Abstract
Ultrasound-guided quadratus lumborum block (QLB) technology has become a widely used perioperative analgesia method during abdominal and pelvic surgeries. Due to the anatomical complexity and individual variability of the quadratus lumborum muscle (QLM) on ultrasound images, nerve blocks heavily rely on anesthesiologist experience. Therefore, using artificial intelligence (AI) to identify different tissue regions in ultrasound images is crucial. In our study, we retrospectively collected 112 patients (3162 images) and developed a deep learning model named Q-VUM, which is a U-shaped network based on the Visual Geometry Group 16 (VGG16) network. Q-VUM precisely segments various tissues, including the QLM, the external oblique muscle, the internal oblique muscle, the transversus abdominis muscle (collectively referred to as the EIT), and the bones. Furthermore, we evaluated Q-VUM. Our model demonstrated robust performance, achieving mean intersection over union (mIoU), mean pixel accuracy, dice coefficient, and accuracy values of 0.734, 0.829, 0.841, and 0.944, respectively. The IoU, recall, precision, and dice coefficient achieved for the QLM were 0.711, 0.813, 0.850, and 0.831, respectively. Additionally, the Q-VUM predictions showed that 85% of the pixels in the blocked area fell within the actual blocked area. Finally, our model exhibited stronger segmentation performance than did the common deep learning segmentation networks (0.734 vs. 0.720 and 0.720, respectively). In summary, we proposed a model named Q-VUM that can accurately identify the anatomical structure of the quadratus lumborum in real time. This model aids anesthesiologists in precisely locating the nerve block site, thereby reducing potential complications and enhancing the effectiveness of nerve block procedures.
Supplementary Information
The online version contains supplementary material available at 10.1007/s10278-024-01267-8.
Keywords: Quadratus lumborum block, Deep learning, Convolutional neural network, Image segmentation, Ultrasonography
Introduction
In recent years, quadratus lumborum block (QLB) technology has greatly improved the quality of perioperative pain management techniques [1]. QLB technology is particularly popular in abdominal and pelvic surgeries, providing targeted pain relief and reducing the need for systemic opioids, thereby improving the postoperative recovery process [1]. Unlike traditional methods that rely on anatomical surface markers, ultrasound guidance enables the visualization of nerves and their surrounding tissue structures in real time, significantly improving the accuracy and safety of nerve block procedures [2]. Therefore, accurately identifying and interpreting meaningful ultrasound images are critical for successfully implementing ultrasound-guided QLB technology [3].
However, identifying meaningful ultrasound images during the ultrasound-guided QLB procedure can be challenging. First, the anatomy of the quadratus lumborum muscle (QLM) is relatively complex. The QLM is located deep and close to other anatomical structures, such as the lumbar vertebrae and transversalis fascia [4]. Distinguishing between these structures on ultrasound images requires a thorough understanding of anatomy [5]. Second, considerable individual variation is observed in ultrasound imaging. The appearance of the QLM in an image can be affected by factors such as the patient's anatomical structure and body mass index (BMI) [6]. Third, the proficiency levels of ultrasound examiners vary significantly. The mastery of ultrasound-guided techniques, particularly the precise identification of different anatomical regions in meaningful ultrasound images, necessitates specialized training and extensive experience [7, 8]. Overall, if a new technology could accurately, objectively, and quantitatively identify the QLM and its surrounding tissue structures on meaningful ultrasound images in real time, it would greatly enhance the effectiveness of the current nerve block techniques.
In recent years, advancements in high-performance computing equipment and artificial intelligence (AI) algorithms have led to the development of AI models for meaningful ultrasound images that can assist clinicians during diagnoses and treatments [9–11]. In the field of image-based tissue segmentation, machine learning, and deep learning techniques, including but not limited to random walks, region growing, convolutional neural networks (CNNs), and vision transformers, have demonstrated the ability to automatically identify anatomical structures that are relevant to nerve blocks in meaningful ultrasound images, such as the nerves, muscles, bones, and blood vessels [12]. On this basis, we conducted searches in databases such as Google Scholar, the Web of Science, and PubMed using keywords such as “artificial intelligence,” “deep learning,” and “quadratus lumborum block.” To date, we have not found any published studies on an application of AI technology for assisting in recognizing the QLB on meaningful ultrasound images. Therefore, we believe that utilizing AI technology to accurately identify the deeper and anatomically specific QLM in meaningful ultrasound images holds significant research value.
In this study, we collected ultrasound data from patients undergoing QLB procedures and proposed a deep learning model named Q-VUM. This model can adaptively extract features related to the QLB and locate them based on meaningful ultrasound images. The design of Q-VUM incorporates various advanced deep learning techniques. First, we employed transfer learning, which speeds up the model training process and reduces its dependency on computing hardware. Additionally, we enhanced the ability of the model to focus on smaller and more intricate regions by assigning different loss function weights to different target areas, thereby improving the segmentation accuracy of the model when addressing various tissue regions. In summary, the aim of our study was to develop a deep learning model that is capable of accurately segmenting the QLM and the surrounding tissue structures in meaningful ultrasound images. This model is intended to assist clinicians in more safely performing nerve block procedures, thereby reducing the risk of organ injuries in patients undergoing QLB treatments.
Materials and Methods
Patients
This research received ethical approval from the Ethics Committee of the National Cancer Center/Cancer Hospital, Chinese Academy of Medical Sciences, and Peking Union Medical College under approval number 23/139–3891. All participants in the study provided written consent after being fully informed about the procedures of the trial. Furthermore, the study was officially registered with the Chinese Clinical Trials Registry under registration number ChiCTR2300076361 before patient recruitment commenced.
During our clinical practice, we collected quadratus lumborum meaningful ultrasound images with a WISONIC ultrasound machine and obtained images from October 15, 2023, to March 15, 2024. The patients included in the study were those aged between 18 and 65 years who were undergoing elective oncological surgery.
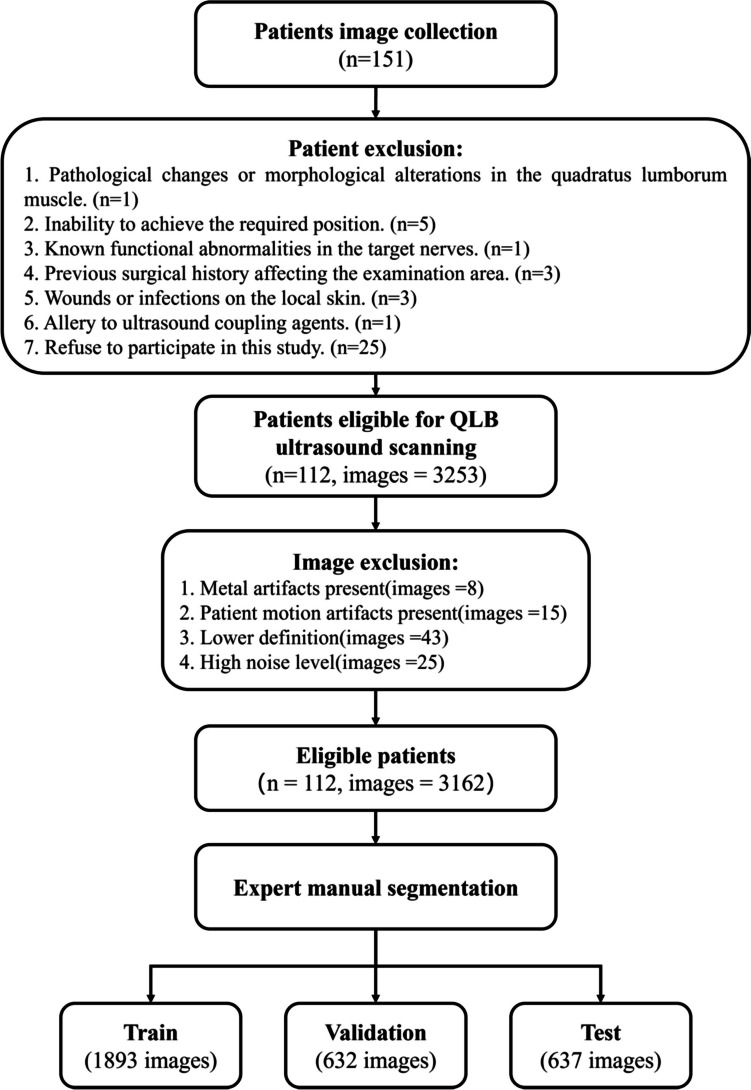
The exclusion criteria for participants in this study were as follows: (1) those with preoperative abdominal and pelvic CT scans indicating pathological changes or morphological alterations in their QLMs; (2) those who were unable to cooperate or assume the required position for the ultrasound examination due to physical reasons; (3) those with known functional abnormalities in the nerves intended for ultrasound examination; (4) those who had previous surgical procedures performed on the area intended for ultrasound examination; (5) those with the presence of wounds or infections on the local skin where the ultrasound was intended to be performed; (6) those who were allergic to the ultrasound coupling agents; and (7) those who refused to participate in this study. A total of 39 individuals who did not meet the inclusion criteria were excluded from this study. Ultimately, 112 individuals, including 62 males and 50 females, met the inclusion criteria. Additionally, we performed image quality exclusion, and the relevant details are shown in Fig. 1.
Fig. 1.
Flowchart of the overall experimental procedure
Data Acquisition
In this study, all ultrasound images used to establish the model were screened and met the inclusion criteria: they were meaningful ultrasound images that clearly displayed the three key structures and their boundaries—the quadratus lumborum muscle, vertebral bones, and transverse abdominis muscle group. The meaningful ultrasound images were collected by an anesthesiologist with more than 5 years of experience related to ultrasound-guided nerve blocks who was unaware of the research design and objectives of this study. For our ultrasound imaging process, we utilized scanners with 3–11-MHz sector ultrasonic probes (WISONIC Navi, Shenzhen, China). The meaningful ultrasound images were collected in a dedicated preoperative ultrasound imaging room with the subjects awake and cooperative. Initially, each subject was positioned on a patient transfer cart in the lateral decubitus position. Subsequently, after warm coupling gel was applied, a low-frequency sector ultrasound probe was placed between the subject’s iliac crest and their lowest rib margin. The probe was then manipulated via swiveling and rotation to clearly display the QLM on the display screen of the ultrasound machine. A slight adjustment or rotation of the probe also enabled the clear visualization of the lumbar transverse processes connected to the QLM, as well as the superficial transversus abdominis muscle group. At this juncture, by pressing the “save” button on the keyboard of the ultrasound machine, images that clearly showed the QLM, lumbar transverse processes, and transversus abdominis muscle group were stored on the hard drive of the ultrasound machine. Finally, the meaningful ultrasound images were transferred to the hard drive of another computer in DICOM format using a data transfer cable.
Data Annotation
Three anesthesiologists, each with 5 years of experience in ultrasound-guided QLB technology, manually segmented the stored meaningful ultrasound images using the ITK-SNAP Toolbox (version 3.6.0). Specifically, each anesthesiologist meticulously traced continuous lines along the perimeters of the specified anatomical features, ensuring that the markings did not extend beyond the actual boundaries of these structures. In ultrasound images, for the lumbar vertebrae boundaries, we precisely trace along the highly echogenic periosteum surface, accurately following the bright periosteum boundary line. For the quadratus lumborum muscle, we precisely trace along the relatively bright fascia that surrounds the muscle. Similarly, for the transverse abdominis muscle group, we accurately trace along the relatively bright fascia that encases the muscle group. This level of precision ensured that the model could avoid noise interference and achieve more accurate segmentation results. In addition, when the three anesthesiologists had differing opinions regarding the annotations of the relevant structures in the QLM ultrasound images, a senior anesthesiologist with over 10 years of clinical practice experience in ultrasound-guided QLB technology was employed to make the final determination. Moreover, the QLM, the bones, the external oblique, the internal oblique, and the transversus abdominis (EIT) were marked with different numeric classes.
Data Preprocessing
All images were resampled to 224 × 224 pixels and randomly divided at the patient level into three groups for this study, a training set (1893 images), a validation set (632 images), and a test set (637 images), at a proportion of 6:2:2. It is important to emphasize that multiple images from each patient were grouped into the same set to prevent potential data bias during modeling. The purpose of the training set was to facilitate the learning process of the model. The validation set consists of images used in training the model and is employed for fine-tuning the parameters. The test set included images that were previously unseen by the model and were used to evaluate the performance of the model. Additionally, to enhance the generalizability of the model, we incorporated data augmentation techniques into the training process, including (1) randomly scaling each image within a range of 0.25 to 2 times; (2) performing image distortion; (3) flipping the image horizontally and vertically; and (4) rotating the image clockwise within a range of 0 to 180°.
Construction of Q-VUM
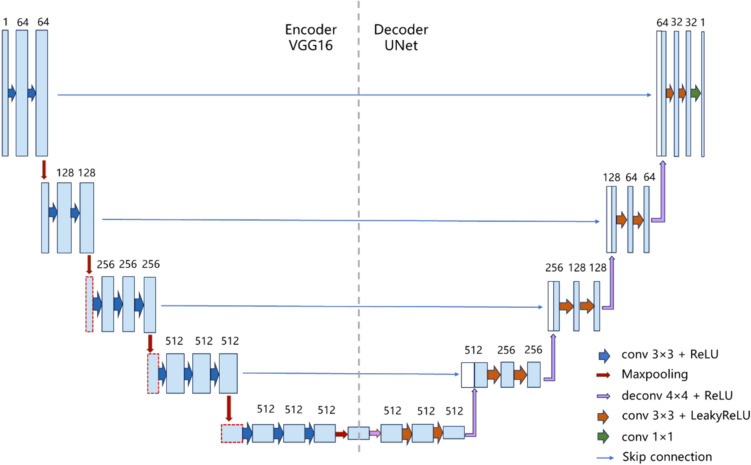
Similar to the classic UNet network [13], the Q-VUM network has an encoder-decoder neural network architecture that combines the advantages of the Visual Geometry Group 16 (VGG16) network [14] and U-shaped network. Q-VUM aims to aid in the identification of anatomical structures that are pertinent to the QLB during continuous scanning processes. For its architecture, we adopted the U-Net, which is effective for medical image segmentation, and optimized its encoder by integrating VGG16. Additionally, we kept the U-Net decoder unchanged, directly concatenating the feature maps from the downsampling layers of VGG16 with the corresponding feature maps in the decoder at each scale. By integrating the excellent feature extraction capabilities of VGG16, this network architecture not only retains the simplicity of its model structure but also enhances the flow of information between features at different levels with the help of the skip connection technology provided by UNet. This design enables the network to more accurately locate and distinguish regions of interest when processing medical images while retaining the key spatial details, which is crucial for accurately segmenting medical images. In addition, due to the existence of skip connections, high-resolution features can be maintained even in the deep layers of the network, which further improves the resulting segmentation quality, especially when edges and small objects need to be identified. In addition, we use a transfer learning method to reduce the required training time. The use of pretrained model parameters significantly reduces the training resources needed for new tasks and reduces the incurred training costs. Simultaneously utilizing a large amount of pretraining data can enhance the robustness of the model. The overall network structure is shown in Fig. 2.
Fig. 2.
The architecture of the Q-VUM
Furthermore, we chose the focal loss and dice loss as the loss functions for model training and assigned them equal weights (0.5) to address the class imbalance issues encountered during segmentation. To ensure that the model focused more on the QLM region requiring the nerve block procedure, we also adjusted the class weights of the background (w = 1), the EIT (w = 2), the QLM (w = 3), and the bones (w = 1). Finally, the detailed training procedures and parameters are provided in Supplementary Section A.
Evaluation of Q-VUM
We calculated evaluation metrics that are commonly used in segmentation tasks to evaluate the model, including the mean intersection over union (mIoU), categorical mean pixel accuracy (mPA), Dice coefficient, and accuracy (Acc). Furthermore, we selected commonly used models in medical imaging, such as U-Net [13] and U-Net + + [15], as well as the currently published studies (BPSegSys [16] and VisTR [17]) on neural block region segmentation as comparison models to evaluate the accuracy and reliability of our proposed Q-VUM model in terms of segmentation.
Results
Clinical Characteristics
The study comprised 112 participants, consisting of 62 men and 50 women. On average, the participants had a height of 167.61 cm with a standard deviation of 8.87, a weight of 68.21 kg with a standard deviation of 13.22, and an age of 51.43 years with a standard deviation of 9.25. Detailed information about the patients in each cohort can be found in Table 1, with no significant differences among the clinical characteristics of different cohorts.
Table 1.
Clinical characteristics of the patients in each cohort
| Characteristics | Total (n = 112) | Training cohort (n = 68) | Validation cohort (n = 22) | P value* | Test cohort (n = 22) | P value** |
|---|---|---|---|---|---|---|
| Age, years, mean ± SD | 51.4 ± 9.3 | 49.5 ± 9.5 | 52.0 ± 9.6 | 0.273 | 52.6 ± 8.6 | 0.174 |
| Gender, n (%) | 0.792 | 0.630 | ||||
| Male | 62 (55.4) | 38 (55.9) | 13 (59.1) | 11 (50.0) | ||
| Female | 50 (44.6) | 30 (44.1) | 9 (40.9) | 11 (50.0) | ||
| Height (cm) | 167.6 ± 8.9 | 166.0 ± 8.7 | 167.1 ± 9.6 | 0.633 | 166.2 ± 9.0 | 0.949 |
| Weight (kg) | 68.2 ± 13.2 | 66.2 ± 12.5 | 65.4 ± 12.4 | 0.793 | 64.7 ± 11.7 | 0.613 |
Categorical data are shown as numbers (%), and continuous data are shown as means ± SDs
*The P value is the test result obtained with respect to the training cohort and the validation cohort
**The P value is the test result obtained with respect to the training cohort and the test cohort
The Performance of Q-VUM
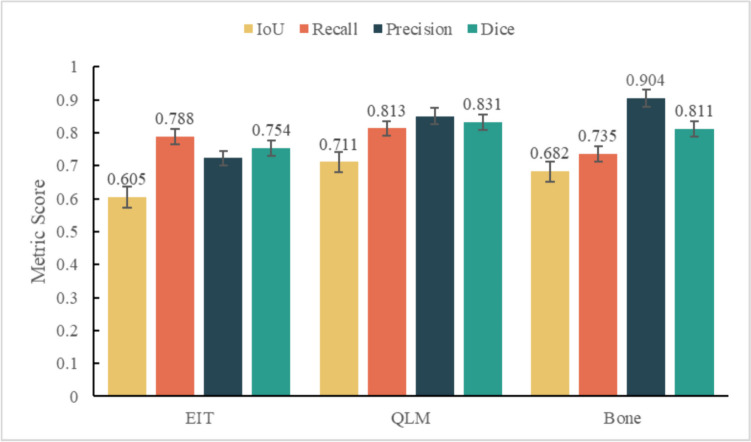
The performance metrics achieved by the proposed model for multitarget segmentation, denoted as the mIoU, mPA, dice, and accuracy, were 0.734, 0.829, 0.841, and 0.944, respectively. In addition, we compiled the performance attained by the model predictions for the EIT, the QLM, and bones and plotted them in a bar chart (Fig. 3). The results indicate that among the three target regions, the QLM exhibited the highest IoU (0.711 vs. 0.605 and 0.682) and recall (0.813 vs. 0.788 and 0.735) values. Although the precision was slightly lower than that attained for bones (0.904), it still reached 0.850.
Fig. 3.
The results of the Q-VUM predictions produced for each region
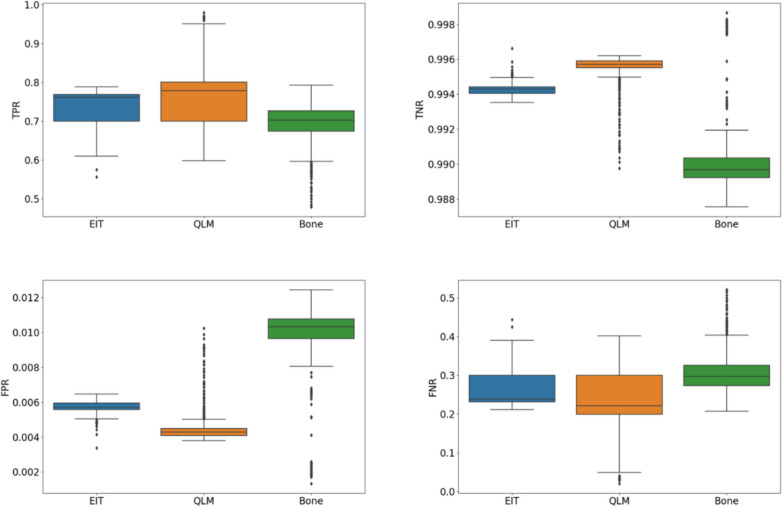
To further assess the usability of the model, we calculated four evaluation indicators: the true-positive rate (TPR), true-negative rate (TNR), false-positive rate (FPR), and false-negative rate (FNR). The relevant results are summarized in Table 2, and we can see that the QLM area yielded the highest TPR and TNR values, which were 0.813 and 0.996, respectively. This shows that our algorithm has a strong ability to correctly identify both samples that are actually QLM areas and samples that are not actually QLM areas. In addition, the QLM area had the lowest FNR, which was 0.163. This means that the number of false positives generated by the algorithm during prediction, that is, non-QLM regions that were incorrectly identified as QLMs, and the number of false negatives, that is, the number of actual QLM regions that could not be identified, were both very small. The TPRs and TNRs produced for the EIT and bone areas reached high levels, while the FPRs and FNRs were low, indicating that the segmentation accuracy and sensitivity of the model were high. At the same time, we also used the Turkey test to calculate the significant differences of each indicator (Fig. 4), and the results showed that there were significant differences.
Table 2.
Evaluation index statistics
| Areas | TPR | TNR | FPR | FNR |
|---|---|---|---|---|
| EIT | 0.788 | 0.994 | 0.232 | 0.163 |
| QLM | 0.813 | 0.996 | 0.126 | 0.163 |
| Bone | 0.735 | 0.990 | 0.072 | 0.245 |
| P value* | < 0.001 | < 0.001 | < 0.001 | < 0.001 |
EIT, external oblique muscle, internal oblique muscle, transversus abdominis muscle; QLM, quadratus lumborum block; TPR, true-positive rate; TNR, true-negative rate; FPR, false-positive rate; FNR, false-negative rate. *The P value is calculated by ANOVA for the test result for the same indicator
Fig. 4.
Significant differences between different groups (Turkey test)
Visualization of Several Cases
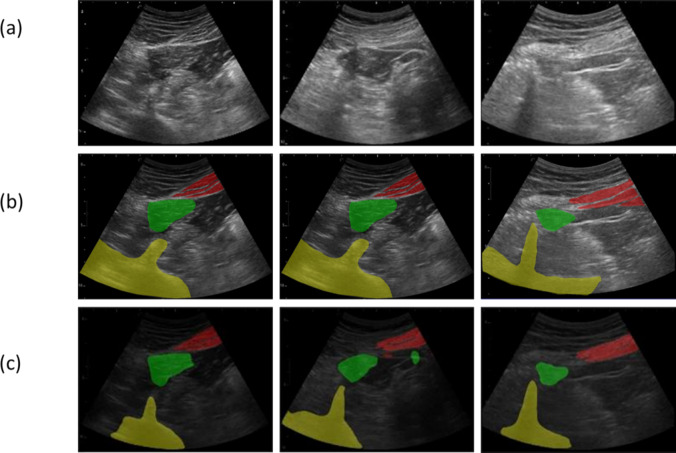
We selected three representative patient images from the test set and conducted visualizations (Fig. 5). The prediction outcomes of our model demonstrated a relatively high level of agreement with the delineation standards set by medical professionals, particularly in the case of the QLM area, where 85% of the pixel areas fell within the target mask. In addition, the Q-SUM was able to accurately segment the EIT region, identify its location in the upper-right quadrant of the QLM, and capture its fine structures. In terms of the overall positions of nonpenetrable structures such as the skeletal system, the segmentation results were generally accurate, with only small areas left unsegmented. These areas are typically located in the lower-left portion of an ultrasonic image, but imprecisely segmenting them has less of an effect on nerve block procedures.
Fig. 5.
Visualization of the segmentation results produced in different areas. The original image (a), the manually segmented image (b), and the prediction result (c) of the Q-VUM. In the ultrasound images, green, red, and yellow represent the QLM, EIT, and bones, respectively
Comparison Experiment
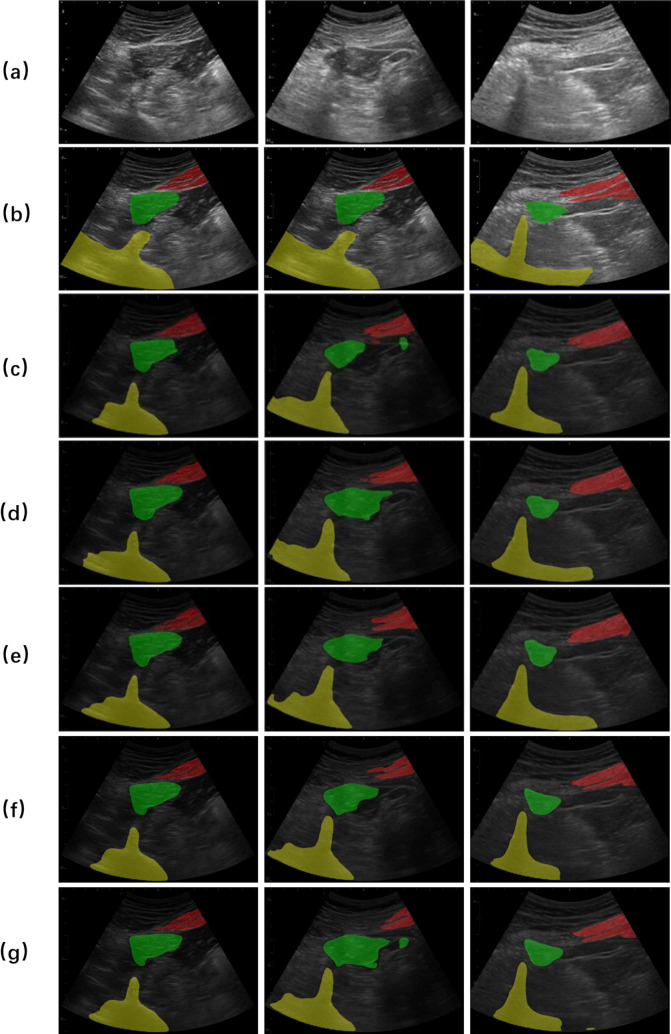
For the model comparison experiment, we first conducted a statistical analysis of the results obtained from different models (Table 3). Compared to the commonly used models in medical image segmentation, Q-VUM demonstrated superior segmentation performance. Additionally, we selected three ultrasound images from the test set for case visualizations (Fig. 6). According to the images, Q-VUM excelled at accurately identifying and segmenting the QLM region while preserving its intricate structure without significantly over-segmenting it. Particularly, in the second image, compared to the images manually delineated by medical professionals, the results of UNet and UNet + + exhibited larger segmentation areas. The other two models also showed different degrees of mis-segmentation.
Table 3.
Summary of the network performance metrics achieved on the test set
| Model | mIoU | t* | p** | mPA | t* | p** | Dice | t* | p** | Acc | t* | p** |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| UNet [13] | 0.720 | 29.35 | < 0.05 | 0.820 | 11.02 | < 0.05 | 0.831 | 29.55 | < 0.05 | 0.942 | 24.67 | < 0.05 |
| UNet + + [15] | 0.720 | 37.65 | < 0.05 | 0.812 | 63.36 | < 0.05 | 0.831 | 39.00 | < 0.05 | 0.942 | 31.90 | < 0.05 |
| BPSegSys [16] | 0.729 | -5.12 | < 0.05 | 0.819 | 2.33 |
0.02 (< 0.05) |
0.837 | 2.01 |
0.04 (< 0.05) |
0.944 | 2.61 | < 0.05 |
| VisTR [17] | 0.727 | 6.23 | < 0.05 | 0.822 | 4.64 | < 0.05 | 0.835 | 8.14 | < 0.05 | 0.944 | 3.20 | < 0.05 |
| Q-VUM | 0.734 | - | - | 0.829 | - | - | 0.841 | - | - | 0.944 | - | - |
*t value is calculated by paired T-test for the test results of Q-VUM and other models
**P value is calculated by ANOVA for the test results of Q-VUM and other models calculated
Fig. 6.
Comparison among the results produced by different models. The original image (a), the corresponding gold standard (b), the prediction result of the Q-VUM (c), the prediction result of UNet (d), the prediction result of UNet + + (e), the prediction result of BPSegSys (f), and is the prediction of VisTR (g)
For real-time segmentation, the Q-VUM model proposed in this study showed significant performance advantages, with a segmentation rate of 42.40 frames per second (fps), which meets the needs of clinical surgery and exceeds other models compared in this study (Table 4). This means that Q-VUM can also meet the needs of real-time processing, and the higher segmentation frame rate ensures that the system can respond and process quickly in dynamic and complex clinical environments.
Table 4.
Comparison on the processing speed of different models
| UNet | UNet + + | BPSegSys | VisTR | Q-VUM | |
|---|---|---|---|---|---|
| Speed(fps) | 40.37 | 38.28 | 40.28 | 38.08 | 42.40 |
Discussion
QLB technology is a nerve blockade analgesia method with promising clinical application prospects [18]. Its analgesic effectiveness in abdominal and pelvic surgeries is even superior to that of the traditional transversus abdominis plane nerve blockade method [19]. However, due to the deep anatomical location of the QLM and its significant shape variations among different patients, accurately identifying and locating the QLM under ultrasound guidance remains a challenging and troublesome issue for clinicians. In this study, we proposed an AI-based recognition model, Q-VUM, to accurately identify the target structures on meaningful ultrasound images for QLB procedures. The proposed Q-VUM model achieved relatively satisfactory performance in terms of identifying the three crucial anatomical structures in the QLB, namely, the QLM, the transverse processes of the lumbar vertebrae, and the lateral abdominal muscle group (the external oblique, internal oblique, and transversus abdominis).
In this study, we implemented several measures to ensure that different annotators could consistently label the same segmentation labels. First, we established a unified ultrasound image segmentation standard that all annotators adhered to. Second, we provided standardized training to ensure that all annotators accurately understood and applied this segmentation standard. Additionally, we appointed several senior doctors with over 10 years of experience in ultrasound-guided quadratus lumborum nerve block techniques as quality control inspectors for the image segmentation process. These experts reviewed each segmentation label marked by the annotators. If any segmented images did not meet the unified segmentation standard, they were re-labeled and re-checked until the segmentation labels conformed to our established standard. The Q-VUM model achieved the best performance with respect to identifying the key target of the QLM (with an mIoU of 0.734, an mPA of 0.829, a dice coefficient of 0.841, and an Acc of 0.944) in comparison with other commonly used models. This indicates that the Q-VUM model could accurately distinguish the QLM from other anatomical structures regardless of the challenges caused deep anatomical locations and shape variations. To our knowledge, no published study has focused on automatically recognizing the quadratus lumborum as a target structure for aiding in the QLB nerve blockade process. Previous studies have employed deep learning models to automatically segment brachial plexus nerves, achieving an IoU of 0.535, which was notably lower than the performance achieved in our study [20]. The segmentation performance on different tissues in quadratus lumborum blocks is affected by the high-level noise and low-contrast characteristics of ultrasound images to a certain extent. As reported in previous studies [21, 22], stochastic resonance constructively used the input noise to enhance the signal, which might be helpful to improve the segmentation performance of the Q-VUM model if employed in future study.
In addition, we further evaluated the performance of the Q-VUM model in terms of identifying the QLM through four key evaluation indicators: the TPR, TNR, FPR, and FNR. The proposed Q-VUM model achieved a TPR of 0.813, a TNR of 0.996, an FPR of 0.126, and an FNR of 0.163 when identifying QLM regions. These high TPR and TNR values demonstrated the strong ability of the Q-VUM model to accurately identify QLM regions and non-QLM regions. The prediction results of our model yielded relatively high consistency according to the delineation standards set by medical professionals, especially in the QLM area, where 85% of the pixel area fell within the target mask. This result demonstrated that the proposed Q-VUM model had high sensitivity and specificity. On the other hand, the low FPR and low FNR metrics showed that the Q-VUM model was highly accurate with respect to identifying QLM regions and rarely incorrectly identified QLM regions as non-QLM areas; at the same time, non-QLM areas were rarely misjudged as QLM areas. This shows that our prediction area was more conservative than the actual nerve block area, thereby reducing the risk of accidentally affecting other areas during nerve block operations and enhancing the safety of the operation.
Our study also improved upon the algorithmic structure employed in previous studies. Huang et al. [23] used UNet to conduct ultrasound-guided femoral nerve block segmentation. Their model achieved an IoU of 0.713 on a training set, 0.633 on a development set, and 0.638 on a test set. The segmentation accuracy achieved on the test set was 83.9%. In a tenfold cross-validation experiment, the median IoU was 0.656, and the accuracy was 88.4%. Miyatake et al. [24] used a dilated UNet architecture for supraclavicular nerve block segmentation, achieving an average dice score of 0.56 on images acquired from six subjects. For some images, the dice scores were less than 0.40, but the network still correctly identified the potential nerve distributions. Compared with UNet and Detailed-UNet, the Q-VUM model proposed in this study used fewer model parameters, which reduces the imposed computational burden and resource consumption levels during the training process, making model training easier and more efficient. In addition, a transfer learning method was adopted to use the knowledge learned by the pretrained model from related tasks to accelerate and improve the training process of the current task. This consideration also explains why the Q-VUM model proposed in this study outperformed the methods developed in previous studies.
Real-time processing is necessary for the clinical application of segmentation model in the operation of quadratus lumborum block, which would limit the computational complexity of the model. Nevertheless, the limited computational complexity of the segmentation model might lead to compromising segmentation accuracy. As indicated in previous studies [25–27], parallel computation would be very useful to simultaneously ensure real-time processing and relatively high computational complexity to apply the Q-VUM model in the actual clinical practice in the future. Additionally, machine learning-based segmentation methods have the advantages of strong interpretability and low computational requirements when dealing with small datasets [28–30], and they may produce results that surpass those of deep learning methods. However, for scenarios like nerve blocks, which demand high precision and real-time performance, deep learning offers better scalability and the potential for multi-modal data integration. In future research, we will continue to optimize Q-VUM and actively promote its clinical translation.
Our study has several limitations. Firstly, due to potential differences in equipment (such as low-frequency and high-frequency ultrasound probe imaging), environment, or patient populations, Q-VUM requires further validation using multi-center data to confirm its effectiveness in real clinical settings. Secondly, the amount of data used to train Q-VUM still has room for improvement. Developing the foundation model for nerve blocks based on large-scale ultrasound images from a broad patient base is an essential step for the future. Lastly, Q-VUM is currently only capable of segmenting the block region within the image. The next research focus will be on integrating anatomical knowledge into the model using 3D imaging technologies like NeRF (Neural Radiance Fields) [31], enabling physicians to more efficiently identify the optimal block angle with an ultrasound probe. Overall, compared to nerve block techniques targeting more distal spinal nerves (such as ultrasound-guided transversus abdominis plane block), ultrasound-guided QLB offers superior surgical analgesia outcomes because it can block the transmission of nociceptive signals closer to the intervertebral foramen [17]. Based on this, we developed Q-VUM, which features low computational weight, fast training speed, and high accuracy, providing a promising outlook for research into ultrasound-guided QLB techniques.
In summary, we developed a deep learning model, Q-VUM, for automatically identifying the anatomical structure of the quadratus lumborum in real time, effectively assisting anesthesiologists in performing nerve block procedures. We firmly believe that AI has substantial potential for providing transformative applications involving ultrasound-guided QLB technology, paving the way for improved patient outcomes and surgical efficiency.
Supplementary Information
Below is the link to the electronic supplementary material.
Author Contribution
Qiang Wang and Bingxi He wrote the manuscript. Zhenchao Tang and Bingxi He developed the network architecture, the data/modeling infrastructure, and the training and testing setups. Zhenchao Tang designed the overall structure of the study. Hui Zheng provided clinical expertise and guidance for the study design. Jie Yu, Bowen Zhang, Jingchao Yang, Jin Liu, Xinwei Ma, and Shijing Wei created the clinical datasets, interpreted the data, and defined the clinical labels. All authors read and approved the final manuscript.
Funding
This work was supported by the Beijing Natural Science Foundation (no. 7232133); the CAMS Innovation Fund for Medical Sciences (CIFMS; no. 2022-I2M-C&T-B-086); the Beijing Hope Run Special Fund of the Cancer Foundation of China (no. LC2022A05); the Young Elite Scientists Sponsorship Program of CAST (no. 2023QNRC001); the National Key R&D Program of China (no. 2021YFA1301603); the Fundamental Research Funds for the Central Universities (no. YWF-23-Q-1074); the Key-Area Research and Development Program of Guangdong Province (no. 2021B0101420005); the National Natural Science Foundation of China (no. 82302317); and the China Postdoctoral Science Foundation (2021M700341).
Data Availability
The data analyzed in the current study are not publicly available for patient privacy purposes but are available from the corresponding authors upon reasonable request.
Declarations
Ethics Approval
This study was performed in line with the principles of the Declaration of Helsinki. Approval was granted by the Ethics Committee of the National Cancer Center/Cancer Hospital, Chinese Academy of Medical Sciences, and Peking Union Medical College under approval number 23/139–3891.
Consent to Participate
Written informed consent was obtained from the parents.
Competing Interests
The authors declare no competing interests.
Footnotes
Qiang Wang, Bingxi He, and Jie Yu are contributed equally to this work as co-first authors.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Contributor Information
Hui Zheng, Email: zhenghui0715@hotmail.com.
Zhenchao Tang, Email: tangzhenchao@buaa.edu.cn.
References
- 1.Jin Z, Liu J, Li R, et al. Single injection quadratus lumborum block for postoperative analgesia in adult surgical population: a systematic review and meta-analysis. J Clin Anesth 62:109715, 2020. [DOI] [PubMed] [Google Scholar]
- 2.Saranteas T, Koliantzaki I, Savvidou O, et al. Acute pain management in trauma: anatomy, ultrasound-guided peripheral nerve blocks and special considerations. Minerva Anestesiol 85:763-773, 2019. [DOI] [PubMed] [Google Scholar]
- 3.Ahmed A, Fawzy M, Nasr MAR, et al. Ultrasound-guided quadratus lumborum block for postoperative pain control in patients undergoing unilateral inguinal hernia repair, a comparative study between two approaches. BMC Anesthesiol 19:184, 2019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Alver S, Bahadir C, Tahta AC, et al. The efficacy of ultrasound-guided anterior quadratus lumborum block for pain management following lumbar spinal surgery: a randomized controlled trial. BMC Anesthesiol 22:394, 2022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Ueshima H, Otake H, Lin JA. Ultrasound-guided quadratus lumborum block: an updated review of anatomy and techniques. Biomed Res Int 2017:2752876, 2017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Balocco AL, López AM, Kesteloot C, et al. Quadratus lumborum block: an imaging study of three approaches. Reg Anesth Pain Med 46:35-40, 2021. [DOI] [PubMed] [Google Scholar]
- 7.Dickson R, Duncanson K, Shepherd S. The path to ultrasound proficiency: a systematic review of ultrasound education and training programmes for junior medical practitioners. Australas J Ultrasound Med 20:5-17, 2017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Kefala-Karli P, Sassis L, Sassi M, et al. Introduction of ultrasound-based living anatomy into the medical curriculum: a survey on medical students’ perceptions. Ultrasound J 13:47, 2021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Salehi AW, Khan S, Gupta G, et al. A study of CNN and transfer learning in medical imaging: advantages, challenges, future scope. Sustainability 15:5930, 2023. [Google Scholar]
- 10.Esteva A, Kuprel B, Novoa RA, et al. Dermatologist-level classification of skin cancer with deep neural networks. Nature 542:115-118, 2017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Beam AL, Kohane IS. Big data and machine learning in health care. JAMA 319:1317-1318, 2018. [DOI] [PubMed] [Google Scholar]
- 12.Lloyd J, Morse R, Taylor A, et al. Artificial intelligence: innovation to assist in the identification of sono-anatomy for ultrasound-guided regional anaesthesia. Advances in Experimental Medicine and Biology 1356:117-140, 2022. [DOI] [PubMed] [Google Scholar]
- 13.Ronneberger O, Fischer P, Brox T. U-net: convolutional networks for biomedical image segmentation. In: Medical Image Computing and Computer-Assisted Intervention – MICCAI 2015: 18th International Conference, Munich, Germany, October 5–9, 2015, Proceedings, Part III, Springer International Publishing, pp 234–241, 2015.
- 14.Simonyan K, Zisserman A. Very deep convolutional networks for large-scale image recognition. arXiv preprint arXiv:1409.1556, 2014.
- 15.Zhou Z, Rahman Siddiquee MM, Tajbakhsh N, et al. Unet++: a nested u-net architecture for medical image segmentation. In: Deep Learning in Medical Image Analysis and Multimodal Learning for Clinical Decision Support: 4th International Workshop, DLMIA 2018, and 8th International Workshop, ML-CDS 2018, Held in Conjunction with MICCAI 2018, Granada, Spain, September 20, 2018, Proceedings, Springer International Publishing, pp 3–11, 2018. [DOI] [PMC free article] [PubMed]
- 16.Wang Y, Zhu B, Kong L, et al. BPSegSys: A brachial plexus nerve trunk segmentation system using deep learning. Ultrasound in Medicine & Biology, 50(3): 374-383, 2024. [DOI] [PubMed] [Google Scholar]
- 17.Gujarati K R, Bathala L, Venkatesh V, et al. Transformer-based automated segmentation of the median nerve in ultrasound videos of wrist-to-elbow region. IEEE Transactions on Ultrasonics, Ferroelectrics, and Frequency Control, 2023. [DOI] [PubMed]
- 18.Elsharkawy H, El-Boghdadly K, Barrington M. Quadratus Lumborum Block: Anatomical Concepts, Mechanisms, and Techniques. Anesthesiology 130:322-335, 2019. [DOI] [PubMed] [Google Scholar]
- 19.Blanco R, Ansari T, Riad W, Shetty N. Quadratus Lumborum Block Versus Transversus Abdominis Plane Block for Postoperative Pain After Cesarean Delivery: A Randomized Controlled Trial. Reg Anesth Pain Med 41:757-762, 2016. [DOI] [PubMed] [Google Scholar]
- 20.Wang Y, Zhu B, Kong L, et al. BPSegSys: A Brachial Plexus Nerve Trunk Segmentation System Using Deep Learning. Ultrasound Med Biol 50:374-383, 2024. [DOI] [PubMed] [Google Scholar]
- 21.Mohanty S. and Dakua SP. Toward Computing Cross-Modality Symmetric Non-Rigid Medical Image Registration, IEEE Access, vol. 10, pp. 24528–24539, 2022.
- 22.Dakua, Sarada Prasad. LV Segmentation using Stochastic Resonance and Evolutionary Cellular Automata, International Journal of Pattern Recognition and Artificial Intelligence, World Scientific, vol. 29, no. 3, pp. 1557002:1–26, 2015.
- 23.Esfahani SS, Zhai X, Chen M, et al. Lattice-Boltzmann Interactive Blood Flow Simulation Pipeline, International Journal of Computer Assisted Radiology and Surgery, Springer, vol.15, pp. 629-639, 2020. [DOI] [PubMed] [Google Scholar]
- 24.Zhai X, Chen M, Fsfahani SS, et al. Heterogeneous System-on-Chip based Lattice- Boltzmann Visual Simulation System," Systems Journal, IEEE, vol. 14, no. 2, pp. 1592-1601, 2020. [Google Scholar]
- 25.Zhai X, Amira A, Bensaali F, et al. Zynq SoC based Acceleration of the Lattice Boltzmann Method, Concurrency and Computation: Practice and Experience, Wiley, col. 31, issue 17, 2019.
- 26.Huang C, Zhou Y, Tan W, et al. Applying deep learning in recognizing the femoral nerve block region on ultrasound images. Ann Transl Med 7:453, 2019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Miyatake M, Nerella S, Simpson D, et al. Automatic Ultrasound Image Segmentation of Supraclavicular Nerve Using Dilated U-Net Deep Learning Architecture. arXiv preprint arXiv:2208.05050, 2022.
- 28.Dakua SP, Sahambi J S. LV Contour Extraction from Cardiac MR Images Using Random Walk Approach, Proc. of IEEE International Advance Computing Conference, Patiala, pp. 228 - 233, 2009.
- 29.Dakua SP, Sahambi JS. Automatic Contour Extraction of Multi-labeled Left Ventricle from CMR Images Using Cantilever Beam and Random Walk Approach, Cardiovascular Engineering, Springer, vol. 10, pp. 30-43, 2010. [DOI] [PubMed]
- 30.Dakua SP, Sahambi JS. Detection of Left Ventricular Myocardial Contours from Ischemic Cardiac MR Images, IETE Journal of Research, Taylor & Francis, vol. 57, pp. 372-384, 2011.
- 31.Mildenhall B, Srinivasan PP, Tancik M, et al. Nerf: Representing scenes as neural radiance fields for view synthesis. Communications of the ACM, 65(1): 99-106, 2021. [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The data analyzed in the current study are not publicly available for patient privacy purposes but are available from the corresponding authors upon reasonable request.