Abstract
Targeted next-generation sequencing (tNGS) offers a high-throughput, culture-independent approach that delivers a comprehensive resistance profile in a significantly shorter turn-around time, making it promising in enhancing tuberculosis (TB) diagnosis and informing treatment decisions. This study aims to evaluate the performance of tNGS in the TB diagnosis and drug resistance detection of Mycobacterium tuberculosis (MTB) using MTB clinical isolates and bronchoalveolar lavage fluid (BALF) samples. A total of 143 MTB clinical isolates were assessed, tNGS, phenotypic antimicrobial susceptibility testing (AST), and AST based on whole genome sequencing (WGS) exhibited high concordance rates, averaging 95.10% and 97.05%. Among 158 BALF samples, culture, Xpert MTB/RIF, and tNGS reported 29, 70 and 111 positives, respectively. In the confirmed cases with etiological evidence (smears, cultures, or molecular test), the positive rate of tNGS (73/83, 87.95%) was higher than that of Xpert MTB (67/83, 80.72%). Additionally, 45% (27/60) of clinically diagnosed cases (with imaging or immunological evidence) were positive for tNGS. Further validation on the discrepant results between tNGS and Xpert MTB/RIF with droplet digital PCR (ddPCR) yielded 35 positives, tNGS detected all, and Xpert MTB/RIF only identified 6 positives. In conclusion, tNGS demonstrates robust and rapid performance in the identification of MTB and its associated drug resistance, and can be directly applied to clinical samples, positioning it as a promising approach for laboratory testing of tuberculosis.
Supplementary Information
The online version contains supplementary material available at 10.1007/s15010-024-02411-w.
Keywords: Mycobacterium tuberculosis, Targeted next-generation sequencing, Diagnosis, Drug resistance
Introduction
Tuberculosis caused by Mycobacterium tuberculosis (MTB) is the second leading cause of death from a single source of infection after COVID-19. Globally, the incidence rate of tuberculosis only declined by 8.7% from 2015 to 2022, and the phased goal of reducing the incidence rate of tuberculosis by 50% by 2025 is difficult to achieve [1]. The World Health Organization (WHO) reported that approximately 10.6 million new TB cases and 1.3 million deaths occurred globally in 2022 [1]. The prompt, precise, and thorough detection of MTB and its drug susceptibility is essential for the effective management and prevention of tuberculosis. Consequently, it is imperative to expedite the advancement and implementation of laboratory diagnostic techniques for TB, particularly those capable of directly detecting pathogens in clinical samples.
Currently, culture testing remains as a major reference standard for TB diagnosis [1]. The slow bacterial growth rate, complex operational procedures, and strict biosafety requirements of MTB make culture-based detection methods incapable of meeting the need for swift clinical detection. The evolution of laboratory diagnostic technologies has fostered the adoption of molecular biology methods in identifying antimicrobial resistance in MTB. While PCR-based molecular diagnostic techniques have shown significant advancements in sensitivity, specificity, and detection speed compared to culture-based methods, their efficacy is constrained by the limited range of drug resistance gene targets they can identify. Furthermore, variations exist in detection targets and performance across different diagnostic technologies [2]. For example, Xpert MTB/RIF Ultra is a detection method recommended by WHO in 2017 presenting a substantially lower limit of detection (LoD) than that of Xpert MTB/RIF due to the use of multi-copy IS1081 and IS6110 insertion elements as MTB target sequences, and both of them have similar performance in detecting rifampicin resistance [3].
Targeted next-generation sequencing (tNGS) is a promising diagnostic technique for tuberculosis, offering the ability to reveal a comprehensive profile of resistance genotyping. This method combines gene-specific amplification with next-generation sequencing to detect resistance to multiple drugs within a single assay. The WHO has released an implementation manual for the use of tNGS in monitoring drug-resistant tuberculosis, as it’s featured by multiple advantages, including, but not limited to, culture-independent, directly applicable to clinical samples, reporting heterogeneous/silent mutations [4]. This test represents a major advancement in diagnostic technology and provides an option for rapid and accurate genetic analysis and detection of mutations associated with resistance, requiring only a small fraction of the time needed for culture-based methods for detecting resistance [5]. Targeted NGS has been used in clinical microbiology due to its ability to detect dozens to hundreds of known pathogenic microorganisms and their virulence and/or resistance genes [6–17], while PCR-based methods could only cover much smaller spectrum. The first-line and second-line anti-tuberculosis drugs and corresponding drug resistance genes found include katG, inhA, kasA, ndh, rpoB, embB, embC, embA, pncA, rpsL, rrs, gyrA, and gyrB. The WHO released a catalog of MTBC resistant mutations in June 2021, and updated it in November 2023, elucidating the association between specific gene mutations and drug resistance in MTB [18, 19].
In this study, we utilized tNGS to identify MTB in clinical isolates and bronchoalveolar lavage fluid (BALF) samples. We compared its efficacy with culture, phenotypic AST, whole-genome sequencing (WGS), and Xpert MTB/RIF. We conducted a thorough evaluation of tNGS’s performance in TB diagnosis and the detection of drug-resistant gene mutations.
Materials and methods
Study design and participants
This study was conducted at Shanghai Pulmonary Hospital (SPH), Tongji University School of Medicine, which is a modern grade A tertiary hospital that integrates medical care, teaching, and research functions. The schematic of the study design is shown in Fig. 1. 60 Rifampicin-resistant and 83 susceptible TB clinical isolates with genome sequencing data and phenotypic AST data were randomly selected from TB strains stored at -80℃ in the clinical laboratory in SPH [20, 21]. A total of 158 BALF samples enrolled in the study were collected from non-repetitive inpatients in the department of tuberculosis of SPH from April 18, 2023 to August 24, 2023. The inclusion criteria are for individuals aged 12 and above, as well as those who tested HIV-negative prior to enrollment and had a volume of collected BALF greater than 6mL. Further information on clinical symptoms, basic disease, tuberculosis history, tuberculosis contact, and results of QuantiFERON®-TB Gold Plus (Qiagen, Hilden, Germany) were collected.This study was approved by the Ethical Committee of SPH (No. K23-265Z).
Fig. 1.
Comparative analysis and workflow schematic of MTB diagnosis and drug resistance detection. (A) Comparison of the performance of tNGS, WGS, and phenotypic AST in detecting drug resistance on MTB isolates. (B) Comparison of the performance in MTB and rifampicin-resistant gene identification between tNGS and Xpert MTB/RIF on BALF samples. (C) The workflow of the tNGS in routine clinical practice
The diagnosis of tuberculosis was based on the “Diagnosis of Pulmonary tuberculosis” standard of the People’s Republic of China Health Industry Standard (WS 288–2017) [22]. The principle of diagnosis was to primarily rely on etiological (including bacteriology, molecular biology) examinations, combined with epidemiological history, clinical manifestations, chest imaging, related auxiliary examinations, and differential diagnosis, to make a comprehensive analysis for the diagnosis. Additionally, patients were divided into four cohorts in reference to this document: confirmed cases, clinically diagnosed cases, suspected cases, and non-tuberculosis cases. Suspected cases were defined as having only abnormal chest imaging consistent with tuberculosis. Clinically diagnosed cases were those that, after differential diagnosis to rule out other lung diseases, have chest imaging abnormalities consistent with tuberculosis and clinical symptoms and signs of tuberculosis or positive tuberculosis immunological tests. Confirmed cases were those with positive tuberculosis etiological tests (smears, cultures, or molecular biology methods) and are clinically diagnosed cases. Non-tuberculosis cases were those that, after differential diagnosis, had ruled out tuberculosis [22].
MTB identification and drug resistance profiling
For the analysis of MTB clinical isolates, the preserved bacterial liquid was directly transferred to Löwenstein-Jensen medium to culture the bacteria. Once colonies were formed, they were collected and resuspended in a 1 mL buffer solution (pH 8.0; containing 10 mM Tris-HCl and 1 mM ethylenediaminetetraacetic acid). This suspension was then heat-inactivated at 95 °C for 10 min to prepare the samples for further testing.
The phenotypic AST was conducted using the Myco TB system (Trek Diagnostic Systems, Thermo Fisher Scientific, USA), which provides detailed information on the drug resistance profiles of the isolates.
For genomic analysis, WGS data were previously analyzed using the TB Profiler tool, as documented in other studies [20, 21]. This tool facilitates the identification of known drug resistance mutations in MTB genomes in reference to the WHO guidelines. It should be noted that the confidence gradings for three mutations (rrs 799C > T, rrs 888G > A, and rrs 1402C > A) were classified as having “Uncertain significance”. Therefore, these resistance loci were excluded and not considered as tuberculosis drug resistance mutations.
The collected BALF was divided into three equal parts for culture, Xpert MTB/RIF and tNGS by professional staff in accordance with standard operating procedures. The Bactec MGIT 960 culture instrument (Becton Dickinson, Cockeysville, MD, USA) and GeneXpert Infinity (Cepheid, Sunnyvale, United States of America [USA]) were used for culture and Xprert MTB/RIF (Cepheid, Sunnyvale, United States of America [USA]), respectively. All MTB isolates were confirmed as MTB using MBP 64 antigen detection kits (Genesis, Hangzhou, China), and the phenotypic AST was performed using Myco TB system (Trek Diagnostic Systems, ThermoFisher Scientific Inc., USA) [23].
Nucleic acid extraction
2mL of BALF samples were collected, total nucleic acid extraction was conducted using the Nucleic Acid Extraction and Purification Kit (KS118-BYTQ-96, KingCreate, Guangzhou, China). This extraction process was automated on the automation platform KingFisher Flex Purification System (24074420, Thermo Fisher Scientific, Waltham, USA). To ensure the quality of pathogen nucleic acid extraction, both positive control (PC) and negative control (NC) were included in the extraction process. These controls were provided within the MTB Premix kit (KS620-JHFZGJ96, KingCreate, Guangzhou, China) and underwent extraction simultaneously with our test samples. UltraPure™ DNase/RNase-Free Distilled Water (10977023, Invitrogen, Waltham, USA) was utilized as non-template-control (NTC) to detect any contamination from foreign nucleic acids during the extraction process.
Library construction and sequencing
Nucleic acids extracted from all samples, including PC, NC, and NTC, were combined with the One-Step Reaction Solution from the MTB Premix kit. The mixture was then subjected to PCR amplification under specific thermal cycling conditions. The amplification protocol commenced with an initial denaturation at 95 °C for 3 min, followed by 25 cycles of DNA denaturation at 95 °C for 30 s and annealing at 68 °C for 1 min. Subsequently, the samples underwent 30 cycles of sequential heating, which included denaturation at 95 °C for 30 s, annealing at 60 °C for 30 s, and extension at 72 °C for 30 s. The final steps involved a prolonged extension at 72 °C for 1 min to ensure the completion of all partially amplified fragments, before cooling the reaction to 4 °C for storage.
Following PCR amplification, the resulting product underwent purification utilizing the MTB Magnetic Beads included in the MTB Premix kit. The concentration of the purified PCR product was then measured with the Equalbit DNA HS Assay Kit (Eq. 121, Vazyme, Nanjing, China), using the Invitrogen™ Qubit™ 3.0/4.0 Fluorometer (Eqs. 121-02, Thermo Fisher Scientific, Waltham, USA). The concentration of all the samples was adjusted through dilution to a uniform level to facilitate library construction. The sequencing library was constructed using the general sequencing kit MR100 (KS107-CXR, KingCreate, Guangzhou, China) and sequenced on the KM MiniSeq Dx-CN Platform (KY301, KingCreate, Guangzhou, China).
Bioinformatics
The raw data from the sequencing process was subjected to a quality control regimen. For the purposes of adaptor removal and discarding subpar quality reads, fastp v0.20.1 was utilized with its standard settings [24]. Samples with raw reads possessing a Q30 ≥ 75% and a minimum threshold of 50,000 reads were deemed acceptable. In instances where these criteria were not met, the samples were re-sequenced to ensure the reliability of the final data. Once filtered, the high-quality reads were then reconstructed into extended sequences via the BWA-mem algorithm [25]. A comprehensive reference database of MTBC and resistance gene target loci was cureated from several sources, mainly from Genbank and Refseq [26, 27]. Positive identification of pathogens was determined by the number of the normalized mapped reads, with a specific threshold of reads per 100,000 (RPhK). A sample was marked positive for a given pathogen if it exhibited relevant RPhK values. A species or taxonomic group was considered to be “present” within a sample if it was detected at specific target loci with an RPhK of 3 or higher. Conversely, it was labeled “absent” if such criteria were not fulfilled. Regarding drug resistance genes, given a single nucleotide polymorphism (SNP) revealed on a specific loci, it will only be reported if the mutation rate on this position is equal and greater than 0.1.
Droplet digital PCR
Droplet digital PCR (ddPCR) was used for verification on BALF samples with inconsistent MTB identification results by tNGS and Xpert MTB/RIF. A liquid reaction system for ddPCR amplification was generated as follows: 11µL of ddPCR Supermix for Probes (1863024, Bio-Rad Laboratories, Hercules, California), primer and probe mix of 1µL, 5µL DNA template, and 5µL of UltraPure™ DNase/RNase-Free Distilled Water (10977023, Invitrogen, Waltham, USA) to reach a final total volume of 22µL. the prepared mix was transferred to the QX200™ Droplet Generator (1864002, Bio-Rad Bio-Rad Laboratories, Hercules, California). Subsequently, 70µL of droplet generation oil was added, and ddPCR amplification was performed on QX200™ Droplet Digital PCR System (1864001, Bio-Rad Laboratories, Hercules, California). The thermal cycling conditions were as follows: initial denaturation at 95 °C for 10 min; followed by 40 cycles of denaturation at 94 °C for 30 s, annealing at 54 °C for 1 min; and a final extension at 72 °C for 10 min. Upon completion of amplification, the reaction was cooled to 4 °C for storage. After PCR amplification was complete, the products were transferred to the droplet reader (QX200™ Droplet Reader, Bio-Rad Laboratories, Hercules, California) for quantification.
Statistical analysis
All statistics, including concordance, sensitivity, specificity, positive predictive value (PPV), negative predictive value (NPV), and 95% confidence interval (CI) were computed in reference to the formulas described in previous studies [28–30], which are implemented in MedCalc statistical software v20.022 (https://www.medcalc.org/).
Results
Antimicrobial susceptibility patterns of MTB clinical isolates
For 143 MTB clinical isolates, the antimicrobial susceptibility patterns based on phenotype, WGS, and tNGS are shown in Table 1. The phenotypic AST showed that the resistance rates of 9 drugs ranged from 2.80% (amikacin) to 48.95% (isoniazid). According to the results of AST based on tNGS, the drug-resistance rates ranged from 4.90% (amikacin and kanamycin) to 46.15% (rifampicin) and the difference between the resistance rate of tNGS and phenotypic AST in each drug does not exceed 5%. For ethambutol and ethionamide, the drug resistance rates based on WGS differed significantly from those based on phenotypic AST and tNGS, at 31.47% and 16.08%, respectively.
Table 1.
The antimicrobial susceptibility patterns based on phenotype, WGS, and tNGS
| phenotypic AST | AST based on WGS | AST based on tNGS | |||||||
|---|---|---|---|---|---|---|---|---|---|
| susceptible | resistant | resistance rate | susceptible | resistant | resistance rate | susceptible | resistant | resistance rate | |
| Rifampicin | 83 | 60 | 41.96% | 74 | 69 | 48.25% | 77 | 66 | 46.15% |
| Isoniazid | 73 | 70 | 48.95% | 76 | 67 | 46.85% | 78 | 65 | 45.45% |
| Ethambutol | 111 | 32 | 22.38% | 98 | 45 | 31.47% | 108 | 35 | 24.48% |
| Streptomycin | 93 | 50 | 34.97% | 90 | 53 | 37.06% | 92 | 51 | 35.66% |
| Moxifloxacin | 108 | 35 | 24.48% | 106 | 37 | 25.87% | 109 | 34 | 23.78% |
| Ofloxacin | 109 | 34 | 23.78% | 106 | 37 | 25.87% | 109 | 34 | 23.78% |
| Amikacin | 139 | 4 | 2.80% | 137 | 6 | 4.20% | 136 | 7 | 4.90% |
| Kanamycin | 137 | 6 | 4.20% | 136 | 7 | 4.90% | 136 | 7 | 4.90% |
| Ethionamide | 136 | 7 | 4.90% | 120 | 23 | 16.08% | 134 | 9 | 6.29% |
AST: antimicrobial susceptibility testing
Performance of tNGS in detecting drug resistance of MTB clinical isolates
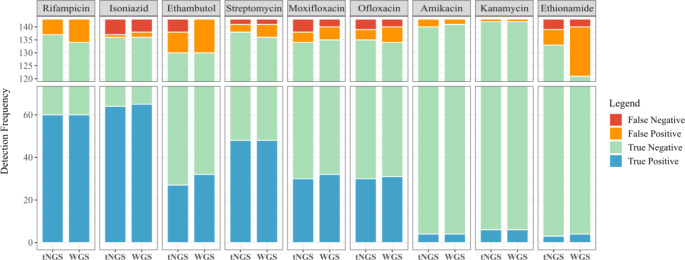
Using phenotypic AST as the reference standard, we evaluated the ability of tNGS and WGS to detect drug resistance. For 143 isolates, an average concordance between tNGS and phenotypic AST of 95.10% was achieved across all nine drugs, ranging from 90.91% (ethambutol) to 99.30% (kanamycin). The overall performance of tNGS and WGS including true positive, true negative, false positive, and false negative for each drug is shown in Fig. 2. The false positives of tNGS, that is, false resistance, ranged from 1 case (isoniazid and kanamycin) to 8 cases (ethambutol). At the same time, the false negatives of tNGS, that is, false sensitivity, ranged from 0 (rifampicin, amikacin, and kanamycin) to 6 cases (isoniazid).
Fig. 2.
The overall performance of tNGS and WGS for each drug using phenotypic AST as the reference standard
The sensitivity and specificity of tNGS compared with WGS in detecting drug resistance of nine drugs are shown in Table 2. The average consistency between tNGS and WGS in detecting the drug resistance of nine drugs was 97.05%. The sensitivity and specificity of tNGS in predicting MTB drug resistance were respectively, 95.65% (87.82-99.09%) and 100.00% (95.14-100.00%) for rifampicin, 97.01% (89.63-99.64%) and 100.00% (95.26-100.00%) for isoniazid, 77.78% (62.91-88.80%) and 100.00% (96.31-100.00%) for ethambutol, 96.23% (87.02-99.54%) and 100.00% (95.98-100.00%) for streptomycin, 91.89% (78.09-98.30%) and 100.00% (96.58-100.00%) for moxifloxacin, 91.89% (78.09-98.30%) and 100.00% (96.58-100.00%) for ofloxacin, 100.00% (54.07-100.00%) and 99.27% (96.00-99.98%) for amikacin, 100.00% (59.04-100.00% ) and 100.00% (97.32-100.00%) for kanamycin, and 39.13% (19.71-61.46%) and 100.00% (96.97-100.00%) for ethionamide. The drug-resistance predictive values, except for amikacin, were all 100%. The drug-susceptibility predictive values were greater than 95% except for ethambutol and ethionamide.
Table 2.
Drug resistance detection performance of tNGS compared with WGS
| Drug | WGS-sensitive | WGS-resistant | Concordance (%) | Sensitivity (%) |
Specificity (%) |
Positive Predictive Value (%) | Negative Predictive Value (%) | ||
|---|---|---|---|---|---|---|---|---|---|
| tNGS- susceptible |
tNGS -resistant |
tNGS- susceptible |
tNGS- resistant |
||||||
| Rifampicin | 74 | 0 | 3 | 66 | 97.90 | 95.65 (87.82–99.09) | 100.00 (95.14–100.00) | 100.00(94.56–100.00) | 96.10 (89.03–99.19) |
| Isoniazid | 76 | 0 | 2 | 65 | 98.60 | 97.01 (89.63–99.64) | 100.00 (95.26–100.00) | 100.00 (94.48–100.00) | 97.44 (91.04–99.69) |
| Ethambutol | 98 | 0 | 10 | 35 | 93.01 | 77.78 (62.91–88.80) | 100.00 (96.31–100.00) | 100.00 (90.00-100.00) | 90.74 (83.63–95.47) |
| Streptomycin | 90 | 0 | 2 | 51 | 98.60 | 96.23 (87.02–99.54) | 100.00 (95.98–100.00) | 100.00 (93.02–100.00) | 97.83 (92.37–99.74) |
| Moxifloxacin | 106 | 0 | 3 | 34 | 97.90 | 91.89 (78.09–98.30) | 100.00 (96.58–100.00) | 100.00 (89.72–100.00) | 97.25 (92.17–99.43) |
| Ofloxacin | 106 | 0 | 3 | 34 | 97.90 | 91.89 (78.09–98.30) | 100.00 (96.58–100.00) | 100.00 (89.72–100.00) | 97.25 (92.17–99.43) |
| Amikacin | 136 | 1 | 0 | 6 | 99.30 | 100.00 (54.07–100.00) | 99.27 (96.00-99.98) | 85.71 (42.13–99.64) | 100.00 (97.32–100.00) |
| Kanamycin | 136 | 0 | 0 | 7 | 100.00 | 100.00 (59.04–100.00) | 100.00 (97.32–100.00) | 100.00 (59.04–100.00) | 100.00 (97.32–100.00) |
| Ethionamide | 120 | 0 | 14 | 9 | 90.21 | 39.13 (19.71–61.46) | 100.00 (96.97–100.00) | 100.00 (66.37–100.00) | 89.55 (83.09–94.17) |
As shown in Appendix Table, the majority of MTB isolates resistant in WGS but susceptible in tNGS were due to drug resistance mutations detected by WGS that were not covered by tNGS. Additionally, one isolate had embB Met306Leu mutation detected by WGS with a mutation frequency of 0.1, but not detected by tNGS. Another isolate had gyrA Asp94Gly mutation detected by WGS with a mutation frequency of 0.2, and this mutation was detected by tNGS with a mutation frequency of 0.08, which was filtered out and not reported. Conversely, there was only one isolate that was resistant in tNGS but susceptible in WGS. This isolate was detected by tNGS to have a mutation of rrs 1402C > T, with a mutation frequency of 0.1, reported as amikacin resistance, but this mutation was not detected by WGS.
Clinical characteristics
A total of 158 cases were included in this study, including 83 confirmed cases, 60 clinically diagnosed cases, 7 suspected cases, and 8 non-tuberculosis cases. The demographic characteristics of the included participants are shown in Table 3. Among them, there were 78 (49.37%) males and 137 (86.71%) married, with ages ranging from 15 to 83 years old (mean: 49 years old). Of all, 58 (36.71%) had symptoms of coughing and expectoration for more than 2 weeks, 15 had hypertension, and 8 had a history of tuberculosis. The number of positive results for QuantiFERON®-TB Gold Plus in the confirmed cases and the clinically diagnosed cases were 53 and 42, respectively.
Table 3.
The demographic characteristics of the included participants in this study
| Suspected cases | Clinically diagnosed cases | Confirmed cases | Non-tuberculosis cases | Total | ||
|---|---|---|---|---|---|---|
| Gender | Male | 5 | 26 | 46 | 1 | 78 |
| Female | 2 | 34 | 37 | 7 | 80 | |
| Age | <20 | 0 | 2 | 0 | 0 | 2 |
| 20–40 | 1 | 19 | 22 | 2 | 44 | |
| 40–60 | 4 | 26 | 36 | 4 | 70 | |
| ≥ 60 | 2 | 13 | 25 | 2 | 42 | |
| Marital status | Unmarried | 0 | 10 | 11 | 0 | 21 |
| Married | 7 | 50 | 72 | 8 | 137 | |
| Symptoms | Cough, expectoration ≥ 2 weeks | 3 | 21 | 31 | 3 | 58 |
| Coughing up blood or spitting blood | 2 | 3 | 3 | 2 | 10 | |
| Systemic symptoms # | 2 | 13 | 24 | 3 | 42 | |
| Basic Disease | Diabetes Mellitus | 0 | 4 | 9 | 0 | 13 |
| Hypertension | 0 | 8 | 6 | 1 | 15 | |
| Tuberculosis history | 0 | 2 | 6 | 0 | 8 | |
| Tuberculosis contact history | 0 | 2 | 2 | 0 | 4 | |
| QuantiFERON®-TB Gold Plus* | 0 | 42 | 53 | 0 | 95 | |
# Night sweats, fatigue, fever, loss of appetite, weight loss;
* There were 16 cases that did not undergo QuantiFERON®-TB Gold Plus testing, including 11 confirmed cases and 5 clinically diagnosed cases
Performance of tNGS in diagnosis and drug resistance detection of MTB on BALF samples
The results of culture, Xpert MTB/RIF and tNGS in different groups are shown in Table 4. Among 158 BALF samples, the number of positive results for culture, Xpert MTB/RIF, and tNGS were 29, 70 and 111, respectively. Among these, in 83 confirmed cases, the number of positive results for culture, Xpert MTB/RIF and tNGS were 29, 67 and 73, respectively. Additionally, there were 27 clinically diagnosed cases with positive tNGS results. In the 29 cases of confirmed cases with positive results of culture, 5 cases were Xpert MTB/RIF-negative, and only one case was tNGS-negative.
Table 4.
The results of culture, Xpert MTB/RIF and tNGS in different groups
| Culture | Xpert MTB/RIF# | tNGS | ||||
|---|---|---|---|---|---|---|
| positive | negative | positive | negative | positive | negative | |
| Confirmed cases (N = 83) |
29 (34.94%) |
54 (65.06%) |
67 (80.72%) |
16 (19.28%) |
73 (87.95%) |
10 (12.05%) |
| Clinically diagnosed cases (N = 60) | 0 |
60 (100.00%) |
0 |
60 (100.00%) |
27 (45.00%) |
33 (55.00%) |
| Suspected cases (N = 7) | 0 |
7 (100.00%) |
2 (28.57%) |
5 (71.43%) |
5 (71.43%) |
2 (28.57%) |
| Non-tuberculosis cases (N = 8) | 0 |
8 (100.00%) |
1 (12.50%) |
7 (87.50%) |
6 (75.00%) |
2 (25.00%) |
# Combining symptoms, physical signs, and imaging studies, two chief physicians made a differential diagnosis. Two individuals with positive Xpert MTB/RIF were identified as “suspected cases”, and one individual with positive Xpert MTB/RIF was identified as “non-tuberculosis case”
The results of MTB identification by tNGS and Xpert MTB/RIF were inconsistent in 51 cases. Among them, 5 cases were reported as “MTB detected” by Xpert MTB/RIF, while tNGS reported negative, and the semi-quantitative results of Xpert MTB/RIF in these 5 cases were “very low”. The remaining 46 cases were reported as “MTB not detected” by Xpert MTB/RIF, but detected MTB by tNGS.
The comparison of tNGS and Xpert MTB/RIF results in 158 BALF samples are shown in Table 5. The results of Xpert MTB/RIF showed that there were 61 cases of “MTB detected, rifampicin resistance not detected”, including 58 confirmed cases, 2 suspected cases, and 1 non-tuberculosis case. At the same time, tNGS detected 29 cases of MTB positive and rifampicin-susceptible, including 26 confirmed cases and 3 clinically diagnosed cases. In 78 tNGS-positive samples, rifampicin resistance could not be detected because the concentration of MTB was too low, and it was reported as “rifampicin not applicable”. For 5 BALF samples with Xpert MTB/RIF results reported as “MTB detected, rifampicin resistance detected”, tNGS results showed that 3 cases were resistant, and 2 cases were not applicable for rifampicin. Among the four cases of tNGS reporting rifampicin resistance, one case was reported by Xpert MTB/RIF as “MTB detected, rifampicin resistance not detected”. The tNGS result of this case showed a rifampicin resistance mutation site at rpoB Leu511Pro with a mutation frequency of 0.6. Upon reviewing the case, we found that the patient was a confirmed case with tuberculosis. The patient had been treated with anti-tuberculosis drugs at another hospital for six months, during which three Xpert MTB/RIF tests all indicated “MTB detected, rifampicin resistance not detected”. Currently, the lesions have progressed, and there is suspicion of multidrug-resistant tuberculosis. In summary, this case may represent heterogeneous drug resistance.
Table 5.
Comparison of tNGS and Xpert MTB/RIF results in 158 BALF samples
| Xpert MTB/RIF | tNGS | Total | |||
|---|---|---|---|---|---|
| Rifampicin- susceptible |
Rifampicin- resistant |
Rifampicin -not applicable# |
MTB not detected | ||
| Rifampicin resistance not detected |
26 (16.46%) |
1 (6.33%) |
29 (18.35%) |
5 (3.17%) |
61 |
| Rifampicin resistance detected | 0 |
3 (1.90%) |
2 (1.27%) |
0 | 5 |
| Rifampicin resistance indeterminate | 0 | 0 |
4 (2.53%) |
0 | 4 |
| MTB not detected |
3 (1.90%) |
0 |
43 (27.22%) |
42 (26.57%) |
88 |
| Total | 29 | 4 | 78 | 47 | 158 |
# indicates that MTB concentration is too low, and the resistance to rifampicin cannot be clearly determined
There were 51 cases where the tuberculosis identification results of Xpert MTB/RIF and tNGS on the BALF samples were inconsistent. We further explored the factors which may possibly lead to such discrepancy. In this study, ddPCR was used for verification on the remaining nucleic acid. First, the tNGS RPhKs presented significantly different among semi-quantitative categories reported by Xpert MTB/RIF, notely the Xpert MTB/RIF negatives were still with low tNGS RPhK instead of no tNGS signal (Fig. 3A). Regarding the 8 non-tuberculosis cases, 6 were concluded as positives supported by tNGS or ddPCR validation (Fig. 3B), suggesting a potential of reporting early infection even in asymptomatic infection. These findings indicate that tNGS presents higher detection sensitivity than Xpert MTB/RIF and may assist in improving aetiology identification. Among the total of 51 cases, there were 13 cases where there was not enough volume for ddPCR. Among the 38 BALF samples with inconsistent MTB identification results between Xpert MTB/RIF and tNGS, 29 were ddPCR positive, of which all were tNGS-positive, while none was Xpert MTB/RIF-positive (Fig. 3C). In addition, 9 cases of ddPCR negative samples, included 1 case of Xpert MTB/RIF positive and 8 cases of tNGS positive.
Fig. 3.
Further validation of the discrepant results
Note: A) Relationship between Xpert MTB/RIF results with varying bacterial loads and the RPhK detected by tNGS. The tNGS RPhK prevalence was plotted based on different Xpert MTB/RIF semi-quantitative positive categories: RPhKXpert MTB/RIF(high) = [74323, 85882]; RPhKXpert MTB/RIF(medium) = [9105,71443]; RPhKXpert MTB/RIF(low) = [34, 37556]; RPhKXpert MTB/RIF(very low) = [0, 3036]; RPhKXpert MTB/RIF(−)−ddPCR(+) = [3, 1973]; RPhKXpert MTB/RIF(−)−ddPCR(−) = [14, 234]. Significant differences were observed in the RPhK values between the Xpert MTB/RIF(high) group and the Xpert MTB/RIF(medium) group (P = 0.02, Wilcoxon rank sum test), between the Xpert MTB/RIF(medium) group and the Xpert MTB/RIF(low) group (P = 0.035, Wilcoxon rank sum test), and between Xpert MTB/RIF(low) group and the Xpert MTB/RIF(very low) group (P = 1.3e-06, Wilcoxon rank sum test). The RPhK values for both ddPCR(+) and ddPCR(-) did not show a significant difference (P = 0.11, Wilcoxon rank sum test). B) Detection results of Non-tuberculosis. C) Validation of the inconsistent MTB identification results between Xpert MTB/RIF and tNGS using ddPCR (N = 38/51#)
Discussion
This study primarily utilized MTB clinical isolates and BALF samples to assess the clinical application value of tNGS in the diagnosis and drug resistance detection of MTB. The “WHO operational handbook on tuberculosis. Module 3: Diagnosis - Rapid diagnostics for tuberculosis detection, third edition” proposes that in people with bacteriologically confirmed pulmonary TB disease, tNGS may be used on respiratory samples to diagnose resistance to rifampicin, isoniazid, fluoroquinolones, pyrazinamide and ethambutol rather than culture-based phenotypic AST [31]. Previous studies have shown that tNGS can be used for the identification, drug resistance detection, and typing of tuberculosis, offering advantages such as cost-effectiveness, efficiency, accuracy, and flexibility [6, 32–44]. To date, there is a lack of studies systematically evaluating the use of tNGS in diagnosing tuberculosis using clinical isolates and samples, in comparison with culture, phenotypic AST, WGS, and Xpert MTB/RIF.
In this study, for 143 MTB clinical isolates, tNGS and phenotypic AST, as well as AST based on WGS, all showed a high agreement, with average concordance rates of 95.10% and 97.05%, respectively, similar to the results of other studies [32, 42, 44]. Nevertheless, a few minor discrepancies were still observed, albeit in a small number of cases. The reasons for the inconsistency between tNGS and phenotypic AST mainly include the following. First, phenotypic AST may miss some low-level resistance to rifampicin, isoniazid, and fluoroquinolones [45, 46]; second, the phenotypic AST of ethambutol may be unreliable [47–49]; finally, tNGS cannot detect other non-resistance gene mutations that cause resistance mechanisms, such as drug efflux. Additionally, the primary reason for the missed detection of drug resistance by tNGS, while it was detected by WGS, is that some rare mutations are not included in the tNGS detection targets. These include rpoB Leu545Met, katG Gly593Asp, ahpC-48G > A, embA -12 C > T, embB Asp354Ala, etc. Among these, rpoB Leu545Met, embA -12 C > T, and embB Asp354Ala have been observed in more than one MTB isolate. These mutation sites may be considered for inclusion in future optimizations of tNGS.
The use of tNGS to detect MTB in clinical samples is common in sputum [33, 34, 37, 42, 44]. Additionally, there are a few studies on cerebrospinal fluid, stool, gastric aspirate, BALF, pus, pleural effusion, and formalin-fixed and paraffin-embedded (FFPE) tissue specimens [6, 36, 38, 41, 43, 44, 50].This study used tNGS to identify MTB in 158 BALF samples, and further validated samples with inconsistent identification results between Xpert MTB/RIF and tNGS using ddPCR. In the confirmed cases, the positive rate of tNGS (73/83, 87.95%) is higher than that of Xpert MTB (67/83, 80.72%).Additionally, 45% of clinically diagnosed cases were detected positive by tNGS. Regarding all 38 samples with discrepant results, further validation with ddPCR yielded 29 positives, of which all tNGS were positive, and none was Xpert MTB/RIF positive. Globally, the bacteriological confirmation rate among people diagnosed with tuberculosis in 2022 was only 63%, and 62% in China [1].The application of new tuberculosis diagnostic technologies, including tNGS, can enhance the bacteriological confirmation rate. Certainly, it is also important to note that the performance of tNGS in detecting rifampicin drug resistance in BALF samples was not as good as Xpert MTB/RIF in our study. There were 29 samples that tested positive for tuberculosis using tNGS, but the results for rifampicin drug resistance were “rifampicin not applicable”, and this situation mostly occurred when the semi-quantitative results of Xpert MTB/RIF were very low. This is because tNGS targets multi-copy genes (IS6110 and IS1081) for detecting MTB, whereas rifampicin resistance is detected using a single-copy gene (rpoB). Additionally, there is inconsistency in amplification efficiency across different detection sites. Therefore, if the concentration of MTB in clinical samples is low, it may result in the inability to detect rifampicin resistance. It is noteworthy that the tNGS results of a BALF sample showed resistance to rifampicin, indicating a rifampicin-resistant mutation site at the rpoB Leu511Pro, with a mutation frequency of 0.6. The Xpert MTB/RIF result of this case showed that rifampicin resistance was not detected, suggesting that in some cases, clinical treatment can be initiated based on tNGS results.
The detection performance of tNGS is closely related to the sequencing platform and nucleic acid extraction methods, which has also been mentioned in the studies by Andrea M Cabibbe and Tara E Ness [34, 50]. In this study, the nucleic acid extraction protocol incorporated both mechanical cell wall disruption and subsequent chemical lysis techniques to achieve optimized overall efficiency and reliability for drug resistance mutation detection. Additionally, this study employed the KM MiniSeq Dx-CN platform, a relatively compact sequencing system adapted to clinical laboratory applications where sample size is relatively small (e.g. MTB). The MiniSeq platform offers low sequencing error rates, thereby ensuring the accurate detection of drug resistance-associated mutations.
Our study has several limitations. Firstly, the anti-tuberculosis drugs involved in this study are limited, such as bedaquiline, pretomanid, linezolid and other drugs that have been included in the treatment of drug-resistant tuberculosis [51]. Secondly, the retrospective study was conducted in the clinical laboratory, and tNGS results were not communicated to clinicians and patients, which did not directly provide reference for clinical diagnosis and treatment. Finally, the targets of tNGS in this study did not fully cover the comprehensive resistance mutations in the WHO document [18]. Future studies that utilize an expanded panel encompassing a wider range of target loci and diverse treatment interventions could provide more comprehensive insights into the diagnosis value of tNGS. Additionally, the overall sensitivity of detecting resistance genotypes is lower than that of MTB identification (which may be due to the lack of multi-copy target loci for primer design and low MTB titer in clinical samples). Consequently, reporting both the presence of MTB and the resistance genotypes may not occur simultaneously. To address this limitation, further research is required to develop methods that enhance detection sensitivity and enable concurrent reporting of both MTB and its resistance profiles. With the accumulation of tNGS clinical data, optimization of detection targets, development of sequencing technology and optimization of nucleic acid extraction, the performance of tNGS in laboratory testing for tuberculosis can be improved in the next version of the assay.
The findings of this study underscore the potential of tNGS in transforming routine practice within the field of tuberculosis management. The high concordance rate between tNGS and WGS/AST, combined with its cost-efficiency, suggests that tNGS may be a competitive alternative and may lead to a shift in the TB diagnosis paradigm in the future: rather than relying on sequential testing for drug resistance, a single-step comprehensive assessment could streamline the diagnostic process. This approach offers several clinical advantages: it enables the early detection of tuberculosis and its resistance profiles, provides broad coverage of resistance targets for commonly used clinical drugs, and supports precision medicine by guiding appropriate therapeutic choices. It should be noted that guidelines initiated by WHO on the use of tNGS provide effective direction for the implementation and application of tNGS, thereby accelerating its clinical application and promotion in the diagnosis of tuberculosis and drug resistance [52]. Ultimately, integrating rapid, multifaceted testing into routine clinical practice could enhance patient outcomes and contribute to more effective public health interventions against tuberculosis.
In conclusion, tNGS demonstrates robust and rapid performance in the identification of MTB and its associated drug resistance, and can be directly applied to clinical samples, positioning it as a promising approach for laboratory testing of tuberculosis. Combined with its cost-efficiency and short turn-around time, tNGS may have an impact on the future TB diagnosis paradigm and treatment decisions.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Author contributions
X.W and F.Y designed the study. X.W and G.T wrote the manuscript. X.W and F.Y modified the manuscript. G.T, C.S and X.W did the statistics. Y.W and C.W provided technical assistance in sequencing and data visualization. X.W and J.Y did laboratory examination. C.H and F.Y supervised the project.
Funding
This project was supported through grants from Shanghai Municipal Health Commission’s Special Youth Project for Clinical Research in the Health Industry (No. 20224Y0129), Shanghai Three-year (2023–2025) Action Plan to Strengthen the Public Health System (No. GWVI-11.2-YQ52, and GWVI-11.1-05), and Shanghai Science and Technology Innovation Action Plan, Medical Innovation Research Special Project (No. 22Y11902600).
Data availability
The sequencing data generated in this study has been deposited in the CNCB Genome Sequence Archive database under the accession number PRJCA026575.
Declarations
Competing interests
The authors declare no competing interests.
Ethical approval
This study was approved by the Ethical Committee of Shanghai Pulmonary Hospital.
Conflict of interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Footnotes
Xiaocui Wu, Guangkun Tan and Chunlei Sun contributed equally to this work.
Contributor Information
Chaohui Hu, Email: huzh@kingcreate.com.cn.
Fangyou Yu, Email: wzjxyfy@163.com.
References
- 1.WHO. Global tuberculosis report 2023. 2023.
- 2.Forbes BA, Hall GS, Miller MB, et al. Practice guidelines for clinical Microbiology Laboratories: Mycobacteria. Clin Microbiol Rev. 2018;31(2):artnoe00038-17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Dorman S, Schumacher S, Alland D, et al. Xpert MTB/RIF Ultra for detection of Mycobacterium tuberculosis and rifampicin resistance: a prospective multicentre diagnostic accuracy study. Lancet Infect Dis. 2018;18(1):76–84. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.WHO. The use of next-generation sequencing for the surveillance of drug-resistant tuberculosis: an implementation manual. 2023.
- 5.WHO. Use of targeted next-generation sequencing to detect drug-resistant tuberculosis: rapid communication, July 2023. 2023.
- 6.Ye J, Huang K, Xu Y, et al. Clinical application of nanopore-targeted sequencing technology in bronchoalveolar lavage fluid from patients with pulmonary infections. Microbiol Spectr. 2024 Apr;30:e0002624. [DOI] [PMC free article] [PubMed]
- 7.Cai S, Yuan J, Li Y, et al. Etiological diagnostic performance of probe capture-based targeted next-generation sequencing in bloodstream infection. J Thorac Dis. 2024;16(4):2539–49. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Lin R, Xing Z, Liu X et al. Performance of targeted next-generation sequencing in the detection of respiratory pathogens and antimicrobial resistance genes for children. J Med Microbiol. 2023;72(11). [DOI] [PubMed]
- 9.Lin A, Singh A, Allred A, et al. Targeted next-generation sequencing assay for direct detection and serotyping of Salmonella from Enrichment. J Food Prot. 2024;87(4):100256. [DOI] [PubMed] [Google Scholar]
- 10.Poulsen SH, Søgaard KK, Fuursted K, Nielsen HL. Evaluating the diagnostic accuracy and clinical utility of 16S and 18S rRNA gene targeted next-generation sequencing based on five years of clinical experience. Infect Dis (Lond). 2023;55(11):767–75. [DOI] [PubMed] [Google Scholar]
- 11.Li J, Zhang L, Yang X, et al. Diagnostic significance of targeted next-generation sequencing in Central Nervous System infections in Neurosurgery of Pediatrics. Infect Drug Resist. 2023;16:2227–36. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Kyi D, Xiao Y, Wang X, et al. Predominance of A2063G mutant strains in the Mycoplasma pneumoniae epidemic in children-a clinical and epidemiological study in 2023 in Wuhan, China. Int J Infect Dis. 2024 May;9:107074. [DOI] [PubMed]
- 13.Hong HL, Flurin L, Thoendel MJ, et al. Targeted Versus Shotgun Metagenomic sequencing-based detection of microorganisms in Sonicate Fluid for Periprosthetic Joint infection diagnosis. Clin Infect Dis. 2023;76(3):e1456–62. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Zhang Y, Jiang X, Ye W, Sun J. Clinical features and outcome of eight patients with Chlamydia psittaci pneumonia diagnosed by targeted next generation sequencing. Clin Respir J. 2023;17(9):915–30. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Li X, Liu Y, Li M, et al. Epidemiological investigation of lower respiratory tract infections during influenza A (H1N1) pdm09 virus pandemic based on targeted next-generation sequencing. Front Cell Infect Microbiol. 2023;13:1303456. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Deng Z, Li C, Wang Y, et al. Targeted next-generation sequencing for pulmonary infection diagnosis in patients unsuitable for bronchoalveolar lavage. Front Med (Lausanne). 2023;10:1321515. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Huang C, Huang Y, Wang Z, et al. Multiplex PCR-based next generation sequencing as a novel, targeted and accurate molecular approach for periprosthetic joint infection diagnosis. Front Microbiol. 2023;14:1181348. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.WHO. Catalogue of mutations in Mycobacterium tuberculosis complex and their association with drug resistance, 2nd. Second ed. 2023 15 November.
- 19.WHO. Catalogue of mutations in Mycobacterium tuberculosis complex and their association with drug resistance, 1nd. 2021.
- 20.Wu X, Gao R, Shen X, et al. Use of whole-genome sequencing to predict Mycobacterium tuberculosis drug resistance in Shanghai, China. Int J Infect Dis. 2020;96:48–53. [DOI] [PubMed] [Google Scholar]
- 21.Wu X, Tan G, Sha W, et al. Use of whole-genome sequencing to Predict Mycobacterium tuberculosis Complex Drug Resistance from early positive liquid cultures. Microbiol Spectr. 2022;10(2):e0251621. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Industry PsRoCH. Diagnosis of Pulmonary Tuberculosis (WS 288–2017). Electron J Emerg Infect Dis2018. pp. 59–61.
- 23.Wu X, Yang J, Tan G, et al. Drug Resistance characteristics of Mycobacterium tuberculosis isolates from patients with tuberculosis to 12 antituberculous drugs in China. Front Cell Infect Microbiol. 2019;9:345. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Chen S, Zhou Y, Chen Y, Gu J. Fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics. 2018;34(17):i884–90. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Li H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. arXiv e-prints. 2013.
- 26.Benson DA, Cavanaugh M, Clark K, et al. GenBank Nucleic Acids Res. 2018;46(D1):D41–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.O’Leary NA, Wright MW, Brister JR, et al. Reference sequence (RefSeq) database at NCBI: current status, taxonomic expansion, and functional annotation. Nucleic Acids Res. 2016;44(D1):D733–45. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Gardner MJ, Altman DG. Confidence intervals rather than P values: estimation rather than hypothesis testing. Br Med J (Clin Res Ed). 1986;292(6522):746–50. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Altman DG, Bland JM. Diagnostic tests 2: predictive values. BMJ. 1994;309(6947):102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Altman DG, Bland JM. Diagnostic tests. 1: sensitivity and specificity. BMJ. 1994;308(6943):1552. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.WHO. WHO operational handbook on tuberculosis. Module 3: Diagnosis - Rapid diagnostics for tuberculosis detection. third edition ed. 2024.
- 32.Wu SH, Xiao YX, Hsiao HC, Jou R. Development and Assessment of a Novel Whole-Gene-based targeted next-generation sequencing assay for detecting the susceptibility of Mycobacterium tuberculosis to 14 drugs. Microbiol Spectr. 2022;10(6):e0260522. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Kambli P, Ajbani K, Kazi M, et al. Targeted next generation sequencing directly from sputum for comprehensive genetic information on drug resistant Mycobacterium tuberculosis. Tuberculosis (Edinb). 2021;127:102051. [DOI] [PubMed] [Google Scholar]
- 34.Cabibbe AM, Spitaleri A, Battaglia S et al. Application of targeted next-generation sequencing assay on a portable sequencing platform for culture-free detection of drug-resistant tuberculosis from clinical samples. J Clin Microbiol. 2020;58(10). [DOI] [PMC free article] [PubMed]
- 35.Schlanderer J, Hoffmann H, Lüddecke J, et al. Two-stage tuberculosis diagnostics: combining centrifugal microfluidics to detect TB infection and inh and rif resistance at the point of care with subsequent antibiotic resistance profiling by targeted NGS. Lab Chip. 2023;24(1):74–84. [DOI] [PubMed] [Google Scholar]
- 36.Song J, Du W, Liu Z, et al. Application of Amplicon-based targeted NGS technology for diagnosis of drug-resistant tuberculosis using FFPE specimens. Microbiol Spectr. 2022;10(1):e0135821. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Iyer A, Ndlovu Z, Sharma J, et al. Operationalising targeted next-generation sequencing for routine diagnosis of drug-resistant TB. Public Health Action. 2023;13(2):43–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Tram TTB, Trieu LPT, Nhat LTH, et al. Targeted sequencing from cerebrospinal fluid for rapid identification of drug-resistant tuberculous meningitis. J Clin Microbiol. 2024;62(4):e0128723. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Zhang H, Dai X, Hu P, et al. Comparison of targeted next-generation sequencing and the Xpert MTB/RIF assay for detection of Mycobacterium tuberculosis in clinical isolates and sputum specimens. Microbiol Spectr. 2024;12(5):e0409823. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Genestet C, Baffert Y, Vallée M et al. Development, evaluation, and implementation of a House-made targeted next-generation sequencing spoligotyping in a French laboratory. Int J Mol Sci. 2022;23(19). [DOI] [PMC free article] [PubMed]
- 41.Zheng H, Yang H, Wang Y, et al. Diagnostic value of tNGS vs Xpert MTB/RIF in childhood TB. Heliyon. 2024;10(1):e23217. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Cloutier Charette W, Rabodoarivelo MS, Point F, et al. Concordance of targeted and whole genome sequencing for Mycobacterium tuberculosis genotypic drug susceptibility testing. Diagn Microbiol Infect Dis. 2024;109(2):116249. [DOI] [PubMed] [Google Scholar]
- 43.Sibandze DB, Kay A, Dreyer V, et al. Rapid molecular diagnostics of tuberculosis resistance by targeted stool sequencing. Genome Med. 2022;14(1):52. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Murphy SG, Smith C, Lapierre P, et al. Direct detection of drug-resistant Mycobacterium tuberculosis using targeted next generation sequencing. Front Public Health. 2023;11:1206056. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Li J, Gao X, Luo T, et al. Association of gyrA/B mutations and resistance levels to fluoroquinolones in clinical isolates of Mycobacterium tuberculosis. Emerg Microbes Infect. 2014;3(3):e19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Angra PK, Taylor TH, Iademarco MF, et al. Performance of tuberculosis drug susceptibility testing in U.S. laboratories from 1994 to 2008. J Clin Microbiol. 2012;50(4):1233–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Nikolayevskyy V, Hillemann D, Richter E, et al. External Quality Assessment for Tuberculosis diagnosis and Drug Resistance in the European Union: a five year multicentre implementation study. PLoS ONE. 2016;11(4):e0152926. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Madison B, Robinson-Dunn B, George I, et al. Multicenter evaluation of ethambutol susceptibility testing of mycobacterium tuberculosis by agar proportion and radiometric methods. J Clin Microbiol. 2002;40(11):3976–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Van Deun A, Wright A, Zignol M, et al. Drug susceptibility testing proficiency in the network of supranational tuberculosis reference laboratories. Int J Tuberc Lung Dis. 2011;15(1):116–24. [PubMed] [Google Scholar]
- 50.Ness TE, Meiwes L, Kay A, et al. Optimizing DNA extraction from Pediatric Stool for diagnosis of tuberculosis and use in next-generation sequencing applications. Microbiol Spectr. 2023;11(1):e0226922. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.WHO. WHO operational handbook on tuberculosis. Module 4: treatment - drug-resistant tuberculosis treatment, 2022 update. 2022. [PubMed]
- 52.WHO. Use of targeted next-generation sequencing to detect drug-resistant tuberculosis: rapid communication. 2023.
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The sequencing data generated in this study has been deposited in the CNCB Genome Sequence Archive database under the accession number PRJCA026575.