Summary
Herpes simplex virus 1 (HSV-1) is a ubiquitous human pathogen that causes serious symptoms and is known for its strong interactions with host immunity. Here, we revealed that the HSV-1-encoded UL38 is a stimulator of interferon genes (STING) antagonist that interacts with STING to abrogate the STING-TANK-binding kinase 1 (TBK1)-interferon regulatory factor 3 (IRF3) interaction, thereby suppressing cyclic GMP-AMP synthase (cGAS)-STING-dependent immune signaling. Losing UL38’s STING antagonist activity made HSV-1 incapable of immune evasion and less replicable and pathogenic in vivo. Moreover, on the basis of the UL38-interacting sequence within STING, we rationally designed a series of peptides to target the STING-UL38 interface of UL38 specifically. Among them, a peptide effectively disrupts the STING-UL38 interaction, which unlocks the UL38-suppressed immune response and shows potent therapeutic efficacy against HSV-1 infection in vivo. Therefore, our findings demonstrate that HSV-1 UL38 is a STING antagonist, and targeting the activity of UL38 is a promising strategy for the development of antivirals against this notorious virus.
Keywords: herpes simplex virus 1, UL38, STING, drug target, antiviral innate immunity, antivirals
Graphical abstract

Highlights
-
•
Herpes simplex virus 1-encoded UL38 is a STING antagonist
-
•
Loss of this UL38 activity made HSV-1 less replicable and pathogenic in vivo
-
•
A STING-derived peptide is designed to target the STING-UL38 interface
-
•
This peptide shows potent therapeutic efficacy against HSV-1 infection in vivo
Wang et al. reveal that herpes simplex virus 1-encoded UL38 is a STING antagonist to evade antiviral immunity by directly binding to STING. They further develop a peptide derived from STING sequence to specifically target the STING-UL38 interface, which shows potent in vivo therapeutic efficacy against this virus.
Introduction
Herpes simplex virus 1 (HSV-1) is an enveloped large double-stranded DNA (dsDNA) virus belonging to the genus Simplexvirus of the Alphaherpesvirinae subfamily within the family Herpesviridae.1 After primary infection, HSV-1 genomes remain latent within infected cells, particularly in neurons in the sensory and autonomic ganglia, which can occasionally be reactivated.2 HSV-1 is highly widespread across the global population, as the World Health Organization estimates that approximately 3.8 billion people under age 50 are infected with HSV-1. Infection caused by HSV-1 is usually asymptomatic or only causes moderate symptoms, such as oral herpes or cold sores, while its recurrence from latency is often associated with painful symptoms, such as lesions in the facial or genital areas.3,4 In addition, HSV-1 infection can be associated with a number of more severe complications, such as herpetic stromal keratitis, a leading cause of infectious corneal blindness, and herpes simplex encephalitis, which can result in severe damage to neurological functions with a high risk of mortality.5,6 Furthermore, individuals with immunocompromised conditions or who are receiving immunosuppressive treatment are more prone to HSV-1 infection, which has increased severity and more frequent recurrence.7
Thus far, all approved antivirals against HSV-1 (e.g., acyclovir and related nucleoside analogs) have the same therapeutic target—viral DNA polymerase—and can only inhibit but not eradicate HSV-1 replication. Moreover, resistance or even cross-resistance to acyclovir and other anti-HSV-1 nucleoside analogs has frequently emerged.8 Therefore, identifying novel therapeutic targets and developing new antiviral strategies against HSV-1 is highly desirable.
The host antiviral immune system intricately modulates primary, latent, and recurrent HSV-1 infections. For DNA viruses, including herpesviruses, cyclic GMP-AMP synthase (cGAS) senses viral dsDNA and produces cyclic GMP-AMP (cGAMP), which then activates stimulator of interferon genes (STING), which bind to TANK-binding kinase 1 (TBK1) to phosphorylate and activate interferon regulatory factor 3 (IRF3), leading to the induction of type I interferons (IFNs) and subsequent IFN-stimulated genes (ISGs).9,10,11,12 The cGAS-STING-IFN pathway plays a central role in controlling the infection, latency, and recurrence of HSV-1 in host cells, while HSV-1 has been reported to use several virus-encoded proteins, such as ICP0, ICP27, UL37, UL41, UL46, and UL56, to evade the cGAS-STING-IFN immune pathway.13,14,15,16,17,18,19,20,21,22,23 Thus, HSV-1 is considered a good model for studying the interactions between viruses and host immunity, and the elucidation of such sophisticated mechanisms could provide valuable hints for identifying new antiviral targets and strategies. However, despite the multiple antagonizing mechanisms of the cGAS-STING pathway employed by HSV-1, few reports have described the development of novel antivirals that target these HSV-1 immune evasion strategies.
HSV-1-encoded UL38 (also named VP19C) is a component of the HSV-1 capsid, which has been reported to have DNA-binding and nucleoplasmic shuttle activities and is critical for viral assembly and replication.24,25,26,27 In this study, we revealed that HSV-1 UL38 could inhibit cGAS-STING-dependent immune signaling. Mechanistically, UL38 interacts with STING to abrogate cGAMP binding to STING and the STING-TBK1-IRF3 interaction, leading to the blockade of subsequent immune signaling. Losing the STING antagonist function of UL38 made HSV-1 incapable of inhibiting innate immunity and less replicable and pathogenic in vivo. Moreover, on the basis of the amino acid sequence within STING responsible for the UL38 interaction, we rationally designed a series of peptides to specifically target the STING-UL38 interface within UL38, which should inhibit UL38 function but not affect STING. We found that a designer peptide effectively disrupted the STING-UL38 interaction, which led to the unlocking of the UL38-suppressed innate immune response and showed potent therapeutic efficacy against both cerebral and systematic infections of HSV-1 in vivo, which is comparable with that of acyclovir. Therefore, our findings demonstrate that HSV-1 UL38 is a STING antagonist that evades host antiviral innate immunity and that targeting UL38 is a promising strategy for the development of antivirals against HSV-1 and associated diseases.
Results
HSV-1 UL38 inhibits DNA- or virus-triggered innate immune signaling
Previous studies have revealed that a number of viral structural proteins, such as coronaviral M and N proteins, Ebola virus VP35 protein, and rabies virus P protein, can inhibit innate immunity.28,29,30,31 To examine whether HSV-1 UL38 can affect innate immune signaling, we measured the promoter activity of IFN-sensitive response element (ISRE) and nuclear factor κB (NF-κB), as well as the transcription of IFNB1, ISG15, ISG56, and RANTES in cultured 293 cells transfected with cGAS and STING expression plasmids together with increasing amounts of the UL38 expression vector. The data revealed that the ectopic expression of UL38 inhibited the cGAS-STING-triggered activation of the ISRE and NF-κB promoters and the transcription of these immune genes in a dose-dependent manner in cells (Figures 1A and S1A). Moreover, UL38 inhibited the transcription of IFNB1, ISG15, and RANTES induced by the transfection of the DNA ligand poly(dA:dT) in HeLa cells (Figure 1B) and primary murine lung fibroblasts (MLFs) (Figure S1B). We then examined the phosphorylation of TBK1 and IRF3, the key downstream signaling events of the cGAS-STING pathway, in both HeLa cells and MLFs treated with poly(dA:dT) in the presence or absence of UL38 expression. Our data showed that UL38 effectively inhibited the DNA-triggered phosphorylation of TBK1 or IRF3 (Figures 1C and S1C).
Figure 1.
HSV-1 UL38 inhibits DNA- or virus-triggered innate immune signaling
(A) A luciferase reporter assay analyzing ISRE or NF-κB promoter activity was conducted in 293 cells transfected with a plasmid encoding FLAG-cGAS plus FLAG-STING or an empty vector together with increasing amounts of FLAG-UL38 expression plasmid for 24 h. The lower blots show the expression levels of these transfected proteins.
(B) HeLa cells were transfected with a FLAG-UL38 plasmid or an empty vector. After 24 h, the cells were left untreated (control) or were transfected with poly(dA:dT) (1 μg/mL). At 6 hr post transfection (h.p.t.), total RNA was extracted and subjected to quantitative reverse-transcription PCR (qRT-PCR) of the indicated antiviral genes.
(C) HeLa cells were transfected with an FLAG-UL38 plasmid or an empty vector. After 24 h, the cells were transfected with poly(dA:dT). The cell lysates were prepared and subjected to immunoblotting with the indicated antibodies.
(D–F) HeLa cells were transfected with the FLAG-UL38 plasmid or an empty vector, followed by infection with HSV-1. At the indicated h.p.i., the transcription of the indicated antiviral genes was measured via qRT-PCR (D), ELISA was performed to measure the secretion of IFN-β (E), and the cell lysates were prepared and subjected to immunoblotting with the indicated antibodies (F).
(G and H) HeLa cells were transfected with the FLAG-UL38 plasmid or empty vector and then infected with HSV-1 (MOI = 0.1). At 12 or 24 h.p.i., the mRNA levels of the HSV-1 UL30 gene were measured via qRT-PCR (G), and the virus titer was detected by the plaque assay (H). All experiments were independently conducted in triplicate and repeated at least twice with reproducible results. The graph shows the means ± SD, n = 3.
p values were calculated via one-way or two-way ANOVA. See also Figure S1.
After that, we examined whether UL38 can inhibit virus-induced immune signaling. The data revealed that the ectopic expression of UL38 inhibited the transcription of IFNB1, ISG15, and RANTES, the secretion of IFN-β, and the phosphorylation of TBK1 or IRF3 in HSV-1-infected HeLa cells (Figures 1D–1F), consistent with the observations in monocytic THP-1 cells and MLFs (Figures S1D–S1F). Moreover, we aimed to examine the effect of virally expressed UL38. Our data revealed that the HSV-1-encoded UL38 gene was transcribed at 12 hr post infection (h.p.i.) (Figure S1I). Moreover, the knockdown of virally expressed UL38 in HSV-1-infected HeLa cells or MLFs significantly increased immune gene transcription (Figures S1J and S1K) and the phosphorylation of TBK1 or IRF3 (Figures S1L–S1M). Moreover, the ectopic expression of UL38 efficiently enhanced the replication of HSV-1 in infected HeLa cells (Figures 1G and 1H) and MLFs (Figures S1G and S1H), whereas the knockdown of UL38 had the opposite effect, as expected (Figures S1N–S1Q).
Our data show that HSV-1 UL38 can inhibit cGAS-STING-dependent innate immune signaling triggered by DNA ligands or viral infection in human or murine cells.
UL38 binds to STING to inhibit cGAS-STING-mediated immune signaling
We then explored how HSV-1 UL38 inhibits the cGAS-STING axis. The data revealed that UL38 suppressed the activation of the ISRE promoter triggered by the overexpression of cGAS plus STING but not TBK1 or IRF3 (Figure 2A), indicating that UL38 functions upstream of TBK1. We then examined whether UL38 can interact with any key component of the cGAS-STING axis and found that UL38 co-immunoprecipitated with STING but not with cGAS, TBK1, or IRF3 (Figure 2B). To characterize the interaction between STING and UL38 further, we generated a series of STING truncations, and the co-immunoprecipitation (co-IP) data revealed that the region of amino acids (aa) 161–190 of STING is responsible for its interaction with UL38 (Figures S2A and 2C).
Figure 2.
UL38 binds to STING to inhibit cGAS-STING-mediated immune signaling
(A) A luciferase reporter assay analyzing ISRE or NF-κB promoter activity was conducted in 293 cells transfected with the ISRE reporter and plasmids encoding FLAG-tagged cGAS plus STING, TBK1, IRF3, or an empty vector for 24 h. The lower blots show the expression levels of these transfected proteins.
(B) 293 cells were transfected with plasmids encoding FLAG-tagged cGAS, STING, TBK1, or IRF3 together with HA-UL38. After 24 h, the cells were prepared and subjected to immunoprecipitation (IP) with an anti-HA monoclonal antibody (ɑH) or IgG, followed by immunoblotting with anti-FLAG and anti-HA antibodies.
(C) 293 cells were transfected with HA-tagged STING or STINGΔ161–190 together with the FLAG-UL38 expression plasmid. After 24 h, the cells were subjected to IP with an anti-FLAG monoclonal antibody (ɑF) or IgG, followed by immunoblotting with anti-FLAG and anti-HA antibodies.
(D) 293 cells were transfected with plasmids containing FLAG-UL38 or the indicated truncations together with HA-STING. After 24 h, the cells were subjected to IP with an anti-FLAG monoclonal antibody or IgG, followed by immunoblotting with anti-FLAG and anti-HA antibodies.
(E) HeLa cells were transfected with plasmids encoding FLAG-UL38, FLAG-UL38Δ81–90 or an empty vector. After 24 h, the cells were infected with HSV-1 (MOI = 0.1); at 24 h.p.i., qRT-PCR was performed to measure the transcription of the HSV-1 UL30 gene.
(F and G) HeLa STING−/− cells were transfected with the ISRE reporter plasmid together with the indicated plasmids. After 24 h, the cell lysates were subjected to a luciferase assay.
(H and I) HeLa cells were infected with HSV-1 (MOI = 1) or HSV-1UL38Δ81–90 (MOI = 1). At 0, 6, and 12 h.p.i., the cell lysates were subjected to immunoblotting with the indicated antibodies (H), and qRT-PCR was performed to measure the transcription of antiviral genes (I).
(J–M) HeLa (J and K) or HeLa STING−/− (L and M) cells were infected with HSV-1 or HSV-1UL38Δ81–90 (MOI = 0.1) as indicated. At 24 h.p.i., qRT-PCR was performed to measure the expression of HSV-1 UL30 (J and L), and the virus titer was detected by the plaque assay (K and M). All experiments were independently conducted in triplicate and repeated at least twice with reproducible results. The graph shows the means ± SD, n = 3.
p values were calculated via one-way or two-way ANOVA. See also Figure S2.
To determine the region of UL38 responsible for the interaction with STING, we constructed a series of UL38 deletion mutants. Our data revealed that UL38Δ61–120 (without aa 61–120 of UL38) completely lost the ability to interact with STING (Figure S2B) or to suppress the cGAS-STING-triggered activation of the ISRE promoter (Figure S2C). To further narrow the critical region within UL38, we generated a series of deletions with aa 61–120 and found that UL38Δ81–90 (with aa 81–90 deleted from UL38) failed to bind to STING (Figure 2D) or inhibit DNA-triggered IFNB1 transcription (Figure S2D). Consistently, this UL38 mutant lost its ability to increase HSV-1 replication in infected cells (Figure 2E). Furthermore, we ectopically expressed STING or STINGΔ161–190 in STING−/− HeLa cells in the presence or absence of UL38 or UL38Δ81–90. The data revealed that UL38 but not UL38Δ81–90 inhibited STING-mediated ISRE promoter activation (Figure 2F); however, although STINGΔ161–190 expression can support ISRE activation, UL38 failed to inhibit this activation because of its inability to bind to STINGΔ161–190 (Figure 2G).
To further assess the impact of the STING-targeting activity of UL38 on HSV-1, we generated an HSV-1 mutant virus with the aa 81–90 deletion of its UL38 (i.e., HSV-1UL38Δ81–90) (Figures S2E and S2F). Moreover, wild-type HSV-1 and HSV-1UL38Δ81–90 showed comparable replication efficiency in IFNAR−/− 293 cells (Figures S2G and S2H), indicating that deletion of these residues within UL38 did not affect the infectivity or replication of HSV-1. Compared with those in wild-type HSV-1-infected cells, the phosphorylation of TBK1 or IRF3 or the transcription of IFNB1, ISG15, and RANTES was significantly greater in HSV-1UL38Δ81–90-infected HeLa cells (Figures 2H and 2I) and MLFs (Figures S2I and S2J). In addition, the loss of STING-targeting activity (i.e., HSV-1UL38Δ81–90) resulted in reduced replication efficiency in normal cells (Figures 2J, 2K, S2K, and S2L) but not in STING−/− cells (Figures 2L and 2M), further confirming that the function of UL38 depends on the presence of STING.
Overall, the results demonstrate that HSV-1 UL38 is a STING antagonist that inhibits cGAS-STING-mediated immune signaling.
UL38 abrogates the interactions of STING with TBK1 and IRF3
We then aimed to reveal the mechanism by which UL38 antagonizes STING. After viral infection, viral DNA triggers cGAS activation and cGAMP production. Then, cGAMP binds to STING, resulting in the endoplasmic reticulum (ER)-Golgi transport of STING and the formation of the STING-TBK1-IRF3 complex, leading to subsequent immune induction. Therefore, we speculate that UL38 may interfere with the binding of cGAMP to STING. To test this possibility, we conducted in vitro binding assay of cGAMP-STING with or without UL38, and the data showed that UL38 efficiently inhibited cGAMP binding to STING (Figure 3A). Considering that UL38 binds to STING at aa 161–190, which are in close proximity to the cGAMP-binding site at STING,32 UL38 inhibits cGAMP-STING binding probably via competition. Moreover, the overexpression of UL38 markedly inhibited the phosphorylation of STING in the presence of cGAMP (Figure 3B), the interaction of STING with inactive rhomboid protein 2 (iRhom2) and translocon-associated protein β (TRAPβ), the two proteins facilitating the ER-Golgi transport of STING9,33 (Figures 3C and 3D), as well as the STING-TBK1 or STING-IRF3 interaction (Figures 3E and 3F). Consistently, the presence of UL38 also substantially impaired the interactions of endogenous TBK1 and IRF3 with STING in HeLa cells (Figure 3G). Moreover, the transfection of DNA ligands promoted the endogenous STING-TBK1-IRF3 interaction, which was dramatically inhibited by the expression of UL38 (Figure 3H).
Figure 3.
UL38 abrogates the interactions of STING with TBK1 and IRF3
(A) In vitro competition assay involving biotin-2′,3′-cGAMP, FLAG-STING, and His-UL38. Biotin-2′3'-cGAMP was incubated with affinity-purified FLAG-STING and His-UL38. After incubation, the reaction mixture was subjected to pull-down using Strep-beads (Str), and empty beads were used in controls (Ctrl). The mixture was then subjected to immunoblotting analysis with indicated antibodies.
(B) HeLa cells were transfected with FLAG-UL38 for 24 h, followed by stimulating with 2′3′-cGAMP for 30 min. Cells were harvested and analyzed by immunoblotting to detect phosphorylated STING (P-STING).
(C–F) 293 cells were transfected with the indicated plasmids. After 24 h, the cell lysates were prepared and subjected to IP with an indicated monoclonal antibody or IgG, followed by immunoblotting with indicated antibodies.
(G) HeLa cells were transfected with the FLAG-UL38 plasmid; after 24 h, the cell lysates were subjected to IP with an anti-STING monoclonal antibody (ɑS) or IgG, followed by immunoblotting with the indicated antibodies.
(H) HeLa cells were transfected with the HA-UL38 plasmid or an empty vector; at 24 h.p.t., the cells were untreated (control) or transfected with poly(dA:dT) (1 μg/mL). After 6 h, the cell lysates were prepared and subjected to IP with an anti-STING monoclonal antibody or IgG, followed by immunoblotting with the indicated antibodies.
(I–K) 293 cells were transfected with the indicated plasmids. After 24 h, the cell lysates were subjected to IP with an anti-HA monoclonal antibody or IgG, followed by immunoblotting with anti-FLAG and anti-HA antibodies.
(L) HeLa cells were infected with HSV-1 (MOI = 1) or HSV-1UL38Δ81–90 (MOI = 1). At 24 h.p.i., the cell lysates were subjected to IP with an anti-STING monoclonal antibody or IgG, followed by immunoblotting with the indicated antibodies. All experiments were independently conducted in triplicate and repeated at least twice with reproducible results.
The loss of the UL38-binding region of STING (i.e., STINGΔ161–190) did not affect its interaction with TBK1 or IRF3, while UL38 failed to inhibit these interactions (Figures 3I and 3J). In addition, unlike wild-type UL38, UL38Δ81–90 lost the ability to inhibit the interaction between STING and TBK1 (Figure 3K), and the formation of the STING-TBK1-IRF3 complex was obviously stronger in HSV-1UL38Δ81–90-infected cells than in HSV-1-infected cells (Figure 3L). Our results demonstrate that UL38 functions as a STING antagonist by inhibiting cGAMP binding to STING, thereby interfering with the interactions of STING with its partner proteins TBK1 and IRF3.
UL38 contributes to the immune evasion, replication, and pathogenesis of HSV-1
After determining the STING antagonist function of UL38, we evaluated the in vivo impact of UL38 on HSV-1. We first assessed the effect of UL38 on the systemic infection of HSV-1 in vivo. To this end, mice were infected with either HSV-1 or HSV-1UL38Δ81–90 via the intraperitoneal (i.p.) route injection. Loss of the STING antagonist function of UL38 (i.e., HSV-1UL38Δ81–90) significantly increased the survival of infected mice (Figure 4A), increased the antiviral immune response, and reduced viral replication in the lungs and spleens of infected mice (Figures 4B–4E).
Figure 4.
HSV-1-expressed UL38 inhibits immune signaling and promotes viral replication and pathogenesis
(A–E) Eight-week-old female C57BL/6J mice (n = 6) were infected with 1 × 107 plaque formation units (PFUs) of HSV-1 or 1 × 107 PFU of HSV-1UL38Δ81–90 by intraperitoneal injection (i.p.). Mouse survival was observed and recorded daily until 14 days post infection (d.p.i.) (A). At 4 d.p.i., total RNA was extracted from the lungs and spleen, and qRT-PCR was performed to measure the transcription of the indicated antiviral genes (B and C) and the HSV-1 UL30 gene (D and E).
(F–H) Eight-week-old female C57BL/6J mice (n = 6) were infected with HSV-1 (1 × 104 PFU) or HSV-1UL38Δ81–90 (1 × 104 PFU) via intracerebroventricular injection (i.c.v.). Mouse survival was observed and recorded daily until 14 d.p.i (F). At 0.5 d.p.i., total RNA was extracted from the brain, and the mRNA levels of the antiviral genes (G) and the HSV-1 UL30 gene (H) were measured via qRT-PCR.
(I) 8-week-old female C57BL/6J mice were i.c.v. infected with HSV-1 (1 × 104 PFU) or HSV-1UL38Δ81–90 (1 × 104 PFU). At 0.5 d.p.i., the murine brain was subjected to IP with anti-STING antibody or IgG, followed by immunoblots with indicated antibodies. All experiments were independently conducted in triplicate and repeated at least twice with reproducible results. The graph shows the means ± SD.
p values were calculated via one-way ANOVA.
Because HSV-1 infection often causes encephalitis, we infected murine brains with HSV-1 or HSV-1UL38Δ81–90 via the intracerebroventricular route injection. Compared with wild-type HSV-1, cerebral infection with HSV-1UL38Δ81–90 resulted in an increased survival rate (Figure 4F), increased innate immune response (Figure 4G), and decreased viral replication in infected murine brains (Figure 4H). Moreover, compared with those in HSV-1UL38Δ81–90-infected murine brains, the interactions among STING, TBK1, and IRF3 were markedly inhibited in the brain tissues of mice infected with wild-type HSV-1, demonstrating that UL38 did impair the interaction between STING-IRF3-TBK1 in vivo (Figure 4I).
Therefore, our results demonstrate that the STING antagonist function of UL38 facilitates HSV-1 evasion of innate immune signaling, which contributes to the replication and pathogenesis of HSV-1 under both systematic and cerebral infection conditions in vivo.
A STING-derived designer peptide targets the STING-UL38 interface
Because UL38 binds to STING to antagonize STING-dependent immune signaling, we speculate that if some peptides can specifically target the UL38-STING interface, they may disrupt this protein-protein interaction, thereby unlocking UL38-suppressed innate immune signaling to inhibit HSV-1 replication. Since we identified aa 161–190 within STING (STING161–190) as the region responsible for the UL38 interaction, we first designed a peptide designated W30L via the STING161–190 sequence, which likely competes with endogenous STING for UL38 interaction but does not affect the function of STING. Notably, the 47-57 a.a. of human immunodeficiency virus-1 trans-activator of transcription protein (TAT47–57) is used as the cell penetrating peptide (CPP) and is conjugated with this peptide and subsequent peptides. We then simulated the structure of the UL38-W30L interaction via AlphaFold3, confirming that the W30L peptide can interact with the aa 81–90 region of UL38 (Figure 5A), which is consistent with previous data (Figure 2D). In particular, the structural simulation suggests that four residues of Leu-Ile-Leu-Pro (LILP) within the STING161–190 region are the core amino acids that interact with UL38 (Figure 5A).
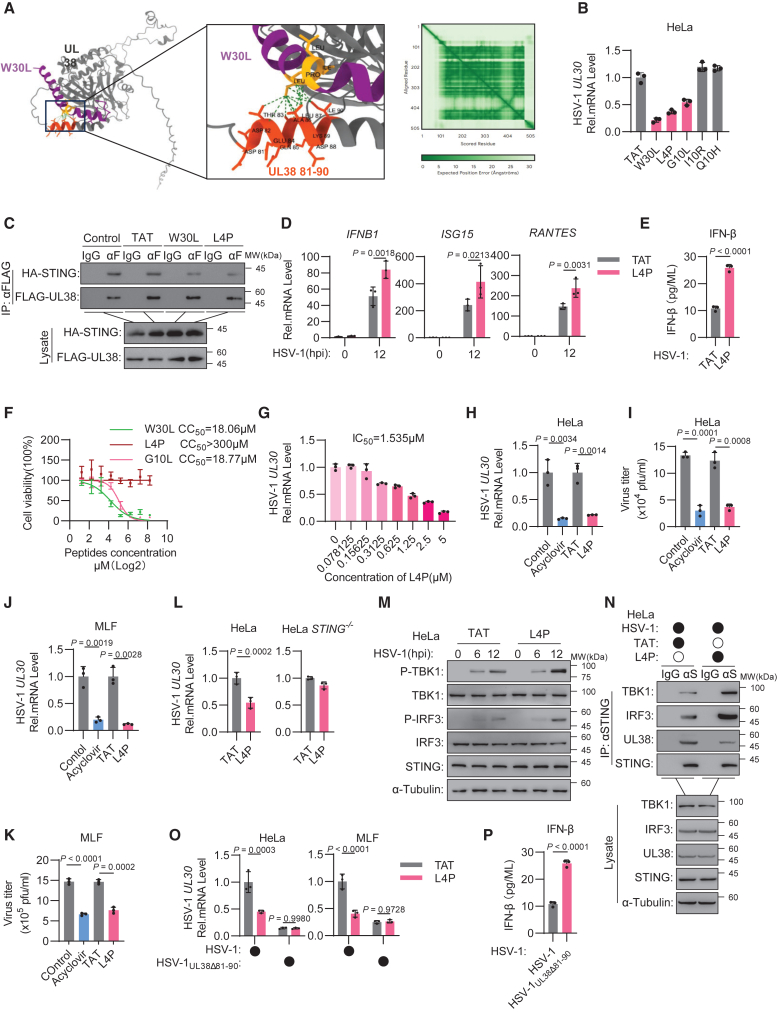
Figure 5.
A STING-derived designer peptide targets the STING-UL38 interface and shows antiviral efficacy in cells
(A) Left: the interaction sites between UL38 (gray) and the peptide W30L (purple) were predicted via AlphaFold3. The binding sites in W30L are shown in yellow, those in UL38 (amino acids 81–90) are shown in orange, and the interaction links are shown in green. Right: predicted aligned error (PAE) per token pair for the prediction on the left with rows and columns labeled by the chain ID and green gradient indicating PAE.
(B) HeLa cells were incubated with the indicated peptides (10 μM). After 24 h, the cells were infected with HSV-1 (MOI = 0.1); at 24 h.p.i., qRT-PCR was performed to measure the transcription of HSV-1 UL30.
(C) HeLa cells were transfected with FLAG-UL38 and HA-STING plasmids. At 24 h.p.t., the cells were incubated with the indicated peptide (10 μM) for 24 h. IP was performed using an anti-FLAG monoclonal antibody or IgG, followed by immunoblotting with anti-FLAG and anti-HA antibodies.
(D and E) HeLa cells were incubated with TAT (10 μM) or L4P (10 μM). After 24 h, the cells were infected with HSV-1 (MOI = 1); at 0, 6, or 12 h.p.i., qRT-PCR was performed to measure the transcription of the indicated genes (D), and ELISA was performed to measure the secretion of IFN-β (E).
(F) Increasing concentrations of L4P in DMEM supplemented with 2% FBS were added to HeLa cells for 12 h at 37°C. The viability of L4P cells was determined via a CCK-8 assay, and the absorbance at 450 nm was measured with a microplate reader (Infinite M200PRO).
(G) HeLa cells were incubated with increasing concentrations of L4P as indicated for 24 h, after which the cells were infected with HSV-1 (MOI = 0.1). At 24 h.p.i., qRT-PCR was performed to measure the transcription of the HSV-1 UL30 gene.
(H–K) HeLa cells (H and I) and MLFs (J and K) were incubated with acyclovir (10 μM) or buffer as a control and L4P (10 μM) or TAT as a control for 24 h. Then, the cells were infected with HSV-1 (MOI = 0.1), and at 24 h.p.i., qRT-PCR was performed to measure the expression of the HSV-1 UL30 gene (H and J), and the virus titer was detected by the plaque assay (I and K).
(L) HeLa and HeLa STING−/− cells were incubated with TAT (10 μM) or L4P (10 μM) for 24 h, after which the cells were infected with HSV-1 (MOI = 0.1). At 24 h.p.i., qRT-PCR was performed to measure the expression of the HSV-1 UL30 gene.
(M) HeLa cells were incubated with TAT (10 μM) or L4P (10 μM) for 24 h, after which the cells were infected with HSV-1 (MOI = 1). At 0, 6, or 12 h.p.i., the cell lysates were subjected to immunoblotting with the indicated antibodies.
(N) HeLa cells were incubated with L4P (10 μM) for 24 h and then infected with HSV-1 (MOI = 1). At 24 h.p.i., the cells were subjected to IP with an anti-STING monoclonal antibody or IgG, followed by immunoblotting with the indicated antibodies.
(O and P) HeLa cells and MLFs were incubated with L4P (10 μM) for 24 h and then infected with HSV-1 (MOI = 0.1) or HSV-1UL38Δ81–90 (MOI = 1). At 24 h.p.i., qRT-PCR was performed to measure the expression of the HSV-1 UL30 gene (O), and ELISA was performed to measure the secretion of IFN-β in HeLa cells (P). All experiments were independently conducted in triplicate and repeated at least twice with reproducible results. The graph shows the means ± SD, n = 3.
p values were calculated via one-way or two-way ANOVA. See also Figure S3.
As shown in Figures S3A and S3B, the peptide W30L showed potent antiviral activity that dramatically inhibited HSV-1 replication in infected HeLa cells. To further optimize the peptide, we designed a series of peptides based on the STING161–190 sequence (Figure S3C) and found that, in addition to W30L, the peptides L4P and G10L also exhibited antiviral activity against HSV-1 (Figure 5B). Notably, all 3 peptides contain the predicted core UL38-interacting residue LILP, among which peptide L4P uses the four residues (LILP) plus the CPP sequence. We investigated whether, as intended, these peptides target STING to disrupt the UL38-STING interaction and unlock antiviral immune signaling. The data revealed that treatment with either W30L or L4P inhibited the coIP of UL38 with STING (Figure 5C) and promoted immune gene transcription and IFN-β secretion in HSV-1-infected HeLa cells (Figures 5D, 5E, S3D, and S3E) or MLFs (Figure S3F).
We then assessed the cytotoxicity of these peptides. We found that the 50% cytotoxicity concentration (CC50) of the peptide L4P was >300 μM. In contrast, the CC50 of W30L or G10L was ∼18 μM in HeLa cells (Figure 5F). The inhibitory concentration (IC50) of the peptide L4P was 1.535 μM in HSV-1-infected HeLa cells (Figure 5G) and 1.637 μM in primary MLFs (Figure S3G). The selectivity index (SI = CC50/IC50) for L4P is greater than 200, indicating a high margin of safety. Therefore, we used the peptide L4P in subsequent experiments. Moreover, L4P treatment resulted in antiviral activity similar to that of acyclovir, a well-known anti-HSV-1 drug, in HSV-1-infected HeLa or MLFs (Figures 5H–5K).
To further study the mechanism of action of this anti-HSV-1 peptide, we examined its antiviral activity in normal or STING−/− HeLa cells. The results showed that the anti-HSV-1 activity of L4P is strictly STING dependent (Figure 5L), excluding the possibility that L4P inhibits HSV-1 through another mechanism. Moreover, treatment with L4P efficiently increased the phosphorylation of TBK1 and IRF3 in HSV-1-infected HeLa cells (Figure 5M) and MLFs (Figure S3H). L4P treatment inhibited the UL38-STING interaction but simultaneously enhanced the STING-TBK1-IRF3 interaction (Figure 5N). Furthermore, L4P treatment inhibited only wild-type HSV-1 but not STING interaction-defective HSV-1UL38Δ81–90 in infected HeLa cells or MLFs (Figures 5O and 5P) or reversed only the immunosuppressive effect of wild-type UL38 but not UL38Δ81–90 on DNA ligand-triggered immune signaling (Figure S3I), verifying that the STING-binding region within UL38 is the target of peptide L4P.
Together, we rationally designed the peptide L4P, which is derived from the UL38-binding region of STING and exerts potent antiviral efficacy against HSV-1 by specifically targeting the STING-UL38 interaction to unlock STING-dependent antiviral immunity.
Peptide L4P treatment shows potent in vivo antiviral efficacy by protecting mice from lethal HSV-1 infection
To explore the preclinical potential of the peptide L4P, we first administered fluorescently labeled L4P via i.p. injection and assessed its distribution in C57BL/6J mice. The data revealed that the fluorescent signals of Cy5-L4P were widely distributed throughout the whole body of the injected mice (Figures S4A–S4C). Compared with those from vehicle-treated mice, the brains, livers, spleens, lungs, kidneys, hearts, and intestines obtained from Cy5-L4P-treated mice presented significantly greater fluorescent signals (Figures S4D–S4K), indicating that the i.p. administration of L4P can result in a systematic distribution of the peptide in vivo.
We then performed an acute toxicity test to examine whether the peptide L4P causes obvious harm to the health of the mice. L4P treatment did not affect hepatic or renal function (Figures S4L and S4M) or survival (Figure S4N) in C57BL/6J mice, even at doses 5 or 10 times greater than those (10 mg/kg) used to protect mice from lethal HSV-1 infection.
To assess the in vivo therapeutic efficacy of the L4P peptide on the systematic infection of HSV-1, we tested the effect of i.p. delivered L4P in an HSV-1 systematic infection mouse model. We treated C57BL/6J mice with L4P at a dose of 10 mg/kg at 2 h post HSV-1 infection or 1 day before HSV-1 infection, followed by treatment twice a day. Notably, the CPP TAT was used as the control in this and subsequent experiments. Compared with TAT treatment, L4P treatment, either before or after infection, significantly increased the survival and reversed the trend of body weight reduction of HSV-1-infected mice (Figures 6A–6D). Consistent with the murine survival data, L4P showed potent anti-HSV-1 efficacy, as its treatment markedly inhibited the replication of HSV-1 in the lungs and spleens of infected mice (Figures 6E and 6F).
Figure 6.
Peptide L4P shows potent in vivo antiviral efficacy against lethal HSV-1 infection
(A–D) Model of the experiment (left of A and C). Groups (n = 10) of 8-week-old female C57BL/6J mice were treated with L4P or TAT at 10 mg/kg twice a day at 2 h post HSV-1 (1 × 107 PFU) infection via i.p. injection (A and B) or 1 day before HSV-1 infection (1 × 107 PFU) (C and D). Mouse survival (right of A and C) and body weight (B and D) were observed and recorded daily until 14 d.p.i.
(E–H) Groups (n = 6) of 8-week-old female C57BL/6J mice were challenged with HSV-1 (1 × 107 PFU) via i.p. injection and treated with L4P or TAT at 10 mg/kg twice a day. At 0, 4, or 7 d.p.i., total RNA was extracted from the lungs (E, G) and spleen (F, H). qRT-PCR was performed to measure the transcription of HSV-1 UL30 (E and F) or the indicated antiviral genes (G and H). All experiments were independently conducted in triplicate and repeated at least twice with reproducible results. The graph shows the means ± SD.
p values were calculated via two-way ANOVA. See also Figures S4 and S5.
On the other hand, L4P did not differ from TAT in terms of the survival rate or viral replication of HSV-1UL38Δ81–90-infected mice (Figures S5A–S5C), confirming that L4P targets the STING-binding interface in HSV-1 UL38 in vivo. Moreover, we treated C57BL/6J mice with acyclovir at a dose of 10 mg/kg at 2 h post HSV-1 infection, followed by treatment twice a day. The data revealed that the antiviral and therapeutic efficacies of L4P were comparable to those of acyclovir in HSV-1-infected mice (Figures S5F–S5I).
We also examined the effects of the peptide L4P on the innate immune signaling triggered by HSV-1 infection. L4P treatment significantly enhanced the transcriptional induction of immune genes, including Ifnb1, Isg15, Rantes, and Tnfα, in both the lungs and spleens of HSV-1-infected mice at 4 days post infection (d.p.i.) (Figures 6G and 6H) but not in those of HSV-1UL38Δ81–90-infected mice (Figures S5D and S5E), which is consistent with the mechanism of action of L4P, which unlocks UL38-suppressed innate immunity to confer antiviral effects (Figures 5D and 5E and 5L). Interestingly, our results also revealed that at 7 d.p.i., the enhancing effect of L4P treatment on innate immune signaling was diminished (Figures 6G and 6H), which should be attributed to restricted viral replication as well as host immune homeostasis, indicating that L4P treatment does not cause lasting immune elevation.
Therefore, our findings demonstrate that the systematic in vivo therapeutic effect of the designer peptide L4P is mediated by antiviral innate immune signaling and that its mechanism of action is via UL38 targeting.
Peptide L4P treatment protects mice from cerebral infection and the pathogenesis of HSV-1
HSV-1 infection in the brain can cause encephalitis, which results in severe neurological damage with a high risk of mortality. Therefore, we aimed to evaluate the therapeutic potential of the i.p. administered peptide L4P in a cerebral infection mouse model of HSV-1. Notably, i.p. administered L4P can enter murine brains (Figures S4D and S4K). To this end, we treated C57BL/6J mice with L4P at a dose of 10 mg/kg 2 h post infection or 1 day before HSV-1 infection, followed by L4P treatment twice a day. Consistent with the results of HSV-1 systematic infection, L4P treatment, either before or after infection, significantly increased the survival and reversed the trend of body weight reduction of mice with cerebral HSV-1 infection (Figures 7A–7D) but did not affect the survival of mice infected with HSV-1UL38Δ81–90 (Figure S6A). Consistent with these findings, L4P showed potent anti-HSV-1 efficacy, as its treatment markedly inhibited the replication of HSV-1 in the brains of infected mice (Figure 7E). On the other hand, L4P did not differ from TAT in terms of viral replication in the HSV-1UL38Δ81–90-infected murine brain (Figure S6B). Moreover, we treated C57BL/6J mice with acyclovir at a dose of 10 mg/kg at 2 h post HSV-1 cerebral infection, followed by treatment twice a day, and found that the antiviral and therapeutic efficacies of L4P were comparable to those of acyclovir in mice with HSV-1 cerebral infection (Figures S7A–S7C).
Figure 7.
L4P treatment protects mice from HSV-1 cerebral infection and mitigates its pathogenesis
(A–D) Model of the experiment (left of A and C). Groups (n = 10) of 8-week-old female C57BL/6J mice were treated with L4P or TAT at 10 mg/kg twice a day via i.p. injection at 2 h post HSV-1 (1 × 104 PFU) i.c.v. infection (A and B) or 1 day before HSV-1 (1 × 104 PFU) i.c.v. infection (C and D). Mouse survival (right of A and C) and body weight (B and D) were observed and recorded daily until 14 d.p.i.
(E–G) Groups (n = 6) of 8-week-old female C57BL/6J mice were i.c.v. infected with HSV-1 (1 × 104 PFU) and treated with L4P or TAT at 10 mg/kg twice a day via i.p. injection. At 0, 0.5, 3, or 5 d.p.i., total RNA was extracted from the brain, and qRT-PCR was performed to measure the transcription of HSV-1 UL30 (E), ELISA was performed to measure the secretion of IFN-β (F) and the indicated antiviral genes (G).
(H and I) 8-week-old female C57BL/6J mice were i.c.v. infected with HSV-1 (1 × 104 PFU). At 2 h.p.i., these mice were treated with L4P or TAT at 10 mg/kg twice a day via i.p. injection. At 0 and 5 d.p.i., the murine brain was subjected to immunofluorescence staining, which revealed 2× views of the striatum sections, views of the piriform cortex, and views of the olfactory tubercle (H). The murine striatum brain slices were subjected to H&E staining, which revealed 20× views of the piriform cortex and olfactory tubercle (I). All experiments were independently conducted in triplicate and repeated at least twice with reproducible results. Graph shows mean ± SD.
p values were calculated via one-way ANOVA or two-way ANOVA. See also Figures S6 and S7.
Consistent with the observations of HSV-1 systematic infection, L4P treatment significantly increased IFN-β secretion or immune gene induction at 0.5 d.p.i. in cerebrally infected mice, whereas this effect diminished at 3 or 5 d.p.i., implying that L4P treatment did not result in harmful immune induction in the brain (Figures 7F and 7G), whereas L4P treatment did not affect immune gene induction in mice infected with HSV-1UL38Δ81–90 (Figure S6C). Furthermore, we evaluated the effect of L4P treatment on HSV-1-induced pathogenesis in murine brains. Our results revealed that HSV-1 infection was associated with the accumulation of inflammatory CD11b+ cells in the striatum, piriform cortex, and olfactory tubercle of infected murine brains, whereas L4P treatment dramatically reduced the accumulation of both viruses and CD11b+ cells (Figure 7H). In addition, the piriform cortex and olfactory tubercle of HSV-1-infected murine brains were marked with cavitations, which were mitigated by L4P treatment (Figure 7I).
Together, our findings demonstrate that the peptide L4P protects mice from cerebral infection and the pathogenesis of HSV-1, indicating that L4P has therapeutic potential for treating HSV-1-associated encephalitis.
Discussion
In this study, we first identified the HSV-1-encoded UL38 as a STING antagonist and then revealed the mechanism by which UL38 inhibits STING and subsequent innate immune signaling. After that, we explored the possibility that the STING-interacting interface in HSV-1 UL38 could be a druggable target. We designed a series of STING-derived peptides to specifically target the UL38-STING interface and found that one of the designed peptides, L4P, effectively disrupts the STING-UL38 interaction to unlock the UL38-suppressed innate immune response and shows potent therapeutic efficacy against HSV-1 infection in both systematic and cerebral infection mouse models.
The life cycles of herpesviruses such as HSV-1 include distinct states, including primary infection, latency, and recurrence, which are sophisticatedly regulated by the interaction between host immunity and viral immune evasion mechanisms.16,34 STING plays a pivotal role in antiviral innate immunity against DNA viruses.35 STING-dependent immune signaling has been reported to tightly control the infection and latency of HSV-1 and other herpesviruses.36,37 Therefore, antagonizing STING or STING-mediated immune signaling is an efficient strategy to confer immune evasion by these viruses. Previous studies have shown that HSV-1-encoded ICP27 suppresses STING-dependent immune signaling by functioning downstream of TBK1 phosphorylation but upstream of IRF3 phosphorylation and that HSV-1 ICP27 is recruited to the TBK1-activated STING-TBK1 complex, thereby inhibiting TBK1-mediated IRF3 activation and subsequent immune induction.15 Similarly, HSV-1-encoded UL46 (also known as VP11/12) binds to both STING and TBK1 via separate domains, which probably interferes with the functions of STING.20
On the other hand, unlike ICP27 or UL46, HSV-1 UL38 directly binds to and inhibits STING itself but not TBK1 and disrupts cGAMP-binding to STING and the formation of the STING-TBK1-IRF3 complex, thereby acting as a direct STING antagonist that functions upstream of TBK1. Moreover, HSV-1-encoded UL37, UL41, UL36, VP22, and VP24 were also reported to antagonize the cGAS-STING pathway via different strategies.16,17,38,39,40 The requirement of multiple HSV-1-encoded proteins to evade cGAS-STING-dependent immune signaling via distinct mechanisms highlights the importance of suppressing STING and its downstream immune events for HSV-1, while UL38, in concert with these viral immune antagonists, should function collaboratively in different spatiotemporal manners to evade antiviral immunity in different stages of viral infection, which should be particularly important for viruses with complicated life cycles like herpesviruses. However, although HSV-1 employs these strategies to evade the cGAS-STING pathway, how to target these mechanisms to develop novel antivirals is unclear.
After uncovering the working mechanism of UL38 as a direct STING antagonist, we determined that the UL38-STING interface could be a potential target for the development of antivirals against HSV-1. Thus, we designed a series of peptides, particularly peptide L4P, on the basis of the STING sequence responsible for the UL38 interaction, which should mimic STING to interact with UL38, thereby disrupting the UL38-STING interaction, leading to the unlocking of UL38-suppressed immune signaling to confer antiviral efficacy. Thus, a concern is that the L4P-mediated disruption of STING signaling will cause some harmful inflammation. Our data show that although L4P treatment did have an immunoenhancing effect at the early stage of HSV-1 infection, the immunoenhancing effect gradually diminished as the peptide restricted the viral load at a low level. This observation is reasonable since the peptide L4P does not function as a STING agonist but unlocks the STING signaling that was originally suppressed by UL38, and the L4P-mediated increase in STING signaling has negative feedback with its antiviral efficacy. Therefore, targeting UL38 by L4P treatment would not cause lasting immune elevation.
Given the importance of immune evasion strategies for viruses, it is rational to consider them as targets for therapeutic interventions. In particular, for virus-encoded immunosuppressors that directly interact with key host immune proteins, targeting the viral protein-host protein interface, such as the UL38-STING interface we identified, should disrupt immune evasion to unlock the suppressed immune response. In this study, L4P and the other peptides we designed are derived from the amino acid sequence of the interface’s STING side. This design strategy not only allows peptides to target the UL38-STING interface without interfering with proper STING functionalities but also provides a unique advantage in avoiding possible drug resistance, at least theoretically. Indeed, if HSV-1 evolves to generate some UL38 mutations to escape peptide targeting, the mutants will also lose the UL38-STING interaction, resulting in the ability of the peptides to achieve this effect, thereby leaving the virus in an unresolvable dilemma. Considering that all the available anti-HSV-1 drugs, such as acyclovir, share the same target, viral DNA polymerase, and the emergence of resistance and cross-resistance to acyclovir and other anti-HSV-1 nucleoside analogs, identifying the STING antagonist function of UL38 as a druggable target and developing such a drug resistance-proof strategy should have particular importance as alternative therapies to HSV-1 and associated diseases. Given that most viruses employ their immune evasion mechanisms, this strategy may be applicable to the development of antivirals against other viruses.
Together, our studies identify HSV-1 UL38 as a STING antagonist and druggable target and design a peptide-based drug candidate that directly targets the STING-interacting interface of UL38 and shows potent in vivo therapeutic efficacy against HSV-1 infection. Our findings shed light on the understanding of the immune evasion mechanisms of HSV-1 and may represent a future direction of antiviral development.
Limitations of the study
First, because the antiviral efficacy of the peptide we developed requires intact STING-dependent immune signaling, this therapy is not applicable to patients who are deficient in the cGAS-STING pathway. Second, there is one amino acid difference between the STING-interacting (aa 81–90) region of HSV-1 UL38 and its corresponding region of HSV-2 UL38, which probably makes peptide L4P applicable only to HSV-1. Third, although peptides have the advantage of disrupting protein-protein interactions, there are several limitations associated with peptide-based drugs, including difficulty in oral administration and relatively short half-lives in vivo. Future studies may further optimize the UL38-targeting peptide, such as the use of D-isoform amino acids to increase its stability or the development of peptidomimetics based on the L4P sequence.
Resource availability
Lead contact
Further information and requests for resources and reagents should be directed to and will be fulfilled by the lead contact, Yujie Ren (renyujie@wh.iov.cn).
Material availability
All reagents generated in this study are available from the lead contact with a completed materials transfer agreement.
Data and code availability
All data reported in this paper will be shared by the lead contact upon request. This paper does not report the original code. Any additional information required to reanalyze the data reported in this paper is available from the lead contact upon request.
Acknowledgments
We are grateful to the Animal Center, Wuhan Institute of Virology, CAS, for their technical assistance. We thank all the members of the Zhou laboratory for their support.
This work was supported by the National Key R&D Program of China (2023YFC23066000 to Y.R.), the National Natural Science Foundation of China (82472274 to Y.R. and U21A20423 and 32225004 to X.Z.), the CAS Youth Innovation Promotion Association (2023351 to Y.R.), the Hubei Province Natural Science Funds (2023AFB582 to Y.R. and 2023AFA008 to X.Z.), the Young Top-Notch Talent Cultivation Program of Hubei Province (Y.R.), and the Fund of the Science and Technology Bureau of Wuhan (2023020201010086 to Y.R.).
Author contributions
A.W. performed most of the experiments. Q.P., H.F., W.J., and J.L. performed specific experiments. Y.R. and X.Z. conceived the idea, supervised the project, designed the experiments, analyzed the data, and wrote the manuscript with input from all the authors.
Declaration of interests
Wuhan Institute of Virology on behalf of Y.R., X.Z., and A.W. has filed a patent application for the use of antiviral peptide (Chinese patent application 202510237389.X).
STAR★Methods
Key resources table
| REAGENT or RESOURCE | SOURCE | IDENTIFIER |
|---|---|---|
| Antibodies | ||
| Mouse control IgG | Protein Tech Group | Cat# B900620 |
| Rabbit control IgG | Millipore | Cat# 12–370 |
| HRP-conjugated goat-anti mouse IgG | Thermo Fisher Scientific | Cat# PA1-86717; RRID:AB_933847 |
| HRP-conjugated goat-anti rabbit IgG | Thermo Fisher Scientific | Cat# SA1-9510; RRID:AB_1086400 |
| Anti-FLAG | Sigma-Aldrich | Cat# F3165; RRID:AB_259529 |
| Anti-HA | ABclonal | Cat# AE008; RRID:AB_2770404 |
| Anti-GFP | ABclonal | Cat# AE012; RRID:AB_2770402 |
| Anti-STING | Cell Signaling Technology | Cat# 13647; RRID: N/A |
| Anti-TBK1 | Cell Signaling Technology | Cat# 3504, RRID:AB_2255663 |
| Anti-IRF3 | Cell Signaling Technology | Cat# 4302, RRID:AB_1904036 |
| Anti-Phosoho-STING(Ser366) | Cell Signaling Technology | Cat# 82083, RRID: N/A |
| Anti-Phospho-TBK1 (Ser172) | Cell Signaling Technology | Cat# 5483; RRID: N/A |
| Anti-Phospho-IRF-3 (Ser396) | Cell Signaling Technology | Cat# 4947, RRID:AB_823547 |
| Anti-α-Tubulin | Cell Signaling Technology | Cat# 2144; RRID: N/A |
| Anti-UL38 Rabbit polyclonal Antibody | Atagenix | N/A |
| Bacterial and virus strains | ||
| Human alphaherpesvirus 1 (HSV-1) strain F | Wang et al., 2024 | N/A |
| HSV-1UL38Δ81-90 | This paper | N/A |
| Chemicals, peptides, and recombinant proteins | ||
| TAT47-57 (YGRKKRRQRRR) | This paper | N/A |
| W30L(YGRKKRRQRRRWSYYIGYLR LILPELQARIRTYNQHYNNLL) |
This paper | N/A |
| G10L(YGRKKRRQRRRGYLRLILPEL) | This paper | N/A |
| I10R(YGRKKRRQRRRILPELQARIR) | This paper | N/A |
| Q10H(YGRKKRRQRRRQARIRTYNQH) | This paper | N/A |
| L4P(YGRKKRRQRRRLILP) | This paper | N/A |
| Cy5-L4P(Cy5-YGRKKRRQRRRLILP) | This paper | N/A |
| Cell Counting Kit-8 (CCK-8) | Dojindo | Cat# CK04 |
| Proteinase K-Agarose | Sigma | Cat# P9290 |
| Protease/phosphatase inhibitor cocktail | TargetMol | Cat# C0002 |
| DMEM | Gibco | Cat# A4192101 |
| RPMI 1640 | Gibco | Cat# 11875119 |
| FBS | Gibco | Cat# 10099141C |
| 3×FLAG peptide | Sigma | Cat# F4799 |
| c[3′-Biotin-16-G(2′,5')pA(3′,5')p] | InvivoGen | Cat# C 196-001 |
| 2′3′-cGAMP | InvivoGen | Cat# tlrl-nacga23-02 |
| Streptomycin-penicillin | Gibco | Cat# 15140122 |
| Low-melting-point agarose | Sigma-Aldrich | Cat# A9414 |
| Type II collagenase | Worthington | Cat# LS005273 |
| DNase I | Thermo Fisher Scientific | Cat# EN0521 |
| Protease inhibitor cocktail | MCE | Cat# HY-K0010 |
| ECL | 4A Biotech | Cat# BN01036 |
| RNase inhibitor | Promega | Cat# N2115 |
| Protein A/G Magnetic Beads | MCE | Cat# HY-K0202 |
| Streptavidin Magnetic beads | MCE | Cat# 563175 |
| qPCR SYBR Green Master Mix | Yeasen | Cat# 11202ES03 |
| Puromycin | InvivoGen | Cat# ant-pr-1 |
| RNAiso plus | Takara | Cat# 9109 |
| EDC | Sigma-Aldrich | Cat# E7750 |
| acyclovir | MCE | Cat# HY-17422 |
| Critical commercial assays | ||
| Cell Total RNA Isolation Kit | Foregene | RE-03111 |
| Hieff® qPCR SYBR Green Master Mix | Yeasen | Cat# 11202ES08 |
| Mouse ALT ELISA Kit | Mlbio | Cat# ml063179-J |
| Mouse Creatinine (Cr) ELISA Kit | Mlbio | Cat# ml037726 |
| Experimental Models: Organisms/Strains | ||
| C57BL/6J | This pape | N/A |
| Experimental models: Cell lines | ||
| Primary MLF | This paper | N/A |
| HEK293 | ATCC | RRID:CVCL_0045 |
| HeLa | ATCC | RRID:CVCL_0030 |
| Vero | ATCC | N/A |
| THP-1 | ATCC | N/A |
| Oligonucleotides | ||
| Primers for qRT-PCR, see Table S1 | This paper | N/A |
| Primers for construction of recombinant plasmids, see Table S1 | This paper | N/A |
| siRNA sequence, see Table S1 | This paper | N/A |
| Software and algorithms | ||
| GraphPad Prism v6.03 | GraphPad Software | www.graphpad.com |
| CLC sequence viewer | CLC Bio | https://clc-sequence-viewer.software.informer.com/ |
| Pymol | Schrödinger | https://pymol.org/2/ |
| ChimeraX 1.8 | UC San Francisco | https://www.cgl.ucsf.edu/chimerax/download.html |
| QuantStudio Design & Analysis Software v2 | Thermo Fisher | https://www.thermofisher.cn |
Experimental model and subject details
Cell culture
HeLa, 293, Vero, and THP-1 cells were obtained from the American Type Culture Collection (ATCC). HeLa, 293 and Vero cells were cultured in DMEM (Gibco, A4192101), and THP-1 cells were cultured in RPMI 1640 (Gibco, 11875119), with 10% fetal bovine serum (FBS) (Gibco, 10099141C) and 1% streptomycin-penicillin (Gibco, 15140122) at 37°C in a CO2 incubator. The cell lines were tested for mycoplasma contamination.
Primary murine lung fibroblast culture
Primary MLFs were isolated from C57BL/6J mice as previously described.41 Briefly, murine lungs were minced and digested in magnesium- and calcium-free Hank’s balanced salt solution (HBSS) (Thermo) supplemented with 20 μg/mL DNase I (Thermo) and 10 mg/mL type II collagenase (Worthington) for 3 h at 37°C with shaking. The cell suspensions were filtered through progressively smaller cell strainers (100 and 40 mm). Filtered cells were plated in DMEM supplemented with 10% FBS, 100 U/mL penicillin, 100 μg/mL streptomycin and 2 mM L-glutamine. After 1 h, adherent fibroblasts were rinsed with HBSS 3 times and cultured for subsequent experiments.
Mice
All the mice were bred and cared under specific pathogen-free (SPF) conditions, and the viral infection experiments in this study were performed at the Biosafety Level 2 facility at the Animal Center of Wuhan Institute of Virology. All experiments were performed according to protocols approved by the Institutional Animal Care and Use Committee of the Wuhan Institute of Virology.
Eight-week-old C57BL/6J mice were randomly allocated to different groups. And sample size of animal experiments was estimated on the basis of similar researches reported in the literature.42 For HSV-1 infection, the mice were i.p. or i.c.v. injected with the indicated virus at 1 × 107 PFU or 1 × 104 PFU, respectively. For drug treatment, the mice were i.p. or i.c.v. injected with L4P or acyclovir at the indicated concentrations twice a day. For the in vivo toxicity assay, the ALT and creatinine levels in the sera from the mice before injection and 3 days after the final injection were measured via ALT and creatinine assay kits (Mlbio).
Viruses
The HSV-1 strain F were obtained from Microbial Culture and Virus Collection Center of Wuhan Institute of Virology, CAS (CSTR:16533.IVCAS6.9705) and propagated at a low MOI in Vero cells, and the virus titer was determined via standard plaque assays.
The HSV-1 recombinant viruses were derived from the pYEbac102 infectious clone of the HSV-1 strain F, and recombination was performed with the GS1783 E. coli strain. HSV-1UL38Δ81-90 was generated via two-step Red-mediated recombination, propagated and titrated in Vero cells.43
Peptides
TAT47-57, W30L, G10L, I10R, Q10H, and L4P were synthesized by GenScript Biochem with 95% purity. The peptides were dissolved in ultrapure water at a concentration of 2 mM stock solution in vitro and 10 mg/mL stock solution for in vivo experiments. The sequences of all the peptides are listed in the key resources table.
Method details
Plasmids, siRNAs and transfection
The DNA fragment of UL38 was amplified from the HSV-1 genome and then digested with restriction enzymes (SalI and NotI, Thermo Fisher, ER0641 and ER0593) and ligated to the pRK-FLAG vector with T4 ligase (Thermo Fisher, EP0061). The wild-type pRK-FLAG-UL38 construct was used as a template to generate the mutant UL38 constructs. The UL38 truncations were generated by subcloning the synthesized (Synbio Technologies) corresponding DNA fragments into the pRK-FLAG plasmid. The pGL3-IFNβ promoter, pGL3-ISRE, and NF-κB luciferase reporter plasmids, as well as the pRK-FLAG and pRK-HA vectors for cGAS, STING, IRF3, IRF-5D, TBK1, and TBK1 truncated mutants, have been previously described.42 The pRK-FLAG or pRK-HA for UL38 and UL38 truncations were generated using the vector for UL38 as a template. The primers used are listed in Table S1. All mutations were confirmed by DNA sequencing.
siRNAs specifically targeting UL38 were synthesized by RiboBio. For transfection, 100 nM siRNA or 500 ng plasmid was transfected with 3 μL of Lipofectamine 2000 reagent and 100 μL of Opti-MEM (Gibco, 22600134) into cells cultured in 24-well plates at 50–70% confluence. For other culture plates, the amounts of siRNA, plasmid, or Lipofectamine 2000 reagent were adjusted based on the bottom area of the well. The cells were collected at 24 h.p.i., and total RNA was extracted. All the siRNAs and oligonucleotides used here are listed in Table S1.
Reporter gene assays
293 cells were transiently transfected with the reporter plasmid (100 ng) and the internal control vector phRL-TK-Renilla luciferase (Promega, E1980) (20 ng) together with the indicated plasmids. The empty vector was added to equalize the total amount of DNA. After 24 h, luciferase assays were performed with a dual-specific luciferase assay kit (Promega, E1980). The firefly luciferase activity was normalized to Renilla luciferase activity to obtain the relative luciferase activity.
RNA extraction and quantitative real-time PCR (qPCR)
These experiments were performed as previously described.42 In brief, the cells were collected, and total RNA was isolated via a total RNA isolation kit (Foregene, RE-03111). The first-strand complementary DNA (cDNA) was reverse-transcribed with 1st strand cDNA Synthesis (Takara, D6110A). Gene expression was examined with a Bio-Rad SFX connection system by qPCR SYBR Green Master Mix (Yeasen, 11198ES03). The data were normalized to the expression of the gene encoding β-actin. All the qPCR primers used here are listed in Table S1. The assays were conducted via an ABI QuantStudio Q3 real-time PCR system, and the data were collected via QuantStudio design and analysis software v1.4.
Cell viability test
A CCK-8 assay was used to evaluate the cytotoxicity of the peptides in Vero cells. Briefly, the cells seeded into 96-well plates were incubated with increasing concentrations of peptides. After incubation at 37°C for 12 h, the cell mixture was replaced with fresh DMEM. After incubation for another 12 h, CCK-8 solution was added to detect the absorbance at 450 nm.
Co-immunoprecipitation and immunoblotting
These experiments were performed as previously described.44 Cells were collected and lysed in lysis buffer containing 20 mM Tris-HCl, 150 mM NaCl, 1 mM EDTA, 1% NP-40 and 1% protease and phosphatase inhibitor cocktail (TargetMol, C0002). The cell lysates were subjected to immunoblot analysis with the appropriate antibodies. For immunoprecipitation assays, the lysates were immunoprecipitated with protein G agarose and a control IgG or the appropriate antibodies (all the antibodies used here are listed in the key resources table), and the precipitates were washed 3 times with lysis buffer, followed by immunoblot analysis with enhanced chemiluminescence (ECL) (4A Biotech, BN01036).
Immunohistochemistry and immunofluorescence staining
Briefly, tissue samples were collected, mounted onto slides from paraffin blocks (5-μm sections), and then deparaffinized in xylene, followed by hydration in a methanol gradient (100%, 95%, 70%, and 50%). Slide samples were treated with 10 mM citrate buffer (pH 6.0) containing 3% H2O2 for antigen retrieval and then incubated with 5% BSA for 30 min. The slides were incubated with the primary antibody at 4°C overnight and then incubated with the biotinylated secondary antibody for 30 min. After that, an avidin-biotin complex kit (Dako/Agilent Technologies) was used for an additional 30 min, and 3,3′-diaminobenzidine tetrahydrochloride hydrate (DAB) containing 5% H2O2 was used as a chromogen. Hematoxylin and eosin (H&E) staining was performed, and the results were visualized via microscopy (Olympus).45 The antibodies used were as follows: anti-HSV-1 gD (Abcam, ab18638) and anti-CD11b (Abcam, ab133357).46
In vivo fluorescence imaging
Eight-week-old C57BL/6J mice were randomly assigned to two groups and i.p. injected with 10 mg/kg Cy5-L4P (n = 3) or vehicle (n = 3) (for background fluorescence measurements). At 1 h post injection, the distribution of Cy5-L4P in the mice was imaged via the IVIS Lumina K series III in vivo imaging system (PerkinElmer). To image and evaluate the fluorescence values of the brains, hearts, lungs, livers and kidneys, sodium pentobarbital was injected to sacrifice the mice, and the indicated tissues were obtained. Living Image 4.4 software was used to calculate the relevant radiant efficiency (Ps−1 cm−2 sr−1) (μW−1 cm2).
In vitro 2′3′-cGAMP binding assay
Reaction mixtures were prepared in 1.5 mL microcentrifuge tubes containing the following components per 50 μL reaction: 10 μL biotin-2′3′-cGAMP (0.1 μM; InvivoGen), 5 μg His-tagged UL38 protein, 10 μg FLAG-STING protein 5 μL 10 × reaction buffer (100 mM Tris-HCl, 500 mM KCl, 10 mM DTT), and 0.5 μL 200 mM EDTA (pH 8.0). The reaction mixtures were incubated at 37°C for 2 h with constant rotation (20 rpm). Subsequently, reaction volumes were adjusted to 800 μL by adding RIPA lysis buffer (50 mM Tris-HCl, 150 mM NaCl, 0.5% sodium deoxycholate, 1% IGEPAL CA-630). For affinity purification, 100 μL pre-washed streptavidin magnetic beads (MCE) were added to each reaction tube. Following overnight incubation at 4°C with rotation, beads were collected by centrifugation (4,000 × g, 5 min) and subjected to three washing cycles with TBST buffer (10 mM Tris-HCl, 150 mM NaCl, 0.1% Tween 20), each consisting of 1 min gentle vortexing followed by magnetic separation. Bound protein complexes were eluted in 2 × Laemmli buffer and resolved by 12% SDS-PAGE for subsequent immunoblot analysis.
Structure prediction
UL38-W30L binding complexes were modeled from sequences via AlphaFold3. AlphaFold3 was run through its public web server (https://www.alphafoldserver.com), which generated five models per complex, ranked by the AlphaFold3 ranking score.47 ChimeraX1.8 was used to analyze the predicted structure.
Quantification and statistical analysis
GraphPad Prism (v8.0.2.263) was used for the statistical analyses. The sample size and number of repeats are indicated in the respective figure legends. Statistical significance was calculated by one-way or two-way analysis of variance (ANOVA), as indicated in the figure legends. No animals or data points were excluded from the analyses.
Published: April 15, 2025
Footnotes
Supplemental information can be found online at https://doi.org/10.1016/j.xcrm.2025.102051.
Contributor Information
Xi Zhou, Email: zhouxi@wh.iov.cn.
Yujie Ren, Email: renyujie@wh.iov.cn.
Supplemental information
References
- 1.Yin Y., Favoreel H.W. Herpesviruses and the Type III Interferon System. Virol. Sin. 2021;36:577–587. doi: 10.1007/s12250-020-00330-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Lee S., Ives A.M., Bertke A.S. Herpes Simplex Virus 1 Reactivates from Autonomic Ciliary Ganglia Independently from Sensory Trigeminal Ganglia To Cause Recurrent Ocular Disease. J. Virol. 2015;89:8383–8391. doi: 10.1128/jvi.00468-15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Antony F., Kinha D., Nowińska A., Rouse B.T., Suryawanshi A. The immunobiology of corneal HSV-1 infection and herpetic stromal keratitis. Clin. Microbiol. Rev. 2024;37 doi: 10.1128/cmr.00006-24. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Gupta R., Warren T., Wald A. Genital herpes. Lancet. 2007;370:2127–2137. doi: 10.1016/s0140-6736(07)61908-4. [DOI] [PubMed] [Google Scholar]
- 5.Yin D., Ling S., Wang D., Dai Y., Jiang H., Zhou X., Paludan S.R., Hong J., Cai Y. Targeting herpes simplex virus with CRISPR-Cas9 cures herpetic stromal keratitis in mice. Nat. Biotechnol. 2021;39:567–577. doi: 10.1038/s41587-020-00781-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Armangue T., Spatola M., Vlagea A., Mattozzi S., Cárceles-Cordon M., Martinez-Heras E., Llufriu S., Muchart J., Erro M.E., Abraira L., et al. Frequency, symptoms, risk factors, and outcomes of autoimmune encephalitis after herpes simplex encephalitis: a prospective observational study and retrospective analysis. Lancet Neurol. 2018;17:760–772. doi: 10.1016/s1474-4422(18)30244-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Forel J.M., Martin-Loeches I., Luyt C.E. Treating HSV and CMV reactivations in critically ill patients who are not immunocompromised: pro. Intensive Care Med. 2014;40:1945–1949. doi: 10.1007/s00134-014-3445-y. [DOI] [PubMed] [Google Scholar]
- 8.Coen D.M. Acyclovir-resistant, pathogenic herpesviruses. Trends Microbiol. 1994;2:481–485. doi: 10.1016/0966-842x(94)90652-1. [DOI] [PubMed] [Google Scholar]
- 9.Ishikawa H., Barber G.N. STING is an endoplasmic reticulum adaptor that facilitates innate immune signalling. Nature. 2008;455:674–678. doi: 10.1038/nature07317. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Hopfner K.P., Hornung V. Molecular mechanisms and cellular functions of cGAS-STING signalling. Nat. Rev. Mol. Cell Biol. 2020;21:501–521. doi: 10.1038/s41580-020-0244-x. [DOI] [PubMed] [Google Scholar]
- 11.Zhang X., Bai X.C., Chen Z.J. Structures and Mechanisms in the cGAS-STING Innate Immunity Pathway. Immunity. 2020;53:43–53. doi: 10.1016/j.immuni.2020.05.013. [DOI] [PubMed] [Google Scholar]
- 12.Ma Z., Ni G., Damania B. Innate Sensing of DNA Virus Genomes. Annu. Rev. Virol. 2018;5:341–362. doi: 10.1146/annurev-virology-092917-043244. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Kalamvoki M., Roizman B. HSV-1 degrades, stabilizes, requires, or is stung by STING depending on ICP0, the US3 protein kinase, and cell derivation. Proc. Natl. Acad. Sci. USA. 2014;111:E611–E617. doi: 10.1073/pnas.1323414111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Orzalli M.H., DeLuca N.A., Knipe D.M. Nuclear IFI16 induction of IRF-3 signaling during herpesviral infection and degradation of IFI16 by the viral ICP0 protein. Proc. Natl. Acad. Sci. USA. 2012;109:E3008–E3017. doi: 10.1073/pnas.1211302109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Christensen M.H., Jensen S.B., Miettinen J.J., Luecke S., Prabakaran T., Reinert L.S., Mettenleiter T., Chen Z.J., Knipe D.M., Sandri-Goldin R.M., et al. HSV-1 ICP27 targets the TBK1-activated STING signalsome to inhibit virus-induced type I IFN expression. Embo j. 2016;35:1385–1399. doi: 10.15252/embj.201593458. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Zhang J., Zhao J., Xu S., Li J., He S., Zeng Y., Xie L., Xie N., Liu T., Lee K., et al. Species-Specific Deamidation of cGAS by Herpes Simplex Virus UL37 Protein Facilitates Viral Replication. Cell Host Microbe. 2018;24:234–248.e5. doi: 10.1016/j.chom.2018.07.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Su C., Zheng C. Herpes Simplex Virus 1 Abrogates the cGAS/STING-Mediated Cytosolic DNA-Sensing Pathway via Its Virion Host Shutoff Protein. J. Virol. 2017;91 doi: 10.1128/jvi.02414-16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Zheng Z.Q., Fu Y.Z., Wang S.Y., Xu Z.S., Zou H.M., Wang Y.Y. Herpes simplex virus protein UL56 inhibits cGAS-Mediated DNA sensing to evade antiviral immunity. Cell Insight. 2022;1 doi: 10.1016/j.cellin.2022.100014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Blest H.T.W., Redmond A., Avissar J., Barker J., Bridgeman A., Fowler G., Chauveau L., Hertzog J., Vendrell I., Fischer R., et al. HSV-1 employs UL56 to antagonize expression and function of cGAMP channels. Cell Rep. 2024;43 doi: 10.1016/j.celrep.2024.114122. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Deschamps T., Kalamvoki M. Evasion of the STING DNA-Sensing Pathway by VP11/12 of Herpes Simplex Virus 1. J. Virol. 2017;91 doi: 10.1128/jvi.00535-17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Jiang Z., Su C., Zheng C. Herpes Simplex Virus 1 Tegument Protein UL41 Counteracts IFIT3 Antiviral Innate Immunity. J. Virol. 2016;90:11056–11061. doi: 10.1128/jvi.01672-16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.You H., Zheng S., Huang Z., Lin Y., Shen Q., Zheng C. Herpes Simplex Virus 1 Tegument Protein UL46 Inhibits TANK-Binding Kinase 1-Mediated Signaling. mBio. 2019;10 doi: 10.1128/mBio.00919-19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Zhu H., Zheng C. The Race between Host Antiviral Innate Immunity and the Immune Evasion Strategies of Herpes Simplex Virus 1. Microbiol. Mol. Biol. Rev. 2020;84 doi: 10.1128/mmbr.00099-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Flanagan W.M., Papavassiliou A.G., Rice M., Hecht L.B., Silverstein S., Wagner E.K. Analysis of the herpes simplex virus type 1 promoter controlling the expression of UL38, a true late gene involved in capsid assembly. J. Virol. 1991;65:769–786. doi: 10.1128/jvi.65.2.769-786.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Zhao L., Zheng C. The first identified nucleocytoplasmic shuttling herpesviral capsid protein: herpes simplex virus type 1 VP19C. PLoS One. 2012;7 doi: 10.1371/journal.pone.0041825. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Li Y., Zhao L., Wang S., Xing J., Zheng C. Identification of a novel NLS of herpes simplex virus type 1 (HSV-1) VP19C and its nuclear localization is required for efficient production of HSV-1. J. Gen. Virol. 2012;93:1869–1875. doi: 10.1099/vir.0.042697-0. [DOI] [PubMed] [Google Scholar]
- 27.Xing J., Wang S., Li Y., Guo H., Zhao L., Pan W., Lin F., Zhu H., Wang L., Li M., et al. Characterization of the subcellular localization of herpes simplex virus type 1 proteins in living cells. Med. Microbiol. Immunol. 2011;200:61–68. doi: 10.1007/s00430-010-0175-9. [DOI] [PubMed] [Google Scholar]
- 28.Lui P.Y., Wong L.Y.R., Fung C.L., Siu K.L., Yeung M.L., Yuen K.S., Chan C.P., Woo P.C.Y., Yuen K.Y., Jin D.Y. Middle East respiratory syndrome coronavirus M protein suppresses type I interferon expression through the inhibition of TBK1-dependent phosphorylation of IRF3. Emerg. Microbes Infect. 2016;5 doi: 10.1038/emi.2016.33. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Zheng Y., Deng J., Han L., Zhuang M.W., Xu Y., Zhang J., Nan M.L., Xiao Y., Zhan P., Liu X., et al. SARS-CoV-2 NSP5 and N protein counteract the RIG-I signaling pathway by suppressing the formation of stress granules. Signal Transduct. Target. Ther. 2022;7:22. doi: 10.1038/s41392-022-00878-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Scrima N., Le Bars R., Nevers Q., Glon D., Chevreux G., Civas A., Blondel D., Lagaudrière-Gesbert C., Gaudin Y. Rabies virus P protein binds to TBK1 and interferes with the formation of innate immunity-related liquid condensates. Cell Rep. 2023;42 doi: 10.1016/j.celrep.2022.111949. [DOI] [PubMed] [Google Scholar]
- 31.Cao Z., Liu C., Peng C., Ran Y., Yao Y., Xiao G., Li E., Chen Z., Chuai X., Chiu S. Ebola virus VP35 perturbs type I interferon signaling to facilitate viral replication. Virol. Sin. 2023;38:922–930. doi: 10.1016/j.virs.2023.10.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Ergun S.L., Fernandez D., Weiss T.M., Li L. STING Polymer Structure Reveals Mechanisms for Activation, Hyperactivation, and Inhibition. Cell. 2019;178:290–301.e10. doi: 10.1016/j.cell.2019.05.036. [DOI] [PubMed] [Google Scholar]
- 33.Luo W.W., Li S., Li C., Lian H., Yang Q., Zhong B., Shu H.B. iRhom2 is essential for innate immunity to DNA viruses by mediating trafficking and stability of the adaptor STING. Nat. Immunol. 2016;17:1057–1066. doi: 10.1038/ni.3510. [DOI] [PubMed] [Google Scholar]
- 34.Muscolino E., Schmitz R., Loroch S., Caragliano E., Schneider C., Rizzato M., Kim Y.H., Krause E., Juranić Lisnić V., Sickmann A., et al. Herpesviruses induce aggregation and selective autophagy of host signalling proteins NEMO and RIPK1 as an immune-evasion mechanism. Nat. Microbiol. 2020;5:331–342. doi: 10.1038/s41564-019-0624-1. [DOI] [PubMed] [Google Scholar]
- 35.Ishikawa H., Ma Z., Barber G.N. STING regulates intracellular DNA-mediated, type I interferon-dependent innate immunity. Nature. 2009;461:788–792. doi: 10.1038/nature08476. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Guo Y., Jiang F., Kong L., Wu H., Zhang H., Chen X., Zhao J., Cai B., Li Y., Ma C., et al. OTUD5 promotes innate antiviral and antitumor immunity through deubiquitinating and stabilizing STING. Cell. Mol. Immunol. 2021;18:1945–1955. doi: 10.1038/s41423-020-00531-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Choi H.J., Park A., Kang S., Lee E., Lee T.A., Ra E.A., Lee J., Lee S., Park B. Human cytomegalovirus-encoded US9 targets MAVS and STING signaling to evade type I interferon immune responses. Nat. Commun. 2018;9:125. doi: 10.1038/s41467-017-02624-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Ye R., Su C., Xu H., Zheng C. Herpes Simplex Virus 1 Ubiquitin-Specific Protease UL36 Abrogates NF-κB Activation in DNA Sensing Signal Pathway. J. Virol. 2017;91 doi: 10.1128/jvi.02417-16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Huang J., You H., Su C., Li Y., Chen S., Zheng C. Herpes Simplex Virus 1 Tegument Protein VP22 Abrogates cGAS/STING-Mediated Antiviral Innate Immunity. J. Virol. 2018;92 doi: 10.1128/jvi.00841-18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Zhang D., Su C., Zheng C. Herpes Simplex Virus 1 Serine Protease VP24 Blocks the DNA-Sensing Signal Pathway by Abrogating Activation of Interferon Regulatory Factor 3. J. Virol. 2016;90:5824–5829. doi: 10.1128/jvi.00186-16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Fang Y., Liu Z., Qiu Y., Kong J., Fu Y., Liu Y., Wang C., Quan J., Wang Q., Xu W., et al. Inhibition of viral suppressor of RNAi proteins by designer peptides protects from enteroviral infection in vivo. Immunity. 2021;54:2231–2244.e6. doi: 10.1016/j.immuni.2021.08.027. [DOI] [PubMed] [Google Scholar]
- 42.Ren Y., Wang A., Wu D., Wang C., Huang M., Xiong X., Jin L., Zhou W., Qiu Y., Zhou X. Dual inhibition of innate immunity and apoptosis by human cytomegalovirus protein UL37x1 enables efficient virus replication. Nat. Microbiol. 2022;7:1041–1053. doi: 10.1038/s41564-022-01136-6. [DOI] [PubMed] [Google Scholar]
- 43.Li Y., Wang S., Zhu H., Zheng C. Cloning of the herpes simplex virus type 1 genome as a novel luciferase-tagged infectious bacterial artificial chromosome. Arch. Virol. 2011;156:2267–2272. doi: 10.1007/s00705-011-1094-9. [DOI] [PubMed] [Google Scholar]
- 44.Ren Y., Wang A., Zhang B., Ji W., Zhu X.X., Lou J., Huang M., Qiu Y., Zhou X. Human cytomegalovirus UL36 inhibits IRF3-dependent immune signaling to counterbalance its immunoenhancement as apoptotic inhibitor. Sci. Adv. 2023;9 doi: 10.1126/sciadv.adi6586. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Yang W., Tang C.Y., Fan D.Y., Wang Y.S., Wang P.G., An J., Luan G.M. Mice with type I interferon signaling deficiency are prone to epilepsy upon HSV-1 infection. Virol. Sin. 2024;39:251–263. doi: 10.1016/j.virs.2024.01.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Liu L., Chen Z., Zhang K., Hao H., Ma L., Liu H., Yu B., Ding S., Zhang X., Zhu M., et al. NSUN2 mediates distinct pathways to regulate enterovirus 71 replication. Virol. Sin. 2024;39:574–586. doi: 10.1016/j.virs.2024.05.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Abramson J., Adler J., Dunger J., Evans R., Green T., Pritzel A., Ronneberger O., Willmore L., Ballard A.J., Bambrick J., et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature. 2024;630:493–500. doi: 10.1038/s41586-024-07487-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
All data reported in this paper will be shared by the lead contact upon request. This paper does not report the original code. Any additional information required to reanalyze the data reported in this paper is available from the lead contact upon request.