Potential role of N6-adenine DNA methylation in alternative splicing and endosymbiosis in Paramecium bursaria. iScience26, 106676 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Bochtler, M. & Fernandes, H. DNA adenine methylation in eukaryotes: enzymatic mark or a form of DNA damage? Bioessays 2000243 10.1002/bies.202000243 (2020). [DOI] [PubMed]
- 42.Wang, Y. et al. A distinct class of eukaryotic MT-A70 methyltransferases maintain symmetric DNA N6-adenine methylation at the ApT dinucleotides as an epigenetic mark associated with transcription. Nucleic Acids Res.47, 11771–11789 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Luo, G.-Z. et al. N6-methyldeoxyadenosine directs nucleosome positioning in Tetrahymena DNA. Genome Biol.19, 200 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Sedaghatalab, M., Nouripour-Sisakht, S., Cheraghzadeh, S., Sabz, G. & Gharaghani, M. Rhizopus microsporus as causative agent of mucormycosis in COVID-19 patient. Clin. Lab. 68, 220232 (2022). [DOI] [PubMed]
- 45.Jeong, W. et al. The epidemiology and clinical manifestations of mucormycosis: a systematic review and meta-analysis of case reports. Clin. Microbiol. Infect.25, 26–34 (2019). [DOI] [PubMed] [Google Scholar]
- 46.Hassan, M. I. A. & Voigt, K. Pathogenicity patterns of mucormycosis: epidemiology, interaction with immune cells and virulence factors. Méd. Mycol.57, S245–S256 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Lax, C., Nicolás, F. E., Navarro, E. & Garre, V. Molecular mechanisms that govern infection and antifungal resistance in Mucorales. Microbiol. Mol. Biol. Rev. e0018822 10.1128/mmbr.00188-22 (2024). [DOI] [PMC free article] [PubMed]
- 48.Lastovetsky, O. A. et al. Lipid metabolic changes in an early divergent fungus govern the establishment of a mutualistic symbiosis with endobacteria. Proc. Natl. Acad. Sci. USA113, 15102–15107 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Partida‐Martinez, L. P., Bandemer, S., Rüchel, R., Dannaoui, E. & Hertweck, C. Lack of evidence of endosymbiotic toxin‐producing bacteria in clinical Rhizopus isolates. Mycoses51, 266–269 (2008). [DOI] [PubMed] [Google Scholar]
- 50.Partida-Martinez, L. P. & Hertweck, C. Pathogenic fungus harbours endosymbiotic bacteria for toxin production. Nature437, 884–888 (2005). [DOI] [PubMed] [Google Scholar]
- 51.Lackner, G. & Hertweck, C. Impact of endofungal bacteria on infection biology, food safety, and drug development. PLoS Pathog.7, e1002096 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Lackner, G., Moebius, N. & Hertweck, C. Endofungal bacterium controls its host by an hrp type III secretion system. ISME J.5, 252–261 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Partida-Martinez, L. P. et al. Rhizonin, the first mycotoxin isolated from the zygomycota, is not a fungal metabolite but is produced by bacterial endosymbionts. Appl. Environ. Microbiol.73, 793–797 (2007). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Mondo, S. J. et al. Bacterial endosymbionts influence host sexuality and reveal reproductive genes of early divergent fungi. Nat. Commun.8, 1843 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Moebius, N., Üzüm, Z., Dijksterhuis, J., Lackner, G. & Hertweck, C. Active invasion of bacteria into living fungal cells. eLife3, e03007 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Carter, M. E. et al. A TAL effector-like protein of an endofungal bacterium increases the stress tolerance and alters the transcriptome of the host. Proc. Natl. Acad. Sci. USA117, 17122–17129 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Lax, C. et al. Stable and reproducible homologous recombination enables CRISPR-based engineering in the fungus Rhizopus microsporus. Cell Rep. Methods 100124. 10.1016/j.crmeth.2021.100124 (2021). [DOI] [PMC free article] [PubMed]
- 58.Lax, C. et al. Transformation and CRISPR-Cas9-mediated homologous recombination in the fungus Rhizopus microsporus. Star. Protoc.3, 101237 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Wu, K.-J. The epigenetic roles of DNA N6-Methyladenine (6mA) modification in eukaryotes. Cancer Lett.494, 40–46 (2020). [DOI] [PubMed] [Google Scholar]
- 60.Wang, Y., Chen, X., Sheng, Y., Liu, Y. & Gao, S. N6-adenine DNA methylation is associated with the linker DNA of H2A.Z-containing well-positioned nucleosomes in Pol II-transcribed genes in Tetrahymena. Nucleic Acids Res.45, gkx883 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Greer, E. L. et al. DNA methylation on N6-adenine in C. elegans. Cell161, 868–878 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Bewick, A. J. et al. Diversity of cytosine methylation across the fungal tree of life. Nat. Ecol. Evol.3, 479–490 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Kong, Y. et al. Critical assessment of DNA adenine methylation in eukaryotes using quantitative deconvolution. Science375, 515–522 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Bernstein, B. E. et al. Genomic maps and comparative analysis of histone modifications in human and mouse. Cell120, 169–181 (2005). [DOI] [PubMed] [Google Scholar]
- 65.Zhang, X., Bernatavichute, Y. V., Cokus, S., Pellegrini, M. & Jacobsen, S. E. Genome-wide analysis of mono-, di- and trimethylation of histone H3 lysine 4 in Arabidopsis thaliana. Genome Biol.10, R62 (2009). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Liu, C. L. et al. Single-nucleosome mapping of histone modifications in S. cerevisiae. Plos Biol.3, e328 (2005). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Barski, A. et al. High-resolution profiling of histone methylations in the human genome. Cell129, 823–837 (2007). [DOI] [PubMed] [Google Scholar]
- 68.Bargaje, R. et al. Proximity of H2A. Z containing nucleosome to the transcription start site influences gene expression levels in the mammalian liver and brain. Nucleic Acids Res.40, 8965–8978 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Gómez-Zambrano, Á., Merini, W. & Calonje, M. The repressive role of Arabidopsis H2A.Z in transcriptional regulation depends on AtBMI1 activity. Nat. Commun.10, 2828 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Guillemette, B. et al. Variant histone H2A.Z is globally localized to the promoters of inactive yeast genes and regulates nucleosome positioning. PloS Biol.3, e384 (2005). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Courtney, A. J. et al. Normal patterns of histone H3K27 methylation require the histone variant H2A.Z in Neurospora crassa. Genetics216, 51–66 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Yao, F. et al. Histone variant H2A.Z is required for the maintenance of smooth muscle cell identity as revealed by single-cell transcriptomics. Circulation138, 2274–2288 (2018). [DOI] [PubMed]
- 73.Ku, M. et al. H2A.Z landscapes and dual modifications in pluripotent and multipotent stem cells underlie complex genome regulatory functions. Genome Biol.13, R85 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Bagchi, D. N., Battenhouse, A. M., Park, D. & Iyer, V. R. The histone variant H2A.Z in yeast is almost exclusively incorporated into the +1 nucleosome in the direction of transcription. Nucleic Acids Res.48, 157–170 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Wang, X. et al. DNA methylation on N6-adenine in lepidopteran Bombyx mori. Biochim. Biophys Acta Gene Regul. Mech.1861, 815–825 (2018). [DOI] [PubMed] [Google Scholar]
- 76.Liu, J. et al. A METTL3-METTL14 complex mediates mammalian nuclear RNA N6-adenosine methylation. Nat. Chem. Biol.10, 93–95 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Iyer, L. M., Zhang, D. & Aravind, L. Adenine methylation in eukaryotes: apprehending the complex evolutionary history and functional potential of an epigenetic modification. BioEssays38, 27–40 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Chen, J. et al. Structural basis for MTA1c-mediated DNA N6-adenine methylation. Nat. Commun.13, 3257 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Navarro-Mendoza, M. I. et al. Early diverging fungus Mucor circinelloides lacks centromeric histone CENP-A and displays a mosaic of point and regional centromeres. Curr. Biol.29, 3791–3802.e6 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Bartlett, A. et al. Mapping genome-wide transcription-factor binding sites using DAP-seq. Nat. Protoc.12, 1659–1672 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.O’Malley, R. C. et al. Cistrome and Epicistrome Features Shape the Regulatory DNA Landscape. Cell166, 1598 (2016). [DOI] [PubMed] [Google Scholar]
- 82.Wang, B. et al. Structural insights into target DNA recognition by R2R3-MYB transcription factors. Nucleic Acids Res.48, 460–471 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Nicolás, F. E. et al. Molecular tools for carotenogenesis analysis in the mucoral Mucor circinelloides. Methods Mol. Biol.1852, 221–237 (2018). [DOI] [PubMed] [Google Scholar]
- 84.Allis, C. D. & Jenuwein, T. The molecular hallmarks of epigenetic control. Nat. Rev. Genet.17, 487–500 (2016). [DOI] [PubMed] [Google Scholar]
- 85.Chang, Z., Yadav, V., Lee, S. C. & Heitman, J. Epigenetic mechanisms of drug resistance in fungi. Fungal Genet. Biol.132, 103253 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Allshire, R. C. & Ekwall, K. Epigenetic regulation of chromatin states in Schizosaccharomyces pombe. Cold Spring Harb. Perspect. Biol.7, a018770 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Kronholm, I., Johannesson, H. & Ketola, T. Epigenetic control of phenotypic plasticity in the filamentous fungus Neurospora crassa. G36, 4009–4022 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Dubey, A. & Jeon, J. Epigenetic regulation of development and pathogenesis in fungal plant pathogens. Mol. Plant Pathol.18, 887–898 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Huang, Y. Transcriptional silencing in Saccharomyces cerevisiae and Schizosaccharomyces pombe. Nucleic Acids Res.30, 1465–1482 (2002). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Luo, G.-Z., Blanco, M. A., Greer, E. L., He, C. & Shi, Y. DNA N6-methyladenine: a new epigenetic mark in eukaryotes? Nat. Rev. Mol. Cell Biol.16, 705–710 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Li, H. et al. DNA N6-methyladenine modification in eukaryotic genome. Front. Genet.13, 914404 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Urig, S. et al. The Escherichia coli dam DNA methyltransferase modifies DNA in a highly processive reaction. J. Mol. Biol.319, 1085–1096 (2002). [DOI] [PubMed] [Google Scholar]
- 93.Liu, J. et al. Abundant DNA 6mA methylation during early embryogenesis of zebrafish and pig. Nat. Commun.7, 13052 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Fernandes, S. B. et al. N6-methyladenine in eukaryotic DNA: tissue distribution, early embryo development, and neuronal toxicity. Front. Genet.12, 657171 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Zhang, G. et al. N6-methyladenine DNA modification in Drosophila. Cell161, 893–906 (2015). [DOI] [PubMed] [Google Scholar]
- 96.Xie, Q. et al. N-methyladenine DNA modification in glioblastoma. Cell175, 1228–1243.e20 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97.Li, X. et al. The exploration of N6-deoxyadenosine methylation in mammalian genomes. Protein Cell 1–13 10.1007/s13238-021-00866-3 (2021). [DOI] [PMC free article] [PubMed]
- 98.Min, B., Park, J. S., Jeong, Y. S., Jeon, K. & Kang, Y.-K. Dnmt1 binds and represses genomic retroelements via DNA methylation in mouse early embryos. Nucleic Acids Res.48, gkaa584 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99.Liang, Z. et al. DNA N6-adenine methylation in Arabidopsis thaliana. Dev. Cell45, 406–416.e3 (2017). [DOI] [PubMed] [Google Scholar]
- 100.Yao, B. et al. Active N6-methyladenine demethylation by DMAD regulates gene expression by coordinating with polycomb protein in neurons. Mol. Cell71, 848–857.e6 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.Koziol, M. J. et al. Identification of methylated deoxyadenosines in vertebrates reveals diversity in DNA modifications. Nat. Struct. Mol. Biol.23, 24–30 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102.Wu, T. P. et al. DNA methylation on N6-adenine in mammalian embryonic stem cells. Nature532, 329–333 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Lee, W. et al. A high-resolution atlas of nucleosome occupancy in yeast. Nat. Genet.39, 1235–1244 (2007). [DOI] [PubMed] [Google Scholar]
- 104.Torres-Garcia, S. et al. Epigenetic gene silencing by heterochromatin primes fungal resistance. Nature585, 453–458 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105.Richards, E. J. & Elgin, S. C. R. Epigenetic codes for heterochromatin formation and silencing. Cell108, 489–500 (2002). [DOI] [PubMed] [Google Scholar]
- 106.Saksouk, N., Simboeck, E. & Déjardin, J. Constitutive heterochromatin formation and transcription in mammals. Epigenet. Chromatin8, 3 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107.Kurup, J. T., Campeanu, I. J. & Kidder, B. L. Contribution of H3K4 demethylase KDM5B to nucleosome organization in embryonic stem cells revealed by micrococcal nuclease sequencing. Epigenet. Chromatin12, 20 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 108.Zhu, Q., Ramakrishnan, M., Park, J. & Belden, W. J. Histone H3 lysine 4 methyltransferase is required for facultative heterochromatin at specific loci. BMC Genom.20, 350 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.Ferraro, A. R. et al. Chromatin accessibility profiling in Neurospora crassa reveals molecular features associated with accessible and inaccessible chromatin. BMC Genom.22, 459 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110.Liu, Y. et al. RNAi-dependent H3K27 methylation is required for heterochromatin formation and DNA elimination in Tetrahymena. Genes Dev.21, 1530–1545 (2007). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111.Roden, M. M. et al. Epidemiology and outcome of zygomycosis: a review of 929 reported cases. Clin. Infect. Dis.41, 634–653 (2005). [DOI] [PubMed] [Google Scholar]
- 112.Lamoth, F. Novel approaches in the management of mucormycosis. Curr. Fungal Infect. Rep.17, 1–10 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113.Dannaoui, E. Antifungal resistance in mucorales. Int. J. Antimicrob. Agents50, 617–621 (2017). [DOI] [PubMed] [Google Scholar]
- 114.Abdel-Hafez, Y. et al. Tolerability and epidemiology of nephrotoxicity associated with conventional amphotericin B therapy: a retrospective study in tertiary care centers in Palestine. BMC Nephrol.23, 132 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 115.Kagan, S. et al. Toxicity mechanisms of amphotericin B and its neutralization by conjugation with arabinogalactan. Antimicrob. Agents Chemother.56, 5603–5611 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 116.Jiang, X. et al. The role of m6A modification in the biological functions and diseases. Signal Transduct. Target. Ther.6, 74 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 117.Eid, J. et al. Real-time DNA sequencing from single polymerase molecules. Science323, 133–138 (2009). [DOI] [PubMed] [Google Scholar]
- 118.Vaser, R., Sović, I., Nagarajan, N. & Šikić, M. Fast and accurate de novo genome assembly from long uncorrected reads. Genome Res.27, 737–746 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 119.Grigoriev, I. V. et al. MycoCosm portal: gearing up for 1000 fungal genomes. Nucleic Acids Res.42, D699–D704 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 120.Manni, M., Berkeley, M. R., Seppey, M., Simão, F. A. & Zdobnov, E. M. BUSCO update: novel and streamlined workflows along with broader and deeper phylogenetic coverage for scoring of eukaryotic, prokaryotic, and viral genomes. Mol. Biol. Evol.38, 4647–4654 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121.Stanke, M. & Morgenstern, B. AUGUSTUS: a web server for gene prediction in eukaryotes that allows user-defined constraints. Nucleic Acids Res.33, W465–W467 (2005). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 122.Chaisson, M. J. & Tesler, G. Mapping single molecule sequencing reads using basic local alignment with successive refinement (BLASR): application and theory. BMC Bioinform.13, 238–238 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 123.Flusberg, B. A. et al. Direct detection of DNA methylation during single-molecule, real-time sequencing. Nat. Methods7, 461–465 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 124.Flynn, J. M. et al. RepeatModeler2 for automated genomic discovery of transposable element families. Proc. Natl. Acad. Sci. USA117, 9451–9457 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 125.Rice, P., Longden, I. & Bleasby, A. EMBOSS: the European molecular biology open software suite. Trends Genet.16, 276–277 (2000). [DOI] [PubMed] [Google Scholar]
- 126.Jones, P. et al. InterProScan 5: genome-scale protein function classification. Bioinformatics30, 1236–1240 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 127.Ramírez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res.44, W160–W165 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 128.Wickham, H. ggplot2, elegant graphics for data analysis. 109–145 10.1007/978-3-319-24277-4_6 (2016).
- 129.Bailey, T. L., Johnson, J., Grant, C. E. & Noble, W. S. The MEME suite. Nucleic Acids Res.43, W39–W49 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 130.Lowe, T. M. & Eddy, S. R. tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res.25, 955–964 (1997). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 131.Lagesen, K. et al. RNAmmer: consistent and rapid annotation of ribosomal RNA genes. Nucleic Acids Res.35, 3100–3108 (2007). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 132.Kalvari, I. et al. Rfam 14: expanded coverage of metagenomic, viral and microRNA families. Nucleic Acids Res.49, D192–D200 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 133.Krueger, F. & Andrews, S. R. Bismark: a flexible aligner and methylation caller for Bisulfite-Seq applications. Bioinformatics27, 1571–1572 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 134.Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol.10, R25–R25 (2009). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 135.Schultz, M. D., Schmitz, R. J. & Ecker, J. R. Leveling’ the playing field for analyses of single-base resolution DNA methylomes. Trends Genet.28, 583–585 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 136.Grabherr, M. G. et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol.29, 644–652 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 137.Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods12, 357–360 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 138.Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics30, 923–930 (2014). [DOI] [PubMed] [Google Scholar]
- 139.Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol.15, 550 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 140.Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J.17, 10–12 (2011). [Google Scholar]
- 141.Chen, K. et al. DANPOS: dynamic analysis of nucleosome position and occupancy by sequencing. Genome Res.23, 341–351 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 142.Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol.29, 24–26 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 143.Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows–Wheeler transform. Bioinformatics26, 589–595 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 144.Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol.9, R137 (2008). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 145.Baumgart, L. A. et al. Persistence and plasticity in bacterial gene regulation. Nat. Methods18, 1499–1505 (2021). [DOI] [PubMed] [Google Scholar]
- 146.Consortium, T. U. et al. UniProt: the universal protein knowledgebase in 2021. Nucleic Acids Res.49, D480–D489 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 147.Fu, L., Niu, B., Zhu, Z., Wu, S. & Li, W. CD-HIT: accelerated for clustering the next-generation sequencing data. Bioinformatics28, 3150–3152 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 148.Camacho, C. et al. BLAST+: architecture and applications. BMC Bioinform.10, 421 (2009). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 149.Mistry, J. et al. Pfam: The protein families database in 2021. Nucleic Acids Res.49, gkaa913- (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 150.Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res.32, 1792–1797 (2004). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 151.Jones, D. T., Taylor, W. R. & Thornton, J. M. The rapid generation of mutation data matrices from protein sequences. Bioinformatics8, 275–282 (1992). [DOI] [PubMed] [Google Scholar]
- 152.Kumar, S., Stecher, G., Li, M., Knyaz, C. & Tamura, K. MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol.35, 1547–1549 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 153.Emms, D. M. & Kelly, S. OrthoFinder: phylogenetic orthology inference for comparative genomics. Genome Biol.20, 238 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 154.Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol.30, 772–780 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 155.Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics30, 1312–1313 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 156.Waterhouse, A. M., Procter, J. B., Martin, D. M. A., Clamp, M. & Barton, G. J. Jalview Version 2—a multiple sequence alignment editor and analysis workbench. Bioinformatics25, 1189–1191 (2009). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 157.Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol.215, 403–410 (1990). [DOI] [PubMed] [Google Scholar]
- 158.Gouy, M., Guindon, S. & Gascuel, O. SeaView version 4: a multiplatform graphical user interface for sequence alignment and phylogenetic tree building. Mol. Biol. Evol.27, 221–224 (2010). [DOI] [PubMed] [Google Scholar]
- 159.Capella-Gutiérrez, S., Silla-Martínez, J. M. & Gabaldón, T. trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics25, 1972–1973 (2009). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 160.Minh, B. Q. et al. IQ-TREE 2: New models and efficient methods for phylogenetic inference in the genomic era. Mol. Biol. Evol.37, 1530–1534 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 161.Letunic, I. & Bork, P. Interactive Tree Of Life (iTOL) v5: an online tool for phylogenetic tree display and annotation. Nucleic Acids Res.49, W293–W296 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 162.Mirdita, M. et al. ColabFold: making protein folding accessible to all. Nat. Methods19, 679–682 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 163.Crooks, G. E., Hon, G., Chandonia, J.-M. & Brenner, S. E. WebLogo: a sequence logo generator. Genome Res.14, 1188–1190 (2004). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 164.Nicolás, F. E., Haro, J. P., de, Torres-Martínez, S. & Ruiz-Vázquez, R. M. Mutants defective in a Mucor circinelloides dicer-like gene are not compromised in siRNA silencing but display developmental defects. Fungal Genet. Biol.44, 504–516 (2007). [DOI] [PubMed] [Google Scholar]
- 165.Lasker, B. A. & Borgia, P. T. High-frequency heterokaryon formation by Mucor racemosus. J. Bacteriol.141, 565–569 (1980). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 166.Gietz, R. D., Schiestl, R. H., Willems, A. R. & Woods, R. A. Studies on the transformation of intact yeast cells by the LiAc/SS‐DNA/PEG procedure. Yeast11, 355–360 (1995). [DOI] [PubMed] [Google Scholar]
- 167.Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics26, 841–842 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Description of Additional Supplementary Files
Data Availability Statement
The raw sequence data that support the findings of this study have been deposited in the Sequence Read Archive (SRA) under the accession numbers SRP496792, SRP496793, SRP496795, SRP496796, SRP496797, SRP496798, SRP496799, SRP496800, SRP496816, SRP496817, SRP496818, SRP496819, SRP496820, SRP496821, SRP496822, SRP496823, SRP496824, SRP496825, SRP496830, SRP496831, SRP496832, SRP496833, SRP497170, SRP497171, SRP497172, and SRP536317, and in the Gene Expression Omnibus (GEO) under the accession numbers GSE285378, GSE276938, and GSE276939. The Whole Genome Shotgun project for R. Microsporus ATCC 11559 has been deposited at DDBJ/ENA/GenBank under the accession JBFXLZ000000000. The version described in this paper is version JBFXLZ010000000 [https://www.ncbi.nlm.nih.gov/nuccore/JBFXLZ000000000.1/]. The HPLC/MS raw data has been deposited in Zenodo under the accession number 15194697 (10.5281/zenodo.15194696; https://zenodo.org/records/15194697). Source data are provided with this paper.