ABSTRACT
Unknown ellipsoid bodies, later classified as apicomplexan cysts, are prevalent in the ovaries of Japanese sea cucumbers ( Apostichopus japonicus ), where they can lead to lower fecundity in infected individuals and adverse effects on wild populations as well as aquaculture efforts for this endangered species. Apicomplexans are widespread and essential to marine environments, where they can affect the health and fitness of host populations. We performed genomic sequencing of recovered cysts to gain more ecological and evolutionary information on this parasite. We recovered this apicomplexan's complete nuclear ribosomal RNA (rrn) operon, the entire mitochondrial genome, and a partial apicoplast (relic chloroplast) genome. The rrn operon phylogeny revealed this parasite as being closely related to coccidian‐like parasites of marine fish (ichthyocolids) and cnidarians (corallicolids), while organelle phylogenomics hint at a closer relation to the protococcidian Eleutheroschizon. Using this new phylogenetic context and previous morphological descriptions, we describe this parasite as Apostichocystis gudetama gen. nov. sp. nov. Mining available microbiomes reveal the presence of Apostichocystis spp. beyond its host range, alluding to other potential hosts or cryptic, closely related lineages. Its phylogenetic placement has important implications concerning the evolution of parasitism within Apicomplexa and the divergence of a marine‐host‐specific clade of coccidian‐like parasites.
Keywords: apicomplexa, aquaculture, coccidia, holobiont, holothuria, host–microbe interactions, metagenomics, parasitism, protococcidia, symbiosis
1. Introduction
The Japanese sea cucumber ( Apostichopus japonicus ) is a commercially valuable species currently endangered according to the International Union for Conservation of Nature Red List of Threatened Species due to overharvesting (Robinson and Lovatelli 2015). Aquaculture of these sea cucumbers has expanded significantly in recent years, especially in Japan and China (Yu et al. 2014; Robinson and Lovatelli 2015; Yang et al. 2015), where they are a popular seafood. To account for over‐harvesting, some Japanese hatcheries restock wild A. japonicus populations by releasing juveniles to fishery grounds (Sakai 2015). Through these hatchery operations, technicians first noted unknown ellipsoidal bodies (UEBs), about ~250 μm in length, within the ovaries of female sea cucumbers (Figure 1). The individuals with UEBs‐filled ovaries did not spawn with the normal temperature cue used for spawning induction of A. japonicus and have been shown to have lower fecundity than their non‐infected counterparts (Unuma et al. 2020). These UEBs were later characterized as coccidian (Apicomplexa; Coccidia) cysts by sequencing of the small subunit ribosomal RNA (18S rRNA gene) (Unuma et al. 2020). Importantly, this parasite has been detected in more than half of the females during the reproductive season (Unuma et al. 2020), which infer a potential ecological trade‐off and co‐evolution between the host and parasite.
FIGURE 1.
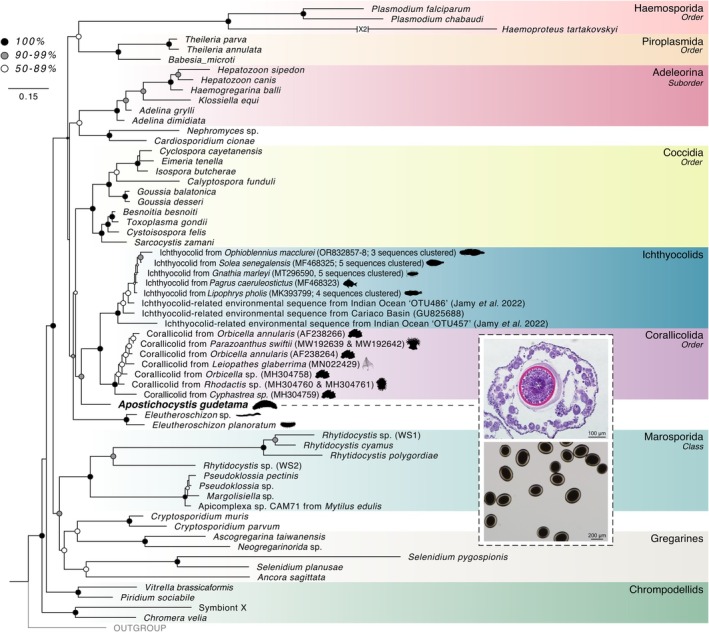
This maximum likelihood tree was constructed using the 18S and 28S subunits of the rRNA operon using 5017 nucleic acid sites with RAxML General Time Reversible model with a GAMMA+P‐Invar model of rate heterogeneity (‘‐m GTRGAMMAIX’) and 1000 bootstrap replicates. Silhouettes of the primary hosts for select coccidians and protococcidians are also shown. (See also Supporting Information S1A). Upper image inset—hematoxylin and eosin‐stained cysts. Lower image inset—stereoscopic micrograph of cysts isolated from the ovaries of the Japanese sea cucumber, Apostichopus japonicus .
This sea cucumber‐infecting parasite belongs to the phylum Apicomplexa, which encompasses many obligate associates of animal hosts. Dozens of apicomplexan‐related lineages (ARLs) (Jan Janouškovec et al. 2012; Janouškovec et al. 2019) have been identified from environmental sequencing data and, along with more genomic work on core apicomplexans, has expanded our knowledge on the origin of parasitism (Mathur et al. 2019), organelle evolution (McFadden and Yeh 2017), and the breadth of known microeukaryotic diversity (del Campo et al. 2019). The corallicolids (previously known as ARL‐V) and ichthyocolids (previously known as ARL‐VI), widespread symbionts of anthozoans and marine teleosts, respectively, are two recently characterized lineages of apicomplexan parasites from marine hosts that have enlightened our understanding of parasite evolution (Jacko‐Reynolds et al. 2025; Bonacolta et al. 2024). New phylogenomic data suggest these parasites represent a more recently divergent lineage in the context of apicomplexan evolution (i.e., late‐branching), possibly with the Protococcidiorida clade (Jacko‐Reynolds et al. 2025). The Protococcidiorida is a poorly described group of apicomplexans, basal to Coccidia, which includes marine representatives such as the polychaeta‐infecting Eleutheroschizon duboscqi (Valigurová et al. 2015). The 18S rRNA gene alone does not resolve the lower taxonomic classification of the coccidian‐like parasite of Japanese sea cucumber. Therefore, more work is needed to place this parasite within an evolutionary context confidently, correlate it to another known ARL, and to investigate its host and geographic distribution further.
To investigate the evolutionary history of this enigmatic parasite, deep genomic sequencing was conducted on isolated cysts recovered from the ovaries of A. japonicus . This approach recovered enough genetic material from this apicomplexan parasite to retrieve the complete nuclear ribosomal RNA (rrn) operon, entire mitochondrial genome, and partial apicoplast (non‐photosynthetic chloroplast) genome. This new genetic information was utilized to learn more about this parasite's evolutionary history and further distribution. Lastly, we formally describe this new species by combining this phylogenetic context with previous morphological descriptions.
2. Materials and Methods
2.1. Isolation of Cysts
Cysts were extracted from A. japonicus specimen as described in Unuma et al. (2020). Briefly, a green morphotype of A. japonicus was collected by gaff fishing at Hamajima (34°29′ N, 136°76′ E), Mie, Japan, on December 17, 2010, then transported to the Mie Prefectural Fish Farming Center. Once in aquaculture, the ovaries were examined under a stereoscopic microscope for the presence of cysts. Once found, cysts were manually extracted and stored in ethanol at −30°C until DNA extraction. Histological imaging was performed as described in Unuma et al. (2020). Briefly, a portion of the infected ovaries was fixed in Davidson's solution (Bell and Lightner 1988). The tissue was then embedded in paraffin wax and stained using hematoxylin and eosin.
2.2. DNA Extraction and Low‐Input Metagenomic Sequencing
DNA was extracted from combined cysts using the QIAGEN DNeasy Blood & Tissue Kit (Germantown, Maryland, USA) with an overnight proteinase K digestion step. Extracted DNA was quantified using a QIAGEN Qubit 4 Fluorometer with the Qubit 1X dsDNA HS Assay Kit. Extracted DNA was then brought to the Center for Genome Technology at the John P. Hussman Institute for Human Genomics (University of Miami Miller School of Medicine), where it was prepared for sequencing using the NEBNext Ultra II DNA Library Prep Kit with 100 ng of input and 3 cycles of PCR. DNA libraries were sequenced on an Illumina Novaseq X (San Diego, California, USA) targeting 50 M 150 × 150 paired‐end reads. Paired‐end reads were trimmed of adapters and low‐quality (< 20 Q‐score) reads using Fastp (Chen et al. 2018). Trimmed reads were assembled into a metagenome using Megahit with default settings (Li et al. 2015).
2.3. Extraction of Rrn Operon and Phylogenetics
To retrieve the complete rrn operon (including the 18S + 28S rRNA genes), a hidden Markov Model (HMM) profile was generated based on an alignment of published microeukaryotic rrn operons (Supporting Information S1A). This profile was then used with nhmmer (Wheeler and Eddy 2013) to extract any matching sequences from the assembled metagenome, resulting in the recovery of the majority of the 28S rRNA and ITS portions of the operon. The 18S rRNA gene was recovered using barrnap v0.9 (Seemann 2018) and then combined with the previously extracted 28S and ITS regions to assemble the complete rrn operon.
A rrn operon tree of Apicomplexa was generated using previously recovered 18S and 28S rRNA genes from representative apicomplexan lineages (Supporting Information S1A). Whole rrn operon sequences longer than 500 bp and shorter than 15,000 bp were sorted and clustered at 99%, then aligned using MAFFT v7.480 (Katoh and Standley 2013). The alignment was then trimmed using Trimal v1.4.rev15 (Capella‐Gutiérrez et al. 2009) with a gap threshold of 30%, a similarity threshold of 0.001, and a minimum conservation of 30%. ModelTest‐NG v0.1.7 (Darriba et al. 2020) was run on the trimmed alignment to determine the optimal substitution model for phylogenetic reconstruction. A maximum‐likelihood tree was then constructed using RAxML v8.2.12 with the General Time Reversible model with a GAMMA+P‐Invar model of rate heterogeneity (‘‐m GTRGAMMAIX’) and 1000 bootstrap replicates (Stamatakis 2014).
2.4. Organelle Assemblies and Phylogenetics
The mitochondrial genome of this parasite was assembled by first identifying mitochondrial contigs within the metagenomic assembly using tblastn v2.13.0 (Camacho et al. 2009) against an apicomplexan mitochondrial gene database from Bonacolta et al. (2024). Using this approach, a contig representing the incomplete mitochondrial genome of the parasite was identified and utilized as an input seed for Novoplasty v4.3.3.pl (Dierckxsens et al. 2017), which assembled a complete mitochondrial genome from the raw sequencing reads. MFANNOT was used for mitochondrial genome annotation (Lang et al. 2023). The cox1, cox3, and cob genes were then extracted and aligned using MUSCLE v3.8.1551 (Edgar 2004) with those from other apicomplexans (plus a dinoflagellate outgroup; Supporting Information S1B). Gene alignments were combined using CONCATENATOR v0.2.1 (Vences et al. 2022) and then trimmed using Trimal v1.4.rev15 (Capella‐Gutiérrez et al. 2009) with a gap threshold of 20%. This gap threshold was chosen by manually inspecting the output from a range of thresholds to ensure sufficient information was retained while removing poorly aligned regions. IQ‐TREE with the C60 + LG + GAMMA model and 1000 bootstrap replicates was run to generate a phylogenetic reconstruction of apicomplexan mitochondrial evolution (Nguyen et al. 2015). This model was chosen because it performed better than other, less complex models (Baños et al. 2024).
Contigs corresponding to apicoplast genes were identified using BLASTn v2.13.0 against the NCBI database. After failing to get an assembly using Novoplasty as detailed for the mitochondrial genome above, these contigs were used as seed input for GetOrganelle v1.7.7 (Jin et al. 2020) to generate a fragmented apicoplast genome. A second round of GetOrganelle v1.7.7 using the fragmented assembly as a seed yielded a less fragmented but still incomplete apicoplast genome. MFANNOT was used to annotate the genes we could recover from this assembly, and then the genes were aligned with their apicomplexan counterparts using MUSCLE v3.8.1551. The alignments were then trimmed using Trimal v1.4.rev15 with a gap threshold of 60% using the same methodology mentioned above. The apicoplast phylogenomic tree was constructed from the concatenated trimmed alignments using IQ‐TREE with the C60 + LG + GAMMA model and 1000 bootstrap replicates.
The plastid 16S rRNA gene (SSU) was recovered using barrnap v0.9 and combined with other apicoplast‐derived SSUs. Sequences between 500 and 15,000 bp were sorted and clustered at 99.5% using VSEARCH v2.25.0 (Rognes et al. 2016). They were then aligned using MAFFT v7.520 and trimmed using Trimal v1.4.rev15 under the following conditions: a gap threshold of 30%, a similarity threshold of 0.001, and a minimum conservation of 30%. ModelTest‐NG v0.1.7 was run on the trimmed alignment to determine the optimal substitution model for phylogenetic reconstruction. Then, a maximum‐likelihood tree was built with RAxML v8.2.13, using the General Time Reversible model with a GAMMA+P‐Invar model of rate heterogeneity (‘‐m GTRGAMMAIX’) and 1000 bootstrap replicates.
2.5. Microbiome Screening and Evolutionary Placement Analysis
The apicoplast SSU of the sea cucumber apicomplexan was searched against the IMNGS server (a massive, continuously updated database of 16S rRNA gene metabarcoding studies; Lagkouvardos et al. 2016) for any sequences with > 90% similarity. Matching sequences were then evolutionarily mapped to the apicoplast SSU tree constructed earlier using RAxML's evolutionary placement algorithm. The General Time Reversible model with a gamma‐distributed rate variation (GTR + CAT) and a minimum bootstrap support threshold of 20% were used to generate the EPA placement tree. Those sequences that clustered with the sea cucumber parasite were considered as likely the same parasite (or cyst stage of the parasite). A small selection of additional studies spanning other echinoderms and ocean regions was screened (Table S1) for the sea cucumber parasite apicoplast SSU using the same approach to ensure a comprehensive microbiome search.
3. Results
The complete rrn operon could be recovered from the metagenomic assembly of the recovered cysts. Phylogenetic analysis using this recovered rrn operon places the sea cucumber parasite, hereafter referred to as Apostichocystis gudetama, as sister to the clade, which includes the corallicolids and ichthyocolids (+90% bootstrap; Figure 1). This clade of strictly marine parasites rests sister to the Eimeriorina suborder of Coccidia (Figure 1). The protococcidians Eleutheroschizon spp. branch basal to those clades (Figure 1).
Contigs corresponding to organelle genomes were successfully recovered for A. gudetama. The complete 6.74 kb mitochondrial genome, containing a potentially trans‐spliced cox1 gene (Figure 2), was assembled with Novoplasty using a mitochondrial contig identified from the metagenomic assembly as a seed. A multi‐gene tree incorporating cox1, cox3, and cob presents a contrary phylogeny to the rrn operon tree with A. gudetama branching sister to E. duboscqui, although with weaker support (87% bootstrap; Figure 2). Together, this clade is sister to coccidians (Figure 2). A fragmented apicoplast genome assembly (22.39 kb) was recovered, containing the following genes: rpoB, sufB, rps4, rnl, rns, rpl4, rpl2, rps19, tufA, rps12, rps11, rpl36, rps5, rpl6, rps8 (Figure 3A). A 31‐gene apicoplast phylogeny generated using this new genetic information showed A. gudetama once again branching sister to the protococcidian E. duboscqi with stronger support than the mitochondrial gene tree (95% bootstrap; Figure 3B). Notably, the corallicolids and ichthyocolids branch distinctly away from A. gudetama and coccidians in this phylogeny (Figure 3B). Taking the SSU of the apicoplast and generating a phylogenetic tree incorporating other recovered apicoplast SSUs as well as environmental sequences correlating to SSU from apicomplexan‐related lineages (ARLs) shows A. gudetama branching deep within Apicomplexa and not correlating to any known ARL (Figure 3C). It still branches near E. duboscqi with sufficient surrounding bootstrap support (Figure 3C). Searching the IMNGS server, which contains operational taxonomic unit (OTU) data from thousands of metabarcoding studies, for any sequences with 90% similarity to the apicoplast SSU derived from the sea cucumber parasite revealed 5 closely related OTUs with low overall abundance in the environment. Of these, 4 OTUs were mapped onto the branch with A. gudetama's apicoplast SSU using an evolutionary placement algorithm (Figure 4A). All 4 OTUs originate from marine ecosystems around Australia. 2 OTUs (representing 4 total sequences) were recovered from the encrusting coral, Plesiastrea versipora , found near Sydney, Australia. The other 2 OTUs (representing 2 total sequences) were retrieved from marine waters around Rottnest Island, Australia (Figure 4B).
FIGURE 2.
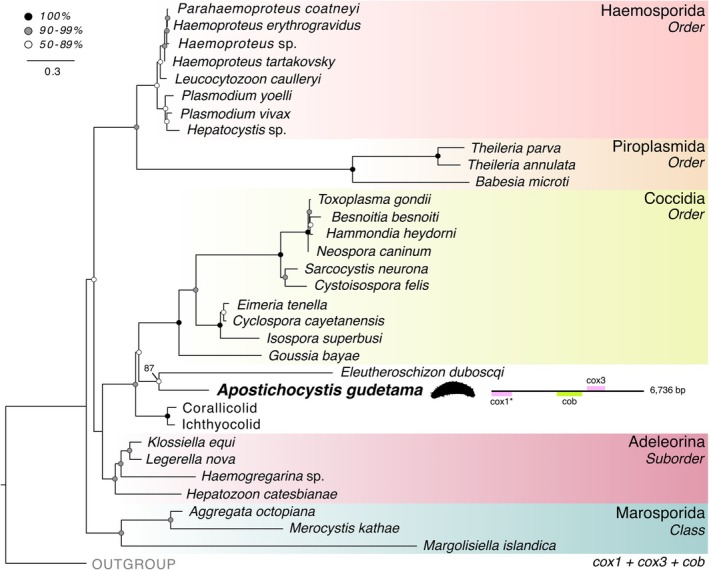
Maximum‐likelihood tree of apicomplexan mitochondria based on cox1, cox3, and cob using 1113 amino acid sites using IQ‐TREE with the C60 + LG + GAMMA model and 1000 bootstrap replicates. Major clades of Apicomplexa and related lineages are color‐coded. Stramenopiles were used as the outgroup. The gene arrangement of the A. gudetama mitochondrial genome alongside its placement on the tree. (See also Supporting Information S1B).
FIGURE 3.
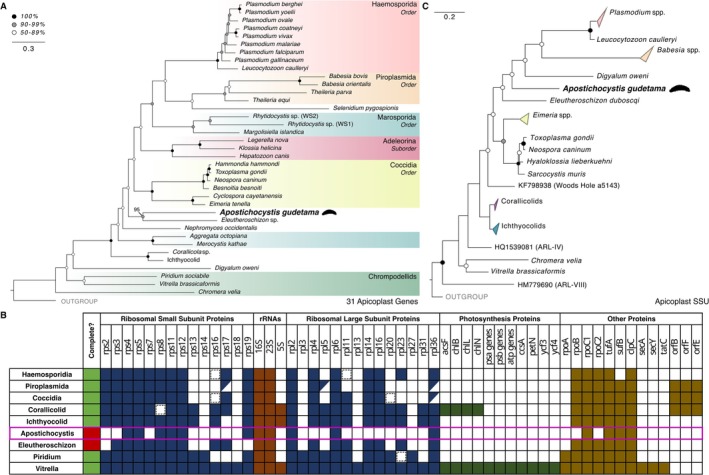
(A) Maximum‐likelihood tree of apicomplexans based on 31 plastid‐encoded genes and 3908 amino acid sites using IQ‐TREE with the C60 + LG + GAMMA model and 1000 bootstrap replicates. Major clades of Apicomplexa and related lineages are color‐coded. Rhodophyte plastids were used as the outgroup. (See also Supporting Information S1C). (B) Plastid gene content of apicomplexans and related lineages, including incomplete assemblies. Variable genes are represented with a slash, while a dashed box represents remnant genes. (C) Maximum‐likelihood tree of Apicomplexa using plastid 16S rRNA gene (16S) based on 1510 nucleic acid sites using RAxML GTR substitution matrix with GAMMA+P‐Invar model of rate heterogeneity and 1000 bootstrap replicates. Rhodophyte plastid 16S rRNA genes were used as the outgroup. (See also Supporting Information S1D).
FIGURE 4.
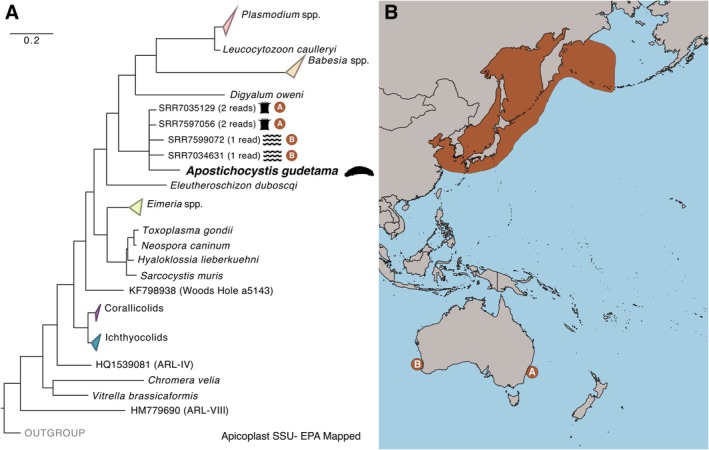
(A) Maximum‐likelihood tree of Apicomplexa using plastid 16S rRNA genes (16S) from Figure 3, with sequences from the Integrated Microbial Next Generation Sequencing (IMNGS) database with 90% similarity results placed to the A. gudetama apicoplast 16S rRNA gene with RAxML's evolutionary placement algorithm (EPA). (B) The geographic distribution of A. japonicus (orange shading) in comparison to the origins of the EPA‐mapped, likely Apostichocystis spp. operational taxonomic units (OTUs), which are denoted with (A) and (B).
4. Discussion
Through deep genomic sequencing of apicomplexan cysts originating from the ovaries of the Japanese sea cucumber ( A. japonicus ), we were able to recover additional genetic information to reveal a rather complicated phylogenetic placement of this commercially relevant parasite, which we have named Apostichocystis gudetama. All phylogenies originating from organelle data consistently show A. gudetama as closely related to but distinct from E. duboscqi (Figures 2 and 3), a protococcidian that infects marine polychaete worms (Valigurová et al. 2015). However, due to the fast‐evolving nature of AT%‐rich apicoplast genomes (Muñoz‐Gómez et al. 2019), along with the more limited taxonomic sampling of deep‐branching apicoplasts, the usefulness of plastid‐encoded genes for resolving phylogenetic relationships within Apicomplexa is limited (Mathur et al. 2021). On the other hand, the rrn operon phylogeny, which includes the broadest taxonomic sampling and more closely matches apicomplexan nuclear‐gene phylogenies (Mathur et al. 2023), shows support for a close relation of A. gudetama to the clade of ichthyocolids and corallicolids (Figures 1 and 2). Given the recent findings that these three clades (Eleutheroschizon spp., corallicolids, and ichthyocolids) are closely related to each other and branch together in phylogenomic trees (Jacko‐Reynolds et al. 2025), likely all as protococcidians, it is safe to assume that Apostichocytsis spp. likely branches within this clade as well. Together, these parasites encompass ecologically important apicomplexans, all found exclusively within marine hosts. Marine parasite host‐divergence in the ocean and the genetic underpinnings of parasite evolution can be investigated within this lineage thanks to this expanded phylogenetic framework.
Diversification of apicomplexan lineages within marine hosts is common, as shown by the vast diversity of gregarines that infect marine hosts (Odle et al. 2024). The ichthyocolids and corallicolids are globally distributed, appearing in hosts from cold, upper latitudes and warm, tropical latitudes (Bonacolta et al. 2024; Trznadel et al. 2024), exemplifying the broad host‐specific diversification common among apicomplexans (Morrison 2009). Relatedly, several sea cucumber‐infecting apicomplexans have been described morphologically but lack molecular data (Massin et al. 1977; Fankboner and Cameron 1985; Jangoux 1986; Pomory and Lares 1998; Eeckhaut et al. 2004; Mueller 2017). Targeted sequencing of these parasites, as well as the gonads of other echinoderms, may lead to the characterization or expansion of the Apostichocystis clade and other existing apicomplexan lineages, as well as our understanding of the diversity of parasites in the ocean. To this end, the detection of Apostichocystis‐related sequences in coral reefs and marine waters of Australia outside the known distribution of A. japonicus (Figure 4) hints at the possibility that other closely related apostichocystids may not be described.
Little is known about the biology and ecology of these closely related lineages of marine apicomplexan parasites. Known host‐specificity varies, with A. gudetama and E. dubosqi only being described in association with a singular host ( A. japonicus and Scoloplos armiger , respectively; Unuma et al. 2020; Valigurová et al. 2015). Corallicolids and ichthyocolids have much broader host ranges with corallicolids being recovered in the majority of sampled anthozoans as well as fireworms (Kwong et al. 2019; Bonacolta et al. 2025), and ichthyocolids being recovered from a variety of marine fish as well as gnathiids (Bonacolta et al. 2024; Sikkel et al. 2020). Morphologically, there is a large difference in size between A. gudetama and the rest of the mentioned apicomplexans. Gamont size is approximately 8 μm in ichthyocolids and 20 μm in E. dubosqi (Bonacolta et al. 2024; Valigurová et al. 2015). Corallicolids are approximately ~10 μm (Kwong et al. 2019). All much smaller than the 250 μm long cysts that A. gudetama forms. Regarding parasite development, A. gudetama grows and divides within the cytoplasm of the host cell, while E. dubosqi develops epicellularly.
Nuclear phylogenomics of Apostichocystis spp. could further support the Protococcidia as a marine‐host‐specific clade of apicomplexan parasites and shed light on our understanding of how the chlorophyll biosynthesis pathway has evolved within Apicomplexa. This pathway is hypothesized to have evolved unparsimoniously (Bonacolta et al. 2024; Jacko‐Reynolds et al. 2025), having been lost multiple times independently throughout apicomplexan evolution. Corallicolids and Eleutheroschizon have retained elements of the chlorophyll biosynthesis pathway from their apicoplast, while all other apicomplexans, including the closely related ichthyocolids, have independently lost this pathway (Jacko‐Reynolds et al. 2025). Future work on A. gudetama should focus on collecting additional nuclear and plastid gene data to determine the exact placement of this lineage in relation to these other lineages and to see whether chlorophyll biosynthesis genes have been lost or retained. This would provide further information as to the mysterious functional role of this pathway in these lineages and the gene loss and co‐option underpinning parasite evolution within Apicomplexa as a whole.
5. Taxonomic Summary
Phylum Apicomplexa Levine 1980, emend. Adl et al. 2005
Class Conoidasida Levine 1988.
Order Conoidasida IS.
Family Apostichocystidae fam. nov. Bonacolta et al. 2025
Genus Apostichocystis gen. nov. Bonacolta et al. 2025
Species Apostichocystis gudetama sp. nov. Bonacolta et al. 2025
Apostichocystidae fam. nov. Bonacolta et al. 2025
Zoobank ID: 7F2E916A‐A9DA‐433A‐9134‐FEE17739C3EF.
Included taxa: Apostichocystis gudetama sp. nov.
Diagnosis: A distinct clade within Apicomplexa based on rrn operon, mitochondrial, and plastid (apicoplast) gene sequences. All recovered genetic data in this family have been recovered from the marine environment, with the only described species thus far being found exclusively within the ovaries of the Japanese sea cucumber ( A. japonicus ).
Type genus: Apostichocystis gen. nov.
Apostichocystis gen. nov. Bonacolta et al. 2025
Zoobank ID: A4ABB7B3‐C69F‐4F6F‐B291‐4DB0E2C7CD5C.
Etymology: N.L. n. Apostichopus, referencing the host species Apostichopus japonicus , and Gr. prefix apo‐ meaning “away from” or “separate,” combined with Gr. adj. sticho‐ meaning “orderly” or “arranged,” reflecting both the association with Apostichopus and the phylogenetic placement of the genus as potentially “disorderly” based on the inconsistencies between the rrn gene and organelle phylogenomics. N.L. n. cystis, from Gr. n. kystis, meaning “bladder” or “sac,” commonly used in the names of cyst‐forming parasitic or symbiotic protists. Together, the name conveys the organism's biological and phylogenetic context.
Diagnosis: A distinct clade within Apicomplexa based on rrn operon, mitochondrial, and plastid (apicoplast) gene sequences.
Type species: Apostichocystis gudetama sp. nov.
Apostichocystis gudetama sp.
nov. Bonacolta et al. 2025
Zoobank ID: 15BFD54E‐478C‐41DD‐99FB‐4879AA354369.
Etymology: N.L. n. gudetama, inspired by the Japanese pop culture figure Gudetama, a character resembling a lazy egg, referencing the parasite's association with the ovaries of the Japanese sea cucumber ( Apostichopus japonicus ). The name reflects both the reproductive context of its host and the cultural connection to Japan.
Diagnosis: An unknown life stage invades the oocytes of female A. japonicus, where it forms ~2 1950 μm multi‐cell cysts surrounded by a 50 μm transparent layer during host maturation. The sporozoites/merozoites (~40 μm in length) are then shed with the eggs from the gonopore during host spawning.
Type sequence: 18S rRNA gene sequence (PV105649) and 28S rRNA gene sequence (PV105650).
Type habitat: Marine; host‐associated.
Type host: Apostichopus japonicus Selenka, 1867 (Metazoa; Echinodermata; Holothuroidea; Synallactida; Stichopodidae).
Type material: Cysts preserved in ethanol are stored at Tohoku University under the supervision of Dr. Tatsuya Unuma.
Conflicts of Interest
The authors declare no conflicts of interest.
Supporting information
Data S1.
Table S1.
Acknowledgments
This study was supported by startup funds from the University of Miami Rosenstiel School of Marine, Atmospheric, and Earth Sciences, project PID2020‐118836GA‐I00 financed by MICIU/AEI/10.13039/501100011033, and project 2021 SGR 00420 financed by the Departament de Recerca i Universitats de la Generalitat de Catalunya.
Funding: This work was supported by University of Miami, Ministerio de Ciencia, Innovación y Universidades, PID2020‐118836GA‐I00; Departament de Recerca i Universitats de la Generalitat de Catalunya, 2021 SGR 00420.
Data Availability Statement
The data that support the findings of this study are openly available in NCBI SRA at https://www.ncbi.nlm.nih.gov/sra, reference number PRJNA1227500.
References
- Adl, S. M. , Simpson A. G., Farmer M. A., et al. 2005. “The New Higher Level Classification of Eukaryotes With Emphasis on the Taxonomy of Protists.” Journal of Eukaryotic Microbiology 52: 399–451. [DOI] [PubMed] [Google Scholar]
- Baños, H. , Susko E., and Roger A. J.. 2024. “Is Over‐Parameterization a Problem for Profile Mixture Models?” Systematic Biology 73: 53–75. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bell, T. A. , and Lightner D. V.. 1988. “A Handbook of Normal Penaeid Shrimp Histology.” World Aqwuaculture Society.
- Bonacolta, A. M. , Krause‐Massaguer J., Smit N. J., Sikkel P. C., and del Campo J.. 2024. “A New and Widespread Group of Fish Apicomplexan Parasites.” Current Biology 34: 2748–2755. [DOI] [PubMed] [Google Scholar]
- Bonacolta, A. M. , Weiler B. A., Grimes C. J., et al. 2025. “Fireworms Are a Reservoir and Potential Vector for Coral‐Infecting Apicomplexans.” ISME Journal 19: wraf078. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Camacho, C. , Coulouris G., Avagyan V., et al. 2009. “BLAST+: Architecture and Applications.” BMC Bioinformatics 10: 421. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Capella‐Gutiérrez, S. , Silla‐Martínez J. M., and Gabaldón T.. 2009. “trimAl: A Tool for Automated Alignment Trimming in Large‐Scale Phylogenetic Analyses.” Bioinformatics 25: 1972–1973. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen, S. , Zhou Y., Chen Y., and Gu J.. 2018. “Fastp: An Ultra‐Fast All‐In‐One FASTQ Preprocessor.” Bioinformatics 34: i884–i890. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Darriba, D. , Posada D., Kozlov A. M., Stamatakis A., Morel B., and Flouri T.. 2020. “ModelTest‐NG: A New and Scalable Tool for the Selection of DNA and Protein Evolutionary Models.” Molecular Biology and Evolution 37: 291–294. [DOI] [PMC free article] [PubMed] [Google Scholar]
- del Campo, J. , Heger T. J., Rodríguez‐Martínez R., et al. 2019. “Assessing the Diversity and Distribution of Apicomplexans in Host and Free‐Living Environments Using High‐Throughput Amplicon Data and a Phylogenetically Informed Reference Framework.” Frontiers in Microbiology 10: 2373. 10.3389/fmicb.2019.02373. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dierckxsens, N. , Mardulyn P., and Smits G.. 2017. “NOVOPlasty: de Novo Assembly of Organelle Genomes From Whole Genome Data.” Nucleic Acids Research 45: e18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Edgar, R. C. 2004. “MUSCLE: Multiple Sequence Alignment With High Accuracy and High Throughput.” Nucleic Acids Research 32: 1792–1797. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Eeckhaut, I. , Parmentier E., Becker P., da Gomez Silva S., and Jangoux M.. 2004. “Parasites and Biotic Diseases in Field and Cultivated Sea Cucumbers.” In Advances in sea cucumber aquaculture and management, edited by Lovatelli A., Conand C., Purcell S. et al., 463. FAO Fisheries Technical Paper. [Google Scholar]
- Fankboner, P. V. , and Cameron J. L.. 1985. “Seasonal Atrophy of the Visceral Organs in a Sea Cucumber.” Canadian Journal of Zoology 63: 2888–2892. [Google Scholar]
- Jacko‐Reynolds, V. K. L. , Kwong W. K., Livingston S. J., et al. 2025. “Phylogenomics of Coral‐Infecting Corallicolids Reveal Multiple Independent Losses of Chlorophyll Biosynthesis in Apicomplexan Parasites.” Current Biology 35: 1156–1163.e4. [DOI] [PubMed] [Google Scholar]
- Jangoux, M. 1986. “Diseases of Echinodermata. I. Agents Microorganisms and Protistans.” Diseases of Aquatic Organisms—DISEASE AQUAT ORG 2: 147–162. 10.3354/dao002147. [DOI] [Google Scholar]
- Janouškovec, J. , Paskerova G. G., Miroliubova T. S., et al. 2019. “Apicomplexan‐Like Parasites Are Polyphyletic and Widely but Selectively Dependent on Cryptic Plastid Organelles.” eLife 8: e49662. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Janouškovec, J. , Horák A., Rohwer F., and Keeling P. J.. 2012. “Global Analysis of Plastid Diversity Reveals Apicomplexan‐Related Lineages in Coral Reefs.” Current Biology 22: R518–R519. [DOI] [PubMed] [Google Scholar]
- Jin, J.‐J. , Yu W.‐B., Yang J.‐B., et al. 2020. “GetOrganelle: A Fast and Versatile Toolkit for Accurate de Novo Assembly of Organelle Genomes.” Genome Biology 21: 241. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Katoh, K. , and Standley D. M.. 2013. “MAFFT Multiple Sequence Alignment Software Version 7: Improvements in Performance and Usability.” Molecular Biology and Evolution 30: 772–780. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kwong, W. K. , del Campo J., Mathur V., Vermeij M. J., and Keeling P. J.. 2019. “A Widespread Coral‐Infecting Apicomplexan With Chlorophyll Biosynthesis Genes.” Nature 568, no. 7750: 103–107. [DOI] [PubMed] [Google Scholar]
- Lagkouvardos, I. , Joseph D., Kapfhammer M., et al. 2016. “IMNGS: A Comprehensive Open Resource of Processed 16S rRNA Microbial Profiles for Ecology and Diversity Studies.” Scientific Reports 6: 33721. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lang, B. F. , Beck N., Prince S., Sarrasin M., Rioux P., and Burger G.. 2023. “Mitochondrial Genome Annotation With MFannot: A Critical Analysis of Gene Identification and Gene Model Prediction.” Frontiers in Plant Science 14: 1222186. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li, D. , Liu C.‐M., Luo R., Sadakane K., and Lam T.‐W.. 2015. “MEGAHIT: An Ultra‐Fast Single‐Node Solution for Large and Complex Metagenomics Assembly via Succinct de Bruijn Graph.” Bioinformatics 31: 1674–1676. [DOI] [PubMed] [Google Scholar]
- Massin, C. , Jangoux M., and Sibuet M.. 1977. “Description D'ixoreis Psychropotae, Nov. Gen., nov. sp., Coccidie Parasite du Tube Digestif de L'holothurie Abyssale Psychropotes Longicauda Theel.” Protist 14: 253–259. [Google Scholar]
- Mathur, V. , Kolísko M., Hehenberger E., et al. 2019. “Multiple Independent Origins of Apicomplexan‐Like Parasites.” Current Biology 29: 2936–2941.e5. [DOI] [PubMed] [Google Scholar]
- Mathur, V. , Kwong W. K., Husnik F., et al. 2021. “Phylogenomics Identifies a New Major Subgroup of Apicomplexans, Marosporida Class nov., With Extreme Apicoplast Genome Reduction.” Genome Biology and Evolution 13: evaa244. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mathur, V. , Salomaki E. D., Wakeman K. C., et al. 2023. “Reconstruction of Plastid Proteomes of Apicomplexans and Close Relatives Reveals the Major Evolutionary Outcomes of Cryptic Plastids.” Molecular Biology and Evolution 40: msad002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McFadden, G. I. , and Yeh E.. 2017. “The Apicoplast: Now You See It, Now You Don't.” International Journal for Parasitology 47: 137–144. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Morrison, D. A. 2009. “Evolution of the Apicomplexa: Where Are We Now?” Trends in Parasitology 25: 375–382. [DOI] [PubMed] [Google Scholar]
- Mueller, K. W. 2017. “Symbionts of the Giant Red Sea Cucumber, Parastichopus Californicus (Stimpson, 1857), With Some Implications for Culture of the Host.” SPC Beche‐de‐Mer Information Bulletin 37: 79–91. [Google Scholar]
- Muñoz‐Gómez, S. A. , Durnin K., Eme L., et al. 2019. “Nephromyces Represents a Diverse and Novel Lineage of the Apicomplexa That Has Retained Apicoplasts.” Genome Biology and Evolution 11: 2727–2740. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nguyen, L.‐T. , Schmidt H. A., von Haeseler A., and Minh B. Q.. 2015. “IQ‐TREE: A Fast and Effective Stochastic Algorithm for Estimating Maximum‐Likelihood Phylogenies.” Molecular Biology and Evolution 32: 268–274. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Odle, E. , Riewluang S., Ageishi K., Kajihara H., and Wakeman K. C.. 2024. “Pacific Marine Gregarines (Apicomplexa) Shed Light on Biogeographic Speciation Patterns and Novel Diversity Among Early Apicomplexans.” European Journal of Protistology 94: 126080. [DOI] [PubMed] [Google Scholar]
- Pomory, C. M. , and Lares M. T.. 1998. “Gonospora Holoflora: A New Species of Gregarine Protozoan Parasite (Apicomplexa) in Holothuria floridana (Echinodermata: Holothuroidea) From the Florida Keys.” Bulletin of Marine Science 62: 213–218. [Google Scholar]
- Robinson, G. , and Lovatelli A.. 2015. “Global Sea Cucumber Fisheries and Aquaculture FAO'S Inputs Over the Past Few Years.” FAO Aquaculture Newsletter 53: 55–57. [Google Scholar]
- Rognes, T. , Flouri T., Nichols B., Quince C., and Mahé F.. 2016. “VSEARCH: A Versatile Open Source Tool for Metagenomics.” PeerJ 4: e2584. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sakai, Y. 2015. “Mass Production of Artificial Seed of the Japanese Common Sea Cucumber (Apostichopus japonicus) in Hokkaido, Japan.” Bulletin of the Fisheries Research Agency 40: 129–134. [Google Scholar]
- Seemann, T. 2018. “Barrnap: BAsic Rapid Ribosomal RNA Predictor.” https://github.com/tseemann/barrnap.
- Sikkel, P. C. , Pagan J. A., Santos J. L., Hendrick G. C., Nicholson M. D., and Xavier R.. 2020. “Molecular Detection of Apicomplexan Blood Parasites of Coral Reef Fishes From Free‐Living Stages of Ectoparasitic Gnathiid Isopods.” Parasitology Research 119: 1975–1980. [DOI] [PubMed] [Google Scholar]
- Stamatakis, A. 2014. “RAxML Version 8: A Tool for Phylogenetic Analysis and Post‐Analysis of Large Phylogenies.” Bioinformatics 30: 1312–1313. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Trznadel, M. , Holt C. C., Livingston S. J., Kwong W. K., and Keeling P. J.. 2024. “Coral‐Infecting Parasites in Cold Marine Ecosystems.” Current Biology 34: 1810–1816.e4. [DOI] [PubMed] [Google Scholar]
- Unuma, T. , Tsuda N., Sakai Y., et al. 2020. “Coccidian Parasite in Sea Cucumber ( Apostichopus japonicus ) Ovaries.” Biological Bulletin 238: 64–71. [DOI] [PubMed] [Google Scholar]
- Valigurová, A. , Paskerova G. G., Diakin A., Kováčiková M., and Simdyanov T. G.. 2015. “Protococcidian Eleutheroschizon Duboscqi, an Unusual Apicomplexan Interconnecting Gregarines and Cryptosporidia.” PLoS One 10: e0125063. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vences, M. , Patmanidis S., Kharchev V., and Renner S. S.. 2022. “Concatenator, a User‐Friendly Program to Concatenate DNA Sequences, Implementing Graphical User Interfaces for MAFFT and FastTree.” Bioinformatics Advances 2: vbac050. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wheeler, T. J. , and Eddy S. R.. 2013. “Nhmmer: DNA Homology Search With Profile HMMs.” Bioinformatics 29: 2487–2489. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang, H. , Hamel J.‐F., and Mercier, eds A.. 2015. The Sea Cucumber Apostichopus japonicus : History, Biology and Aquaculture (Vol. 39). Academic Press. [Google Scholar]
- Yu, Z. , Zhou Y. I., Yang H., and Hu C.. 2014. “Survival, Growth, Food Availability and Assimilation Efficiency of the Sea Cucumber Apostichopus japonicus Bottom‐Cultured Under a Fish Farm in Southern China.” Aquaculture 426: 238–248. [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data S1.
Table S1.
Data Availability Statement
The data that support the findings of this study are openly available in NCBI SRA at https://www.ncbi.nlm.nih.gov/sra, reference number PRJNA1227500.


