Abstract
Background
The rapid advancement of Artificial Intelligence (AI) has led to its widespread application across various domains, showing encouraging outcomes. Many studies have utilized AI to forecast emergency department (ED) disposition, aiming to forecast patient outcomes earlier and to allocate resources better; however, a dearth of comprehensive review literature exists to assess the objective performance standards of these predictive models using quantitative evaluations. This study aims to conduct a meta-analysis to assess the diagnostic accuracy of AI in predicting ED disposition, encompassing admission, critical care, and mortality.
Methods
Multiple databases, including Scopus, Springer, ScienceDirect, PubMed, Wiley, Sage, and Google Scholar, were searched until December 31, 2023, to gather relevant literature. Risk of bias was assessed using the Prediction Model Risk of Bias Assessment Tool. Pooled estimates of sensitivity, specificity, and area under the receiver operating characteristic curve (AUROC) were calculated to evaluate AI’s predictive performance. Sub-group analyses were performed to explore covariates affecting AI predictive model performance.
Results
The study included 88 articles possessed with 117 AI models, among which 39, 45, and 33 models predicted admission, critical care, and mortality, respectively. The reported statistics for sensitivity, specificity, and AUROC represent pooled summary measures derived from the component studies included in this meta-analysis. AI’s summary sensitivity, specificity, and AUROC for predicting admission were 0.81 (95% Confidence Interval [CI] 0.74–0.86), 0.87 (95% CI 0.81–0.91), and 0.87 (95% CI 0.84–0.93), respectively. For critical care, the values were 0.86 (95% CI 0.79–0.91), 0.89 (95% CI 0.83–0.93), and 0.93 (95% CI 0.89–0.95), respectively, and for mortality, they were 0.85 (95% CI 0.80–0.89), 0.94 (95% CI 0.90–0.96), and 0.93 (95% CI 0.89–0.96), respectively. Emergent sample characteristics and AI techniques showed evidence of significant covariates influencing the heterogeneity of AI predictive models for ED disposition.
Conclusions
The meta-analysis indicates promising performance of AI in predicting ED disposition, with certain potential for improvement, especially in sensitivity. Future research could explore advanced AI techniques such as ensemble learning and cross-validation with hyper-parameter tuning to enhance predictive model efficacy.
Trial registration
This systematic review was not registered with PROSPERO or any other similar registry because the review was completed prior to the opportunity for registration, and PROSPERO currently does not accept registrations for reviews that are already completed. We are committed to transparency and have adhered to best practices in systematic review methodology throughout this study.
Supplementary Information
The online version contains supplementary material available at 10.1186/s12911-025-03010-x.
Keywords: Artificial intelligence, Deep learning, Diagnostic test accuracy, Emergency department disposition, Predictive models, Machine learning
Background
The emergency department (ED) of most hospital facilities serve as the frontline of medical care, its importance remains undeniable. Despite its crucial role, predicting the demand for emergency services is challenging, as it is both random and unpredictable. Compounded by finite medical staffing and resources, sudden influxes of patients can overwhelm the ED, making it difficult to meet medical needs promptly. This scenario often leads to ED crowding, where the demand for emergency services exceeds the ED’s capacity to provide timely care [1]. ED over-crowding has a significant negative impact on overall healthcare quality, such as a negative patient experience [2], reduced care quality [3], and unpleasant staff experiences [4]. Despite the adoption of various ED crowding intervention techniques, including technology-based, physical-based, or flow modification methods [5], ED crowding remains a considerable challenge for most EDs worldwide.
The use of artificial intelligence (AI) has been applied across various fields. Advancements promoted by AI have been especially felt in healthcare, for predictive purposes, to yield promising results. One such application is the utilization of machine learning in predicting ED dispositions [6–10], which has shown significant achievements. The logical next step is to systematically and objectively consolidate those study findings in order to provide reference for both medical practice and academia alike. However, existing reviews on the application of machine learning in predicting ED dispositions maintain certain shortcomings. For instance, most studies only analyze the performance of prediction models through systematic review, lacking more objective quantitative analysis through meta-analysis to consolidate the stated performance of those models. Additionally, there is a lack of comprehensive analysis regarding different ED dispositions (e.g., admission, critical care, and mortality). Dispositions in this study are limited to the outcomes of patients’ visits to the ED, specifically focusing on hospital admission, critical care, and mortality. Diagnoses or presenting conditions (e.g., sepsis) are not included in the scope of this analysis.
Therefore, this study aims to systematically evaluate the diagnostic performance of AI models in predicting key ED dispositions—hospital admission, critical care, and mortality—through a comprehensive meta-analysis. Specifically, the study seeks to: (1) Quantify the overall diagnostic accuracy of AI models in predicting ED dispositions, enabling a clearer understanding of their general capabilities, (2) Identify and analyze covariates (e.g., data characteristics, model types, and study settings) that contribute to heterogeneity in AI performance across studies, and (3) Provide actionable insights and practical recommendations for stakeholders—including clinicians, researchers, and administrators—on how AI applications can be better utilized to improve ED workflows and decision-making processes. The research questions of this study include: (1) What is the performance of AI applications in predicting admission, critical care, and mortality?; and, (2) What covariates can account for the heterogeneity between studies? Given the varied application settings of each predictive model, this meta-analysis is intended to present an overall view of AI applications rather than offering tailored recommendations for individual clinical situations. By synthesizing results from diverse contexts, this meta-analysis aims to highlight general trends, identify strengths and limitations, and outline key areas for future research. While this study offers insights into the overall status of AI applications in predicting ED dispositions, specific applications in particular settings would still require further development to meet unique contextual needs.
The primary contributions of this study are as follows: (1) Providing an objective, quantitative evaluation of AI performance in predicting ED dispositions, helping readers understand the general capabilities, advantages, and limitations of current AI applications in this context. (2) Identifying covariates from different data and technical perspectives that may influence AI performance to offer strategies to enhance the performance of AI predictive models. (3) Compiling and synthesizing existing knowledge on AI predictions in ED dispositions, creating a practical resource to guide clinicians, administrators, and researchers considering AI adoption to improve ED workflows.
Related works
To provide a comprehensive evaluation of the application of AI in predicting ED dispositions, this study adopts both a macro and a micro perspective. The macro perspective focuses on synthesizing findings from existing review studies, which offer a high-level understanding of trends, methodologies, and limitations in the field. Complementing this, the micro perspective examines original studies to provide a granular view of AI model performance, including key metrics such as sensitivity, specificity, and area under the receiver operating characterisitc curve. By integrating both perspectives, this study aims to bridge the gap between broad trends and specific evidence, offering a holistic understanding of AI’s role in predicting ED dispositions.
Macro perspective - Review studies
Previous literature has systematically reviewed and/or meta-analyzed the dispositions of ED patients, thereby contributing to a better understanding of the topic. As such, these studies also indicate areas for further research and improvement. For example, Shung et al. [11] conducted a systematic review analyzing machine learning’s use in predicting outcomes of acute gastrointestinal-bleeding patients, finding an area under the curve of approximately 0.84 for mortality, interventions, or re-bleeding prediction. However, the study lacked meta-analysis, limiting comprehensive evaluation, especially regarding ED disposition. Guo et al. [12] reviewed machine-learning applications in predicting heart failure diagnoses, readmissions, and mortality, affirming its effectiveness. Yet, the review summarized results without comprehensive statistical analyses, particularly regarding heart failure patients.
Kareemi et al. [13] reviewed machine-learning’s diagnostic and prognostic applications in ED patients, showing superior performance but lacking meta-analyses for a more objective assessment. Naemi et al. [14] reviewed studies predicting in-hospital mortality among ED patients using vital signs and noting reporting shortcomings. Despite proposing future directions, it lacked meta-analysis to objectively quantify predictive capabilities. Buttia et al. [15] focused on machine-learning predictions of COVID-19 outcomes, highlighting limitations in model generalizability. But, as noted, it didn’t deeply assess machine-learning performance.
Chen et al. [16] reviewed studies predicting ICU transfers among ED patients, demonstrating promising performance but lacking both a broader perspective and meta-analytic techniques. Issaiy et al. [17] reviewed machine-learning predictions for acute appendicitis, emphasizing high accuracy but lacking meta-analysis for comprehensive assessment. Larburu et al. [18] systematically reviewed studies predicting ED patient hospitalizations, noting logistic regression’s common usage but lacking meta-analytical synthesis. Olender et al. [19] reviewed studies predicting mortality among older adults using machine learning, conducting meta-analyses to quantify predictive abilities. While providing valuable insights into mortality prediction, the review’s focus on mortality and lack of specificity regarding in-hospital mortality limits its comprehensive assessment capability. Zhang et al. [20] systematically reviewed and meta-analyzed studies predicting sepsis patients’ mortality using machine learning, demonstrating superior predictive performance compared to existing scoring systems. Despite its comprehensive analysis, the review’s limitation lies in its focus solely on the combination of sepsis and mortality prediction.
Based on the existing review literature, there are notable areas for improvement in synthesizing studies that apply machine learning to predict ED dispositions. To begin with, there is a lack of comprehensive review studies on ED dispositions. Among the ten reviewed studies, only a few focused on specific aspects such as admission, mortality, or critical care, rather than providing a holistic view. Secondly, the meta-analytical approach is under-utilized, with only a minority of the reviewed studies employing them. Meta-analysis has the potential to provide a more objective evaluation of machine-learning predictive models, benefiting both practitioners and academics alike. Detailed information on existing reviews is shown in Table 1.
Table 1.
Emergency department disposition-related review studies
| Source | ED disposition | Database | Analytic strategy | Articles included | Specific disease/condition | Major findings |
|---|---|---|---|---|---|---|
| Shung et al. [11] | Admission and mortality | Embase, Medline, Cochrane, Central, Web of Science, WHO COVID-19 Global Literature on Coronavirus Disease, and Google Scholar | Systematic review | 14 |
Gastrointestinal bleeding |
The median AUC for mortality was 0.84, with AI yielding higher AUCs. Machine learning demonstrated superior performance compared to clinical risk scores for mortality in cases of upper gastrointestinal bleeding. |
| Guo et al. [12] | Mortality | PubmMed | Systematic review | 335 | Heart failure | Machine learning helps identify heart failure patients and assess their risk for readmission and mortality accurately. |
| Kareemi et al. [13] | Admission and mortality |
Medline, Embase, Central, and CINAHL |
Systematic review | 23 | No | Machine learning might surpass standard care in predicting outcomes for emergency department patients in diverse clinical scenarios. |
| Naemi et al. [14] | Mortality | PubMed, Scopus, and Embase | Systematic review | 15 | No | Eight recommendations for future research to enhance the practical implementation of machine learning in various domains. |
| Buttia et al. [15] | ICU admission, intubation, high-flow nasal therapy, extracorporeal membrane oxygenation, and mortality | Embase, Medline, Cochrane Central, Web of Science, WHO COVID-19 Global Literature on Coronavirus Disease, and Google Scholar | Systematic review | 314 | COVID-19 | Several clinical prognostic models for COVID-19, described in the literature, suffer from limited generalizability and applicability due to unresolved statistical and methodological concerns. |
| Chen et al. [16] | ICU admission |
PubMed, Embase, Cochrane Library, and Web of Science |
Systematic review | 10 | No | Machine learning excels in identifying and predicting critically ill patients in emergency department triage. |
| Issaiy et al. [17] | ICU admission | PubMed, Embase, Scopus, and Web of Science | Systematic review | 29 | Acute appendicitis | The artificial neural network exhibited high performance across the majority of cases. |
| Larburu et al. [18] | Admission | Scopus, PubMed, and Google Scholar | Systematic review | 14 | No | Artificial intelligence models improve emergency department care and ease healthcare system burdens. |
| Olender et al. [19] | Mortality | PubMed, Embase, Web of Science, Scopus, and Proquest | Systematic review and meta-analysis | 37 |
Older adults (+ 65) |
Machine learning models demonstrate strong discriminatory power in predicting mortality. |
| Zhang et al. [20] | Mortality | PubMed, Embase, Cochrane Library, and Web of Science | Systematic review and meta-analysis | 50 | Sepsis | Machine learning methods exhibit notably high accuracy in predicting mortality risk among sepsis patients. |
Note: AI = Artificial Intelligence and AUC = Area Under Curve
Micro perspective - Original studies
From a micro perspective, in studies predicting admission disposition, most utilized private datasets, while only a few studies [21–24] developed prediction models using public datasets, such as the National Hospital and Ambulatory Medical Care Survey (NHAMCS) ED data and the Medical Information Mart for Intensive Care IV (MIMIC-IV) ED database. Most studies relied on structured features, with some [25–29] combining structured and unstructured features (e.g., free text), while others [30–32] used only unstructured features. Regarding model validation, the majority employed internal validation, with only a small number of studies [33] using external validation. Studies that applied cross-validation (e.g., K-fold cross-validation) outnumbered those that did not. Additionally, studies using traditional machine learning methods for ED disposition prediction slightly outnumbered those employing deep learning techniques [21–24, 26, 30–33]. A large portion of studies [21, 28, 29, 34–48] adopted ensemble methods for building predictive models.
In studies predicting critical care disposition, research utilizing public datasets [9, 21, 23, 24] remained limited, with NHAMCS and MIMIC-IV being the most commonly used public datasets. These studies predominantly relied on structured features, with only a few [49–57] combining structured and unstructured features, or solely using unstructured features [58]. Similarly, external validation was infrequently employed [51, 52, 59], with most studies relying on internal validation. The number of studies applying cross-validation was greater than the number of studies that did not. Most predictive models were built using traditional machine learning methods, while the use of ensemble methods in this context was less common [54, 57, 60].
For studies predicting mortality, research utilizing public datasets [6, 61] was significantly less prevalent compared to those using private datasets. These studies primarily relied on structured features, with fewer studies incorporating unstructured features, including free text and imaging data [52, 58, 62, 63]. Model construction predominantly relied on internal validation, with fewer studies adopting external validation [52]. Studies employing cross-validation still outnumbered those that did not. Traditional machine learning approaches were more commonly used than deep learning methods [52, 58, 61, 62, 64–67]. However, for mortality prediction, studies utilizing ensemble methods outnumbered those that did not.
From the above analysis, it is evident that studies predicting admission, critical care, and mortality dispositions differ significantly in terms of sample sources, feature structuredness, and algorithm choices, leading to varied performance outcomes. Therefore, conducting a meta-analysis to summarize the overall performance of these predictive models is essential. Furthermore, the differences in sample sources and methodologies may contribute to between-study heterogeneity, making it equally important to identify potential factors causing this heterogeneity.
Methods
This study adheres to the reporting guidelines outlined in the Preferred Reporting Items for Systematic Reviews and Meta-Analyses [68, 69] (see Additional file 1 and Additional file 2). Additionally, the research protocol for this study has the approval of E-Da Hospital (EMRP-109-158).
Search strategy and selection process
This study utilized a combination of keywords to search across seven electronic databases, including Scopus, SpringerLink, ScienceDirect, PubMed, Wiley, Sage, and Google Scholar, until December 31, 2023. The primary focus was on three types of ED dispositions: Admission, critical care, or mortality. ‘Admission’ refers to when patients seek treatment at the ED and then transfer to a general ward: ‘critical care’ involves patients with critical conditions requiring ICU transfer with or without the use of intubation or mechanical ventilation; and, ‘mortality’ refers to patients who expire before leaving the ED. Due to potentially diverse keywords for expressing these three ED dispositions, the study employed the keyword combination ‘emergency department’ AND (‘machine learning’ OR ‘deep learning’ OR ‘artificial intelligence’), followed by manual filtering by the researchers.
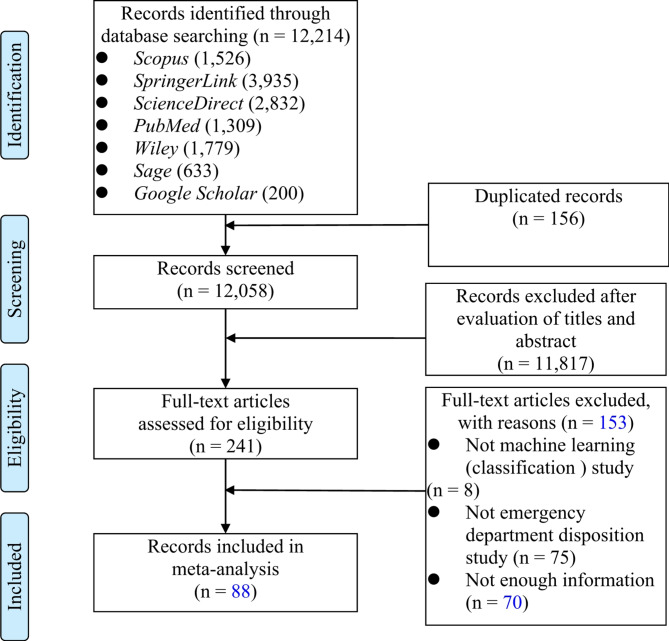
The inclusion criteria consisted of (1) studies focusing on ED dispositions, (2) studies reported in English, and (3) studies utilizing machine learning or deep learning methods. Exclusion criteria included (1) studies not employing machine learning or deep learning, (2) studies lacking sufficient information on outcome measures, and (3) studies not related to the prediction of ED dispositions. Following these criteria, 12,214 potential articles were identified. After excluding 156 duplicate records and the screening of titles and abstracts, 241 full-text articles remained. These were independently reviewed by two researchers, resulting in the exclusion of 153 articles not meeting the inclusion criteria. Ultimately, 88 articles [6–10, 21–67, 70–105]were selected for subsequent meta-analysis. The literature screening process is illustrated in Fig. 1. The studies included in this review are listed in Additional file 3 and Additional file 4.
Fig. 1.
Article selection process
Data extraction
For the included articles, this study extracted the following information: author(s), publication year, sample size, type of ED disposition (admission, critical care, or mortality), data source (private or public dataset), data structure for features (structured, unstructured, or combined), type of unstructured feature (free text or image), age group of samples (adult, mixed, youth, elder, or unclear), type of AI techniques adopted (machine learning or deep learning), whether cross-validation was used, and whether ensemble learning was adopted. Additionally, this study captured the numbers of true/false positives and true/false negatives. If not directly provided, this study performed conversions based on existing data located in the articles. As the same article may develop multiple ED disposition models simultaneously, this study treated them as distinct ED disposition models for purposes of inclusion.
Methodological analysis
This study assessed the risk of bias and applicability of the evidence based on the Prediction model risk of bias assessment tool (PROBAST) [106, 107]. It primarily focuses on four domains: participants, predictors, outcomes, and analysis.
Statistical analysis
This study followed recommendations from prior diagnostic test accuracy literature [108] to calculate the following measures for test accuracy: sensitivity, specificity, area under the receiver operating characteristic curve (AUROC), diagnostic odds ratio (DOR), positive likelihood ratio (+ LR), and negative likelihood ratio (-LR). Additionally, forest plots were utilized to depict the variability among the included literature, along with the hierarchical summary receiver operating characteristic curve (HSROC) with 95% confidence intervals (CI) and 95% prediction intervals. Further, to identify potential factors influencing heterogeneity, meta-regression analysis was conducted, incorporating variables such as type of ED disposition, data source, type of feature, type of unstructured feature, type of sample, type of AI techniques employed, whether cross-validation was undertaken, and whether ensemble learning was adopted. All analyses were conducted using R Statistical Software v4.3.2 [109] with the lime4 v1.1-35.1 [110] and the mada 0.5.11 [111] package. The MetaDTA was utilized to create the HSROC [112, 113].
Results
General study characteristics
As illustrated in Table 2, among the 88 included articles, there were a total of 117 models present (see Supplementary file C and D), with 39, 45, and 33 models predicting admission, critical care, and mortality, respectively. The majority of models sourced their data from private sources (88.89%). Features used to construct predictive models for ED dispositions were predominantly structured data (78.63%), while among models employing unstructured features, approximately 68% and 32% utilized free text and image data, respectively. Notably, free text data were processed using natural language processing (NLP) techniques to extract meaningful features for model development. Most models (94.02%) only adopted internal validation (the same dataset) instead of external validation (a completely independent dataset) to develop predictive models. Most models were based on samples of adult (70.94%), and the majority employed machine learning techniques (71.79%) rather than deep-learning techniques. Approximately 63% of models utilized cross validation during the training process, while about 38% employed ensemble learning.
Table 2.
Characteristics of included studies
| Characteristic | Value | Frequency | % |
|---|---|---|---|
| Disposition | Admission | 39 | 33.33 |
| Critical care | 45 | 38.46 | |
| Mortality | 33 | 28.21 | |
| Data source | Private | 104 | 88.89 |
| Public | 13 | 11.11 | |
| Data structure for feature | Structured | 92 | 78.63 |
| Unstructured | 21 | 17.95 | |
| Combined | 4 | 3.42 | |
| Type of unstructured feature | Free text | 17 | 68.00 |
| Image | 8 | 32.00 | |
| Type of validation | External | 7 | 5.98% |
| Internal | 110 | 94.02% | |
| Age group of sample | Adult | 83 | 70.94 |
| Mixed | 13 | 11.11 | |
| Youth | 10 | 8.55 | |
| Elder | 6 | 5.13 | |
| Unclear | 5 | 4.27 | |
| Type of artificial intelligence techniques adopted | Deep learning | 33 | 28.21 |
| Machine learning | 84 | 71.79 | |
| Cross validation | No | 43 | 36.75% |
| Yes | 74 | 63.25% | |
| Ensemble | No | 45 | 38.46 |
| Yes | 72 | 61.54 |
This study further categorizes machine learning and deep learning approaches. From Table 3, it is evident that Random forest (RF) (19.66%) and eXtreme gradient boosting (XGB) (18.80%) are the most commonly used algorithms, followed by Gradient boosting machine (GBM) (11.97%) and LightGBM (5.13%). In the realm of deep learning, the Deep neural network (DNN) (18.80%) has a higher usage rate than the Convolutional neural network (CNN) (6.84%) and Recurrent neural network (RNN) (2.56%).
Table 4.
Performance of predicting ED dispositions by artificial intelligence
| Metric | Admission | Critical care | Mortality |
|---|---|---|---|
| AUROC | 0.866 (0.836–0.929) | 0.928 (0.893–0.951) | 0.932 (0.894–0.956) |
| Sensitivity | 0.81 (0.74–0.86) | 0.86 (0.79–0.91) | 0.85 (0.80–0.89) |
| Specificity | 0.87 (0.81–0.91) | 0.89 (0.84–0.93) | 0.94 (0.90–0.96) |
| DOR | 17.3 (12.40–23.50) | 44.5 (24.70–74.20) | 74.6 (37.70–133.00) |
| +LR | 4.76 (3.73–6.04) | 7.73 (5.06–11.50) | 12.8 (7.77–20.10) |
| -LR | 0.277 (0.23–0.33) | 0.18 (0.12–0.25) | 0.177 (0.13–0.23) |
Note: AUROC = Area Under Receiver Operating Characteristic curve, CI = Confidence Interval, DOR = Diagnostic Odds Ratio, ED = Emergency Department, +LR = Positive Likelihood Ratio, and –LR = Negative Likelihood Ratio
Table 3.
Type of artificial intelligence techniques adopted
| Technique | Algorithm | Frequency | % |
|---|---|---|---|
| Machine learning | Random forest | 23 | 19.66 |
| eXtreme Gradient Boosting | 22 | 18.80 | |
| Gradient boosting machine | 14 | 11.97 | |
| LightGBM | 6 | 5.13 | |
| Logistic regression | 4 | 3.42 | |
| Support vector machine | 4 | 3.42 | |
| Stacking (or other ensembles) | 3 | 2.56 | |
| CatBoost | 2 | 1.71 | |
| Decision tree | 2 | 1.71 | |
| Neural network | 1 | 0.85 | |
| INC2.5 | 1 | 0.85 | |
| LASSO | 1 | 0.85 | |
| AutoScore | 1 | 0.85 | |
| Deep learning | Deep neural network | 22 | 18.80 |
| Convolutional neural network | 11 | 6.84 | |
| Recurrent neural network | 3 | 2.56 |
Quality assessment
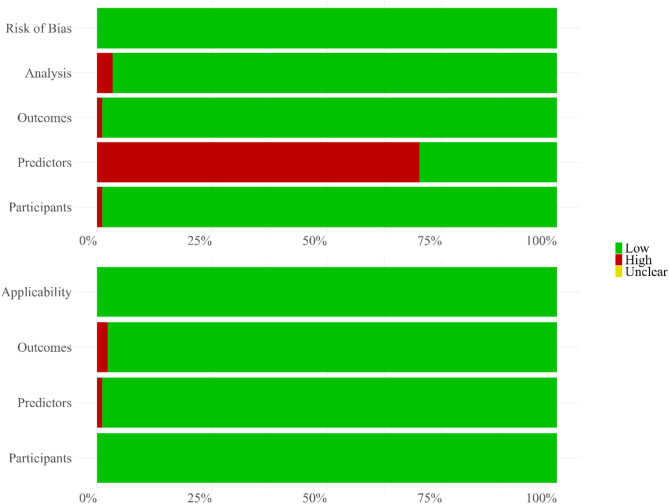
In terms of risk of bias, among the 87 included articles, approximately 70.11% were classified as having a high risk of bias regarding the predictors domain. For the other three domains—participants, outcomes, and analysis—98.85%, 100%, and 97.70% of articles, respectively, were assessed as having a low risk of bias. Overall, 96.55% of articles were assessed as having a low risk of bias, while 3.45% were classified as high risk.
Regarding the risk of applicability, nearly all articles were assessed as having a low risk across the three domains. Specifically, all articles demonstrated low risk concerning participant applicability, while 98.85% showed low risk for predictors and outcomes. Overall, 100% of the articles were assessed as having a low risk of applicability. This assessment of risk of bias and applicability based on the PROBAST tool is summarized in Fig. 2.
Fig. 2.
Quality assessment by PROBAST
Diagnostic accuracy
Among the three major types of ED disposition prediction models, those forecasting mortality achieved the highest area under the receiver operating characteristic curve (AUROC), followed by models predicting critical care, with admission prediction models exhibiting the lowest performance (see Table 4). The reported statistics for sensitivity, specificity, and AUROC represent pooled summary measures derived from the component studies included in this meta-analysis. The pooled summary AUROC for predicting admission, critical care, and mortality were 0.866 (95% CI 0.836–0.929), 0.928 (95% CI 0.893–0.951), and 0.932 (95% CI 0.894–0.956), respectively. In terms of sensitivity, admission prediction models showed the lowest sensitivity at 0.81 (95% CI 0.74–0.86), followed by critical care models at 0.86 (95% CI 0.79–0.91), and mortality models at 0.85 (95% CI 0.80–0.89). Regarding specificity, admission models exhibited the lowest specificity at 0.87 (95% CI 0.81–0.91), followed by critical care models at 0.89 (95% CI 0.84–0.93), and mortality models at 0.94 (95% CI 0.90–0.96). These statistics are primarily based on models utilizing internal validation, as only 5.98% of the included models performed external validation. In terms of sensitivity, critical care prediction models performed the best, closely followed by mortality prediction models, while admission prediction models showed the lowest sensitivity. Regarding specificity, mortality prediction models demonstrated the highest specificity, followed by critical care prediction models, with admission prediction models exhibiting the lowest specificity.
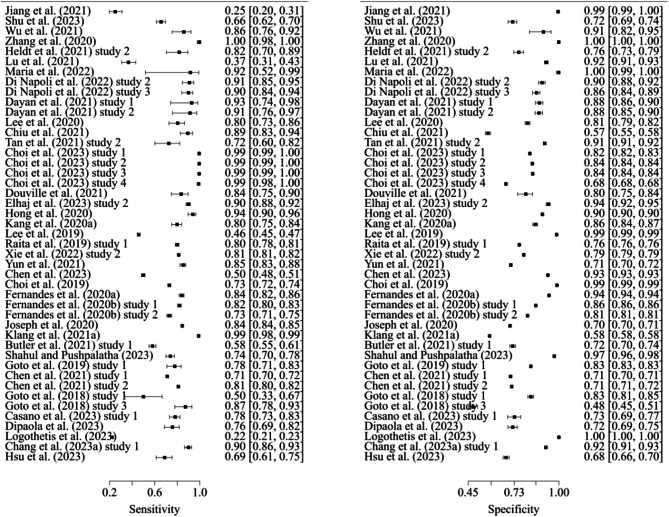
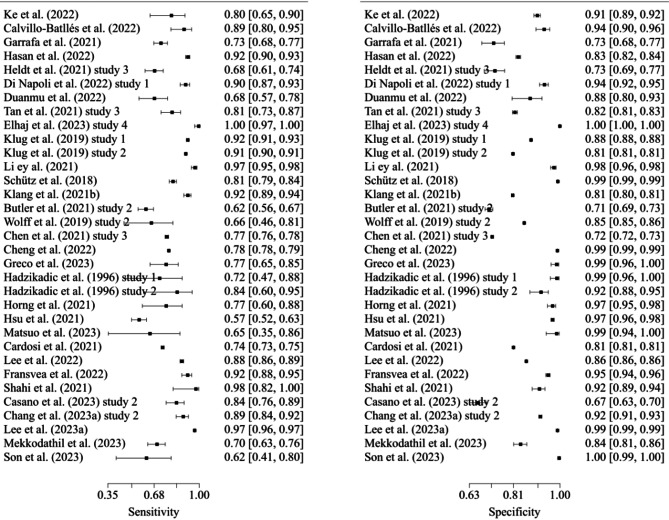
Analysis of the DOR revealed that models predicting mortality exhibited the highest discriminatory performance [114], while models predicting admission had the lowest DOR. Considering that the + LR, models predicting mortality were better at identifying true mortality cases due to their highest + LR, whereas models predicting critical care had a lower -LR, indicating their better ability to identify non-critical care patients [115]. Forest plots of sensitivity and specificity for models predicting admission, critical care, and mortality are illustrated in Figs. 3, 4 and 5.
Fig. 3.
Sensitivity and specificity of models predicting admission (n = 39)
Fig. 4.
Sensitivity and specificity of models predicting critical care (n = 45)
Fig. 5.
Sensitivity and specificity of models predicting mortality (n = 33)
Plausible covariates to explain between-study heterogeneity
Overall, machine learning models for predicting ED disposition demonstrate a sensitivity of approximately 0.84 (95% CI 0.80–0.87) and a specificity of around 0.90 (95% CI 0.87–0.92) (see Table 5), indicating their higher ability to correctly identify negative cases. When differentiating ED disposition into the categories of admission, critical care, and mortality, using admission as the reference category for comparison (as depicted in Table 5), the sensitivity of admission prediction models (0.81, 95% CI 0.74–0.86) is slightly lower than that of critical care and mortality prediction models (0.86, 95% CI 0.79–0.91 and 0.85, 95% CI 0.80–0.89, respectively), although these differences are not statistically significant. Similarly, the specificity of admission prediction models (0.87, 95% CI 0.81–0.91) is also slightly lower than that of critical care and mortality prediction models (0.89, 95% CI 0.84–0.93 and 0.94, 95% CI 0.90–0.96, respectively), with a statistically significant difference observed between admission prediction models and mortality prediction models’ specificity (0.87 vs. 0.94, p = 0.027).
Table 5.
Summary estimates for sensitivity and specificity
| Covariate | Metric | n | Estimate | 95% | C.I. | p value |
|---|---|---|---|---|---|---|
| Overall | Sens | 117 | 0.84 | 0.80 | 0.87 | |
| Overall | Specs | 117 | 0.90 | 0.87 | 0.92 | |
| Admission | Sens | 39 | 0.81 | 0.74 | 0.86 | [Reference] |
| Admission | Specs | 39 | 0.87 | 0.81 | 0.91 | [Reference] |
| Critical care | Sens | 45 | 0.86 | 0.79 | 0.91 | 0.289 |
| Critical care | Specs | 45 | 0.89 | 0.84 | 0.93 | 0.503 |
| Mortality | Sens | 33 | 0.85 | 0.80 | 0.89 | 0.193 |
| Mortality | Specs | 33 | 0.94 | 0.90 | 0.96 | 0.027 |
Note: C.I. = Confidence Interval, Sens = Sensitivity, and Specs = Specificity
Plausible covariates for admission predictive models
This study compares the predictive abilities of different ED dispositions (admission, critical care, and mortality) to assess whether or not they are influenced by variables such as data characteristics, sample properties, or machine-learning methods.
Firstly, regarding the prediction of admission models, Table 6 shows that models using public datasets exhibit higher sensitivity (0.94) and specificity (0.90) when compared to those using private datasets (sensitivity = 0.80, specificity = 0.86), although the differences are not statistically significant. In terms of data structuring, both sensitivity (0.84) and specificity (0.88) of structured data models surpass those of unstructured data models (sensitivity = 0.72, specificity = 0.80) and those models combining unstructured and structured data (sensitivity = 0.74, specificity = 0.82). However, the differences among the three types of data are not statistically significant. Notably, among unstructured data, while some studies utilize image and free-text data, only eight models use free text as a feature for predicting admission, with no models using image data, thus this variable was not included for purposes of analysis. Among the eight models using free text data, the pooled sensitivity was 0.73 (95% CI 0.65–0.80), and the pooled specificity was 0.81 (95% CI 0.73–0.88), indicating moderate diagnostic accuracy.
Table 6.
Summary estimates for sensitivity and specificity of admission studies
| Characteristic | Covariate | Metric | n | Estimate | 95% C.I. | p value | |
|---|---|---|---|---|---|---|---|
| Data | Public dataset | Sens | 6 | 0.94 | 0.57 | 0.99 | 0.284 |
| Public dataset | Specs | 6 | 0.90 | 0.70 | 0.97 | 0.581 | |
| Private dataset | Sens | 33 | 0.79 | 0.73 | 0.84 | [Reference] | |
| Private dataset | Specs | 33 | 0.86 | 0.79 | 0.91 | [Reference] | |
| Unstructured | Sens | 3 | 0.72 | 0.70 | 0.74 | 0.387 | |
| Unstructured | Specs | 3 | 0.80 | 0.74 | 0.86 | 0.521 | |
| Combined | Sens | 5 | 0.74 | 0.61 | 0.84 | 0.362 | |
| Combined | Specs | 5 | 0.82 | 0.68 | 0.91 | 0.510 | |
| Structured | Sens | 31 | 0.83 | 0.75 | 0.89 | [Reference] | |
| Structured | Specs | 31 | 0.88 | 0.81 | 0.93 | [Reference] | |
| Sample | Adult | Sens | 21 | 0.81 | 0.72 | 0.87 | 0.308 |
| Adult | Specs | 21 | 0.87 | 0.76 | 0.93 | 0.448 | |
| Youth | Sens | 6 | 0.82 | 0.69 | 0.90 | 0.218 | |
| Youth | Specs | 6 | 0.80 | 0.67 | 0.88 | 0.064 | |
| Elder | Sens | 3 | 0.76 | 0.74 | 0.79 | 0.541 | |
| Elder | Specs | 3 | 0.75 | 0.70 | 0.79 | 0.027 | |
| Mixed | Sens | 5 | 0.70 | 0.55 | 0.82 | [Reference] | |
| Mixed | Specs | 5 | 0.92 | 0.84 | 0.96 | [Reference] | |
| Artificial intelligence technique | Machine learning | Sens | 28 | 0.80 | 0.72 | 0.86 | 0.535 |
| Machine learning | Specs | 28 | 0.87 | 0.80 | 0.92 | 0.770 | |
| Deep learning | Sens | 11 | 0.85 | 0.71 | 0.93 | [Reference] | |
| Deep learning | Specs | 11 | 0.85 | 0.73 | 0.92 | [Reference] | |
| Random forest | Sens | 10 | 0.82 | 0.61 | 0.93 | 0.147 | |
| Random forest | Specs | 10 | 0.92 | 0.77 | 0.98 | 0.720 | |
| eXtreme Gradient Boosting | Sens | 8 | 0.77 | 0.69 | 0.84 | 0.022 | |
| eXtreme Gradient Boosting | Specs | 8 | 0.86 | 0.77 | 0.91 | 0.788 | |
| Deep neural network | Sens | 7 | 0.70 | 0.65 | 0.75 | 0.002 | |
| Deep neural network | Specs | 7 | 0.78 | 0.74 | 0.82 | 0.360 | |
| Recurrent neural network | Sens | 2 | 0.97 | 0.31 | 1.00 | 0.604 | |
| Recurrent neural network | Specs | 2 | 0.93 | 0.58 | 0.99 | 0.899 | |
| Convolutional neural network | Sens | 2 | 0.99 | 0.15 | 1.00 | [Reference] | |
| Convolutional neural network | Specs | 2 | 0.99 | 0.02 | 1.00 | [Reference] | |
| Ensemble | Sens | 26 | 0.78 | 0.71 | 0.84 | 0.301 | |
| Ensemble | Specs | 26 | 0.86 | 0.78 | 0.91 | 0.713 | |
| No-ensemble | Sens | 13 | 0.86 | 0.73 | 0.93 | [Reference] | |
| No-ensemble | Specs | 13 | 0.88 | 0.76 | 0.95 | [Reference] | |
| Cross validation | Sens | 26 | 0.84 | 0.76 | 0.90 | 0.170 | |
| Cross validation | Specs | 26 | 0.86 | 0.76 | 0.92 | 0.541 | |
| No cross validation | Sens | 13 | 0.74 | 0.65 | 0.82 | [Reference] | |
| No cross validation | Specs | 13 | 0.88 | 0.82 | 0.93 | [Reference] | |
Note: C.I. = Confidence Interval, Sens = Sensitivity, and Specs = Specificity
Regarding sample properties, models using mixed samples (all age groups) demonstrate lower sensitivity compared to models using adult, youth, and elder samples, yet the specificity of models using mixed samples is higher than those using the other three types of samples. Sensitivity among the four different samples does not reach statistical significance, but the specificity of models using mixed samples (0.92 vs. 0.75, p = 0.027) is significantly higher than that of models using elderly samples.
In terms of machine-learning methods, models utilizing deep learning exhibit higher sensitivity (0.86) when compared to those generated using traditional machine-learning methods (sensitivity = 0.80), but the difference is not statistically significant (p = 0.541). The specificity of models built using traditional machine learning (0.87) is slightly higher than that of models generated using deep learning (0.86), also not statistically significant (p = 0.868). To further compare the performance of different machine-learning algorithms, models utilizing CNN show both higher sensitivity and specificity than those using RF, XGB, DNN, and RNN. Among these, the differences in sensitivity compared with XGB (p = 0.022) and DNN (p = 0.002) are statistically significant. Models built using ensemble learning demonstrate lower sensitivity (0.78) and specificity (0.86) when compared to models not using ensemble learning (sensitivity = 0.86, specificity = 0.88), but the differences are not statistically significant (p = 0.301 and 0.713). Additionally, models not using cross-validation exhibit a lower level of sensitivity (0.74) and a higher level of specificity (0.88) when compared to models using cross-validation (sensitivity = 0.84, specificity = 0.86), yet both sensitivity and specificity do not reach statistical significance (p = 0.170 and 0.541).
Plausible covariates for critical care predictive models
In predicting critical care models, the sensitivity (0.87) and specificity (0.90) of models using private datasets are higher than those using public datasets (sensitivity = 0.76 and specificity = 0.73), but the differences do not reach statistical significance (p = 0.316 and 0.103). As there is only one model in the critical care prediction category that solely uses unstructured data, comparisons are made only between models using structured and combined data types. From Table 7, it is evident that the sensitivity (0.86) and specificity (0.90) of models using structured data are higher than or equal to those of models using combined data (sensitivity = 0.86, specificity = 0.87), but none of the differences are statistically significant (p = 0.865 and 0.626). In these prediction models, some models combine image and free-text data. This study further compares the impact of these two formats on prediction models, revealing that models combining image have higher sensitivity (0.87) compared to those combining free text (0.83), while models combining image have lower sensitivity (0.86) as compared to those combining free text (0.87), but neither difference reaches statistical significance (p = 0.530 and 0.861).
Table 7.
Summary estimates for sensitivity and specificity of critical care studies
| Characteristic | Covariate | Metric | n | Estimate | 95% | C.I. | p value |
|---|---|---|---|---|---|---|---|
| Data | Public dataset | Sens | 5 | 0.76 | 0.64 | 0.85 | 0.316 |
| Public dataset | Specs | 5 | 0.73 | 0.62 | 0.82 | 0.103 | |
| Private dataset | Sens | 40 | 0.87 | 0.79 | 0.92 | [Reference] | |
| Private dataset | Specs | 40 | 0.90 | 0.85 | 0.94 | [Reference] | |
| Combined | Sens | 12 | 0.86 | 0.77 | 0.92 | 0.865 | |
| Combined | Specs | 12 | 0.87 | 0.80 | 0.92 | 0.626 | |
| Structured | Sens | 32 | 0.86 | 0.77 | 0.92 | [Reference] | |
| Structured | Specs | 32 | 0.90 | 0.83 | 0.95 | [Reference] | |
| Image | Sens | 5 | 0.87 | 0.76 | 0.94 | 0.530 | |
| Image | Specs | 5 | 0.86 | 0.80 | 0.90 | 0.861 | |
| Free text | Sens | 8 | 0.83 | 0.67 | 0.92 | [Reference] | |
| Free text | Specs | 8 | 0.87 | 0.74 | 0.94 | [Reference] | |
| Sample | Adult | Sens | 38 | 0.86 | 0.78 | 0.91 | 0.664 |
| Adult | Specs | 38 | 0.90 | 0.83 | 0.94 | 0.616 | |
| Youth | Sens | 2 | 0.85 | 0.73 | 0.92 | 0.762 | |
| Youth | Specs | 2 | 0.72 | 0.51 | 0.86 | 0.204 | |
| Mixed | Sens | 3 | 0.90 | 0.74 | 0.96 | [Reference] | |
| Mixed | Specs | 3 | 0.84 | 0.75 | 0.90 | [Reference] | |
| Artificial intelligence technique | Machine learning | Sens | 31 | 0.88 | 0.80 | 0.93 | 0.205 |
| Machine learning | Specs | 31 | 0.91 | 0.84 | 0.95 | 0.171 | |
| Deep learning | Sens | 14 | 0.78 | 0.69 | 0.85 | [Reference] | |
| Deep learning | Specs | 14 | 0.83 | 0.78 | 0.87 | [Reference] | |
| Random forest | Sens | 9 | 0.91 | 0.77 | 0.97 | 0.465 | |
| Random forest | Specs | 9 | 0.96 | 0.83 | 0.99 | 0.297 | |
| eXtreme gradient boosting | Sens | 11 | 0.95 | 0.85 | 0.99 | 0.302 | |
| eXtreme gradient boosting | Specs | 11 | 0.84 | 0.71 | 0.92 | 0.965 | |
| LightGBM | Sens | 3 | 0.69 | 0.29 | 0.92 | 0.439 | |
| LightGBM | Specs | 3 | 0.98 | 0.59 | 1.00 | 0.259 | |
| Logistic regression | Sens | 2 | 0.78 | 0.71 | 0.84 | 0.673 | |
| Logistic regression | Specs | 2 | 0.84 | 0.80 | 0.87 | 0.922 | |
| Deep neural network | Sens | 10 | 0.75 | 0.64 | 0.83 | 0.442 | |
| Deep neural network | Specs | 10 | 0.83 | 0.77 | 0.88 | 0.932 | |
| Convolutional neural network | Sens | 3 | 0.84 | 0.63 | 0.94 | [Reference] | |
| Convolutional neural network | Specs | 3 | 0.84 | 0.74 | 0.90 | [Reference] | |
| Ensemble | Sens | 25 | 0.91 | 0.82 | 0.95 | 0.032 | |
| Ensemble | Specs | 25 | 0.91 | 0.82 | 0.96 | 0.339 | |
| No ensemble | Sens | 20 | 0.76 | 0.69 | 0.82 | [Reference] | |
| No ensemble | Specs | 20 | 0.86 | 0.81 | 0.90 | [Reference] | |
| Cross validation | Sens | 27 | 0.86 | 0.76 | 0.92 | 0.952 | |
| Cross validation | Specs | 27 | 0.87 | 0.79 | 0.92 | 0.288 | |
| No cross validation | Sens | 18 | 0.87 | 0.75 | 0.93 | [Reference] | |
| No cross validation | Specs | 18 | 0.92 | 0.83 | 0.96 | [Reference] |
Note: C.I. = Confidence Interval, Sens = Sensitivity, and Specs = Specificity
Regarding sample properties, since there is only one data point for unclear and the elderly in the models predicting critical care, they were not included in the analysis. Only adult, youth, and mixed samples were compared. From Table 7, it is observed that the sensitivity (0.90) of models using mixed samples is higher than those using adult (0.86) and youth (0.85), but the differences are not statistically significant (p = 0.664 and 0.762). Furthermore, the specificity (0.84) of models using mixed samples is lower than that of models using adult (0.90) but higher than that of models using youth (0.72), yet none of the differences are statistically significant (p = 0.616 and 0.204).
Regarding machine-learning methods, the sensitivity (0.88) and specificity (0.91) of models using traditional machine learning are higher than those using deep learning (sensitivity = 0.78 and specificity = 0.83), but the differences are not statistically significant (p = 0.205 and 0.171). When comparing specific machine-learning algorithms, CNN algorithm achieves both a sensitivity and specificity of 0.84. Compared to RF models, which have a higher sensitivity (0.91) and specificity (0.96), CNN models perform slightly lower on both metrics, though these differences are not statistically significant. Similarly, CNN models outperform LightGBM and LR in sensitivity (LightGBM: 0.69, LR: 0.78) but are on par with LR in specificity (0.84) and below LightGBM (0.98), without statistically significant differences. When compared to DNN, CNN models achieve higher sensitivity (DNN: 0.75) but perform similarly in specificity (DNN: 0.83), with no statistically significant differences. Overall, CNN models show a balanced performance in sensitivity and specificity compared to other algorithms. Concerning the use of ensemble learning, the sensitivity (0.91) and specificity (0.91) of models using ensemble learning are higher than those not using it (sensitivity = 0.69 and specificity = 0.81), but only sensitivity reaches statistical significance (p = 0.032). Lastly, the sensitivity (0.86) of models using cross-validation is slightly lower than those not using it (0.87), and the specificity (0.87) of models using cross-validation is also lower compared to models not using it (0.92). Neither difference reaches statistical significance (p = 0.952 and 0.288, respectively).
Plausible covariates for mortality predictive models
In models predicting mortality, both sensitivity (0.90) and specificity (0.95) are higher for models using public datasets than those using private datasets (sensitivity = 0.85, specificity = 0.94), but none of the differences are statistically significant (see Table 8). Regarding data structuring, since no models solely use unstructured data, comparisons were made only between models using structured and combined data. From Table 8, it is observed that both sensitivity (0.86) and specificity (0.95) of models using structured data are higher than those using combined data (sensitivity = 0.82, specificity = 0.85), but none of the differences are statistically significant (p = 0.591, 0.189). Although some models using combined data incorporate image, only one model utilizes free text, achieving a sensitivity of 0.92 (95% CI: 0.89–0.94) and a specificity of 0.81 (95% CI: 0.80–0.81). Therefore, a comparison between these two types of unstructured data was not conducted.
Table 8.
Summary estimates for sensitivity and specificity of mortality studies
| Characteristic | Covariate | Metric | n | Estimate | 95% | C.I. | p value |
|---|---|---|---|---|---|---|---|
| Data | Public dataset | Sens | 2 | 0.90 | 0.64 | 0.98 | 0.446 |
| Public dataset | Specs | 2 | 0.95 | 0.70 | 0.99 | 0.799 | |
| Private dataset | Sens | 31 | 0.85 | 0.80 | 0.88 | [Reference] | |
| Private dataset | Specs | 31 | 0.94 | 0.90 | 0.96 | [Reference] | |
| Combined | Sens | 4 | 0.82 | 0.66 | 0.91 | 0.591 | |
| Combined | Specs | 4 | 0.85 | 0.74 | 0.92 | 0.189 | |
| Structured | Sens | 29 | 0.86 | 0.80 | 0.90 | [Reference] | |
| Structured | Specs | 29 | 0.95 | 0.91 | 0.97 | [Reference] | |
| Sample | Adults | Sens | 24 | 0.85 | 0.79 | 0.89 | 0.853 |
| Adults | Specs | 24 | 0.94 | 0.89 | 0.97 | 0.759 | |
| Youths | Sens | 2 | 0.99 | 0.00 | 1.00 | 0.663 | |
| Youths | Specs | 2 | 0.89 | 0.83 | 0.92 | 0.458 | |
| Elders | Sens | 2 | 0.88 | 0.77 | 0.94 | 0.689 | |
| Elders | Specs | 2 | 0.90 | 0.77 | 0.96 | 0.540 | |
| Mixed | Sens | 5 | 0.84 | 0.67 | 0.93 | [Reference] | |
| Mixed | Specs | 5 | 0.96 | 0.82 | 0.99 | [Reference] | |
| Artificial intelligence technique | Machine learning | Sens | 25 | 0.86 | 0.80 | 0.90 | 0.709 |
| Machine learning | Specs | 25 | 0.95 | 0.90 | 0.97 | 0.442 | |
| Deep learning | Sens | 8 | 0.83 | 0.74 | 0.90 | [Reference] | |
| Deep learning | Specs | 8 | 0.91 | 0.84 | 0.96 | [Reference] | |
| Random forest | Sens | 4 | 0.91 | 0.56 | 0.99 | 0.516 | |
| Random forest | Specs | 4 | 1.00 | 0.04 | 1.00 | 0.476 | |
| eXtreme Gradient boosting | Sens | 3 | 0.73 | 0.61 | 0.82 | 0.562 | |
| eXtreme Gradient boosting | Specs | 3 | 0.95 | 0.77 | 0.99 | 0.839 | |
| LightGBM | Sens | 3 | 0.91 | 0.78 | 0.96 | 0.205 | |
| LightGBM | Specs | 3 | 0.92 | 0.75 | 0.98 | 0.748 | |
| Logistic regression | Sens | 2 | 0.80 | 0.73 | 0.86 | 0.938 | |
| Logistic regression | Specs | 2 | 0.95 | 0.68 | 0.99 | 0.866 | |
| Deep neural network | Sens | 5 | 0.86 | 0.76 | 0.92 | 0.418 | |
| Deep neural network | Specs | 5 | 0.89 | 0.83 | 0.93 | 0.436 | |
| Convolutional neural network | Sens | 3 | 0.79 | 0.63 | 0.89 | [Reference] | |
| Convolutional neural network | Specs | 3 | 0.94 | 0.74 | 0.99 | [Reference] | |
| Ensemble | Sens | 21 | 0.88 | 0.82 | 0.92 | 0.095 | |
| Ensemble | Specs | 21 | 0.95 | 0.89 | 0.98 | 0.620 | |
| No ensemble | Sens | 12 | 0.79 | 0.71 | 0.86 | [Reference] | |
| No ensemble | Specs | 12 | 0.93 | 0.87 | 0.96 | [Reference] | |
| Cross validation | Sens | 21 | 0.85 | 0.78 | 0.90 | 0.926 | |
| Cross validation | Specs | 21 | 0.96 | 0.92 | 0.98 | 0.032 | |
| No cross validation | Sens | 12 | 0.85 | 0.78 | 0.90 | [Reference] | |
| No cross validation | Specs | 12 | 0.87 | 0.81 | 0.91 | [Reference] |
Note: C.I. = Confidence Interval, Sens = Sensitivity, and Specs = Specificity
Regarding sample properties, as there are no models classified as ‘unclear,’ comparisons were made among models with samples classified as mixed, adult, youth, and elder. The results show that sensitivity (0.84) of models using mixed samples is lower than those using the other three types of samples (0.85 for adult, 0.99 for youth, and 0.88 for elder), but none of the differences are statistically significant. However, specificity (0.96) of models using mixed samples is higher than those using the other three types of samples (0.94 for adult, 0.89 for youth, and 0.90 for elder), yet none of the differences are statistically significant.
Regarding machine-learning methods, both sensitivity (0.86) and specificity (0.95) of models using machine learning are higher than those using deep learning (sensitivity = 0.83, specificity = 0.91), but neither difference reaches statistical significance (p = 0.709 and 0.442). To further compare the performance of different algorithms, models using the RF and LR algorithms have both higher sensitivity and specificity than those using CNN, though these differences are not statistically significant. Models employing LightGBM and DNN have higher sensitivity but lower specificity compared to CNN, with none of these differences reaching statistical significance. Additionally, models using XGB exhibit lower sensitivity than CNN, but their specificity is higher, also without statistical significance. Models using ensemble methods have higher sensitivity (0.88) and specificity (0.95) than those not using ensemble methods (sensitivity = 0.79 and specificity = 0.93), but none of the differences reach statistical significance (p = 0.095 and 0.620).
Regarding the use of cross-validation in prediction models, sensitivity is the same for models with and without cross-validation (0.85). However, models using cross-validation have significantly higher specificity compared to those not using it (0.96 vs. 0.87, p = 0.032). The summary sensitivity and specificity performance of mortality prediction models are presented in Table 8.
Summarization of plausible covariates for three predictive models
Summarizing the performance of the three disposition prediction models (see Table 9), First off, regarding data sources, models utilizing public data sources perform better in predicting admission and mortality when compared to those using private data sources, but the opposite trend is observed for predicting critical care. Secondly, concerning data structuring, models using structured data outperform those using both structured and unstructured data in predicting admission, critical care, and mortality. In admission-prediction models, those solely using unstructured data exhibit the poorest performance. As for sample properties, no distinct pattern emerges favoring any particular sample combination among the four different types.
Table 9.
Summarization of the performance of predictive models for three emergency department dispositions
| Disposition | Metric | Characteristic | ||||||
|---|---|---|---|---|---|---|---|---|
| Data | Sample | Artificial-intelligence technique | ||||||
| Source | Feature Type | Group | Approach | Top algorithm | Ensemble | Cross validation | ||
| Admission | Sens | Public > Private | S > B > U | A = Y > E > M | DL > ML | CNN | No ensemble > Ensemble | No CV > CV |
| Spec | Public > Private | S > B > U | M > A > Y > E | ML > DL | CNN | No ensemble > Ensemble | No CV > CV | |
| Critical care | Sens | Private > Public | S > B | M > A > Y | ML > DL | XGB | Ensemble > No ensemble | CV > No CV |
| Spec | Private > Public | S > B | A > M > Y | ML > DL | LightGBM | Ensemble > No ensemble | No CV > CV | |
| Mortality | Sens | Public > Private | S > B | Y > E > A > M | ML > DL | RF/LightGBM | Ensemble > No ensemble | CV = No CV |
| Spec | Public > Private | S > B | A > M > Y > E | ML > DL | RF | Ensemble > No ensemble | CV > No CV | |
Notes:
1. Metric: Sens = Sensitivity and Spec = Specificity
2. Source: Public = public data source and Private = private data source
3. Type of feature: S = structured data, B = structured and unstructured data, and U = unstructured data
4. Sample: A = adults, Y = youths, E = elders, and M = mixed samples
5. Approach: ML = machine learning and DL = deep learning
6. CNN = Convolutional neural network, XGB = eXtreme gradient boosting, and RF = Random forest
7. Cross-validation: CV = cross-validation
In terms of machine learning methods, except for admission-prediction models where sensitivity is better with deep learning, both sensitivity and specificity in the other two prediction models favor machine learning. To further compare the performance of different machine-learning algorithms, CNN demonstrates superior sensitivity and specificity for admission prediction compared to other algorithms. For critical care prediction, XGB shows the highest sensitivity, while LightGBM excels in specificity. For mortality prediction, RF and LightGBM yield the best sensitivity, with RF also showing the highest specificity. However, when employing ensemble learning, the sensitivity and specificity of admission-prediction models are both lower as compared to models not using ensemble learning. For critical care and mortality-prediction models, the sensitivity and specificity of models without ensemble learning are higher than those with ensemble learning. Additionally, the use of cross-validation does not consistently guarantee better model performance; while admission-prediction models without cross-validation exhibit higher sensitivity and specificity, the sensitivity and specificity of critical care- or mortality-prediction models vary.
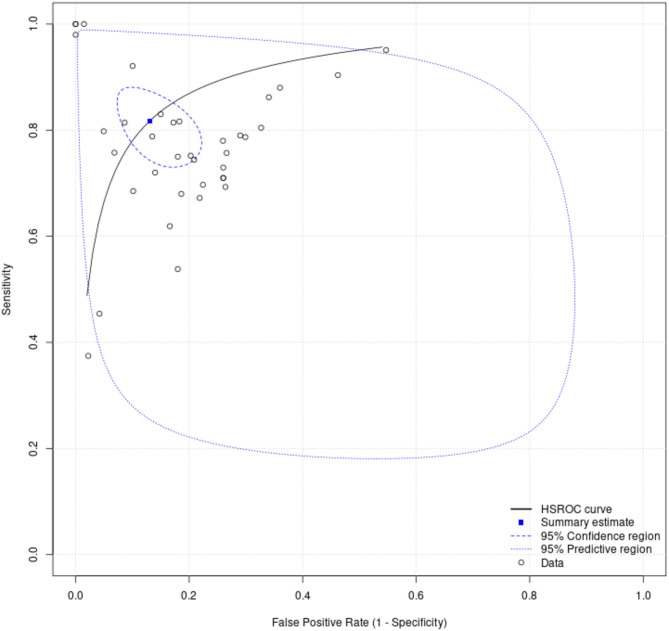
Finally, this study employed HSROC to evaluate model performance, with the HSROC curves for admission, critical care, and mortality depicted in Figs. 6 and 7, and 8, respectively. It is evident from these figures that the mortality-prediction model exhibits higher precision compared to the admission and critical care prediction models. This is indicated by the smaller 95% prediction-interval region and 95% confidence region for the mortality-prediction model, as observed in Figs. 6, 7 and 8.
Fig. 6.
Summary receiver operating-characteristic curve for models predicting admission
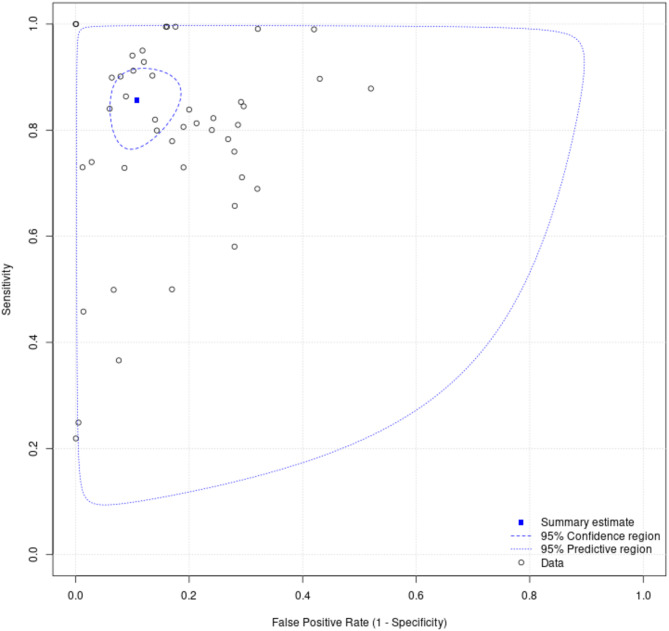
Fig. 7.
Summary receiver operating-characteristic curve for models predicting critical care
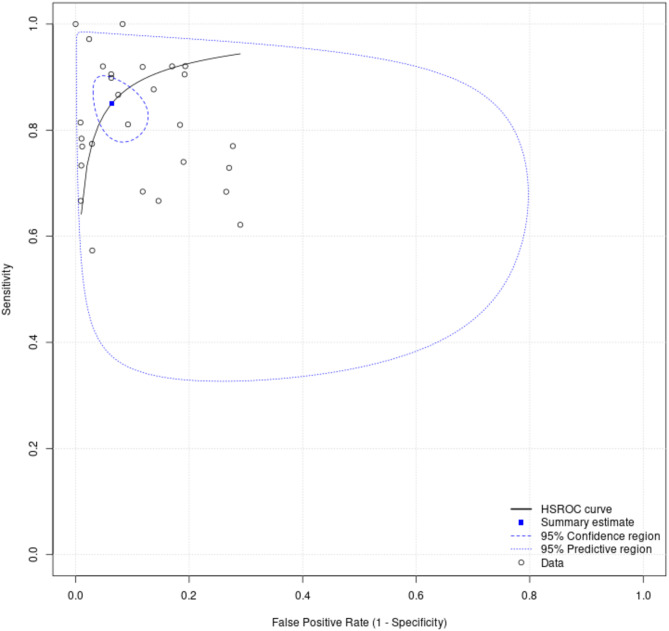
Fig. 8.
Summary receiver operating-characteristic curve for models predicting mortality
Discussion
Based on the meta-analysis of 117 models extracted from 87 articles included in this study, the overall sensitivity and specificity for predicting ED disposition patterns were determined to be 0.84 and 0.90, respectively. These results indicate that the utilization of machine learning in predicting the discharge disposition of ED patients shows acceptable predictive capabilities. This capability allows for the early acquisition of patient disposition information, which can greatly aid in the effective allocation of medical personnel and resources within modern healthcare institutions.
Type of ED dispositions predicted
Upon further examination, among the 117 predictive models, 39 are focused on admission, 45 on critical care, and 33 on mortality. The meta-analysis reveals that mortality-prediction models exhibit the highest AUROC, followed by critical-care prediction models, with admission prediction models demonstrating the lowest performance. Sensitivity analysis indicates that critical care-prediction models have the highest sensitivity, followed by mortality-prediction models, while admission-prediction models have the lowest. Similarly, regarding specificity, mortality-prediction models show the highest, followed by critical care-prediction models, with admission-prediction models again displaying the lowest specificity.
Notably, when admission prediction serves as the reference category, sensitivity and specificity among these three types of prediction models generally do not exhibit statistically significant differences, except for the specificity of mortality-prediction models, which is significantly higher than that of admission-prediction models. Additionally, the specificity of these prediction models tends to outweigh sensitivity, suggesting a stronger ability to correctly identify true negatives but potentially missing some true positives. Future research may necessitate an iterative refinement process to enhance the sensitivity of ED disposition models.
Clarification of research purpose and scope
While this meta-analysis provides a quantitative overview of AI performance in predicting ED patient dispositions, it is important to recognize the heterogeneity among the predictive models included. These models vary in terms of the disposition types predicted, data used, machine learning methods, and patient conditions, which limits the generalizability of our meta-analysis results to specific clinical situations. The primary goal of this meta-analysis is to offer insights into general trends, strengths, and challenges in AI applications for ED disposition prediction, rather than to provide tailored recommendations for individual contexts. Additional research and development tailored to the unique demands of each clinical setting may therefore be necessary.
Public or private data source
In terms of the data utilized, the majority of the data are proprietary rather than publicly available. However, the analysis results show that when publicly available data are used for predicting admission and mortality, both sensitivity and specificity are higher compared to predictive models using private data. Conversely, when publicly available data are used for predicting critical care, sensitivity and specificity are lower than those of private datasets. The meta-analysis results of this study indicate an improvement in the predictive ability of publicly available data for predicting admission and mortality. In the realm of machine learning for skin image recognition, Tschandl et al. [116] argue for the importance of making skin image data publicly available, suggesting that by doing so would enhance skin image recognition performance. Since the number of evidences included in this study from public datasets is limited (n = 13), whether this argument applies to non-image-based data necessitates further research for validation.
Data structure of features
Based on the findings of this study, predictive models utilizing solely structured data consistently demonstrate higher sensitivity and specificity across all prediction categories: admission, critical care, and mortality. Notably, the lowest sensitivity and specificity are observed in models predicting admission solely based on structured features. This observation may stem from the necessity of feature extraction in handling unstructured data, wherein differences in extraction methods could impact prediction performance variability, ultimately leading to less effective performance than models using structured features alone.
Although the integration of both structured and unstructured features theoretically offers better informative data for purposes of model development, this assertion remains unconfirmed by the current meta-analysis. Specifically regarding unstructured data, while the use of image data for critical care prediction yields higher sensitivity compared to free text, the opposite is observed for specificity. However, neither type of unstructured data significantly influences the predictive outcomes of ED disposition.
Sample type
Regarding sample selection, models utilizing adult samples consistently demonstrate higher sensitivity and specificity across all three ED dispositions—admission, critical care, and mortality— when compared to models using other sample types. Models employing mixed samples exhibit superior sensitivity and specificity in predicting admission and critical-care dispositions compared to other sample types. Furthermore, models utilizing youth samples demonstrate higher specificity in predicting mortality compared to other samples. However, there is no discernible pattern in the performance of prediction models based on sample utilization, suggesting that the choice of sample may not substantially impact prediction model performance.
Machine learning vs. deep learning
Among the included models, machine learning remains predominant. Generally, in predicting the three ED dispositions, models built using machine-learning methods demonstrate higher sensitivity and specificity compared to those employing deep learning methods, except for admission-prediction models, where sensitivity is higher in deep learning-based models. This study infers that since the included research data primarily consist of structured data rather than images, the complexity may be lower, thus making machine-learning methods adequate for handling the task. Conversely, deep-learning methods may not effectively leverage their image processing capabilities in this context. Further analysis of different machine learning algorithms reveals that Convolutional neural networks perform best for predicting ED dispositions related to admission. For critical care disposition predictions, eXtreme Gradient Boosting and LightGBM models show superior performance, while for mortality predictions, Random forest and LightGBM models demonstrate the highest performance.
Ensemble-learning technique
Using ensemble learning is generally believed to improve the predictive capability of models [117]. However, according to the results of this study, the situation is not entirely straightforward. For the prediction of admission, models not utilizing ensemble learning performed better, while for predicting critical care and mortality, models employing ensemble learning outperformed in both sensitivity and specificity. The discrepancy in performance may be attributed to the fact that the ensemble learning techniques employed in the study were not identical. This suggests that the selection of appropriate methods and parameter configurations is crucial when utilizing ensemble learning.
Cross-validation technique
Regarding the use of cross-validation, the results of this study show a mixed picture. For predicting admission, models without cross-validation demonstrated superior sensitivity and specificity compared to those with cross-validation. However, for predicting critical care and mortality, models utilizing cross-validation exhibited higher sensitivity and specificity than those without. There was no clear pattern indicating that adopting cross-validation consistently enhanced model performance across all prediction categories. An inference drawn from this study is that among the 51 models employing cross-validation, 26 did not undergo hyper-parameter tuning to find the optimal settings, potentially leading to sub-optimal performance improvements. Finally, our review highlights that most models predominantly relied on internal validation rather than external validation, raising concerns about potential overfitting.
Risk of bias assessment
Our study used PROBAST to assess the risk of bias across four domains: participants, predictors, outcomes, and analysis. Overall, the results show that most studies have a low risk of bias, indicating that these studies are well-designed with adequate sample sizes and appropriate handling of missing data. This finding differs from previous PROBAST-based assessments [118, 119], which often identified a high risk of bias in most prediction models. This discrepancy may be due to the fact that PROBAST was not specifically designed for AI applications, and some signaling questions may not fully apply in this context. This limitation could be addressed once the PROBAST + AI tool is officially released.
Future research directions
The analysis results of this study suggest that the specificity of the included models for predicting admission, critical care, and mortality is higher than their sensitivity. This implies that the models excel in correctly identifying non-cases of ED disposition, but may struggle to identify all relevant cases. While the sensitivity of these prediction models exceeds 80%, there is room for improvement in their predictive capabilities. Recommendations for enhancement could be explored in the following areas.
Define standard features for predicting ED disposition at various stages
Throughout the ED visitation process, a wide array of data is generated, encompassing physiological signs, injury records, diagnoses, laboratory findings, radiographic images, and far more. In the models analyzed in this meta-analysis, the utilization of data varies considerably across stages, making it challenging to identify over-arching patterns for comparison. Future research could explore constructing predictive models in stages based on the data generated during patients’ ED visits and evaluate the performance of these models at each stage. Additionally, recommendations for features and timeframes applicable to each stage of ED visits could be proposed to facilitate further model development. Early prediction of patient disposition in the ED holds significant potential for optimizing emergency medical resource management, service capabilities, and overall allocations.
Build a public dataset for predicting ED disposition
Once suggested features are identified, relevant data can be collected based on these features to construct predictive models. Another logical step is to establish a public dataset by leveraging collaborative efforts from EDs worldwide. This shared dataset aims to support hospitals in developing their own models for predicting ED patient disposition. Furthermore, with access to these shared datasets, different models become comparative, potentially enhancing predictive performance. The results of this meta-analysis also suggest that models using public datasets outperform those using private datasets in predicting admission and mortality.
Structure the nature of features for predicting ED disposition
In theory, unstructured data may contain more crucial information, suggesting that utilizing unstructured data could lead to better predictive model performance. However, the analysis results of this study do not support this argument. Instead, models built using structured data outperformed those using both structured and unstructured data in predicting admission, critical care, and mortality. Models solely based on unstructured data performed the least satisfactorily in predicting admission. One possible explanation for this finding may be that the unstructured data were completed using templates, resulting in uniform content and reducing the significance of the information contained within. It is suggested that future research prioritize structured data as they contain primary features, with the simultaneous use of structured and unstructured data as additional features.
Sample dataset for predicting ED disposition
Due to the possibility of incomplete physiological maturity in younger emergency department patients, such as with infants, their response to illness may differ significantly from that of adults, particularly the elderly. Therefore, it is suggested that future studies consider distinguishing between age groups when constructing predictive models for emergency department patient disposition. This approach would better cater to the clinical needs of emergency departments. In the studies included in this meta-analysis, certain models were specifically tailored for the elderly [25, 33, 45, 47, 48, 65] or for adolescents/infants [44, 82].
Utilize tailored artificial intelligence techniques for predicting ED disposition
Based on the results of this meta-analysis, it appears that the predictive performance of deep- learning models is generally lower than that of machine-learning models, contrary to the common belief among the general public that deep learning outperforms. Subsequent research should further investigate possible reasons for this discrepancy and subsequently enhance the predictive capabilities of models built using deep-learning methods. Additionally, ensemble learning in this meta-analysis demonstrated superior performance in predicting critical care and mortality compared to models that did not utilize ensemble learning. Future research may consider employing different types of ensemble learning to identify more effective model architectures. Moreover, while cross-validation theoretically aids in improving model predictive ability, it is recommended that future studies utilize hyper-parameter tuning alongside cross-validation to enhance model performance. Lastly, future studies are strongly encouraged to adopt external validation to minimize the risk of overfitting.
Limitations
This review has several limitations that warrant acknowledgment. Firstly, caution is needed when interpreting the pooled sensitivity and specificity of this study due to the presence of between-studies heterogeneity. Secondly, 71 articles were excluded due to insufficient quantitative information. It is recommended that future research on ED disposition using machine learning provide sufficient metric information to enhance profile study characteristics.
Conclusions
The main aim of this study is to meta-analyze the performance of artificial intelligence techniques used in predicting ED dispositions. Due to the lack of objective assessments in existing review literature on this topic, a comprehensive understanding of how artificial intelligence performs in predicting ED disposition is limited. This limitation may hinder the effective utilization of this technology, which could be crucial for optimizing emergency medical resources and for addressing vital issues such as ED overcrowding.
The primary findings of this study indicate that machine-learning techniques applied to predict different ED dispositions, including admission, critical care, or mortality, achieve AUROC scores ranging from 0.87 to 0.93. Models predicting mortality perform the best, with sensitivity and specificity ranging from 0.81 to 0.94. However, the specificity for each of the three ED dispositions is higher than sensitivity, suggesting room for improvement in predicting positive cases of ED disposition. Feasible approaches to address this matter include:
To establish standardized feature sets for predicting ED dispositions;
To create shared datasets for training predictive models, accessible to both emergency medical practitioners and researchers;
To integrate structured and unstructured datasets; and,
To leverage machine-learning techniques such as cross-validation with hyper-parameter tuning and ensemble learning to enhance performance.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Acknowledgements
None.
Abbreviations
- AUC
Area under curve
- AUROC
Area under receiver operating characteristic curve
- CI
Confidence interval
- CNN
Convolutional neural network
- DNN
Deep neural network
- DOR
Diagnostic odds ratio
- DT
Decision tree
- ED
Emergency department
- GBM
Gradient boosting machine
- HSROC
Hierarchical summary receiver operating characteristic curve
- ICU
Intensive care unit
- -LR
Negative likelihood ratio
- +LR
Positive likelihood ratio
- LR
Logistic regression
- NN
Neural network
- PROBAST
The Prediction Model Risk of Bias Assessment Tool
- RF
Random forest
- RNN
Recurrent neural network
- SVM
Support vector machine
- XGB
EXtreme gradient boosting
Author contributions
KMK and CSC conceived of this study and participated in the design and administration of the study. KMK and CSC drafted the manuscript and performed the statistical analysis. All authors read and approved the final manuscript.
Funding
This study was supported by the National Science and Technology Council, Taiwan, under grant number MOST-110-2410-H-239-015. The funder had no role in the study design, data collection, analysis, interpretation, or manuscript preparation.
Data availability
The datasets utilized and analyzed in this study are provided in the Additional file 3 and Additional file 4.
Declarations
Ethics approval and consent to participate
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards. This article does not contain any studies with animals performed by any of the authors. The experimental protocols, approved by the Institutional Review Board of E-DA Hospital (IRB No. EMRP-109-158), included waived informed-consent requirements.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Footnotes
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
- 1.Affleck A, Parks P, Drummond A, Rowe BH, Ovens HJ. Emergency department overcrowding and access block. Can J Emerg Med. 2013;15(6):359–84. 10.1017/s1481803500002451. [DOI] [PubMed] [Google Scholar]
- 2.Berlyand Y, Copenhaver MS, White BA, Dutta S, Baugh JJ, Wilcox SR, Yun BJ, Raja AS, Sonis JD. Impact of emergency department crowding on discharged patient experience. Western J Emerg Med. 2022;24(2):185–92. 10.5811/westjem.2022.10.58045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Parvaresh-Masoud M, Cheraghi MA, Imanipour M. Nurses’ perception of emergency department overcrowding: A qualitative study. J Educ Health Promotion. 2023;12(1). 10.4103/jehp.jehp_1789_22. [DOI] [PMC free article] [PubMed]
- 4.Eriksson J, Gellerstedt L, Hillerås P, Craftman Åsa G. Registered nurses’ perceptions of safe care in overcrowded emergency departments. J Clin Nurs. 2018;27(5–6):e1061–7. 10.1111/jocn.14143. [DOI] [PubMed] [Google Scholar]
- 5.Azari L, Turner K, Hong Y-R, Alishahi Tabriz A. Adoption of emergency department crowding interventions among US hospitals between 2007 and 2020. Am J Emerg Med. 2023;70:127–32. 10.1016/j.ajem.2023.05.034. [DOI] [PubMed] [Google Scholar]
- 6.Lee S, Kang WS, Kim DW, Seo SH, Kim J, Jeong ST, Yon DK, Lee J. An artificial intelligence model for predicting trauma mortality among emergency department patients in South Korea: retrospective cohort study. J Med Internet Res. 2023;25. 10.2196/49283. [DOI] [PMC free article] [PubMed]
- 7.Chang C-H, Chen C-J, Ma Y-S, Shen Y-T, Sung M-I, Hsu C-C, Lin H-J, Chen Z-C, Huang C-C, Liu C-F. Real-time artificial intelligence predicts adverse outcomes in acute pancreatitis in the emergency department: comparison with clinical decision rule. Academic Emergency Medicine 2023a, n/a(n/a). 10.1111/acem.14824 [DOI] [PubMed]
- 8.Son B, Myung J, Shin Y, Kim S, Kim SH, Chung JM, Noh J, Cho J, Chung HS. Improved patient mortality predictions in emergency departments with deep learning data-synthesis and ensemble models. Sci Rep. 2023;13(1). 10.1038/s41598-023-41544-0. [DOI] [PMC free article] [PubMed]
- 9.Shu T, Huang J, Deng J, Chen H, Zhang Y, Duan M, Wang Y, Hu X, Liu X. Development and assessment of scoring model for ICU stay and mortality prediction after emergency admissions in ischemic heart disease: a retrospective study of MIMIC-IV databases. Intern Emerg Med. 2023;18(2):487–97. 10.1007/s11739-023-03199-7. [DOI] [PubMed] [Google Scholar]
- 10.Shahul M, Pushpalatha KP. Machine Learning Based Patient Classification In Emergency Department. In: 2023 International Conference on Advances in Intelligent Computing and Applications. Kochi, India: IEEE; 2023.
- 11.Shung D, Simonov M, Gentry M, Au B, Laine L. Machine learning to predict outcomes in patients with acute Gastrointestinal bleeding: A systematic review. Dig Dis Sci. 2019;64(8):2078–87. 10.1007/s10620-019-05645-z. [DOI] [PubMed] [Google Scholar]
- 12.Guo A, Pasque M, Loh F, Mann DL, Payne PRO. Heart failure diagnosis, readmission, and mortality prediction using machine learning and artificial intelligence models. Curr Epidemiol Rep. 2020;7(4):212–9. 10.1007/s40471-020-00259-w. [Google Scholar]
- 13.Kareemi H, Vaillancourt C, Rosenberg H, Fournier K, Yadav K. Machine learning versus usual care for diagnostic and prognostic prediction in the emergency department: A systematic review. Acad Emerg Med. 2021;28(2):184–96. 10.1111/acem.14190. [DOI] [PubMed] [Google Scholar]
- 14.Naemi A, Schmidt T, Mansourvar M, Naghavi-Behzad M, Ebrahimi A, Wiil UK. Machine learning techniques for mortality prediction in emergency departments: a systematic review. BMJ Open. 2021;11(11):e052663. 10.1136/bmjopen-2021-052663. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Buttia C, Llanaj E, Raeisi-Dehkordi H, Kastrati L, Amiri M, Meçani R, Taneri PE, Ochoa SAG, Raguindin PF, Wehrli F, et al. Prognostic models in COVID-19 infection that predict severity: a systematic review. Eur J Epidemiol. 2023;38(4):355–72. 10.1007/s10654-023-00973-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Chen Y, Chen H, Sun Q, Zhai R, Liu X, Zhou J, Li S. Machine learning model identification and prediction of patients’ need for ICU admission: A systematic review. Am J Emerg Med. 2023;73:166–70. 10.1016/j.ajem.2023.08.043. [DOI] [PubMed] [Google Scholar]
- 17.Issaiy M, Zarei D, Saghazadeh A. Artificial intelligence and acute appendicitis: A systematic review of diagnostic and prognostic models. World J Emerg Surg. 2023;18(1):59. 10.1186/s13017-023-00527-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Larburu N, Azkue L, Kerexeta J. Predicting hospital ward admission from the emergency department: A systematic review. J Personalized Med. 2023;13(5):849. 10.3390/jpm13050849. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Olender RT, Roy S, Nishtala PS. Application of machine learning approaches in predicting clinical outcomes in older adults – a systematic review and meta-analysis. BMC Geriatr. 2023;23(1). 10.1186/s12877-023-04246-w. [DOI] [PMC free article] [PubMed]
- 20.Zhang Y, Xu W, Yang P, Zhang A. Machine learning for the prediction of sepsis-related death: a systematic review and meta-analysis. BMC Med Inf Decis Mak. 2023;23(1). 10.1186/s12911-023-02383-1. [DOI] [PMC free article] [PubMed]
- 21.Goto T, Camargo CA, Faridi MK, Yun BJ, Hasegawa K. Machine learning approaches for predicting disposition of asthma and COPD exacerbations in the ED. Am J Emerg Med. 2018;36(9):1650–4. 10.1016/j.ajem.2018.06.062. [DOI] [PubMed] [Google Scholar]
- 22.Ozer I, Cetin O, Gorur K, Temurtas F. Improved machine learning performances with transfer learning to predicting need for hospitalization in arboviral infections against the small dataset. Neural Comput Appl. 2021;33(21):14975–89. 10.1007/s00521-021-06133-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Raita Y, Goto T, Faridi MK, Brown DFM, Camargo CA, Hasegawa K. Emergency department triage prediction of clinical outcomes using machine learning models. Crit Care. 2019;23(1). 10.1186/s13054-019-2351-7. [DOI] [PMC free article] [PubMed]
- 24.Xie F, Zhou J, Lee JW, Tan M, Li S, Rajnthern LSO, Chee ML, Chakraborty B, Wong A-KI, Dagan A, et al. Benchmarking emergency department prediction models with machine learning and public electronic health records. Sci Data. 2022;9(1):658. 10.1038/s41597-022-01782-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Bunney G, Tran S, Han S, Gu C, Wang H, Luo Y, Dresden S. Using machine learning to predict hospital disposition with geriatric emergency department innovation intervention. Ann Emerg Med. 2023;81(3):353–63. 10.1016/j.annemergmed.2022.07.026. [DOI] [PubMed] [Google Scholar]
- 26.Chen C-H, Hsieh J-G, Cheng S-L, Lin Y-L, Lin P-H, Jeng J-H. Emergency department disposition prediction using a deep neural network with integrated clinical narratives and structured data. Int J Med Informatics. 2020;139:104146. 10.1016/j.ijmedinf.2020.104146. [DOI] [PubMed] [Google Scholar]
- 27.Li J, Guo L, Handly N. Hospital Admission Prediction Using Pre-hospital Variables. In: 2009 IEEE International Conference on Bioinformatics and Biomedicine. 2009: 283–286.
- 28.Patel D, Cheetirala SN, Raut G, Tamegue J, Kia A, Glicksberg B, Freeman R, Levin MA, Timsina P, Klang E. Predicting adult hospital admission from emergency department using machine learning: an inclusive gradient boosting model. J Clin Med. 2022;11(23). 10.3390/jcm11236888. [DOI] [PMC free article] [PubMed]
- 29.Roquette BP, Nagano H, Marujo EC, Maiorano AC. Prediction of admission in pediatric emergency department with deep neural networks and triage textual data. Neural Netw. 2020;126:170–7. 10.1016/j.neunet.2020.03.012. [DOI] [PubMed] [Google Scholar]
- 30.Akhlaghi H, Freeman S, Vari C, McKenna B, Braitberg G, Karro J, Tahayori B. Machine learning in clinical practice: evaluation of an artificial intelligence tool after implementation. Emerg Med Australasia. 2023;n/a(n/a). 10.1111/1742-6723.14325. [DOI] [PubMed]
- 31.Sterling NW, Patzer RE, Di M, Schrager JD. Prediction of emergency department patient disposition based on natural Language processing of triage notes. Int J Med Informatics. 2019;129:184–8. 10.1016/j.ijmedinf.2019.06.008. [DOI] [PubMed] [Google Scholar]
- 32.Tahayori B, Chini-Foroush N, Akhlaghi H. Advanced natural Language processing technique to predict patient disposition based on emergency triage notes. Emerg Med Australasia. 2021;33(3):480–4. 10.1111/1742-6723.13656. [DOI] [PubMed] [Google Scholar]
- 33.Goto T, Camargo CA Jr., Faridi MK, Freishtat RJ, Hasegawa K. Machine Learning–Based prediction of clinical outcomes for children during emergency department triage. JAMA Netw Open. 2019;2(1):e186937–186937. 10.1001/jamanetworkopen.2018.6937. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Barak-Corren Y, Agarwal I, Michelson KA, Lyons TW, Neuman MI, Lipsett SC, Kimia AA, Eisenberg MA, Capraro AJ, Levy JA, et al. Prediction of patient disposition: comparison of computer and human approaches and a proposed synthesis. J Am Med Inform Assoc. 2021;28(8):1736–45. 10.1093/jamia/ocab076. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Cusidó J, Comalrena J, Alavi H, Llunas L. Predicting hospital admissions to reduce crowding in the emergency departments. Appl Sci (Switzerland). 2022;12(21). 10.3390/app122110764.
- 36.Dadabhoy FZ, Driver L, McEvoy DS, Stevens R, Rubins D, Dutta S. Prospective external validation of a commercial model predicting the likelihood of inpatient admission from the emergency department. Ann Emerg Med. 2023;81(6):738–48. 10.1016/j.annemergmed.2022.11.012. [DOI] [PubMed] [Google Scholar]
- 37.De Hond A, Raven W, Schinkelshoek L, Gaakeer M, Ter Avest E, Sir O, Lameijer H, Hessels RA, Reijnen R, De Jonge E, et al. Machine learning for developing a prediction model of hospital admission of emergency department patients: hype or hope? Int J Med Informatics. 2021;152:104496. 10.1016/j.ijmedinf.2021.104496. [DOI] [PubMed] [Google Scholar]
- 38.Feretzakis G, Sakagianni A, Kalles D, Loupelis E, Panteris V, Tzelves L, Chatzikyriakou R, Trakas N, Kolokytha S, Batiani P et al. Using Machine Learning for Predicting the Hospitalization of Emergency Department Patients. In: Studies in Health Technology and Informatics: 2022; 2022: 405–408. [DOI] [PubMed]
- 39.Graham B, Bond R, Quinn M, Mulvenna M. Using data mining to predict hospital admissions from the emergency department. IEEE Access. 2018;6:10458–69. 10.1109/ACCESS.2018.2808843. [Google Scholar]
- 40.Heldt FS, Vizcaychipi MP, Peacock S, Cinelli M, McLachlan L, Andreotti F, Jovanović S, Dürichen R, Lipunova N, Fletcher RA, et al. Early risk assessment for COVID-19 patients from emergency department data using machine learning. Sci Rep. 2021;11(1):4200. 10.1038/s41598-021-83784-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Kim E, Han KS, Cheong T, Lee SW, Eun J, Kim SJ. Analysis on benefits and costs of machine Learning-Based early hospitalization prediction. IEEE Access. 2022;10:32479–93. 10.1109/ACCESS.2022.3160742. [Google Scholar]
- 42.Lam C, Calvert J, Siefkas A, Barnes G, Pellegrini E, Green-Saxena A, Hoffman J, Mao Q, Das R. Personalized stratification of hospitalization risk amidst COVID-19: A machine learning approach. Health Policy Technol. 2021;10(3):100554. 10.1016/j.hlpt.2021.100554. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Logaras E, Billis A, Kyparissidis Kokkinidis I, Ketseridou SN, Fourlis A, Tzotzis A, Imprialos K, Doumas M, Bamidis P. Risk assessment of COVID-19 cases in emergency departments and clinics with the use of Real-World data and artificial intelligence: observational study. JMIR Formative Res. 2022;6(11):e36933. 10.2196/36933. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Luo G, Stone BL, Nkoy FL, He S, Johnson MD. Predicting appropriate hospital admission of emergency department patients with bronchiolitis: secondary analysis. JMIR Med Inf. 2019;7(1). 10.2196/12591. [DOI] [PMC free article] [PubMed]
- 45.Mowbray F, Zargoush M, Jones A, de Wit K, Costa A. Predicting hospital admission for older emergency department patients: insights from machine learning. Int J Med Informatics. 2020;140:104163. 10.1016/j.ijmedinf.2020.104163. [DOI] [PubMed] [Google Scholar]
- 46.Patel SJ, Chamberlain DB, Chamberlain JM. A machine learning approach to predicting need for hospitalization for pediatric asthma exacerbation at the time of emergency department triage. Acad Emerg Med. 2018;25(12):1463–70. 10.1111/acem.13655. [DOI] [PubMed] [Google Scholar]
- 47.Tan TH, Hsu CC, Chen CJ, Hsu SL, Liu TL, Lin HJ, Wang JJ, Liu CF, Huang CC. Predicting outcomes in older ED patients with influenza in real time using a big data-driven and machine learning approach to the hospital information system. BMC Geriatr. 2021;21(1):280. 10.1186/s12877-021-02229-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Wolff P, Ríos SA, Graña M. Setting up standards: A methodological proposal for pediatric triage machine learning model construction based on clinical outcomes. Expert Syst Appl. 2019;138:112788. 10.1016/j.eswa.2019.07.005. [Google Scholar]
- 49.Chen M-C, Huang T-Y, Chen T-Y, Boonyarat P, Chang Y-C. Clinical narrative-aware deep neural network for emergency department critical outcome prediction. J Biomedical Inf. 2023;138:104284. 10.1016/j.jbi.2023.104284 [DOI] [PubMed]
- 50.Choi SW, Ko T, Hong KJ, Kim KH. Machine Learning-Based prediction of Korean triage and acuity scale level in emergency department patients. Healthc Inf Res. 2019;25(4):305–12. 10.4258/hir.2019.25.4.305. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Dayan I, Roth HR, Zhong A, Harouni A, Gentili A, Abidin AZ, Liu A, Costa AB, Wood BJ, Tsai C-S, et al. Federated learning for predicting clinical outcomes in patients with COVID-19. Nat Med. 2021;27(10):1735–43. 10.1038/s41591-021-01506-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Di Napoli A, Tagliente E, Pasquini L, Cipriano E, Pietrantonio F, Ortis P, Curti S, Boellis A, Stefanini T, Bernardini A, et al. 3D CT-Inclusive Deep-Learning model to predict mortality, ICU admittance, and intubation in COVID-19 patients. J Digit Imaging. 2022. 10.1007/s10278-022-00734-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Dipaola F, Gatti M, Giaj Levra A, Menè R, Shiffer D, Faccincani R, Raouf Z, Secchi A, Rovere Querini P, Voza A, et al. Multimodal deep learning for COVID-19 prognosis prediction in the emergency department: a bi-centric study. Sci Rep. 2023;13(1):10868. 10.1038/s41598-023-37512-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Fernandes M, Mendes R, Vieira SM, Leite F, Palos C, Johnson A, Finkelstein S, Horng S, Celi LA. Risk of mortality and cardiopulmonary arrest in critical patients presenting to the emergency department using machine learning and natural Language processing. PLoS ONE. 2020;15(4). 10.1371/journal.pone.0230876. [DOI] [PMC free article] [PubMed]
- 55.Fernandes M, Mendes R, Vieira SM, Leite F, Palos C, Johnson A, Finkelstein S, Horng S, Celi LA. Predicting intensive care unit admission among patients presenting to the emergency department using machine learning and natural Language processing. PLoS ONE. 2020b;15(3):e0229331. 10.1371/journal.pone.0229331. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Joseph JW, Leventhal EL, Grossestreuer AV, Wong ML, Joseph LJ, Nathanson LA, Donnino MW, Elhadad N, Sanchez LD. Deep-learning approaches to identify critically ill patients at emergency department triage using limited information. J Am Coll Emerg Physicians Open. 2020;1(5):773–81. 10.1002/emp2.12218. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Klang E, Kummer BR, Dangayach NS, Zhong A, Kia MA, Timsina P, Cossentino I, Costa AB, Levin MA, Oermann EK. Predicting adult neuroscience intensive care unit admission from emergency department triage using a retrospective, tabular-free text machine learning approach. Sci Rep. 2021;11(1). 10.1038/s41598-021-80985-3. [DOI] [PMC free article] [PubMed]
- 58.Butler L, Karabayir I, Samie Tootooni M, Afshar M, Goldberg A, Akbilgic O. Image and structured data analysis for prognostication of health outcomes in patients presenting to the ED during the COVID-19 pandemic. Int J Med Informatics. 2021;158:104662. 10.1016/j.ijmedinf.2021.104662. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Kang D-Y, Cho K-J, Kwon O, Kwon J-m, Jeon K-H, Park H, Lee Y, Park J, Oh B-H. Artificial intelligence algorithm to predict the need for critical care in prehospital emergency medical services. Scand J Trauma Resusc Emerg Med. 2020;28(1). 10.1186/s13049-020-0713-4. [DOI] [PMC free article] [PubMed]
- 60.Choi A, Choi SY, Chung K, Chung HS, Song T, Choi B, Kim JH. Development of a machine learning-based clinical decision support system to predict clinical deterioration in patients visiting the emergency department. Sci Rep. 2023;13(1). 10.1038/s41598-023-35617-3. [DOI] [PMC free article] [PubMed]
- 61.Cardosi JD, Shen H, Groner JI, Armstrong M, Xiang H. Machine learning for outcome predictions of patients with trauma during emergency department care. BMJ Health Care Inf. 2021;28(1). 10.1136/bmjhci-2021-100407. [DOI] [PMC free article] [PubMed]
- 62.Duanmu H, Ren T, Li H, Mehta N, Singer AJ, Levsky JM, Lipton ML, Duong TQ. Deep learning of longitudinal chest X-ray and clinical variables predicts duration on ventilator and mortality in COVID-19 patients. Biomed Eng Online. 2022;21(1). 10.1186/s12938-022-01045-z. [DOI] [PMC free article] [PubMed]
- 63.Klang E, Levin MA, Soffer S, Zebrowski A, Glicksberg BS, Carr BG, McGreevy J, Reich DL, Freeman R. A simple free-text-like method for extracting semi-structured data from electronic health records: exemplified in prediction of in-hospital mortality. Big Data Cogn Comput. 2021;5(3). 10.3390/bdcc5030040.
- 64.Cheng CY, Kung CT, Chen FC, Chiu IM, Lin CHR, Chu CC, Kung CF, Su CM. Machine learning models for predicting in-hospital mortality in patient with sepsis: analysis of vital sign dynamics. Front Med. 2022;9. 10.3389/fmed.2022.964667. [DOI] [PMC free article] [PubMed]
- 65.Fransvea P, Fransvea G, Liuzzi P, Sganga G, Mannini A, Costa G. Study and validation of an explainable machine learning-based mortality prediction following emergency surgery in the elderly: A prospective observational study. Int J Surg. 2022;107:106954. 10.1016/j.ijsu.2022.106954. [DOI] [PubMed] [Google Scholar]
- 66.Lee S, Kang WS, Seo S, Kim DW, Ko H, Kim J, Lee S, Lee J. Model for predicting In-Hospital mortality of physical trauma patients using artificial intelligence techniques: nationwide Population-Based study in Korea. J Med Internet Res. 2022;24(12). 10.2196/43757. [DOI] [PMC free article] [PubMed]
- 67.Shahi N, Shahi AK, Phillips R, Shirek G, Bensard D, Moulton SL. Decision-making in pediatric blunt solid organ injury: A deep learning approach to predict massive transfusion, need for operative management, and mortality risk. J Pediatr Surg. 2021;56(2):379–84. 10.1016/j.jpedsurg.2020.10.021. [DOI] [PubMed] [Google Scholar]
- 68.Page MJ, McKenzie JE, Bossuyt PM, Boutron I, Hoffmann TC, Mulrow CD, Shamseer L, Tetzlaff JM, Akl EA, Brennan SE, et al. The PRISMA 2020 statement: an updated guideline for reporting systematic reviews. BMJ. 2021a;372:n71. 10.1136/bmj.n71. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Page MJ, McKenzie JE, Bossuyt PM, Boutron I, Hoffmann TC, Mulrow CD, Shamseer L, Tetzlaff JM, Akl EA, Brennan SE et al. The PRISMA 2020 statement: an updated guideline for reporting systematic reviews. BMJ 2021b, 372:n71. 10.1136/bmj.n71 [DOI] [PMC free article] [PubMed]
- 70.Hadzikadic M, Hakenewerth A, Bohren B, Norton J, Mehta B, Andrews C. Concept formation vs. logistic regression: predicting death in trauma patients. Artif Intell Med. 1996;8(5):493–504. 10.1016/S0933-3657(96)00356-9. [DOI] [PubMed] [Google Scholar]
- 71.Berikol GB, Yildiz O, Özcan İT. Diagnosis of acute coronary syndrome with a support vector machine. J Med Syst. 2016;40(4). 10.1007/s10916-016-0432-6. [DOI] [PubMed]
- 72.Golmohammadi D. Predicting hospital admissions to reduce emergency department boarding. Int J Prod Econ. 2016;182:535–44. 10.1016/j.ijpe.2016.09.020. [Google Scholar]
- 73.Schütz N, Leichtle AB, Riesen K. A comparative study of pattern recognition algorithms for predicting the inpatient mortality risk using routine laboratory measurements. Artif Intell Rev. 2018;52(4):2559–73. 10.1007/s10462-018-9625-3. [Google Scholar]
- 74.Hong JC, Niedzwiecki D, Palta M, Tenenbaum JD. Predicting emergency visits and hospital admissions during radiation and chemoradiation: an internally validated pretreatment machine learning algorithm. JCO Clin Cancer Inf 2018a, 2:1–11. 10.1200/cci.18.00037 [DOI] [PubMed]
- 75.Hong WS, Haimovich AD, Taylor RA. Predicting hospital admission at emergency department triage using machine learning. PLoS One. 2018;13(7). 10.1371/journal.pone.0201016 [DOI] [PMC free article] [PubMed]
- 76.Klug M, Barash Y, Bechler S, Resheff YS, Tron T, Ironi A, Soffer S, Zimlichman E, Klang E. A gradient boosting machine learning model for predicting early mortality in the emergency department triage: devising a Nine-Point triage score. J Gen Intern Med. 2019;35(1):220–7. 10.1007/s11606-019-05512-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Lee S-Y, Chinnam RB, Dalkiran E, Krupp S, Nauss M. Prediction of emergency department patient disposition decision for proactive resource allocation for admission. Health Care Manag Sci. 2019;23(3):339–59. 10.1007/s10729-019-09496-y. [DOI] [PubMed] [Google Scholar]
- 78.Hong S, Lee S, Lee J, Cha WC, Kim K. Prediction of cardiac arrest in the emergency department based on machine learning and sequential characteristics: model development and retrospective clinical validation study. JMIR Med Inf. 2020;8(8):e15932. 10.2196/15932. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Lee S, Hong S, Cha WC, Kim K. Predicting adverse outcomes for febrile patients in the emergency department using sparse laboratory data: development of a time adaptive model. JMIR Med Inf. 2020;8(3). 10.2196/16117. [DOI] [PMC free article] [PubMed]
- 80.Zhang P-I, Hsu C-C, Kao Y, Chen C-J, Kuo Y-W, Hsu S-L, Liu T-L, Lin H-J, Wang J-J, Liu C-F, et al. Real-time AI prediction for major adverse cardiac events in emergency department patients with chest pain. Scand J Trauma Resusc Emerg Med. 2020;28(1). 10.1186/s13049-020-00786-x. [DOI] [PMC free article] [PubMed]
- 81.Chen Y-M, Kao Y, Hsu C-C, Chen C-J, Ma Y-S, Shen Y-T, Liu T-L, Hsu S-L, Lin H-J, Wang J-J, et al. Real-time interactive artificial intelligence of things–based prediction for adverse outcomes in adult patients with pneumonia in the emergency department. Acad Emerg Med. 2021;28(11):1277–85. 10.1111/acem.14339. [DOI] [PubMed] [Google Scholar]
- 82.Chiu IM, Cheng CY, Zeng WH, Huang YH, Lin CR. Using machine learning to predict invasive bacterial infections in young febrile infants visiting the emergency department. J Clin Med. 2021;10(9). 10.3390/jcm10091875. [DOI] [PMC free article] [PubMed]
- 83.Douville NJ, Douville CB, Mentz G, Mathis MR, Pancaro C, Tremper KK, Engoren M. Clinically applicable approach for predicting mechanical ventilation in patients with COVID-19. Br J Anaesth. 2021;126(3):578–89. 10.1016/j.bja.2020.11.034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Garrafa E, Vezzoli M, Ravanelli M, Farina D, Borghesi A, Calza S, Maroldi R. Early prediction of in-hospital death of covid-19 patients: A machine-learning model based on age, blood analyses, and chest x-ray score. eLife. 2021;10. 10.7554/eLife.70640. [DOI] [PMC free article] [PubMed]
- 85.Horng G-J, Lin T-C, Lee K-C, Chen K-T, Hsu C-C. Prediction of prognosis in emergency trauma patients with optimal limit gradient based on grid search optimal parameters. Wireless Pers Commun. 2021;120(2):1741–51. 10.1007/s11277-021-08532-x. [Google Scholar]
- 86.Hsu SD, Chao E, Chen SJ, Hueng DY, Lan HY, Chiang HH. Machine learning algorithms to predict in-hospital mortality in patients with traumatic brain injury. J Personalized Med. 2021;11(11). 10.3390/jpm11111144. [DOI] [PMC free article] [PubMed]
- 87.Jiang H, Mao H, Lu H, Lin P, Garry W, Lu H, Yang G, Rainer TH, Chen X. Machine learning-based models to support decision-making in emergency department triage for patients with suspected cardiovascular disease. Int J Med Informatics. 2021;145:104326. 10.1016/j.ijmedinf.2020.104326. [DOI] [PubMed] [Google Scholar]
- 88.Li C, Zhang Z, Ren Y, Nie H, Lei Y, Qiu H, Xu Z, Pu X. Machine learning based early mortality prediction in the emergency department. Int J Med Informatics. 2021;155:104570. 10.1016/j.ijmedinf.2021.104570. [DOI] [PubMed] [Google Scholar]
- 89.Lu JQ, Musheyev B, Peng Q, Duong TQ. Neural network analysis of clinical variables predicts escalated care in COVID-19 patients: A retrospective study. PeerJ. 2021;9. 10.7717/peerj.11205. [DOI] [PMC free article] [PubMed]
- 90.Wu TT, Zheng RF, Lin ZZ, Gong HR, Li H. A machine learning model to predict critical care outcomes in patient with chest pain visiting the emergency department. BMC Emerg Med. 2021;21(1). 10.1186/s12873-021-00501-8. [DOI] [PMC free article] [PubMed]
- 91.Yun H, Choi J, Park JH. Prediction of critical care outcome for adult patients presenting to emergency department using initial triage information: an XGBoost algorithm analysis. JMIR Med Inf. 2021;9(9). 10.2196/30770. [DOI] [PMC free article] [PubMed]
- 92.Lee JT, Hsieh CC, Lin CH, Lin YJ, Kao CY. Prediction of hospitalization using artificial intelligence for urgent patients in the emergency department. Sci Rep. 2021;11(1). 10.1038/s41598-021-98961-2. [DOI] [PMC free article] [PubMed]
- 93.Calvillo-Batllés P, Cerdá-Alberich L, Fonfría-Esparcia C, Carreres-Ortega A, Muñoz-Núñez CF, Trilles-Olaso L, Martí-Bonmatí L. Development of severity and mortality prediction models for covid-19 patients at emergency department including the chest x-ray. Radiologia. 2022;64(3):214–27. 10.1016/j.rxeng.2021.09.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Chang Y-H, Shih H-M, Wu J-E, Huang F-W, Chen W-K, Chen D-M, Chung Y-T, Wang CCN. Machine learning–based triage to identify low-severity patients with a short discharge length of stay in emergency department. BMC Emerg Med. 2022;22(1). 10.1186/s12873-022-00632-6. [DOI] [PMC free article] [PubMed]
- 95.Hasan M, Bath PA, Marincowitz C, Sutton L, Pilbery R, Hopfgartner F, Mazumdar S, Campbell R, Stone T, Thomas B, et al. Pre-hospital prediction of adverse outcomes in patients with suspected COVID-19: development, application and comparison of machine learning and deep learning methods. Comput Biol Med. 2022;151. 10.1016/j.compbiomed.2022.106024. [DOI] [PMC free article] [PubMed]
- 96.Ke J, Chen Y, Wang X, Wu Z, Zhang Q, Lian Y, Chen F. Machine learning-based in-hospital mortality prediction models for patients with acute coronary syndrome. Am J Emerg Med. 2022;53:127–34. 10.1016/j.ajem.2021.12.070. [DOI] [PubMed] [Google Scholar]
- 97.Maria A, Dimitrios V, Ioanna M, Charalampos M, Gerasimos M, Constantinos K. Clinical Decision Making and Outcome Prediction for COVID-19 Patients Using Machine Learning. In: 15th EAI International Conference, Pervasive Health 2021. Virtual Event; 2022.
- 98.Casano N, Santini SJ, Vittorini P, Sinatti G, Carducci P, Mastroianni CM, Ciardi MR, Pasculli P, Petrucci E, Marinangeli F, et al. Application of machine learning approach in emergency department to support clinical decision making for SARS-CoV-2 infected patients. J Integr Bioinform. 2023;20(2). 10.1515/jib-2022-0047. [DOI] [PMC free article] [PubMed]
- 99.Elhaj H, Achour N, Tania MH, Aciksari K. A comparative study of supervised machine learning approaches to predict patient triage outcomes in hospital emergency departments. Array. 2023;17:100281. 10.1016/j.array.2023.100281. [Google Scholar]
- 100.Greco M, Caruso PF, Spano S, Citterio G, Desai A, Molteni A, Aceto R, Costantini E, Voza A, Cecconi M. Machine learning for early outcome prediction in septic patients in the emergency department. Algorithms. 2023;16(2). 10.3390/a16020076.
- 101.Hsu C-C, Kao Y, Hsu C-C, Chen C-J, Hsu S-L, Liu T-L, Lin H-J, Wang J-J, Liu C-F, Huang C-C. Using artificial intelligence to predict adverse outcomes in emergency department patients with hyperglycemic crises in real time. BMC Endocr Disorders. 2023;23(1). 10.1186/s12902-023-01437-9. [DOI] [PMC free article] [PubMed]
- 102.Matsuo K, Aihara H, Hara Y, Morishita A, Sakagami Y, Miyake S, Tatsumi S, Ishihara S, Tohma Y, Yamashita H, et al. Machine learning to predict three types of outcomes after traumatic brain injury using data at admission: A Multi-Center study for development and validation. J Neurotrauma. 2023. 10.1089/neu.2022.0515. [DOI] [PubMed] [Google Scholar]
- 103.Mekkodathil A, El-Menyar A, Naduvilekandy M, Rizoli S, Al-Thani H. Machine learning approach for the prediction of In-Hospital mortality in traumatic brain injury using Bio-Clinical markers at presentation to the emergency department. Diagnostics. 2023;13(15). 10.3390/diagnostics13152605. [DOI] [PMC free article] [PubMed]
- 104.Pai DR, Rajan B, Jairath P, Rosito SM. Predicting hospital admission from emergency department triage data for patients presenting with fall-related fractures. Intern Emerg Med. 2023;18(1):219–27. 10.1007/s11739-022-03100-y. [DOI] [PubMed] [Google Scholar]
- 105.Logothetis SB, Green D, Holland M, Al Moubayed N. Predicting acute clinical deterioration with interpretable machine learning to support emergency care decision making. Sci Rep. 2023;13(1). 10.1038/s41598-023-40661-0. [DOI] [PMC free article] [PubMed]
- 106.Moons KGM, Wolff RF, Riley RD, Whiting PF, Westwood M, Collins GS, Reitsma JB, Kleijnen J, Mallett S. PROBAST: A tool to assess risk of Bias and applicability of prediction model studies: explanation and elaboration. Ann Intern Med. 2019;170(1):W1–33. 10.7326/M18-1377. [DOI] [PubMed] [Google Scholar]
- 107.Wolff RF, Moons KGM, Riley RD, Whiting PF, Westwood M, Collins GS, Reitsma JB, Kleijnen J, Mallett S. PROBAST: A tool to assess the risk of Bias and applicability of prediction model studies. Ann Intern Med. 2019;170(1):51–8. 10.7326/M18-1376. [DOI] [PubMed] [Google Scholar]
- 108.Takwoingi Y, Riley RD, Deeks JJ. Meta-analysis of diagnostic accuracy studies in mental health. Evid Based Mental Health. 2015;18(4):103. 10.1136/eb-2015-102228. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.R Core Team. In: Vienna, editor. R: A Language and environment for statistical computing. Austria: R Foundation for Statistical Computing; 2023. [Google Scholar]
- 110.Bates D, Mächler M, Bolker B, Walker S. Fitting linear Mixed-Effects models using lme4. J Stat Softw. 2015;67. 10.18637/jss.v067.i01.
- 111.Doebler P. mada: Meta-Analysis of Diagnostic Accuracy. In R package version 0.5.9. edn; 2019.
- 112.Freeman SC, Kerby CR, Patel A, Cooper NJ, Quinn T, Sutton AJ. Development of an interactive web-based tool to conduct and interrogate meta-analysis of diagnostic test accuracy studies: MetaDTA. BMC Med Res Methodol. 2019;19(1):81. 10.1186/s12874-019-0724-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113.Patel A, Cooper N, Freeman S, Sutton A. Graphical enhancements to summary receiver operating characteristic plots to facilitate the analysis and reporting of meta-analysis of diagnostic test accuracy data. Res Synthesis Methods. 2021;12(1):34–44. 10.1002/jrsm.1439. [DOI] [PubMed] [Google Scholar]
- 114.Glas AS, Lijmer JG, Prins MH, Bonsel GJ, Bossuyt PMM. The diagnostic odds ratio: a single indicator of test performance. J Clin Epidemiol. 2003;56(11):1129–35. 10.1016/S0895-4356(03)00177-X. [DOI] [PubMed] [Google Scholar]
- 115.Deeks JJ, Altman DG. Diagnostic tests 4: likelihood ratios. BMJ. 2004;329(7458):168–9. 10.1136/bmj.329.7458.168. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 116.Tschandl P, Codella N, Akay BN, Argenziano G, Braun RP, Cabo H, Gutman D, Halpern A, Helba B, Hofmann-Wellenhof R, et al. Comparison of the accuracy of human readers versus machine-learning algorithms for pigmented skin lesion classification: an open, web-based, international, diagnostic study. Lancet Oncol. 2019;20(7):938–47. 10.1016/S1470-2045(19)30333-X. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 117.Brownlee J. Ensemble learning algorithms with Python. Melbourne, Australia: Machine Learning Mastery; 2020. [Google Scholar]
- 118.Dhiman P, Ma J, Andaur Navarro CL, Speich B, Bullock G, Damen JAA, Hooft L, Kirtley S, Riley RD, Van Calster B, et al. Risk of bias of prognostic models developed using machine learning: a systematic review in oncology. Diagn Prognostic Res. 2022;6(1):13. 10.1186/s41512-022-00126-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 119.Wynants L, Van Calster B, Collins GS, Riley RD, Heinze G, Schuit E, Albu E, Arshi B, Bellou V, Bonten MMJ, et al. Prediction models for diagnosis and prognosis of covid-19: systematic review and critical appraisal. BMJ. 2020;369:m1328. 10.1136/bmj.m1328. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The datasets utilized and analyzed in this study are provided in the Additional file 3 and Additional file 4.