ABSTRACT
Uropathogenic Escherichia coli (UPEC) is one of the leading causes of nosocomial infections and urinary tract infections (UTIs), capable of inducing a spectrum of conditions ranging from acute bladder cystitis to chronic pyelonephritis. The virulence arsenal of UPEC includes factors, such as curli fimbriae, hemolysin, motility elements and metallophores. Moreover, UPEC can form biofilm‐like communities and quiescent intracellular reservoirs within host tissues, contributing to recurrent and persistent infections. This study investigates the antibiofilm and antimicrobial activities of two halogen‐substituted aniline derivatives, 4‐bromo‐3‐chloroaniline (4B3CA) and 3,5‐dibromoaniline (3,5‐DBA), against UPEC. The compounds demonstrated minimum inhibitory concentrations (MICs) of 200 μg/mL for 4B3CA and 100 μg/mL for 3,5‐DBA, with both exhibiting a biofilm inhibition IC50 value of 10 μg/mL. Additionally, these derivatives showed antibiofilm activity against ESKAPE pathogens. Treatment with 4B3CA and 3,5‐DBA led to significant downregulation of UPEC virulence‐ and biofilm‐related genes, including those involved in curli production, fimbrial adhesion, motility, iron acquisition, quiescent colony formation, and stress response. Interestingly, a mild upregulation of hlyA, csrA and uvrY was noted, alongside a marked downregulation of the adenylate cyclase genes cyaA and crp. These findings suggest that inhibition of adenylate cyclase activity may be a primary mode of action, leading to both antimicrobial and antibiofilm effects. The presence of halogen atoms in these compounds appears to enhance their binding affinity to adenylate cyclase through stabilising halogen bond interactions. Furthermore, 3D‐QSAR analysis indicates that electrostatic favourability at the third and fourth positions of the aniline ring is critical for bioactivity. Finally, in silico ADMET profiling and cytotoxicity assessments using Caenorhabditis elegans suggest that these aniline derivatives hold promise as therapeutic candidates, warranting further investigation.
Keywords: aniline, biofilm, E. coli , ESKAPE, halogen, UPEC, virulence
This study evaluates the antimicrobial and antibiofilm activities of halogen‐substituted aniline derivatives, 4‐bromo‐3‐chloroaniline (4B3CA) and 3,5‐dibromoaniline (3,5‐DBA), against uropathogenic E. coli (UPEC) and some ESKAPE pathogens, with MICs of 200 and 100 μg/mL, respectively, and a common biofilm IC50 of 10 μg/mL. These derivatives inhibited key UPEC virulence factors, potentially disrupting adenylate cyclase, whereas 3D‐QSAR analysis and in silico ADMET profiling suggest their therapeutic potential pending further toxicity assessments.

1. Introduction
Urinary tract infections (UTIs) are one of the most common bacterial and nosocomial infections, with estimated cases up to 400 million resulting in 230,000 mortalities worldwide. The sheer number of outpatient cases and the associated morbidities are a grave cause for alarm, with an annual medical burden of 1.6 billion USD in the United States alone, and EUR 5700 per case of complicated UTI caused by multidrug resistant (MDR) bacteria in Europe (Whelan et al. 2023). Uropathogenic Escherichia coli (UPEC) is the foremost contributor, accounting for 50%–90% of UTI infections among other pathogens, such as Enterococcus faecalis , Klebsiella pneumoniae , Pseudomonas aeruginosa and Proteus mirabilis (Shah et al. 2019; Zhang et al. 2022; Moreland et al. 2023). UPEC is the primary focus of this study and is an opportunistic Gram‐negative bacterium, causing symptoms like urosepsis syndrome and bladder cystitis, which can lead to pyelonephritis in chronic and unattended infections (García et al. 2023).
UPEC pathogenesis is capable of host‐immune defence evasion, as well as actively invading and infecting the urinary tract with its host of virulence factors. UPEC pathogenesis begins with colonisation of the urethra and subsequent propagation to the bladder lumen wherein the bacteria multiply in urine as planktonic cells. These cells then adhere to the bladder epithelial layer and interact with host defences, leading to heightened proteinaceous polysaccharide matrix production and replication, forming biofilm clusters called intracellular bacterial communities (IBCs). IBCs aid in further invasion of the urothelium to form quiescent intracellular reservoirs (QIRs), causing recurrent infections, bacteremia or septicemia, kidney ascension and ultimately pyelonephritis (Terlizzi et al. 2017).
The vast virulence factor armamentarium of UPEC includes fimbriae, which facilitate attachment to various surfaces like the type 1 fimbriae (bladder) and P fimbriae (Kidney), encoded by the fim and pap operons respectively (García et al. 2023). An important surface factor, antigen 43 (Ag43), is an autotransporter protein that promotes cell aggregation for biofilm formation (Ageorges et al. 2023). Curli are amyloid fibres that play a crucial role in the biofilm architecture, lending structural support to the extracellular matrix along with cellulose, and are regulated by the csg operon (García et al. 2023). UPEC releases toxins like α‐hemolysin, which are cytotoxic to epithelium, leading to hematuria for ancillary nutrient procurement (Hreha et al. 2024). Metal acquisition systems involve siderophores like enterobactin and aerobactin that aid in the chelation of iron in the iron‐deficient urinary tract along with yersiniabactin, which can act as a copper scavenging chalkophore during starvation or to mitigate copper‐induced cytotoxicity (Heffernan 2023). UPEC also employs flagella for motility and ascension to upper urinary tracts, with their biosynthesis primarily regulated by the fliC gene (Hirakawa et al. 2023). Additionally, UPEC utilises several two‐component phosphotransfer systems, with the BarA‐UvrY system being the chief regulator of several virulence factors like lipopolysaccharide and hemolysin production, flagellar biosynthesis and biofilm formation (Brannon et al. 2024). UPEC also has a functioning AI‐2 autoinducer quorum sensing system, whose mechanism and interaction pathways are yet to be completely elucidated but it can also influence biofilm formation in UPEC (Capper‐Parkin et al. 2023). In UPEC, cyaA encodes adenylate cyclase, which synthesises cAMP, a key secondary messenger. cAMP binds to cAMP receptor protein (CRP), forming a complex that regulates the transcription of numerous genes involved in UPEC pathogenesis. The cyaA‐crp regulatory system can control the expression of fimbriae, motility, biofilm formation, iron acquisition systems and metabolic adaptation during infection. These factors are essential for UPEC colonisation, survival in nutrient‐limited environments and evasion of host immune responses (Donovan et al. 2013).
Biofilm formation and the aforementioned virulence factors significantly enhance the pathogenesis portfolio of UPEC, affording it great survival in various environments (Long et al. 2024). Novel avant‐garde treatment options, such as vaccines, curlicides and pilicides are several years away from feasible use in clinical settings, leaving traditional chemotherapy as the only short‐term viable option to treat UTIs. The caveat of traditional chemotherapy is the significant and indispensable use of antibiotics to treat UTIs, leading to antimicrobial resistance. This results in several classes of antibiotics, such as carbapenems, aminoglycosides, and fluoroquinolones, being ineffective in treating recalcitrant pathogens (Liu et al. 2022). Therefore, a novel approach to counteract UTIs that mitigates multiple virulence factors and evasion strategies of the pathogens is of paramount need. This strategy can also prove to be fruitful in similarly thwarting ESKAPE pathogen infections, which have been designated as high priority by the World Health Organization (WHO).
Aniline is an aromatic amine compound consisting of a phenyl ring with an amino group substitution. Carbon atoms in the phenyl ring contribute to resonance, and the electronegative amino group contributes to the inductive effect. The amino group in the aniline scaffold plays a pivotal role from a supramolecular chemistry perspective, enabling aniline to form hydrogen bonds. Many supramolecular structures used in dye, antioxidants, rubber and pharmaceutical industries are synthesised with aniline as a reactant or intermediate (Mary et al. 2020). Aniline is an important synthesis intermediate or backbone of several drugs, such as phenacetin, paracetamol, procainamide and sulfonamides owing to its simple structure and ease of hydrogen bond formation (Atapalkar and Kulkarni 2023; Liang et al. 2023; Pervaiz et al. 2024). Halogenation is a key strategy to improve the bioactivity of organic chemicals (Islam et al. 2024). Recently, halogenation has been touted to increase antimicrobial efficacy especially against MDR pathogens (Faleye et al. 2024). Although halogenated anilines were researched in the early 1970s for their antimicrobial efficacies in a limited capacity, no substantial progress or application was reported thereafter. Anilines are considered a privileged scaffold in medicinal chemistry due to their intrinsic bioactivity and the ease with which they can be functionalised with various chemical groups. Several aniline derivatives have demonstrated the ability to interfere with key bacterial targets, including dihydrofolate reductase (Tripathi et al. 2017) and DNA gyrase (Zidar et al. 2015). Additionally, they can compromise bacterial membrane integrity in Rubrivivax benzoatilyticus (Mohammed et al. 2020), inhibit efflux pumps and suppress biofilm formation in Pseudomonas aeruginosa (Moore et al. 2014; Gallarato et al. 2017), all of which are key mechanisms used by UPEC to resist antimicrobial therapy. Halogenation has been shown to further enhance the antimicrobial potency of aniline derivatives. Moreover, these compounds have recently exhibited effectiveness against other Gram‐negative pathogens such as Vibrio species (Sathiyamoorthi et al. 2025). These combined attributes prompted us to select halogenated aniline derivatives for evaluation against UPEC.
Therefore, this study explores halogenated aniline derivatives for their antimicrobial, antibiofilm and antivirulence potential against uropathogens, primarily focusing on UPEC and assessing their potential as antimicrobial and antivirulence chemotherapeutic agents to treat UTIs. In this study, we evaluated aniline and 69 of its halogenated derivatives for their dual antibiofilm and antimicrobial activity against UPEC, of which 4‐bromo 3‐chloroaniline (4B3CA) and 3,5‐dibromoaniline (3,5‐DBA) were found to be the most potent. Their efficacy against other virulence factors of UPEC, such as motility, curli production, quiescence, iron scavenging and hemolysis, was gauged. These hits were also subjected to in silico ADME and toxicity profiling as well as Caenorhabditis elegans toxicity assay.
2. Materials and Methods
2.1. Aniline and Its Derivatives
Aniline and its 69 halogenated derivatives were purchased from Combi Blocks (San Diego, CA, USA). The chemicals were dissolved in DMSO (dimethyl sulfoxide) to prepare 100 mg/mL stocks and were stored at −20°C. DMSO has no effect on bacterial growth or biofilm at this dilution percentage (< 0.1%). All the aniline derivatives used in the study are listed in Table S1.
2.2. Microbial Growth Conditions
The clinical isolate of uropathogenic E. coli O6:H1 strain CFT073 (ATCC 700928) was isolated from the blood of a woman with acute pyelonephritis. The bacterium was streaked from glycerol stock stored at −80°C onto Luria‐Bertani (LB) agar plates, incubated at 37°C, and single colonies were always cultured in nutrient broth (NB—3 g of beef extract and 5 g of peptone in 1 L of water) at 37°C unless mentioned otherwise. S. aureus ATCC6538, Enterococcus faecalis ATCC35058, P. aeruginosa PAO1, A. baumannii ATCC17978 and K. pneumoniae ATCC 13883 were cultured in Luria‐Bertani broth (LB), brain‐heart infusion broth with 0.5% glucose (w/v), tryptic‐soy broth (TSB) with 0.5% glucose (w/v), TSB and TSB respectively, at 37°C. Serratia marcescens was cultured in LB at 30°C. Overnight cultures for all bacteria refer to cultures grown to a stationary phase with ~107–108 CFU/mL. All experiments were performed with two independent cultures in triplicate.
2.3. MIC, Antibiofilm, Planktonic Cell Growth Curve, Cell Survival Assessments of Halogenated Aniline Derivatives Against UPEC
Aniline and its halogenated derivatives were screened for their minimal inhibitory concentrations (MIC) and biofilm inhibition IC50's by adding overnight cultures of UPEC to 96‐well plates (SPL Life Sciences, Korea) at 1:100 dilution with or without aniline and halogenated aniline derivatives at concentrations 5, 10, 20, 50, 100, 200, 400, 600 and 800 μg/mL in NB for 24 h at 37°C without agitation. For biofilm assay, the plate was washed thrice with distilled water, stained with 0.1% crystal violet for 20 min, rewashed thrice with distilled water, solubilised in 95% ethanol and absorbance was measured at 570 nm using Multiskan EX microplate reader (Thermo Fisher Scientific, Waltham, MA, USA). MIC and biofilm IC50 were considered the lowest concentration that inhibited visible growth of bacteria post incubation and the lowest concentration that inhibited 50% of biofilm biomass, respectively. The hit compounds, along with aniline at concentrations of 50 and 100 μg/mL and positive control amoxicillin at 5 and 10 μg/mL, were assayed for biofilm inhibition to confirm efficacy.
MICs and biofilm inhibition of other pathogens viz. E. faecalis , P. aeruginosa , K. pneumoniae , A. baumannii and S. aureus were assayed similarly with their respective media (Table S2). UPEC biofilm dispersal or disruption of mature biofilms was assayed as previously described, with or without aniline and halogenated aniline derivatives at concentrations of 100, 200, 400 and 800 μg/mL (Boya et al. 2022). For the planktonic cell growth of UPEC, overnight culture was reinoculated at 1:100 dilution in NB with or without aniline and halogenated aniline derivatives at concentrations of 50, 100 and 200 μg/mL in 250‐mL flasks at 250 rpm and 37°C. Absorbances were measured at 600 nm for every 2 h up to 14 h and at 24 h using Ultrospec 7000 spectrophotometer (GE, Boston, USA). The cell survival assay was assayed as previously described, using aniline and halogenated aniline derivatives at concentrations of 50, 100 and 200 μg/mL (Boya et al. 2024).
2.4. Microscopy of Biofilms
3D representations of UPEC biofilms were produced as previously described, with or without aniline and halogenated aniline derivatives at 50 μg/mL, and amoxicillin at 5 μg/mL (Boya et al. 2024). For scanning electron microscopy (SEM), 96‐well biofilm plates were prepared as described in Section 2.3, with or without aniline at 400 μg/mL, halogenated aniline derivatives at 100 μg/mL and amoxicillin at 10 μg/mL. Nitrocellulose membranes of 3 × 3 mm dimensions were placed vertically in the wells of the plate and incubated for 24 h at 37°C. Biofilms on the membranes were fixed with a solution of 2.5% glutaraldehyde/2% formaldehyde, incubated overnight at 4°C, and dehydrated using graded ethanol series (50%, 70%, 90%, 95%, 99% and 100%). Membranes were critically point‐dried and sputter‐coated with gold, and SEM was performed at 5 kV using a field emission‐scanning electron microscope (S4800, Hitachi, Tokyo).
2.5. Swimming and Swarming Motilities
Swimming motilities were assessed using 1% tryptone (w/v), 0.25% NaCl (w/v), and 0.3% agar plates, whereas swarming motilities were assessed using 1% peptone (w/v), 0.5% yeast extract (w/v), 0.5% glucose and 0.5% agar plates. Fresh colonies of UPEC were inoculated in 2 mL, grown to an OD of 1 at 600 nm, and 5 μL of this culture was spotted on agar plates with or without aniline and halogenated aniline derivatives at 50 and 100 μg/mL, and amoxicillin at 5 and 10 μg/mL. The plates were incubated for 24 h at 37°C.
2.6. Curli and Rugose Colony Formation Assay
Curli production was assayed using the Congo red agar assay as previously described (Boya et al. 2024). Muller‐Hinton agar plates were prepared with 0.04% (w/v) Congo red and 5% (w/v) sucrose, and supplemented with or without aniline and halogenated aniline derivatives at concentrations of 50 μg/mL and 100 μg/mL, as well as amoxicillin at 5 μg/mL and 10 μg/mL. The plates were inoculated with 25 μL of an overnight UPEC culture and incubated at 37°C for 48 h. The colonies were examined for their rough texture and red colour indicating curli formation.
Rugose colony formation was assayed using 1% acid hydrolyzed casein (w/v) and 0.2% yeast extract (w/v) agar plates with or without aniline and halogenated aniline derivatives at 50 and 100 μg/mL and amoxicillin at 5 and 10 μg/mL. Twenty microlitre of overnight culture of UPEC was spotted on the plates and incubated at 25°C for 5 days. The colonies were visually examined for a veined and wrinkled morphology, indicating rugose colony formation.
2.7. Cell Surface Hydrophobicity by MATH (Microbial Adhesion to Hydrocarbons) Assay and Hemolysis Assay
Cell surface hydrophobicity was performed as previously described (Boya et al. 2024). Overnight UPEC cultures were reinoculated at a dilution of 1:100 and incubated with or without aniline and halogenated aniline derivatives at 50 and 100 μg/mL and amoxicillin at 5 and 10 μg/mL for 24 h at 37°C at 250 rpm. After centrifugation and two PBS washes, cells were resuspended in PBS, mixed with xylene (250 μL), vortexed, and left for 30 min. OD values at 600 nm before (A 0) and after (A i) vortexing were recorded. Percent hydrophobicity (%H) was calculated using:
Hemolysis was performed as previously described (Boya et al. 2024). Overnight UPEC cultures were diluted 1:100 and grown to an OD of 1 at 600 nm. Cultures with or without aniline and halogenated aniline derivatives at 50 and 100 μg/mL and amoxicillin at 5 and 10 μg/mL were incubated at 250 rpm for 36 h at 37°C, then diluted to an OD of 1 at 600 nm. Equal volumes of these cultures and 5% erythrocyte solution (prepared from sheep blood) were mixed and incubated at 37°C for 3 h without agitation. After centrifugation at 11,000 g, supernatant absorbance was measured at 545 nm. Sheep blood was sourced from MBcell (Seoul, Korea).
2.8. Autoinducer‐2 (AI‐2) Production
AI‐2 production in UPEC was determined by prodigiosin and pyocyanin production in S. marcescens and P. aeruginosa , respectively. Overnight cultures of UPEC were reinoculated and grown to an OD600 of 0.5 at 37°C and 250 rpm. These cultures were then incubated for 3 h at 37°C and 250 rpm with or without aniline and halogenated aniline derivatives at 50 and 100 μg/mL and amoxicillin at 5 and 10 μg/mL. Cultures were centrifuged at 14,000 g, and the supernatants were added to 1:100 reinoculated media cultures of S. marcescens and P. aeruginosa and incubated at 250 rpm at 30°C and 37°C, respectively, for 16 h.
For the prodigiosin assay, the cultures were centrifuged at 14,000 g for 10 min, and acidified ethanol (1 mL of 4% (v/v) 1 M HCl in ethanol) was added to the cell pellets and vortexed vigorously to extract prodigiosin. After centrifugation, supernatant absorbance was measured at 534 nm. For the pyocyanin assay, the cultures were centrifuged at 14,000 g for 10 min, and 3 mL of chloroform was added to 5 mL of supernatant to extract pyocyanin in the bottom chloroform phase. This bottom phase was added to an equal volume of 0.2 M HCl, and the absorbances of the top aqueous phase were measured at 520 nm. Both prodigiosin and pyocyanin absorbances were normalised with the absorbance of the UPEC cultures to eliminate cell count bias.
2.9. Siderophore, Chalkophore and Quiescence Assays
Siderophore production was assayed using chrome azurol S (CAS) agar as previously described (Boya et al. 2024). CAS agar plates were prepared by adding 20% v/v CAS reagent in LB or M9 minimal agar containing or not containing aniline and halogenated aniline derivatives at 50 and 100 μg/mL, and amoxicillin at 5 and 10 μg/mL. Plates were stabbed with 20 μL of UPEC overnight cultures and incubated at 37°C for 72 h, when the diameters of orange‐yellow halos were recorded.
Quiescence was assayed as previously described, with or without aniline and halogenated aniline derivatives at 200 and 400 μg/mL and amoxicillin at 20 and 40 μg/mL (Boya et al. 2024). Chalkophore production was measured using the complexometric indicator Eriochrome Black T (EBT). Overnight cultures of UPEC were reinoculated and grown to an OD600 of 0.5 at 37°C and 250 rpm. These cultures were then incubated for 3 h at 37°C and 250 rpm with or without aniline and halogenated aniline derivatives at 50 and 100 μg/mL and amoxicillin at 5 and 10 μg/mL. Absorbances at 600 nm were recorded, and 1 mM CuSO4.5H2O solution was added to the cultures in equal volume measure and incubated for 2 h at 37°C and 250 rpm. Copper ion concentration indicating chalkophore production was determined by adding 1% EBT solution in equal measure to the supernatants, vortexed for 5 min and absorbances were recorded at 555 nm. Concentrations were determined using a linear regression curve of standard concentrations of copper sulfate solution and EBT complex and further normalised with OD600 values of culture before copper sulfate treatment. The methodology for lactose fermentation using MacConkey agar is provided in the Supporting Information.
2.10. In Silico Analysis‐ 3D QSAR and ADME Profiling
For 3D QSAR, 3D structures of the aniline and its 69 derivatives were downloaded from PubChem, aligned and energy minimised using the small molecules tool. MIC and biofilm IC50 values were given manually as attributes and converted to the activity bin. Training and test sets were prepared with a random assignment of derivatives at an 80:20 split ratio. A 3D‐QSAR model based on partial least squares regression was built and statistically validated. Positive and negative electrostatically favourable regions were visualised on the core aniline scaffold. Biovia Discovery Studio 2022 was used for all functions and tools to build the 3D‐QSAR model.
The adsorption, distribution, metabolism, elimination, and toxicity profile of aniline and its halogenated derivative hits was evaluated using online webservers, viz. Swiss ADME, Protox 2, and PreADMET (accessed on 10th March 2024). Property validation was performed using web server bioassay parameters and published literature. The methodology for molecular docking, protein quality check and density functional theory (DFT) analysis is mentioned in the Supporting Information.
2.11. Cytotoxicity Using a Nematode Model
Cytotoxicity of the halogenated aniline derivatives was determined using an in vitro nematode model using Caenorhabditis elegans as the model organism. M9 buffer with or without aniline and its halogenated derivatives at concentrations 50, 100, 200 and 400 μg/mL were added to the 96‐well plate with synchronised adult C. elegans fer‐15; fem‐1 worms (non‐infected, adult‐L4 stage; n = ~30 per well). Plates were incubated at 25°C for 7 days and worm survival was evaluated by exposing worms to 20 s of blue LED light pulse (450–490 nm) using the iRiS Digital Cell Imaging System (Logos BioSystems, Anyang, Korea).
2.12. Gene Expression Profile
RNA was isolated as previously described (Boya et al. 2024). Briefly, overnight cultures were reinoculated at a 1:100 dilution into 25 mL of NB media, incubated at 37°C and 250 rpm to an OD600 of 1.0, and treated with or without halogenated aniline derivatives at their MIC concentrations for 2 h at 37°C and 250 rpm. RNAse inhibitor (RNAlater, Ambion, TX, USA) was added to the cultures, which were then centrifuged at 15,000 g for 10 min, and RNA was isolated using Qiagen RNeasy Mini Kits (Valencia, CA, USA). RNA concentrations and purities were determined using a NanoVue Plus nanodrop spectrophotometer (GE, Chicago, IL, USA). The primer names, sequences, and functions are provided in Table S3, with the rrsG gene as the housekeeping endogenous control. The ABI StepOne Real‐Time PCR instrument (Applied Biosystems, Foster City, CA, USA) and SYBR Green master mix were used to run PCR for two independent cultures amounting to four reactions performed per gene. Gene expression levels were quantified using the method (Livak's method) to calculate fold changes relative to the control group. All qPCR experiments were performed with two biological replicates, and the resulting fold change values were used for statistical analysis. The statistical significance of differential expression between groups was assessed using the Student's t‐test.
2.13. Statistical Analysis
All experiments were performed using at least two independent biological cultures in triplicates unless stated otherwise. Results are presented as means ± standard deviation (SD). Statistical significance was assessed using Student's t‐test and Fisher's Exact test with *p < 0.05; **p < 0.01; ***p < 0.001.
3. Results
3.1. Biofilm Inhibition, Cell Growth and Cell Survival
Aniline and its halogenated derivatives were screened for their MIC and biofilm IC50 values using the crystal violet biofilm assay. The hits from the screening with the lowest combination of MIC and biofilm IC50 values were 4‐bromo 3‐chloroaniline (4B3CA) and 3,5‐dibromoaniline (3,5‐DBA) (Table S1). The MIC and biofilm IC50 values of 4B3CA were 200 and 10 μg/mL, respectively, whereas those of 3,5‐DBA were 100 and 10 μg/mL, respectively. Both the MIC and biofilm IC50 of aniline were > 800 μg/mL. The MICs of the other derivatives ranged from 200 to > 800 μg/mL, whereas the biofilm IC50 values ranged from 20 to > 800 μg/mL. The hit compounds 4B3CA and 3,5‐DBA were further tested on other pathogens of the ESKAPE group. The hit compounds dose‐dependently inhibited biofilm formation and completely abolished biofilm formation in all the tested pathogens at 400 μg/mL, except for P. aeruginosa , where the same effect was observed at a higher concentration of 800 μg/mL (Figure 1A). The MICs of the hit compounds against these pathogens ranged from 100 to 800 μg/mL (Table S2). The two hits and aniline were chosen for further testing on antivirulence properties against UPEC. Amoxicillin is commonly recommended for clinical use in cases of pyelonephritis or complicated UTI (Kot 2019). Amoxicillin, with an in vitro minimum inhibitory concentration (MIC) of 10 μg/mL, was selected as a positive control to benchmark the antibiofilm and antivirulence efficacy of a clinically relevant therapeutic agent against the tested aniline derivatives. The hit derivatives did not significantly inhibit mature biofilm dispersal even at higher concentrations of 800 μg/mL (Figure S1).
FIGURE 1.
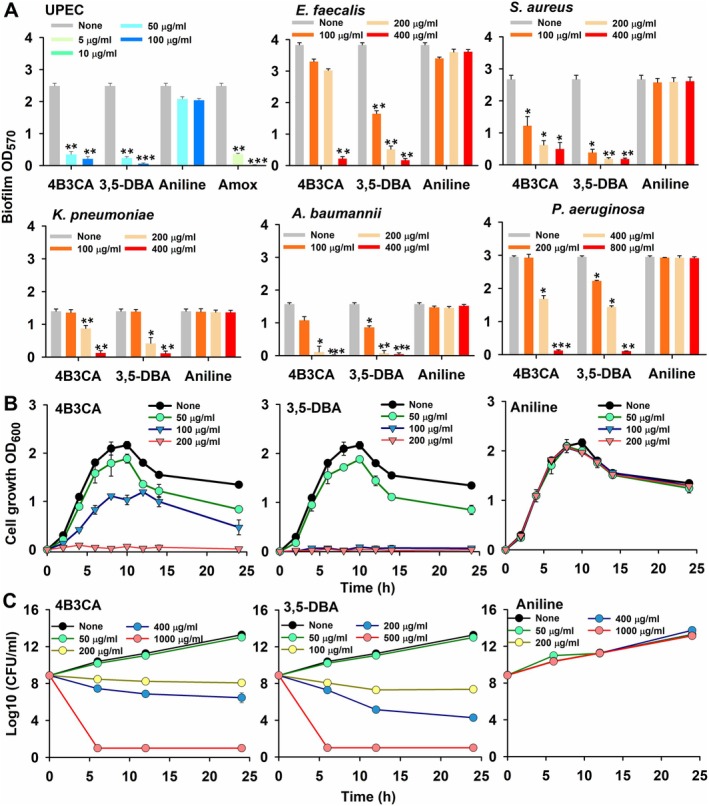
Crystal violet biofilm assays of UPEC and various pathogens in the presence of aniline and its derivatives (A). Biofilm OD570 represents the absorbance of CV‐stained biofilm. Cell growth curves of UPEC in the presence of aniline and its derivatives, wherein cell growth OD600 represents turbidity of the culture (B) and cell survival assay of UPEC in the presence of aniline and its derivatives (C), wherein log10 (CFU/mL) represents the log values of the colony‐forming units per millilitre. Incubation time of agar plates is 24 h at 37°C. *p < 0.05, **p < 0.01 and ***p < 0.001 versus non‐treated controls. All data from the cell growth and cell survival assays were statistically significant, with p < 0.05 compared to non‐treated controls.
The cell growth of UPEC was dose‐dependently inhibited by the aniline derivatives, whereas aniline was ineffective at all tested concentrations (Figure 1B). 4B3CA completely inhibited cell growth at 200 μg/mL, whereas 3,5‐DBA negated cell growth at both 100 and 200 μg/mL owing to its lower MIC (Figure 1B). The hit derivatives demonstrated dose‐dependent killing of UPEC cells in the cell survival assay, whereas aniline remained ineffective (Figure 1C). At MIC, both the hit derivatives were bacteriostatic, whereas complete bactericidal activity of the derivatives was observed at 5xMIC (Figure 1C). At 50 μg/mL, where 90% of the biofilm was inhibited, the hit derivatives did not affect cell survival.
The 3D representations of the biofilm architectures exhibited a range of 180–190 scale in the untreated samples, whereas treatment with hit derivatives at 50, and amoxicillin at 5 μg/mL plummeted the biofilm architecture to a range of 20–50 with ~79% inhibition (Figure 2A). Aniline did not have any effect on the biofilm architectures (Figure 2A). SEM revealed a dramatic reduction of the cells as well as diminished extracellular polymeric substance (EPS) on visual inspection in the hit derivatives‐treated samples at 100 μg/mL (Figure 2B). Aniline did not have any effect on the biofilm formation, whereas amoxicillin treatment at 10 μg/mL decreased cell counts, but EPS production on visual inspection was not significantly affected (Figure 2B). Hit derivatives‐treated samples showed an altered rough and craggy morphology in comparison to the smoother morphology of the untreated samples (Figure 2B).
FIGURE 2.
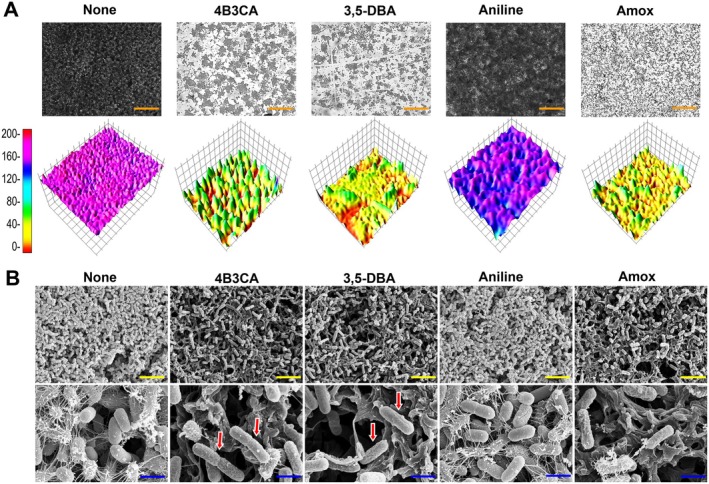
2D‐3D representation of UPEC biofilms in the presence of aniline and its derivatives at 50 μg/mL, and amoxicillin at 5 μg/mL (A). SEM of UPEC biofilms in the presence of 4B3CA, 3,5‐DBA, aniline and amoxicillin at concentrations of 200, 100, 400 and 10 μg/mL, respectively, for 24 h and 37°C (B). All SEM images were taken at 5 kV. The orange, yellow and blue scale bars represent 50, 10, and 2 μm, respectively. The red arrows indicate altered membrane morphology with cell elongation in comparison to non‐treated controls.
3.2. Motility, Hemolysis, Cell Surface Hydrophobicity, Curli, Rugose Colony and AI‐2 Production
Motility plays a vital role in UPEC colonisation by facilitating movement through host environments, enabling surface attachment and promoting biofilm formation for persistent infection (Sudarshan et al. 2021). The hit derivatives dose‐dependently inhibited swarming and swimming motility of UPEC at 50 and 100 μg/mL (Figure 3). Interestingly, aniline also significantly inhibited swarming motility with a minor reduction in swimming motility at similar concentrations, whereas amoxicillin inhibited both types of motility at 5 and 10 μg/mL (Figure 3). The hit derivatives also curtailed curli production in the Congo red agar assay at 50 and 100 μg/mL, where the colony morphology was pale and smooth compared to the untreated control's red and rough morphology (Figure 4A). Aniline at 50 and 100 μg/mL also altered colony morphology to pale and white, although not in the magnitude of the hit derivatives. Amoxicillin treatment at 5 and 10 μg/mL made the colony morphology texture smoother, but the colony retained the distinct red colour (Figure 4A).
FIGURE 3.
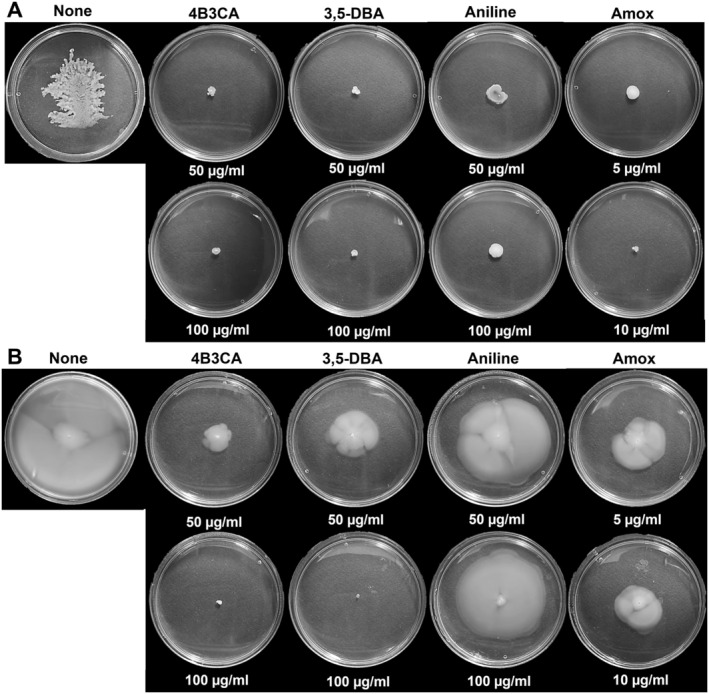
Swarming motility (A) and swimming motility (B) of UPEC in the presence of aniline derivatives, incubated for 24 h at 37°C. Fisher's exact test revealed a statistically significant association between compound treatment and motility inhibition (p < 0.05).
FIGURE 4.
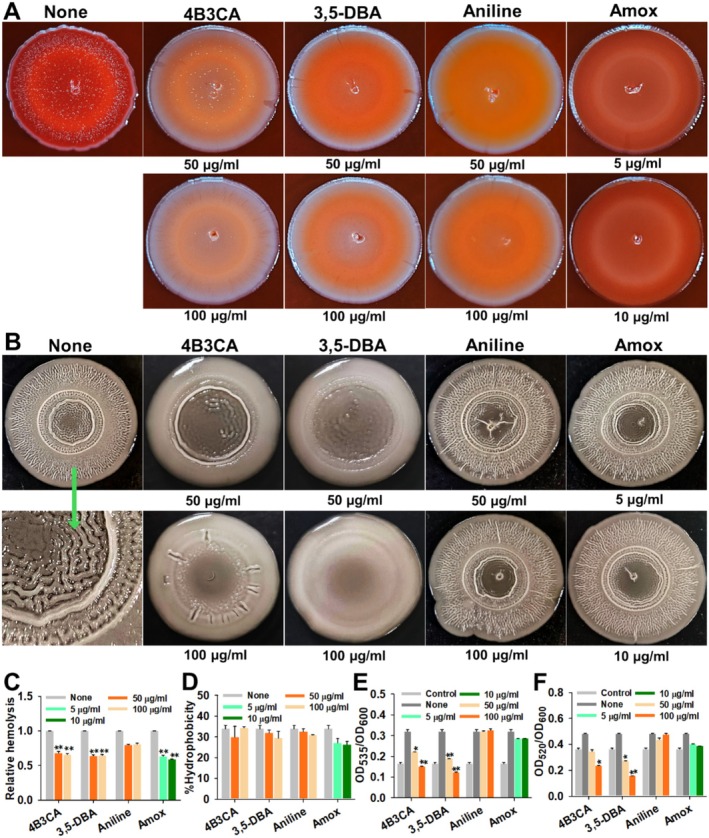
Congo red assay (A), rugose colony formation (B), hemolysis (C) cell surface hydrophobicity (D), prodigiosin assay (E), and procyanin assay (F). All the experiments were performed at 37°C for 24 h, except rugose colony formation, which was incubated for 5 days at 25°C. *p < 0.05, **p < 0.01 and ***p < 0.001 versus non‐treated controls. Fisher's exact test demonstrated a statistically significant association between compound treatment and the inhibition of curli production and colony rugosity (p < 0.05).
Rugose colony formation in UPEC facilitates enhanced colonisation by promoting robust biofilm architecture, increasing adhesion to surfaces, and providing heightened resistance to desiccation, antibiotics and host immune responses (Hufnagel et al. 2013). The hit derivatives dose‐dependently inhibited rugosity at 50 and 100 μg/mL. The treated colonies had substantially smoother and less wrinkled morphology in comparison to the non‐treated controls (Figure 4B). Aniline and amoxicillin did not have any effect on the rugosity at the tested concentrations. Both hemolysis and cell surface hydrophobicity were unaffected by hit derivatives, aniline, and amoxicillin at their treated concentrations (Figure 4C,D). Autoinducer‐2 production in UPEC was assayed by prodigiosin and pyocyanin assays of S. marcescens and P. aeruginosa , respectively, and the hit derivatives significantly inhibited AI‐2 production at concentrations of 50 and 100 μg/mL (Figure 4E,F). Both aniline and amoxicillin did not significantly inhibit AI‐2 production in UPEC (Figure 4E,F).
3.3. Siderophore, Chalkophore Production, Quiescence and Lactose Fermentation
The hit derivatives inhibited siderophore production as assayed by CAS agar at 50 and 100 μg/mL, characterised by the reduced orange‐yellow halo around the colony, signifying reduced uptake of iron from the blue CAS‐iron complex (Figure 5A). Aniline at 50 and 100 μg/mL and amoxicillin at 5 and 10 μg/mL were unable to inhibit siderophore production (Figure 5A). The hit derivatives dose‐dependently inhibited quiescent colony formation at 200 and 400 μg/mL. At 400 μg/mL, the colonies were completely abolished (Figure 5B,C). Aniline and amoxicillin did not have any effect on quiescent colony formation at their tested concentrations (Figure 5B,C). Chalkophore production was measured by the copper ion concentration using EBT dye, and the hit derivatives inhibited chalkophore production, evidenced by the increased copper ion concentration in the culture (Figure 5D). Aniline and amoxicillin did not inhibit chalkophore production (Figure 5D). On visual inspection of MacConkey agar plates, the hit derivatives dose‐dependently inhibited lactose fermentation at 50 and 100 μg/mL, changing the colony colour and medium halo from pale pink to yellow, whereas aniline remained ineffective (Figure S2).
FIGURE 5.
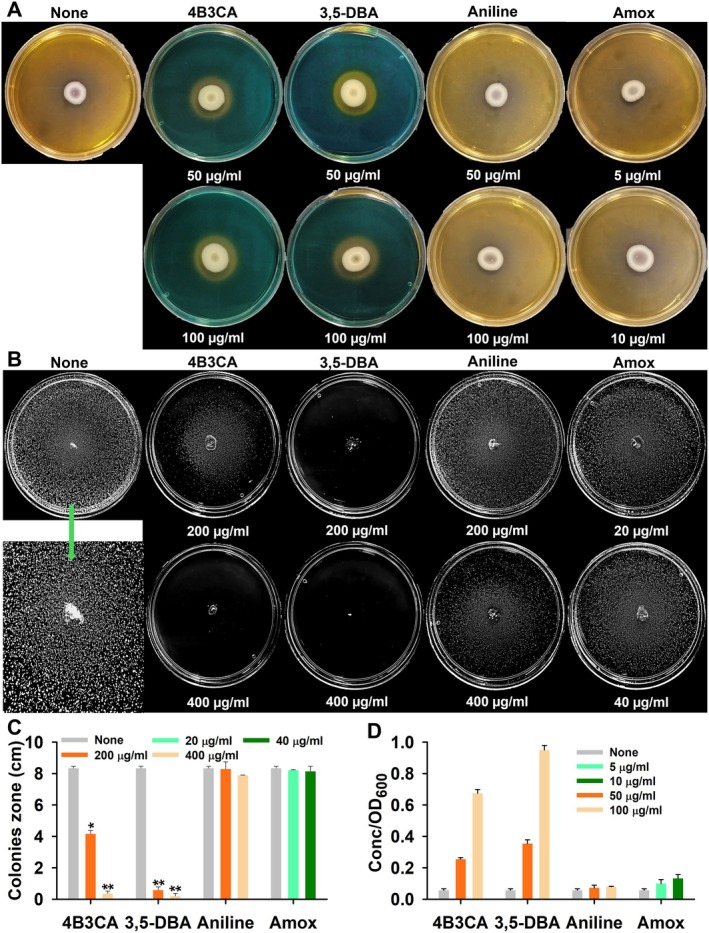
CAS agar assay incubated for 72 h (A), in vitro quiescence assay incubated for 36 h (B), quiescent colony diameters (C), and chalkophore assay incubated for 3 h (D), in the presence of aniline derivatives. All the experiments were performed at 37°C. *p < 0.05 and **p < 0.01 versus non‐treated controls. Fisher's exact test demonstrated a statistically significant association between compound treatment and the inhibition of siderophore production (p < 0.05).
3.4. Gene Expression Analysis
The expression profiles of various virulence and biofilm genes in UPEC were assessed following treatment with two hit derivatives (Figure 6). The average fold change of both hit derivatives is discussed in this section. The curli genes csgA, csgB and csgG were downregulated in expression by 2, 7.1 and 3.8‐fold, respectively. Expression of the fimbriae genes fimA, fimH, papA and papG was downregulated by 16.3, 12.3, 1.7 and 11.1‐fold, respectively. The average downregulation for motility‐related genes, including flagellin and chemotaxis genes (fliC, motA and motB), was 13.5, 7.8 and 31.5, respectively. Iron acquisition genes, such as the metal transport and siderophore genes sitA, entE and iucB, were downregulated by averages of 13.1, 7.1 and 4.8‐fold, respectively. Stress regulation genes oxyR and rpoS saw average downregulations of 9.1 and 7.6‐fold. The antigen 43 adhesin gene flu and the K1 capsule‐associated gene kpsD were downregulated by 7.3 and 16.6‐fold, respectively. Conversely, the α‐hemolysin gene hlyA, the carbon storage regulator csrA, and the two‐component system regulator uvrY saw upregulation in expression by 1.1, 2.6 and 1.2‐fold, respectively. The cAMP adenylate cyclase gene cyaA and the CRP gene crp were downregulated by 18.9 and 5.8‐fold, respectively.
FIGURE 6.
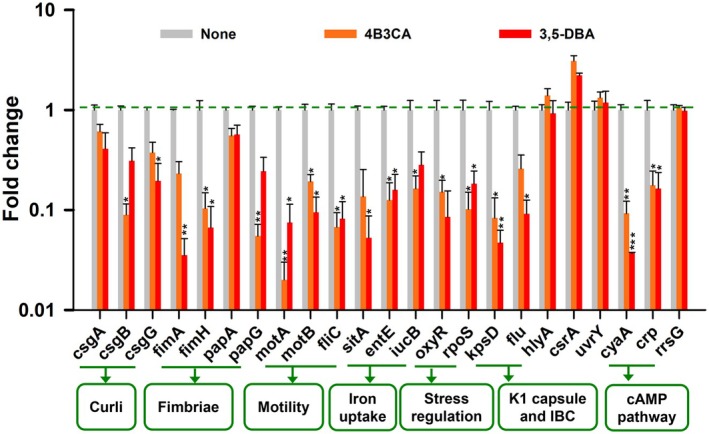
Gene expression profiles of UPEC following treatment with 4B3CA (200 μg/mL) and 3,5‐DBA (100 μg/mL) for 2 h at 37°C. The rrsG gene was used as a housekeeping control. *p < 0.05, **p < 0.01 and ***p < 0.001 versus non‐treated controls.
3.5. In Silico Analysis and Nematode Model Cytotoxicity
The 3D‐QSAR analysis for MIC (Figure 7A–C) and biofilm IC50 (Figure 7D–F) values of aniline and its 69 halogenated derivatives against UPEC was performed. Both antimicrobial and antibiofilm 3D‐QSAR analyses revealed that positions 1 and 2 of the aniline ring were electrostatically unfavourable for substitutions, whereas positions 3 and 4 were favourable. Halogen substitution on the C1 position displacing the amine group is detrimental to both antimicrobial and antibiofilm activities (Figure 7A–F). Based on the affinity of the aniline scaffold to adenylate cyclase, aniline, hit derivatives along with the native ligand adenosine triphosphate (ATP), and known inhibitors of adenylate cyclase viz. arabinosyl‐ATP, 2′,5′‐dideoxy adenosine (2′,5′‐DDA) and 9‐(tetrahydrofuran‐2‐yl)‐9h‐purin‐6‐amine (SQ22536) were docked to the cyaA protein of UPEC. As the 3D structure of cyaA has not been resolved, the structure from AlphaFold was chosen. The residue confidence analysis indicates high pLDDT scoring values in the binding site of the protein, further confirmed with the alignment error analysis (Figure S3A,B). Errat tool and Ramachandran plot indicated most residues were found in the favoured regions and no residues were found in disallowed regions (Figure S3C,D).
FIGURE 7.
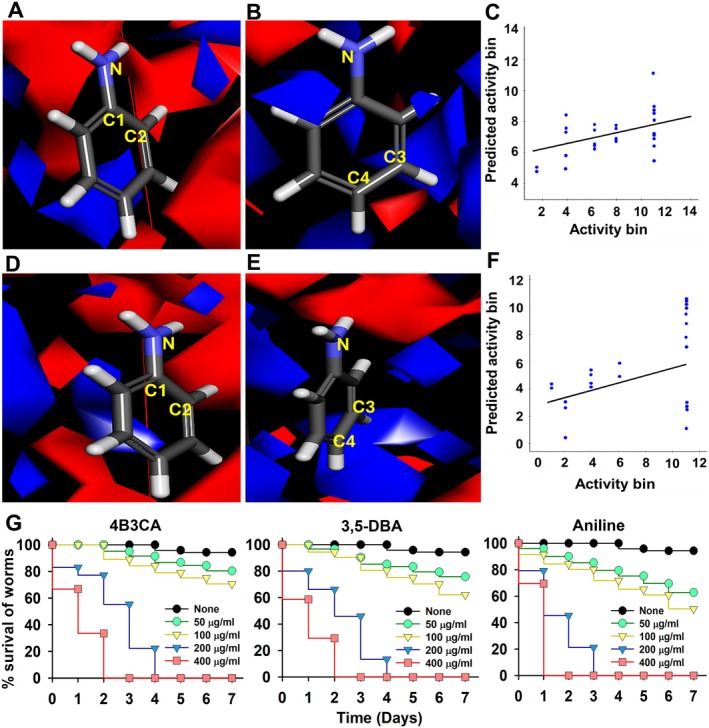
Substitutions on positions 1 and 2 are electrostatically unfavourable for antimicrobial activity (A), substitutions on positions 3 and 4 are electrostatically favourable for antimicrobial activity (B) and scatter plot of predicted antimicrobial activity bin vs. activity bin (C). Substitutions on positions 1 and 2 are electrostatically unfavourable for antibiofilm activity (D), substitutions on positions 3 and 4 are electrostatically favourable for antibiofilm activity (E), scatter plot of predicted antibiofilm activity bin vs. activity bin (F) and Caenorhabditis elegans cytotoxicity assay of halogenated anilines (G). All data from the C. elegans cytotoxicity assay showed statistically significance compared to the non‐treated controls (p < 0.05).
The combination of molecular docking scores and molecular mechanics with generalised born and surface area solvation (MM/GBSA) values indicates that inhibitor ara‐ATP is the best ligand, followed by the order of 3,5‐DBA, 4B3CA, SQ22536, 2′,5′‐DDA, ATP and aniline (Table S4). Interaction poses revealed that the halogens present in the halogenated derivatives formed a halogen bond with tryptophan 118 residue of the binding site and, in the case of 4B3CA, the amine group formed a hydrogen bond with glutamic acid 185 (Figure S4). 3,5‐DBA formed an additional pi‐pi stack with phenylalanine 154, which was also observed in aniline, along with a hydrogen bond with aspartic acid 116 with aniline. In the native ligand ATP, the phosphate group formed various hydrogen bonds with serine 103, Lysine 260,264 and aspartic acid 116. The amine group in ATP formed a hydrogen bond with glycine 175 (Figure S4).
The molecular electrostatic potential of aniline is distinct in comparison to its halogenated derivatives. Aniline did not show a high distinction of positive or negative electrostatic potential regions, whereas the halogenated derivatives showed a high positive electrostatic potential along the axis of halogen substitution, explained by the characteristic sigma‐hole of halogens, whereas the remaining scaffold exhibited high negative electrostatic potential (Figure S5). DFT for molecular orbitals revealed highest occupied molecular orbitals around the benzene ring and amine groups in aniline as well as the hit derivatives, whereas lowest unoccupied molecular orbitals (LUMO) were present near chlorine atom of 4B3CA and both halogen substitutions in 3,5‐DBA (Figure S5). Quantum chemical parameter calculations indicate higher chemical reactivity and electron affinities of hit derivatives in comparison to aniline with good stability (Table S5).
In silico ADMET profile showed high druggability of the hit derivatives with minimal toxicity and carcinogenicity, high absorption and bioavailability, and no Lipinski, Veber or Egan rule violations. The hit derivatives also had no hepatotoxicity and increased toxicity LD50 values in comparison to aniline (Table S6). The hit derivatives and aniline showed minimal lethality to worms in the C. elegans cytotoxicity model at the concentrations of 50 and 100 μg/mL. At a higher concentration of 400 μg/mL, the worms were killed on day 3 by the hit derivatives, whereas the same was observed on day 2 for aniline (Figure 7G).
4. Discussion
This study evaluated the potential of aniline and its halogenated derivatives to quell UPEC biofilms and other associated virulence factors. Both hit derivatives—4B3CA and 3,5‐DBA were—able to inhibit UPEC biofilm at both sub‐MIC and MIC concentrations (Figures 1A and 2). The hit derivatives were bactericidal at 5xMIC concentrations (Figure 1C). SEM images of the hit derivatives‐treated samples at MIC concentration demonstrated altered membrane morphology with a rough texture and cell elongation (Figure 2). Elevated intracellular nitrosative stress can be a likely cause for the altered membrane morphology, as aniline is capable of escalating reactive nitrogen species (Fan et al. 2011; Wang et al. 2016), and a similarly altered morphology was observed in another study wherein E. coli was subjected to nitrosative stress by nitric‐oxide water (Borkar et al. 2023). Disruption of the cAMP‐CRP complex increases sensitivity to nitrosative stress (Donovan et al. 2013) and can contribute to the antimicrobial activity of the hit derivatives (Figure 6). The hit derivatives demonstrated potent activity against various nosocomial pathogens from the ESKAPE group, suggesting a broad mechanism of action for aniline derivatives that targets global regulatory systems across diverse microbial species (Figure 1A).
The hit derivatives inhibited both swimming and swarming motility (Figure 3), attributed to the downregulation of flagellin fliC and chemotaxis genes (motA and motB) leading to disruption of flagella biosynthesis (Figure 6). Curli amyloid fibres and cellulose constitute a significant portion of the extracellular matrix of the biofilm, promoting adhesion, cell aggregation and structural support (Luna‐Pineda et al. 2019). The hit derivatives completely diminished curli formation as evidenced by Congo red agar assay along with rugose colony formation (Figure 4A,B). The downregulated curli genes (csgA, csgB and csgG) and regulatory sigma factor (rpoS) explain the metamorphosed colonies (Figure 6). Interestingly, both cell surface hydrophobicity and hemolysis remained unaffected by hit‐derivative treatment (Figure 4C,D). The biophysical phenomenon of cell surface hydrophobicity can be negatively regulated by cAMP levels in some bacteria (Ono et al. 2014), and downregulated cyaA and crp genes might partially explain the inefficacy of the hit derivatives to lower hydrophobicity (Figure 6). The α‐hemolysin gene hlyA remained unaffected, explaining the lack of impact on hemolysis (Figure 6). The quorum sensing AI‐2 of UPEC contributes to biofilm formation, stress regulation and virulence. The hit derivatives inhibited AI‐2 production (Figure 4E,F), which is explained by the upregulation of carbon storage regulator A (csrA) (Figure 6). This upregulation negatively impacts luxS expression, resulting in decreased accumulation of AI‐2 in the extracellular milieu of E. coli (Mitra et al. 2016; Parker et al. 2017).
UPEC employs several metallophores like enterobactin, aerobactin, and yersiniabactin among others, to sequester iron or other metals like copper from the iron‐deficient urothelium. The hit derivatives inhibited siderophore production (Figure 5A), corroborated by the downregulation of the enterobactin entE, aerobactin iucB and iron transport sitA genes. The hit derivatives also inhibited yersiniabactin production, supported by the chalkophore assay (Figure 5D). IBCs and QIRs afford UPEC survival and evasion from the host immune response and antibiotic treatments, leading to recurrent infections. In vitro quiescent colony formation was completely abolished by the treatment of hit derivatives (Figure 5B,C). The hit derivatives significantly downregulated K1 capsule (kpsD) and antigen 43 auto aggregation adhesin (flu) genes (Figure 6), which play pivotal roles in IBC formation and prolonged persistence in the urinary tract (Ulett et al. 2007).
The cAMP‐CRP complex is a key global regulator involved in alternative carbon source utilisation, virulence and bacterial metabolism (Donovan et al. 2013). Treatment with hit derivatives downregulates this complex (Figure 6). It controls curli‐cellulose and K1 capsule expression via csgD and CRP binding upstream of the kpsMT operon (Hufnagel et al. 2016), and also regulates flagellar biosynthesis essential for motility (Sudarshan et al. 2021). Additionally, it influences type‐1 fimbriae phase variation by repressing fimB and is necessary for pap operon expression (Müller et al. 2009). Although its role in UPEC's iron regulon is unclear, cAMP downregulates fur genes, affecting enterobactin and salmochelin synthesis (Biville et al. 2003; Banerjee et al. 2020), and also interferes with aerobactin synthesis in Vibrio vulnificus (Kim et al. 2012). Notably, cAMP‐CRP does not affect uvrY, csrA or hlyA expression. Deleting cyaA increases hydrogen peroxide resistance via rpoS upregulation but heightens sensitivity to superoxide and nitrosative stress (Donovan et al. 2013). It also partially regulates the TCA cycle via the PEP‐glyoxylate shunt (Nanchen et al. 2008) and is essential for lactose fermentation, which is disrupted by hit derivatives (Figure S2) (Donovan et al. 2013).
The observed trends of downregulation and disruption of the above‐listed virulence factors lead us to hypothesise that cyaA adenylate cyclase might be the likely target of the aniline derivatives. Several commercial and synthesised adenylate cyclase inhibitors, like 2′,5′‐dideoxyadenosine (2′,5′‐DDA) and 9‐(tetrahydrofuran‐2‐yl)‐9h‐purin‐6‐amine (SQ22536), contain aniline moiety, further supporting our hypothesis. The dual activity of antibiofilm and antimicrobial activity of the hit derivatives can be attributed to the disruption of cyaA, which influences both csgD (master curli and biofilm regulator) and DNA gyrase A (DNA replication), which is positively regulated by cAMP (Gomez‐Gomez et al. 1996; Bessaiah et al. 2022). Molecular docking analysis performed with cyaA further confirmed efficient binding of the halogenated aniline derivatives in the catalytic domain and forming halogen bonds with Trp118 (Figure S4), potentially blocking binding to other key residues of Ser103, Ser113, Asp114, Asp116 and Trp118 (Linder 2008). The above results partially validate the hypothesis that the hit derivatives inhibit UPEC adenylate cyclase to manifest antimicrobial and antibiofilm activity. Further studies, including gene knock out studies, are required to conclusively prove this hypothesis. The hit derivatives might also inhibit adenylate cyclase or its homologues in other tested pathogens, warranting further analysis.
Electrostatic potential maps and DFT analysis reveal positive electrostatic potential and LUMO orbitals along the halogen substitution enabling bonds with nucleophiles like tryptophan residues (Figure S5). Therefore, halogenation might be key to the elevated activity of the derivatives in comparison to aniline. The base aniline scaffold lacked antimicrobial and antibiofilm activity, whereas halogenated aniline derivatives exhibited both. Molecular docking supports the hypothesis that these derivatives inhibit UPEC adenylate cyclase (cyaA), showing enhanced binding within the catalytic domain via halogen bonds with Trp118 (Figure S4), potentially disrupting access to key residues such as Ser103, Ser113, Asp114 and Asp116 (Linder 2008). Electrostatic potential maps and DFT analysis reveal that halogen substitution introduces a positive electrostatic potential and favourable LUMO distribution, promoting interactions with nucleophilic residues like tryptophan (Figure S5). 3D‐QSAR analysis (Figure 7A–F) indicates that halogenation at the third and fourth positions of the aniline ring enhances activity, aligning with the docking results (Figure S4). Together, these findings suggest that strategic halogenation significantly contributes to the antimicrobial and antibiofilm potential of aniline derivatives by enhancing binding affinity to the adenylate cyclase enzyme. The in silico ADMET profile of the derivatives was favourable for clinical use (Table S6) and the nematode cytotoxicity model shows good worm survival at the antibiofilm range of 50–100 μg/mL (Figure 7G). Halogenated anilines demonstrated suitability for human use based on the in silico ADMET evaluations (Table S6). These compounds showed no notable toxicity, exhibited favourable logP (lipophilicity) and TPSA values, and displayed high skin and gastrointestinal permeability. Additionally, they did not violate key drug‐likeness criteria such as Lipinski, Veber and Egan rules. Collectively, these properties partially support their pharmacokinetic and pharmacodynamic viability for therapeutic use. More preclinical toxicological studies, including those using cell lines and murine models, are required to conclusively prove clinical fitness and potential drug use. The distinguished biofilm inhibition capability of these hit derivatives makes them a lucrative prophylactic option for coating medical devices such as indwelling catheters. Halogenation of compounds generally enhances their potency against resistant pathogens and reduces the likelihood of resistance development (Faleye et al. 2024). Halogenated aniline derivatives have also shown effectiveness against various MDR pathogens (Halim et al. 2024). Recently, compounds such as 4‐amino‐3‐chloro‐5‐nitrobenzotrifluoride (ACNBF) and 2‐iodo‐4‐trifluoromethylaniline (ITFMA) have been reported to exhibit low potential for resistance development (Sathiyamoorthi et al. 2025). Notably, the risk of resistance can be minimal at lower antibiofilm concentrations (10–50 μg/mL), which do not affect planktonic cell growth. However, at higher bactericidal concentrations, selective pressure may promote resistance. Although halogen anilines might develop less resistance compared to conventional antibiotics, judicious use of these derivatives is still advised.
5. Conclusion
The halogenated aniline derivatives 4B3CA and 3,5‐DBA exhibited antibiofilm and antimicrobial activity against UPEC and some of the ESKAPE pathogens tested. Unlike the traditional antibiotic amoxicillin, these derivatives inhibited various virulence factors associated with UPEC, such as curli, motility, iron acquisition and quiescence. Based on the gene expression pattern, the adenylate cyclase could be a potential target for these derivatives. Disruption of the cAMP‐CRP pathway by these derivatives might lead to antibiofilm activity through inhibition of curli and antimicrobial activity, possibly by DNA gyrase A inhibition. ADMET profile and nematode toxicity model deem these derivatives appropriate for clinical use. These results indicate the viability of these derivatives as broad‐spectrum therapeutic options against uropathogenic and nosocomial microbes.
Author Contributions
Bharath Reddy Boya: methodology, software, data curation, investigation, validation, formal analysis, visualization, writing – original draft, writing – review and editing. Jin‐Hyung Lee: methodology, software, supervision, writing – review and editing. Jintae Lee: conceptualization, writing – review and editing, project administration, resources, funding acquisition, supervision.
Disclosure
Declaration of Generative AI in Scientific Writing: No artificial intelligence tool was used to write the manuscript.
Conflicts of Interest
The authors declare no conflicts of interest.
Supporting information
Data S1
Acknowledgements
This research was supported by the National Research Foundation of Korea (NRF) funded by the Korean government (MSIT) (RS‐2025‐00513239 to J. Lee and RS‐2025‐00553409 to J.‐H. Lee).
Funding: This work was supported by the National Research Foundation of Korea, RS‐2025‐00513239, RS‐2025‐00553409.
Data Availability Statement
All data generated or analysed during this study are included in this published article and its Supporting Information files.
References
- Ageorges, V. , Wawrzyniak I., Ruiz P., et al. 2023. “Genome‐Wide Analysis of Antigen 43 (Ag43) Variants: New Insights in Their Diversity, Distribution and Prevalence in Bacteria.” International Journal of Molecular Sciences 24: 5500. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Atapalkar, R. S. , and Kulkarni A. A.. 2023. “Direct Amidation of Acids in a Screw Reactor for the Continuous Flow Synthesis of Amides.” Chemical Communications 59: 9231–9234. [DOI] [PubMed] [Google Scholar]
- Banerjee, R. , Weisenhorn E., Schwartz K. J., et al. 2020. “Tailoring a Global Iron Regulon to a Uropathogen.” MBio 11, no. 2: e00351‐20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bessaiah, H. , Anamalé C., Sung J., and Dozois C. M.. 2022. “What Flips the Switch? Signals and Stress Regulating Extraintestinal Pathogenic Escherichia coli Type 1 Fimbriae (Pili).” Microorganisms 10: 5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Biville, F. , Oshima T., Mori H., et al. 2003. “ Escherichia coli Response to Exogenous Pyrophosphate and Analogs.” Journal of Molecular Microbiology and Biotechnology 5, no. 1: 37–45. [DOI] [PubMed] [Google Scholar]
- Borkar, S. B. , Negi M., Kaushik N., et al. 2023. “Plasma‐Generated Nitric Oxide Water Mediates Environmentally Transmitted Pathogenic Bacterial Inactivation via Intracellular Nitrosative Stress.” International Journal of Molecular Sciences 24: 1901. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boya, B. R. , Lee J.‐H., and Lee J.. 2022. “Antibiofilm and Antimicrobial Activities of Chloroindoles Against Uropathogenic Escherichia coli .” Frontiers in Microbiology 13: 872943. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boya, B. R. , Lee J.‐H., and Lee J.. 2024. “Antimicrobial and Antibiofilm Activities of Chromone Derivatives Against Uropathogenic Escherichia coli .” Microbiological Research 278: 127537. [DOI] [PubMed] [Google Scholar]
- Brannon, J. R. , Reasoner S. A., Bermudez T. A., et al. 2024. “Mapping Niche‐Specific Two‐Component System Requirements in Uropathogenic Escherichia coli .” Microbiology Spectrum 12: e02236‐02223. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Capper‐Parkin, K. L. , Nichol T., Smith T. J., Lacey M. M., and Forbes S.. 2023. “Antimicrobial and Cytotoxic Synergism of Biocides and Quorum‐Sensing Inhibitors Against Uropathogenic Escherichia coli .” Journal of Hospital Infection 134: 138–146. [DOI] [PubMed] [Google Scholar]
- Donovan, G. T. , Norton J. P., Bower J. M., and Mulvey M. A.. 2013. “Adenylate Cyclase and the Cyclic AMP Receptor Protein Modulate Stress Resistance and Virulence Capacity of Uropathogenic Escherichia coli .” Infection and Immunity 81: 249–258. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Faleye, O. S. , Boya B. R., Lee J.‐H., Choi I., and Lee J.. 2024. “Halogenated Antimicrobial Agents to Combat Drug‐Resistant Pathogens.” Pharmacological Reviews 76: 90–141. [DOI] [PubMed] [Google Scholar]
- Fan, X. , Wang J., Soman K. V., Ansari G. A. S., and Khan M. F.. 2011. “Aniline‐Induced Nitrosative Stress in Rat Spleen: Proteomic Identification of Nitrated Proteins.” Toxicology and Applied Pharmacology 255: 103–112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gallarato, L. A. , Mulko L. E., Dardanelli M. S., Barbero C. A., Acevedo D. F., and Yslas E. I.. 2017. “Synergistic Effect of Polyaniline Coverage and Surface Microstructure on the Inhibition of Pseudomonas aeruginosa Biofilm Formation.” Colloids and Surfaces B: Biointerfaces 150: 1–7. [DOI] [PubMed] [Google Scholar]
- García, V. , Lestón L., Parga A., et al. 2023. “Genomics, Biofilm Formation and Infection of Bladder Epithelial Cells in Potentially Uropathogenic Escherichia coli (UPEC) From Animal Sources and Human Urinary Tract Infections (UTIs) Further Support Food‐Borne Transmission.” One Health 16: 100558. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gomez‐Gomez, J. M. , Baquero F., and Blazquez J.. 1996. “Cyclic AMP Receptor Protein Positively Controls gyrA Transcription and Alters DNA Topology After Nutritional Upshift in Escherichia coli .” Journal of Bacteriology 178: 3331–3334. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Halim, A. N. A. , Diosing D. N., Zamakshshari N. H., Ngaini Z., Wei Y. K., and Hussin A. S. M.. 2024. “Synthesis, In Vitro and In Silico Bacteriostatic Evaluation of Halogenated Triazene Derivatives.” Russian Journal of General Chemistry 94: 2068–2079. [Google Scholar]
- Heffernan, J. R. 2023. Function and Regulation of Virulence‐Associated Siderophores in Uropathogenic Enterobacterales, 114. Washington University in St. Louis. [Google Scholar]
- Hirakawa, H. , Shimokawa M., Noguchi K., et al. 2023. “The PapB/FocB Family Protein TosR Acts as a Positive Regulator of Flagellar Expression and Is Required for Optimal Virulence of Uropathogenic Escherichia coli .” Frontiers in Microbiology 14: 1185804. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hreha, T. N. , Gilbert N. M., and Hunstad D. A.. 2024. “Chapter 61 ‐ Uropathogenic Escherichia coli in Urinary Tract Infections.” In Molecular Medical Microbiology, edited by Tang Y.‐W., Hindiyeh M. Y., Liu D., Sails A., Spearman P., and Zhang J.‐R., 3rd ed., 1271–1297. Academic Press. [Google Scholar]
- Hufnagel, D. A. , Evans M. L., Greene S. E., Pinkner J. S., Hultgren S. J., and Chapman M. R.. 2016. “The Catabolite Repressor Protein‐Cyclic AMP Complex Regulates csgD and Biofilm Formation in Uropathogenic Escherichia coli .” Journal of Bacteriology 198: 3329–3334. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hufnagel, D. A. , Tükel Ç., and Chapman M. R.. 2013. “Disease to Dirt: The Biology of Microbial Amyloids.” PLoS Pathogens 9: e1003740. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Islam, M. , Hossain A., Yamari I., et al. 2024. “Synthesis, Antimicrobial, Molecular Docking Against Bacterial and Fungal Proteins and In Silico Studies of Glucopyranoside Derivatives as Potent Antimicrobial Agents.” Chemistry & Biodiversity 21: e202400932. [DOI] [PubMed] [Google Scholar]
- Kim, C.‐M. , Kim S.‐J., and Shin S.‐H.. 2012. “Cyclic AMP‐Receptor Protein Activates Aerobactin Receptor IutA Expression in Vibrio vulnificus .” Journal of Microbiology 50: 320–325. [DOI] [PubMed] [Google Scholar]
- Kot, B. 2019. “Antibiotic Resistance Among Uropathogenic Escherichia coli .” Polish Journal of Microbiology 68: 403–415. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liang, D.‐D. , Lional N., Scheepmaker B., et al. 2023. “Enantiospecific Synthesis of Aniline‐Derived Sulfonimidamides.” Organic Letters 25: 5666–5670. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Linder, J. U. 2008. “Structure–Function Relationships in Escherichia coli Adenylate Cyclase.” Biochemical Journal 415: 449–454. [DOI] [PubMed] [Google Scholar]
- Liu, L. , Ma X., Bilal M., et al. 2022. “Mechanistic Insight Into Phenolic Compounds Toxicity and State‐Of‐The‐Art Strategies for Enhancing the Tolerance of Escherichia coli to Phenolic Compounds.” Biotechnology and Bioprocess Engineering 27: 533–542. [Google Scholar]
- Long, J. , Yang C., Liu J., et al. 2024. “Tannic Acid Inhibits Escherichia coli Biofilm Formation and Underlying Molecular Mechanisms: Biofilm Regulator CsgD.” Biomedicine & Pharmacotherapy 175: 116716. [DOI] [PubMed] [Google Scholar]
- Luna‐Pineda, V. M. , Moreno‐Fierros L., Cázares‐Domínguez V., et al. 2019. “Curli of Uropathogenic Escherichia coli Enhance Urinary Tract Colonization as a Fitness Factor.” Frontiers in Microbiology 10: 2063. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mary, A. , Kanagathara N., and Baby Suganthi A. R.. 2020. “A Brief Review on Aniline and Its Derivatives.” Materials Today Proceedings 33: 4751–4755. [Google Scholar]
- Mitra, A. , Herren C. D., Patel I. R., Coleman A., and Mukhopadhyay S.. 2016. “Integration of AI‐2 Based Cell‐Cell Signaling With Metabolic Cues in Escherichia coli .” PLoS One 11: e0157532. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mohammed, M. , Mekala L. P., Chintalapati S., and Chintalapati V. R.. 2020. “New Insights Into Aniline Toxicity: Aniline Exposure Triggers Envelope Stress and Extracellular Polymeric Substance Formation in Rubrivivax benzoatilyticus JA2.” Journal of Hazardous Materials 385: 121571. [DOI] [PubMed] [Google Scholar]
- Moore, J. D. , Gerdt J. P., Eibergen N. R., and Blackwell H. E.. 2014. “Active Efflux Influences the Potency of Quorum Sensing Inhibitors in Pseudomonas aeruginosa .” Chembiochem 15, no. 3: 435–442. 10.1002/cbic.201300701. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moreland, R. B. , Choi B. I., Geaman W., et al. 2023. “Beyond the Usual Suspects: Emerging Uropathogens in the Microbiome Age.” Frontiers in Urology 3: 1212590. [Google Scholar]
- Müller, C. M. , Åberg A., Straseviçiene J., Emődy L., Uhlin B. E., and Balsalobre C.. 2009. “Type 1 Fimbriae, a Colonization Factor of Uropathogenic Escherichia coli , Are Controlled by the Metabolic Sensor CRP‐cAMP.” PLoS Pathogens 5: e1000303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nanchen, A. , Schicker A., Revelles O., and Sauer U.. 2008. “Cyclic AMP‐Dependent Catabolite Repression Is the Dominant Control Mechanism of Metabolic Fluxes Under Glucose Limitation in Escherichia coli .” Journal of Bacteriology 190: 2323–2330. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ono, K. , Oka R., Toyofuku M., et al. 2014. “cAMP Signaling Affects Irreversible Attachment During Biofilm Formation by Pseudomonas aeruginosa PAO1.” Microbes and Environments 29: 104–106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Parker, A. , Cureoglu S., De Lay N., Majdalani N., and Gottesman S.. 2017. “Alternative Pathways for Escherichia coli Biofilm Formation Revealed by sRNA Overproduction.” Molecular Microbiology 105: 309–325. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pervaiz, M. , Shahin M., Ejaz A., et al. 2024. “An Overview of Aniline‐Based Schiff Base Metal Complexes: Synthesis, Characterization and Biological Activities—a Review.” Inorganic Chemistry Communications 159: 111851. [Google Scholar]
- Sathiyamoorthi, E. , Boya B. R., Lee J.‐H., and Lee J.. 2025. “Antimicrobial Efficacy of Trifluoro‐Anilines Against Vibrio Species.” International Journal of Molecular Sciences 26: 623. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shah, C. , Baral R., Bartaula B., and Shrestha L. B.. 2019. “Virulence Factors of Uropathogenic Escherichia coli (UPEC) and Correlation With Antimicrobial Resistance.” BMC Microbiology 19: 204. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sudarshan, S. , Hogins J., Ambagaspitiye S., Zimmern P., and Reitzer L.. 2021. “Nutrient and Energy Pathway Requirements for Surface Motility of Nonpathogenic and Uropathogenic Escherichia coli .” Journal of Bacteriology 203: 10–1128. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Terlizzi, M. E. , Gribaudo G., and Maffei M. E.. 2017. “Uropathogenic Escherichia coli (UPEC) Infections: Virulence Factors, Bladder Responses, Antibiotic, and Non‐Antibiotic Antimicrobial Strategies.” Frontiers in Microbiology 8: 1566. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tripathi, M. , Khan S. I., Ponnan P., Kholiya R., and Rawat D. S.. 2017. “Aminoquinoline‐Pyrimidine‐Modified Anilines: Synthesis, In Vitro Antiplasmodial Activity, Cytotoxicity, Mechanistic Studies and ADME Predictions.” ChemistrySelect 2: 9074–9083. [Google Scholar]
- Ulett, G. C. , Valle J., Beloin C., Sherlock O., Ghigo J.‐M., and Schembri M. A.. 2007. “Functional Analysis of Antigen 43 in Uropathogenic Escherichia coli Reveals a Role in Long‐Term Persistence in the Urinary Tract.” Infection and Immunity 75: 3233–3244. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang, Y. , Gao H., Na X.‐L., et al. 2016. “Aniline Induces Oxidative Stress and Apoptosis of Primary Cultured Hepatocytes.” International Journal of Environmental Research and Public Health 13: 1188. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Whelan, S. , Lucey B., and Finn K.. 2023. “Uropathogenic Escherichia coli (UPEC)‐Associated Urinary Tract Infections: The Molecular Basis for Challenges to Effective Treatment.” Microorganisms 11: 2169. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang, X. , Yan Y., Lv Y., et al. 2022. “ Dendrobium officinale Polysaccharides Attenuate Uropathogenic Escherichia coli (UPEC)‐Induced Pyroptosis in Macrophage Cells.” Biomedicine & Pharmacotherapy 151: 113098. [DOI] [PubMed] [Google Scholar]
- Zidar, N. , Macut H., Tomašič T., et al. 2015. “N‐Phenyl‐4,5‐Dibromopyrrolamides and N‐Phenylindolamides as ATP Competitive DNA Gyrase B Inhibitors: Design, Synthesis, and Evaluation.” Journal of Medicinal Chemistry 58: 6179–6194. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data S1
Data Availability Statement
All data generated or analysed during this study are included in this published article and its Supporting Information files.


