Abstract
Whipple's disease is a rare multisystemic bacterial infection with variable clinical manifestations. For decades, the laboratory diagnosis was based on the demonstration of periodic acid Schiff-positive inclusions in macrophages of gastrointestinal biopsies. PCR has improved the diagnosis of Whipple's disease due to its increased sensitivity compared to histopathological analysis. To avoid invasive procedures for taking specimens, we have investigated the possibility of detecting Tropheryma whipplei DNA in feces rather than in biopsies or gastric aspirate of patients with and without Whipple's disease. Total bacterial DNA was isolated from stool specimens using Qiagen columns followed by a T. whipplei-specific hybridization step with a biotinylated capture probe and streptavidin-coated magnetic particles. The captured DNA was then amplified using the same seminested PCR targeting the 16S rRNA gene of the organism that had been applied to other specimens without capturing. For five of eight patients with Whipple's disease, duodenal biopsies and stool samples were PCR positive, whereas for the three other patients, both specimens were PCR negative. Of 84 patients without Whipple's disease, 75 tested negative in the duodenal biopsy and in the stool sample. For four, both specimens were positive. Five patients tested positive in the stool sample but not in the biopsy. However, for three of these five patients, the gastric aspirate had been PCR positive, indicating that the stool PCR result was true rather than false positive. Compared to PCR from duodenal biopsies, stool PCR has a sensitivity of 100% and a specificity of 97.3%. Additionally, 15 PCR-positive and 22 PCR-negative stool samples were extracted using the Invisorb Spin Stool DNA kit. The simplified stool extraction showed 93.3% sensitivity and 95.5% specificity compared to the target capture method. We conclude that PCR with stool specimens with either extraction method is a sensitive and specific diagnostic tool for the detection of T. whipplei DNA and one not requiring invasive sampling procedures.
Whipple's disease is a multisystemic bacterial infection which may involve any organ system in the body (5). It is caused by Tropheryma whipplei, a gram-positive rod-shaped bacterium which is classified within the actinobacteria (22). Several attempts to culture the bacterium have failed, until very recently (21, 25). The diagnosis of Whipple's disease is usually based on the presence of diastase-resistant non-acid-fast periodic acid Schiff (PAS)-positive inclusions in macrophages of intestinal biopsies or by electron microscopy showing the trilamellar membrane of T. whipplei (6). Since the sequences of the rRNA genes are known (14, 15, 22, 28), several PCR systems have been developed to identify parts of its 16S ribosomal DNA (rDNA) (2, 4, 23), of the 16S-23S rDNA internal transcribed spacer (ITS) (12), or of the 23S rDNA (13). These DNA amplification methods are more sensitive than histopathological analysis. However, T. whipplei DNA has also been found in duodenal biopsies, gastric aspirates, and saliva specimens of patients without clinical evidence of Whipple's disease (7, 26; H. U. Ehrbar, P. Bauerfeind, F. Dutly, H. R. Koelz, and M. Altwegg, Letter, Lancet 353:2214, 1999).
To avoid invasive procedures for obtaining suitable specimens, e.g., duodenal biopsies, we have investigated the use of stool specimens for the detection of T. whipplei DNA by PCR. Because multiple PCR inhibitors, such as hemoglobin, bilirubin, bile salts, urea, and heparin, may be present in stool samples (27), a suitable DNA extraction protocol which efficiently eliminates these inhibitors had to be established. For this purpose we have adapted a method previously described by Mangiapan et al. (16) which removes the inhibitors by capturing target DNA with specific probes. Large amounts of other DNA (human, bacterial) which may also reduce the sensitivity of PCR can simultaneously be removed with this method. In addition, we tested a new commercially available kit (Invisorb Spin Stool DNA Kit; Invitek, Berlin, Germany) and compared the results to those of the target capture method. The simplified stool extraction method is based on removing PCR inhibitors with InviAdsorb using a single centrifugation step, followed by the adsorption of the DNA to a column matrix and elution with a low-salt buffer.
(Parts of this study were presented at the 101st General Meeting of the American Society for Microbiology, Orlando, Florida, 20 to 24 May 2001 [R. C. Maibach, F. Dutly, and M. Altwegg, Abstr. 101st Gen. Meet. Am. Soc. Microbiol., abstr. C-456, p. 257, 2001]).
MATERIALS AND METHODS
Patients and specimens.
Stool samples of patients with or without confirmed Whipple's disease were analyzed, and results were compared to PCR from other specimens (duodenal biopsies, gastric juices, heart valves, and saliva). Patients with a positive PCR result in any specimen were contacted through their physician and asked for a stool sample. Most samples of asymptomatic persons originated from the previous prospective study done in collaboration with the University Hospital of Zurich (Ehrbar et al., letter). These stool samples were taken simultaneously with gastric aspirates and biopsies. Patients were divided into three different groups (Tables 1 and 2) as follows: I, patients with confirmed Whipple's disease and typical manifestations (endocarditis [11]), intestinal manifestations, spondylodiscitis [1]); II, patients with suspicion of Whipple's disease; and III, persons from the prospective study mentioned above without clinical signs of Whipple's disease. Stool samples were kept at −20°C until analysis.
TABLE 1.
Compilation of all results for patients with confirmed (group I) and suspected (group II) Whipple's diseasej
| Whipple's disease status | Patient No. | Specimen | Histology | PCR resulti
|
Reference | |||||
|---|---|---|---|---|---|---|---|---|---|---|
| 16S rDNA fragment
|
23S rDNA fragmentc | ITS Spacer typed | hsp65e | |||||||
| TW1/TW3a | TW1/TW2b | TW4/TW2b | ||||||||
| Confirmed | 1 | Duodenal biopsy | PAS-negg | pos | pos | pos | 1 | pos | ||
| Mononuclear cells | neg | |||||||||
| Stool | pos | pos | pos | |||||||
| 2f | Aortic valve | PAS-neg | pos | pos | pos | 1 | pos | |||
| Gastric juice | neg | neg | ||||||||
| Paraffin-embedded duodenal biopsy | neg | neg | ||||||||
| Paraffin-embedded heart valve | pos | pos | pos | 1 | pos | |||||
| Duodenal biopsy | neg | neg | ||||||||
| Stool | neg | neg | neg | |||||||
| 3 | Terminal ileum | pos | pos | pos | 2 | pos | ||||
| Duodenal biopsy | neg | |||||||||
| Stool | pos | pos | pos | |||||||
| 4 | Intestinal biopsy | PAS-neg | pos | pos | pos | pos | 1 | pos | Brühlmann et al. (3) (patient 2) | |
| Synovial fluid | pos | pos | pos | |||||||
| Joint fluid knee left | pos | neg | neg | neg | neg | neg | ||||
| Synovia knee joint (Synovia Kniegelenk li.) | PAS-neg | neg | neg | neg | ||||||
| Ileum fluid | neg | |||||||||
| Biopsy | neg | |||||||||
| Stool | pos | pos | pos | |||||||
| 5 | Intervertebral disc biopsy | pos | pos | neg | 2 | Altwegg et al., 1996 (1) | ||||
| Terminal ileum biopsy | PAS-pos | pos | ||||||||
| Colon asc. biopsy | PAS-neg | pos | ||||||||
| Pars III duodenal biopsy | PAS-neg | pos | ||||||||
| Pars II biopsy | PAS-neg | pos | ||||||||
| CSF | neg | |||||||||
| Stool | pos | pos | pos | |||||||
| 6 | Duodenal biopsy | PAS-neg | pos | pos | 1 | pos | Brühlmann et al. (3) (patient 1) | |||
| Synovial fluid | pos | pos | 1 | pos | ||||||
| Synovial biopsy | PAS-neg | neg | ||||||||
| Plasma | neg | |||||||||
| Leukocyte sediment | neg | |||||||||
| Stool | pos | pos | pos | |||||||
| 7f | Heart valve | PAS-pos | pos | pos | pos | 2 | Gubler et al. (11) | |||
| Paraffin-embedded duodenal biopsy | PAS-neg | neg | (patient 1) | |||||||
| Paraffin-embedded heart valve | PAS-pos | neg | ||||||||
| Stool | neg | neg | neg | |||||||
| 8f | Duodenal biopsy | PAS-neg | neg | neg | Gubler et al. (11) (patient 4) | |||||
| Mitral valve | PAS-pos | pos | pos | |||||||
| CSF | neg | neg | ||||||||
| Stool | neg | neg | neg | |||||||
| Suspected | 9 | Paraffin-embedded intestinal biopsy | neg | |||||||
| Paraffin-embedded sigma | neg | |||||||||
| Paraffin-embedded colon | neg | |||||||||
| Paraffin-embedded ileum | neg | |||||||||
| Stool | neg | neg | neg | |||||||
| 10 | Terminal ileum | pos | pos | pos | pos | 2/3h | pos | |||
| Duodenal biopsy | neg | neg | neg | pos | neg | |||||
| Blood | neg | neg | neg | |||||||
| Stool | pos | pos | pos | /PICK>{tt} | ||||||
| 11 | Terminal ileum | neg | ||||||||
| Duodenal biopsy | neg | pos | 1 | |||||||
| Stool | neg | neg | neg | |||||||
| 12 | Duodenal biopsy | pos | neg | pos | 1 | |||||
| CSF | neg | |||||||||
| Stool | neg | pos | pos | |||||||
Primers TW1 and TW3 (23).
Species-specific seminested PCR with primers TW1 and TW2 followed by TW4 and TW2 (2).
23S rRNA Insertion PCR (13).
ITS-PCR (12).
hsp65-PCR (18).
Patients with Whipple endocarditis.
Before diastase reaction: PAS-positive.
Spacertype not unambigously identified.
pos, positive; neg, negative.
Blanks, no data available.
TABLE 2.
Patients without Whipple's disease (group III)a
| Results by 16S rRNA fragment PCR
|
No. of patients | ||
|---|---|---|---|
| Duodenum | Gastric aspirate | Stool | |
| neg | neg | neg | 70 |
| neg | neg | posb | 2 |
| neg | pos | neg | 5 |
| neg | pos | pos | 3 |
| pos | pos | pos | 4 |
pos, positive; neg, negative.
Both were again positive upon repeat analysis.
Controls.
As a positive control, a lyophilized Escherichia coli strain with a cloned T. whipplei PCR fragment was used (2). As a negative control, either a normal lyophilized E. coli strain or water was used. They were treated as a clinical specimen.
Extraction of DNA.
One milliliter of stool was suspended in 1 ml of 0.85% NaCl and then centrifuged at low speed (500 × g) for 10 min. The pellet was discarded, and the supernatant was again centrifuged at high speed (10,000 × g) for 10 min to pellet the bacteria. The pellet was then washed once with 1 ml of 0.85% NaCl, resuspended in 200 μl of digestion buffer (50 mM Tris-HCl [pH 8.5], 1 mM EDTA, 0.5% sodium dodecyl sulfate) containing 400 μg of proteinase K/ml and incubated at 55°C for 1 to 2 h with agitation. Total DNA was extracted with QIAamp DNA binding columns (QIAGEN, Basel, Switzerland) according to the manufacturer's protocol except that in the final step only 100 μl (instead of 200 μl) of elution buffer AE was used.
Oligonucleotides.
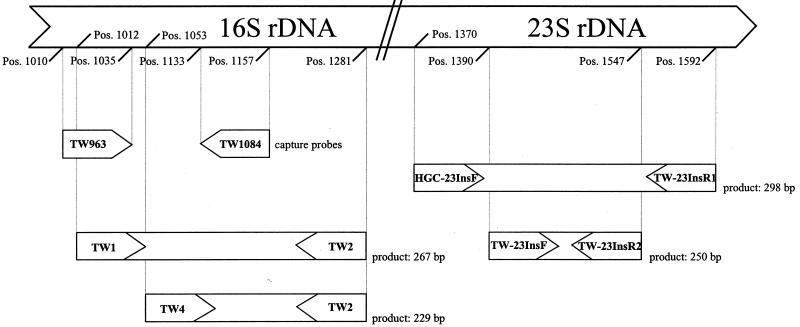
For sequence capturing of the T. whipplei DNA, 5′-biotinylated oligonucleotides TW963f (5′-GTAGAGATACGCCCCCCGCAAGGT) and TW1084r (5′-GTCTCCTGTGAGTCCCCGCCATTAC) designed using the software Oligo 4.1 (Wojciech Rychlik, National Biosciences) were used. These oligonucleotides are complementary to the 16S rDNA specific for T. whipplei in the area between primers TW1 and TW3 (Fig. 1). For amplification of the 16S rRNA, primers TW-1 and TW-2 were used, resulting in a 267-bp fragment, whereas seminested reamplification with TW-4 and TW-2 produced a 229-bp fragment (2). For amplification of the hypervariable region of domain III of the 23S rDNA, primers HGC-23InsF and TW-23InsR1 and for nested reamplification TW-23InsF and TW-23InsR2 were used (13). All primers and capture oligonucleotides were synthesized by Microsynth (Balgach, Switzerland).
FIG. 1.
Schematic overview of the positions of primers and probes used for amplification and capturing, respectively. Positions are shown according to the rRNA genes of E. coli.
Sequence capture.
The method used was based on that of Mangiapan et al. (16) with the following modifications. All eluate (100 μl) from the Qiagen columns was used for capture hybridization. First, the eluate was heated at 95°C for 10 min and cooled on ice. Then, 36.4 μl of 3.75 M NaCl containing 0.125 pmol each of the biotinylated oligonucleotides TW963f and TW1084r was added. Tubes were incubated with agitation at 60°C for 3 h on a thermomixer to allow hybridization. Two microliters of M-280 Streptavidin Dynabeads (Dynal, Oslo, Norway), washed according to the manufacturer's instructions, was added, and the incubation was continued for 2 h at 20°C. Magnetic beads with the oligonucleotides and the hybridized DNA were washed twice with 100 μl of TE wash buffer (10 mM Tris-HCl, 0.1 mM EDTA [pH 8]) and resuspended in 50 μl of TE resuspend buffer (10 mM Tris-HCl, 1 mM EDTA [pH 8]). Five microliters of this suspension was directly used for PCR without removing the magnetic particles.
Invisorb Spin Stool DNA kit.
DNA was extracted from 15 PCR-positive and 22 PCR-negative stool specimens (180 to 300 mg) according to the manufacturer's protocol (Invitek, Berlin, Germany), except that in the final step only 100 μl (instead of 200 μl) of elution buffer was used. Again, 5 μl of the eluate was used for PCR.
Amplification of T. whipplei DNA.
PCR was performed on a Gene Amp PCR System 9600 (Perkin-Elmer) and included an initial activation-denaturation step of 9 min at 95°C and 2 min at 96°C. Amplification and reamplification were both done in a volume of 50 μl containing dATP, dCTP, dGTP, and dUTP at a concentration of 200 μM each, 25 pmol of the corresponding primers, 2.5 U of AmpliTaq Gold polymerase including the appropriate amount of its optimized buffer (Perkin-Elmer, Norwalk, Conn.), and 5 μl of DNA template. For amplification, but not for reamplification, 0.25 U of thermolabile uracil-N-glycosylase (Epicentre Technologies, Madison, Wis.) was added to prevent carryover contamination from previous amplifications. For reamplification, 5 μl of the first amplification mixture was used as a template. Amplification consisted of 40 cycles at 95°C for 1 min, at 60°C for 1 min, and at 72°C for 1 min. Seminested reamplification was performed for 15 cycles at 95°C for 1 min, at 60°C for 1 min, and at 72°C for 1 min. To avoid degradation of the amplified products because of the uracil-N-glycosylase (24), reaction tubes were held at 72 or −20°C after cycling and then stored at −20°C. Amplifications and reamplifications with primers targeting the hypervariable region of domain III of the 23S rDNA were performed as previously described (13). PCR products were separated by electrophoresis on a 2% (wt/vol) agarose gel (UltraPure agarose; Life Technologies, Paisley, Scotland), stained with ethidium bromide, and visualized under UV light.
Amplification control.
All PCR-negative DNA extracts were tested for inhibition of the Taq polymerase by adding positive control DNA to the amplification mix. When still negative, extracts were diluted 1:5 and both PCR and amplification control were repeated.
RESULTS
Patients with confirmed Whipple's disease (group I).
All available information for these eight patients is summarized in Table 1. For five patients, both the duodenal biopsy and the stool specimen were positive, but only one of these had PAS-positive inclusions in one of four duodenal biopsy samples. Three patients had endocarditis caused by T. whipplei and tested positive in the heart valve only. In two of these three patients, the heart valve showed typical PAS-positive structures. The results of the PCR targeting the hypervariable region of the 23S rRNA gene always matched those of the 16S rDNA.
Patients with suspected Whipple's disease (group II).
For two of these four patients, the duodenal biopsy as well as the stool specimen had a positive PCR result. These two patients also tested positive in the stool by the 23S rRNA PCR system. For the other two patients, all specimens were negative with the 16S rRNA PCR system. However, patient 10 tested positive in the duodenal biopsy with the 23S rRNA and the ITS PCR systems (Table 1).
Patients without Whipple's disease (group III).
Of the 84 patients with elective gastroscopy (Ehrbar et al., letter) but no clinical signs suggestive of Whipple's disease, 70 tested PCR negative in the duodenal biopsy, in the gastric aspirate, and in the stool, whereas for 14, at least one of the specimens analyzed was positive (Table 2). All results of the 23S rRNA PCR system corresponded to those of the 16S rRNA PCR system except for the following three patients: (i) the two patients with positive results only for the stool but not for the biopsy or the gastric aspirate were considered as false positive, although they were again positive after reextraction and PCR with the 16S rRNA PCR system but were negative with the 23S rRNA PCR system; (ii) one of the three patients testing PCR positive with the 16S rRNA PCR system for the stool and the gastric aspirate but not for the duodenal biopsy was also negative with the 23S rRNA PCR system for the stool.
Amplification control.
Among all 96 stool samples analyzed, 10 were shown to be inhibited when DNA extracts were tested undiluted. At a dilution of 1:5, all these samples showed a positive amplification control.
Sensitivity and specificity.
Based on the above-described results, the sensitivity of the stool PCR in combination with the target capture DNA extraction method is 100% compared to PCR-positive results in the duodenal biopsies irrespective of the presence of Whipple's disease (Table 3). For calculation of the specificity of the assay, only specimens from patients without clinical evidence of Whipple's disease and negative PCR results in duodenal biopsies and gastric aspirates were included. Only 2 of 72 stool results were considered false positive, resulting in a specificity of 97.3%. However, both these specimens again tested positive upon repeat analysis (including a new DNA extraction from the same stool specimen) (see above).
TABLE 3.
Performance of stool PCR using the target capture methoda
| Patient group | No. of patients | Results
|
Sensitivityc | Specificity | |||
|---|---|---|---|---|---|---|---|
| Duodenal biopsy
|
PCR gastric aspirate | No. positive by stool PCR | |||||
| Histology | PCR | ||||||
| With WD | 1 | pos | pos | ndb | 1 | 5/5d | |
| 4 | neg | pos | nd | 4 | |||
| 3 | neg | neg | nd | 0 | |||
| Without WD | 7 | neg | pos | pos | 7 | 7/7d | |
| 8 | neg | neg | pos | 3 | |||
| 72 | neg | neg | neg | 2 | 70/72e | ||
The sensitivity was calculated based on specimens that were PCR positive in duodenal biopsies, whereas only specimens from patients without Whipple's disease (WD) that had been negative by PCR in duodenal biopsies and gastric aspirate were considered for specificity calculations. Total sensitivity, 100%; total specificity, 97.3%.
nd, no data available.
Compared to positive PCR from duodenal biopsies.
Number of patients with positive stool PCR/number with positive PCR in duodenal biopsy.
Number of patients with negative stool PCR/number with negative PCR in both duodenal biopsy and gastric juice.
DNA extraction with the Invisorb Spin Stool DNA kit.
Among 15 PCR-positive stool samples extracted with the sequence capture method, 13 were also positive with the Invisorb Spin Stool DNA kit. From 22 PCR-negative stool samples, 20 were also negative when extracted with the Invisorb kit. This results in a sensitivity of 87% and a specificity of 91%. These DNA extracts were amplified only with the 16S rRNA PCR system.
DISCUSSION
The variable and nonspecific manifestations of Whipple's disease make its clinical diagnosis very difficult. Laboratory confirmation is hampered by the necessity for specimens requiring invasive procedures (e.g., duodenal biopsies, cerebrospinal/joint fluid) and by the fact that gastrointestinal specimens may be negative by both histopathology (PAS stain) and PCR, although PCR has been shown to be more sensitive than PAS staining (8). Considering that Whipple's disease is rare, a sensitive screening test not requiring invasive specimens would be extremely helpful. Since no serodiagnostic tests are routinely available, we have evaluated the usefulness of PCR in detecting DNA of T. whipplei in stool specimens of patients with and without Whipple's disease. In contrast to a previous study with one single patient, which used three consecutive PCR amplifications with the inherent risk of carry-over contamination (10), we established a new DNA extraction-purification procedure based on target capturing by hybridization to enrich target sequences and to simultaneously remove inhibitors for the Taq polymerase abundantly present in stool specimens (16, 27).
A total of 12 patients with confirmed or suspected Whipple's disease and 84 persons undergoing elective endoscopy but without symptoms suggestive for Whipple's disease (e.g., absence of diarrhea, weight loss, and arthralgias) (Ehrbar et al., letter) were included in the study. Results of the stool PCR were primarily compared to PCR results from duodenal biopsies and gastric juice, data available not only for most patients (Table 1) but also for all controls.
As shown in Table 3, all persons for whom duodenal biopsies were PCR positive were also positive by 16S rRNA PCR with their stool specimens irrespective of the presence or absence of Whipple's disease. In addition, PCR targeting domain III of the 23S rDNA was also positive with all these specimens. Thus, the sensitivity of our stool PCR is 100% if compared to PCR from duodenal biopsies (Table 3). However, the clinical sensitivity, i.e., the sensitivity with regard to the detection of patients with Whipple's disease, is only 63% but still significantly higher than that of histopathology (4, 17, 19, 23), with a clinical sensitivity of only 12.5% in our small series. This confirms that PCR from duodenal biopsies and stool specimens is significantly more sensitive than histopathology and that a number of patients with Whipple's disease show no gastrointestinal evidence (clinical or laboratory) whatsoever for the presence of this disease (9, 11, 20).
In contrast, there are persons without Whipple's disease who carry T. whipplei DNA in their gastrointestinal tract. All seven patients without Whipple's disease but PCR positive in their duodenal biopsies were also positive by 16S rRNA stool PCR. This and the fact that these specimens were also positive by 23S rRNA PCR support the view that asymptomatic carriage of T. whipplei, or at least its DNA, indeed exists, as suggested previously (7, 26; Ehrbar et al., letter). Among the eight persons without Whipple's disease that were PCR positive with gastric juice but not with duodenal biopsies, only three were positive by stool PCR as well. This may reflect a quantitative phenomenon influenced by whether or not the organism penetrates the intestinal mucosa.
To circumvent problems with asymptomatic carriage, the specificity of the stool PCR was analyzed with reference to those persons without evidence for Whipple's disease who were PCR negative with both gastric juice and duodenal biopsy and was calculated to be 97.3%. However, both false-positive specimens were again positive when the entire procedure, i.e., including DNA extraction, was repeated. In contrast, both were negative by 23S rRNA PCR. However, when the DNA extraction was performed with the Invisorb kit, one of the two was also positive. If this sample is considered as being correctly positive, the specificity of the 16S rRNA stool PCR rises to 98.6%. Whether or not very low amounts of T. whipplei DNA at the limits of its detectability for the 23S but not the 16S rRNA PCR system or true false positivity is responsible for these contradictory results remains unsolved.
We were astonished that the results of the 23S rRNA PCR system are comparable to those of the 16S rRNA PCR system, because T. whipplei DNA was captured after column purification with a probe targeting the 16S rRNA gene. The hypervariable region in domain III of the 23S rDNA, where the 23S rRNA primers anneal, is located approximately 2,000 bp downstream from the capturing positions within the 16S rRNA gene. This shows that the captured sequences are at least 2,000 bp long. Thus, the capturing method is suitable for the extraction of long DNA sequences out of difficult samples such as stool or possibly also environmental samples.
The results with the Invisorb kit are comparable to those with the target capture method. One of the two false-negative stool samples using this simplified procedure was positive with the capture method but had been classified as false positive because the duodenal biopsy, the gastric aspirate, and the 23S rRNA PCR from stool were PCR negative. Thus, the result with the Invisorb kit is probably correctly negative. The other sample which was considered false positive with the capture method was also positive with the Invisorb kit (see above). Therefore, the sample has to be considered a true positive. A third sample only weakly positive with the capture method was negative with the Invisorb kit. This may be a problem of storing the stool samples for months at −20°C with a concomitant deterioration of the DNA quality. Another sample negative with the capture method but positive with the Invisorb kit was considered a true positive because the gastric aspirate of the same patient had been positive with the TW1-TW3 PCR system (23) as well as the TW4-TW2 PCR system (2). The last discrepancy involved a specimen which was negative with the capture method but weakly positive with the Invisorb kit. Since no information is available that would suggest otherwise, the Invisorb result has to be considered a true false positive. These adjustments result in a sensitivity of 93.3% and a specificity of 95.5% for PCR from Invisorb-treated stool specimens. Due to the easier handling and because it is less time consuming (i.e., 3 to 4 h for extraction time, versus 9 h for the capturing method), the Invisorb kit is a suitable alternative for use in the routine diagnostic laboratory.
Based on the above-described results, we conclude that PCR from stool specimens is more sensitive than histopathology and about as sensitive as PCR from duodenal biopsies for patients with Whipple's disease. However, positive results do not confirm Whipple's disease and require further evaluation of the patients, as is the case for PCR results from duodenal biopsies and gastric juice. Similar results have meanwhile been obtained with healthy control persons, with a prevalence of positive PCR assays in stool specimens of about 7% (L. Amsler, P. Bauerfeind, Ch. Nigg, R. C. Maibach, R. Steffen, and M. Altwegg, unpublished data). Finally, DNA extraction from stool specimens with the Invisorb kit is a suitable alternative to target capturing in a diagnostic laboratory.
Acknowledgments
This work was supported by a grant from the Swiss National Science Foundation (No. 32-61602.00) to M. Altwegg.
REFERENCES
- 1.Altwegg, M., A. Fleisch-Marx, D. Goldenberger, S. Hailemariam, A. Schaffner, and R. Kissling. 1996. Spondylodiscitis caused by Tropheryma whippelii. Schweiz. Med. Wochenschr. 126:1495-1499. [PubMed] [Google Scholar]
- 2.Brändle, M., P. Ammann, G. A. Spinas, F. Dutly, R. L. Galeazzi, C. Schmid, and M. Altwegg. 1999. Relapsing Whipple's disease presenting with hypopituitarism. Clin. Endocrinol. 50:399-403. [PubMed] [Google Scholar]
- 3.Brühlmann, P., B. A. Michel, and M. Altwegg. 2000. Diagnosis and therapy monitoring of Whipple's arthritis by polymerase chain reaction. Rheumatology 39:1427-1428. [DOI] [PubMed] [Google Scholar]
- 4.Dauga, C., I. Miras, and P. A. Grimont. 1997. Strategy for detection and identification of bacteria based on 16S rRNA genes in suspected cases of Whipple's disease. J. Med. Microbiol. 46:340-347. [DOI] [PubMed] [Google Scholar]
- 5.Dobbins, W. O., III. 1987. Whipple's disease. Charles C. Thomas, Springfield, Ill.
- 6.Dobbins, W. O., III. 1995. The diagnosis of Whipple's disease. N. Engl. J. Med. 332:390-392. [DOI] [PubMed] [Google Scholar]
- 7.Dutly, F., H. P. Hinrikson, T. Seidel, S. Morgenegg, M. Altwegg, and P. Bauerfeind. 2000. “Tropheryma whippelii” DNA in saliva but not in dental plaques of patients without Whipple's disease. Infection 28:219-222. [DOI] [PubMed] [Google Scholar]
- 8.Dutly, F., and M. Altwegg. 2001. Whipple's disease and “Tropheryma whippelii.” Clin. Microbiol. Rev. 14:561-583. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Geissdörfer, W., I. Wittmann, G. Seitz, R. Cesnjevar, M. Röllinghoff, C. Schoerner, and C. Bogdan. 2001. A case of aortic valve disease associated with Tropheryma whippelii infection in the absence of other signs of Whipple's disease. Infection 29:44-47. [DOI] [PubMed] [Google Scholar]
- 10.Gross, M., C. Jung, and W. G. Zoller. 1999. Detection of Tropheryma whippelii DNA (Whipple's disease) in faeces. Ital. J. Gastroenterol. Hepatol. 31:70-72. [PubMed] [Google Scholar]
- 11.Gubler, J. G. H., M. Kuster, F. Dutly, F. Bannwart, M. Krause, H. P. Vögelin, G. Garzoli, and M. Altwegg. 1999. Whipple endocarditis without gastrointestinal disease: report of four cases. Ann. Intern. Med. 131:112-116. [DOI] [PubMed] [Google Scholar]
- 12.Hinrikson, H. P., F. Dutly, S. Nair, and M. Altwegg. 1999. Detection of three different types of “Tropheryma whippelii” directly from clinical specimens by sequencing, single-strand conformation polymorphism (SSCP) analysis and type-specific PCR of their 16S-23S ribosomal intergenic spacer region. Int. J. Syst. Bacteriol. 49:1701-1706. [DOI] [PubMed] [Google Scholar]
- 13.Hinrikson, H. P., F. Dutly, and M. Altwegg. 2000. Analysis of the actinobacterial insertion in domain III of the 23S ribosomal RNA gene of uncultured variants of the bacterium associated with Whipple's disease using broad-range and “ Tropheryma whippelii”-specific PCR. Int. J. Syst. Bacteriol. 50:1007-1011. [DOI] [PubMed] [Google Scholar]
- 14.Maiwald, M., H.-J. Ditton, A. von Herbay, F. A. Rainey, and E. Stackebrand. 1996. Reassessment of the phylogenetic position of the bacterium associated with Whipple' s disease and determination of the 16S-23S ribosomal intergenic spacer sequence. Int. J. Syst. Bacteriol. 46:1078-1082. [DOI] [PubMed] [Google Scholar]
- 15.Maiwald, M., A. von Herbay, P. W. Lepp, and D. A. Relman. 2000. Organization, structure, and variability of the rRNA operon of the Whipple's disease bacterium (Tropheryma whippelii). J. Bacteriol. 182:3292-3297. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Mangiapan, G., M. Vokurka, L. Schouls, J. Cadranel, D. Lecossier, J. van Embden, and A. J. Hance. 1996. Sequence capture-PCR improves detection of mycobacterial DNA in clinical specimens. J. Clin. Microbiol. 34:1209-1215. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Misbah, S. A., B. Ozols, A. Franks, and N. Mapstone. 1997. Whipple's disease without malabsorption: new atypical features. QJM 90:765-772. [DOI] [PubMed] [Google Scholar]
- 18.Morgenegg, S., F. Dutly, and M. Altwegg. 2000. Cloning and sequencing of a part of the heat shock protein 65 gene (hsp65) of “Tropheryma whippelii” and its use for the detection of “ T. whippelii” in clinical specimens by PCR. J. Clin. Microbiol. 38:2248-2253. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Müller, C., D. Petermann, C. Stain, H. Riemer, H. Vogelsang, P. Schnider, K. Zeiler, and F. Wrba. 1997. Whipple' disease: comparison of histology with diagnosis based on polymerase chain reaction in four consecutive cases. Gut 40:425-427. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.O'Duffy, J. D., W. L. Griffing, C.-Y. Li, M. F. Abdelmalek, and D. H. Persing. 1999. Whipple arthritis. Direct detection of Tropheryma whippelii in synovial fluid and tissue. Arthritis Rheum. 42:812-817. [DOI] [PubMed] [Google Scholar]
- 21.Raoult, D., M. L. Birg, B. La Scola, P. E. Fournier, M. Enea, H. Lepidi, V. Roux, J.-C. Piette, F. Vandenesch, D. Vital-Durand, and T. J. Marrie. 2000. Cultivation of the bacillus of Whipple's disease. N. Engl. J. Med. 342:620-625. [DOI] [PubMed] [Google Scholar]
- 22.Relman, D. A., T. M. Schmidt, R. P. MacDermott, and S. Falkow. 1992. Identification of the uncultured bacillus of Whipple's disease. N. Engl. J. Med. 327:293-301. [DOI] [PubMed] [Google Scholar]
- 23.Rickman, L. S., W. R. Freeman, W. R. Green, S. Feldman, J. Sullivan, V. Russack, and D. A. Relman. 1995. Brief report: uveitis caused by Tropheryma whippelii (Whipple's bacillus). N. Engl. J. Med. 322:363-366. [DOI] [PubMed] [Google Scholar]
- 24.Ritzler, M., I. Perschil, and M. Altwegg. 1999. Influence of residual uracil-DNA glycosylase activity on the electrophoretic migration of dUTP-containing PCR products. J. Microbiol. Methods 35:73-76. [DOI] [PubMed] [Google Scholar]
- 25.Schoedon, G., D. Goldenberger, R. Forrer, A. Gunz, F. Dutly, M. Höchli, M. Altwegg, and A. Schaffner. 1997. Deactivation of macrophages with interleukin-4 is the key to the isolation of Tropheryma whippelii. J. Infect. Dis. 176:672-677. [DOI] [PubMed] [Google Scholar]
- 26.Street, S., H. D. Donoghue, and G. H. Neild. 1999. Tropheryma whippelii DNA in saliva of healthy people. Lancet 354:1178-1179. [DOI] [PubMed] [Google Scholar]
- 27.Wilson, I. G. 1997. Inhibition and facilitation of nucleic acid amplification. Appl. Environ. Microbiol. 59:1972-1974. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Wilson, K. H., R. Blitchington, R. Frothingham, and J. A. P. Wilson. 1991. Phylogeny of the Whipple's disease-associated bacterium. Lancet 338:474-475. [DOI] [PubMed] [Google Scholar]