Abstract
A novel Cas13d-based gene-editing approach has been developed to target viral RNAs in infected cells, reducing the replication of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) and Zika virus (ZIKV) by up to 90% compared with controls. Despite its potential, the use of Cas13d as an antiviral faces several challenges that limit its effectiveness before reaching target cells. This study presents a proof-of-concept strategy for constructing Cas13d with gold nanoparticles (Au_NPs) to destroy SARS-CoV-2 and ZIKV genomes into cells. The Au_NPs Cas13d complexes were administered to Huh-7 cells infected with either virus, in single or multiple doses. The study demonstrated that Au_NPs Cas13d cuts target RNAs with comparable efficiency as lipofected ribonucleoprotein (RNP). Additionally, we found that Au_NPs Cas13d can spontaneously enter cells by endocytosis or diffusion, before the first 4 h of treatment. Au_NPs Cas13d co-localized with SARS-CoV-2 virions in early endosomes and reduced SARS-CoV-2 replication after a single administration, unlike RNPs, which showed no antiviral activity. However, Au_NPs Cas13d was less efficient at reducing ZIKV replication compared with lipofected Cas13d-RNPs, likely due to different intracellular localization. These results suggest that Au_NPs can be adapted as a new antiviral strategy, highlighting an innovative delivery method of Cas13d against viruses without the need for transfecting, providing a new gene-editing-based approach against emerging RNA viruses.
Keywords: MT: RNA/DNA Editing, SARS-CoV-2, Zika virus, Cas13d, gold nanoparticles, Au_NPs Cas13d, emerging viruses, RNA viruses
Graphical abstract

Lai and colleagues show that the administration of Au_NPs Cas13d results in the spontaneous entry of the enzyme into cells, overcoming limitations of current strategies. The co-localization with SARS-CoV-2 and ZIKV in the early endosome, albeit with different antiviral efficacy, highlights the importance of the nanoparticle entry route against viruses.
Introduction
Since December 2019, the global community has faced significant challenges due to the coronavirus disease 2019 pandemic, caused by the RNA virus known as severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2).1 Alongside SARS-CoV-2, outbreaks of various RNA viruses in recent years, such as Zika (ZIKV), Ebola, Marburg, and Middle East respiratory syndrome viruses, have prompted the World Health Organization to declare these public health emergencies. While vaccines and treatment options against SARS-CoV-2 have progressed to market, most RNA viruses remain without approved therapies.1,2,3,4,5,6,7
In search of potential antivirals, several researchers highlighted gene editing as a novel weapon against emerging viruses. Indeed, Cas, initially recognized for its role in prokaryotic adaptive immunity against bacteriophages, has emerged as a potential therapeutic tool to combat viral diseases such as HIV-1, human papillomavirus, and herpes simplex virus-1.8,9,10,11,12,13,14 While CRISPR-Cas9 has been shown to be effective against viruses that produce double-stranded DNA (dsDNA) intermediates in their replication, its utility against RNA viruses like SARS-CoV-2, ZIKV, and Dengue virus has been limited.15 However, the discovery of CRISPR-Cas13, a ribonuclease capable of targeting and cleaving single-stranded RNA molecules, offers new prospects for intervention against RNA viruses.15,16,17 CRISPR-Cas13d has demonstrated efficacy in suppressing SARS-CoV-2, ZIKV, and influenza A virus replication in cell culture, suggesting its potential for mutagenizing critical genomic elements or degrading viral RNAs.18,19 This highlights the promise of CRISPR-based approaches for combating RNA virus infections, with a gene-editing weapon that can be easily adapted to emerging RNA viruses with pandemic potential.
Recently, Lin et al.20 conducted a comprehensive in silico analysis utilizing CRISPR-Cas13 to target prominent pathogenic viral species and 16 families of human or animal RNA viruses within the ViPR database (www.viprbrc.org). They observed that a mini-pool of 14 CRISPR RNAs (crRNAs) can flexibly target 10 families of human viruses with greater than 90% coverage. This platform, called PAC-MAN, may serve as a broad-spectrum antiviral strategy for viruses infecting humans and domestic livestock in the next years. Most notably, some of the crRNAs designed by Abbott et al.18 effectively target all sequenced new SARS-CoV-2 strains to date. Indeed, the Cas13 enzyme can utilize crRNA mini-pools to simultaneously target different viral genomic regions in the same administration. Unfortunately, the translation to human application will require addressing technical challenges, such as Cas13d delivery and immunogenicity.18,20,21,22,23,24 Moreover, for broad-spectrum targeting, it remains uncertain whether the current CRISPR-Cas13d system can effectively handle the scale of 14 crRNAs simultaneously for efficient knockdown of each target.
The present work explores a new delivery system of Cas13d using Au_NPs capable of spontaneously entering cells without any transfectants while maintaining the same level of Cas13d activity as by transfection of its canonical ribonucleoprotein (RNP). The selection of Au_NPs as a delivery system was based on several advantages. First, they are biocompatible due to their shape, size, and charge. Second, their plasmonic surface enables the use of the photothermal effect,25,26,27,28 where they release heat when exposed to a specific wavelength of light. Because of these properties, Au_NPs are already widely used in research, diagnostics, and therapy.25,26,27,28,29 Additionally, they can be aerosolized, opening up new possibilities for combating respiratory viruses.30,31,32 Furthermore, Au_NPs easily enter cells through various pathways like endocytosis and diffusion, based on their protein coating, size, and charge, offering a safer and more efficient alternative to other transfection methods like electroporation, which can be toxic, costly, and invasive.33,34,35,36,37 Finally, their surface can be customized with a range of molecules, such as drugs, enzymes, and antibodies.28,38,39,40,41,42
We tested Au_NPs against SARS-CoV-2 and ZIKV, two RNA viruses that use the same entry mechanism but replicate differently in the cytosol.43,44,45,46,47,48,49 Our data show that Au_NPs with Cas13d co-localize with both viruses in early endosomes. Additionally, a single dose of Au_NPs with Cas13d significantly reduced SARS-CoV-2 replication, while RNP administration showed no antiviral activity. However, Au_NPs with Cas13d were much less effective in reducing ZIKV replication compared with lipofected Cas13d RNPs. This difference may be due to Au_NPs being able to reach the replication sites of SARS-CoV-2 but not the viroplasms where ZIKV replicates.
Results
Au_NPs Cas13d functionalization does not affect Au_NPs dynamic light scattering and their Z-potential
We analyzed the Au_NPs dynamic light scattering (DLS) and Z-potential before and after conjugation with Cas13d to assess whether the functionalization process influenced the nanoparticles properties. As shown in Figure S1, the naked Au_NPs exhibited a DLS of 25.37 ± 10.67 nm and a Z-potential of −22.6 ± 4.54 mV (Figures S1A and S1B). Following the conjugation protocol, these values shifted to 23.76 ± 11.04 nm and −24.3 ± 8.48 mV, respectively (Figures S1A and S1B).
Guide RNA selection against SARS-CoV-2 and ZIKV RNA genomes
The choice of an effective guide RNA (gRNA) is crucial for the optimal function of CRISPR-Cas systems, particularly when targeting rapidly mutating viral RNA genomes. To identify the most effective gRNAs, we analyzed the most conserved regions of the SARS-CoV-2 and ZIKV genomes, as illustrated in Figure S2. For ZIKV, we selected four gRNAs targeting the capsid protein (C), non-structural protein 1 (NS1), NS4A, and NS4B. For SARS-CoV-2, we focused on the RNA-dependent RNA polymerase (RdRP), nucleocapsid (N), and NSP3. The selected gRNAs were cloned into lentiviral vectors and transduced into Huh-7 cells, as shown in Figure 1A. Cas13b-expressing Huh-7 cells were then infected with either SARS-CoV-2 or ZIKV at a multiplicity of infection (MOI) of 1, and viral genomes were quantified in both the supernatants (released virions) and cell lysates (replicating genomes) at 24 h post infection (Figures 1B and 1C).
Figure 1.
Assessing the best gRNAs against SARS-CoV-2 and ZIKV
(A) Representative illustration of the experimental workflow. Briefly, eight lentiviral vectors were produced and used to obtain stable cell lines expressing a single gRNA and Cas13. Transduced cells were then infected with SARS-CoV-2 or ZIKV and analyzed by RT-qPCR or HCS. (B) On the left, RT-qPCR quantification of ZIKV genomes released in supernatants. On the right, RT-qPCR quantification on ZIKV viral genomes detected in cell lysates. Data are expressed as mean ± SD and analyzed by one-way ANOVA (∗∗∗∗p < 0.0001; multiple comparison Dunnett’s test: ∗p < 0.05, ∗∗p < 0.01, ∗∗∗p < 0.001, and ∗∗∗∗p < 0.0001). (C) On the left, RT-qPCR quantification of SARS-CoV-2 genomes released in supernatants. On the right, RT-qPCR quantification of SARS-CoV-2 viral genomes detected in cell lysates. Data are expressed as mean ± SD and analyzed by one-way ANOVA (∗∗∗∗p < 0.0001; multiple comparison Dunnett’s test: ∗p < 0.05, ∗∗p < 0.01, ∗∗∗p < 0.001, and ∗∗∗∗p < 0.0001). (D) Representative images of HCS of ZIKV-infected cells, transduced or not with the 4 gRNAs selected. Cells were stained for nuclei (DAPI, blue), F-actin (Phalloidin 488, green), and ZIKV (NS1, red). (E) Automated count of ZIKV infected cells on samples shown in (D). Data are expressed as mean ± SD and analyzed by one-way ANOVA (∗∗p < 0.01; multiple comparison Dunnett’s test: ∗p < 0.05, ∗∗p < 0.01, ∗∗∗p < 0.001, and ∗∗∗∗p < 0.0001). (F) Representative images of HCS on SARS-CoV-2-infected cells, transduced or not with the four gRNAs selected. Cells were stained for Nuclei (DAPI, blue), F-actin (Phalloidin 488, green), and SARS-CoV-2 (N protein, red). (F) Automated count of SARS-CoV-2-infected cells on samples shown in (G). Data are expressed as mean ± SD and analyzed by one-way ANOVA (∗∗p < 0.01; multiple comparison Dunnett’s test: ∗p < 0.05, ∗∗p < 0.01, ∗∗∗p < 0.001, and ∗∗∗∗p < 0.0001).
The most effective gRNAs were identified as NS1. However, we decided to maintain also the gRNA targeting C to assess the effect of the editing against a structural protein for ZIKV. RdRP and N were chosen for SARS-CoV-2, each reducing viral copies by approximately 1 log10 for each virus. We further validated these findings using high-content confocal imaging to automatically count the number of infected cells. Notably, a moderate 25% reduction in ZIKV yield was observed in cells transduced with the NS1 gRNA, compared with the untreated counterparts, while significant reductions of 90% and 80% were observed for SARS-CoV-2 when targeting RdRP and N, respectively (Figures 1D–1G).
Au_NPs Cas13d spontaneously penetrate cells following the endocytic pathway
Naked Au_NPs are well known to enter cells via various pathways, including endocytosis,50,51,52,53,54 which is also the primary entry route for ZIKV and SARS-CoV-2.43,47 Given this, we sought to investigate whether our Au_NPs Cas13d exploit this pathway efficiently. If both viruses and Au_NPs Cas13d utilize the same endocytic vesicles, this could potentially result in the rapid disruption of the viral genome early in the infection process. First, we evaluated cell viability by assessing the number of nuclei over time by performing high-content live confocal microscopy assays, as shown in Figure S3A. This assay demonstrated that Au_NPs Cas13d did not affect cell viability when administered to 109 NPs/mL roughly 14 h after administration. To evaluate the entry processes, we then functionalized Au_NPs Cas13d with Alexa 546-labeled gRNAs. As shown in Figures S3B and S3C, Cas13d enter cells after 15 min, without any differences depending on the delivery method used. Notably, we observed an increased amount of Cas13d delivered by Au_NPs after 4 h.
To better understand this difference, we stained the cells for EEA1, a well-established early endosome marker,55,56 to assess co-localization with Au_NPs Cas13d (Figures 2A and 2B). The analysis showed that Au_NPs Cas13d entered 78.5% of treated cells (Figure S4B) and co-localized with 21% of EEA1+ vesicles, which was three times more efficient than the standard RNP transfection method (7%), used as a reference control (Figure 2C). To further understand the intracellular distribution and availability of Au_NPs Cas13d in Huh-7 cells, we conducted a second high-content confocal analysis, tracking live cells for 14 h, starting 4 h after administration. This experiment revealed that 40% of the administered Au_NPs Cas13d successfully entered the cells. Additionally, 99.9% of the Au_NPs Cas13d that entered the cytosol remained at their maximum concentration for 12 h, 42 min, and 58 s post administration (Figures 2D and 2E).
Figure 2.
Au_NPs Cas13d co-localize with early endosomes in the cytoplasm
(A) Representative scheme of the high-content confocal 3D acquisition. Briefly, Huh-7-mGFP cells treated or not with Au_NPs Cas13d marked in orange (Alexa 546) were stained for nuclei (DAPI, blue) and early endosomes (EEA1-Alexa 647). Then cells were 3D reconstructed taking 1 image every 0.5 μm, for 30 μm. (B) Representative images of 3D reconstructed Huh-7 cells described in (A). (C) Statistical analysis of Au_NPs Cas13d and EEA1 co-localization. Data are expressed as mean ± SD and analyzed by one-way ANOVA (∗∗∗p < 0.001; multiple comparison Dunnett’s test: ∗p < 0.05, ∗∗p < 0.01, ∗∗∗p < 0.001, and ∗∗∗∗p <0.0001). (D, top) Representative image of the experimental workflow. The live imaging acquisition was performed for 18 h, taking a picture every 30 min. (Bottom) Representative images of the experiment described in (C). (E) Statistical analysis of Au_NPs Cas13d localization over time. Data are expressed as mean ± SD and analyzed by one-way ANOVA (∗∗∗∗p < 0.01; multiple comparison Tukey’s test: ∗p < 0.05, ∗∗p < 0.01, ∗∗∗p < 0.001, and ∗∗∗∗p < 0.0001).
To further investigate the cellular entry routes utilized by Au_NPs Cas13d, we systematically explored different mechanisms of uptake. First, we inhibited all energy-dependent pathways by reducing the temperature to 4°C, leaving passive diffusion as the only possible entry route. After 4 h, we observed a significant reduction in cellular uptake, with only 4 Au_NPs Cas13d per cell compared with 9 Au_NPs Cas13d per cell at 37°C. In our experimental settings, diffusion represents an approximately 50% of entry (Figures S4A–S4E). Next, we examined the specific pathways involved by using a series of inhibitors targeting distinct endocytic routes. As shown in Figures S4A and S4F, pre-treatment with nystatin or 7-keto-cholesterol—targeting caveolin-mediated endocytosis and clathrin- and dynamin-independent endocytosis (CLIC/GEEC), respectively—did not result in any significant reduction in Au_NPs Cas13d uptake compared with the untreated control.57 However, pre-treatment with chloroquine, Dynasore, or a combination of Dynasore and nystatin significantly reduced cellular uptake. These compounds inhibit clathrin-mediated endocytosis (CME), fast endophilin-mediated endocytosis (FEME), and macro-/micro-pinocytosis, respectively. Specifically, reductions of 35%, 83%, and 75% were observed, respectively, demonstrating that CME, FEME, and pinocytosis are the predominant pathways for Au_NPs Cas13d cellular entry by endocytosis (FigureS4F).57
Assessing the Au_NPs Cas13d cutting efficiency
After confirming that Au_NPs Cas13d can spontaneously enter cells, we evaluated whether their cleavage activity was affected by the nanoformulation. To do this, we compared the cleavage efficiency of a SARS-CoV-2 RNA fragment by Cas13d fused to Au_NPs or as canonical RNP formulation. We mixed either RNP or Au_NPs Cas13d with a SARS-CoV-2 RNA fragment at different concentrations, then reverse transcribed the digested RNA and analyzed it using RT-qPCR, as outlined in Figure S5A. In this assay, two primer sets were designed: the first amplified the region where Cas13d editing occurred, and an HEX-labeled fluorescent probe recognized the same target sequence as Cas13d. The second primer set amplified an unedited control sequence (FAM) to measure the total number of RNA fragments in the reaction. The results showed that Au_NPs Cas13d had the same cleavage efficiency (∼99.8%) as the standard RNP approach (Figure S5B). Furthermore, the optimal cleavage activity was observed at a 1 viral RNA:60 RNP ratio (Figure S5B).
Au_NPs Cas13d co-localizes with both ZIKV and SARS-CoV-2 during the entry phase
Once probed that Au_NPs Cas13d can spontaneously enter cells and efficiently cleave target RNA like standard RNPs, we investigated whether SARS-CoV-2 virions co-localize with Au_NPs Cas13d during viral entry. To test this, we infected Huh-7 cells with SARS-CoV-2 at a MOI of 100, with or without the addition of Au_NPs Cas13d at a 1:100 viral RNA to Cas13d ratio. Two hours post infection, the cells were fixed and analyzed using transmission electron microscopy (TEM), as shown in Figure 3A. TEM images (Figure 3B) revealed that SARS-CoV-2 virions were present inside endosomes, surrounded by Au_NPs Cas13d (visible as black spots). Interestingly, Au_NPs Cas13d co-localized with ZIKV only during the entry phase, prompting us to investigate why Au_NPs Cas13d was more effective against SARS-CoV-2 than ZIKV. We hypothesized that the difference in efficacy might be due to the nanoformulation’s inability to co-localize with ZIKV in its specific replication compartments. SARS-CoV-2 replicates in double-membrane vesicles, while ZIKV replicates in hijacked endoplasmic reticulum-Golgi compartments. To evaluate this, we infected Huh-7 cells with ZIKV at an MOI of 100, with or without Au_NPs Cas13d. Huh-7 cells were treated either during or 20 h post infection, as shown in Figure 4A. Initially, Au_NPs Cas13d was administered during ZIKV infection, and Huh-7 cells were fixed and processed for TEM 2 h post infection. As seen in Figure 4B, Au_NPs Cas13d (depicted as black dots) surrounded ZIKV virions within early endosomes. To further explore the intracellular localization, we repeated the experiment with Huh-7 cells infected with ZIKV at an MOI of 5. This time, cells were treated with Au_NPs Cas13d 20 h post infection and fixed for TEM imaging 24 h later. As shown in Figure 4B (right), ZIKV virions were found inside the Golgi cisternae, but only a few Au_NPs Cas13d were present. Most Au_NPs Cas13d were seen localized independently within the Golgi, not co-localized with the ZIKV virions.
Figure 3.
Au_NPs Cas13d co-localization with SARS-CoV-2 during the entry phase
(A) Representative scheme of TEM experiment. Briefly, Huh-7 cells were first infected with SARS-CoV-2 at 100 MOI, in the presence or not of Au_NPs Cas13d in a 1:100 ratio. Then cells were fixed and analyzed by TEM 2 h after infection. (B) Representative images taken by TEM, black circles highlight SARS-CoV-2 virions, Au_NPs Cas13d appear as black spots, co-localizing in the same vesicles containing SARS-CoV-2 virions.
Figure 4.
Au_NPs Cas13d do not co-localize with ZIKV virions during replication and egress
(A) Representative scheme of the TEM experiment. Briefly, Huh-7 cells were first infected with ZIKV at 100 MOI, in the presence or absence of Au_NPS Cas13d at a 1:100 ratio. Cells were then fixed and probed for TEM 2 h post infection. In parallel, to assess the co-localization during the replication-egress phase, we administered Au_NPs Cas13d 20 h post infection, fixing cells 24 h post treatment. (B) Representative images taken by TEM. Black circles highlight ZIKV virions while Au_NPs Cas13d appear as black spots.
Au_NPs Cas13d exerts stronger antiviral activity against SARS-CoV-2 than RNP delivery
To evaluate the effectiveness of Au_NPs Cas13d against SARS-CoV-2, we first infected Huh-7 cells with the virus at a MOI of 0.1. Au_NPs Cas13d was administered 1.5 h post infection. The supernatants were analyzed by RT-qPCR to measure the amount of SARS-CoV-2 virions released, and Huh-7 cells were probed for the SARS-CoV-2 N protein to assess the number of infected cells, as outlined in Figure 5A.
Figure 5.
Au_NPs Cas13d reduce SARS-CoV-2 replication
(A) Representative scheme of the experimental workflow. Briefly, cells were infected with SARS-CoV-2 at 1 MOI, then treated or not with Au_NPs or RNP Cas13d. Cell supernatants were then analyzed for virions released by RT-qPCR on or immunostaining, with the aim to automatically count the number of infected cells out of the total. (B) Statistical analysis of SARS-CoV-2 virion release after a single administration of Cas13d (RNP). Data are expressed as mean ± SD and analyzed by One-Way ANOVA: NS > 0.05. (C) Statistical analysis of SARS-CoV-2 virion release after a single administration of Au_NPs Cas13d. Data are expressed as mean ± SD and analyzed by One-Way ANOVA: NS > 0.05; Multiple comparison Dunnett’s test: ∗p < 0.05, ∗∗p < 0.01, ∗∗∗p < 0.001, and ∗∗∗∗p < 0.0001. (D) Representative images of HCS performed on Huh-7 cells treated or not with lipofected Cas13d. Cells were then infected with SARS-CoV-2 and stained for nuclei (DAPI, blue), F-actin (Phalloidin 488, green), and SARS-CoV-2 nucleocapsid protein (N protein 647, red). Then SARS-CoV-2+ Huh-7 cells were counted using the following building blocks: Find Nuclei>Find Cytoplasm (488+)>Calculate Intensity Properties> Select Population (infected cells)> formula output infected cells/total. (E) Automated count of SARS-CoV-2 infected cells treated with Cas13d RNP. Data are expressed as mean ± SD and analyzed by one-way ANOVA (∗p < 0.05; multiple comparison Dunnett’s test: ∗p < 0.05, ∗∗p < 0.01, ∗∗∗p < 0.001, and ∗∗∗∗p < 0.0001). (F) Representative images of HCS performed on Huh-7 cells treated or not with Au_NPs Cas13d as described in D. (G) Automated count of SARS-CoV-2 infected cells treated or not with Cas13d delivered by Au_NPs.
Interestingly, and in contrast with the results seen with ZIKV, Au_NPs Cas13d reduced SARS-CoV-2 released virions by 5-fold, while the canonical RNP transfection showed no antiviral activity (Figures 5B and 5C). This significant difference was further confirmed by counting the number of SARS-CoV-2-infected cells 24 h post infection (Figures 5D–5G). This suggests that Au_NPs Cas13d is notably more effective in reducing SARS-CoV-2 replication than standard RNP transfection.
Au_NPs Cas13d administration moderately reduces ZIKV replication
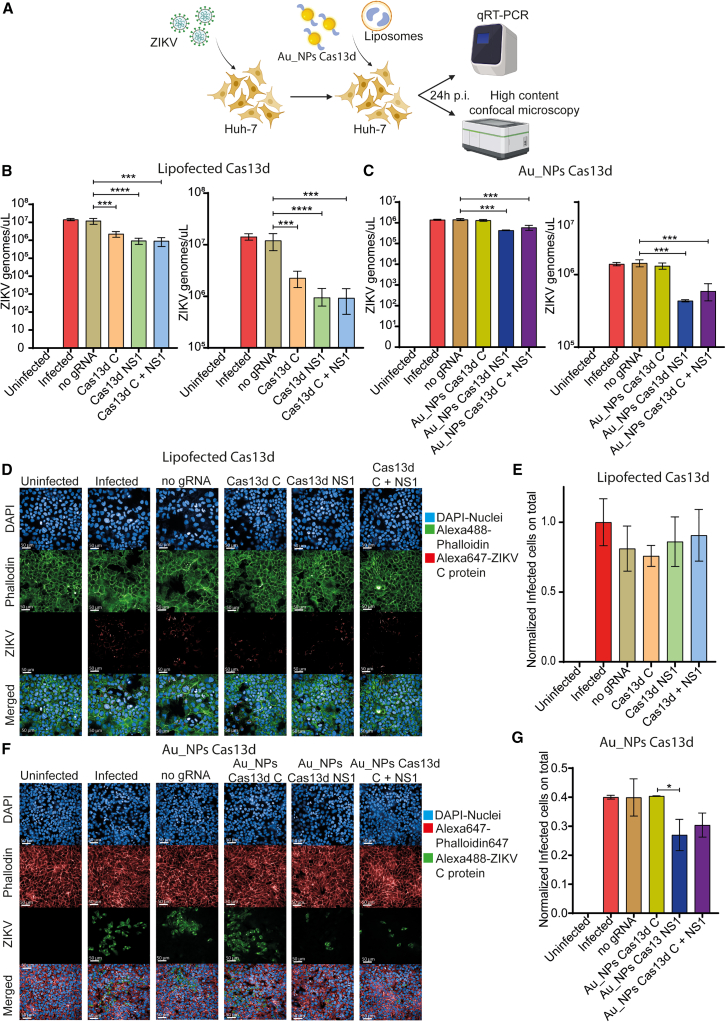
To investigate the antiviral potency of Au_NPs Cas13d in vitro, we infected Huh-7 cells with ZIKV at an MOI of 0.1 and added Au_NPs Cas13d to the culture media, using lipofected Cas13d RNPs as a control, as shown in Figure 6A. We then measured ZIKV genomes in the supernatants via RT-qPCR and assessed the number of infected cells using high content screenings (HCS). As seen in Figures 6B and 6C, Au_NPs Cas13d reduced ZIKV release by about 5-fold, while lipofected Cas13d RNPs decreased replication by 1 log10. Then, we counted the number of infected cells in each condition 48h after infection. A moderate reduction in infected cells was observed with both RNPs and Au_NPs Cas13d (Figures 6D–6G). We hypothesized that this modest antiviral activity could be due to the Cas13d enzyme itself, which is less efficient at cleaving multiple targets compared with Cas9. To test this, we monitored ZIKV replication over time after Au_NPs Cas13d administration, as outlined in Figure 7A. Cells were lysed at 0, 8, 12, and 24 h post infection, and viral RNA was analyzed by RT-qPCR. Interestingly, ZIKV replication resumed in both treatments, with viral genomes increasing similarly in Cas13d-treated and untreated cells after the initial reduction (Figures 7B and 7C). Based on these findings, we explored the effect of multiple administrations of Au_NPs Cas13d during ZIKV infection. Huh-7 cells were infected with ZIKV at 0.1 MOI, and Au_NPs Cas13d was administered at 1.5, 8, and 24 h post infection (Figure 8A). As shown in Figures 8B and 8C, multiple doses of lipofected Cas13d RNPs led to a 2 log10 reduction in ZIKV replication, while Au_NPs Cas13d resulted in a moderate 4-fold decrease in ZIKV genomic content (Figures 8D and 8E).
Figure 6.
Au_NPs Cas13d administration reduces ZIKV replication
(A) Representative illustration of the experimental workflow. (B) RT-qPCR quantification of ZIKV genomes released in supernatants after Cas13d lipofection. One-Way ANOVA: ∗∗∗p < 0.001; multiple comparison Dunnett’s test: ∗p < 0.05, ∗∗p < 0.01, ∗∗∗p < 0.001, and ∗∗∗∗p < 0.0001. (C) RT-qPCR quantification on ZIKV genomes released in supernatant after Cas13d administration delivered by Au_NPs. One-way ANOVA: ∗∗∗p < 0.001; multiple comparison Dunnett’s test: ∗p < 0.05, ∗∗p < 0.01, ∗∗∗p < 0.001, and ∗∗∗∗p < 0.0001. (D) Representative images of HCS on ZIKV-infected cells, treated with Cas13d delivered by lipofection. Cells were stained for nuclei (DAPI, blue), F-actin (Phalloidin 488, green), and ZIKV (C protein1, red). (E) Automated count of ZIKV-infected cells on samples shown in (D). Data are expressed as mean ± SD and analyzed by one-way ANOVA (p < NS; multiple comparison Dunnett’s test: ∗p < 0.05, ∗∗p < 0.01, ∗∗∗p < 0.001, and ∗∗∗∗p < 0.0001). (F) Representative images of HCS on ZIKV-infected cells, treated with Cas13d delivered by Au_NPs. Cells were stained for nuclei (DAPI, blue), F-actin (Phalloidin 647, red), and ZIKV (C protein1, green). (G) Automated count of ZIKV infected cells on samples shown in (D). Data are expressed as mean ± SD and analyzed by one-way ANOVA (∗p < 0.05; multiple comparison Dunnett’s test: ∗p < 0.05, ∗∗p < 0.01, ∗∗∗p < 0.001, and ∗∗∗∗p < 0.0001).
Figure 7.
Single Au_NPs administration slows down but does not abolish ZIKV replication
(A) Representative illustration of the experimental workflow. (B) RT-qPCR quantification on ZIKV genomes detected in cell lysates collected at different time points after single administration via lipofection of Cas13d. Data are expressed as mean ± SD and analyzed by one-way ANOVA: ∗∗∗∗p < 0.001; Multiple comparison Dunnett’s test: ∗p < 0.05, ∗∗p < 0.01, ∗∗∗p < 0.001, and ∗∗∗∗p < 0.0001. (C) RT-qPCR quantification on ZIKV genomes detected in cell lysates collected at different time points after single administration via Au_NPs of Cas13d. Data are expressed as mean ± SD and analyzed by one-way ANOVA: ∗∗∗p < 0.001; multiple comparison Dunnett’s test: ∗p < 0.05, ∗∗p < 0.01, ∗∗∗p < 0.001, and ∗∗∗∗p < 0.0001.
Figure 8.
Multiple administrations of Cas13d blocks ZIKV replication
(A) Representative scheme of the experimental workflow. (B) RT-qPCR quantification of virions released at different time points. (C) RT-qPCR quantification of virions released after 48 h from ZIKV-infected cells after 3 administrations of Cas13d delivered by lipofection. Data are expressed as mean ± SD and analyzed by one-way ANOVA: ∗∗∗p < 0.001; multiple comparison Dunnett’s test: ∗p < 0.05, ∗∗p < 0.01, ∗∗∗p < 0.001, and ∗∗∗∗p < 0.0001. (D) RT-qPCR quantification of virions released at different time points. (E) RT-qPCR quantification of virions released after 48 h from ZIKV infected cells after three administrations of Cas13d delivered by Au_NPs. Data are expressed as mean ± SD and analyzed by one-way ANOVA: ∗p < 0.001; multiple comparison Dunnett’s test: ∗p < 0.05, ∗∗p < 0.01, ∗∗∗p < 0.001, and ∗∗∗∗p <0.0001.
Au_NPs-Cas13d exhibits significantly reduced off-target effects compared with in silico predictions
We evaluated the off-target effects of Au_NPs Cas13d by performing in silico predictions using the bioinformatics pipeline previously described by Tong et al.58 This analysis identified 187 potential off-targets, which are listed in supplementary material 2 (Table S3). To validate these predictions experimentally, we conducted a transcriptomic analysis on cells treated with Au_NPs Cas13d loaded with a gRNA targeting the SARS-CoV-2 RdRP (Figure S6A). The analysis was specifically filtered to focus on the 187 predicted off-target genes to assess whether these were affected by the treatment. As shown in Figure S6B, only 9 of the 187 predicted off-target genes were found to be downregulated in the presence of the Au_NPs Cas13d nanoformulation. This demonstrates that Au_NPs Cas13d significantly minimizes off-target effects compared with the initial in silico predictions, highlighting its precision and potential for targeted therapeutic applications.
Discussion
The CRISPR-Cas systems have emerged as powerful tools in both medicine and basic research, particularly in combating viral infections. Recently, the newly discovered RNA-targeting enzyme Cas13d has been proposed as a promising candidate against RNA viruses, with proof-of-concept studies demonstrating its ability to rapidly degrade viral RNA genomes.18,20 However, several limitations must be addressed before Cas13d can be translated into clinical applications. First, Cas13d is less capable of processing multiple RNA targets compared with Cas9, which poses a challenge for combating rapidly replicating viruses.59,60,61 This limitation becomes critical when dealing with viruses that exhibit high mutation rates and robust replication strategies. Additionally, the delivery of Cas13d remains a significant hurdle, particularly in the context of viral infections where the timing of RNA degradation is crucial for effective antiviral responses.18,20
In this study, we introduced a novel approach for delivering Cas13d into infected cells without relying on traditional transfectants, thereby preserving its enzymatic activity. This was achieved by conjugating spherical Au_NPs with a carboxylic acid background (Au_12_PEG3000NTA_C) to Cas13d, creating a stable and oriented bond that maintains the enzyme functionality. The choice of Au_NPs as a delivery system is supported by several key advantages: their biocompatibility due to optimal shape, size, and charge, and their established use in various diagnostic and therapeutic applications.25,26,27,28 Furthermore, Au_NPs can be aerosolized,30,31,32 presenting a novel strategy for targeting respiratory viruses.
First, we examined whether the functionalization process had any significant effect on the properties of our Au_NPs. Our results demonstrate that the conjugation of Cas13d does not affect the DLS measurements or the Z-potential of Au_NPs. This indicates that there is no difference in cellular uptake between naked Au_NPs and Au_NPs conjugated with Cas13d. Additionally, our findings show that Au_NPs can spontaneously enter cells through multiple pathways, eliminating the need for potentially toxic, expensive, and time-consuming transfection protocols.33,34,35,36,37 Moreover, this system offers a dynamic therapeutic platform, allowing for the attachment of various endonucleases from the Cas protein family, to expand the range of antiviral applications. Importantly, we observed no difference in RNA cleavage efficiency between Cas13d delivered via Au_NPs and free Cas13d. Having established the effectiveness of Au_NPs Cas13d in RNA cleavage, we investigated their cellular entry mechanisms. Our results indicate that Au_NPs Cas13d can efficiently penetrate cells and co-localize with early endosomes (EEA1+) in 21% of cases, reaching peak cytosolic concentration within 4 h post administration. In particular, using inhibitors for different entry routes we demonstrated that 50% of Au_NPs Cas13d entered via passive diffusion; the rest of the Au_NPs Cas13ds enter via CME, FEME, and micro- and macro-pinocytosis. Subsequently, Au_NPs Cas13d accumulates in the cytoplasm (99.9%) and remains stable for at least 18 h before undergoing degradation and exocytosis via various processes, including macro-autophagy and vesicle secretion pathways.62,63
Upon confirming the entry and persistence of Au_NPs Cas13d in cells, we evaluated their antiviral activity against both ZIKV and SARS-CoV-2. Interestingly, we found that Au_NPs exhibited reduced efficacy against ZIKV compared with lipofected Cas13d, likely due to differences in intracellular distribution and the limited ability of Cas13d to target multiple RNAs before inactivation.59,60,61 To address this limitation, we administered Au_NPs Cas13d at various time points post infection resulted in a modest reduction in ZIKV replication. Subsequent experiments with SARS-CoV-2 demonstrated a reduction in viral replication with Au_NPs Cas13d, while the lipofected counterpart showed no activity. This disparity may be attributed to the varying intracellular localization of Cas enzymes relative to viral replication sites. Notably, while Au_NPs Cas13d co-localized with SARS-CoV-2 virions during the entry phase, it did not effectively co-localize with ZIKV in the Golgi cisternae, a virus-induced reshaped site that is mandatory for ZIKV replication.64
In conclusion, our study presents a novel strategy for delivering Cas enzymes using functionalized Au_NPs, which circumvents the need for traditional transfection methods and opens avenues for aerosolized delivery in respiratory viral infections. This proof-of-concept study lays the groundwork for leveraging gene-editing technologies against emerging coronaviruses, potentially intercepting viral infections at their onset. To enhance the efficacy of our approach, addressing the reduced RNA editing window of Cas13d is essential. Future efforts could focus on developing recombinant Cas13d proteins or identifying other Cas proteins with superior RNA cleavage capabilities.65 The next step involves aerosolizing our nanoformulation with a novel high-fidelity Cas13 recently developed for administration to hACE2 transgenic mice. This experiment will unveil the efficacy of mitigating SARS-CoV-2 infection at its onset, thereby reducing transmission. Before the in vivo studies, we will have to solve several challenges that decrease the antiviral potency of the system. First, the current functionalization and synthesis techniques may not yield sufficient quantities of both the enzyme and nanoparticles required to effectively treat the entire respiratory tract. Scaling up the production of Au_NPs and optimizing their functionalization will be necessary. Additionally, since our nanoformulation targets both infected and uninfected cells, achieving adequate concentrations of Au_NPs Cas13d in viral replication sites is crucial for maximizing efficacy. Moreover, the off-target effects, particularly regarding the cleavage of human RNAs needs to be thoroughly evaluated. In fact, we observed only 9 downregulated genes of 187 predicted off-targets when Au_NPs Cas13d was administered against SARS-CoV-2 RdRP, compared with the controls. It is possible that the intravesicular accumulation facilitated by Au_NPs reduces the likelihood of cleavage activity against human RNAs, which are typically not found in endosomes or double-membrane vesicles. Exploring this possibility in greater depth would be important in future studies, potentially by evaluating other gRNAs capable of targeting both viral and human genes.
While our findings indicate that Au_NPs Cas13d may co-localize with SARS-CoV-2, potentially reducing unintended interactions with human mRNAs, there remains a risk of localizing in uninfected cells or unintended regions within infected cells, leading to non-specific RNA degradation. Conversely, this co-localization could also enable the use of collateral activity to enhance the overall antiviral effect. We hypothesize that the antiviral activity of Au_NPs depends on the disruption of the +RNA fragment, as it has been recently published that Cas13d does not efficiently cleave the SARS-CoV-2 −RNA strand. This implies that −RNA strands produced in the replication facilities are not affected by the editing, while the complete SARS-CoV-2 genome is cleaved.66
Finally, using a nebulizer for aerosolizing our nanoformulation could mitigate opsonization, a common challenge when administering Au_NPs systemically. However, this approach may introduce other complications, such as precipitation in the lungs and adhesion to mucus. Future studies will need to address these issues, possibly through co-treatment strategies involving mucolytic agents to ensure optimal delivery and effectiveness of our nanoformulation.
Materials and methods
Au_NPs synthesis and functionalization
Au_NPs were stabilized with citrate and prepared as described by Turkevich et al.67 Trisodium citrate was used for the reduction of gold chloride trihydrate solution in an aqueous solution near the boiling point. Then, Au_NPs with 12 nm size were prepared and stabilized with polyethylene thiol terminated with a carboxylic group (molecular weight of 3 kDa). After purification, terminal carboxylic groups were activated by formation of active esters (sulfoNHS) and treated with thiolated NTA. Size and zeta potential were under investigation at each step of the synthesis. Subsequent treatment with nickel (Ni2+) salt and Cas13d with His tag allowed attachment of the proteins to Au_NPs (Figure S7). The final Au_NPs possessed some free carboxylic acid groups to provide an appropriate background and improved solubility in aqueous solution.
Determination of Au_NPs size and zeta potential
A DLS Zetasizer Ultra (Malvern Panalytical Ltd., Malvern, UK) analyzer was used to determine the hydrodynamic particle size and zeta potential of Au_NPs. The detection angle was set at 90°, the temperature was set at 25°C, and the refractive index was set at 1.33. A helium-neon laser beam and a clear polystyrene cuvette (3 mL, 10 × 10 × 45 mm) were used for sample analysis. The size and morphology of the prepared Au_NPs were determined using a JEM 1400 (JEOL Co., Tokyo, Japan, 2008) with an energy-dispersive full range X-ray microanalysis system (EDS INCA Energy TEM, Oxford Instruments, London, UK), a tomographic holder, and an 11-megapixel TEM Camera MORADA G2 (EMSIS, Münster, Germany) (JEOL, Japan; Laboratory of Electron Microscopy, the Nencki Institute of Experimental Biology Polish Academy of Science, Warsaw, Poland). The samples were prepared by placing them on carbon-coated copper and leaving them to air dry before imaging.
gRNA design and synthesis
First, we selected gRNAs for Cas13b using the online tools http://crispr-pacman.stanford.ed and https://cas13design.nygenome.org/. Then tested for off-target activity in human mRNAs using nucleotide blast as schematically reported in Table S1. Then, gRNAs were adapted for Cas13d, a newly discovered Cas13 with greater precision and reduced dimension than Cas13b.18 Selected gRNAs are reported in Table S2.
gRNA cloning and lentiviral vectors
We cloned selected gRNAs into pLentiRNACRISPR_004 - hU6-DR_BsmBI-EFS-PspCas13b-NES-2A-Puro-WPRE plasmid (Addgene, Watertown, MA, USA, #138146), using the canonical BbsI (Thermo Fisher Scientific, Waltham, MA, USA, FD1014) digestion protocol described elsewhere. These plasmids were then used to produce lentiviral particles by transfecting 107 293T cells with psPAX2 (Addgene, #12260), pCMV-Rev (Addgene, #115776), pMD2.G (Addgene, #12259) and pLentiRNACRISPR_004 - hU6-DR_BsmBI-EFS-PspCas13b-NES-2A-Puro-WPRE (Addgene, #138146) plasmids using Polyetylenimine (Sigma-Aldrich, St. Missouri, MO, USA, 408727) as transfection agent. Supernatants were then collected and used to transduce Huh-7 cells 72 h after transfection, using polybrene (Sigma-Aldrich, TR-1003-g) at 10 μg/mL to maximize transduction efficiency. Transduced cells were then selected by puromycin (Thermo Fisher Scientific, Waltham, MA, USA, A1113803) at 3 μg/μL.
Infections
Infections were performed using 0.1, 1, 5, or 100 MOI of ZIKV (1308258v, strain MP1751) or SARS-CoV-2 (clinical isolate coming from the Pisa University Hospital) for 1.5 h administered to Huh-7 cells using FBS-free DMEM (Thermo Fisher Scientific, 11960044). Then, cell media were replaced 1.5 h after infection. Cells were then treated with Au_NPs Cas13d or lipofected with Cas13d (MCLAB, South San Francisco, CA, USA, CAS13d-300) using phenol red-free media and Lipofectamine RNAiMAX Transfection Reagent (Thermo Fisher Scientific, 13778075).
Au_NPs Cas13d and free Cas13d were mixed with gRNA in a 1:2 ratio in Cas13d reaction buffer (20 mM HEPES, 5 mM MgCl2, 0.1M NaCl, 0.1 mM EDTA, pH 6.5) for 5 min at room temperature (RT). Au_NPs Cas13d were then administered directly to the cells in a final volume of 150 μL of DMEM without fetal bovine serum (FBS) and phenol red. Free Cas13 was mixed with 1.2 μL Lipofectamine RNAiMAX in a total volume of 50 μL DMEM without FBS and phenol red and incubated for 20 min at RT to form liposomes. The liposomes were then applied to the cells in a final volume of 150 μL of DMEM without FBS and phenol red.
Evaluation of entry routes
To evaluate the different entry routes exploited by Au_NPs Cas13 we seeded 1.5 × 104 Huh-7 cell lines. After 24 h, we administered the Au_NPs Cas13d to cells placed at 4°C and incubated them 4 h before to fix cells. In parallel, we administered 2 h before the treatment with Au_NPs Cas13d chloroquine (25 μM), nystatin (20 μM), 7-keto-cholesterol (25 μM), Dynasore (40 μM), and combination of Dynasore and nystatin to inhibit CME, CLIC/GEEC, FEME, and macro- and micro-pinocytosis, respectively.57 At 4 h after administration of Au_NPs Cas13d, we fixed cells to perform immunostaining.
Immunostainings
Cells were fixed using paraformaldehyde (PFA) 4% and permeabilized with PBS Ca2+Mg2+ 0.5% Triton X-100. Then cells were blocked with PBS Ca2+Mg2+ Triton X-100 0.1% and FBS 3% and stained for ZIKV NS1 (Genetex, Irvine, CA, USA, GTX133323) or Zika capsid protein (Genetex), 1:200; SARS-CoV-2 N (Sino Biologica, Beijing, China, 40588-RC02-100) 1:1000; EEA1 (Thermo Fisher Scientific, #MA5-14794) 1:250. To label primary antibodies, we used goat anti-rabbit IgG (H + L) cross-adsorbed secondary antibody, Alexa Fluor 647 (Thermo Fisher Scientific) 1:1,000, goat anti-rabbit IgG (H + L) cross-adsorbed secondary antibody, Alexa Fluor 488 (Thermo Fisher Scientific) 1:1,000, goat anti-mouse IgG (H + L) cross-adsorbed secondary antibody, Alexa Fluor 647 (Thermo Fisher Scientific) 1:1,000. DAPI (Thermo Fisher Scientific) 1:1,000 and Alexa Fluor 488 Phalloidin or Alexa fluor 647 Phalloidin (Thermo Fisher Scientific) 1:2,000 were used to stain nuclei and cytoplasms, respectively. Images were acquired and analyzed by Operetta CLS platform (Revvity, Waltham, MA, USA).
High-content confocal imaging
Operetta CLS platform (Revvity), using water 40× magnification. Images were then processed following these building blocks using Harmony 4.9 software (Revvity): Find Nuclei -DAPI>Find cytoplasm-Alexa488>Calculate Alexa 647 intensity (ZIKV/SARS-CoV-2) >Select population: Infected cells.
RNA extraction and RT-qPCR
Viral genomes were extracted from supernatants by QIAamp Viral RNA Mini Kit (QIAGEN, Hilden, Germany, 52904). Subsequently, RNA was reverse transcribed and amplified by One Step PrimeScript III RT-PCR (Takara, Kyoto, Japan RR601A). Here the complete list of primers and probes: ZIKV- F: 5′-TGAGATCAACCACTGCAAGY-3′, R: 5′- GCCTTATCTCCATTCCATACCA -3′, Probe: 5′- FAM-ATCGAGGAATGGTGCTGCAGGGA-BHQ1-3′. SARS-CoV-2 -F: 5′-TCACCTAATTTAGCATGGCCTCT-3′, R: 5′-CGTAGTGCAACAGGACTAAGC-3′ probes: 5′-FAM/ACAGCAGAATTGGCCCTTAAA/BHQ1-3′.
Au_NPs production and functionalization
Au_NPs Cas13d production and functionalization are schematically reported in Figure S7. Briefly, we added 8 mM NiSO4 to 1013 Au_NPs in 1 mL water for 20 min at RT in the dark. Subsequently, we centrifugated the sample in 15 mL Amicon 50 kDa (Millipore, Burlington, MA, USA, UFC805024) at 1,100×g, 10 min, RT. Au_NPs were then washed twice using 3 mL sterile water, centrifugating the sample at 1,100×g, 10 min after each passage. Au_NPs were then resuspended on Cas13d storage buffer (0.3 M NaCl, 20 mM Tris-HCl, pH 8.0, 0.5 nM EDTA, 1 mM DTT) in 250 μL and centrifugated at 1,100×g 10 min twice. Then, 212 μg of Cas13d were mixed to charged Au_NPs for 3 h at 4°C with gentle shaking. Finally, we purified Au_NPs with Ni-NTA resin. Briefly, the resin was first activated by adding Equilibration buffer (20 mM Na3PO4, 300 mM NaCl, 10 mM imidazole). Then, Au_NPs Cas13d were added to the resin incubated for 1 h at 4°C in gentle shacking. Samples were then centrifuged at 500×g, 4°C for 1 min, repeating this step twice. Then, resin contamination is removed by centrifugating the Au_NPs solution with a microtube filter at 500×g for 1 min. Au_NPs Cas13d were quantified using Qubit 4 fluorometer (Thermo Fisher Scientific) by following the Qubit protein assay kit (Thermo Fisher Scientific) manufacturer’s instruction.
Cell cultures
Huh-7 cells were purchased from American Type Culture Collection (Manassas, VA, USA), and cultured at 37°C, 5% CO2 using DMEM 10% FBS, 1% L-glutamine (Thermo Fisher Scientific), and 1% sodium pyruvate media (Thermo Fisher Scientific). These cells were used to generate Cas13b-Huh-7 and Huh-7-mGFP. Transduced cells were maintained with DMEM with 3 μg/μL puromycin.
High-content confocal screenings
Cell entry assay
We seeded 1.5 × 104 Huh-7-mGFP cells in CellCarrier 96-ultra plates (Revvity). Then, nuclei were labeled using Hoechst 33342 (Thermo Fisher Scientific) and Alexa 546-labeled Au_NPs Cas13d gRNA was administered to cells for 4 h at 37°C, 5% CO2. We performed a live imaging acquisition using Operetta CLS (Revvity) acquiring 1 image every 30 min for 4 h and 1 image every hour for 10 h. Images were then processed with the following building blocks: Find nuclei, DAPI>Find cytoplasm, EGFP>Find spots, Alexa 546>calculate intensity properties in spots>select population: internalized Au_NPs >select population: external Au_NPs >select population: Au_NPs in the cytoplasm>select population: Au_NPs in nuclei.
Au_NPs Cas13d localization assays
We seeded and stained Huh-7 cells as described above. Then, we treated cells with Alexa 546-labeled Au_NPs Cas13d and fixed cells with PFA 4% 4 h after. Then, cells were stained for EEA1and analyzed as follows: Find nuclei, DAPI>Find Cytoplasm, EGFP>Find spot, Alexa 546>calculate intensity properties – spot >Select population: Au_NPs Cas13d>Find spots, Alexa 647>calculate intensity properties>Select population: Early endosomes>calculate intensity properties>Select population: Early endosomes (EEA1+) and Au_NPs Cas13d+.
Au_NPs Cas13d entry route assays
We seeded and treated with inhibitor Huh-7 cells as described above. Then, we treated cells with Alexa 546-labeled Au_NPs Cas13d and fixed cells with PFA 4% 4 h after. Then, cells were stained with DAPI and Phalloidin 647 (Thermo Fisher Scientific) and analyzed as follows: Find nuclei, DAPI>Find cytoplasm, Alexa 647 >Find spot, Alexa 546>calculate intensity properties – spot >Select population: Au_NPs Cas13d.
TEM
We infected 106 Huh-7 cells with 5 or 100 MOI of SARS-CoV-2 or ZIKV in the presence or not of Au_NPs Cas13d, provided simultaneously or 20 h post infection. Then, cells were fixed, 2 h or 24 h after infection, with 2.5% glutaraldehyde in 0.1 M cacodylate buffer, pH 7.2, for 2 h at 4°C, and postfixed in 1% osmium tetroxide in 0.1 M cacodylate buffer for 1 h at RT. Samples were then dehydrated with ethanol washings and transferred into propylene oxide for 15 min. Finally, they were embedded in Epon-Araldite, and polymerized at 60°C. Ultrathin sections (60–90 nm), obtained with a Reichert-Jung ultracut E (Reichert-Jung, Wien, Austria) equipped with a diamond knife, were collected on 200-mesh formvar/carbon-coated copper grids, double stained with aqueous uranyl acetate and lead citrate, and examined with a Jeol 100SX transmission electron microscope (Jeol, Tokyo, Japan) operating at 80 kV. Micrographs were obtained with an AMTXR80b Camera system.
In vitro cleavage assay
We synthetized a 1,672-bp RNA starting from SARS-CoV-2 viral RNA by following the protocol supplied by the MEGAscript T7 Transcription Kit (Thermo Fisher Scientific, AM1334). Here the primers used for the reaction:5′-TAATACGACTCACTATAGGGGGTGCTCAGTGGACTTCTC-3′; 5′- CCTCACAGCTGTCAATCCCA-3′. Then, we mixed Cas13d or Au_NPs Cas13d with the same gRNA and used to digest the SARS-CoV-2 RNA fragment at various concentrations. RNA substrates and Cas13d or Au_NPs Cas13d were mixed in Cas13d reaction buffer (20 mM HEPES, 5 mM MgCl2, 0.1 M NaCl, 0.1 mM EDTA, pH 6.5). Then, 1 μg/μL of Proteinase K (QIAGEN, 19131) was added for 1h at 37°C and subsequently inactivated at 56°C for 15 min following 5 min at 98°C. Then, we evaluated the amount of unedited/edited RNAs by RT-qPCR, using the following primers: F1: 5′-CCCCACTATCGTTTCGAGCA-3′, R1: 5′-TTCTTCAACCCCTCCTGCAC-3′, Probe 1: 5′-HEX-CTTTGGCCTGGTTGGCAATTCGAGCA-BHQ1-3′; F2: 5′-CTGCAAACTGCGATCTCTGC-3′, R2: 5′-GCCTCGAGCTAGTGGTGTTA-3′, Probe 2: 5′-FAM-ACGGCTGCTGGTATGGAATGGAGATAAGG-BHQ1-3′.
RNA sequencing and in silico analysis of off-target
Total RNA was extracted from cells using the RNeasy Mini Kit (QIAGEN) according to the manufacturer’s instructions. The RNA library was prepared using the Illumina Stranded mRNA Prep, Ligation, following the manufacturer’s instructions with 500 ng of RNA input. Each library was quantified using the Qubit dsDNA HS Assay Kit on a Qubit 4.0 Fluorometer (Thermo Fisher Scientific) and the library size was checked using Agilent Bioanalyzer and Agilent Bioanalyzer DNA 1000 kit. Then the library was normalized to 1.3 nM and pooled together. The pooled library was denatured and 1.2 pM was sequenced on NextSeq 550 Illumina platform using the NextSeq 500/550 High Output Kit v2.5 (150 Cycles) with a 2 × 75 bp paired-end layout.
Demultiplexed and trimmed reads were aligned to the GRCh38 human reference genome using HISAT2 software.68 FeatureCounts was used to quantify gene-specific read counts for each sample. Differential analysis among groups (untreated, naked Au_Nps, no gRNA and Au_NPs Cas13d) was performed using limma-voom, a count-based method implemented in R with voom for normalization.69,70 Benjamini-Hochberg adjusted p < 0.05 and at least 1.5 fold-change in expression were the criteria used to screen the significantly expressed genes.
Off-target sites for the gRNAs were predicted using an in silico analysis. Initially, gRNAs were aligned against the human transcriptome (GRCh38, Ensembl release 112) with blastn (megablast), configured to detect potential off-targets by allowing up to seven mismatches. The alignment parameters were set as follows: max_target_seqs = 10,000, evalue = 10,000, word_size = 4, and perc_identity = 0.6. Subsequently, candidate off-targets were refined by requiring each alignment to contain a minimum of 10 consecutive base pairs of perfect matches. Finally, HOMER (v4.11) was used to annotate the identified off-target sites.58
The differentially expressed genes (DEGs), supplementary data, were filtered based on the previous described in silico analysis, only those genes with a maximum of five mismatches with the gRNA were considered. Ggplot2, package of R, was applied to generate Volcano Plot of DEGs.
Statistical analysis
All data were analyzed on software GraphPad Prism (GraphPad Software, La Jolla, CA, USA). Statistical analysis used were one-way ANOVA, multiple comparison Dunnett’s test or multiple comparison Tukey’s test. Significant p value are: ∗≤0.05, ∗∗ ≤0.01, ∗∗∗ ≤0.001, and ∗∗∗∗≤0.0001.
Data availability
Data are available upon request to corresponding author.
Acknowledgments
This work was supported by “I-GENE, In-vivo Gene Editing by Nanotransducers,” European call identifier H2020-FETOPEN-2018-2020, Proposal ID 862714. Progetti di Ricerca di Rilevante Interesse Nazionale PRIN 2020, Prot. n. 2020KSY3KL. Progetto di Ricerca di Ateneo 2020, PRA_2020_32 and PRA_2020_37 of the University of Pisa. PNRR - Tuscany Health Ecosystem (THE) – CUP I53C22000780001 – Project code ECS00000017- spoke n. 7 - “Innovating Translational Medicine”. - Sub-project 5 - Innovative models for management of infections caused by antibiotic-resistant bacteria. PRIN 2020 “Development of VEGFR/Tubulin and Efflux Pump inhibitors loaded on stimuli-responsive cell membrane coated Nanocarriers for the treatment of Metastatic Cancers” - 20200239N53. “Unraveling the molecular and immunologic mechanisms of intrahost persistence in emerging and reemerging arboviral infections – GENESIS”, ID S1.p0002, CUP i53c24001430007, funded by bando a cascata COC-1-2023-UNIPV, using the resources from Piano Nazionale Ripresa e Resilienza (PNRR) missione 4, “Istruzione e Ricerca” – Componente 2, “dalla Ricerca all’Impresa” – linea di investimento 1.3, finanziato dall’Unione Europea - NextGenerationEU”. PNRR-MAD-2022-12376570 “Identification of common pathogenic mechanisms driving squamous cell carcinomas of the anogenital tract and head&neck region to develop overarching therapeutic strategies” “Funded by the European Union - Next Generation EU - NRRP M6C2 - Investment 2.1 Enhancement and strengthening of biomedical research in the NHS” PNRR-MAD-2022-12376570- CUP D53C22004170006. We acknowledge CISUP—Centre for Instrumentation Sharing University of Pisa for the use of Operetta CLS imaging facility. We would like to express our sincere gratitude to Soultana Konstantinidou for her invaluable assistance with the functionalization and characterization of Au_NPs.
Author contributions
Conceptualization, M.L., A.D.C., and M.P.; Methodology, A.D.C, D.F., C.F., F.F., R.F., E.P., S.N., A.AL., E.H.C., and A.L.; Investigation, A.D.C., D.F., P.B., D.W., A.F., G. Frenzilli, G. Freer., and A.L.; Writing – original draft, A.D.C. and D.F.; Writing – review & editing, M.L., M.P., and A.D.C; Funding acquisition, M.P, V.R., and M.L.; Supervision, M.L., M.P., and V.R.
Declaration of interests
The authors declare that they have no competing interests.
Footnotes
Supplemental information can be found online at https://doi.org/10.1016/j.omtn.2025.102540.
Supplemental information
References
- 1.Yuan Y., Jiao B., Qu L., Yang D., Liu R. The development of COVID-19 treatment. Front. Immunol. 2023;14 doi: 10.3389/fimmu.2023.1125246. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Chinta S., Rodriguez-Guerra M., Shaban M., Pandey N., Jaquez-Duran M., Vittorio T.J. COVID-19 therapy and vaccination: a clinical narrative review. Drugs Context. 2023;12:1–11. doi: 10.7573/dic.2022-7-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Fiolet T., Kherabi Y., MacDonald C.J., Ghosn J., Peiffer-Smadja N. Comparing COVID-19 vaccines for their characteristics, efficacy and effectiveness against SARS-CoV-2 and variants of concern: a narrative review. Clin. Microbiol. Infect. 2022;28:202–221. doi: 10.1016/j.cmi.2021.10.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Wilder-Smith A., Ooi E.E., Horstick O., Wills B. Dengue. Lancet. 2019;393:350–363. doi: 10.1016/S0140-6736(18)32560-1. [DOI] [PubMed] [Google Scholar]
- 5.Huang Z., Zhang Y., Li H., Zhu J., Song W., Chen K., Zhang Y., Lou Y. Vaccine development for mosquito-borne viral diseases. Front. Immunol. 2023;14 doi: 10.3389/fimmu.2023.1161149. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Tai W., Zhang X., Yang Y., Zhu J., Du L. Advances in mRNA and other vaccines against MERS-CoV. Transl. Res. 2022;242:20–37. doi: 10.1016/j.trsl.2021.11.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Tomori O., Kolawole M.O. Ebola virus disease: current vaccine solutions. Curr. Opin. Immunol. 2021;71:27–33. doi: 10.1016/j.coi.2021.03.008. [DOI] [PubMed] [Google Scholar]
- 8.Lai M., Maori E., Quaranta P., Matteoli G., Maggi F., Sgarbanti M., Crucitta S., Pacini S., Turriziani O., Antonelli G., et al. CRISPR/Cas9 Ablation of Integrated HIV-1 Accumulates Proviral DNA Circles with Reformed Long Terminal Repeats. J. Virol. 2021;95 doi: 10.1128/JVI.01358-21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Herrera-Carrillo E., Gao Z., Berkhout B. CRISPR therapy towards an HIV cure. Brief. Funct. Genomics. 2020;19:201–208. doi: 10.1093/bfgp/elz021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Jubair L., Fallaha S., McMillan N.A.J. Systemic Delivery of CRISPR/Cas9 Targeting HPV Oncogenes Is Effective at Eliminating Established Tumors. Mol. Ther. 2019;27:2091–2099. doi: 10.1016/j.ymthe.2019.08.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Zhen S., Qiang R., Lu J., Tuo X., Yang X., Li X. CRISPR/Cas9-HPV-liposome enhances antitumor immunity and treatment of HPV infection-associated cervical cancer. J. Med. Virol. 2023;95:e28144. doi: 10.1002/jmv.28144. [DOI] [PubMed] [Google Scholar]
- 12.Yin D., Ling S., Wang D., Dai Y., Jiang H., Zhou X., Paludan S.R., Hong J., Cai Y. Targeting herpes simplex virus with CRISPR–Cas9 cures herpetic stromal keratitis in mice. Nat. Biotechnol. 2021;39:567–577. doi: 10.1038/s41587-020-00781-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Ying M., Wang H., Liu T., Han Z., Lin K., Shi Q., Zheng N., Ye T., Gong H., Xu F. CLEAR Strategy Inhibited HSV Proliferation Using Viral Vectors Delivered CRISPR-Cas9. Pathogens. 2023;12:814. doi: 10.3390/pathogens12060814. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Wei A., Yin D., Zhai Z., Ling S., Le H., Tian L., Xu J., Paludan S.R., Cai Y., Hong J. In vivo CRISPR gene editing in patients with herpetic stromal keratitis. Mol. Ther. 2023;31:3163–3175. doi: 10.1016/j.ymthe.2023.08.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Cui J., Techakriengkrai N., Nedumpun T., Suradhat S. Abrogation of PRRSV infectivity by CRISPR-Cas13b-mediated viral RNA cleavage in mammalian cells. Sci. Rep. 2020;10:9617. doi: 10.1038/s41598-020-66775-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Cox D.B.T., Gootenberg J.S., Abudayyeh O.O., Franklin B., Kellner M.J., Joung J., Zhang F. RNA editing with CRISPR-Cas13. Science. 2017;358:1019–1027. doi: 10.1126/science.aaq0180. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Xue Y., Chen Z., Zhang W., Zhang J. Engineering CRISPR/Cas13 System against RNA Viruses: From Diagnostics to Therapeutics. Bioengineering. 2022;9:291. doi: 10.3390/bioengineering9070291. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Abbott T.R., Dhamdhere G., Liu Y., Lin X., Goudy L., Zeng L., Chemparathy A., Chmura S., Heaton N.S., Debs R., et al. Development of CRISPR as an Antiviral Strategy to Combat SARS-CoV-2 and Influenza. Cell. 2020;181:865–876.e12. doi: 10.1016/j.cell.2020.04.020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Chen P., Chen M., Chen Y., Jing X., Zhang N., Zhou X., Li X., Long G., Hao P. Targeted inhibition of Zika virus infection in human cells by CRISPR-Cas13b. Virus Res. 2022;312 doi: 10.1016/j.virusres.2022.198707. [DOI] [PubMed] [Google Scholar]
- 20.Lin X., Liu Y., Chemparathy A., Pande T., La Russa M., Qi L.S. A comprehensive analysis and resource to use CRISPR-Cas13 for broad-spectrum targeting of RNA viruses. Cell Rep. Med. 2021;2 doi: 10.1016/j.xcrm.2021.100245. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Babalola B.A., Akinsuyi O.S., Folajimi E.O., Olujimi F., Otunba A.A., Chikere B., Adewumagun I.A., Adetobi T.E. Exploring the future of SARS-CoV-2 treatment after the first two years of the pandemic: A comparative study of alternative therapeutics. Biomed. Pharmacother. 2023;165 doi: 10.1016/j.biopha.2023.115099. [DOI] [PubMed] [Google Scholar]
- 22.Alves E., Taifour S., Dolcetti R., Chee J., Nowak A.K., Gaudieri S., Blancafort P. Reprogramming the anti-tumor immune response via CRISPR genetic and epigenetic editing. Mol. Ther. Methods Clin. Dev. 2021;21:592–606. doi: 10.1016/j.omtm.2021.04.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Chaves L.C.S., Orr-Burks N., Vanover D., Mosur V.V., Hosking S.R., Kumar E K P., Jeong H., Jung Y., Assumpção J.A.F., Peck H.E., et al. mRNA-encoded Cas13 treatment of Influenza via site-specific degradation of genomic RNA. PLoS Pathog. 2024;20 doi: 10.1371/journal.ppat.1012345. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Yu D., Han H.J., Yu J., Kim J., Lee G.H., Yang J.H., Song B.M., Tark D., Choi B.S., Kang S.M., Heo W.D. Pseudoknot-targeting Cas13b combats SARS-CoV-2 infection by suppressing viral replication. Mol. Ther. 2023;31:1675–1687. doi: 10.1016/j.ymthe.2023.03.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Sakthi Devi R., Girigoswami A., Siddharth M., Girigoswami K. Applications of Gold and Silver Nanoparticles in Theranostics. Appl. Biochem. Biotechnol. 2022;194:4187–4219. doi: 10.1007/s12010-022-03963-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Anik M.I., Mahmud N., Al Masud A., Hasan M. Gold nanoparticles (GNPs) in biomedical and clinical applications: A review. Nano Select. 2022;3:792–828. doi: 10.1002/nano.202100255. [DOI] [Google Scholar]
- 27.Zhang R., Kiessling F., Lammers T., Pallares R.M. Clinical translation of gold nanoparticles. Drug Deliv. Transl. Res. 2023;13:378–385. doi: 10.1007/s13346-022-01232-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Oliveira B.B., Ferreira D., Fernandes A.R., Baptista P.V. Engineering gold nanoparticles for molecular diagnostics and biosensing. WIREs Nanomedicine and Nanobiotechnology. 2023;15:e1836. doi: 10.1002/wnan.1836. [DOI] [PubMed] [Google Scholar]
- 29.Tang H., Zhang X., Bao Y., Shen H., Fan M., Wang Y., Xiang S., Ran X. Nucleic acid-functionalized gold nanoparticles as intelligent photothermal therapy agents for precise cancer treatment. Nanotechnology. 2024;35 doi: 10.1088/1361-6528/ad6fa7. [DOI] [PubMed] [Google Scholar]
- 30.Urso A., Meloni F., Malatesta M., Latorre R., Damoci C., Crapanzano J., Pandolfi L., Giustra M.D., Pearson M., Colombo M., et al. Endotracheal Nebulization of Gold Nanoparticles for Noninvasive Pulmonary Drug Delivery. Nanomedicine. 2023;18:317–330. doi: 10.2217/nnm-2022-0179. [DOI] [PubMed] [Google Scholar]
- 31.Shrestha A., Haque M.A., Mattheolabakis G. Pulmonary Delivery for miRs: Present and Future Potential. Processes. 2023;11:1788. doi: 10.3390/pr11061788. [DOI] [Google Scholar]
- 32.De Berardis B., Marchetti M., Risuglia A., Ietto F., Fanizza C., Superti F. Exposure to airborne gold nanoparticles: a review of current toxicological data on the respiratory tract. J. Nanoparticle Res. 2020;22:235. doi: 10.1007/s11051-020-04966-9. [DOI] [Google Scholar]
- 33.Cheng X., Tian X., Wu A., Li J., Tian J., Chong Y., Chai Z., Zhao Y., Chen C., Ge C. Protein Corona Influences Cellular Uptake of Gold Nanoparticles by Phagocytic and Nonphagocytic Cells in a Size-Dependent Manner. ACS Appl. Mater. Interfaces. 2015;7:20568–20575. doi: 10.1021/acsami.5b04290. [DOI] [PubMed] [Google Scholar]
- 34.Foroozandeh P., Aziz A.A. Insight into Cellular Uptake and Intracellular Trafficking of Nanoparticles. Nanoscale Res. Lett. 2018;13:339. doi: 10.1186/s11671-018-2728-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Sousa de Almeida M., Taladriz-Blanco P., Drasler B., Balog S., Yajan P., Petri-Fink A., Rothen-Rutishauser B. Cellular Uptake of Silica and Gold Nanoparticles Induces Early Activation of Nuclear Receptor NR4A1. Nanomaterials. 2022;12:690. doi: 10.3390/nano12040690. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Ding L., Yao C., Yin X., Li C., Huang Y., Wu M., Wang B., Guo X., Wang Y., Wu M. Size, Shape, and Protein Corona Determine Cellular Uptake and Removal Mechanisms of Gold Nanoparticles. Small. 2018;14:e1801451. doi: 10.1002/smll.201801451. [DOI] [PubMed] [Google Scholar]
- 37.Peter B., Kanyo N., Kovacs K.D., Kovács V., Szekacs I., Pécz B., Molnár K., Nakanishi H., Lagzi I., Horvath R. Glycocalyx Components Detune the Cellular Uptake of Gold Nanoparticles in a Size- and Charge-Dependent Manner. ACS Appl. Bio Mater. 2023;6:64–73. doi: 10.1021/acsabm.2c00595. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Dockery L.T., Daniel M.C. Targeted Doxorubicin-Loaded Dendronized Gold Nanoparticles. Pharmaceutics. 2023;15:2103. doi: 10.3390/pharmaceutics15082103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Wang L., Kulthinee S., Yano N., Wen H., Zhang L.X., Saleeba Z.S.S.L., Jin N., Chen O., Zhao T.C. Gold nanoparticles-conjugation of irisin enhances therapeutic effect by improving cardiac function and attenuating inflammation in sepsis. Mol. Divers. 2025;29:1557–1568. doi: 10.1007/s11030-024-10933-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Takayanagi T., Miyake K., Seto M., Mizuguchi H., Okabe H., Matsuda N. Conjugation monitoring of gold nanoparticles with alkanedithiols by capillary zone electrophoresis. Anal. Sci. 2023;39:1033–1039. doi: 10.1007/s44211-023-00299-4. [DOI] [PubMed] [Google Scholar]
- 41.Pourali P., Dzmitruk V., Benada O., Svoboda M., Benson V. Conjugation of microbial-derived gold nanoparticles to different types of nucleic acids: evaluation of transfection efficiency. Sci. Rep. 2023;13 doi: 10.1038/s41598-023-41567-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Astorga-Gamaza A., Vitali M., Borrajo M.L., Suárez-López R., Jaime C., Bastus N., Serra-peinado C., Luque-Ballesteros L., Blanch-Lombarte O., Prado J.G., et al. Antibody cooperative adsorption onto AuNPs and its exploitation to force natural killer cells to kill HIV-infected T cells. Nano Today. 2021;36 doi: 10.1016/j.nantod.2020.101056. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Jackson C.B., Farzan M., Chen B., Choe H. Mechanisms of SARS-CoV-2 entry into cells. Nat. Rev. Mol. Cell Biol. 2022;23:3–20. doi: 10.1038/s41580-021-00418-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Lai M., De Carli A., Filipponi C., Iacono E., La Rocca V., Lottini G., Piazza C.R., Quaranta P., Sidoti M., Pistello M., Freer G. Lipid balance remodelling by human positive-strand RNA viruses and the contribution of lysosomes. Antiviral Res. 2022;206 doi: 10.1016/j.antiviral.2022.105398. [DOI] [PubMed] [Google Scholar]
- 45.Wolff G., Melia C.E., Snijder E.J., Bárcena M. Double-Membrane Vesicles as Platforms for Viral Replication. Trends Microbiol. 2020;28:1022–1033. doi: 10.1016/j.tim.2020.05.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Roingeard P., Eymieux S., Burlaud-Gaillard J., Hourioux C., Patient R., Blanchard E. The double-membrane vesicle (DMV): a virus-induced organelle dedicated to the replication of SARS-CoV-2 and other positive-sense single-stranded RNA viruses. Cell. Mol. Life Sci. 2022;79:425. doi: 10.1007/s00018-022-04469-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Agrelli A., de Moura R.R., Crovella S., Brandão L.A.C. ZIKA virus entry mechanisms in human cells. Infect. Genet. Evol. 2019;69:22–29. doi: 10.1016/j.meegid.2019.01.018. [DOI] [PubMed] [Google Scholar]
- 48.van den Elsen K., Quek J.P., Luo D. Molecular Insights into the Flavivirus Replication Complex. Viruses. 2021;13:956. doi: 10.3390/v13060956. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Buchwalter R.A., Ogden S.C., York S.B., Sun L., Zheng C., Hammack C., Cheng Y., Chen J.V., Cone A.S., Meckes D.G., Jr., et al. Coordination of Zika Virus Infection and Viroplasm Organization by Microtubules and Microtubule-Organizing Centers. Cells. 2021;10:3335. doi: 10.3390/cells10123335. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Talarska P., Boruczkowski M., Żurawski J. Current Knowledge of Silver and Gold Nanoparticles in Laboratory Research—Application, Toxicity, Cellular Uptake. Nanomaterials. 2021;11:2454. doi: 10.3390/nano11092454. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Okoampah E., Mao Y., Yang S., Sun S., Zhou C. Gold nanoparticles–biomembrane interactions: From fundamental to simulation. Colloids Surf. B Biointerfaces. 2020;196 doi: 10.1016/j.colsurfb.2020.111312. [DOI] [PubMed] [Google Scholar]
- 52.Schwartz-Duval A.S., Sokolov K.V. Prospecting Cellular Gold Nanoparticle Biomineralization as a Viable Alternative to Prefabricated Gold Nanoparticles. Adv. Sci. 2022;9 doi: 10.1002/advs.202105957. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Dragoni S., Franco G., Regoli M., Bracciali M., Morandi V., Sgaragli G., Bertelli E., Valoti M. Gold Nanoparticles Uptake and Cytotoxicity Assessed on Rat Liver Precision-Cut Slices. Toxicol. Sci. 2012;128:186–197. doi: 10.1093/toxsci/kfs150. [DOI] [PubMed] [Google Scholar]
- 54.Climent N., García I., Marradi M., Chiodo F., Miralles L., Maleno M.J., Gatell J.M., García F., Penadés S., Plana M. Loading dendritic cells with gold nanoparticles (GNPs) bearing HIV-peptides and mannosides enhance HIV-specific T cell responses. Nanomedicine. 2018;14:339–351. doi: 10.1016/j.nano.2017.11.009. [DOI] [PubMed] [Google Scholar]
- 55.van der Beek J., de Heus C., Liv N., Klumperman J. Quantitative correlative microscopy reveals the ultrastructural distribution of endogenous endosomal proteins. J. Cell Biol. 2022;221 doi: 10.1083/jcb.202106044. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.van der Beek J., de Heus C., Sanza P., Liv N., Klumperman J. Loss of the HOPS complex disrupts early-to-late endosome transition, impairs endosomal recycling and induces accumulation of amphisomes. Mol. Biol. Cell. 2024;35 doi: 10.1091/mbc.E23-08-0328. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Rennick J.J., Johnston A.P.R., Parton R.G. Key principles and methods for studying the endocytosis of biological and nanoparticle therapeutics. Nat. Nanotechnol. 2021;16:266–276. doi: 10.1038/s41565-021-00858-8. [DOI] [PubMed] [Google Scholar]
- 58.Tong H., Huang J., Xiao Q., He B., Dong X., Liu Y., Yang X., Han D., Wang Z., Wang X., et al. High-fidelity Cas13 variants for targeted RNA degradation with minimal collateral effects. Nat. Biotechnol. 2023;41:108–119. doi: 10.1038/s41587-022-01419-7. [DOI] [PubMed] [Google Scholar]
- 59.Gupta R., Ghosh A., Chakravarti R., Singh R., Ravichandiran V., Swarnakar S., Ghosh D. Cas13d: A New Molecular Scissor for Transcriptome Engineering. Front. Cell Dev. Biol. 2022;10 doi: 10.3389/fcell.2022.866800. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Ai Y., Liang D., Wilusz J.E. CRISPR/Cas13 effectors have differing extents of off-target effects that limit their utility in eukaryotic cells. Nucleic Acids Res. 2022;50:e65. doi: 10.1093/nar/gkac159. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Bot J.F., van der Oost J., Geijsen N. The double life of CRISPR–Cas13. Curr. Opin. Biotechnol. 2022;78 doi: 10.1016/j.copbio.2022.102789. [DOI] [PubMed] [Google Scholar]
- 62.Tlotleng N., Vetten M.A., Keter F.K., Skepu A., Tshikhudo R., Gulumian M. Cytotoxicity, intracellular localization and exocytosis of citrate capped and PEG functionalized gold nanoparticles in human hepatocyte and kidney cells. Cell Biol. Toxicol. 2016;32:305–321. doi: 10.1007/s10565-016-9336-y. [DOI] [PubMed] [Google Scholar]
- 63.Oh N., Park J.H. Endocytosis and exocytosis of nanoparticles in mammalian cells. Int. J. Nanomed. 2014;9:51–63. doi: 10.2147/IJN.S26592. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Rossignol E.D., Peters K.N., Connor J.H., Bullitt E. Zika virus induced cellular remodelling. Cell. Microbiol. 2017;19 doi: 10.1111/cmi.12740. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Gruber C., Krautner L., Bergant V., Grass V., Ma Z., Rheinemann L., Krus A., Reinhardt F., Mazneykova L., Rocha-Hasler M., et al. Engineered, nucleocytoplasmic shuttling Cas13d enables highly efficient cytosolic RNA targeting. Cell Discov. 2024;10:42. doi: 10.1038/s41421-024-00672-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Hussein M., Andrade dos Ramos Z., Vink M.A., Kroon P., Yu Z., Enjuanes L., Zuñiga S., Berkhout B., Herrera-Carrillo E. Efficient CRISPR-Cas13d-Based Antiviral Strategy to Combat SARS-CoV-2. Viruses. 2023;15:686. doi: 10.3390/v15030686. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Turkevich J., Stevenson P.C., Hillier J. A study of the nucleation and growth processes in the synthesis of colloidal gold. Discuss. Faraday Soc. 1951;11:55. doi: 10.1039/DF9511100055. [DOI] [Google Scholar]
- 68.Kim D., Paggi J.M., Park C., Bennett C., Salzberg S.L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 2019;37:907–915. doi: 10.1038/s41587-019-0201-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Law C.W., Chen Y., Shi W., Smyth G.K. voom: precision weights unlock linear model analysis tools for RNA-seq read counts. Genome Biol. 2014;15:R29. doi: 10.1186/gb-2014-15-2-r29. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Ritchie M.E., Phipson B., Wu D., Hu Y., Law C.W., Shi W., Smyth G.K. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015;43:e47. doi: 10.1093/nar/gkv007. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
Data are available upon request to corresponding author.